Details of the Target
General Information of Target
Probe(s) Labeling This Target
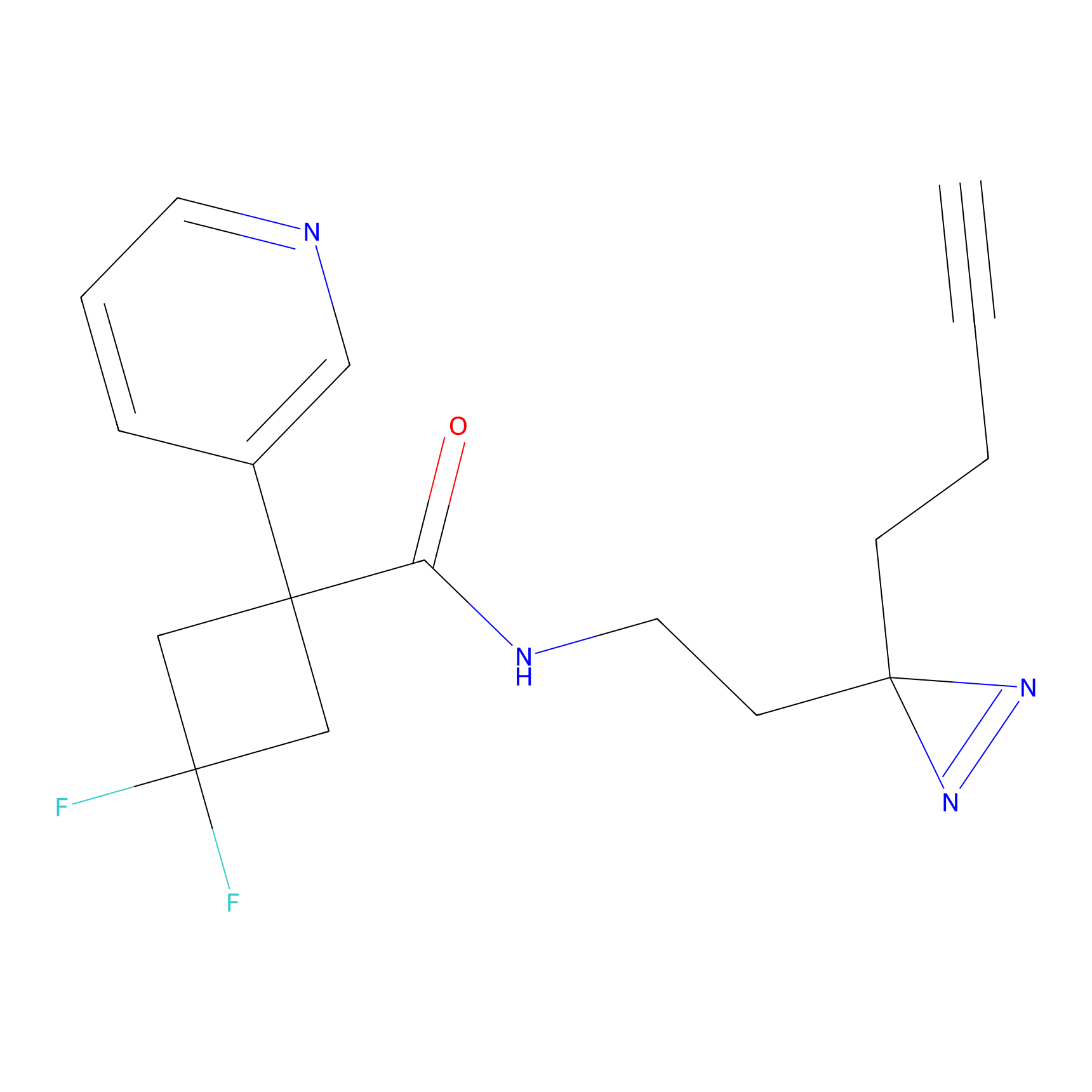
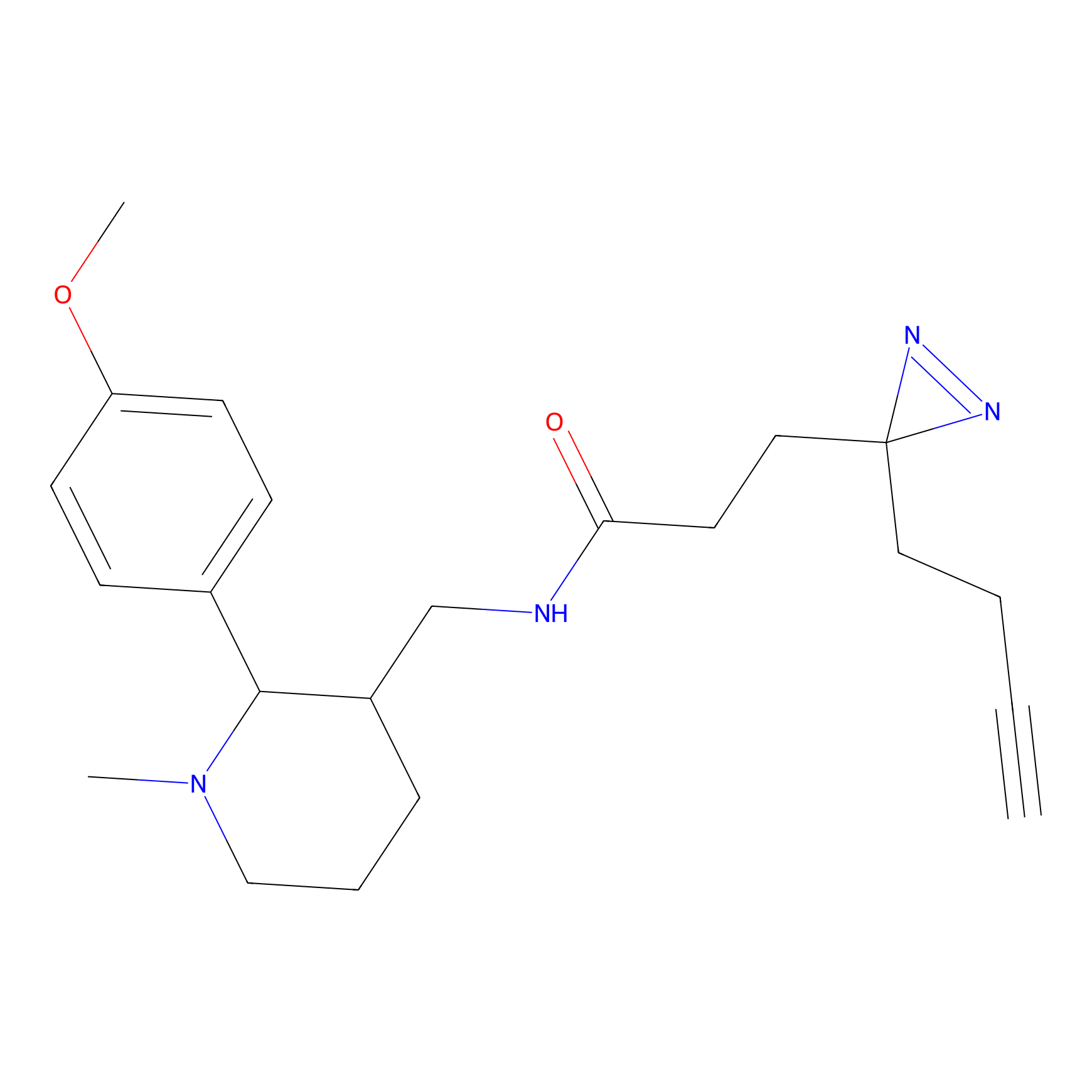
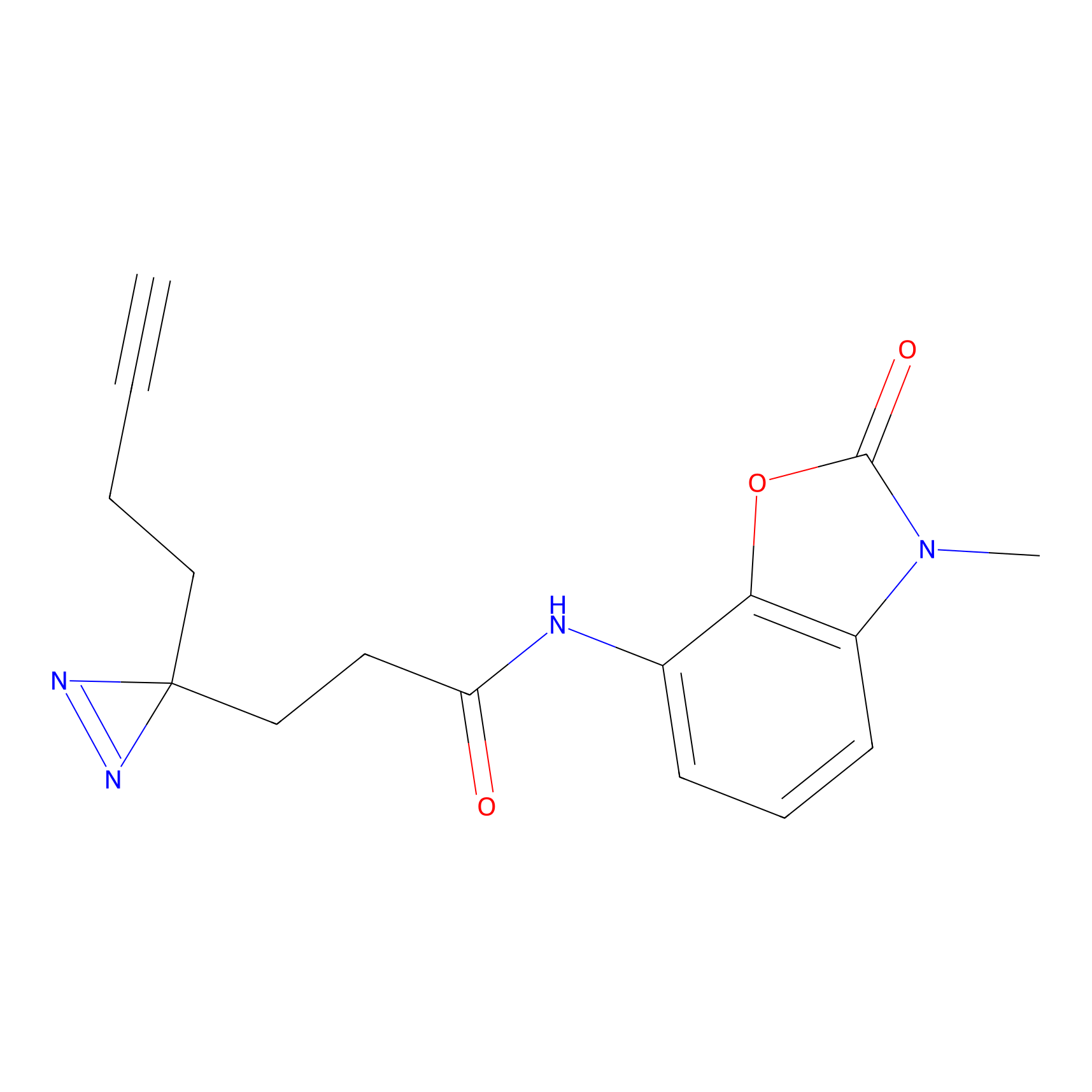
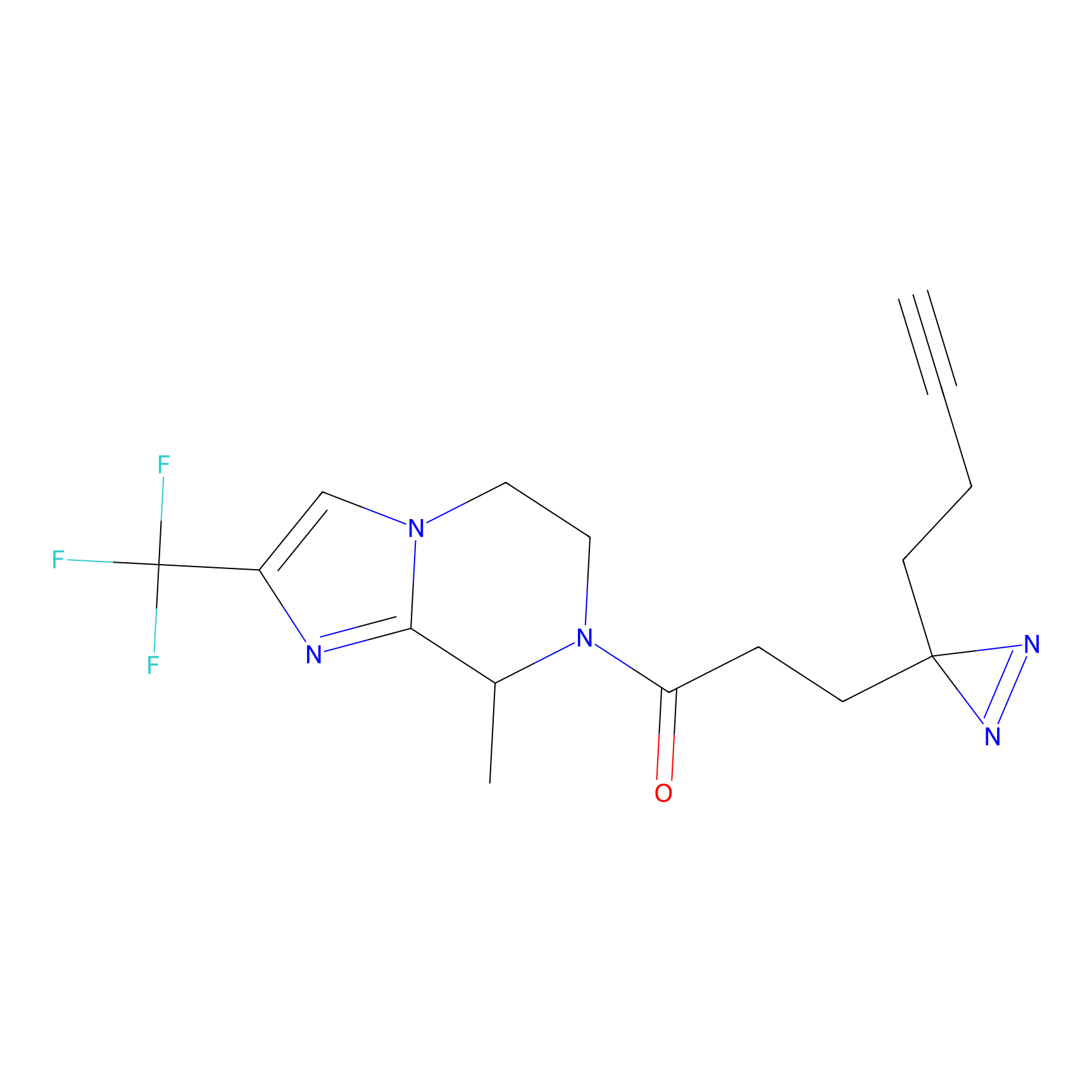
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
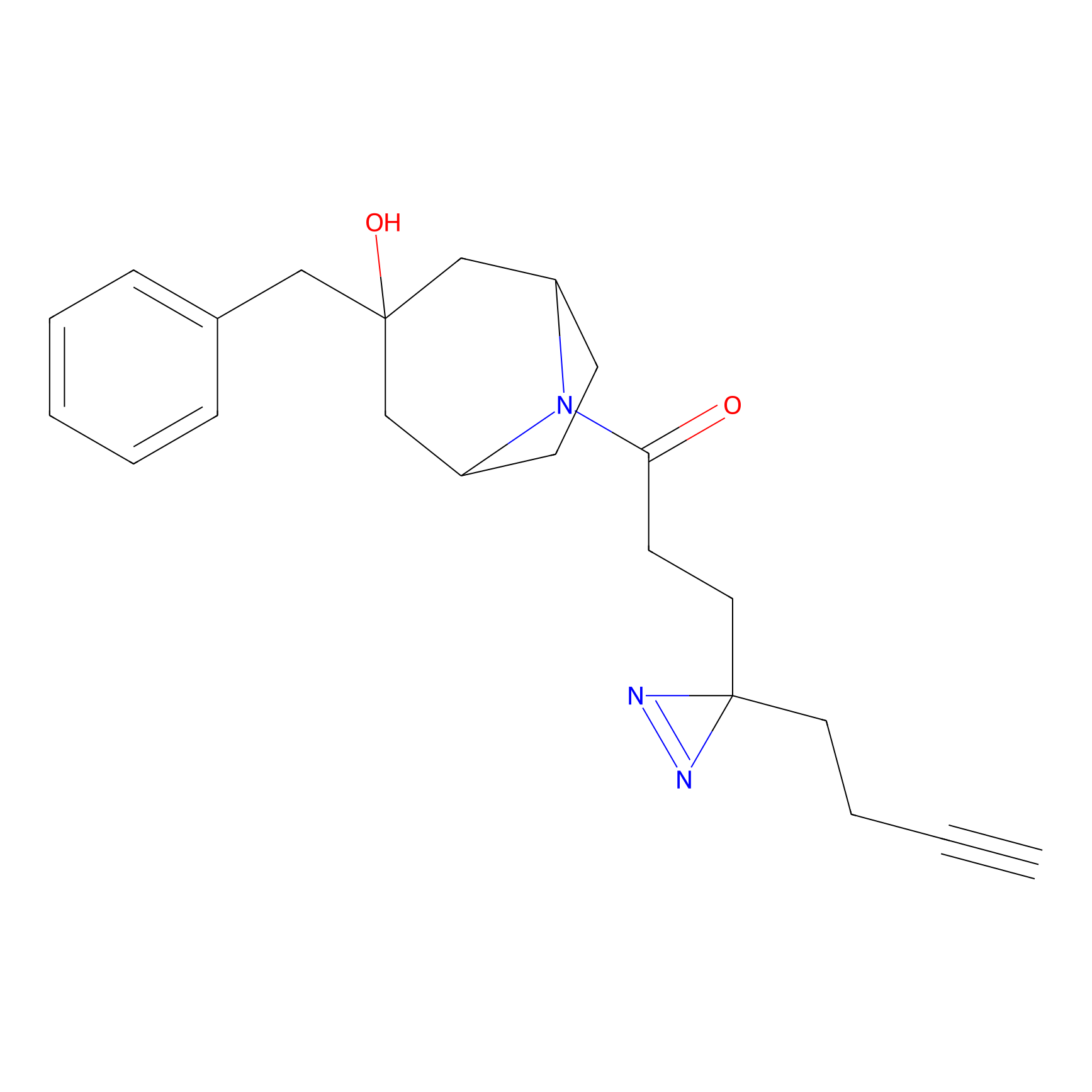
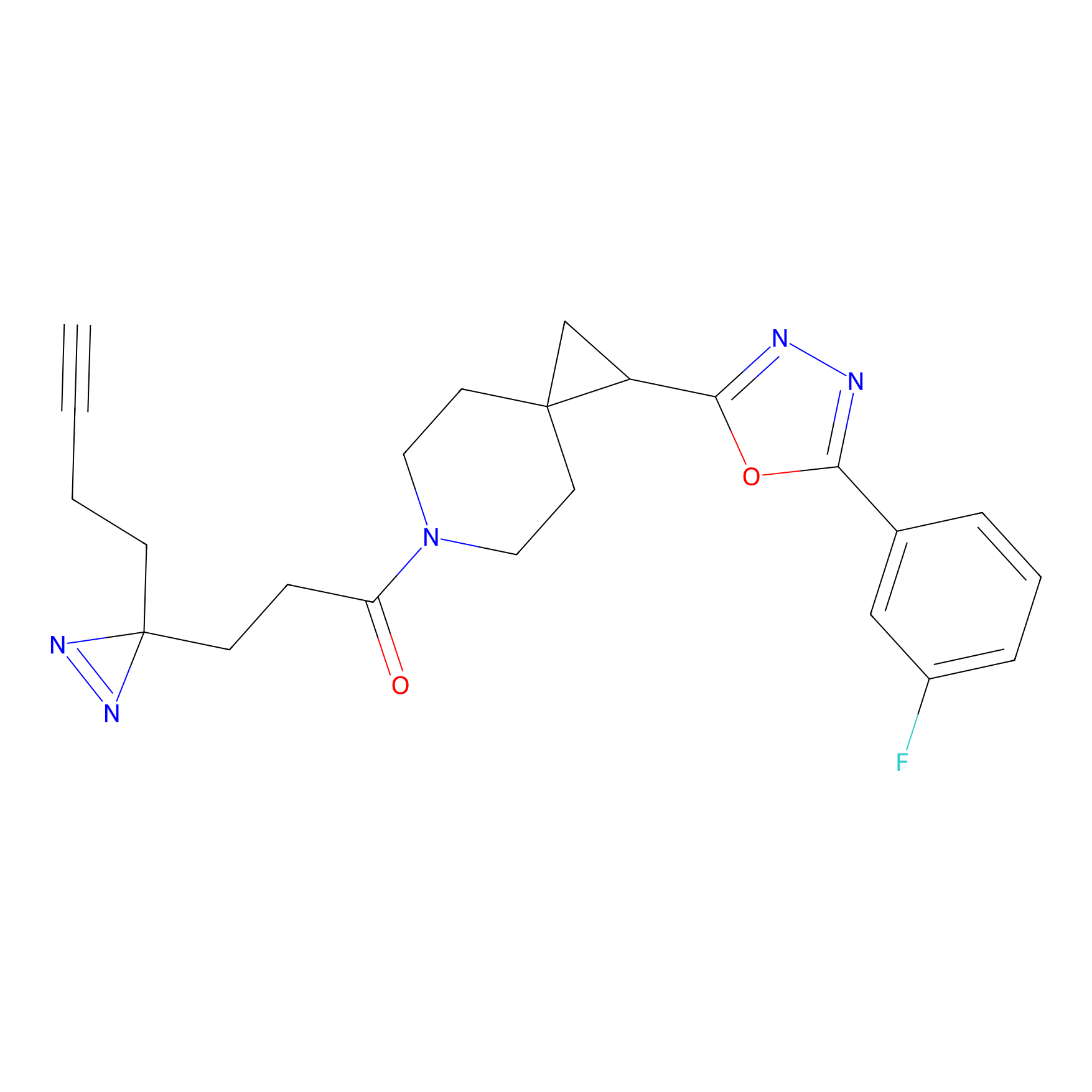
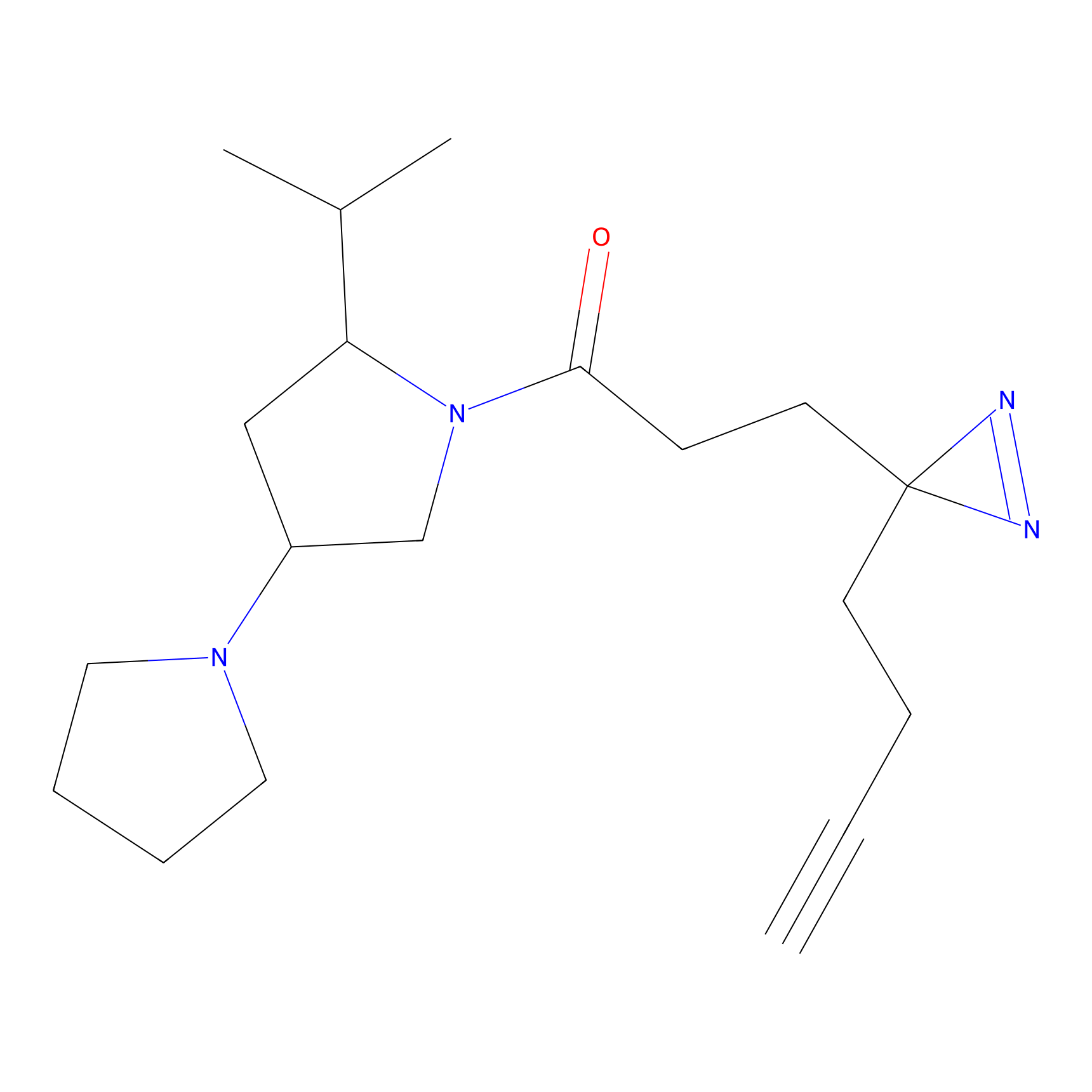
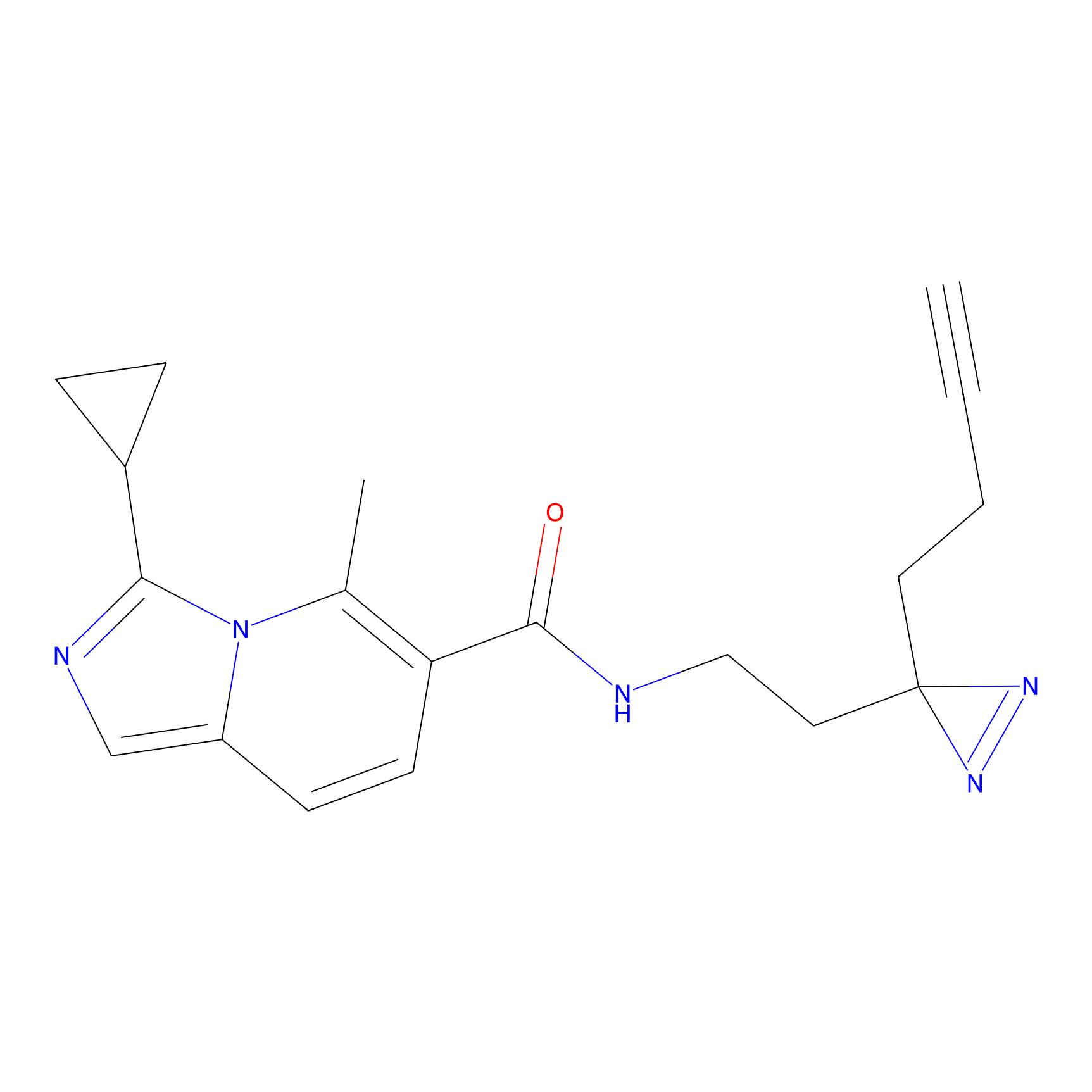
FBPP2 Probe Info |
 |
14.23 | LDD0324 | [1] | |
|
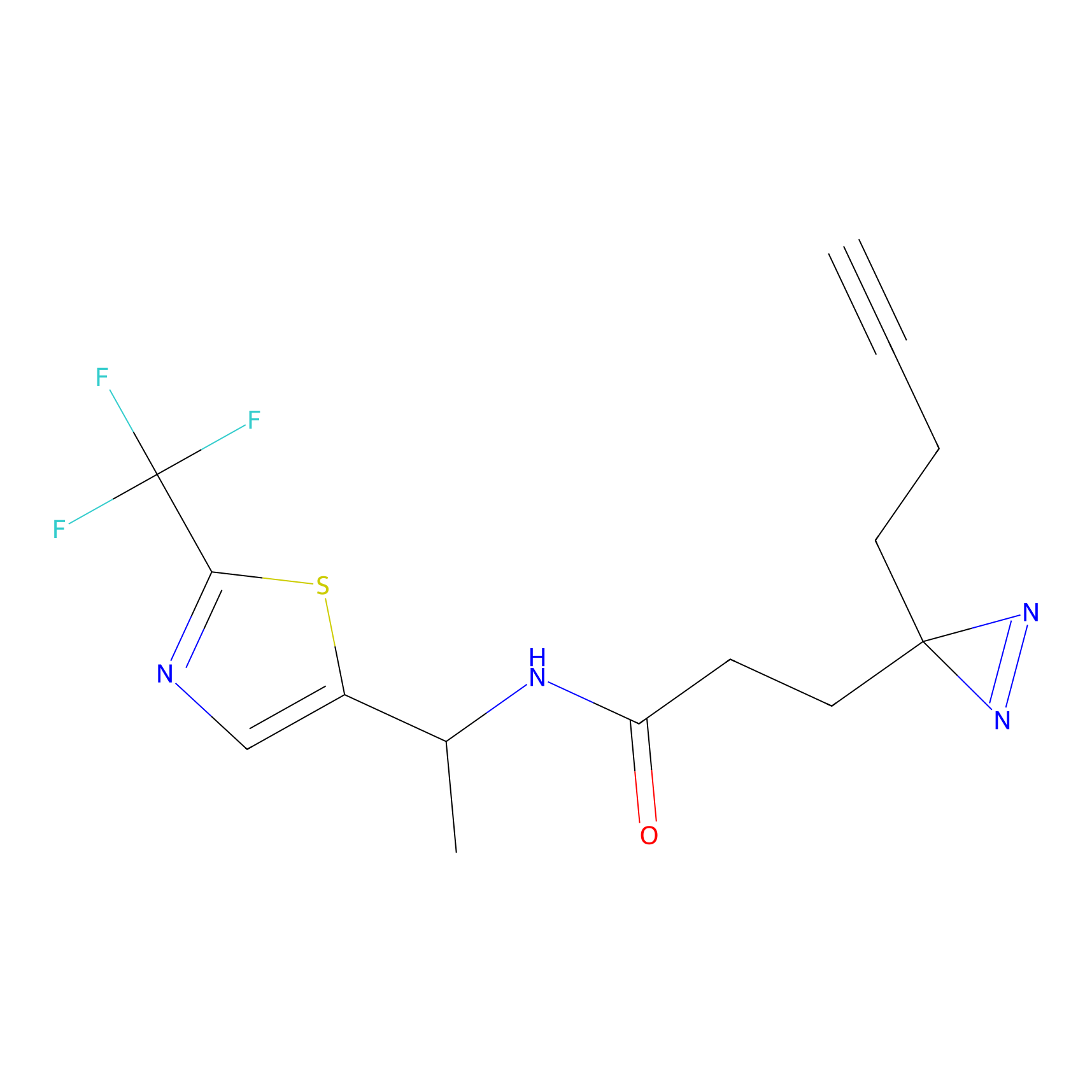
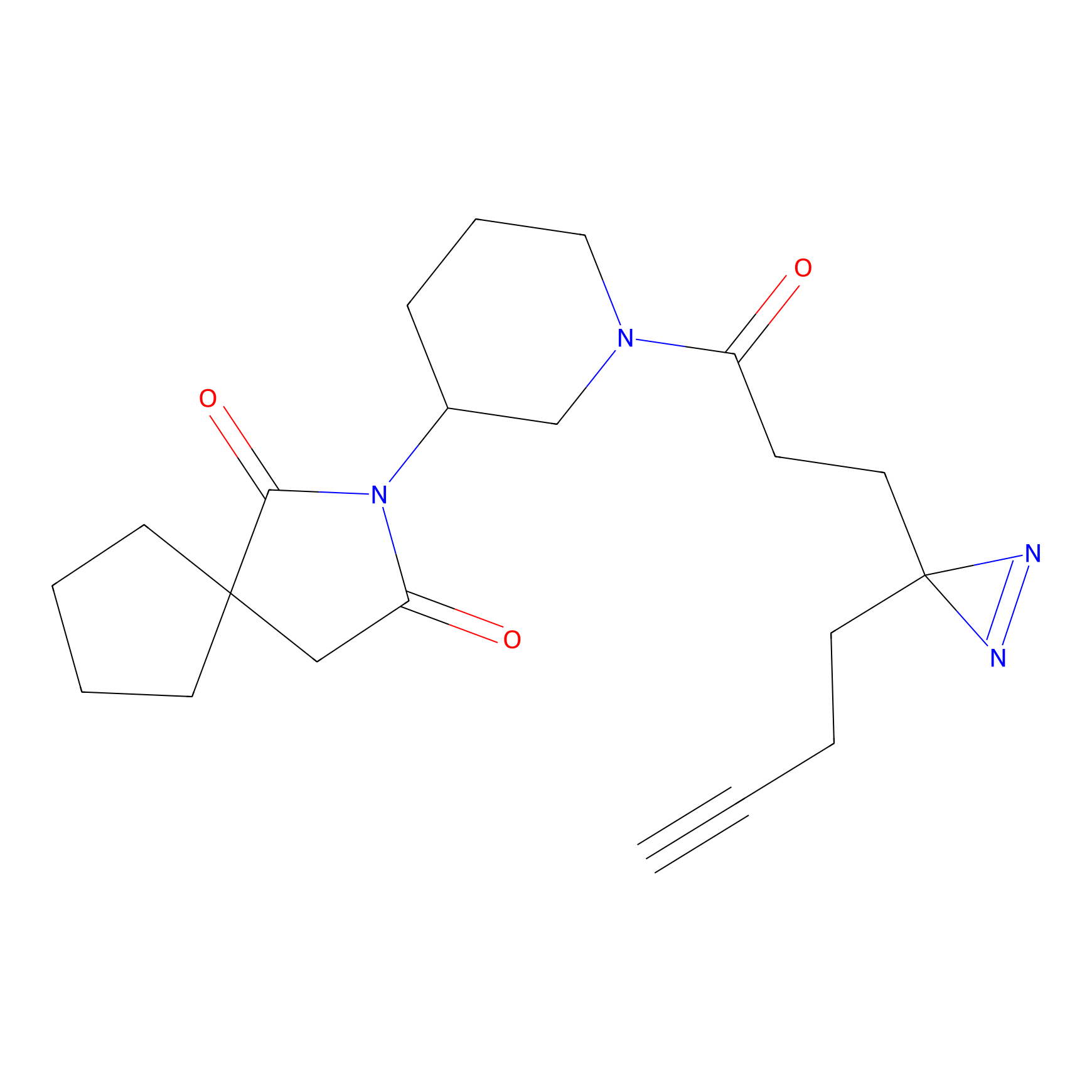
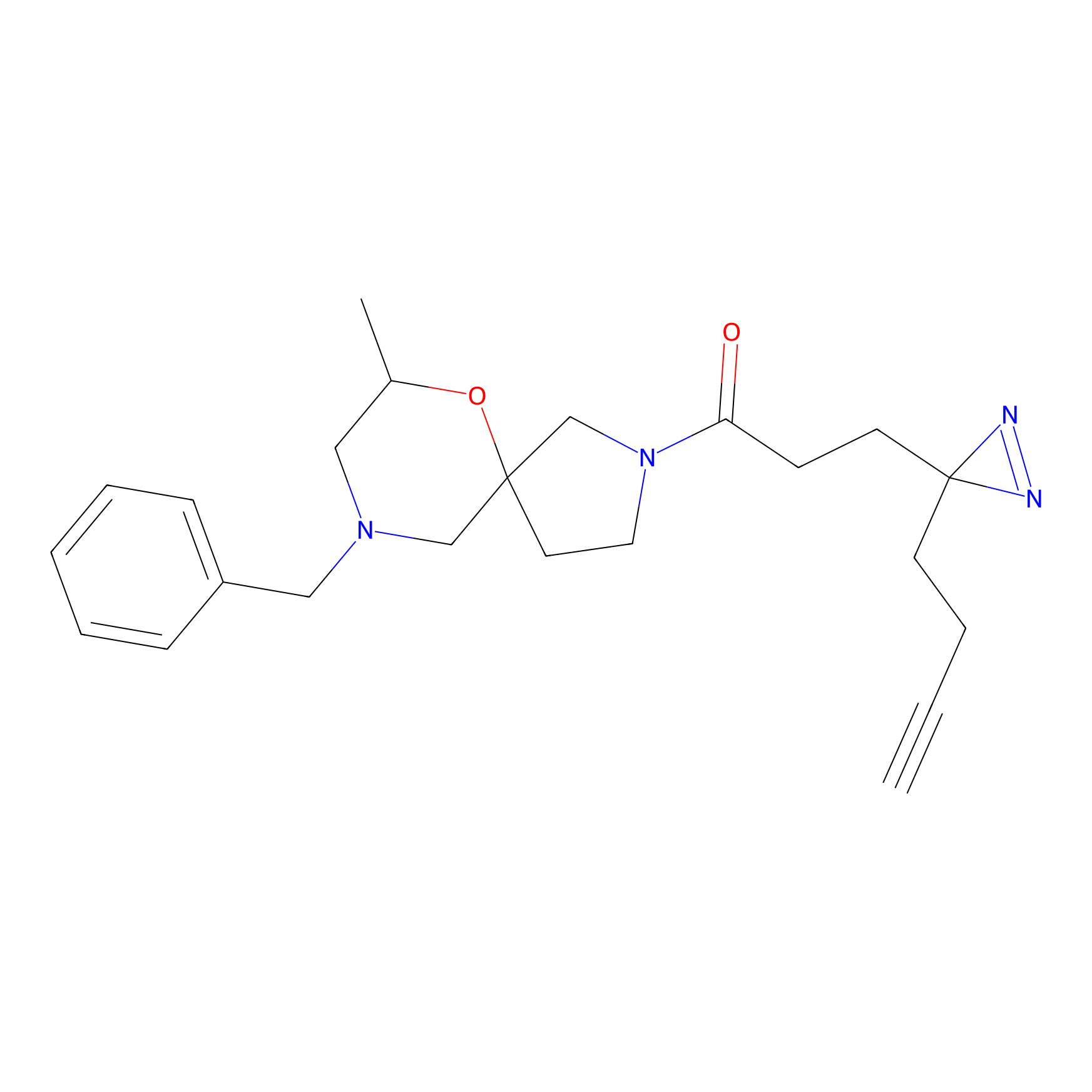
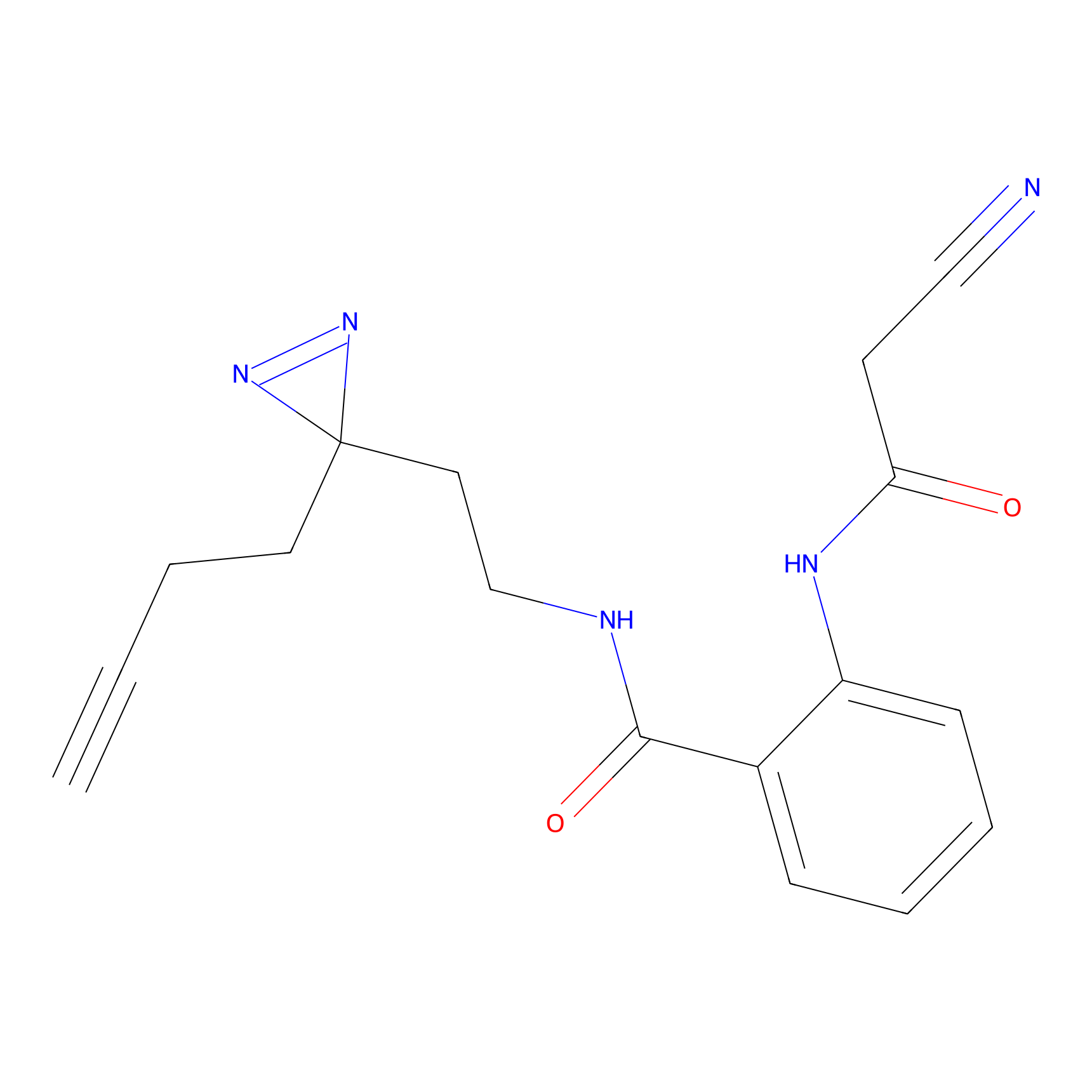
DA-P2 Probe Info |
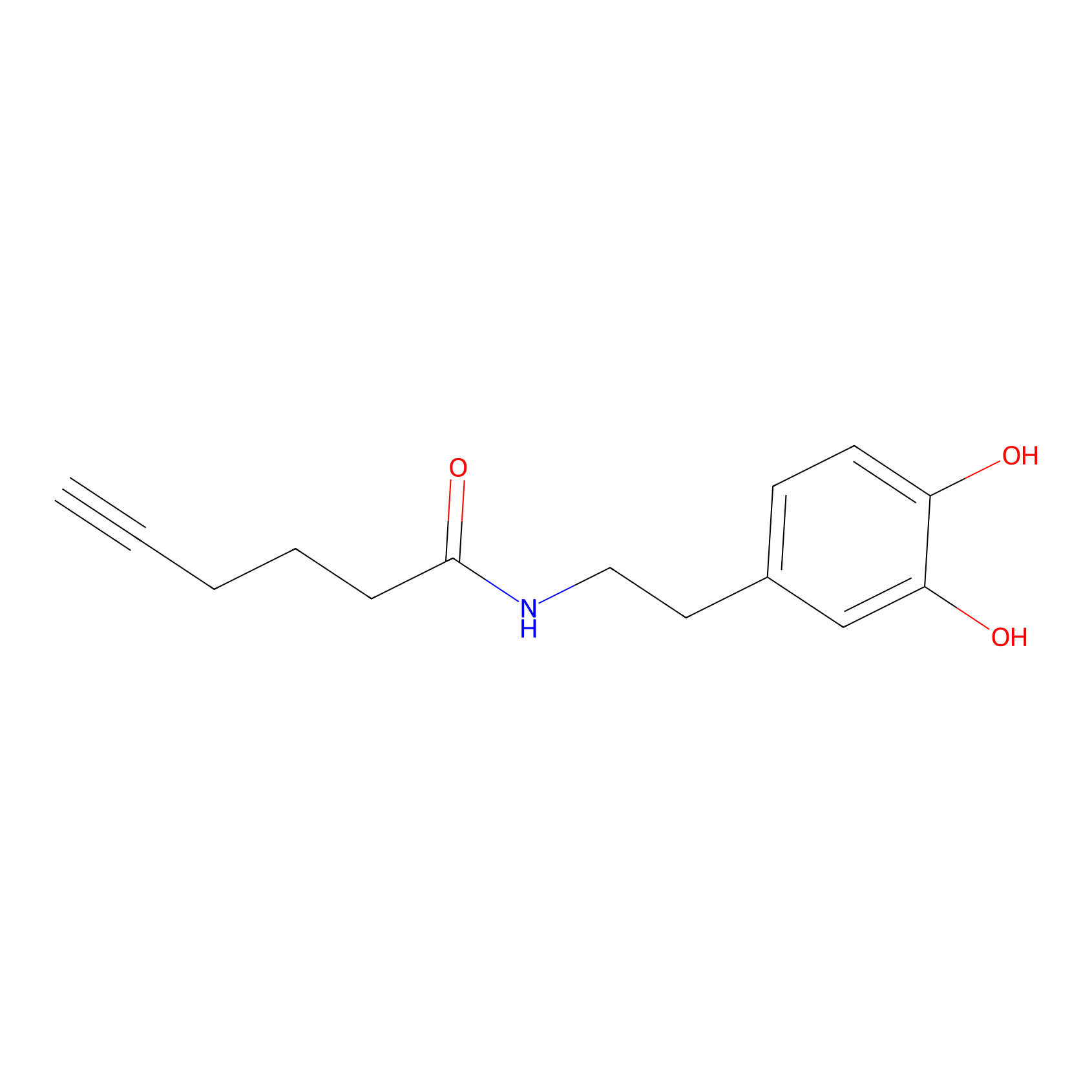 |
4.96 | LDD0348 | [2] | |
|
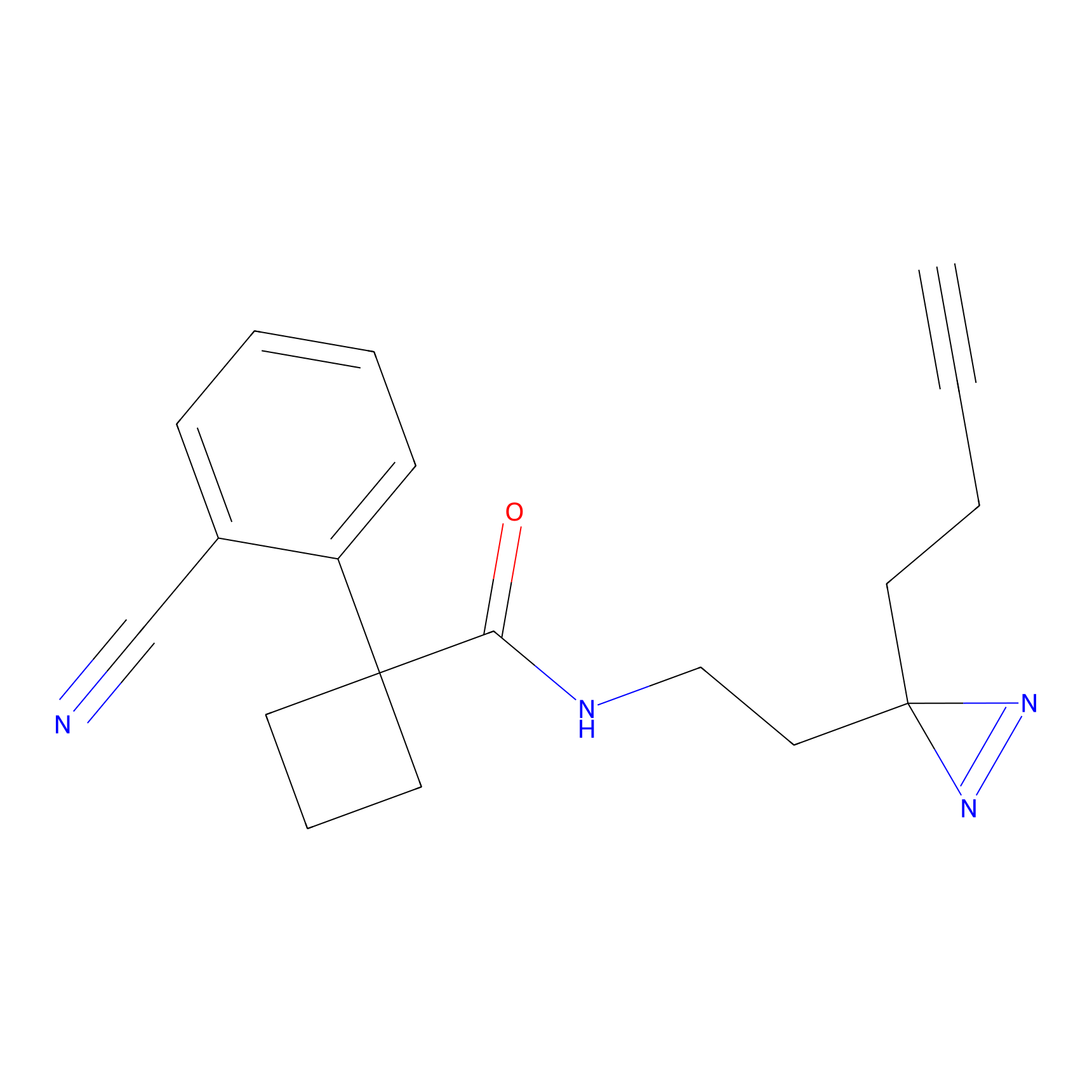
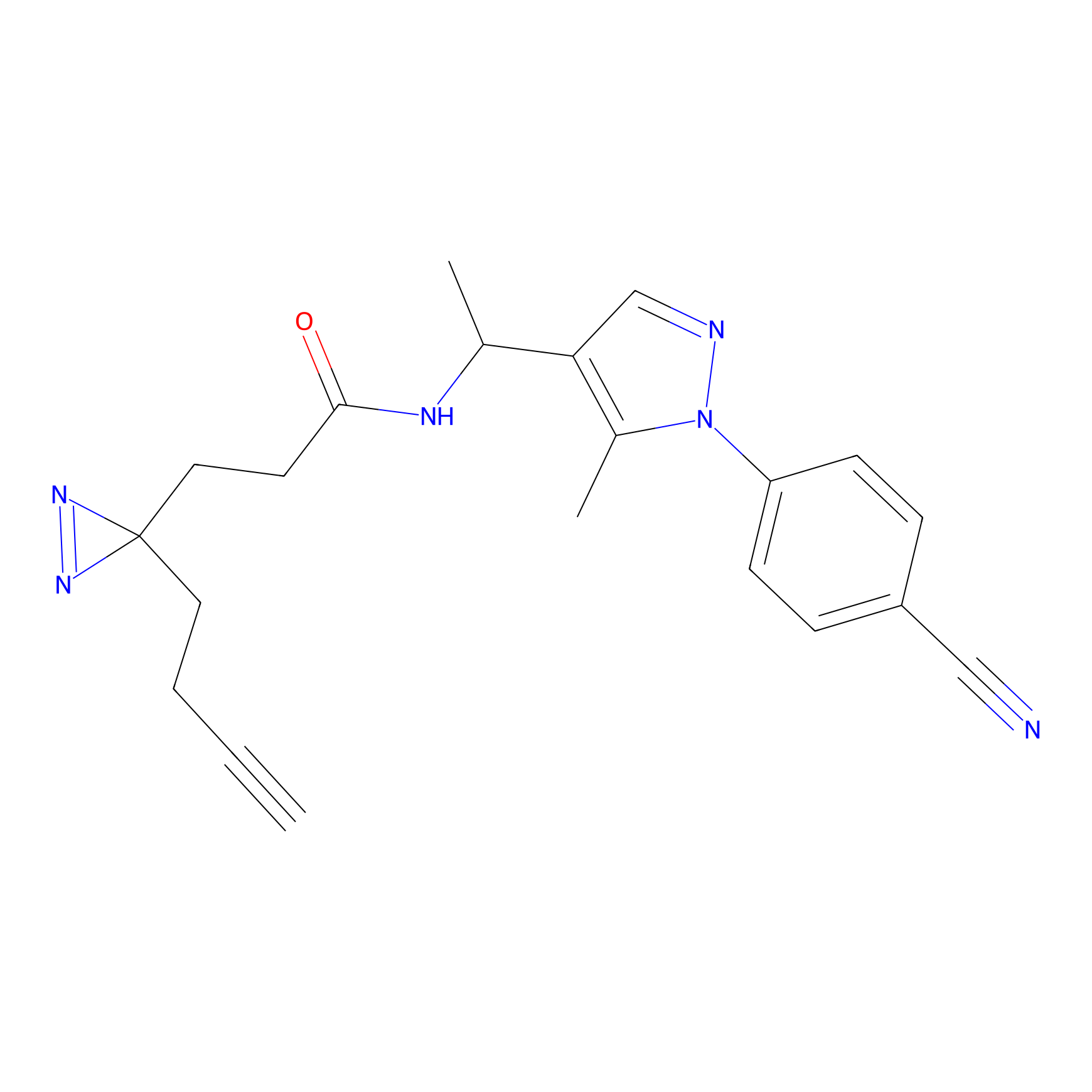
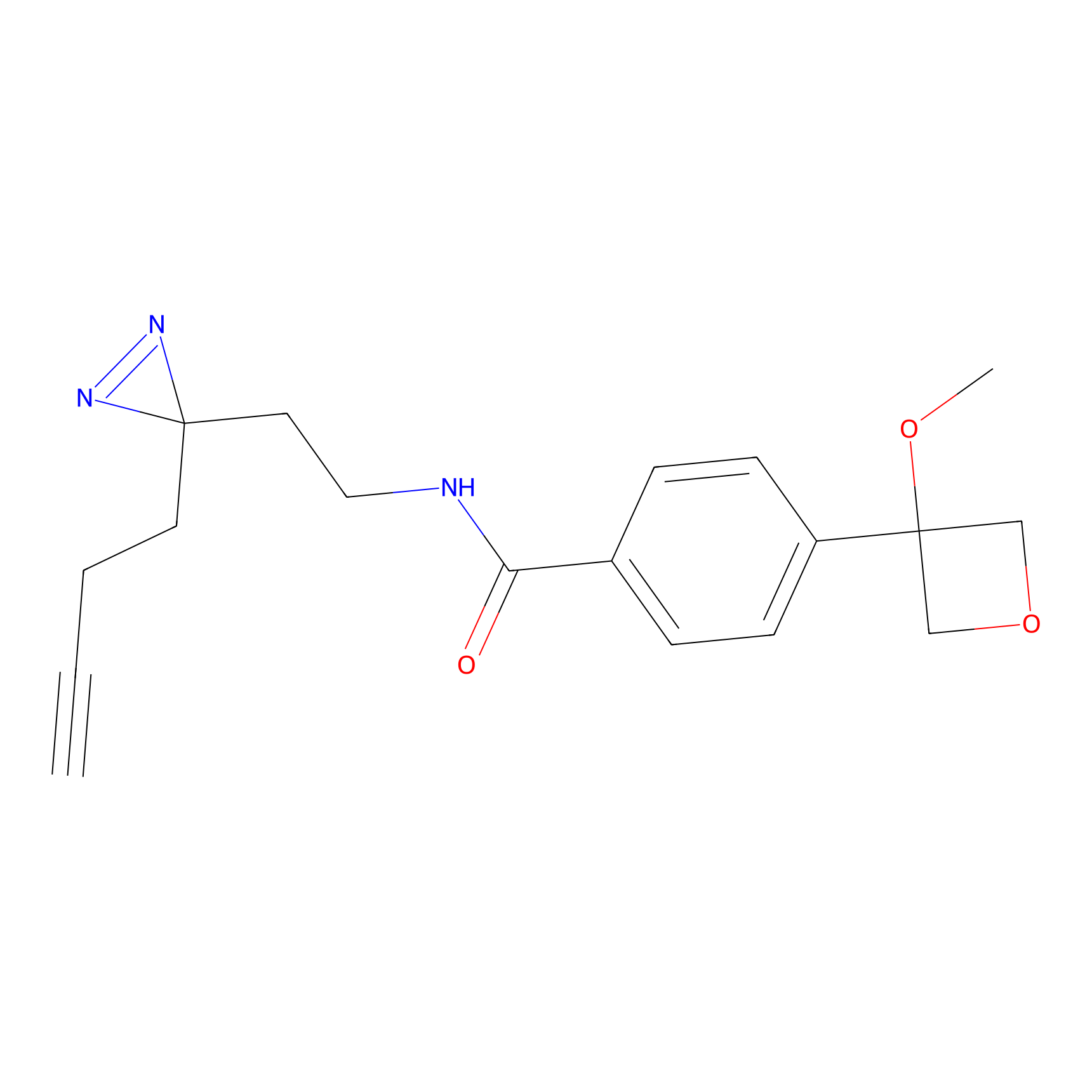
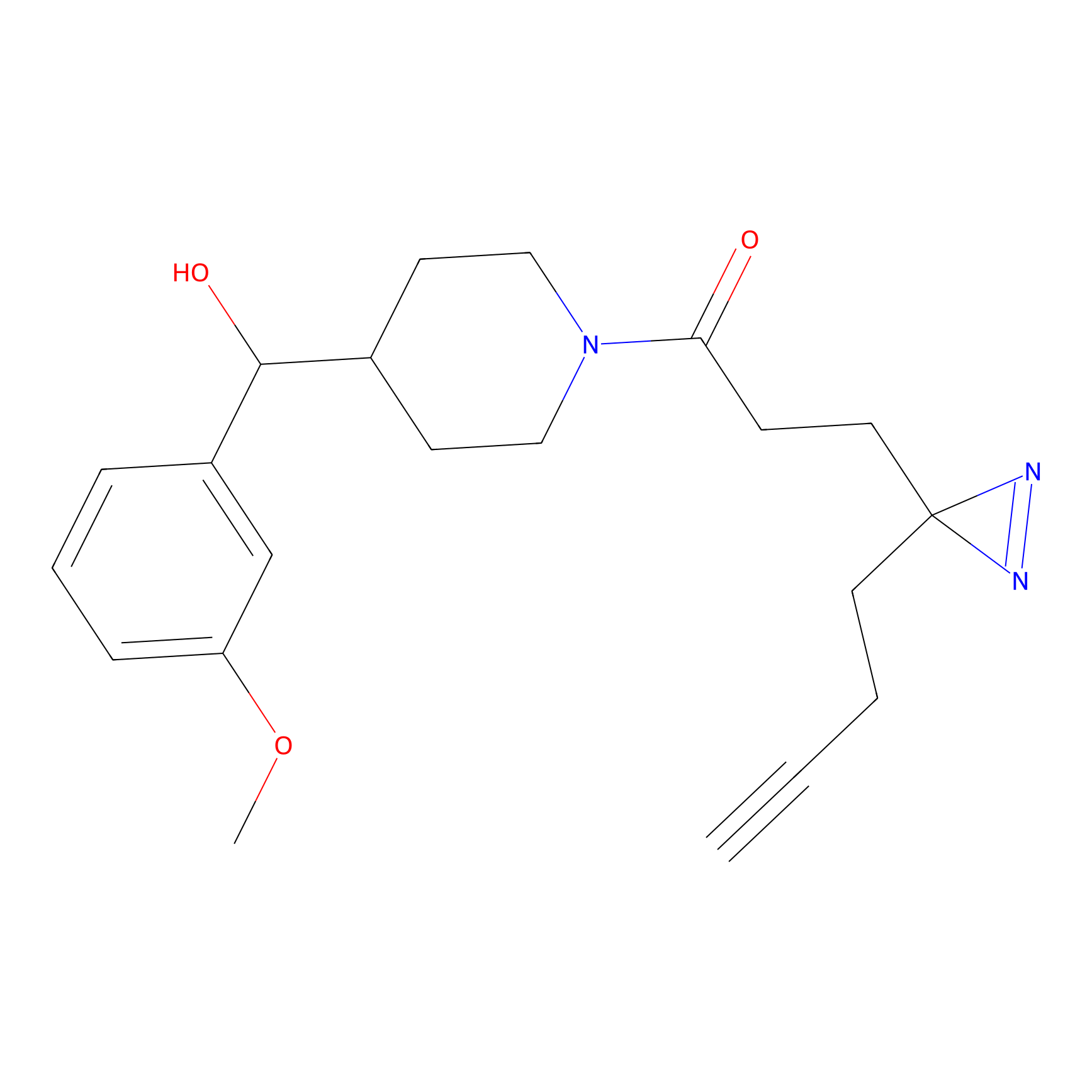
CHEMBL5175495 Probe Info |
 |
10.59 | LDD0196 | [3] | |
|
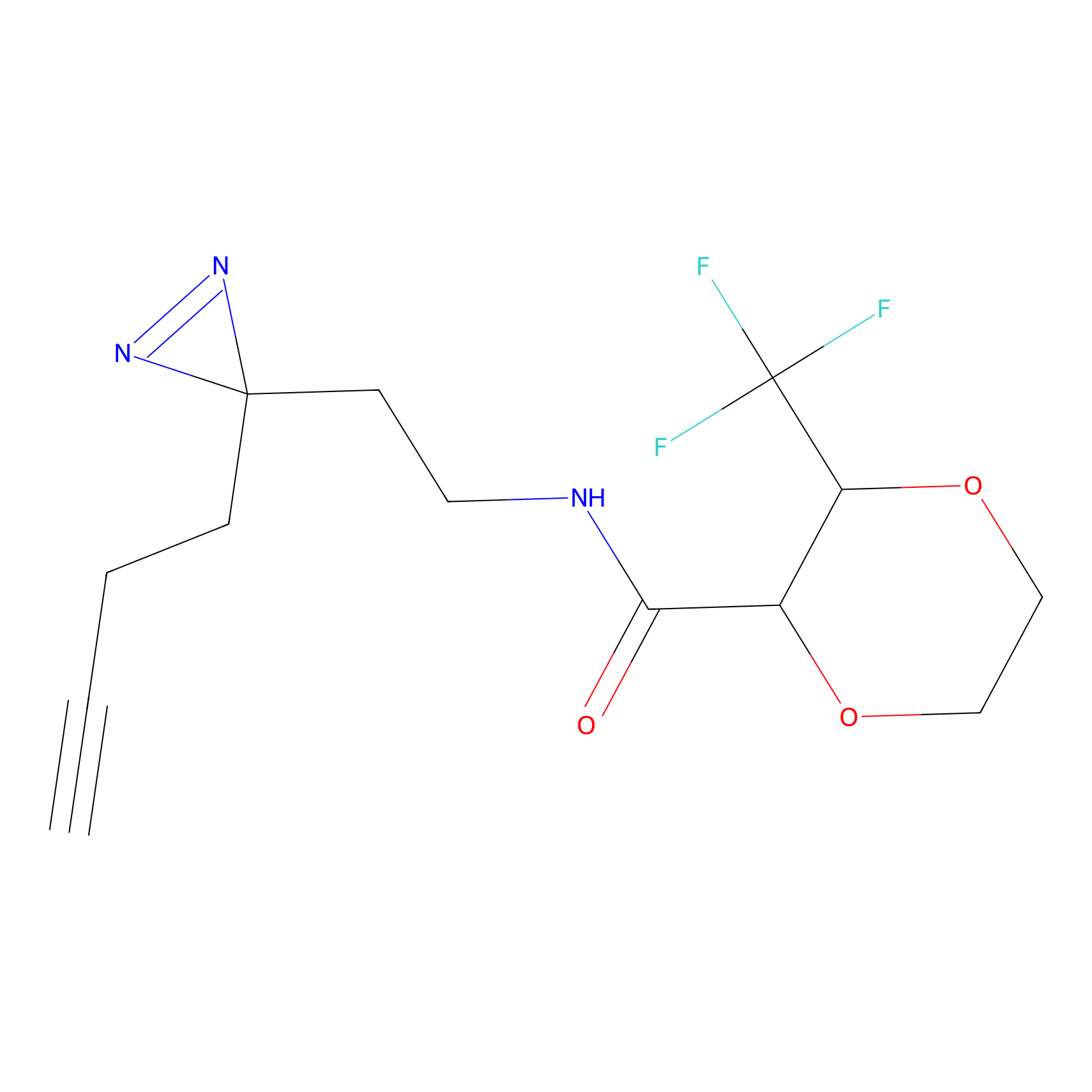
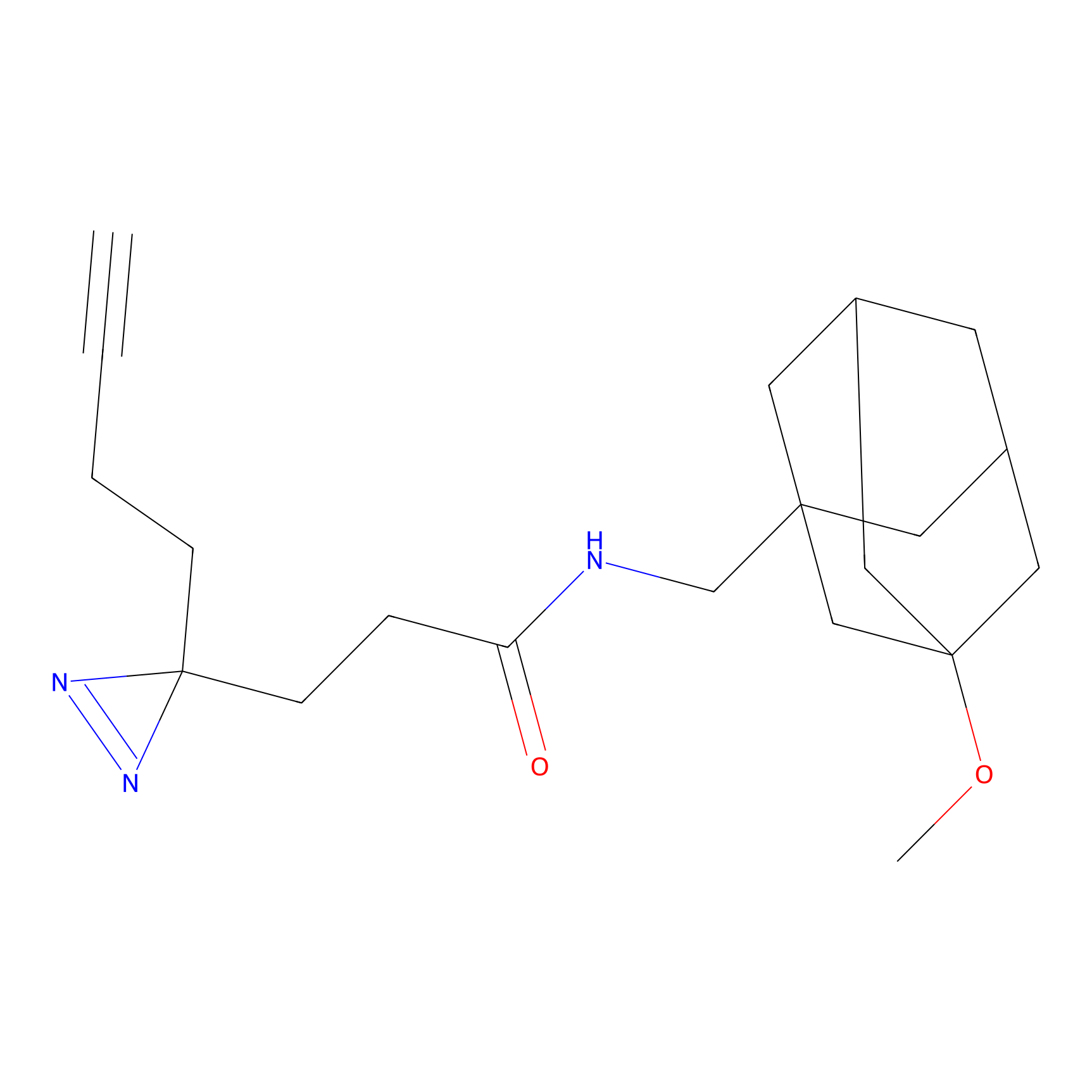
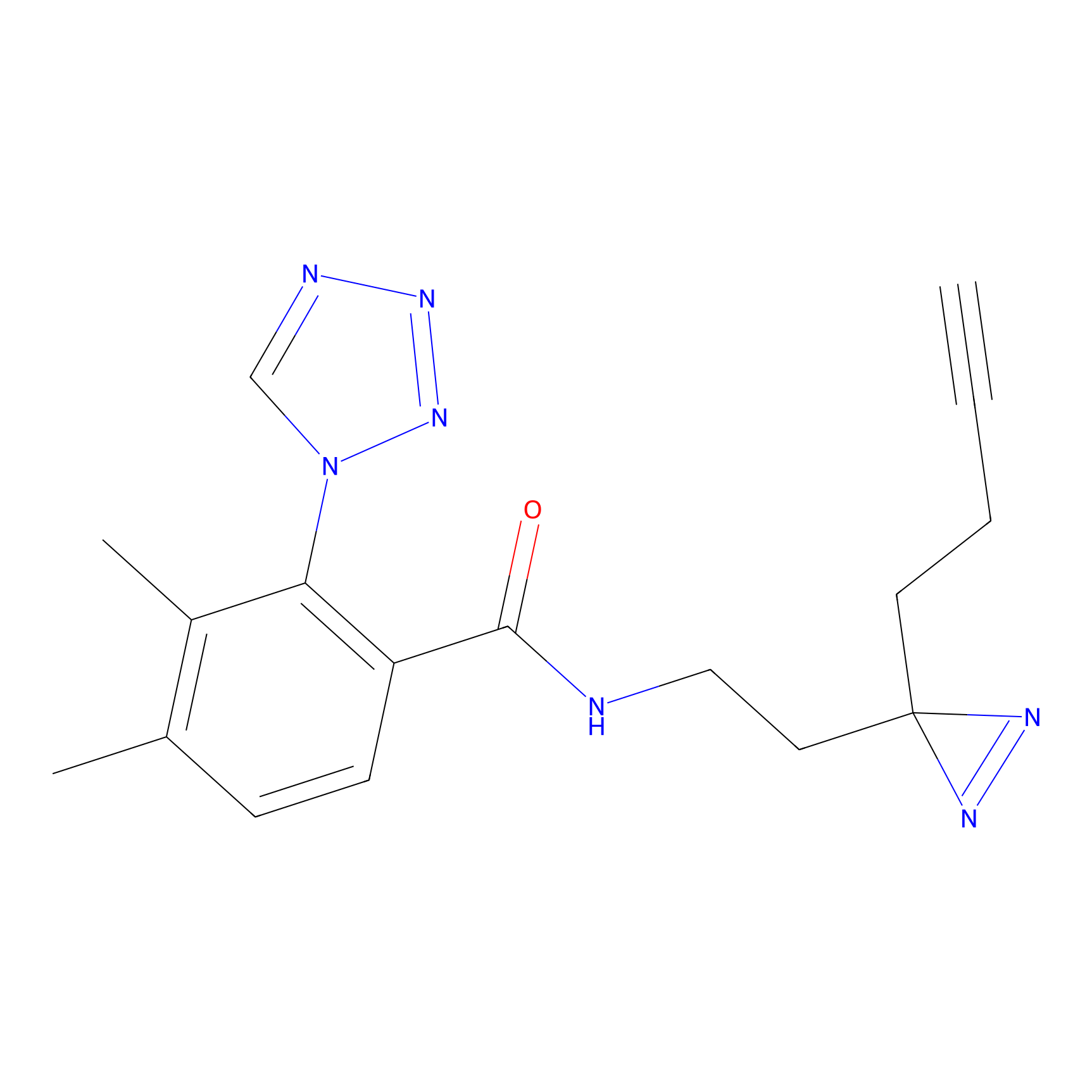
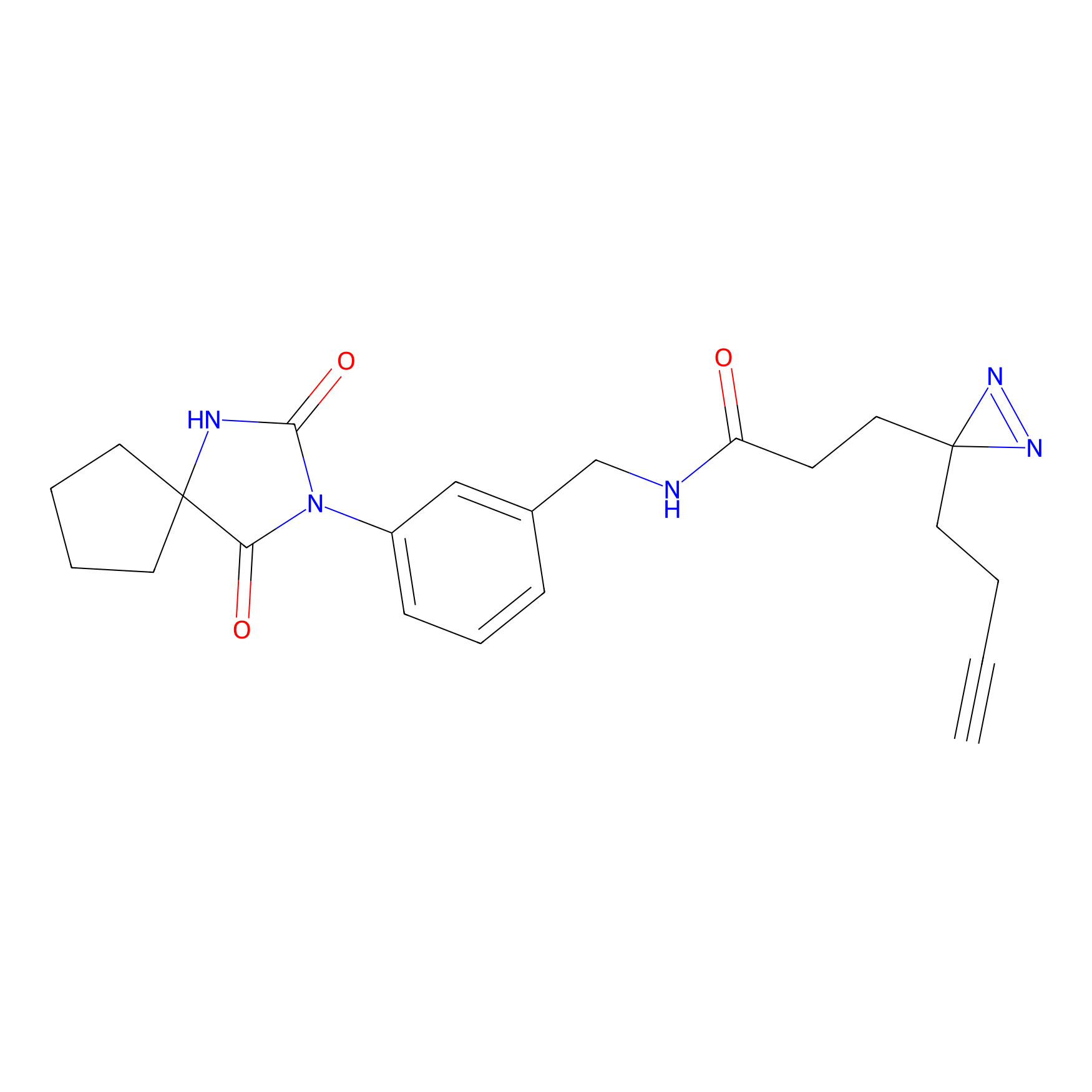
TG42 Probe Info |
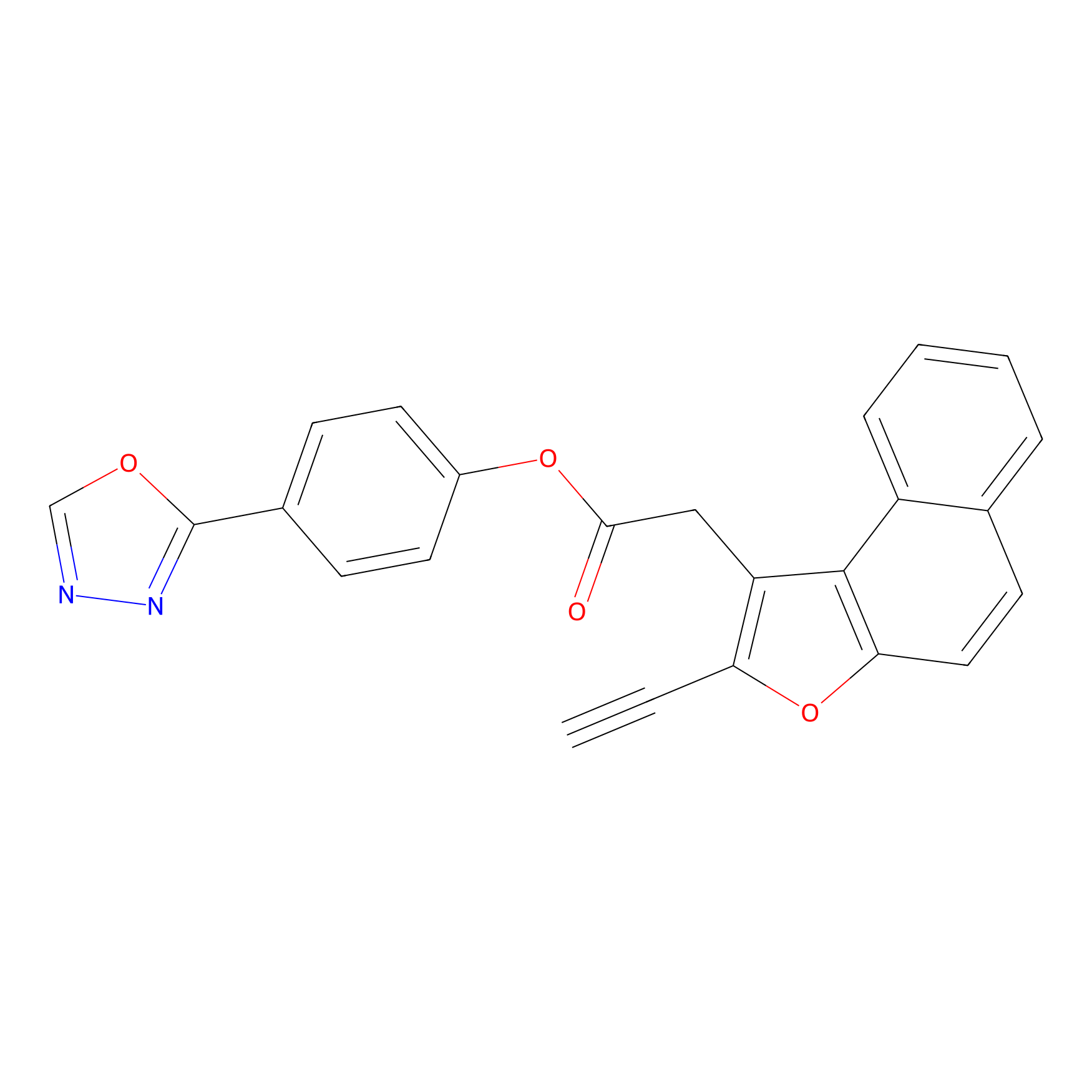 |
12.96 | LDD0326 | [4] | |
|
FBP2 Probe Info |
 |
3.08 | LDD0323 | [1] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
STPyne Probe Info |
 |
K288(3.35) | LDD0277 | [6] | |
|
ONAyne Probe Info |
 |
K42(1.54) | LDD0275 | [6] | |
|
OPA-S-S-alkyne Probe Info |
 |
K42(0.27); K288(0.70) | LDD3494 | [7] | |
|
Propranolol-CA-4PAP Probe Info |
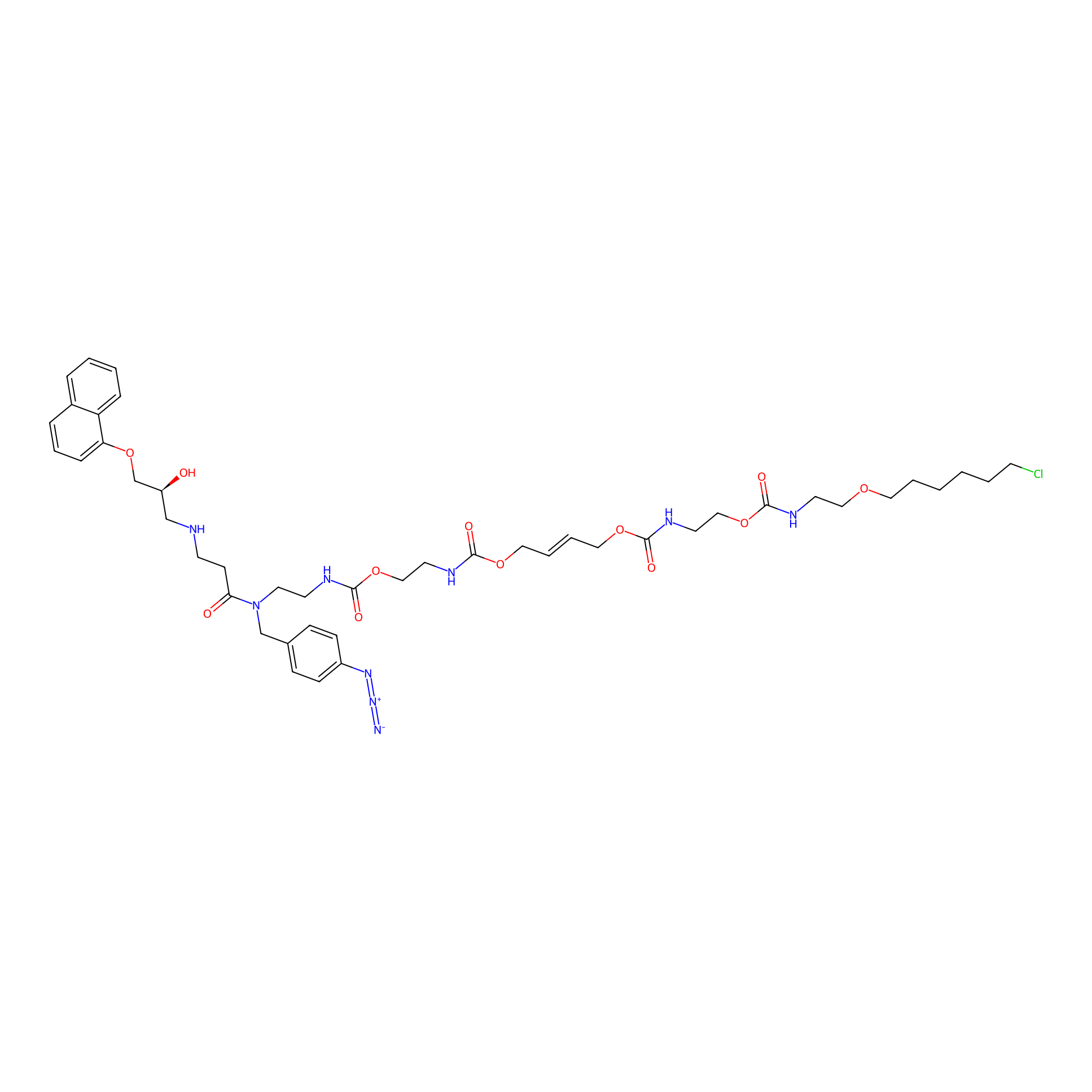 |
4.20 | LDD0271 | [8] | |
|
Jackson_14 Probe Info |
 |
2.22 | LDD0123 | [9] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0166 | [10] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0189 | Compound 16 | HEK-293T | 5.51 | LDD0492 | [12] |
| LDCM0185 | Compound 17 | HEK-293T | 7.30 | LDD0512 | [12] |
| LDCM0184 | Compound 20 | HEK-293T | 14.60 | LDD0503 | [12] |
| LDCM0191 | Compound 21 | HEK-293T | 6.62 | LDD0508 | [12] |
| LDCM0187 | Compound 32 | HEK-293T | 19.61 | LDD0507 | [12] |
| LDCM0190 | Compound 34 | HEK-293T | 11.30 | LDD0497 | [12] |
| LDCM0192 | Compound 35 | HEK-293T | 4.40 | LDD0491 | [12] |
| LDCM0193 | Compound 36 | HEK-293T | 11.05 | LDD0511 | [12] |
| LDCM0194 | Compound 37 | HEK-293T | 5.85 | LDD0498 | [12] |
| LDCM0195 | Compound 38 | HEK-293T | 5.26 | LDD0499 | [12] |
| LDCM0196 | Compound 39 | HEK-293T | 8.85 | LDD0496 | [12] |
| LDCM0197 | Compound 40 | HEK-293T | 14.49 | LDD0495 | [12] |
| LDCM0181 | Compound 41 | HEK-293T | 5.31 | LDD0502 | [12] |
| LDCM0183 | Compound 42 | HEK-293T | 6.99 | LDD0500 | [12] |
| LDCM0098 | Propranolol | K562 | 4.20 | LDD0271 | [8] |
| LDCM0016 | Ranjitkar_cp1 | MDA-MB-231 | 2.22 | LDD0123 | [9] |
| LDCM0084 | Ro 48-8071 | A-549 | 4.77 | LDD0145 | [14] |
| LDCM0137 | SR-4995 | HEK-293T | 3.54 | LDD0334 | [15] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| 1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase gamma-1 (PLCG1) | . | P19174 | |||
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Transcription factor Jun (JUN) | BZIP family | P05412 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Keratin-associated protein 10-9 (KRTAP10-9) | KRTAP type 10 family | P60411 | |||
| Keratin-associated protein 5-9 (KRTAP5-9) | KRTAP type 5 family | P26371 | |||
References