Details of the Target
General Information of Target
| Target ID | LDTP12412 | |||||
|---|---|---|---|---|---|---|
| Target Name | Reticulon-4 (RTN4) | |||||
| Gene Name | RTN4 | |||||
| Gene ID | 57142 | |||||
| Synonyms |
KIAA0886; NOGO; Reticulon-4; Foocen; Neurite outgrowth inhibitor; Nogo protein; Neuroendocrine-specific protein; NSP; Neuroendocrine-specific protein C homolog; RTN-x; Reticulon-5 |
|||||
| 3D Structure | ||||||
| Sequence |
MEEKRRKYSISSDNSDTTDSHATSTSASRCSKLPSSTKSGWPRQNEKKPSEVFRTDLITA
MKIPDSYQLSPDDYYILADPWRQEWEKGVQVPAGAEAIPEPVVRILPPLEGPPAQASPSS TMLGEGSQPDWPGGSRYDLDEIDAYWLELINSELKEMERPELDELTLERVLEELETLCHQ NMARAIETQEGLGIEYDEDVVCDVCRSPEGEDGNEMVFCDKCNVCVHQACYGILKVPTGS WLCRTCALGVQPKCLLCPKRGGALKPTRSGTKWVHVSCALWIPEVSIGCPEKMEPITKIS HIPASRWALSCSLCKECTGTCIQCSMPSCVTAFHVTCAFDHGLEMRTILADNDEVKFKSF CQEHSDGGPRNEPTSEPTEPSQAGEDLEKVTLRKQRLQQLEEDFYELVEPAEVAERLDLA EALVDFIYQYWKLKRKANANQPLLTPKTDEVDNLAQQEQDVLYRRLKLFTHLRQDLERVR NLCYMVTRRERTKHAICKLQEQIFHLQMKLIEQDLCRGLSTSFPIDGTFFNSWLAQSVQI TAENMAMSEWPLNNGHREDPAPGLLSEELLQDEETLLSFMRDPSLRPGDPARKARGRTRL PAKKKPPPPPPQDGPGSRTTPDKAPKKTWGQDAGSGKGGQGPPTRKPPRRTSSHLPSSPA AGDCPILATPESPPPLAPETPDEAASVAADSDVQVPGPAASPKPLGRLRPPRESKVTRRL PGARPDAGMGPPSAVAERPKVSLHFDTETDGYFSDGEMSDSDVEAEDGGVQRGPREAGAE EVVRMGVLAS |
|||||
| Target Type |
Clinical trial
|
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Required to induce the formation and stabilization of endoplasmic reticulum (ER) tubules. They regulate membrane morphogenesis in the ER by promoting tubular ER production. They influence nuclear envelope expansion, nuclear pore complex formation and proper localization of inner nuclear membrane proteins. However each isoform have specific functions mainly depending on their tissue expression specificities (Probable).; [Isoform A]: Developmental neurite growth regulatory factor with a role as a negative regulator of axon-axon adhesion and growth, and as a facilitator of neurite branching. Regulates neurite fasciculation, branching and extension in the developing nervous system. Involved in down-regulation of growth, stabilization of wiring and restriction of plasticity in the adult CNS. Regulates the radial migration of cortical neurons via an RTN4R-LINGO1 containing receptor complex. Acts as a negative regulator of central nervous system angiogenesis. Inhibits spreading, migration and sprouting of primary brain microvascular endothelial cells (MVECs). Also induces the retraction of MVECs lamellipodia and filopodia in a ROCK pathway-dependent manner.; [Isoform B]: Mainly function in endothelial cells and vascular smooth muscle cells, is also involved in immune system regulation (Probable). Modulator of vascular remodeling, promotes the migration of endothelial cells but inhibits the migration of vascular smooth muscle cells. Regulates endothelial sphingolipid biosynthesis with direct effects on vascular function and blood pressure. Inhibits serine palmitoyltransferase, SPTLC1, the rate-limiting enzyme of the novo sphingolipid biosynthetic pathway, thereby controlling production of endothelial sphingosine-1-phosphate (S1P). Required to promote macrophage homing and functions such as cytokine/chemokine gene expression involved in angiogenesis, arteriogenesis and tissue repair. Mediates ICAM1 induced transendothelial migration of leukocytes such as monocytes and neutrophils and acute inflammation. Necessary for immune responses triggered by nucleic acid sensing TLRs, such as TLR9, is required for proper TLR9 location to endolysosomes. Also involved in immune response to LPS. Plays a role in liver regeneration through the modulation of hepatocytes proliferation. Reduces the anti-apoptotic activity of Bcl-xl and Bcl-2. This is likely consecutive to their change in subcellular location, from the mitochondria to the endoplasmic reticulum, after binding and sequestration. With isoform C, inhibits BACE1 activity and amyloid precursor protein processing.; [Isoform C]: Regulates cardiomyocyte apoptosis upon hypoxic conditions. With isoform B, inhibits BACE1 activity and amyloid precursor protein processing.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 22RV1 | Insertion: p.V100RfsTer36 | DBIA Probe Info | |||
| AN3CA | SNV: p.L49Q | . | |||
| CAL78 | SNV: p.D700A | DBIA Probe Info | |||
| CCK81 | Deletion: p.V670YfsTer4 | DBIA Probe Info | |||
| CHL1 | SNV: p.L788F | DBIA Probe Info | |||
| HCC70 | SNV: p.S1030L | DBIA Probe Info | |||
| HT | Insertion: p.P658TfsTer3 | . | |||
| HT115 | SNV: p.E30G | . | |||
| MCC13 | SNV: p.P598S | DBIA Probe Info | |||
| MFE319 | SNV: p.D761N | DBIA Probe Info | |||
| MOLT4 | SNV: p.F85L; p.V972I | IA-alkyne Probe Info | |||
| SG231 | SNV: p.V1039I | DBIA Probe Info | |||
| SNU5 | SNV: p.R91G | DBIA Probe Info | |||
| SUPT1 | Insertion: p.P658TfsTer3 | DBIA Probe Info | |||
| TE10 | SNV: p.S114L | DBIA Probe Info | |||
| TGBC1TKB | SNV: p.S129W | DBIA Probe Info | |||
| VMRCRCW | SNV: p.E56D | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
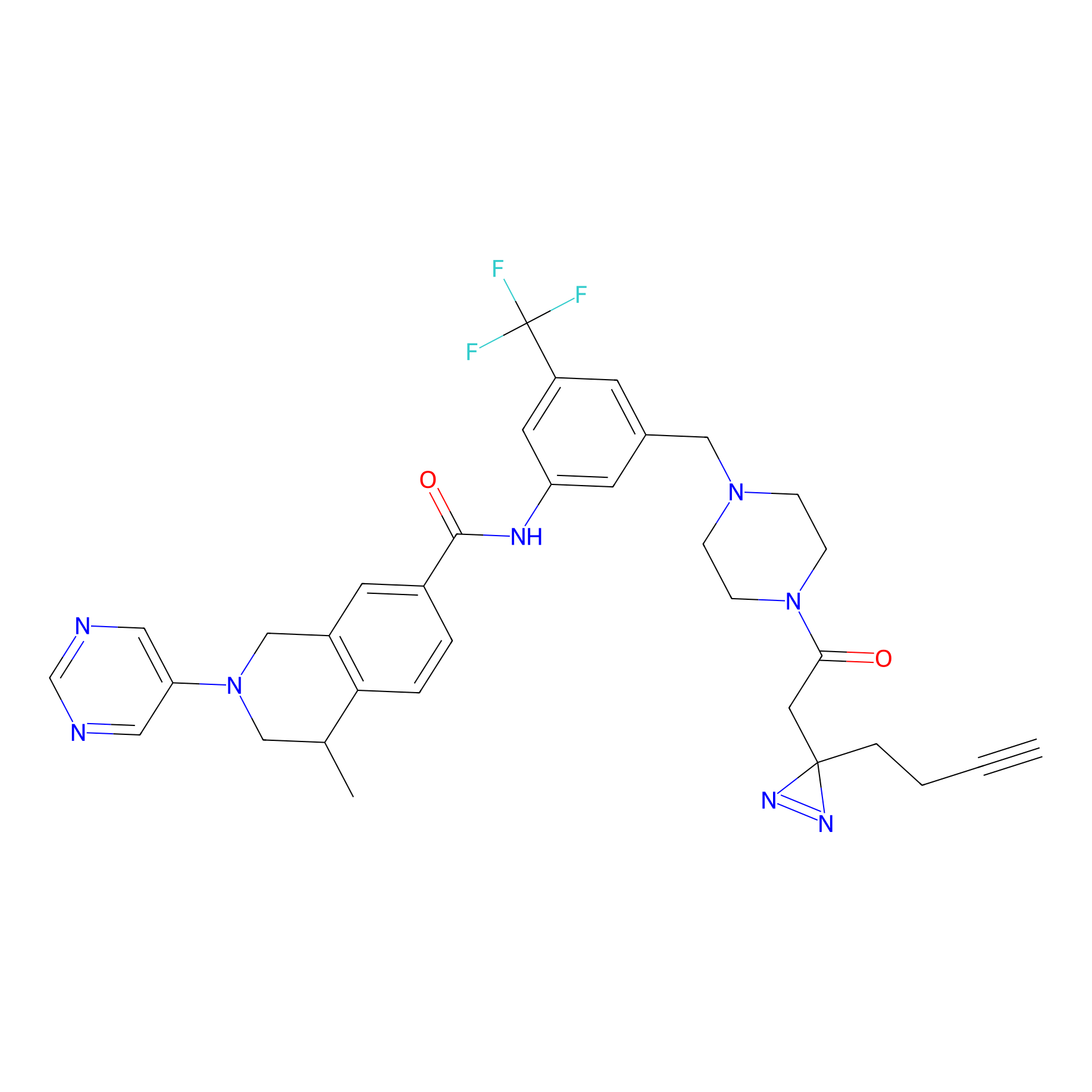
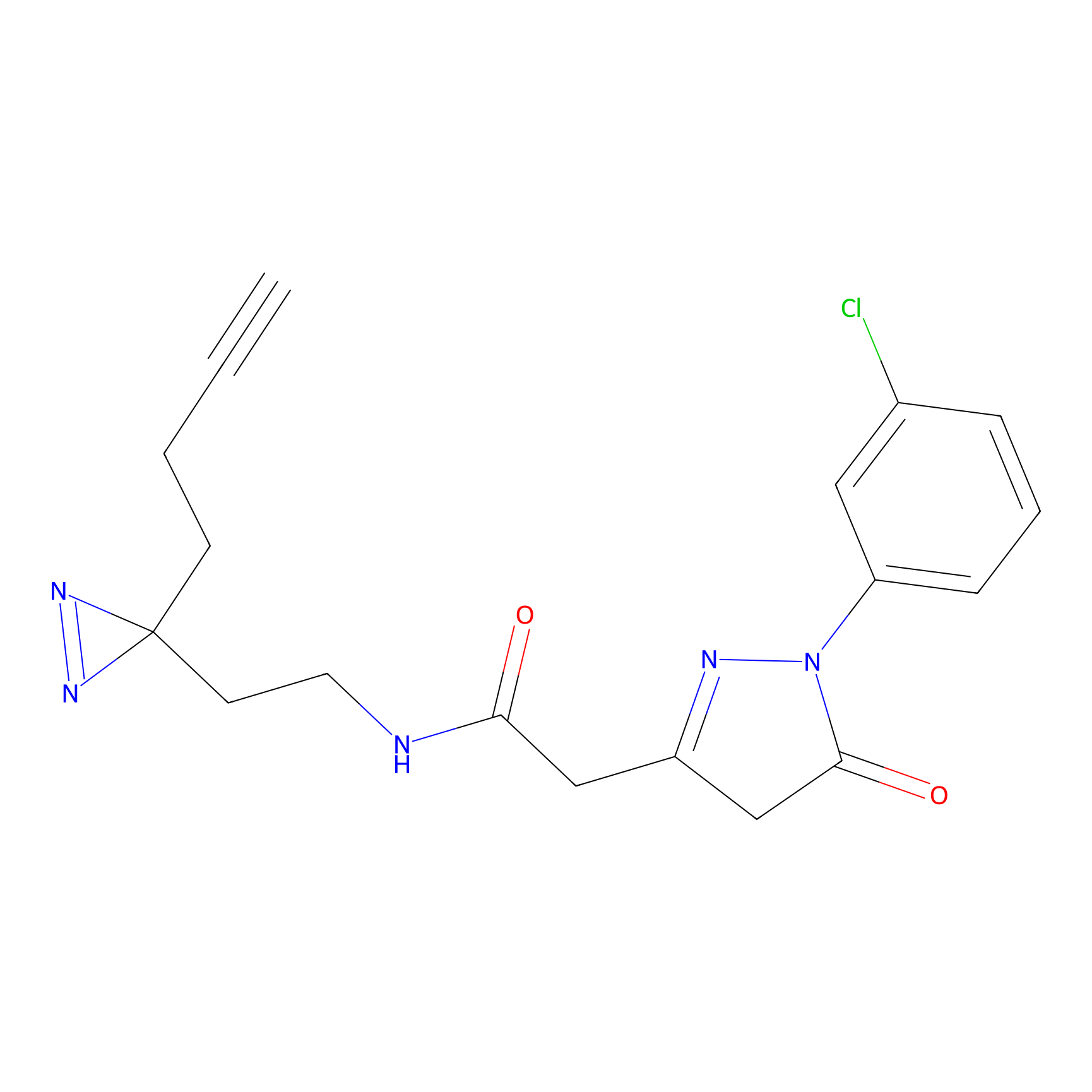
m-APA Probe Info |
 |
12.52 | LDD0402 | [2] | |
|
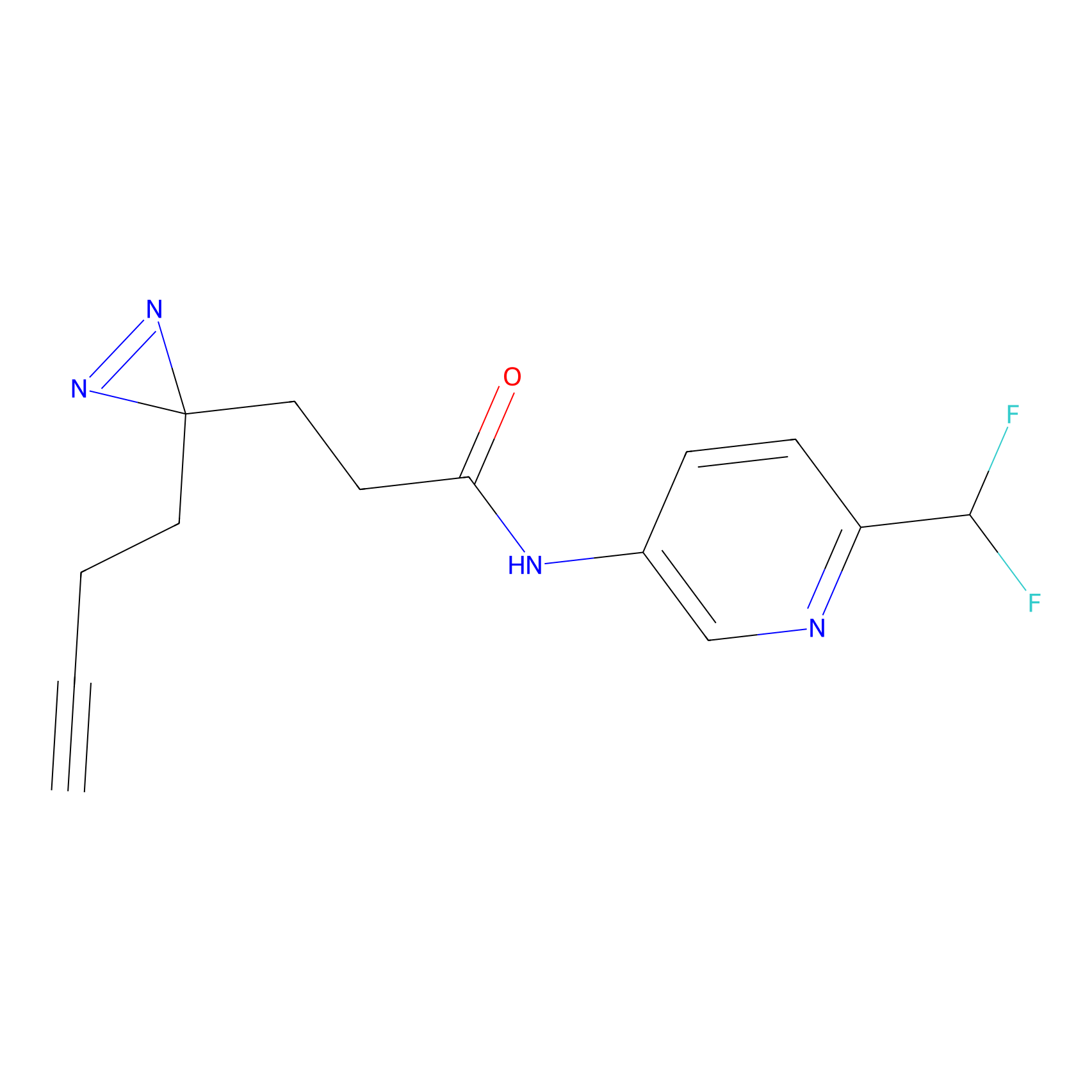
HDSF-alk Probe Info |
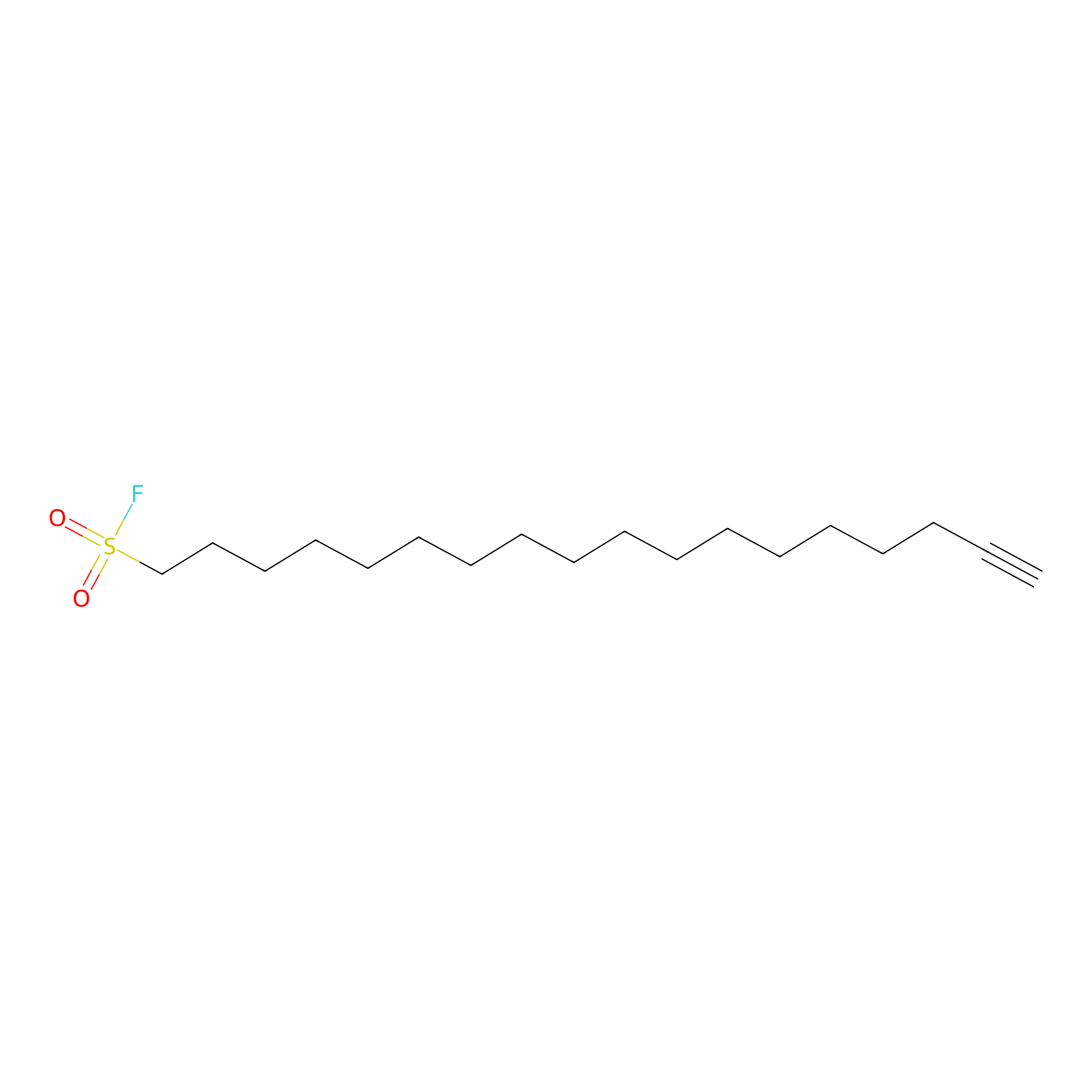 |
2.94 | LDD0197 | [3] | |
|
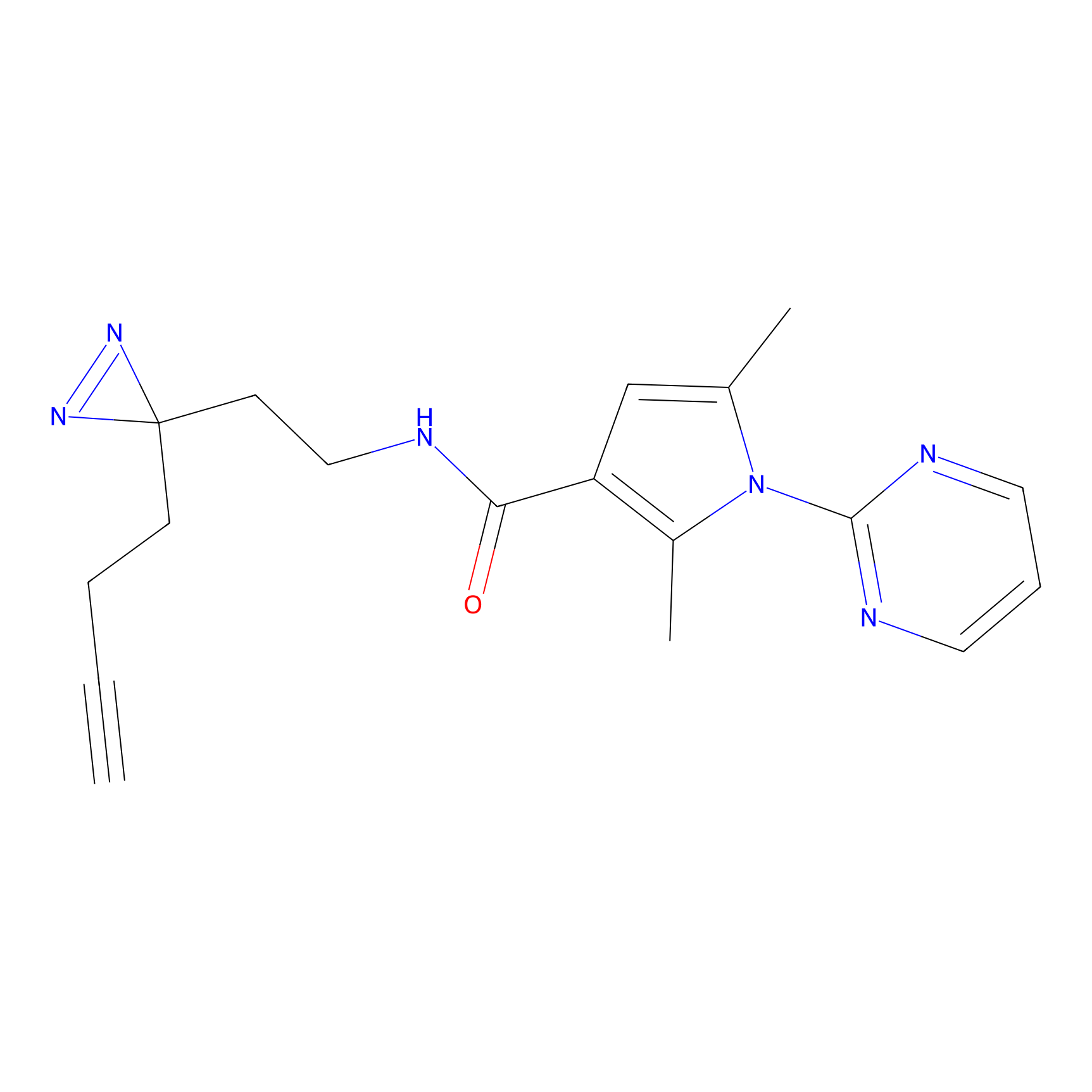
P3 Probe Info |
 |
10.00 | LDD0450 | [4] | |
|
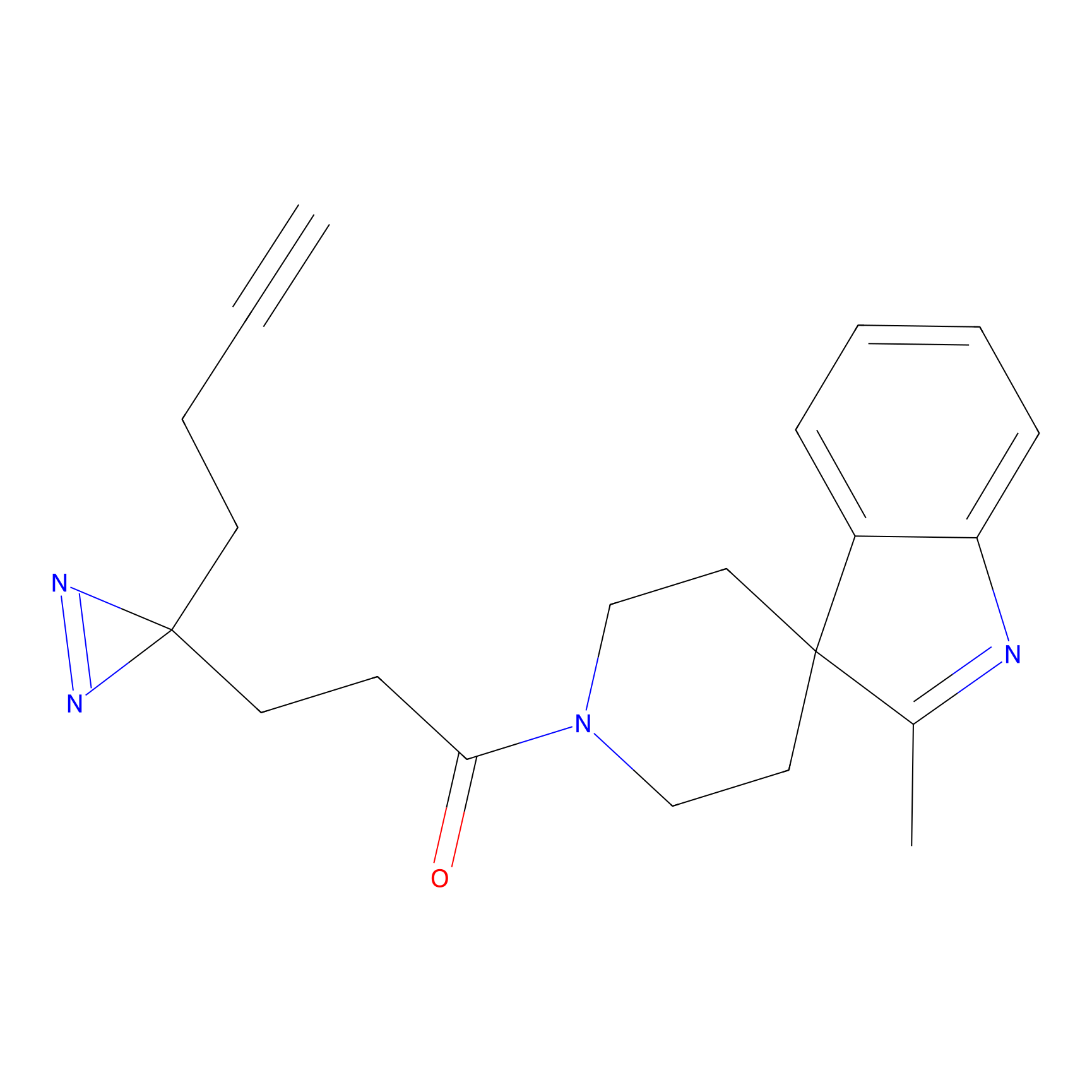
P8 Probe Info |
 |
10.00 | LDD0451 | [4] | |
|
A-EBA Probe Info |
 |
3.71 | LDD0215 | [5] | |
|
EN219-alkyne Probe Info |
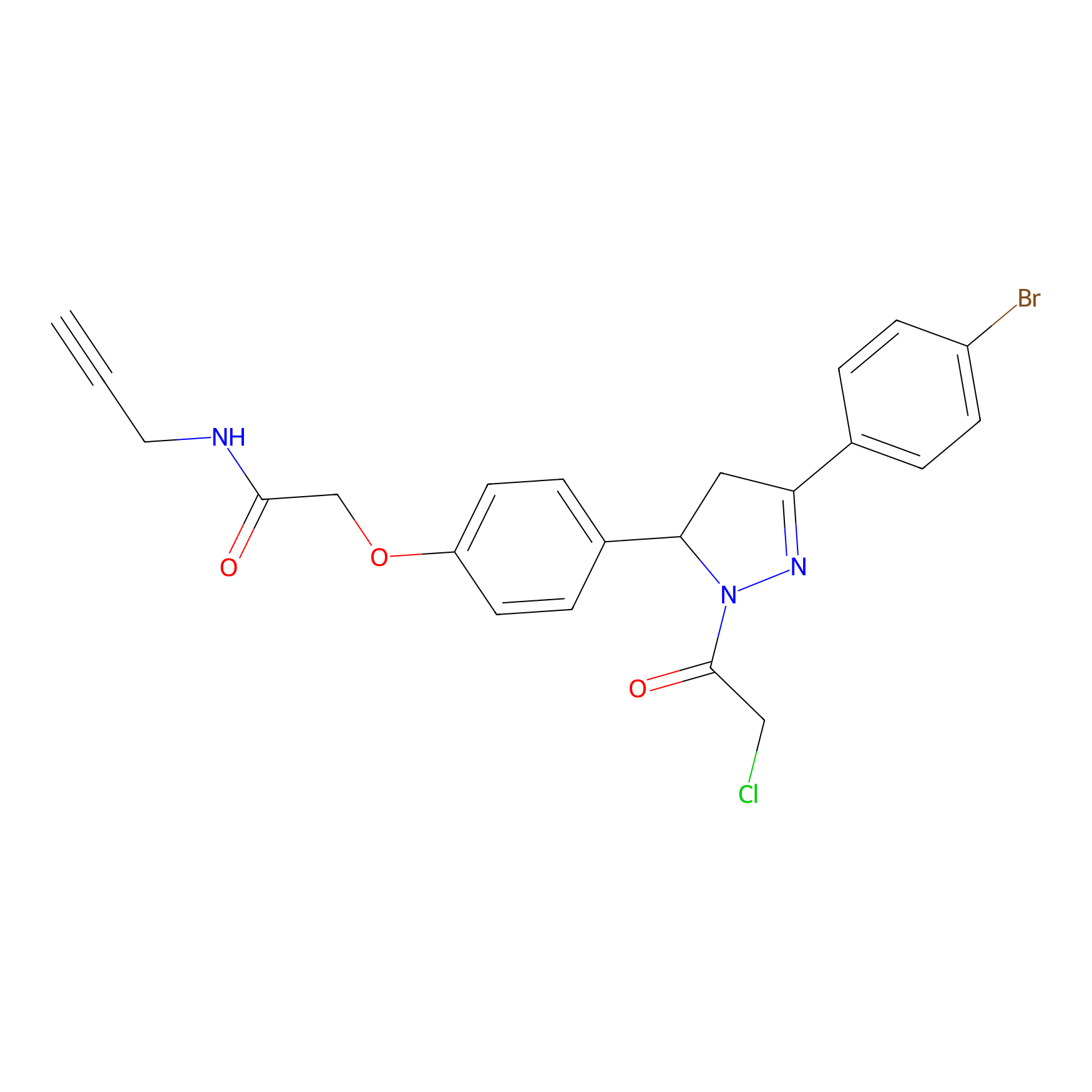 |
3.37 | LDD0297 | [6] | |
|
ILS-2 Probe Info |
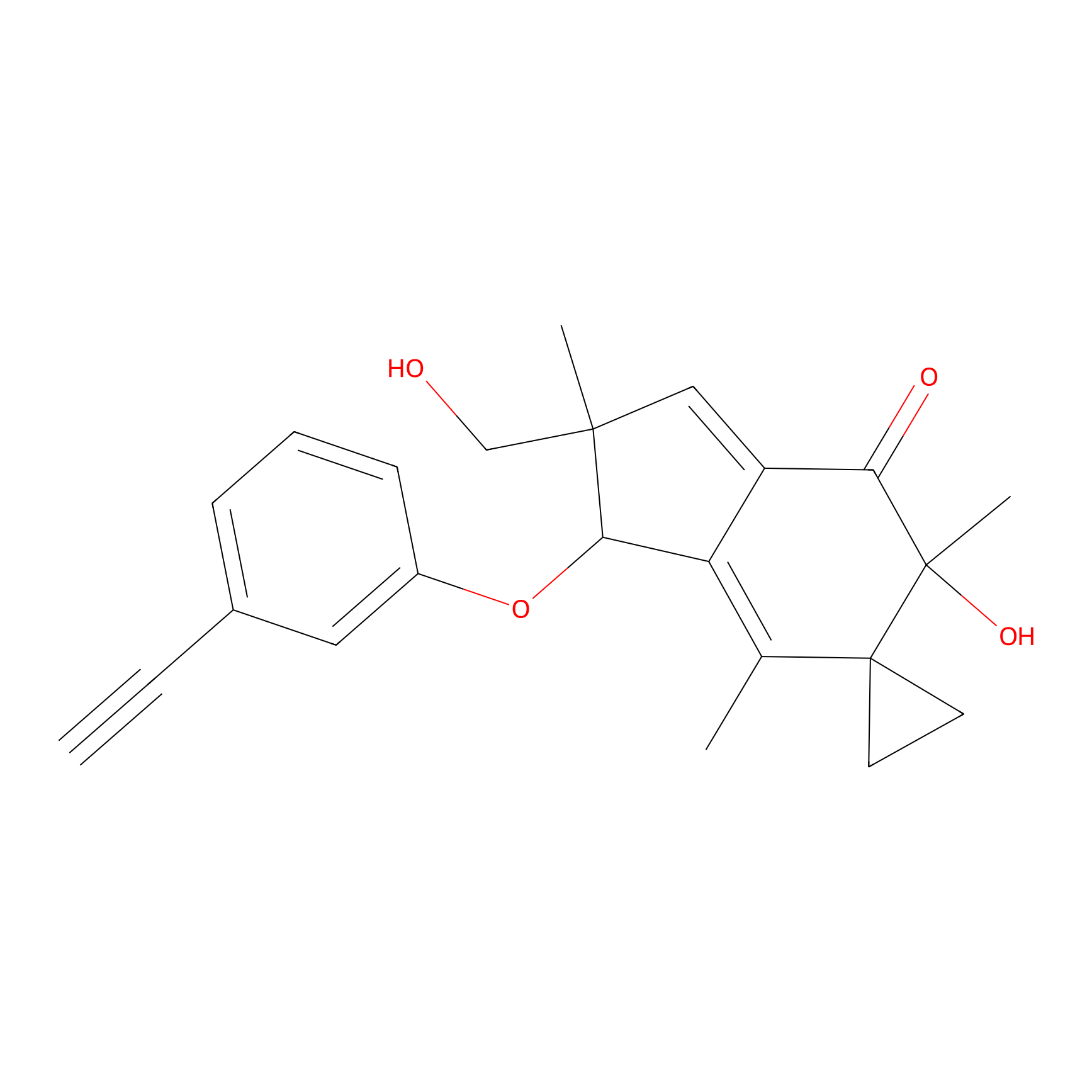 |
2.40 | LDD0416 | [7] | |
|
AZ-9 Probe Info |
 |
2.36 | LDD0393 | [8] | |
|
FBPP2 Probe Info |
 |
5.35 | LDD0318 | [9] | |
|
CHEMBL5175495 Probe Info |
 |
16.08 | LDD0196 | [10] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [11] | |
|
SAA-alkyne Probe Info |
 |
1.11 | LDD0252 | [12] | |
|
TG42 Probe Info |
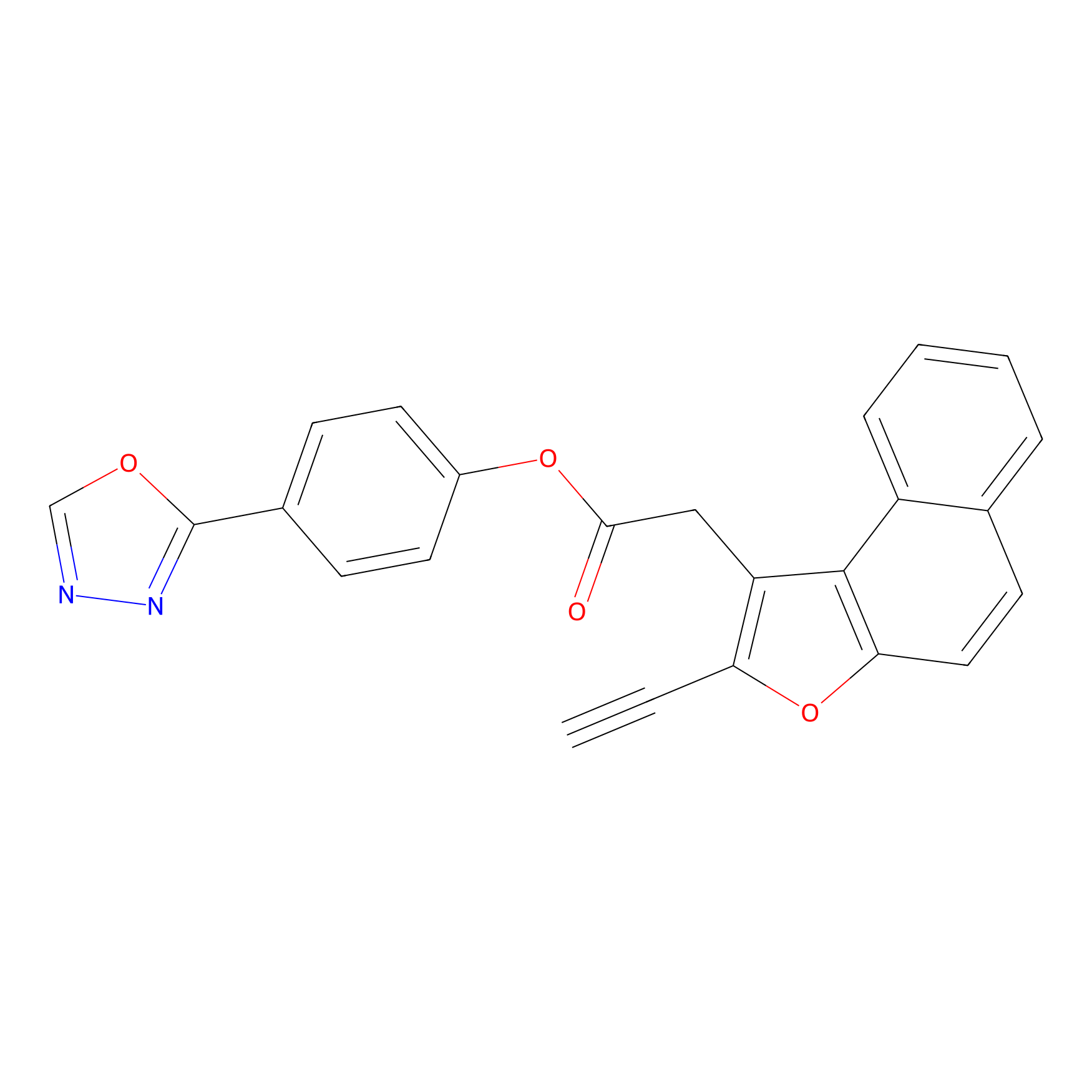 |
12.20 | LDD0043 | [13] | |
|
W1 Probe Info |
 |
17.27 | LDD0235 | [14] | |
|
FBP2 Probe Info |
 |
5.99 | LDD0317 | [9] | |
|
TH211 Probe Info |
 |
Y1165(9.23); Y1076(5.68) | LDD0257 | [15] | |
|
TH214 Probe Info |
 |
Y1091(20.00) | LDD0258 | [15] | |
|
TH216 Probe Info |
 |
Y1091(20.00); Y1165(12.62) | LDD0259 | [15] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [16] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [16] | |
|
ONAyne Probe Info |
 |
K1066(0.18); K1174(0.92); K1179(1.07); K1183(6.55) | LDD0274 | [17] | |
|
OPA-S-S-alkyne Probe Info |
 |
K1104(1.32); K1066(3.40) | LDD3494 | [18] | |
|
Alkylaryl probe 2 Probe Info |
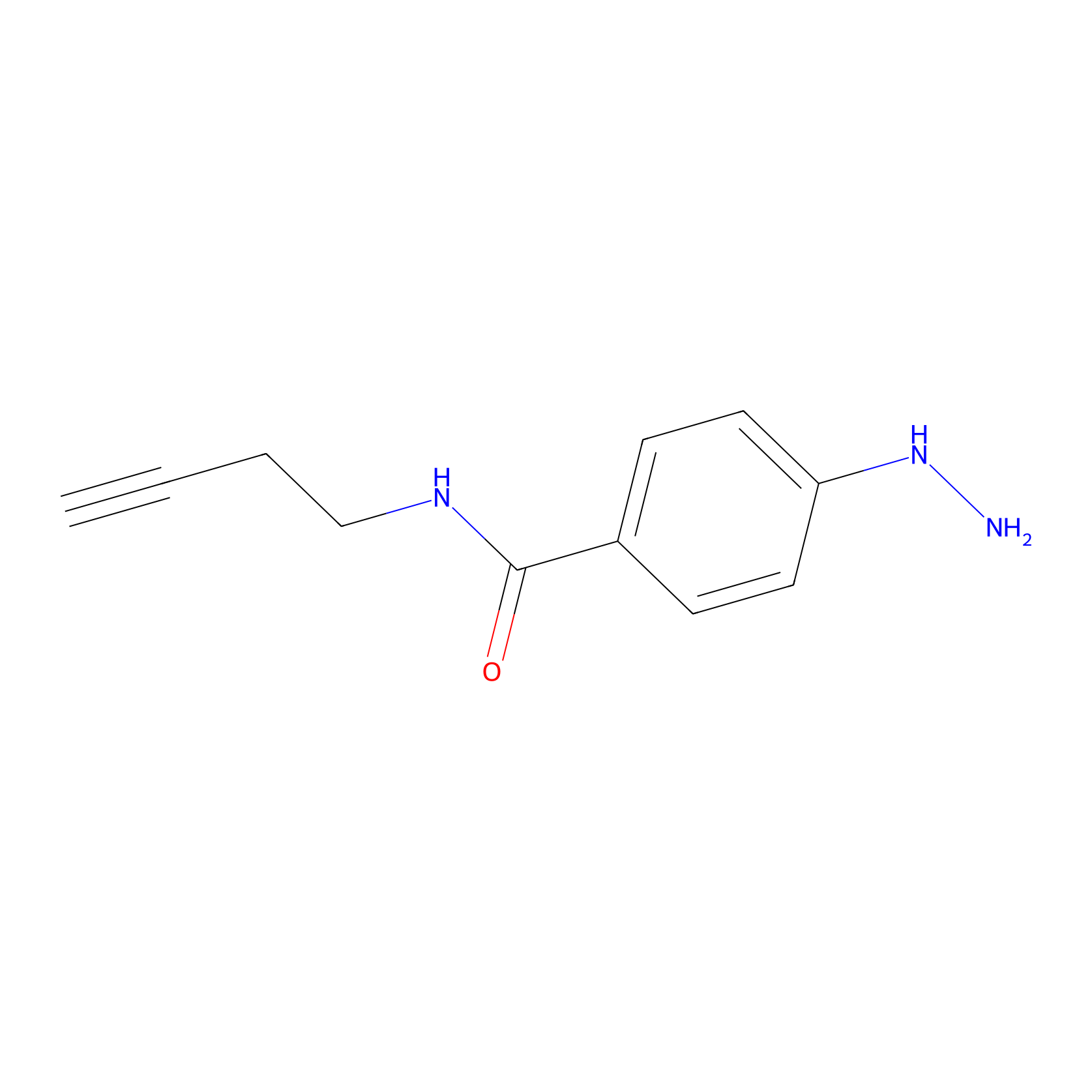 |
11.00 | LDD0390 | [19] | |
|
P13 Probe Info |
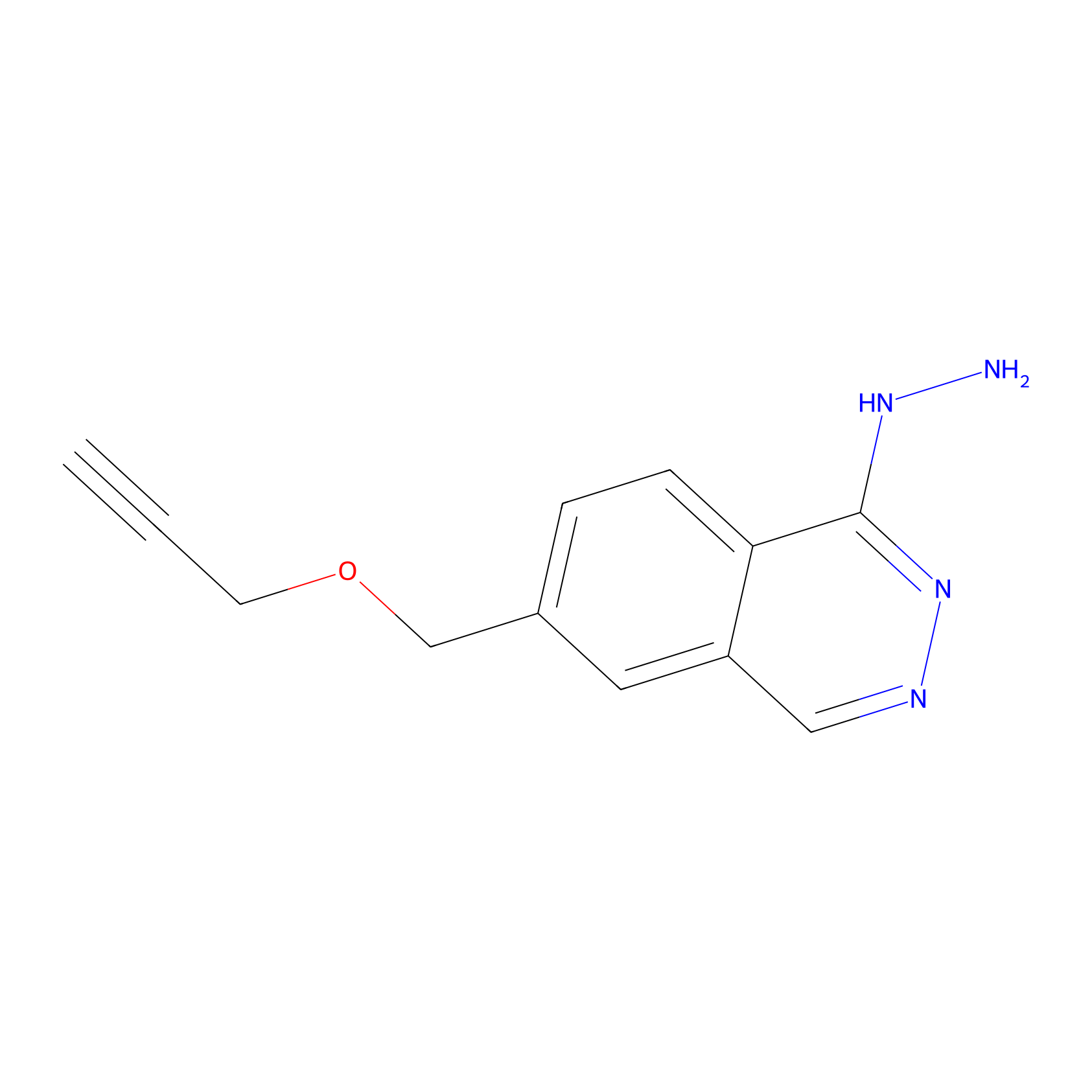 |
6.04 | LDD0203 | [20] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [21] | |
|
THZ1-DTB Probe Info |
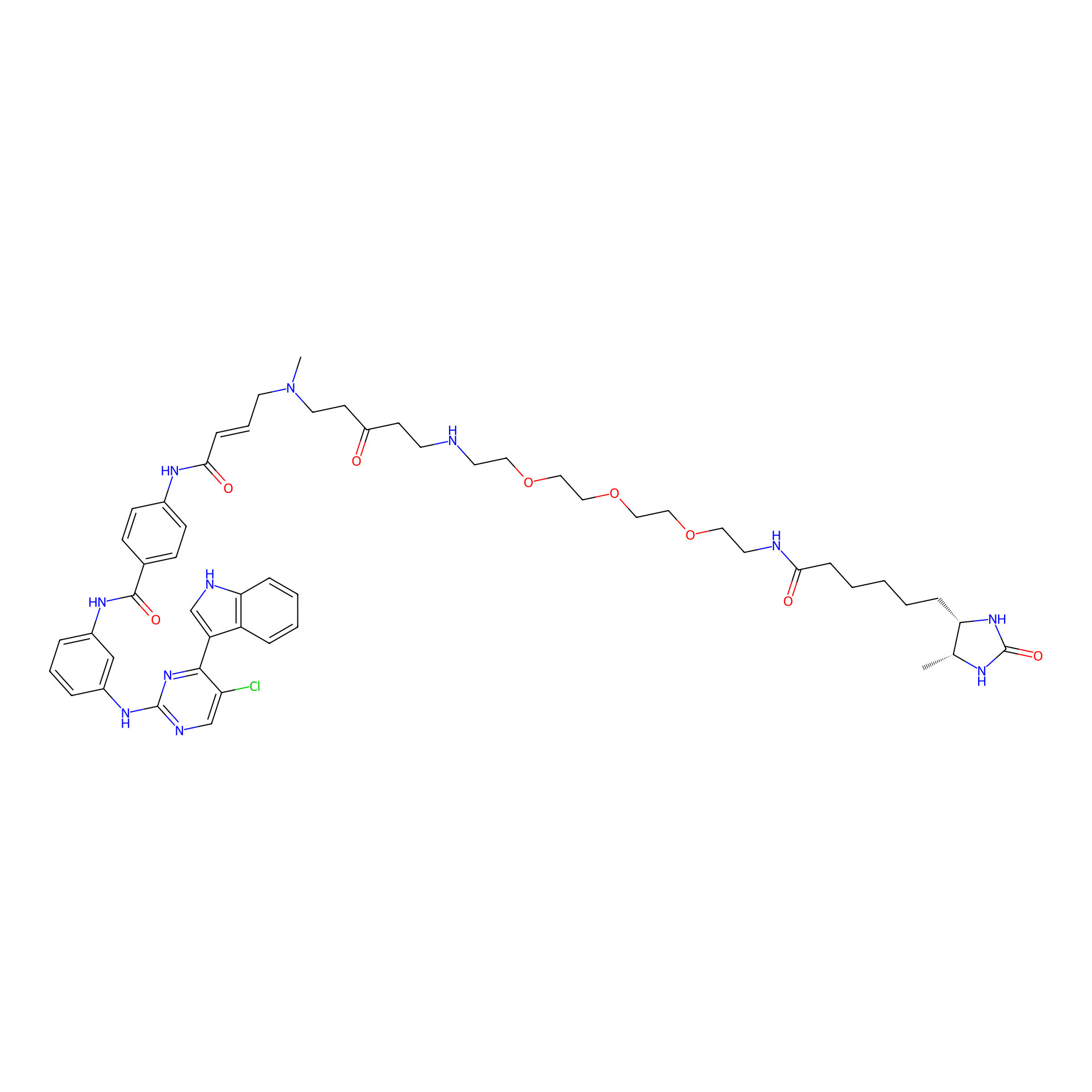 |
C1110(1.18) | LDD0460 | [21] | |
|
Johansson_61 Probe Info |
 |
_(15.99) | LDD1489 | [22] | |
|
YY4-yne Probe Info |
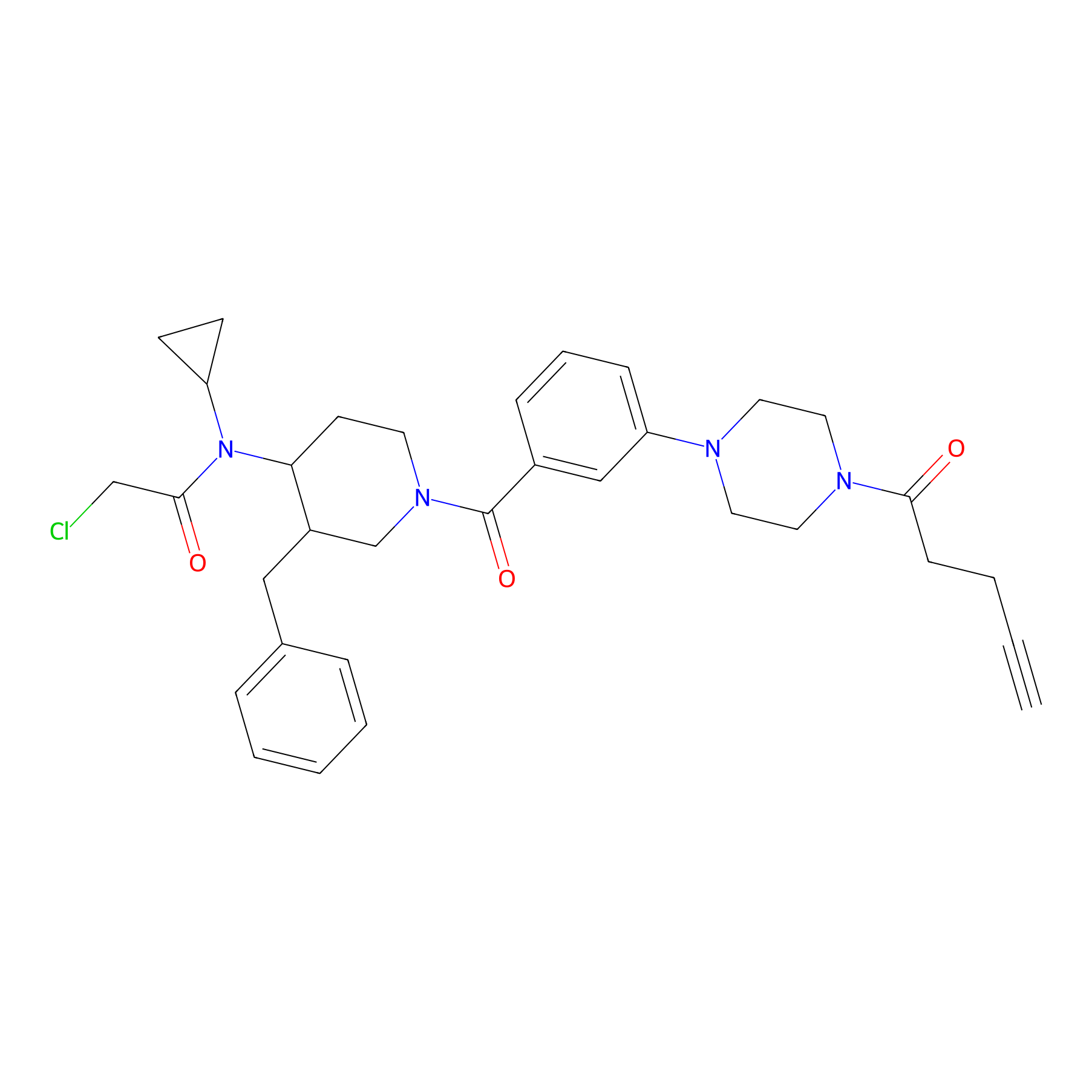 |
2.92 | LDD0400 | [23] | |
|
DA-P3 Probe Info |
 |
36.56 | LDD0179 | [24] | |
|
AHL-Pu-1 Probe Info |
 |
C1101(2.06) | LDD0170 | [25] | |
|
HPAP Probe Info |
 |
5.72 | LDD0064 | [26] | |
|
HHS-482 Probe Info |
 |
Y1076(0.19) | LDD0292 | [27] | |
|
DBIA Probe Info |
 |
C1101(1.77) | LDD0078 | [28] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [29] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0358 | [30] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C912(0.00); C1101(0.00) | LDD0038 | [31] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [32] | |
|
Lodoacetamide azide Probe Info |
 |
C912(0.00); C464(0.00); C1101(0.00) | LDD0037 | [31] | |
|
ATP probe Probe Info |
 |
K1183(0.00); K1171(0.00) | LDD0035 | [33] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [34] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [35] | |
|
NAIA_4 Probe Info |
 |
C699(0.00); C912(0.00); C1101(0.00) | LDD2226 | [36] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2224 | [36] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [35] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [34] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [34] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [37] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [37] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [34] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [34] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [38] | |
|
PPMS Probe Info |
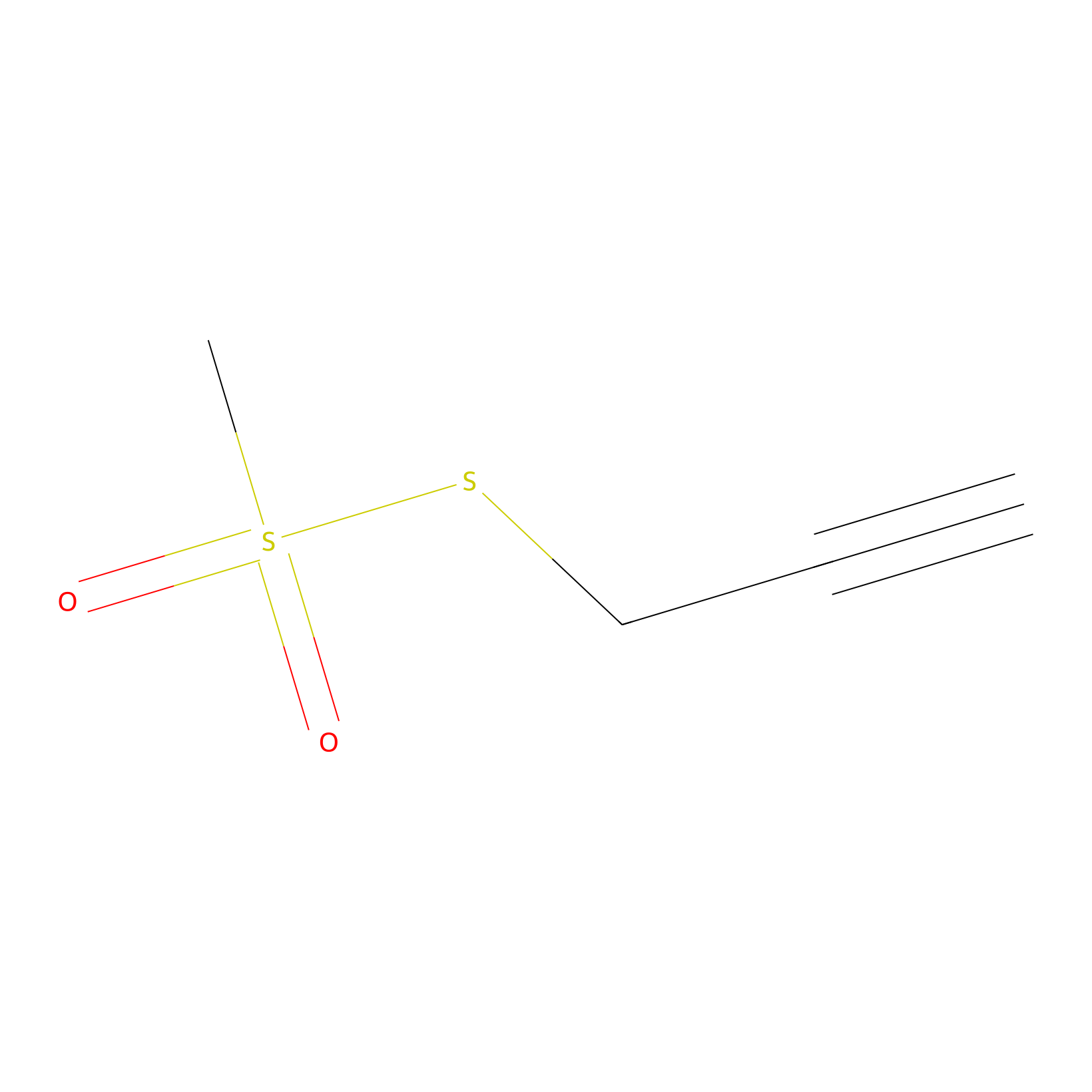 |
N.A. | LDD0008 | [34] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [34] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [34] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [39] | |
|
Acrolein Probe Info |
 |
C1101(0.00); H1164(0.00); H1158(0.00); H1098(0.00) | LDD0217 | [40] | |
|
Cinnamaldehyde Probe Info |
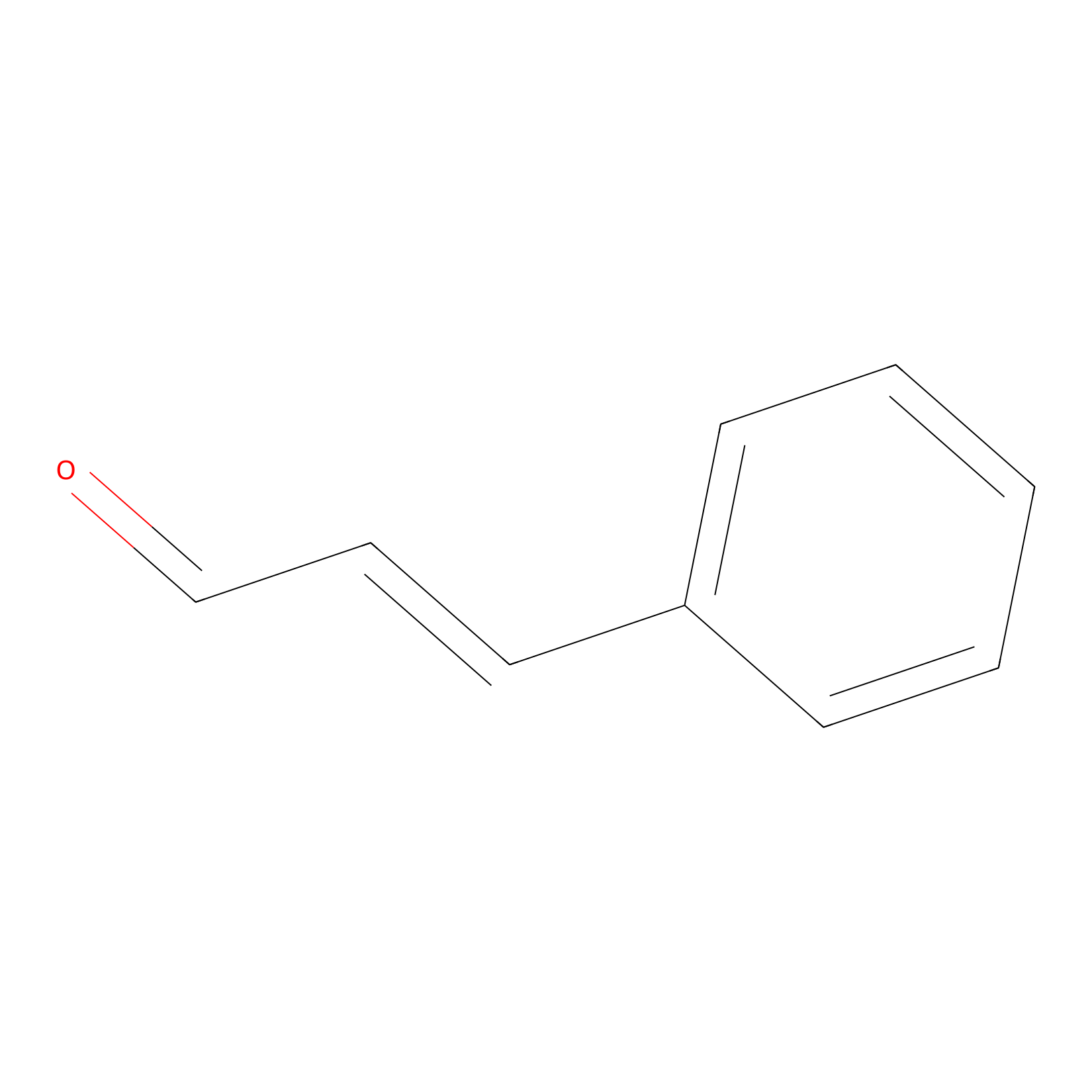 |
N.A. | LDD0220 | [40] | |
|
Crotonaldehyde Probe Info |
 |
C1101(0.00); H1098(0.00); H1071(0.00) | LDD0219 | [40] | |
|
Methacrolein Probe Info |
 |
C1101(0.00); H1098(0.00) | LDD0218 | [40] | |
|
AOyne Probe Info |
 |
12.30 | LDD0443 | [41] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C1101(0.93) | LDD2142 | [47] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C1101(1.24) | LDD2112 | [47] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C1101(0.82) | LDD2095 | [47] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C1101(1.24) | LDD2130 | [47] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C1101(0.99) | LDD2117 | [47] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C1101(1.11) | LDD2152 | [47] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C1101(1.10) | LDD2103 | [47] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C1101(0.96) | LDD2131 | [47] |
| LDCM0025 | 4SU-RNA | DM93 | C1101(2.06) | LDD0170 | [25] |
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C1101(5.14) | LDD0171 | [25] |
| LDCM0214 | AC1 | HEK-293T | C1101(1.29) | LDD0812 | [28] |
| LDCM0215 | AC10 | HEK-293T | C1101(1.18) | LDD0813 | [28] |
| LDCM0226 | AC11 | HEK-293T | C1101(1.02) | LDD0824 | [28] |
| LDCM0230 | AC113 | HEK-293T | C1101(1.04) | LDD0828 | [28] |
| LDCM0231 | AC114 | HEK-293T | C1101(1.16) | LDD0829 | [28] |
| LDCM0232 | AC115 | HEK-293T | C1101(1.18) | LDD0830 | [28] |
| LDCM0233 | AC116 | HEK-293T | C1101(1.00) | LDD0831 | [28] |
| LDCM0234 | AC117 | HEK-293T | C1101(1.02) | LDD0832 | [28] |
| LDCM0235 | AC118 | HEK-293T | C1101(0.94) | LDD0833 | [28] |
| LDCM0236 | AC119 | HEK-293T | C1101(1.18) | LDD0834 | [28] |
| LDCM0237 | AC12 | HEK-293T | C1101(1.12) | LDD0835 | [28] |
| LDCM0238 | AC120 | HEK-293T | C1101(1.17) | LDD0836 | [28] |
| LDCM0239 | AC121 | HEK-293T | C1101(0.95) | LDD0837 | [28] |
| LDCM0240 | AC122 | HEK-293T | C1101(1.12) | LDD0838 | [28] |
| LDCM0241 | AC123 | HEK-293T | C1101(1.11) | LDD0839 | [28] |
| LDCM0242 | AC124 | HEK-293T | C1101(1.07) | LDD0840 | [28] |
| LDCM0243 | AC125 | HEK-293T | C1101(1.19) | LDD0841 | [28] |
| LDCM0244 | AC126 | HEK-293T | C1101(1.26) | LDD0842 | [28] |
| LDCM0245 | AC127 | HEK-293T | C1101(1.44) | LDD0843 | [28] |
| LDCM0246 | AC128 | HEK-293T | C1101(0.83) | LDD0844 | [28] |
| LDCM0247 | AC129 | HEK-293T | C1101(0.85) | LDD0845 | [28] |
| LDCM0249 | AC130 | HEK-293T | C1101(0.88) | LDD0847 | [28] |
| LDCM0250 | AC131 | HEK-293T | C1101(0.87) | LDD0848 | [28] |
| LDCM0251 | AC132 | HEK-293T | C1101(0.61) | LDD0849 | [28] |
| LDCM0252 | AC133 | HEK-293T | C1101(0.98) | LDD0850 | [28] |
| LDCM0253 | AC134 | HEK-293T | C1101(0.92) | LDD0851 | [28] |
| LDCM0254 | AC135 | HEK-293T | C1101(0.88) | LDD0852 | [28] |
| LDCM0255 | AC136 | HEK-293T | C1101(0.80) | LDD0853 | [28] |
| LDCM0256 | AC137 | HEK-293T | C1101(1.12) | LDD0854 | [28] |
| LDCM0257 | AC138 | HEK-293T | C1101(1.18) | LDD0855 | [28] |
| LDCM0258 | AC139 | HEK-293T | C1101(0.91) | LDD0856 | [28] |
| LDCM0259 | AC14 | HEK-293T | C1101(0.96) | LDD0857 | [28] |
| LDCM0260 | AC140 | HEK-293T | C1101(0.91) | LDD0858 | [28] |
| LDCM0261 | AC141 | HEK-293T | C1101(1.00) | LDD0859 | [28] |
| LDCM0262 | AC142 | HEK-293T | C1101(0.92) | LDD0860 | [28] |
| LDCM0263 | AC143 | HCT 116 | C1101(0.97) | LDD0580 | [28] |
| LDCM0264 | AC144 | HCT 116 | C1101(1.05) | LDD0581 | [28] |
| LDCM0265 | AC145 | HCT 116 | C1101(1.10) | LDD0582 | [28] |
| LDCM0266 | AC146 | HCT 116 | C1101(0.89) | LDD0583 | [28] |
| LDCM0267 | AC147 | HCT 116 | C1101(1.23) | LDD0584 | [28] |
| LDCM0268 | AC148 | HCT 116 | C1101(0.89) | LDD0585 | [28] |
| LDCM0269 | AC149 | HCT 116 | C1101(0.97) | LDD0586 | [28] |
| LDCM0270 | AC15 | HEK-293T | C1101(0.95) | LDD0868 | [28] |
| LDCM0271 | AC150 | HCT 116 | C1101(0.95) | LDD0588 | [28] |
| LDCM0272 | AC151 | HCT 116 | C1101(0.92) | LDD0589 | [28] |
| LDCM0273 | AC152 | HCT 116 | C1101(0.92) | LDD0590 | [28] |
| LDCM0274 | AC153 | HCT 116 | C1101(0.69) | LDD0591 | [28] |
| LDCM0621 | AC154 | HCT 116 | C1101(0.98) | LDD2158 | [28] |
| LDCM0622 | AC155 | HCT 116 | C1101(0.75) | LDD2159 | [28] |
| LDCM0623 | AC156 | HCT 116 | C1101(0.98) | LDD2160 | [28] |
| LDCM0624 | AC157 | HCT 116 | C1101(0.92) | LDD2161 | [28] |
| LDCM0276 | AC17 | HCT 116 | C1101(0.70) | LDD0593 | [28] |
| LDCM0277 | AC18 | HCT 116 | C1101(0.61) | LDD0594 | [28] |
| LDCM0278 | AC19 | HCT 116 | C1101(0.58) | LDD0595 | [28] |
| LDCM0279 | AC2 | HEK-293T | C1101(0.90) | LDD0877 | [28] |
| LDCM0280 | AC20 | HCT 116 | C1101(0.85) | LDD0597 | [28] |
| LDCM0281 | AC21 | HCT 116 | C1101(0.73) | LDD0598 | [28] |
| LDCM0282 | AC22 | HCT 116 | C1101(0.59) | LDD0599 | [28] |
| LDCM0283 | AC23 | HCT 116 | C1101(1.05) | LDD0600 | [28] |
| LDCM0284 | AC24 | HCT 116 | C1101(0.63) | LDD0601 | [28] |
| LDCM0285 | AC25 | HEK-293T | C1101(1.03); C912(1.07); C559(1.02) | LDD1524 | [48] |
| LDCM0286 | AC26 | HEK-293T | C1101(1.07); C464(0.96) | LDD1525 | [48] |
| LDCM0287 | AC27 | HEK-293T | C1101(1.02) | LDD1526 | [48] |
| LDCM0288 | AC28 | HEK-293T | C1101(1.07) | LDD1527 | [48] |
| LDCM0289 | AC29 | HEK-293T | C1101(1.00); C464(0.94) | LDD1528 | [48] |
| LDCM0290 | AC3 | HEK-293T | C1101(1.03) | LDD0888 | [28] |
| LDCM0291 | AC30 | HEK-293T | C1101(1.03) | LDD1530 | [48] |
| LDCM0292 | AC31 | HEK-293T | C1101(1.04); C464(1.07) | LDD1531 | [48] |
| LDCM0293 | AC32 | HEK-293T | C1101(0.93) | LDD1532 | [48] |
| LDCM0294 | AC33 | HEK-293T | C1101(1.07); C912(0.98); C559(0.93) | LDD1533 | [48] |
| LDCM0295 | AC34 | HEK-293T | C1101(1.05); C464(1.05) | LDD1534 | [48] |
| LDCM0296 | AC35 | HEK-293T | C1101(0.86) | LDD0894 | [28] |
| LDCM0297 | AC36 | HEK-293T | C1101(0.66) | LDD0895 | [28] |
| LDCM0298 | AC37 | HEK-293T | C1101(0.82) | LDD0896 | [28] |
| LDCM0299 | AC38 | HEK-293T | C1101(0.72) | LDD0897 | [28] |
| LDCM0300 | AC39 | HEK-293T | C1101(0.65) | LDD0898 | [28] |
| LDCM0301 | AC4 | HEK-293T | C1101(1.00) | LDD0899 | [28] |
| LDCM0302 | AC40 | HEK-293T | C1101(0.70) | LDD0900 | [28] |
| LDCM0303 | AC41 | HEK-293T | C1101(1.02) | LDD0901 | [28] |
| LDCM0304 | AC42 | HEK-293T | C1101(0.89) | LDD0902 | [28] |
| LDCM0305 | AC43 | HEK-293T | C1101(1.01) | LDD0903 | [28] |
| LDCM0306 | AC44 | HEK-293T | C1101(0.58) | LDD0904 | [28] |
| LDCM0307 | AC45 | HEK-293T | C1101(0.77) | LDD0905 | [28] |
| LDCM0308 | AC46 | HEK-293T | C1101(0.97) | LDD0906 | [28] |
| LDCM0309 | AC47 | HEK-293T | C1101(1.01) | LDD0907 | [28] |
| LDCM0310 | AC48 | HEK-293T | C1101(0.95) | LDD0908 | [28] |
| LDCM0311 | AC49 | HEK-293T | C1101(1.00) | LDD0909 | [28] |
| LDCM0312 | AC5 | HEK-293T | C1101(0.87) | LDD0910 | [28] |
| LDCM0313 | AC50 | HEK-293T | C1101(0.92) | LDD0911 | [28] |
| LDCM0314 | AC51 | HEK-293T | C1101(0.84) | LDD0912 | [28] |
| LDCM0315 | AC52 | HEK-293T | C1101(0.78) | LDD0913 | [28] |
| LDCM0316 | AC53 | HEK-293T | C1101(0.98) | LDD0914 | [28] |
| LDCM0317 | AC54 | HEK-293T | C1101(0.80) | LDD0915 | [28] |
| LDCM0318 | AC55 | HEK-293T | C1101(0.87) | LDD0916 | [28] |
| LDCM0319 | AC56 | HEK-293T | C1101(0.95) | LDD0917 | [28] |
| LDCM0320 | AC57 | HEK-293T | C1101(0.96); C912(1.05); C559(0.81) | LDD1559 | [48] |
| LDCM0321 | AC58 | HEK-293T | C1101(0.93); C464(1.09) | LDD1560 | [48] |
| LDCM0322 | AC59 | HEK-293T | C1101(0.90) | LDD1561 | [48] |
| LDCM0323 | AC6 | HEK-293T | C1101(0.89) | LDD0921 | [28] |
| LDCM0324 | AC60 | HEK-293T | C1101(0.98) | LDD1563 | [48] |
| LDCM0325 | AC61 | HEK-293T | C1101(0.94); C464(0.93) | LDD1564 | [48] |
| LDCM0326 | AC62 | HEK-293T | C1101(0.94) | LDD1565 | [48] |
| LDCM0327 | AC63 | HEK-293T | C1101(0.94); C464(0.91) | LDD1566 | [48] |
| LDCM0328 | AC64 | HEK-293T | C1101(0.90) | LDD1567 | [48] |
| LDCM0332 | AC68 | HCT 116 | C1101(0.88) | LDD0649 | [28] |
| LDCM0333 | AC69 | HCT 116 | C1101(0.66) | LDD0650 | [28] |
| LDCM0334 | AC7 | HEK-293T | C1101(0.99) | LDD0932 | [28] |
| LDCM0335 | AC70 | HCT 116 | C1101(0.64) | LDD0652 | [28] |
| LDCM0336 | AC71 | HCT 116 | C1101(0.82) | LDD0653 | [28] |
| LDCM0337 | AC72 | HCT 116 | C1101(0.99) | LDD0654 | [28] |
| LDCM0338 | AC73 | HCT 116 | C1101(0.93) | LDD0655 | [28] |
| LDCM0339 | AC74 | HCT 116 | C1101(0.84) | LDD0656 | [28] |
| LDCM0340 | AC75 | HCT 116 | C1101(0.60) | LDD0657 | [28] |
| LDCM0341 | AC76 | HCT 116 | C1101(0.76) | LDD0658 | [28] |
| LDCM0342 | AC77 | HCT 116 | C1101(1.02) | LDD0659 | [28] |
| LDCM0343 | AC78 | HCT 116 | C1101(0.90) | LDD0660 | [28] |
| LDCM0344 | AC79 | HCT 116 | C1101(0.63) | LDD0661 | [28] |
| LDCM0345 | AC8 | HEK-293T | C1101(0.81) | LDD0943 | [28] |
| LDCM0346 | AC80 | HCT 116 | C1101(0.72) | LDD0663 | [28] |
| LDCM0347 | AC81 | HCT 116 | C1101(0.81) | LDD0664 | [28] |
| LDCM0348 | AC82 | HCT 116 | C1101(0.82) | LDD0665 | [28] |
| LDCM0349 | AC83 | HEK-293T | C1101(1.07) | LDD0947 | [28] |
| LDCM0350 | AC84 | HEK-293T | C1101(0.79) | LDD0948 | [28] |
| LDCM0351 | AC85 | HEK-293T | C1101(0.81) | LDD0949 | [28] |
| LDCM0352 | AC86 | HEK-293T | C1101(0.95) | LDD0950 | [28] |
| LDCM0353 | AC87 | HEK-293T | C1101(0.83) | LDD0951 | [28] |
| LDCM0354 | AC88 | HEK-293T | C1101(0.76) | LDD0952 | [28] |
| LDCM0355 | AC89 | HEK-293T | C1101(0.73) | LDD0953 | [28] |
| LDCM0357 | AC90 | HEK-293T | C1101(0.74) | LDD0955 | [28] |
| LDCM0358 | AC91 | HEK-293T | C1101(0.93) | LDD0956 | [28] |
| LDCM0359 | AC92 | HEK-293T | C1101(0.86) | LDD0957 | [28] |
| LDCM0360 | AC93 | HEK-293T | C1101(0.88) | LDD0958 | [28] |
| LDCM0361 | AC94 | HEK-293T | C1101(0.81) | LDD0959 | [28] |
| LDCM0362 | AC95 | HEK-293T | C1101(0.81) | LDD0960 | [28] |
| LDCM0363 | AC96 | HEK-293T | C1101(0.80) | LDD0961 | [28] |
| LDCM0364 | AC97 | HEK-293T | C1101(1.17) | LDD0962 | [28] |
| LDCM0545 | Acetamide | MDA-MB-231 | C1101(0.66) | LDD2138 | [47] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C1101(0.77) | LDD2113 | [47] |
| LDCM0248 | AKOS034007472 | HEK-293T | C1101(0.94) | LDD0846 | [28] |
| LDCM0356 | AKOS034007680 | HEK-293T | C1101(0.92) | LDD0954 | [28] |
| LDCM0275 | AKOS034007705 | HEK-293T | C1101(0.79) | LDD0873 | [28] |
| LDCM0156 | Aniline | NCI-H1299 | 11.92 | LDD0403 | [2] |
| LDCM0020 | ARS-1620 | HCC44 | C1101(1.77) | LDD0078 | [28] |
| LDCM0083 | Avasimibe | A-549 | 2.59 | LDD0143 | [46] |
| LDCM0199 | BPK-21 | T cell | C1101(9.76) | LDD0517 | [49] |
| LDCM0200 | BPK-25 | T cell | C1101(9.13) | LDD0516 | [49] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C1101(0.55) | LDD2091 | [47] |
| LDCM0088 | C45 | HEK-293T | 6.04 | LDD0203 | [20] |
| LDCM0630 | CCW28-3 | 231MFP | C1101(1.59) | LDD2214 | [50] |
| LDCM0108 | Chloroacetamide | HeLa | C1101(0.00); H1164(0.00); H1158(0.00); H1071(0.00) | LDD0222 | [40] |
| LDCM0367 | CL1 | HEK-293T | C1101(1.31); C912(0.90) | LDD1571 | [48] |
| LDCM0368 | CL10 | HEK-293T | C1101(7.73) | LDD1572 | [48] |
| LDCM0369 | CL100 | HEK-293T | C1101(1.01) | LDD0967 | [28] |
| LDCM0370 | CL101 | HEK-293T | C1101(0.94) | LDD0968 | [28] |
| LDCM0371 | CL102 | HEK-293T | C1101(0.92) | LDD0969 | [28] |
| LDCM0372 | CL103 | HEK-293T | C1101(0.82) | LDD0970 | [28] |
| LDCM0373 | CL104 | HEK-293T | C1101(0.92) | LDD0971 | [28] |
| LDCM0374 | CL105 | HCT 116 | C1101(0.73) | LDD0691 | [28] |
| LDCM0375 | CL106 | HCT 116 | C1101(0.64) | LDD0692 | [28] |
| LDCM0376 | CL107 | HCT 116 | C1101(0.66) | LDD0693 | [28] |
| LDCM0377 | CL108 | HCT 116 | C1101(0.61) | LDD0694 | [28] |
| LDCM0378 | CL109 | HCT 116 | C1101(0.93) | LDD0695 | [28] |
| LDCM0379 | CL11 | HEK-293T | C1101(1.37); C464(0.92) | LDD1583 | [48] |
| LDCM0380 | CL110 | HCT 116 | C1101(1.10) | LDD0697 | [28] |
| LDCM0381 | CL111 | HCT 116 | C1101(1.25) | LDD0698 | [28] |
| LDCM0382 | CL112 | HEK-293T | C1101(1.08) | LDD1586 | [48] |
| LDCM0383 | CL113 | HEK-293T | C1101(0.86); C912(0.97) | LDD1587 | [48] |
| LDCM0384 | CL114 | HEK-293T | C1101(1.93) | LDD1588 | [48] |
| LDCM0385 | CL115 | HEK-293T | C1101(1.00); C464(1.12) | LDD1589 | [48] |
| LDCM0386 | CL116 | HEK-293T | C1101(0.95) | LDD1590 | [48] |
| LDCM0387 | CL117 | HEK-293T | C1101(0.80) | LDD0985 | [28] |
| LDCM0388 | CL118 | HEK-293T | C1101(0.84) | LDD0986 | [28] |
| LDCM0389 | CL119 | HEK-293T | C1101(0.61) | LDD0987 | [28] |
| LDCM0390 | CL12 | HEK-293T | C1101(1.17) | LDD1594 | [48] |
| LDCM0391 | CL120 | HEK-293T | C1101(0.83) | LDD0989 | [28] |
| LDCM0392 | CL121 | HEK-293T | C1101(0.96) | LDD0990 | [28] |
| LDCM0393 | CL122 | HEK-293T | C1101(0.87) | LDD0991 | [28] |
| LDCM0394 | CL123 | HEK-293T | C1101(0.96) | LDD0992 | [28] |
| LDCM0395 | CL124 | HEK-293T | C1101(0.88) | LDD0993 | [28] |
| LDCM0396 | CL125 | HEK-293T | C1101(0.85); C912(0.95) | LDD1600 | [48] |
| LDCM0397 | CL126 | HEK-293T | C1101(1.03) | LDD1601 | [48] |
| LDCM0398 | CL127 | HEK-293T | C1101(0.81); C464(0.95) | LDD1602 | [48] |
| LDCM0399 | CL128 | HEK-293T | C1101(0.79) | LDD1603 | [48] |
| LDCM0400 | CL13 | HEK-293T | C1101(0.94); C912(0.89) | LDD1604 | [48] |
| LDCM0401 | CL14 | HEK-293T | C1101(0.99) | LDD1605 | [48] |
| LDCM0402 | CL15 | HEK-293T | C1101(2.29); C464(1.16) | LDD1606 | [48] |
| LDCM0403 | CL16 | HEK-293T | C1101(0.93) | LDD1607 | [48] |
| LDCM0404 | CL17 | HCT 116 | C1101(1.97) | LDD0721 | [28] |
| LDCM0405 | CL18 | HCT 116 | C1101(0.81) | LDD0722 | [28] |
| LDCM0406 | CL19 | HCT 116 | C1101(1.10) | LDD0723 | [28] |
| LDCM0407 | CL2 | HEK-293T | C1101(1.01) | LDD1611 | [48] |
| LDCM0408 | CL20 | HCT 116 | C1101(1.12) | LDD0725 | [28] |
| LDCM0409 | CL21 | HCT 116 | C1101(1.52) | LDD0726 | [28] |
| LDCM0410 | CL22 | HCT 116 | C1101(1.19) | LDD0727 | [28] |
| LDCM0411 | CL23 | HCT 116 | C1101(1.38) | LDD0728 | [28] |
| LDCM0412 | CL24 | HCT 116 | C1101(0.80) | LDD0729 | [28] |
| LDCM0413 | CL25 | HCT 116 | C1101(1.59) | LDD0730 | [28] |
| LDCM0414 | CL26 | HCT 116 | C1101(0.97) | LDD0731 | [28] |
| LDCM0415 | CL27 | HCT 116 | C1101(0.84) | LDD0732 | [28] |
| LDCM0416 | CL28 | HCT 116 | C1101(0.93) | LDD0733 | [28] |
| LDCM0417 | CL29 | HCT 116 | C1101(0.98) | LDD0734 | [28] |
| LDCM0418 | CL3 | HEK-293T | C1101(1.17); C464(0.79) | LDD1622 | [48] |
| LDCM0419 | CL30 | HCT 116 | C1101(1.11) | LDD0736 | [28] |
| LDCM0420 | CL31 | HCT 116 | C1101(0.90) | LDD0737 | [28] |
| LDCM0421 | CL32 | HEK-293T | C1101(0.84) | LDD1019 | [28] |
| LDCM0422 | CL33 | HEK-293T | C1101(0.93) | LDD1020 | [28] |
| LDCM0423 | CL34 | HEK-293T | C1101(0.86) | LDD1021 | [28] |
| LDCM0424 | CL35 | HEK-293T | C1101(0.86) | LDD1022 | [28] |
| LDCM0425 | CL36 | HEK-293T | C1101(0.90) | LDD1023 | [28] |
| LDCM0426 | CL37 | HEK-293T | C1101(0.76) | LDD1024 | [28] |
| LDCM0428 | CL39 | HEK-293T | C1101(0.77) | LDD1026 | [28] |
| LDCM0429 | CL4 | HEK-293T | C1101(1.18) | LDD1633 | [48] |
| LDCM0430 | CL40 | HEK-293T | C1101(0.67) | LDD1028 | [28] |
| LDCM0431 | CL41 | HEK-293T | C1101(0.88) | LDD1029 | [28] |
| LDCM0432 | CL42 | HEK-293T | C1101(0.85) | LDD1030 | [28] |
| LDCM0433 | CL43 | HEK-293T | C1101(0.75) | LDD1031 | [28] |
| LDCM0434 | CL44 | HEK-293T | C1101(0.67) | LDD1032 | [28] |
| LDCM0435 | CL45 | HEK-293T | C1101(0.75) | LDD1033 | [28] |
| LDCM0436 | CL46 | HEK-293T | C1101(1.04) | LDD1034 | [28] |
| LDCM0437 | CL47 | HEK-293T | C1101(0.97) | LDD1035 | [28] |
| LDCM0438 | CL48 | HEK-293T | C1101(0.87) | LDD1036 | [28] |
| LDCM0439 | CL49 | HEK-293T | C1101(1.00) | LDD1037 | [28] |
| LDCM0440 | CL5 | HEK-293T | C1101(0.92); C912(1.25); C559(1.09) | LDD1644 | [48] |
| LDCM0441 | CL50 | HEK-293T | C1101(1.09) | LDD1039 | [28] |
| LDCM0442 | CL51 | HEK-293T | C1101(0.96) | LDD1040 | [28] |
| LDCM0443 | CL52 | HEK-293T | C1101(1.06) | LDD1041 | [28] |
| LDCM0444 | CL53 | HEK-293T | C1101(1.04) | LDD1042 | [28] |
| LDCM0445 | CL54 | HEK-293T | C1101(1.06) | LDD1043 | [28] |
| LDCM0446 | CL55 | HEK-293T | C1101(1.03) | LDD1044 | [28] |
| LDCM0447 | CL56 | HEK-293T | C1101(0.92) | LDD1045 | [28] |
| LDCM0448 | CL57 | HEK-293T | C1101(1.00) | LDD1046 | [28] |
| LDCM0449 | CL58 | HEK-293T | C1101(0.91) | LDD1047 | [28] |
| LDCM0450 | CL59 | HEK-293T | C1101(0.98) | LDD1048 | [28] |
| LDCM0451 | CL6 | HEK-293T | C1101(1.57); C464(1.01) | LDD1654 | [48] |
| LDCM0452 | CL60 | HEK-293T | C1101(1.17) | LDD1050 | [28] |
| LDCM0453 | CL61 | HEK-293T | C1101(0.87); C912(1.09) | LDD1656 | [48] |
| LDCM0454 | CL62 | HEK-293T | C1101(0.99) | LDD1657 | [48] |
| LDCM0455 | CL63 | HEK-293T | C1101(0.91); C464(0.93) | LDD1658 | [48] |
| LDCM0456 | CL64 | HEK-293T | C1101(1.21) | LDD1659 | [48] |
| LDCM0457 | CL65 | HEK-293T | C1101(1.06); C912(1.14); C559(1.03) | LDD1660 | [48] |
| LDCM0458 | CL66 | HEK-293T | C1101(1.07); C464(1.07) | LDD1661 | [48] |
| LDCM0459 | CL67 | HEK-293T | C1101(1.02) | LDD1662 | [48] |
| LDCM0460 | CL68 | HEK-293T | C1101(1.13) | LDD1663 | [48] |
| LDCM0461 | CL69 | HEK-293T | C1101(1.23); C464(0.94) | LDD1664 | [48] |
| LDCM0462 | CL7 | HEK-293T | C1101(1.05) | LDD1665 | [48] |
| LDCM0463 | CL70 | HEK-293T | C1101(1.16) | LDD1666 | [48] |
| LDCM0464 | CL71 | HEK-293T | C1101(1.13); C464(1.07) | LDD1667 | [48] |
| LDCM0465 | CL72 | HEK-293T | C1101(1.04) | LDD1668 | [48] |
| LDCM0466 | CL73 | HEK-293T | C1101(1.24); C912(1.05) | LDD1669 | [48] |
| LDCM0467 | CL74 | HEK-293T | C1101(0.97) | LDD1670 | [48] |
| LDCM0469 | CL76 | HEK-293T | C1101(0.93) | LDD1672 | [48] |
| LDCM0470 | CL77 | HEK-293T | C1101(1.49); C912(1.34); C559(0.82) | LDD1673 | [48] |
| LDCM0471 | CL78 | HEK-293T | C1101(0.94); C464(0.97) | LDD1674 | [48] |
| LDCM0472 | CL79 | HEK-293T | C1101(1.00) | LDD1675 | [48] |
| LDCM0473 | CL8 | HEK-293T | C1101(5.55) | LDD1676 | [48] |
| LDCM0474 | CL80 | HEK-293T | C1101(0.95) | LDD1677 | [48] |
| LDCM0475 | CL81 | HEK-293T | C1101(0.98); C464(0.94) | LDD1678 | [48] |
| LDCM0476 | CL82 | HEK-293T | C1101(1.12) | LDD1679 | [48] |
| LDCM0477 | CL83 | HEK-293T | C1101(1.36); C464(1.01) | LDD1680 | [48] |
| LDCM0478 | CL84 | HEK-293T | C1101(1.24) | LDD1681 | [48] |
| LDCM0479 | CL85 | HEK-293T | C1101(0.82); C912(0.94) | LDD1682 | [48] |
| LDCM0480 | CL86 | HEK-293T | C1101(0.79) | LDD1683 | [48] |
| LDCM0481 | CL87 | HEK-293T | C1101(0.84); C464(1.16) | LDD1684 | [48] |
| LDCM0482 | CL88 | HEK-293T | C1101(0.78) | LDD1685 | [48] |
| LDCM0483 | CL89 | HEK-293T | C1101(0.87); C912(1.09); C559(0.95) | LDD1686 | [48] |
| LDCM0484 | CL9 | HEK-293T | C1101(1.19); C464(0.99) | LDD1687 | [48] |
| LDCM0485 | CL90 | HEK-293T | C1101(5.59); C464(0.91) | LDD1688 | [48] |
| LDCM0486 | CL91 | HEK-293T | C1101(1.20) | LDD1084 | [28] |
| LDCM0487 | CL92 | HEK-293T | C1101(1.03) | LDD1085 | [28] |
| LDCM0488 | CL93 | HEK-293T | C1101(0.88) | LDD1086 | [28] |
| LDCM0489 | CL94 | HEK-293T | C1101(1.13) | LDD1087 | [28] |
| LDCM0490 | CL95 | HEK-293T | C1101(0.91) | LDD1088 | [28] |
| LDCM0491 | CL96 | HEK-293T | C1101(0.95) | LDD1089 | [28] |
| LDCM0492 | CL97 | HEK-293T | C1101(0.87) | LDD1090 | [28] |
| LDCM0493 | CL98 | HEK-293T | C1101(1.11) | LDD1091 | [28] |
| LDCM0494 | CL99 | HEK-293T | C1101(0.99) | LDD1092 | [28] |
| LDCM0189 | Compound 16 | HEK-293T | 58.82 | LDD0492 | [44] |
| LDCM0185 | Compound 17 | HEK-293T | 9.76 | LDD0512 | [44] |
| LDCM0184 | Compound 20 | HEK-293T | 6.59 | LDD0503 | [44] |
| LDCM0191 | Compound 21 | HEK-293T | 4.88 | LDD0493 | [44] |
| LDCM0186 | Compound 31 | HEK-293T | 5.33 | LDD0506 | [44] |
| LDCM0187 | Compound 32 | HEK-293T | 132.33 | LDD0507 | [44] |
| LDCM0190 | Compound 34 | HEK-293T | 37.04 | LDD0497 | [44] |
| LDCM0192 | Compound 35 | HEK-293T | 7.55 | LDD0491 | [44] |
| LDCM0193 | Compound 36 | HEK-293T | 5.49 | LDD0511 | [44] |
| LDCM0194 | Compound 37 | HEK-293T | 42.55 | LDD0498 | [44] |
| LDCM0196 | Compound 39 | HEK-293T | 4.69 | LDD0496 | [44] |
| LDCM0197 | Compound 40 | HEK-293T | 8.00 | LDD0495 | [44] |
| LDCM0027 | Dopamine | HEK-293T | 36.56 | LDD0179 | [24] |
| LDCM0495 | E2913 | HEK-293T | C1101(1.00); C464(0.84) | LDD1698 | [48] |
| LDCM0202 | EV-93 | T cell | C1101(15.97) | LDD0527 | [49] |
| LDCM0203 | EV-96 | T cell | C1101(4.00) | LDD0522 | [49] |
| LDCM0204 | EV-97 | T cell | C1101(14.00) | LDD0525 | [49] |
| LDCM0625 | F8 | Ramos | C1101(0.99) | LDD2187 | [51] |
| LDCM0572 | Fragment10 | MDA-MB-231 | C1101(11.86) | LDD1389 | [22] |
| LDCM0573 | Fragment11 | MDA-MB-231 | C1101(1.08) | LDD1467 | [22] |
| LDCM0574 | Fragment12 | Ramos | C1101(20.00) | LDD1394 | [22] |
| LDCM0575 | Fragment13 | MDA-MB-231 | C1101(3.93) | LDD1395 | [22] |
| LDCM0576 | Fragment14 | MDA-MB-231 | C1101(3.74) | LDD1397 | [22] |
| LDCM0579 | Fragment20 | MDA-MB-231 | C1101(5.63) | LDD1402 | [22] |
| LDCM0580 | Fragment21 | MDA-MB-231 | C1101(8.56) | LDD1404 | [22] |
| LDCM0581 | Fragment22 | MDA-MB-231 | C1101(1.08) | LDD1406 | [22] |
| LDCM0582 | Fragment23 | MDA-MB-231 | C1101(3.16) | LDD1408 | [22] |
| LDCM0578 | Fragment27 | MDA-MB-231 | C1101(0.91) | LDD1401 | [22] |
| LDCM0586 | Fragment28 | MDA-MB-231 | C1101(1.40) | LDD1415 | [22] |
| LDCM0587 | Fragment29 | MDA-MB-231 | C1101(0.95) | LDD1417 | [22] |
| LDCM0588 | Fragment30 | MDA-MB-231 | C1101(1.60) | LDD1419 | [22] |
| LDCM0589 | Fragment31 | MDA-MB-231 | C1101(20.00) | LDD1421 | [22] |
| LDCM0590 | Fragment32 | MDA-MB-231 | C1101(20.00) | LDD1423 | [22] |
| LDCM0468 | Fragment33 | MDA-MB-231 | C1101(1.21) | LDD1425 | [22] |
| LDCM0592 | Fragment34 | MDA-MB-231 | C1101(0.84) | LDD1427 | [22] |
| LDCM0593 | Fragment35 | MDA-MB-231 | C1101(1.38) | LDD1429 | [22] |
| LDCM0594 | Fragment36 | MDA-MB-231 | C1101(2.64) | LDD1431 | [22] |
| LDCM0596 | Fragment38 | MDA-MB-231 | C1101(7.50) | LDD1433 | [22] |
| LDCM0566 | Fragment4 | MDA-MB-231 | C1101(11.88) | LDD1378 | [22] |
| LDCM0599 | Fragment41 | MDA-MB-231 | C1101(1.35) | LDD1438 | [22] |
| LDCM0600 | Fragment42 | Ramos | C1101(0.98) | LDD1440 | [22] |
| LDCM0601 | Fragment43 | Ramos | C1101(10.94) | LDD1442 | [22] |
| LDCM0602 | Fragment44 | MDA-MB-231 | C1101(0.72) | LDD1443 | [22] |
| LDCM0603 | Fragment45 | MDA-MB-231 | C1101(20.00) | LDD1444 | [22] |
| LDCM0604 | Fragment46 | MDA-MB-231 | C1101(1.36) | LDD1445 | [22] |
| LDCM0606 | Fragment48 | MDA-MB-231 | C1101(1.76) | LDD1447 | [22] |
| LDCM0607 | Fragment49 | MDA-MB-231 | C1101(11.17) | LDD1448 | [22] |
| LDCM0608 | Fragment50 | MDA-MB-231 | C1101(20.00) | LDD1449 | [22] |
| LDCM0427 | Fragment51 | HEK-293T | C1101(0.86) | LDD1025 | [28] |
| LDCM0610 | Fragment52 | Ramos | C1101(3.89) | LDD2204 | [51] |
| LDCM0611 | Fragment53 | MDA-MB-231 | C1101(1.38) | LDD1454 | [22] |
| LDCM0612 | Fragment54 | MDA-MB-231 | C1101(1.06) | LDD1456 | [22] |
| LDCM0613 | Fragment55 | MDA-MB-231 | C1101(1.02) | LDD1457 | [22] |
| LDCM0614 | Fragment56 | MDA-MB-231 | C1101(0.93) | LDD1458 | [22] |
| LDCM0616 | Fragment61 | Jurkat | _(15.99) | LDD1489 | [22] |
| LDCM0617 | Fragment63-S | Jurkat | _(11.94) | LDD1490 | [22] |
| LDCM0569 | Fragment7 | Ramos | C1101(8.18) | LDD2186 | [51] |
| LDCM0570 | Fragment8 | MDA-MB-231 | C1101(1.54) | LDD1385 | [22] |
| LDCM0571 | Fragment9 | MDA-MB-231 | C1101(15.03) | LDD1387 | [22] |
| LDCM0107 | IAA | HeLa | H1164(0.00); H1158(0.00); C1101(0.00); H1098(0.00) | LDD0221 | [40] |
| LDCM0130 | JWB211 | DM93 | Y1076(0.19) | LDD0292 | [27] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [21] |
| LDCM0022 | KB02 | Ramos | 14.77 | LDD0431 | [52] |
| LDCM0023 | KB03 | Jurkat | C1101(4.76) | LDD0315 | [32] |
| LDCM0024 | KB05 | COLO792 | C464(0.86); C895(4.05) | LDD3310 | [53] |
| LDCM0169 | KB63 | Ramos | 10.70 | LDD0432 | [52] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C1101(1.24) | LDD2102 | [47] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C1101(0.82) | LDD2121 | [47] |
| LDCM0109 | NEM | HeLa | H1098(0.00); H1164(0.00); H1158(0.00); H1071(0.00) | LDD0223 | [40] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C1101(0.83) | LDD2089 | [47] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C1101(2.01) | LDD2090 | [47] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C1101(1.16) | LDD2092 | [47] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C1101(1.50) | LDD2093 | [47] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C1101(1.54) | LDD2094 | [47] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C1101(0.74) | LDD2096 | [47] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C1101(1.03) | LDD2097 | [47] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C1101(1.54) | LDD2098 | [47] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C1101(0.92) | LDD2099 | [47] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C1101(1.82) | LDD2100 | [47] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C1101(0.95) | LDD2101 | [47] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C1101(1.57) | LDD2104 | [47] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C1101(1.53) | LDD2105 | [47] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C1101(1.57) | LDD2106 | [47] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C1101(0.76) | LDD2107 | [47] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C1101(0.68) | LDD2108 | [47] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C1101(0.87) | LDD2109 | [47] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C1101(5.25) | LDD2110 | [47] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C1101(1.05) | LDD2111 | [47] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C1101(1.19) | LDD2114 | [47] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C1101(0.52) | LDD2115 | [47] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C1101(1.07) | LDD2116 | [47] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C1101(1.39) | LDD2118 | [47] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C1101(2.34) | LDD2119 | [47] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C1101(0.75) | LDD2120 | [47] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C1101(0.97) | LDD2122 | [47] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C1101(0.88) | LDD2123 | [47] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C1101(1.21) | LDD2124 | [47] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C1101(0.85) | LDD2125 | [47] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C1101(3.08) | LDD2126 | [47] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C1101(1.28) | LDD2127 | [47] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C1101(0.75) | LDD2128 | [47] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C1101(1.00) | LDD2129 | [47] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C1101(0.72) | LDD2133 | [47] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C1101(0.99) | LDD2135 | [47] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C1101(1.47) | LDD2136 | [47] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C1101(0.90) | LDD2137 | [47] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C1101(1.50) | LDD1700 | [47] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C1101(0.83) | LDD2140 | [47] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C1101(0.56) | LDD2141 | [47] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C1101(0.96) | LDD2143 | [47] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C1101(2.06) | LDD2144 | [47] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C1101(9.33) | LDD2145 | [47] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C1101(0.85) | LDD2146 | [47] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C1101(2.44) | LDD2147 | [47] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C1101(0.58) | LDD2148 | [47] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C1101(1.15) | LDD2149 | [47] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C1101(0.66) | LDD2150 | [47] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C1101(3.69) | LDD2153 | [47] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C1101(0.30); C1101(0.07) | LDD2206 | [54] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C1101(1.08); C1101(0.49) | LDD2207 | [54] |
| LDCM0014 | Panhematin | hPBMC | 5.72 | LDD0064 | [26] |
| LDCM0099 | Phenelzine | HEK-293T | 11.00 | LDD0390 | [19] |
| LDCM0131 | RA190 | MM1.R | C1101(6.15) | LDD0304 | [55] |
| LDCM0084 | Ro 48-8071 | A-549 | 2.84 | LDD0145 | [46] |
| LDCM0021 | THZ1 | HeLa S3 | C1110(1.18) | LDD0460 | [21] |
| LDCM0110 | W12 | Hep-G2 | C1101(10.39) | LDD0237 | [14] |
| LDCM0111 | W14 | Hep-G2 | C1101(12.24) | LDD0238 | [14] |
| LDCM0112 | W16 | Hep-G2 | C1101(0.60) | LDD0239 | [14] |
| LDCM0113 | W17 | Hep-G2 | H1098(8.42); C1101(8.50) | LDD0240 | [14] |
| LDCM0154 | YY4 | T cell | 2.92 | LDD0400 | [23] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Atlastin-1 (ATL1) | GB1/RHD3 GTPase family | Q8WXF7 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Sorting nexin-1 (SNX1) | Sorting nexin family | Q13596 | |||
| Sorting nexin-15 (SNX15) | Sorting nexin family | Q9NRS6 | |||
| Ceramide transfer protein (CERT1) | . | Q9Y5P4 | |||
Other
References