Details of the Target
General Information of Target
| Target ID | LDTP01328 | |||||
|---|---|---|---|---|---|---|
| Target Name | CAAX prenyl protease 1 homolog (ZMPSTE24) | |||||
| Gene Name | ZMPSTE24 | |||||
| Gene ID | 10269 | |||||
| Synonyms |
FACE1; STE24; CAAX prenyl protease 1 homolog; EC 3.4.24.84; Farnesylated proteins-converting enzyme 1; FACE-1; Prenyl protein-specific endoprotease 1; Zinc metalloproteinase Ste24 homolog |
|||||
| 3D Structure | ||||||
| Sequence |
MGMWASLDALWEMPAEKRIFGAVLLFSWTVYLWETFLAQRQRRIYKTTTHVPPELGQIMD
SETFEKSRLYQLDKSTFSFWSGLYSETEGTLILLFGGIPYLWRLSGRFCGYAGFGPEYEI TQSLVFLLLATLFSALTGLPWSLYNTFVIEEKHGFNQQTLGFFMKDAIKKFVVTQCILLP VSSLLLYIIKIGGDYFFIYAWLFTLVVSLVLVTIYADYIAPLFDKFTPLPEGKLKEEIEV MAKSIDFPLTKVYVVEGSKRSSHSNAYFYGFFKNKRIVLFDTLLEEYSVLNKDIQEDSGM EPRNEEEGNSEEIKAKVKNKKQGCKNEEVLAVLGHELGHWKLGHTVKNIIISQMNSFLCF FLFAVLIGRKELFAAFGFYDSQPTLIGLLIIFQFIFSPYNEVLSFCLTVLSRRFEFQADA FAKKLGKAKDLYSALIKLNKDNLGFPVSDWLFSMWHYSHPPLLERLQALKTMKQH |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Peptidase M48A family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Transmembrane metalloprotease whose catalytic activity is critical for processing lamin A/LMNA on the inner nuclear membrane and clearing clogged translocons on the endoplasmic reticulum. Proteolytically removes the C-terminal three residues of farnesylated proteins. Plays also an antiviral role independently of its protease activity by restricting enveloped RNA and DNA viruses, including influenza A, Zika, Ebola, Sindbis, vesicular stomatitis, cowpox, and vaccinia. Mechanistically, controls IFITM antiviral pathway to hinder viruses from breaching the endosomal barrier by modulating membrane fluidity.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
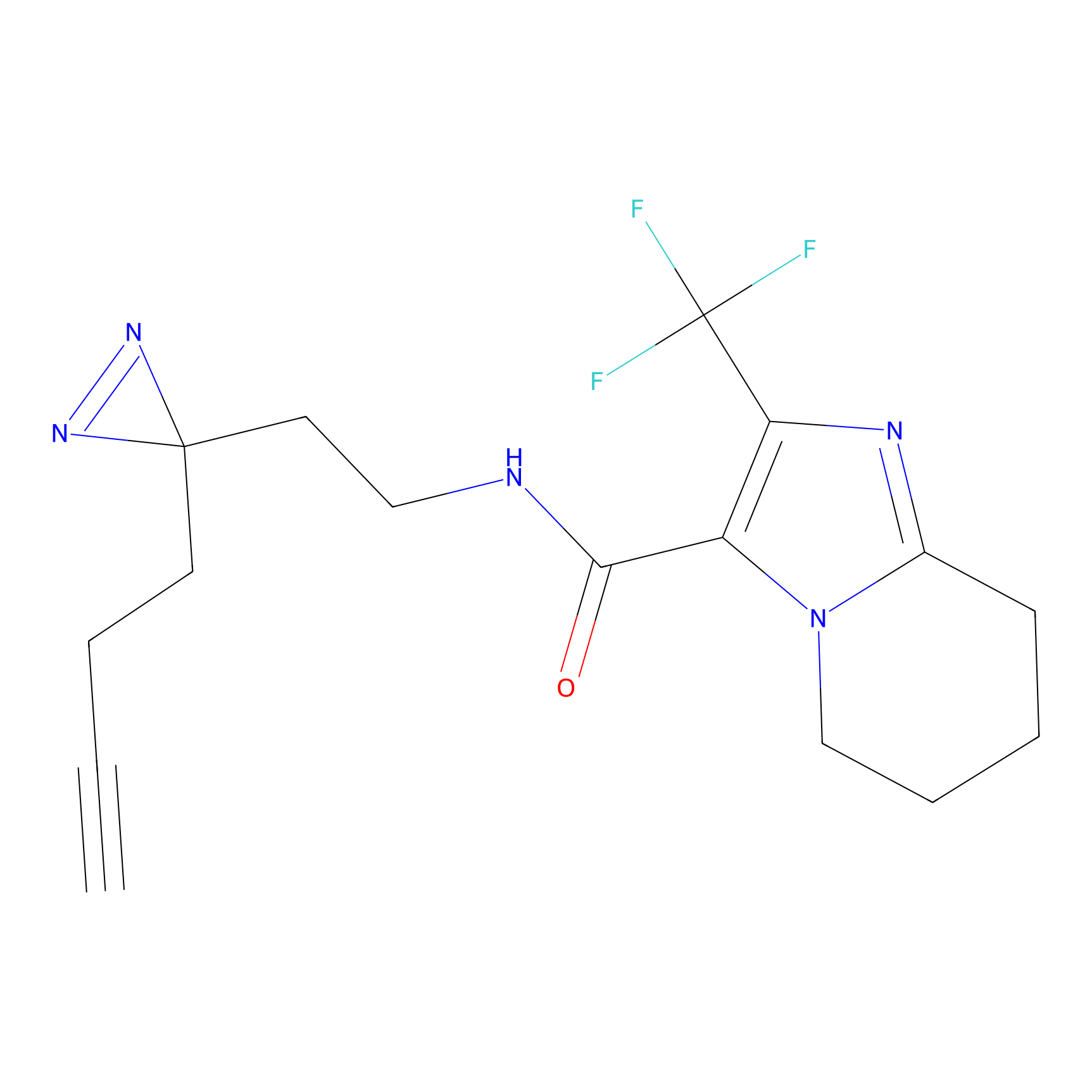
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
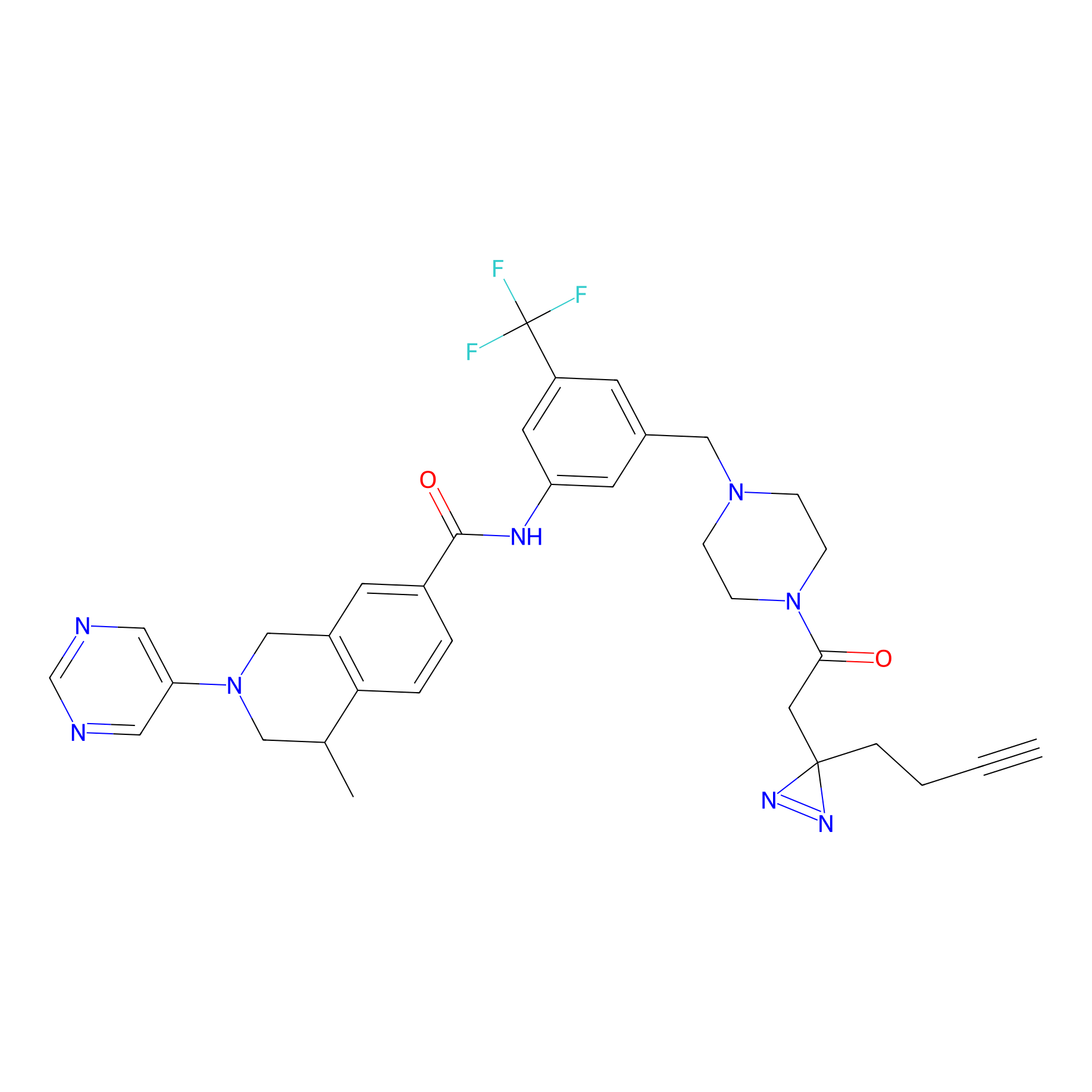
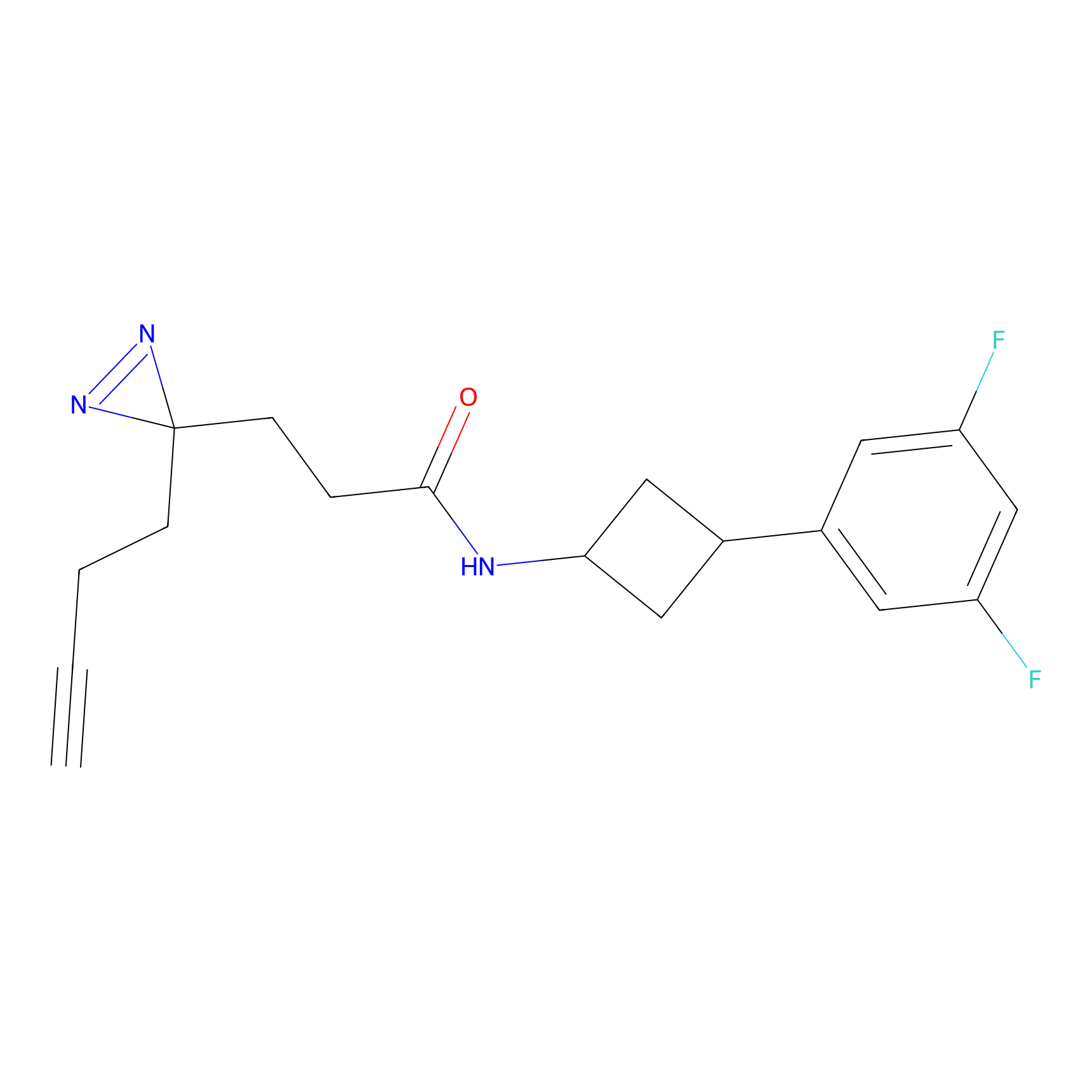
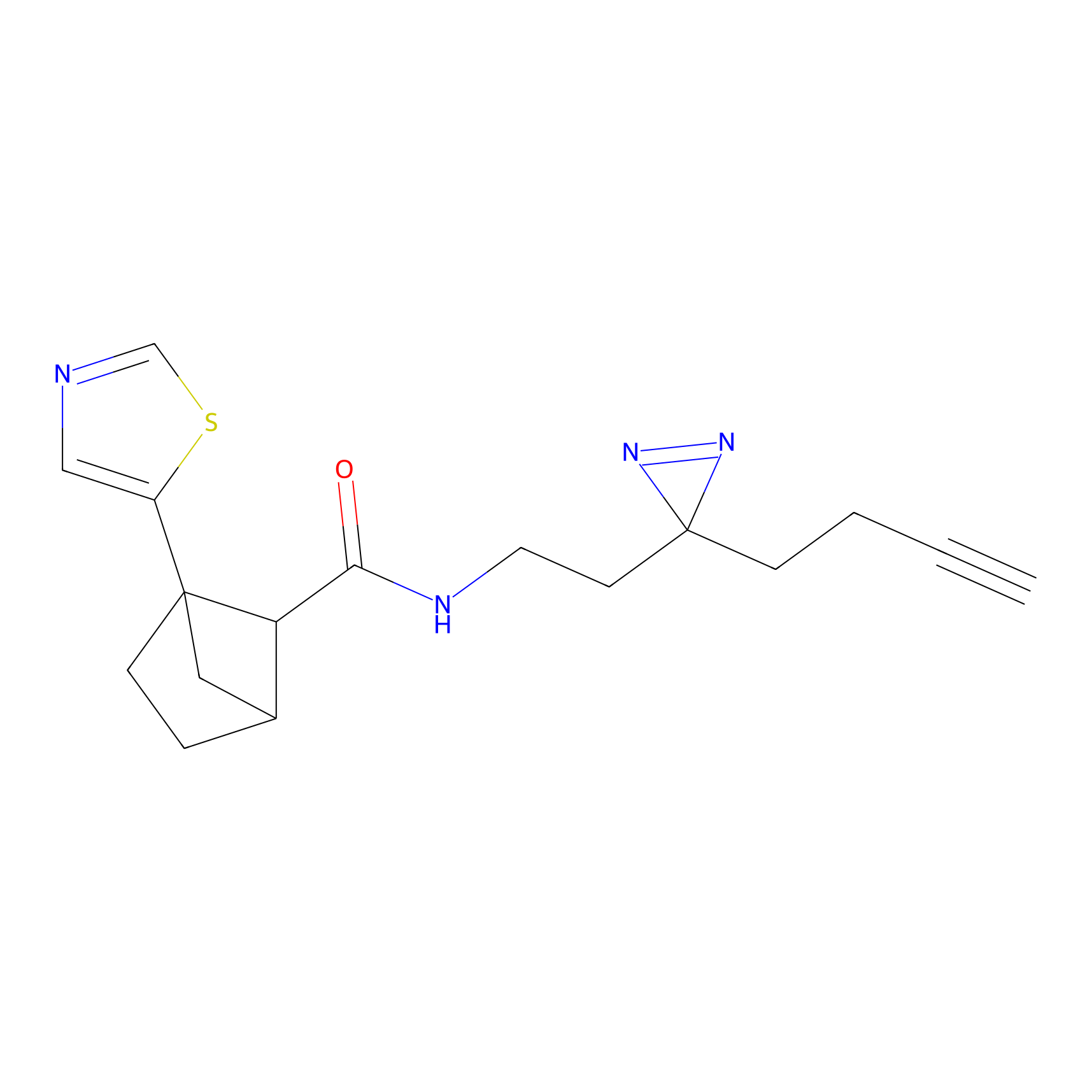
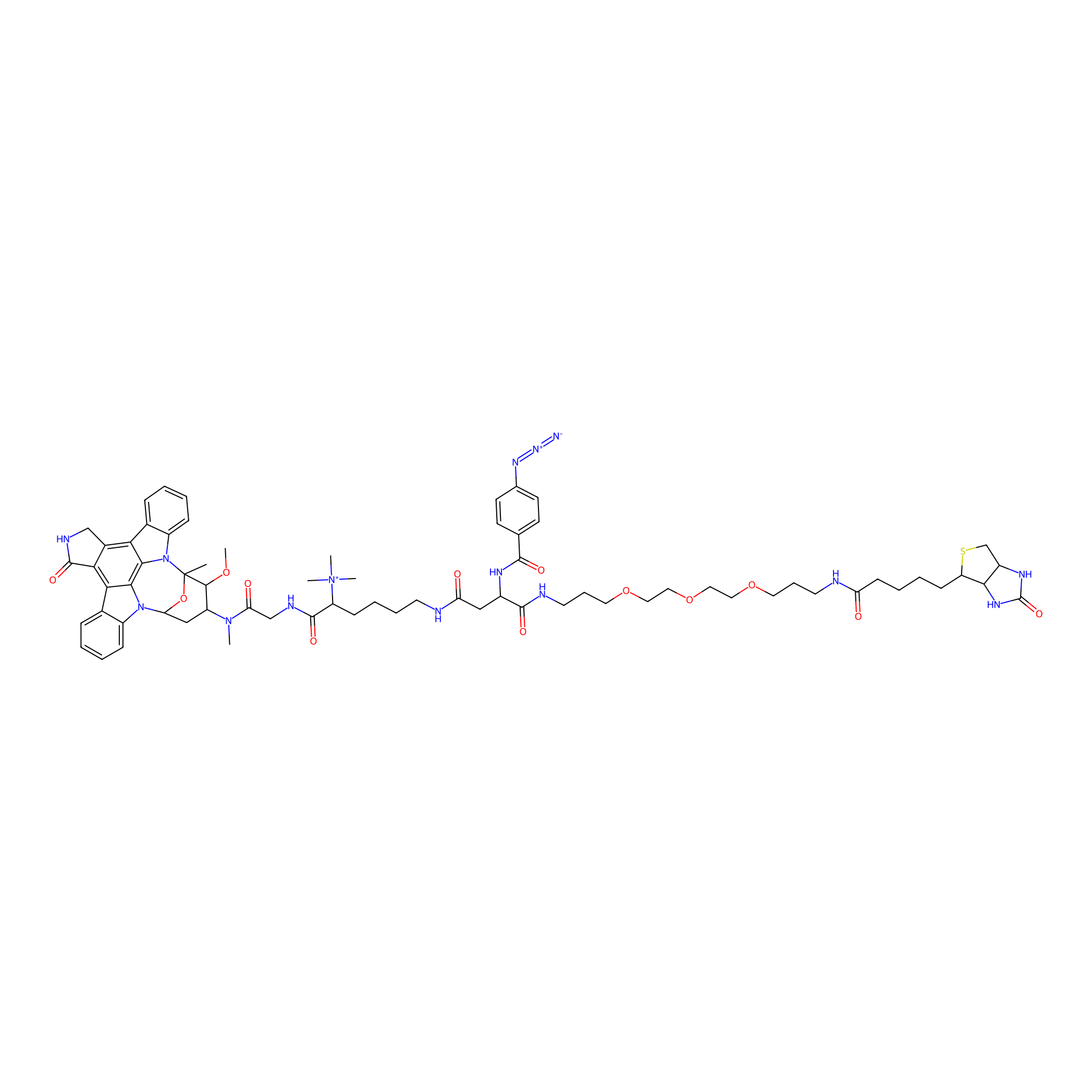
HDSF-alk Probe Info |
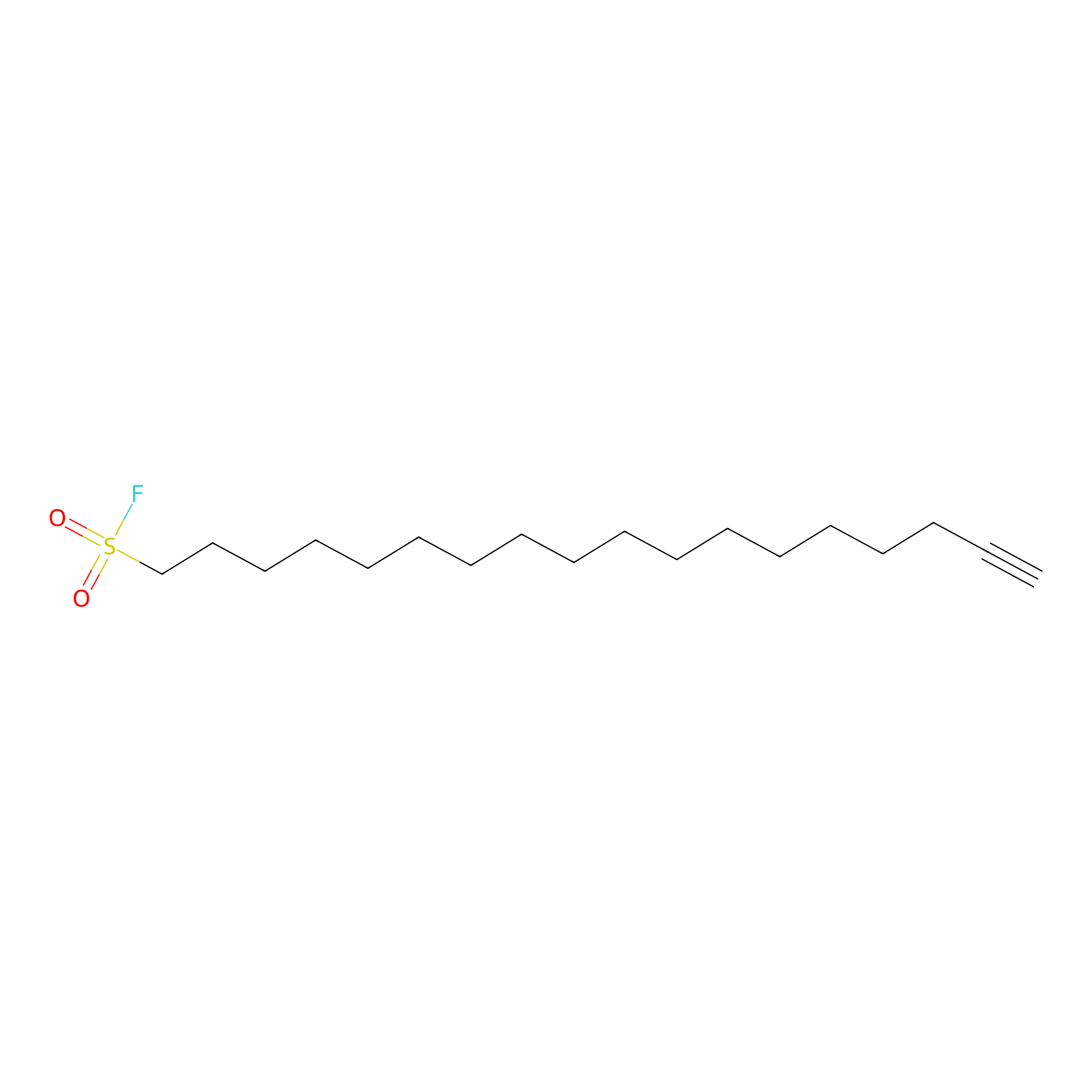 |
2.56 | LDD0197 | [1] | |
|
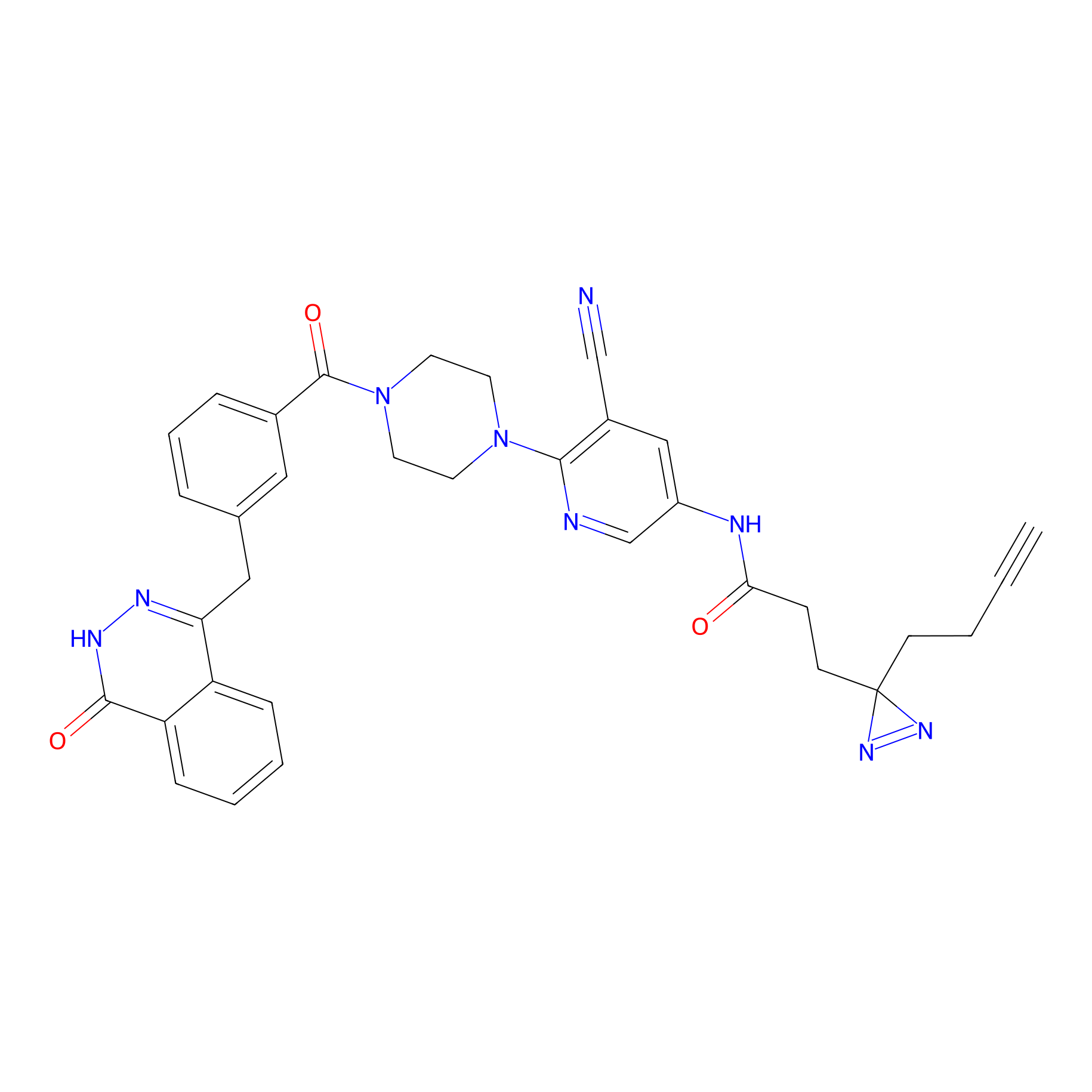
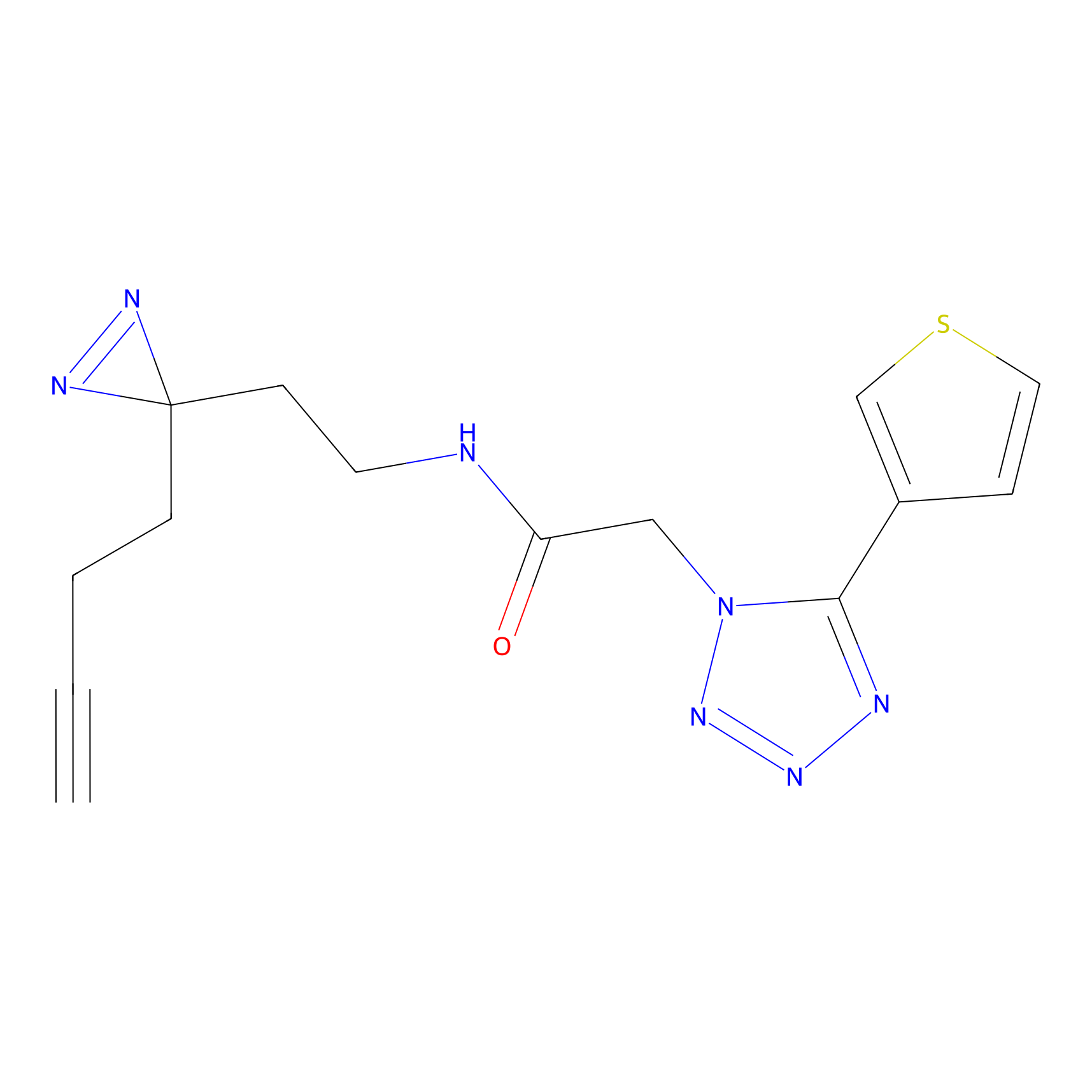
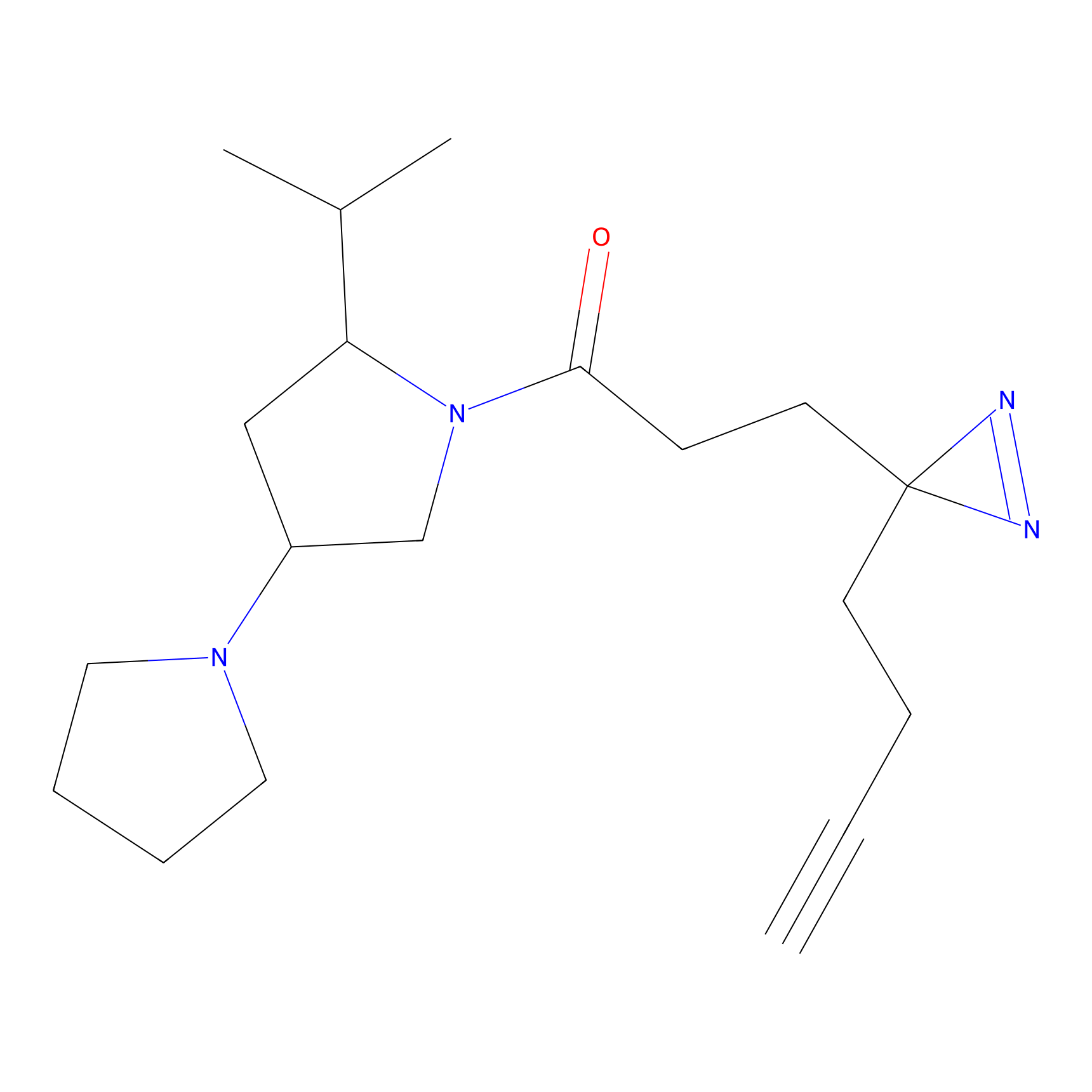
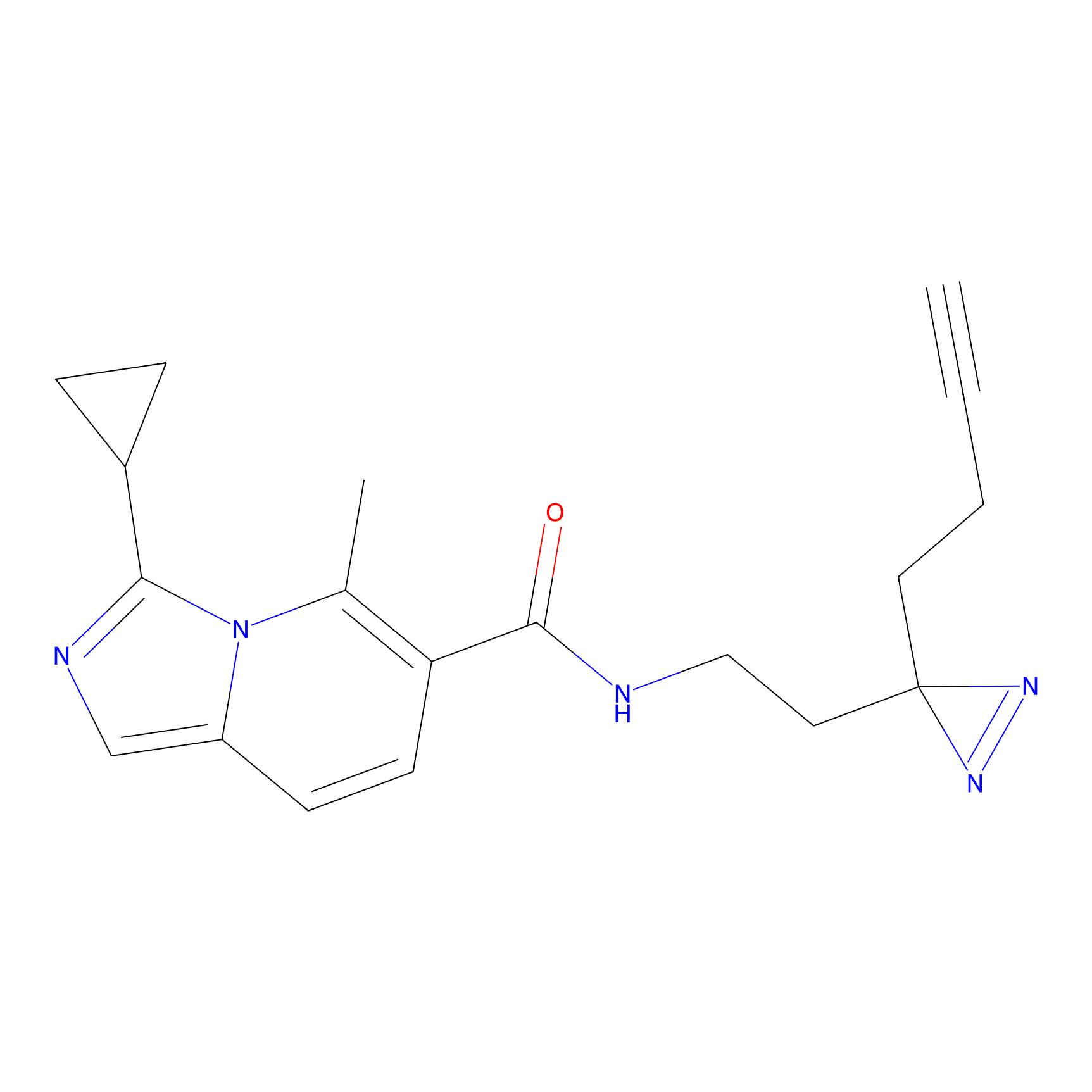
CHEMBL5175495 Probe Info |
 |
9.57 | LDD0196 | [2] | |
|
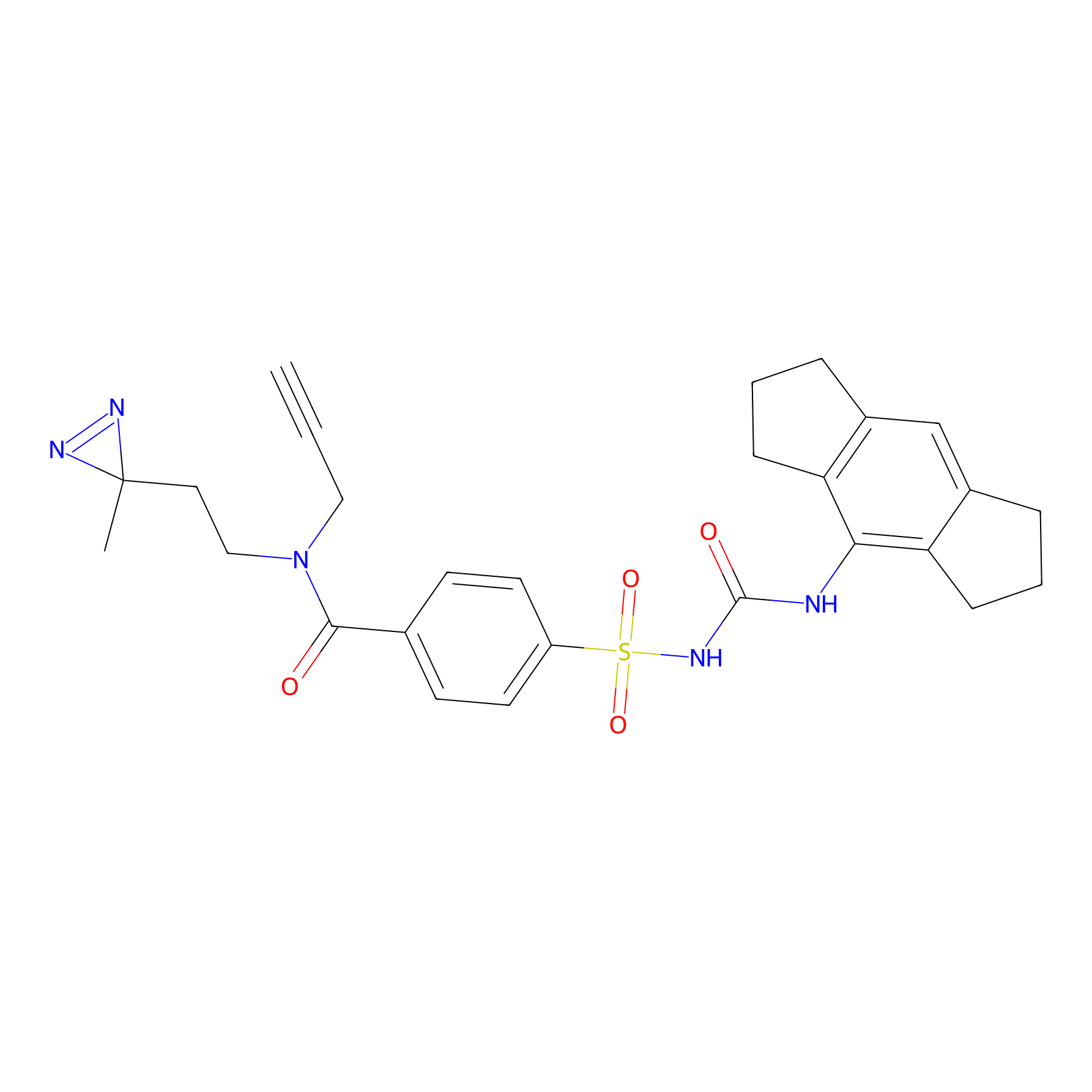
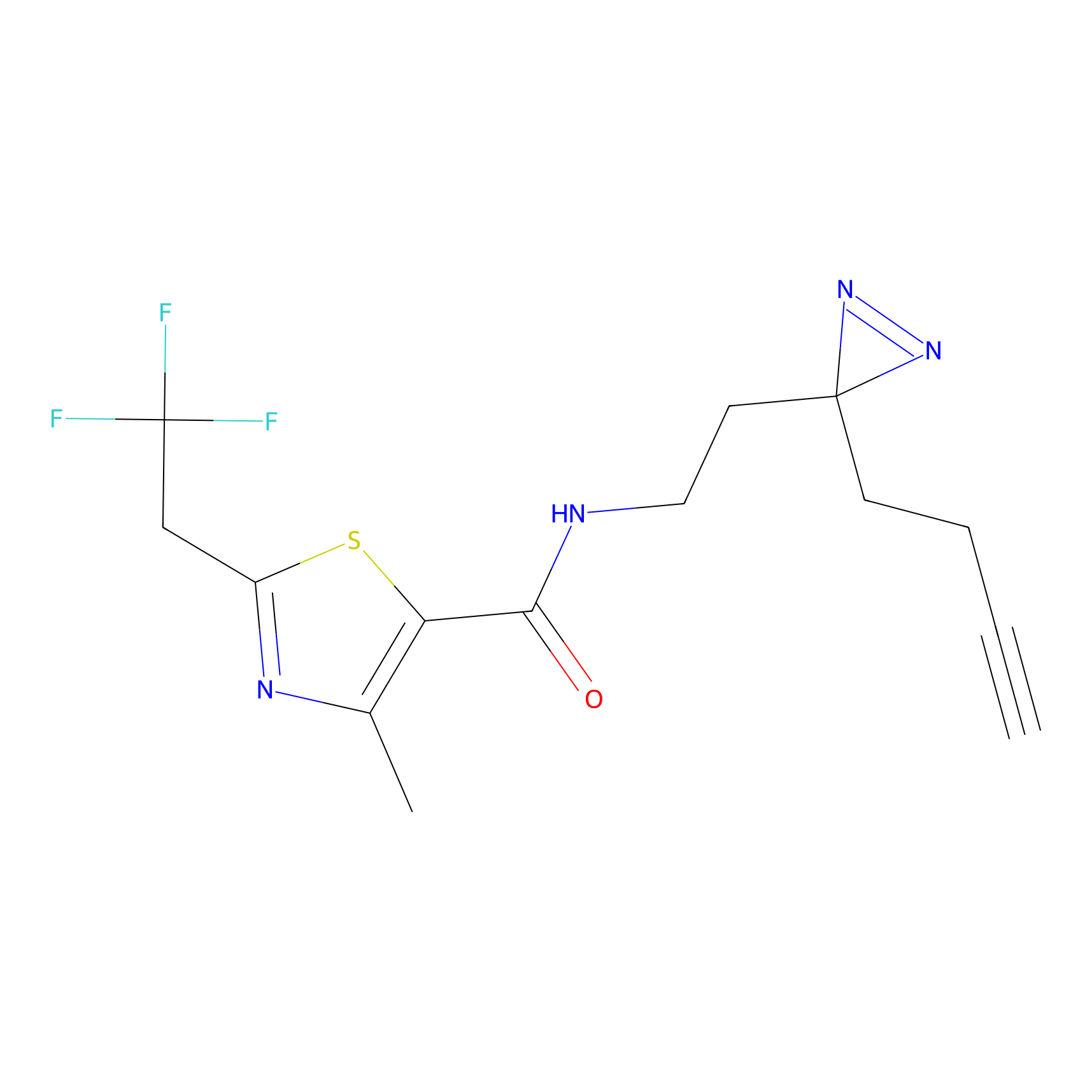
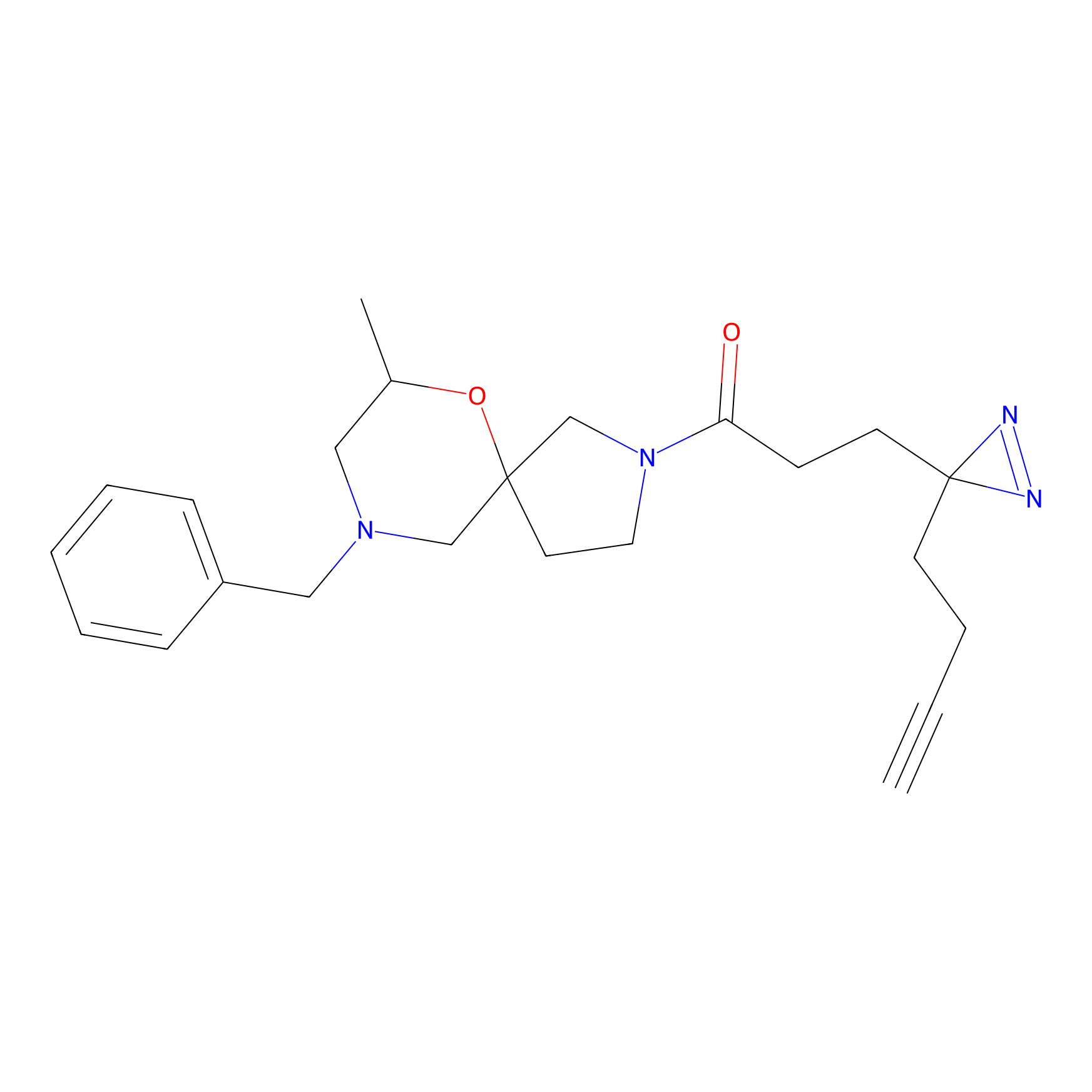
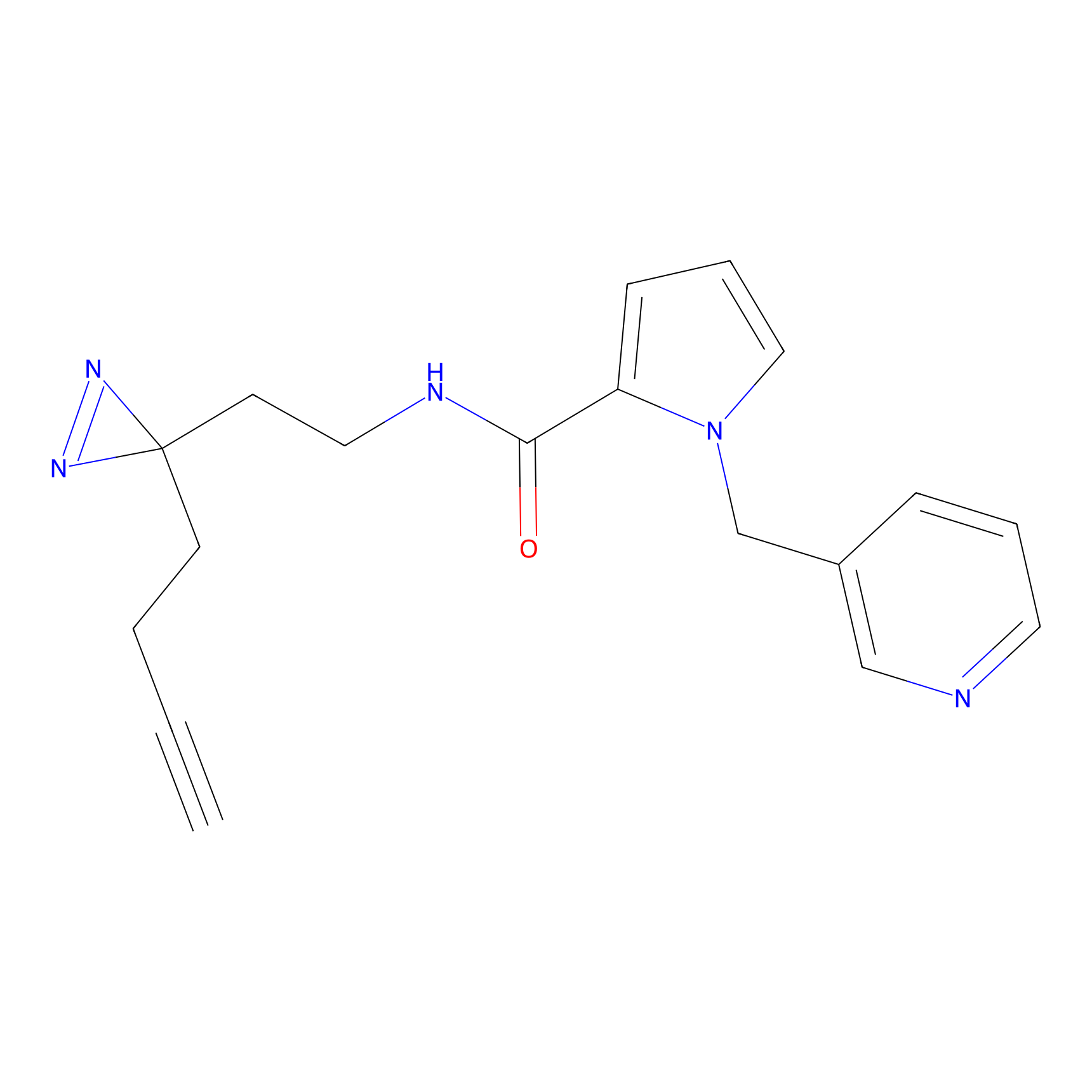
CY4 Probe Info |
 |
100.00 | LDD0244 | [3] | |
|
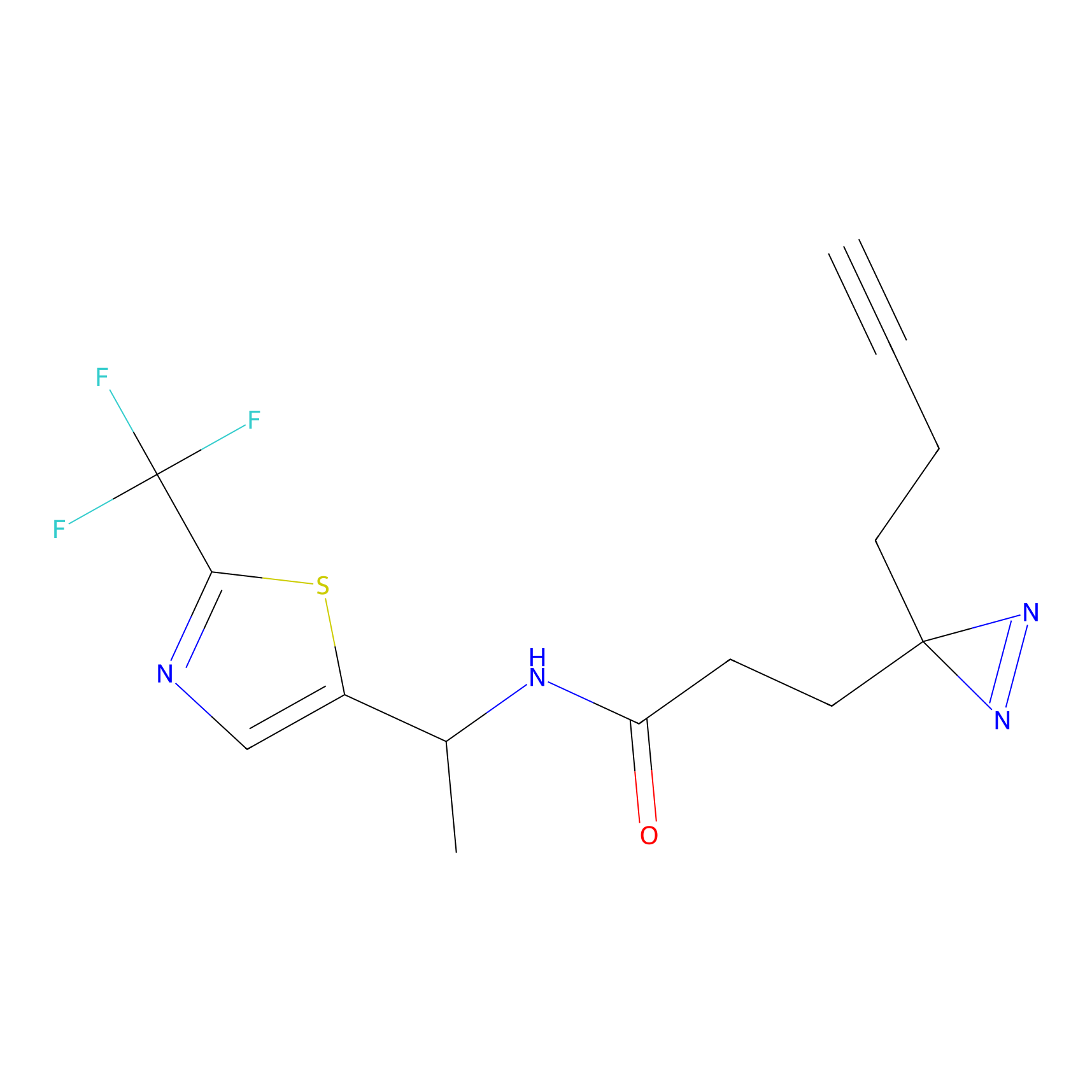
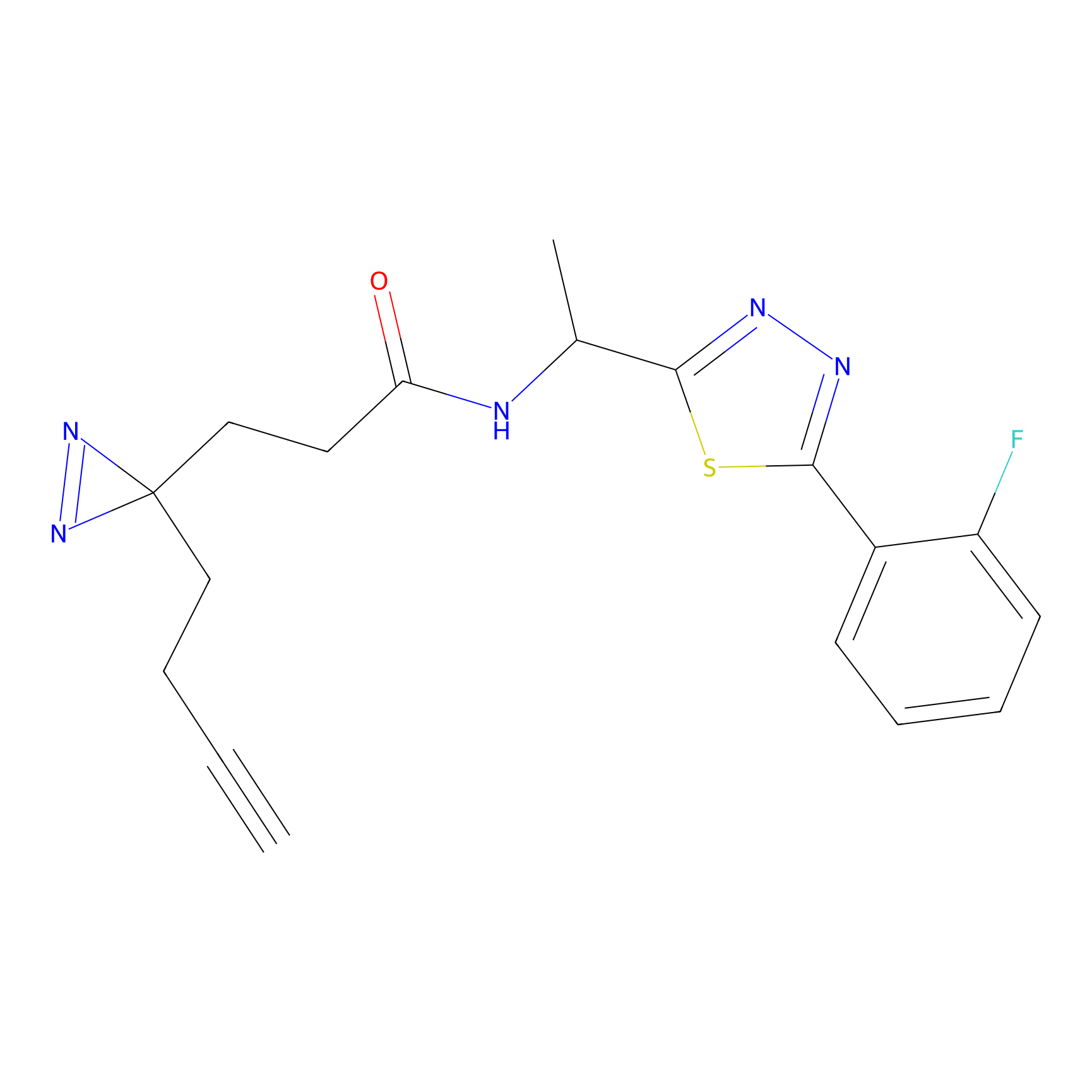
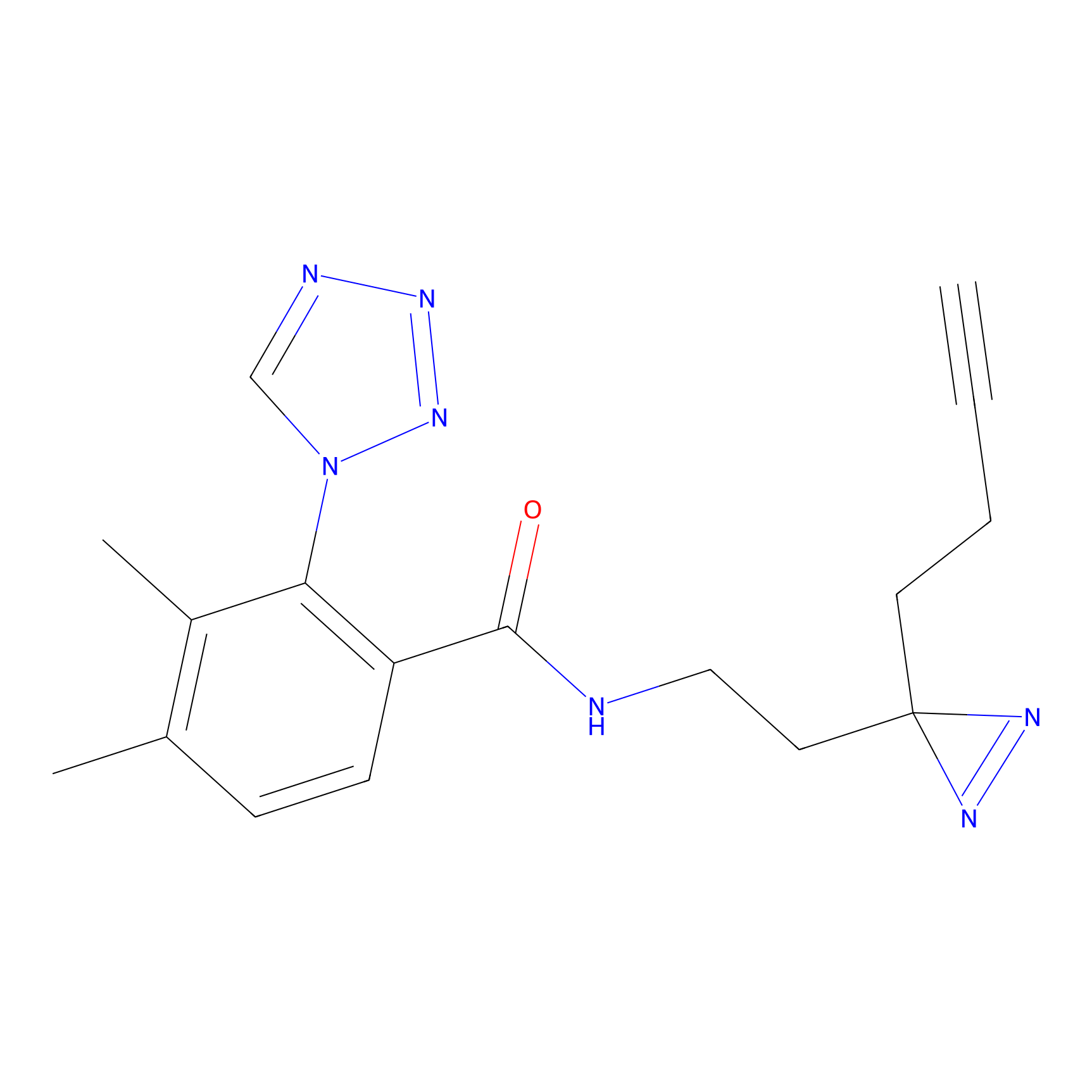
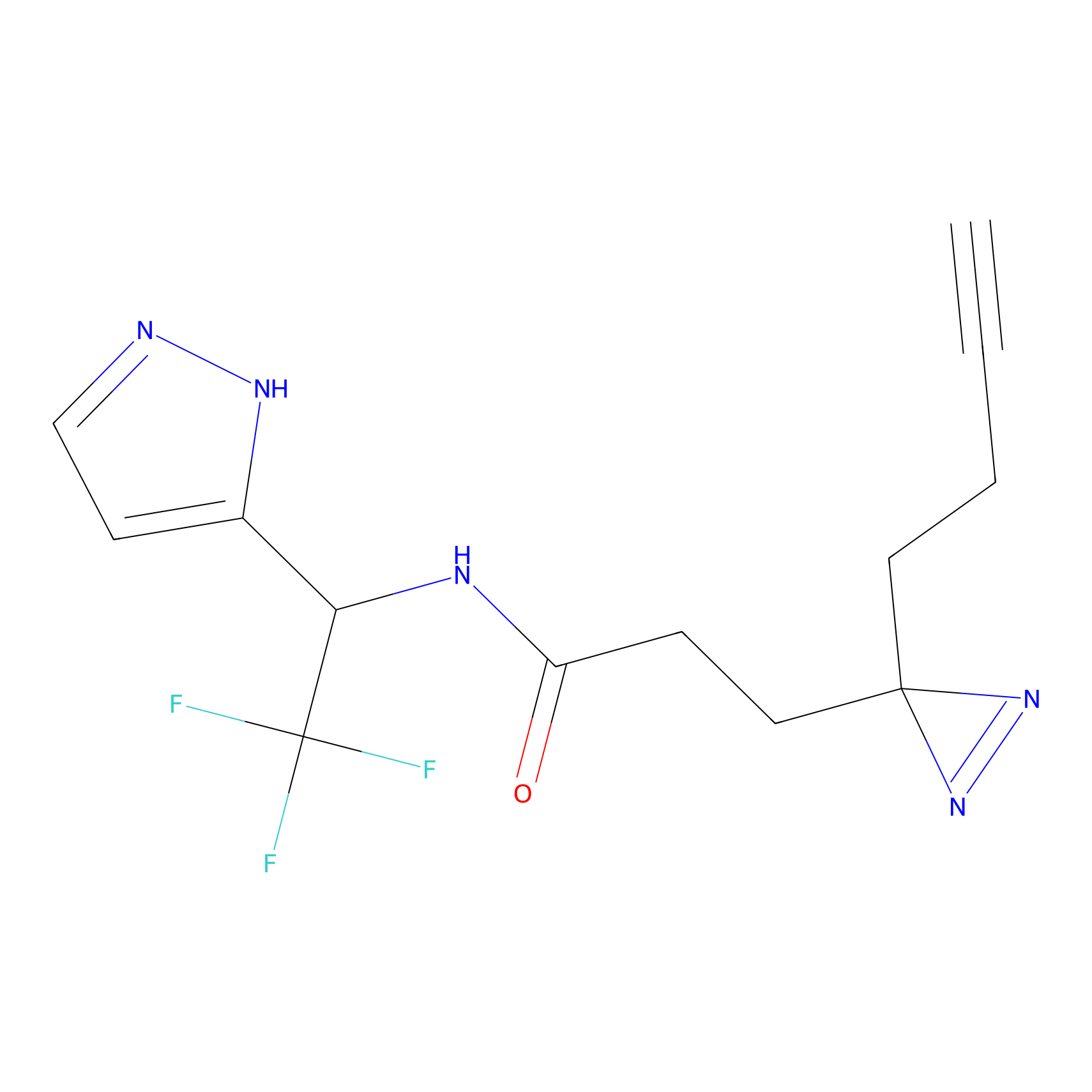
TH211 Probe Info |
 |
Y253(8.19) | LDD0260 | [4] | |
|
C-Sul Probe Info |
 |
3.93 | LDD0066 | [5] | |
|
TH214 Probe Info |
 |
Y253(13.54) | LDD0258 | [4] | |
|
TH216 Probe Info |
 |
Y253(7.03) | LDD0259 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [6] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [6] | |
|
ONAyne Probe Info |
 |
K470(0.68) | LDD0274 | [7] | |
|
STPyne Probe Info |
 |
K251(3.58); K259(7.69); K423(7.14); K429(0.39) | LDD0277 | [7] | |
|
OPA-S-S-alkyne Probe Info |
 |
K259(2.08) | LDD3494 | [8] | |
|
Probe 1 Probe Info |
 |
Y253(9.06); Y432(13.10) | LDD3495 | [9] | |
|
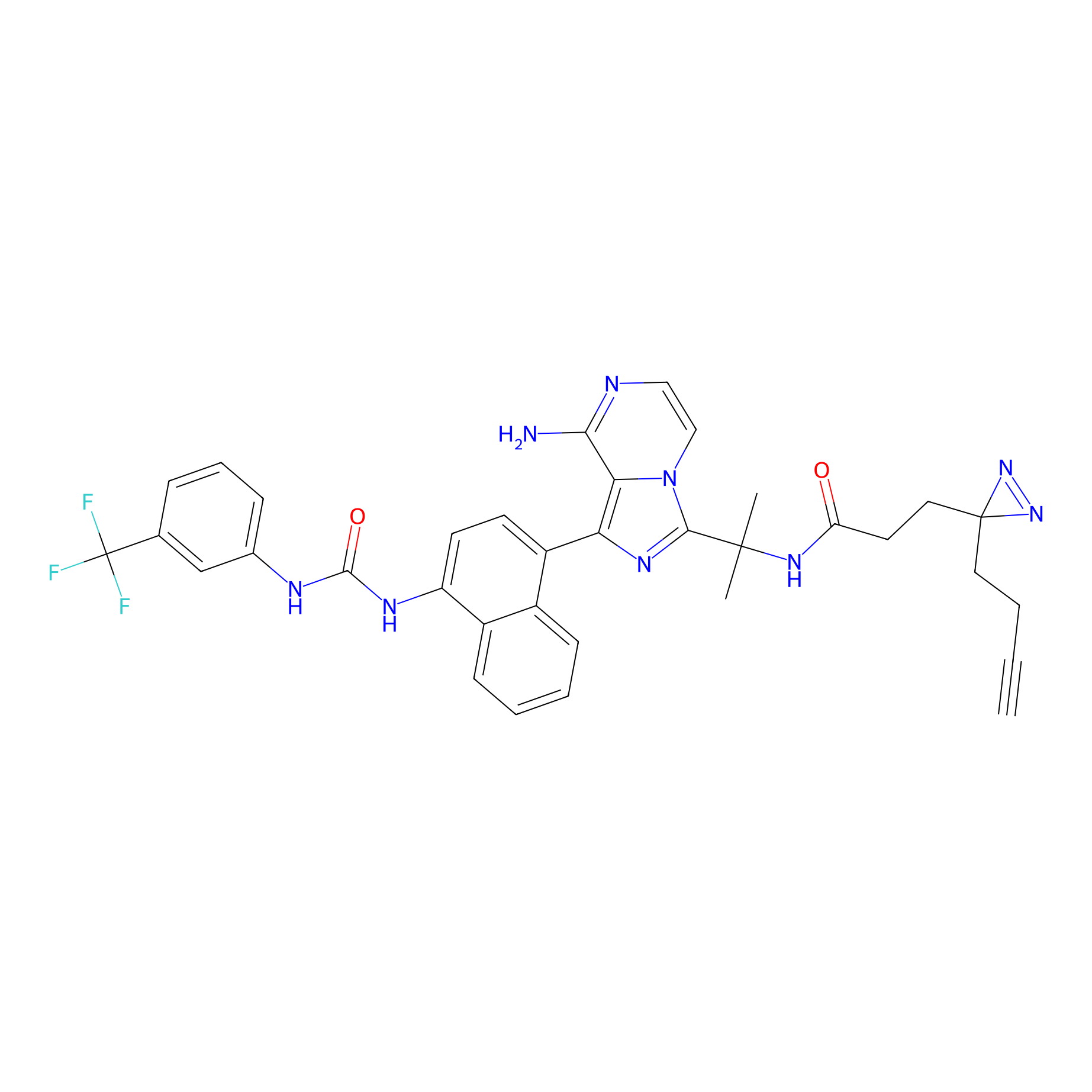
P26 Probe Info |
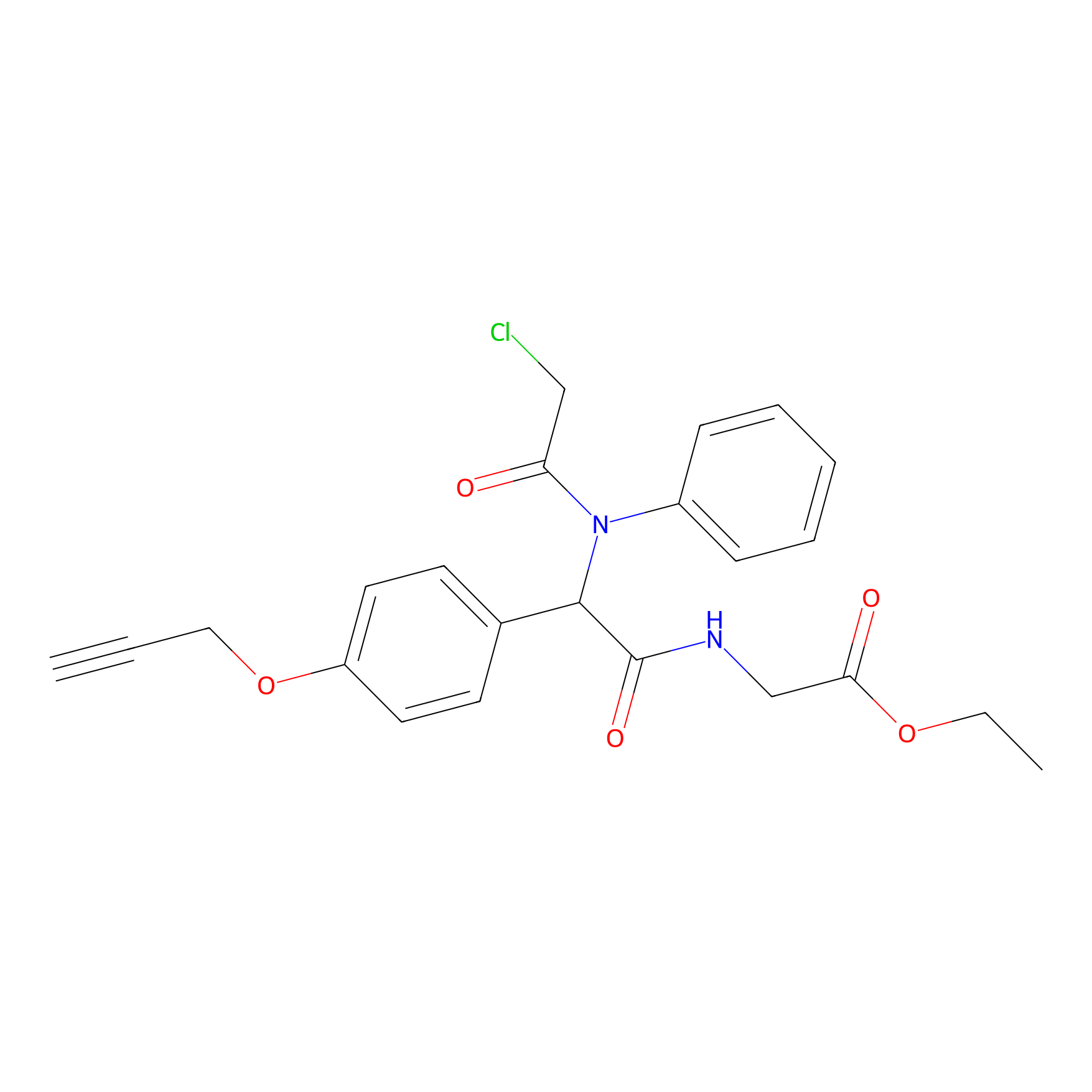 |
4.18 | LDD0409 | [10] | |
|
HPAP Probe Info |
 |
5.27 | LDD0064 | [11] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [12] | |
|
NHS Probe Info |
 |
K251(0.00); K429(0.00) | LDD0010 | [13] | |
|
SF Probe Info |
 |
Y253(0.00); Y432(0.00) | LDD0028 | [14] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [15] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [16] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H50(0.00); H335(0.00) | LDD0222 | [15] |
| LDCM0107 | IAA | HeLa | H339(0.00); H50(0.00); H335(0.00) | LDD0221 | [15] |
| LDCM0109 | NEM | HeLa | H50(0.00); H339(0.00) | LDD0223 | [15] |
| LDCM0159 | P28 | BxPC-3 | 4.18 | LDD0409 | [10] |
| LDCM0014 | Panhematin | hPBMC | 5.27 | LDD0064 | [11] |
| LDCM0090 | Rapamycin | JHH-7 | 7.09 | LDD0213 | [24] |
| LDCM0019 | Staurosporine | Hep-G2 | N.A. | LDD0083 | [25] |
References