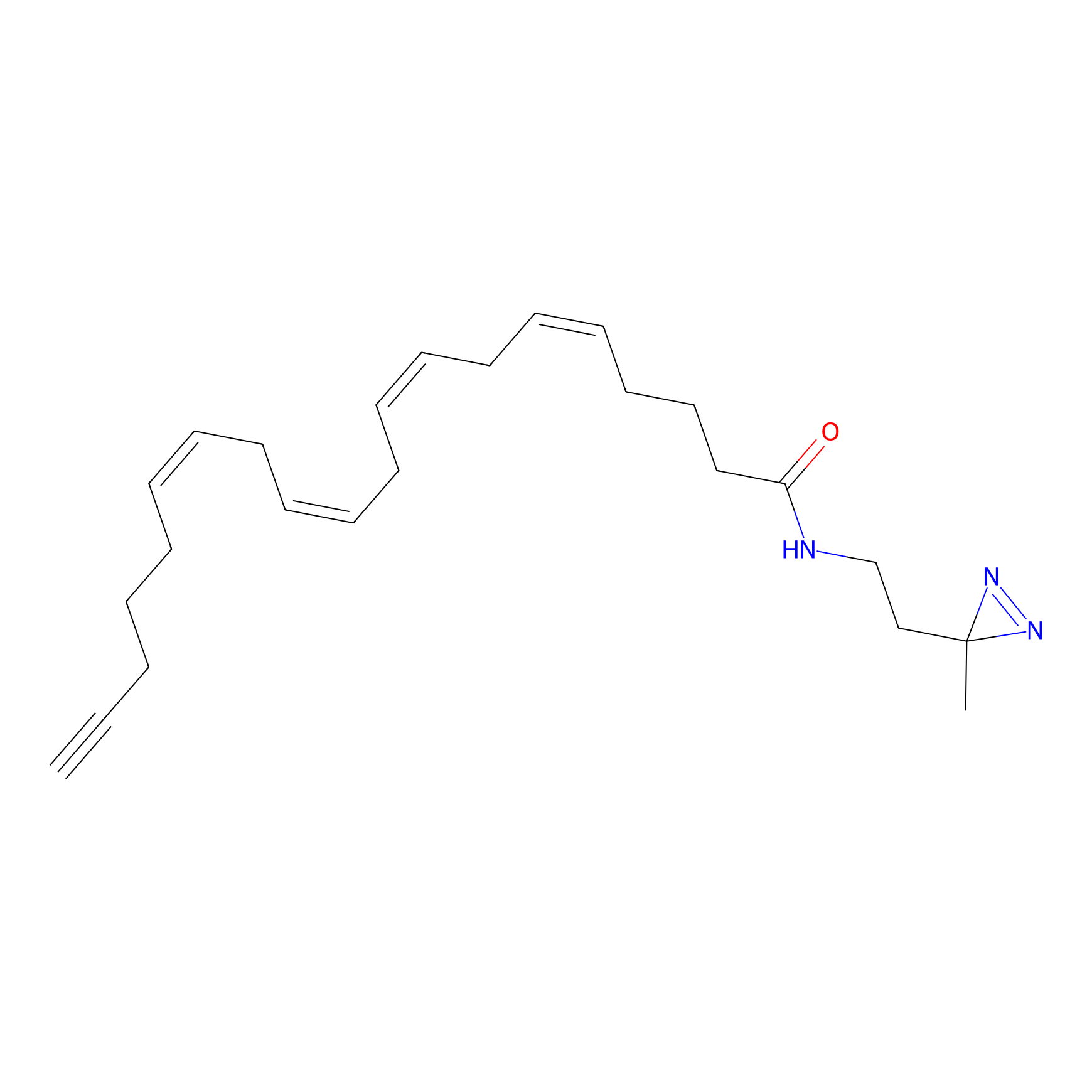
Details of the Target
General Information of Target
Probe(s) Labeling This Target
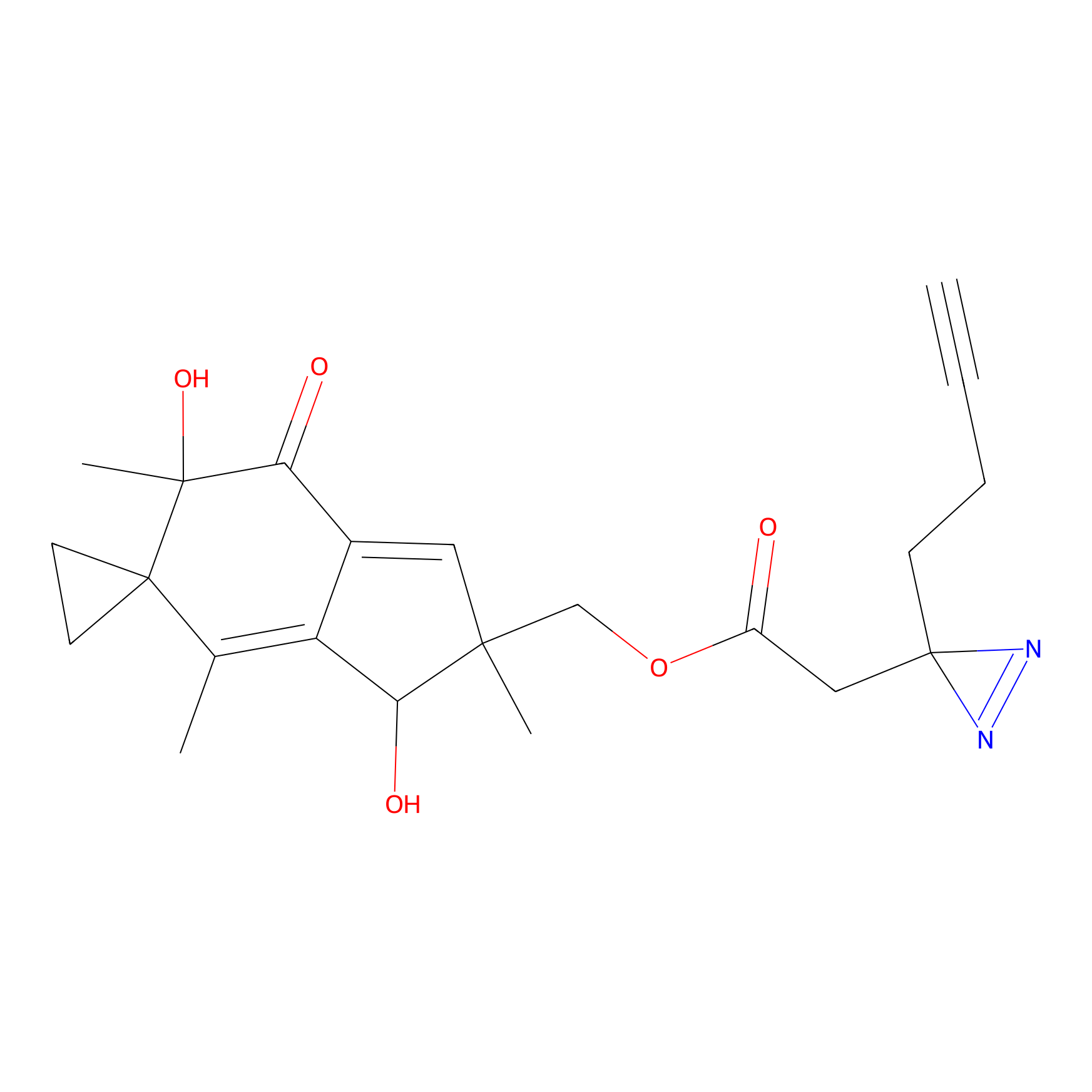
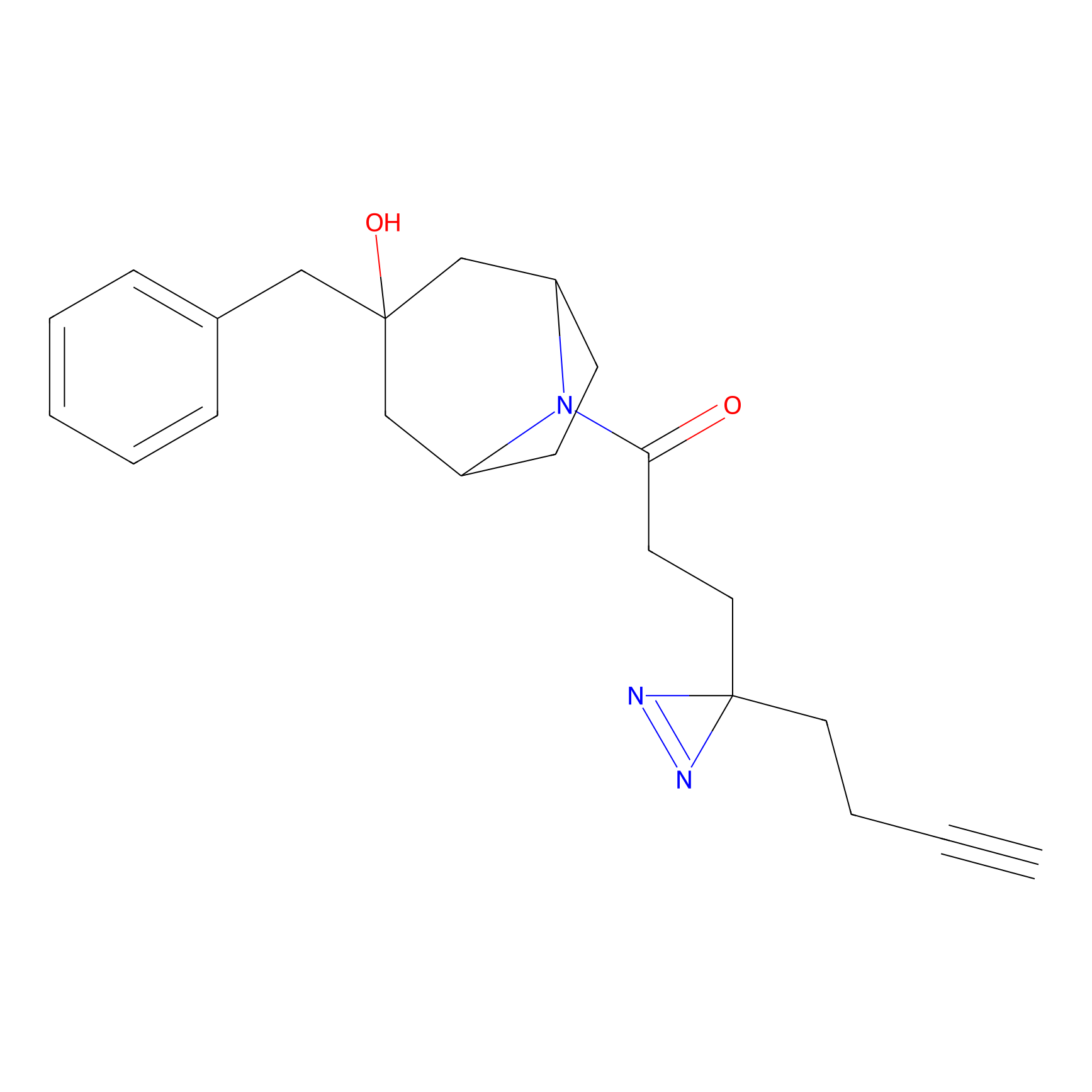
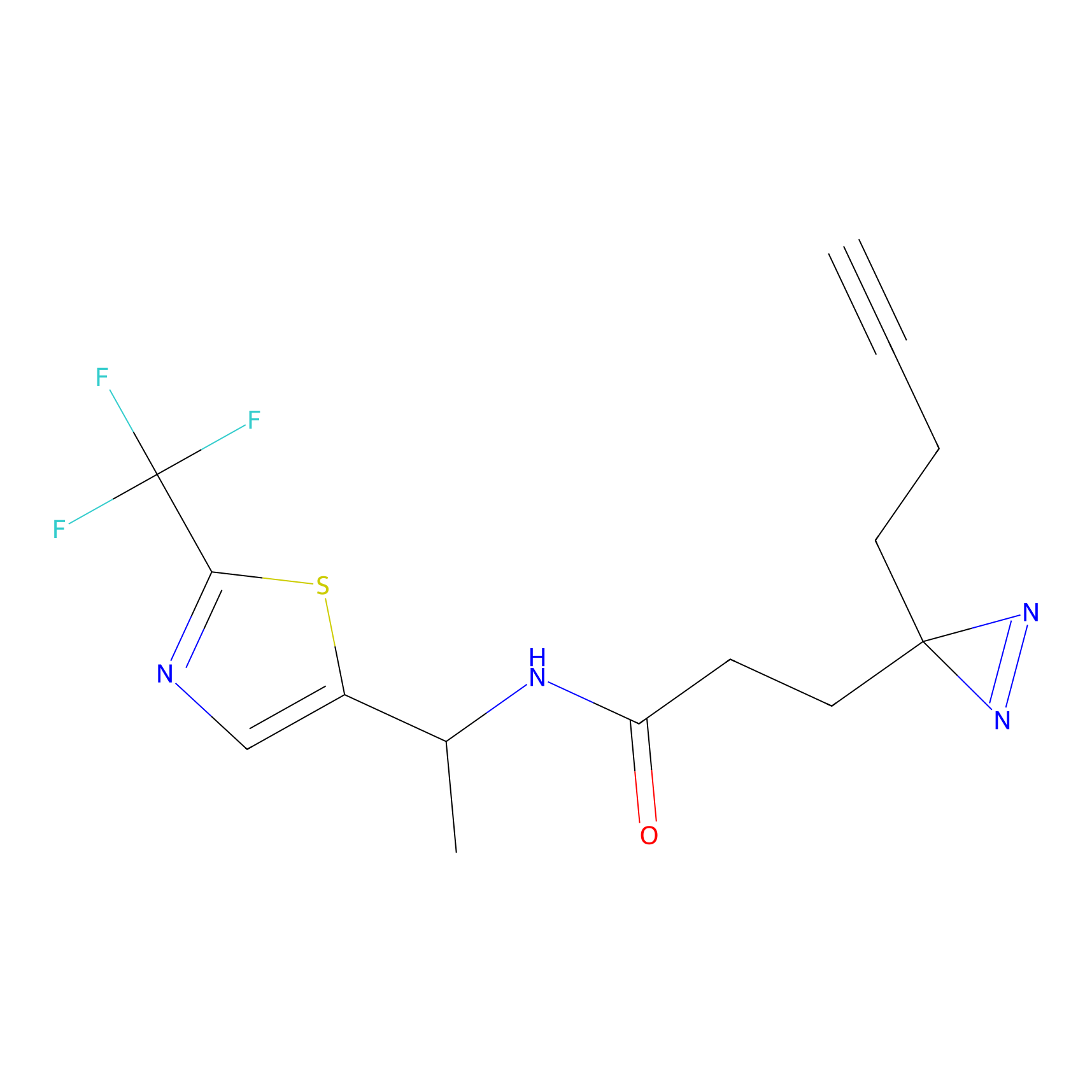
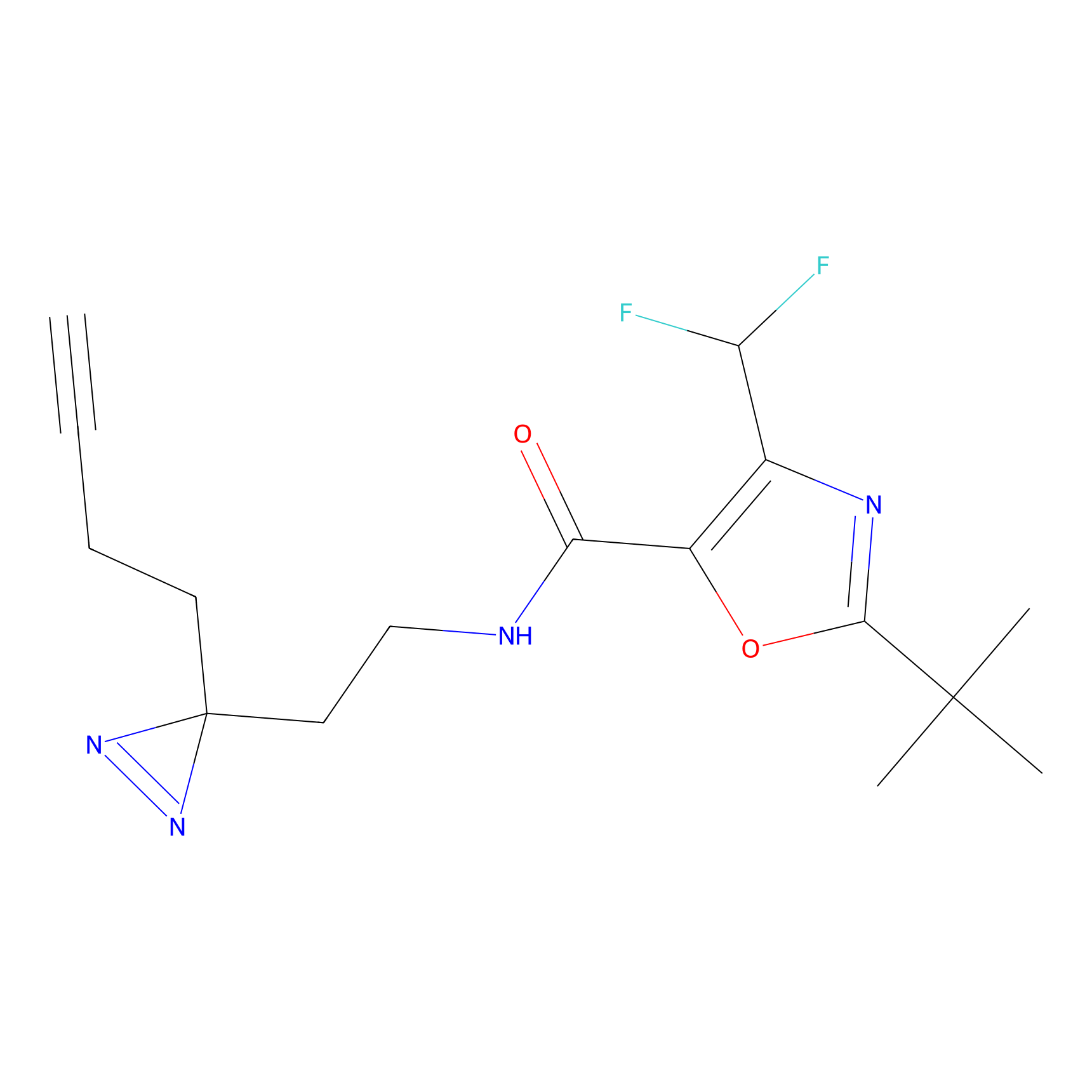
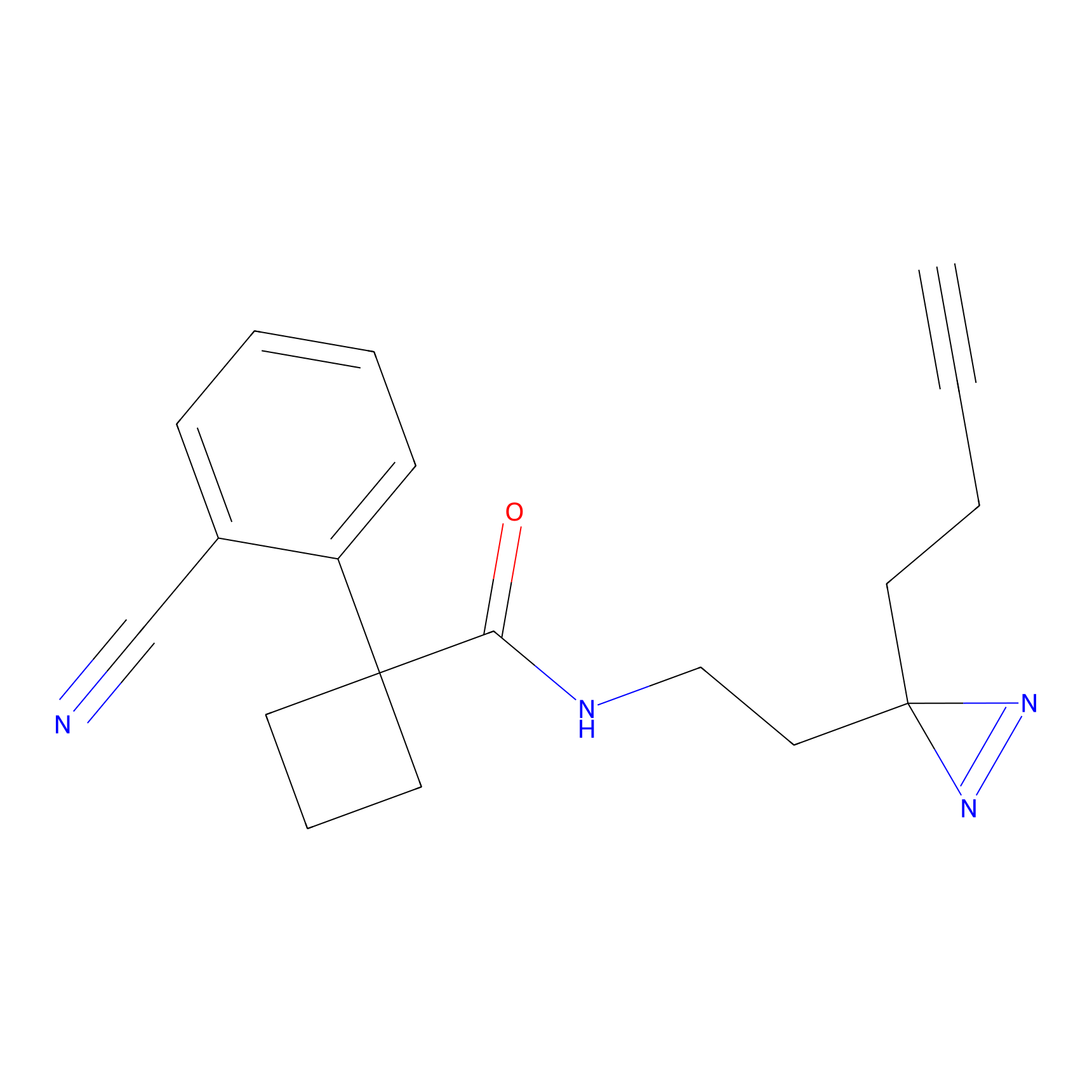
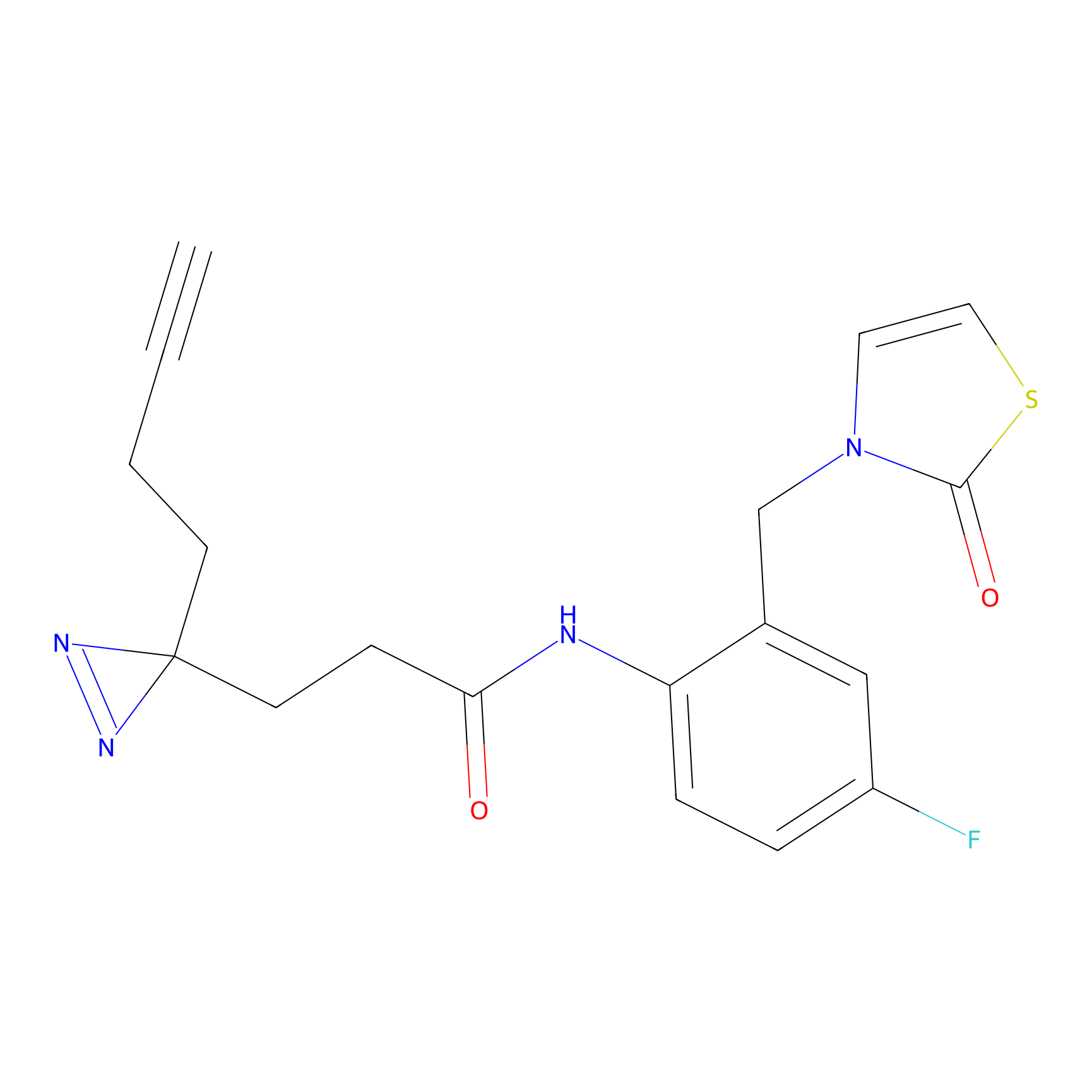
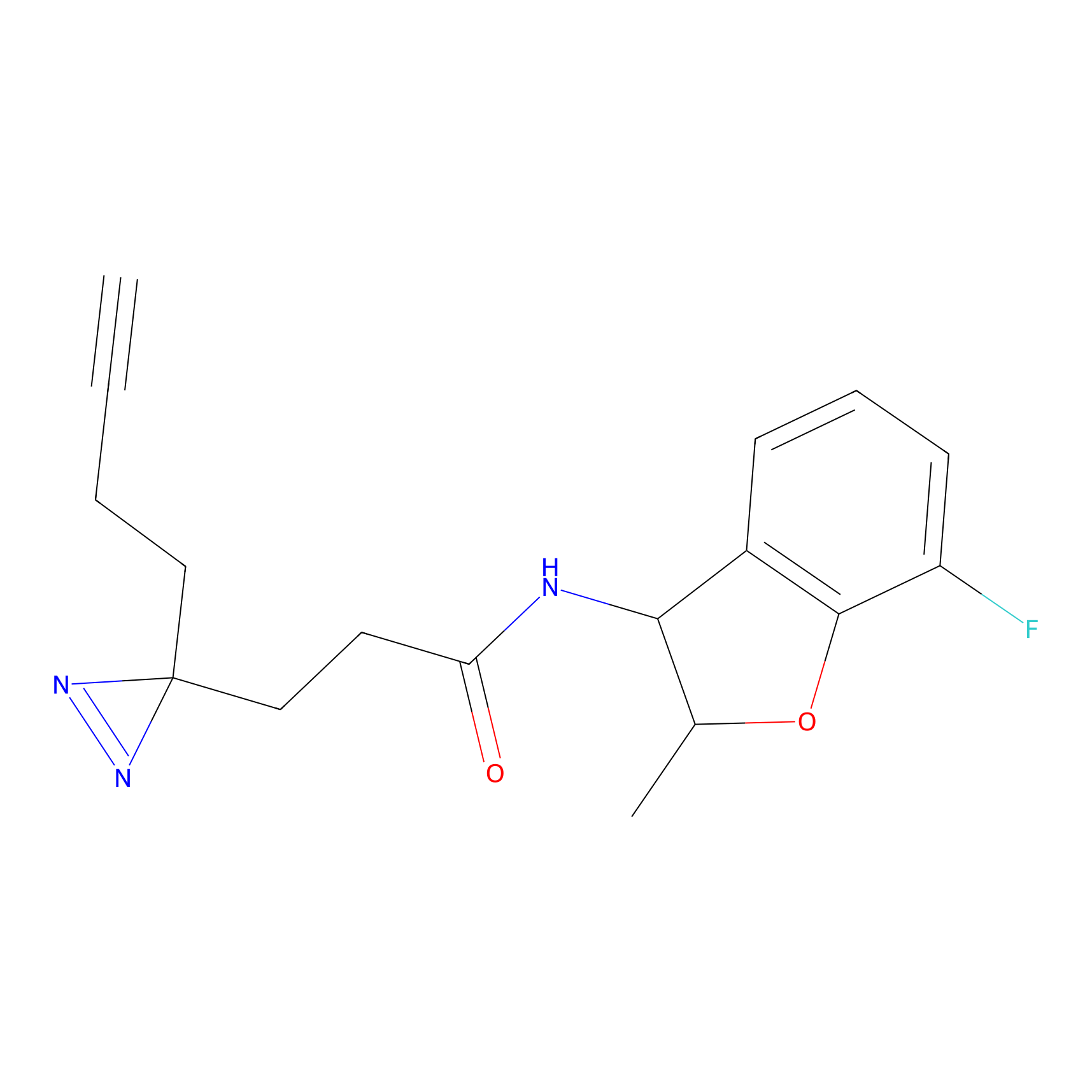
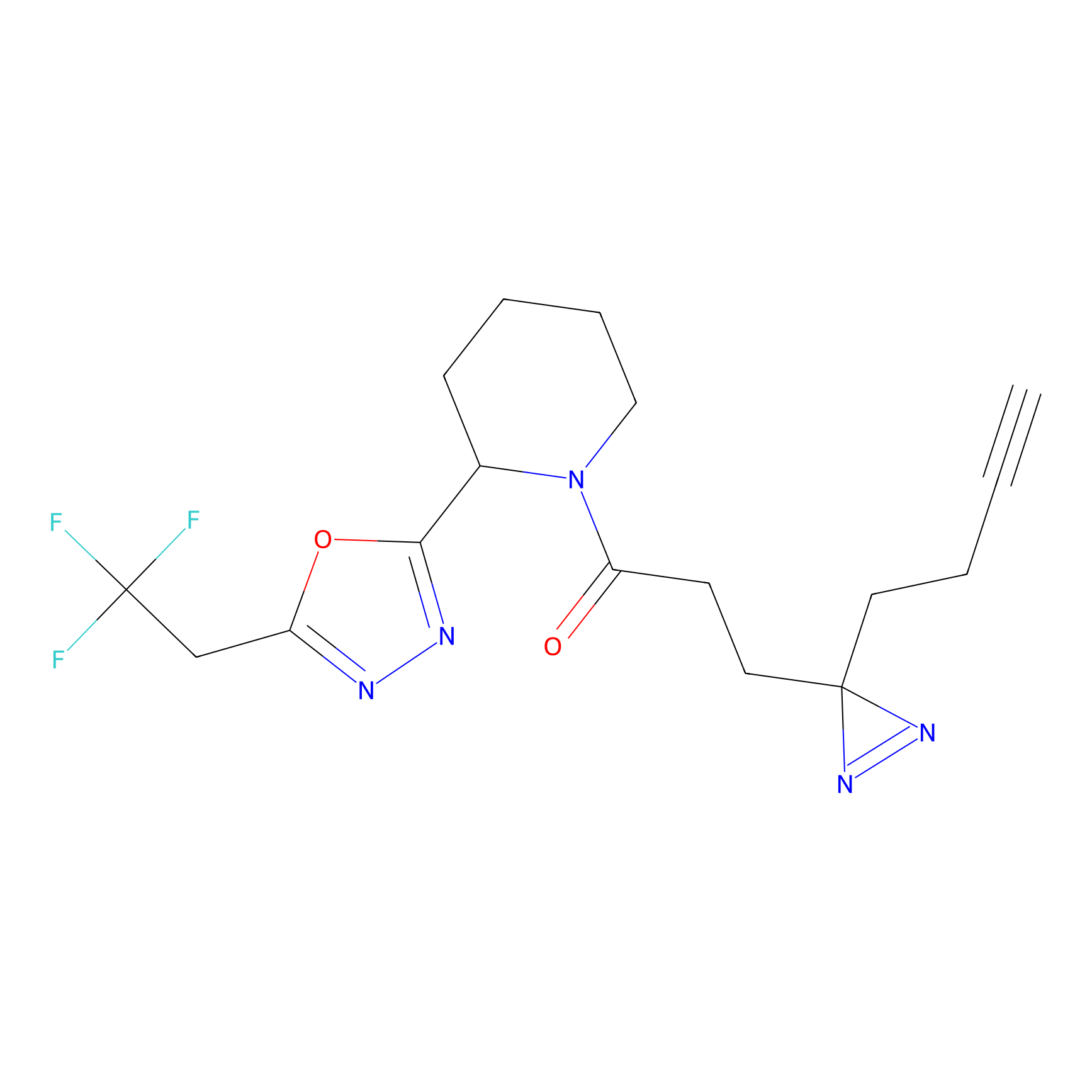
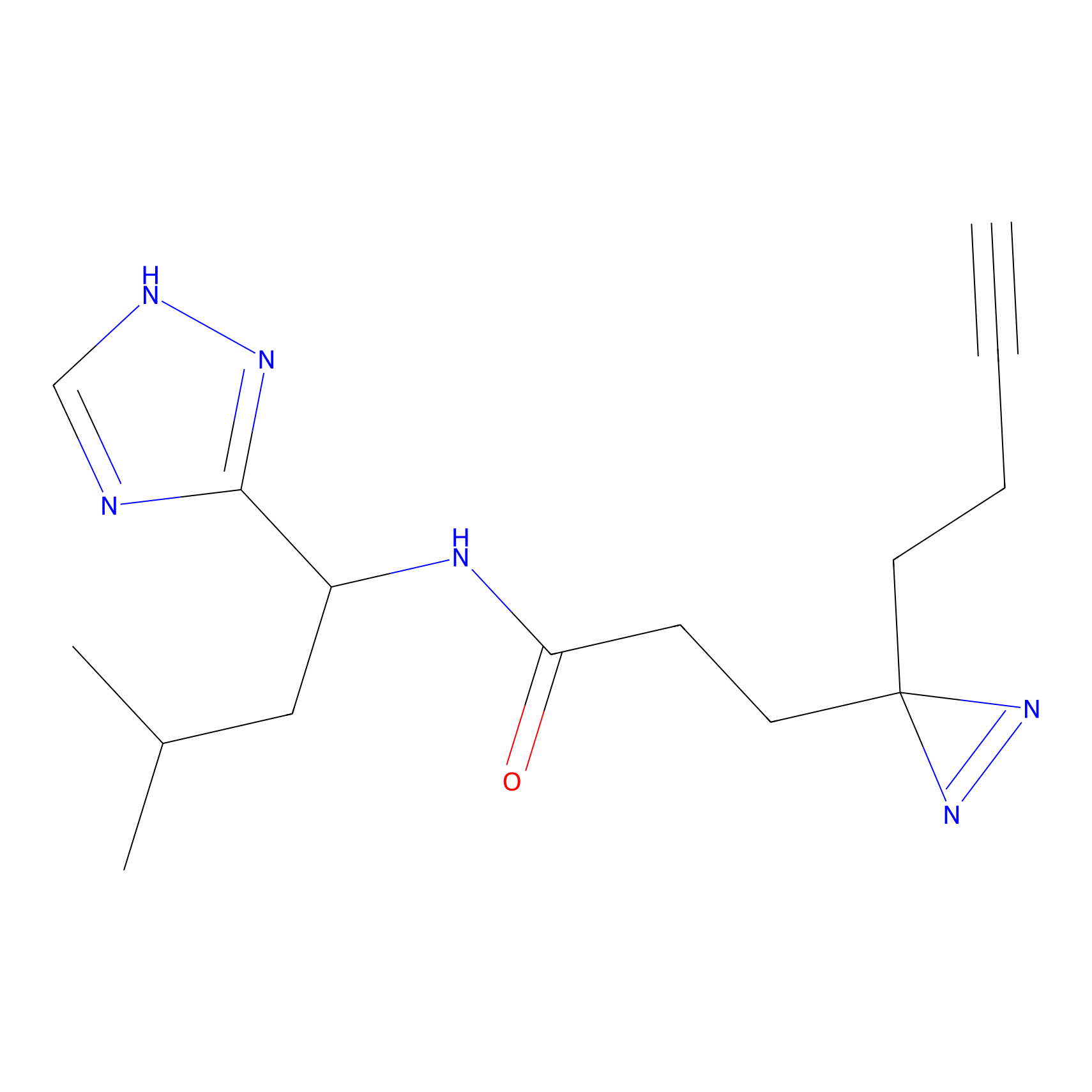
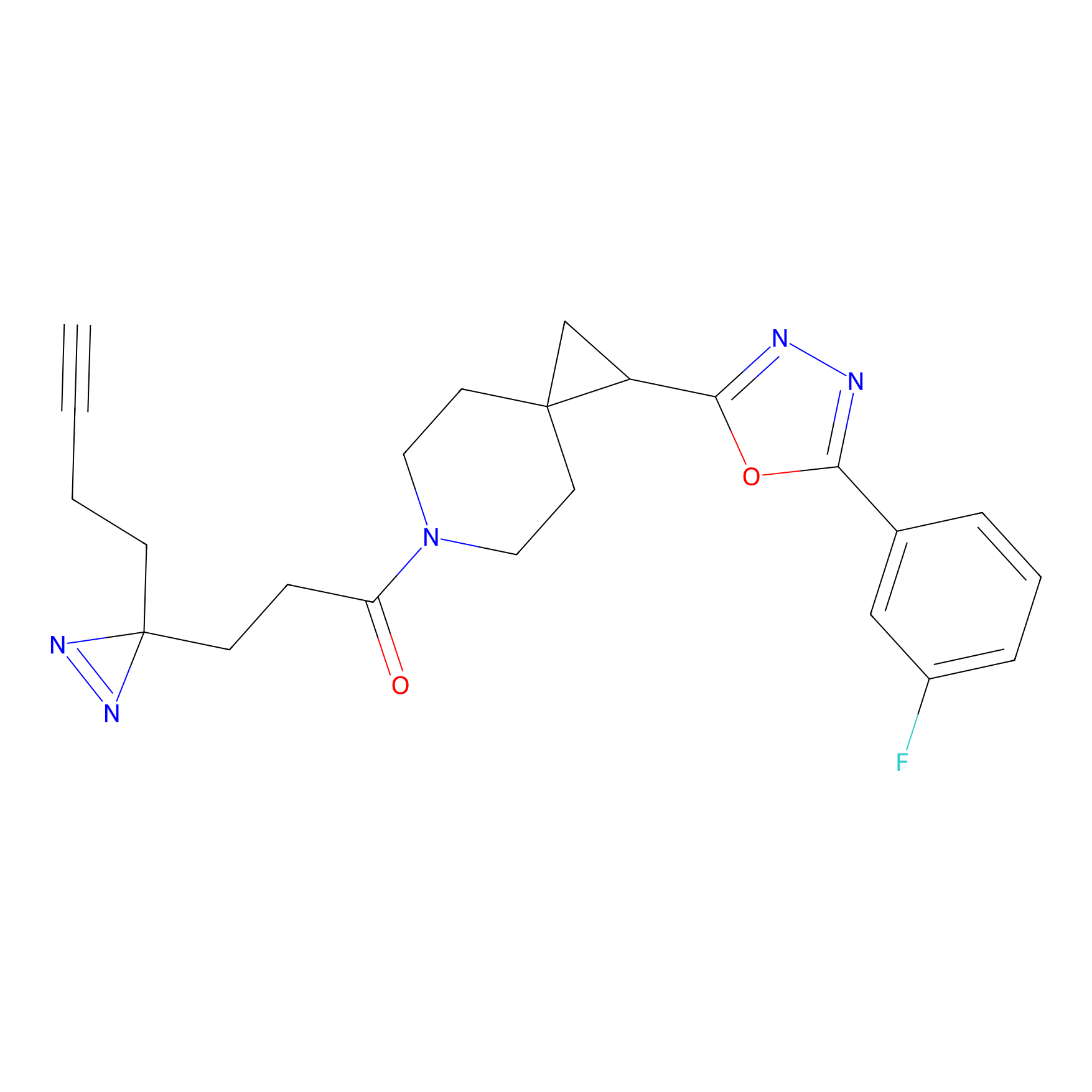
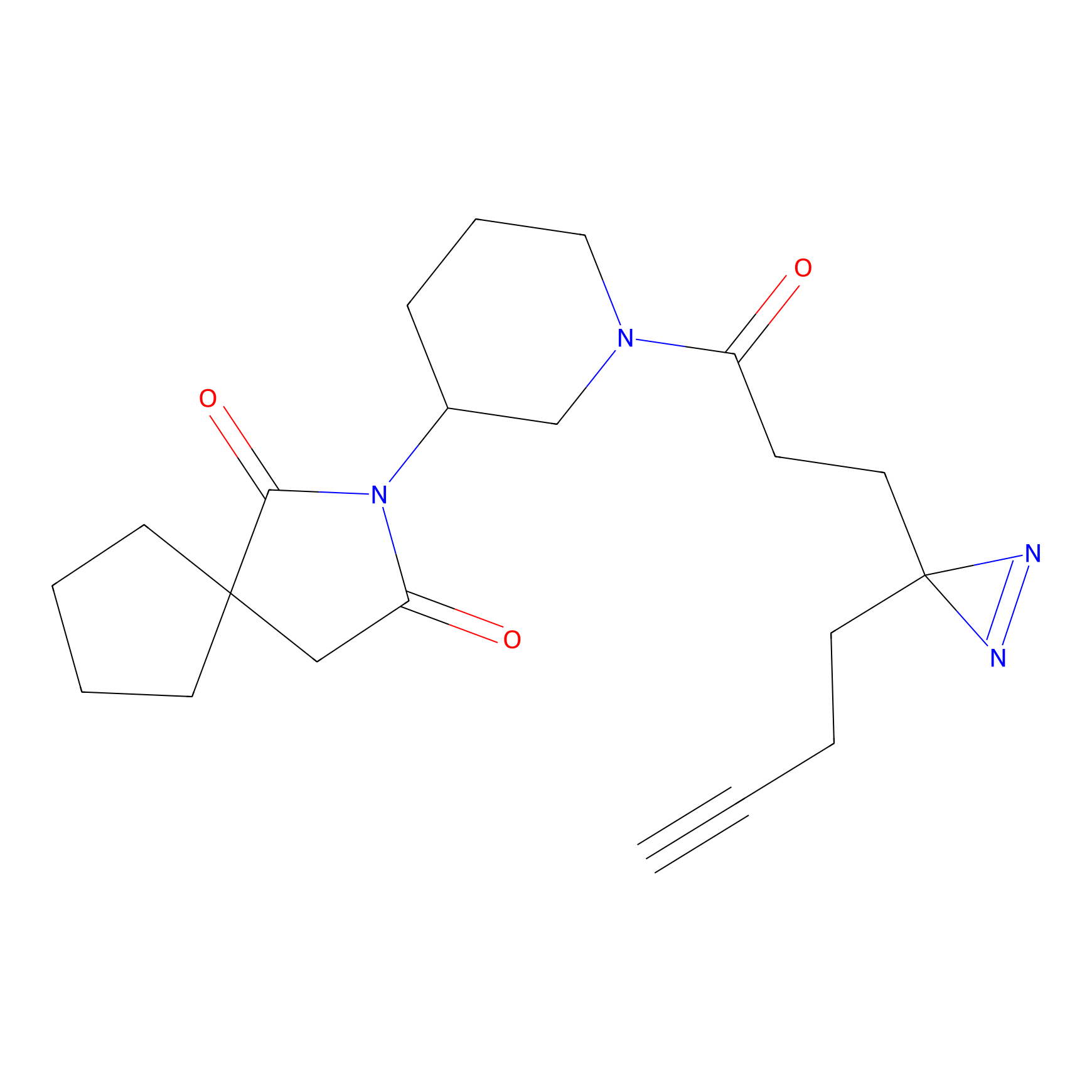
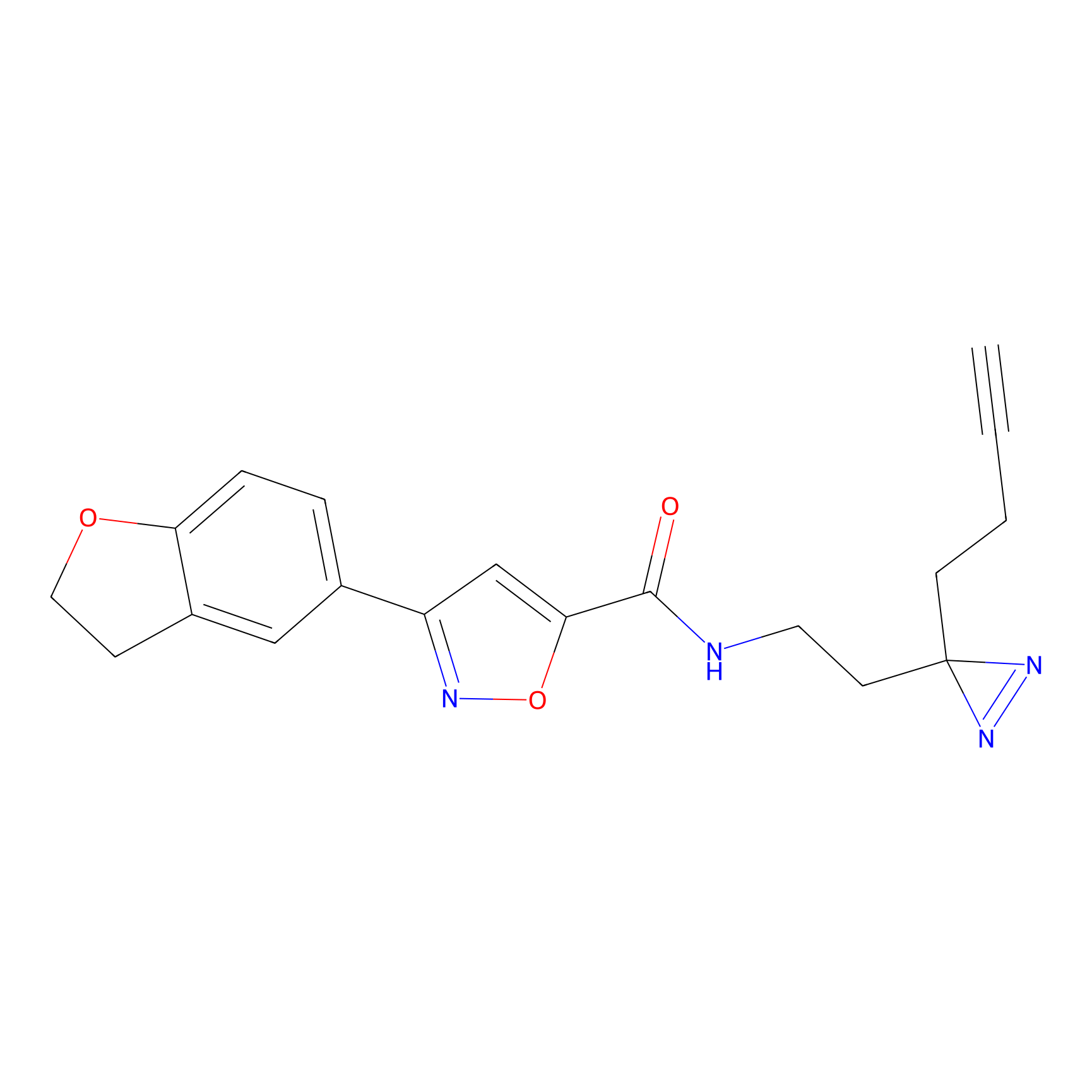
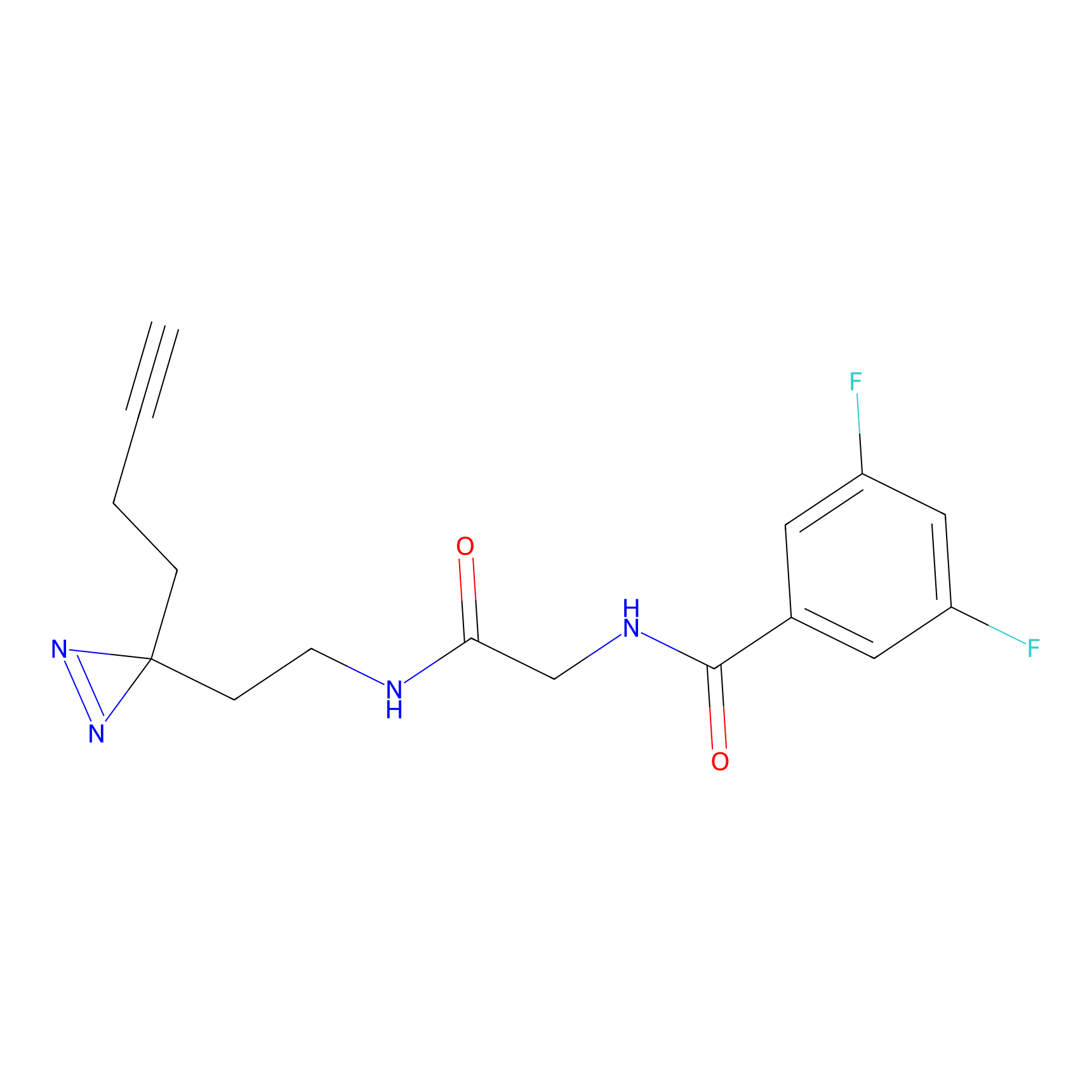
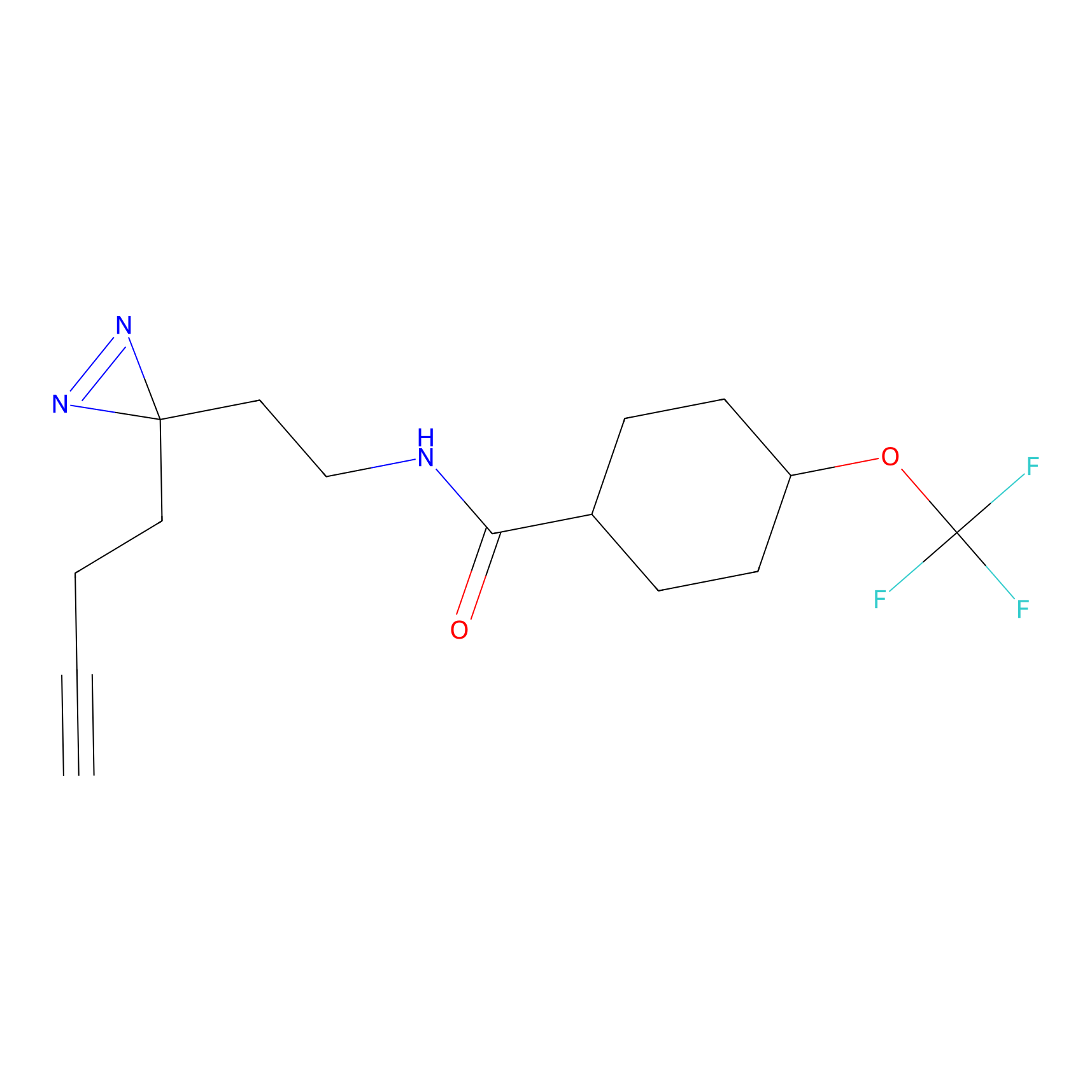
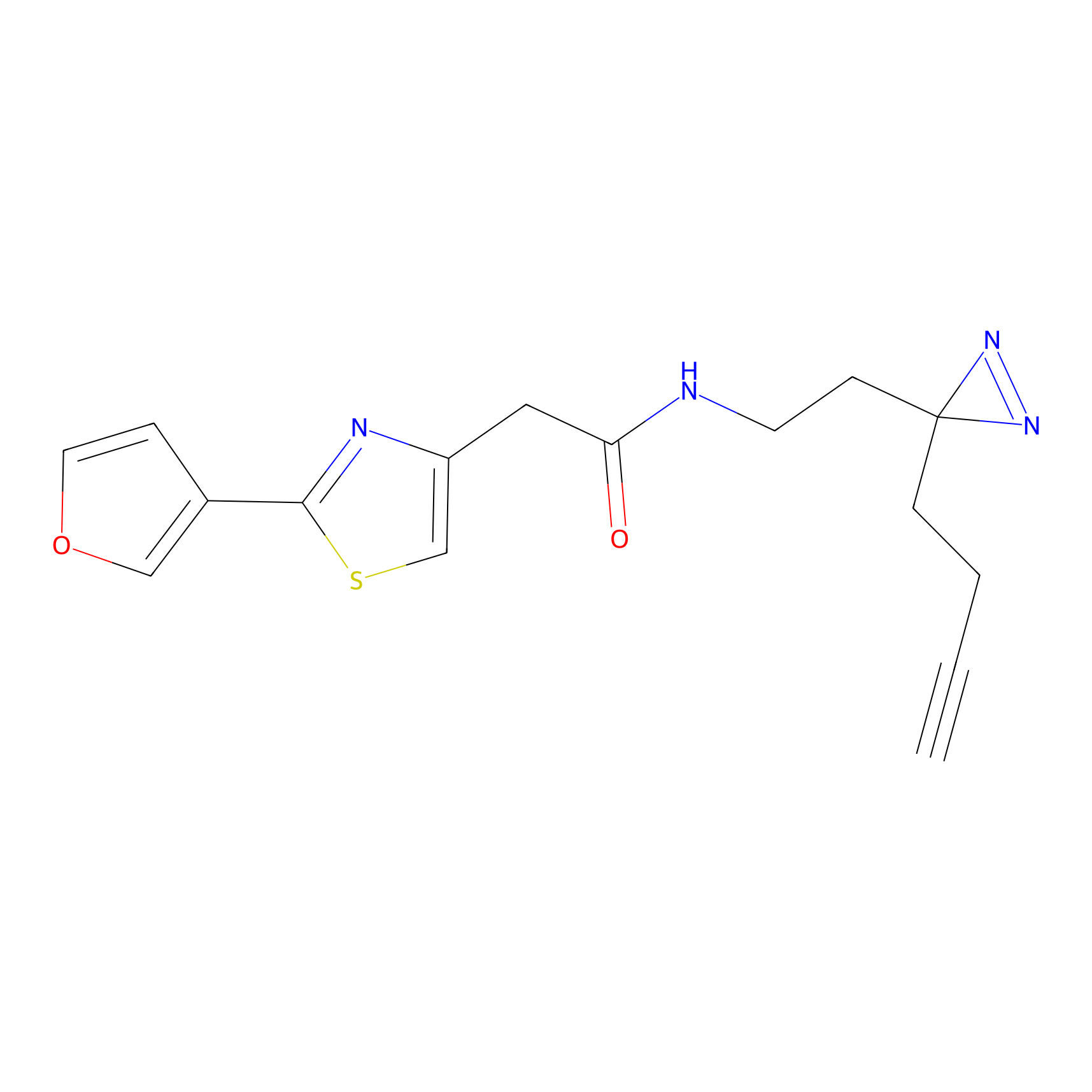
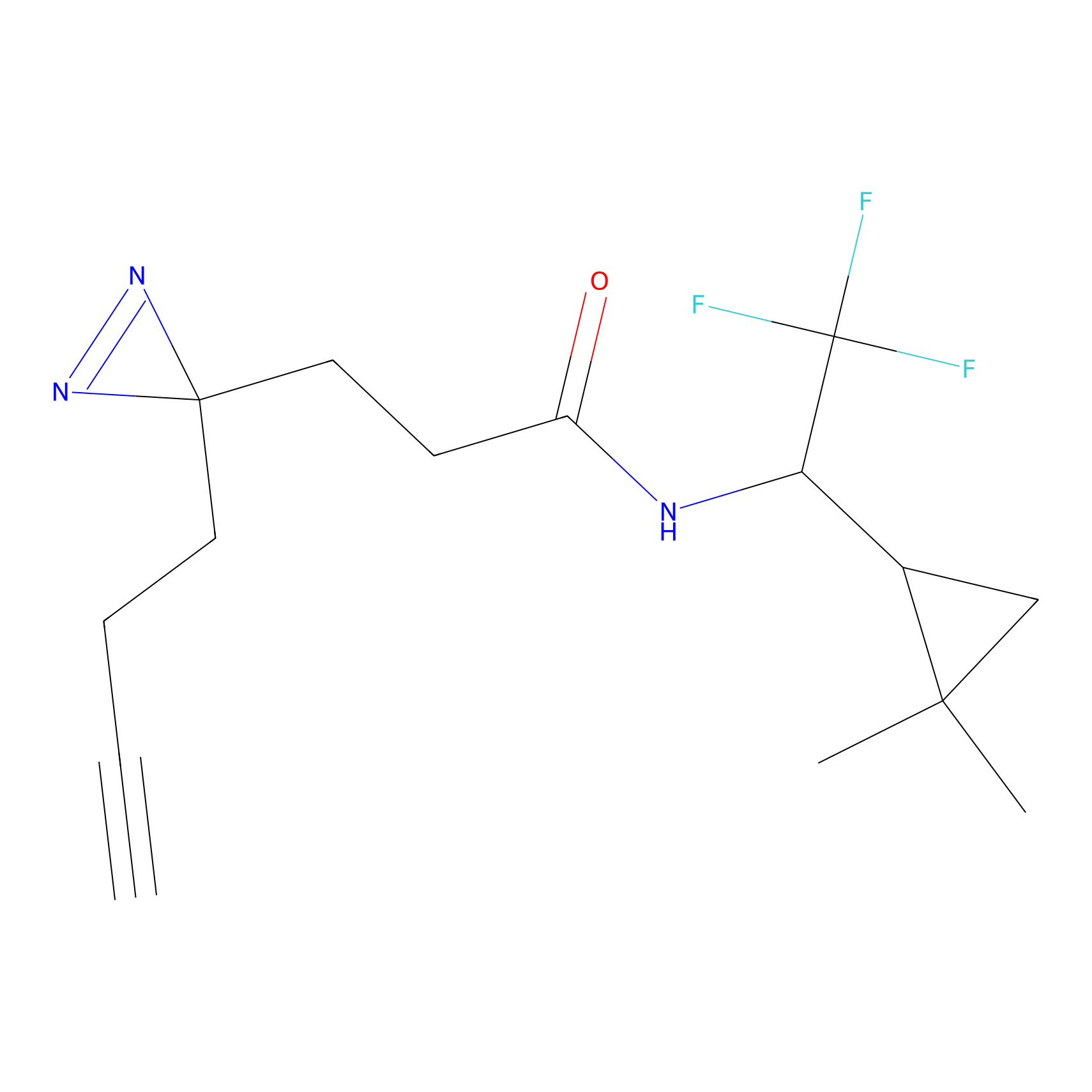
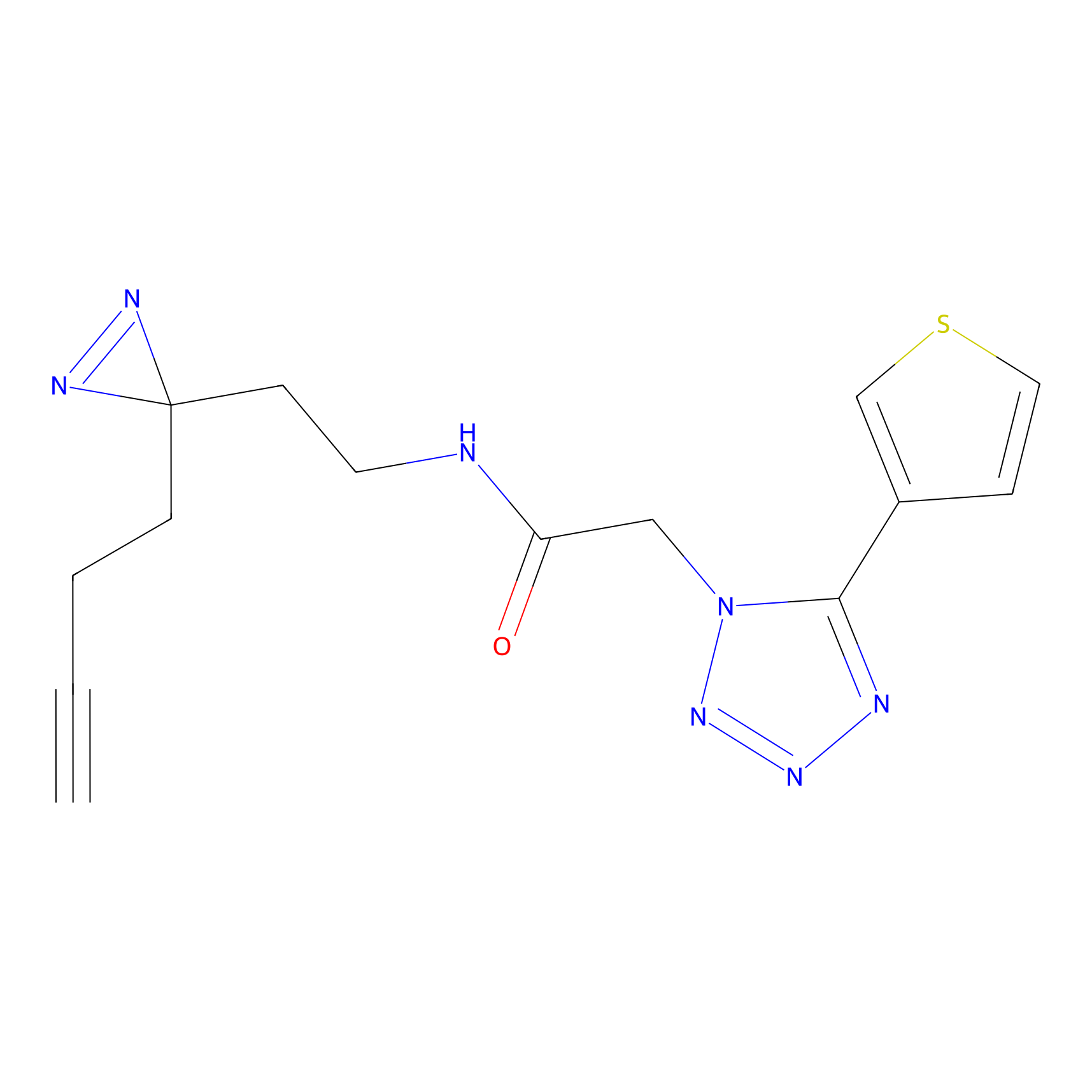
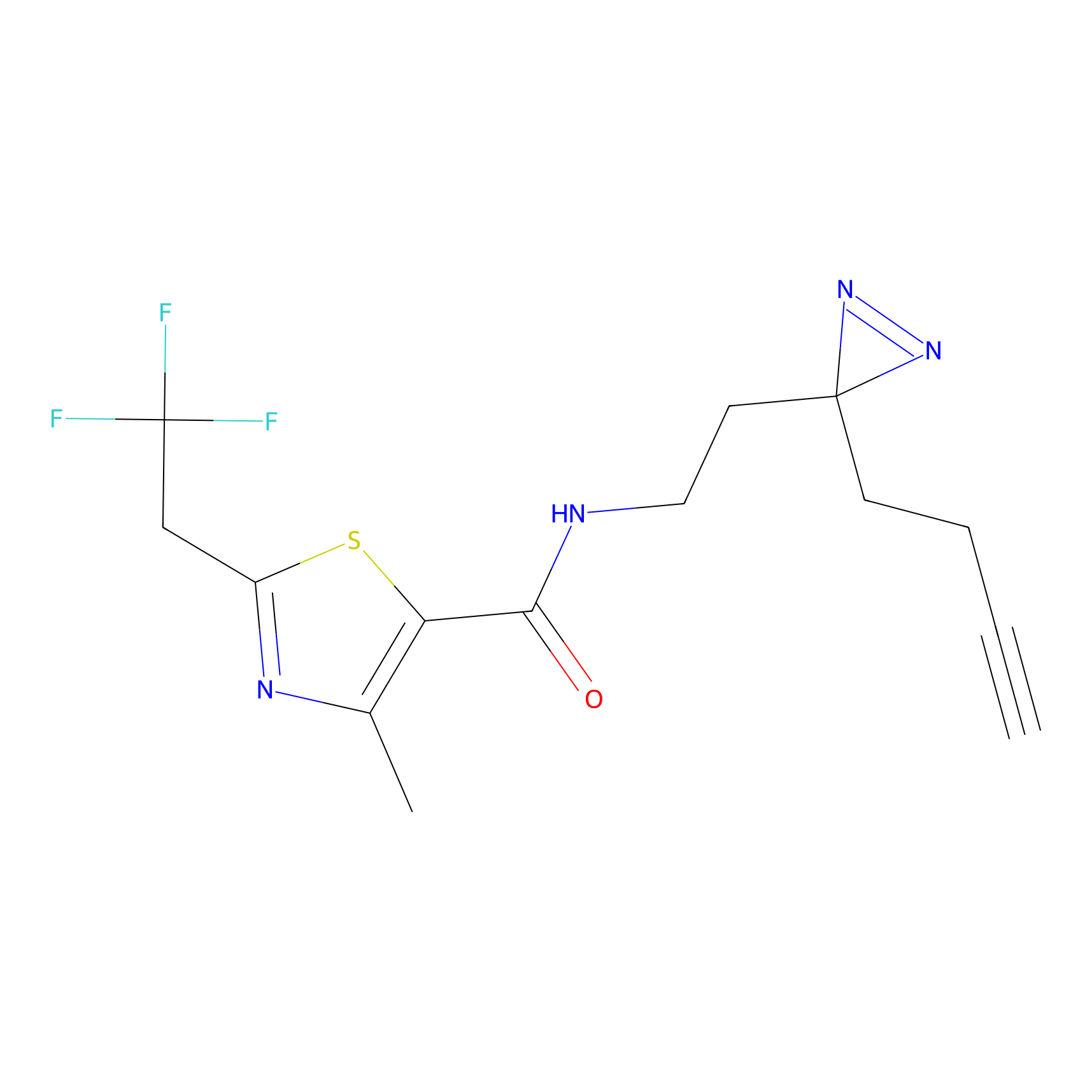
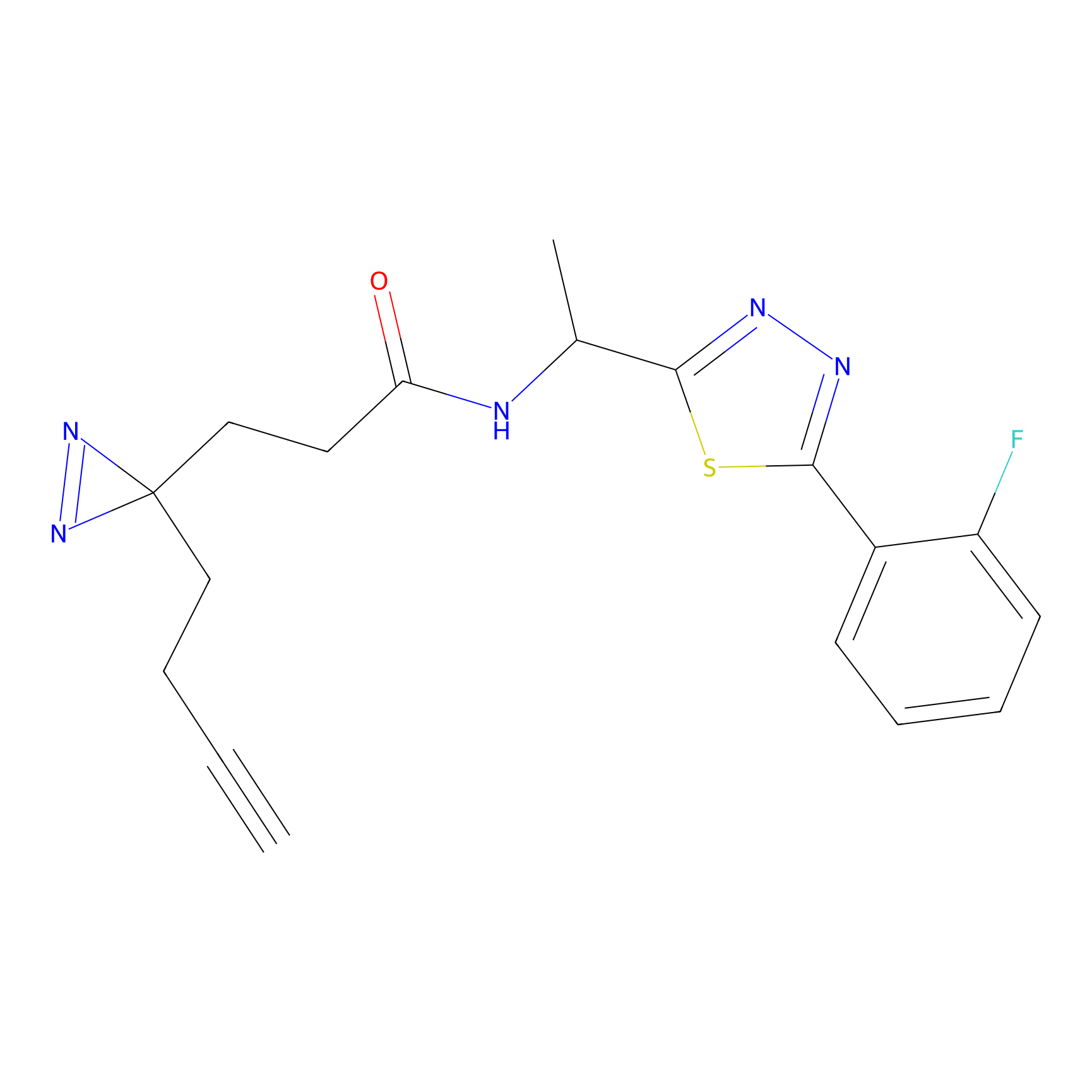
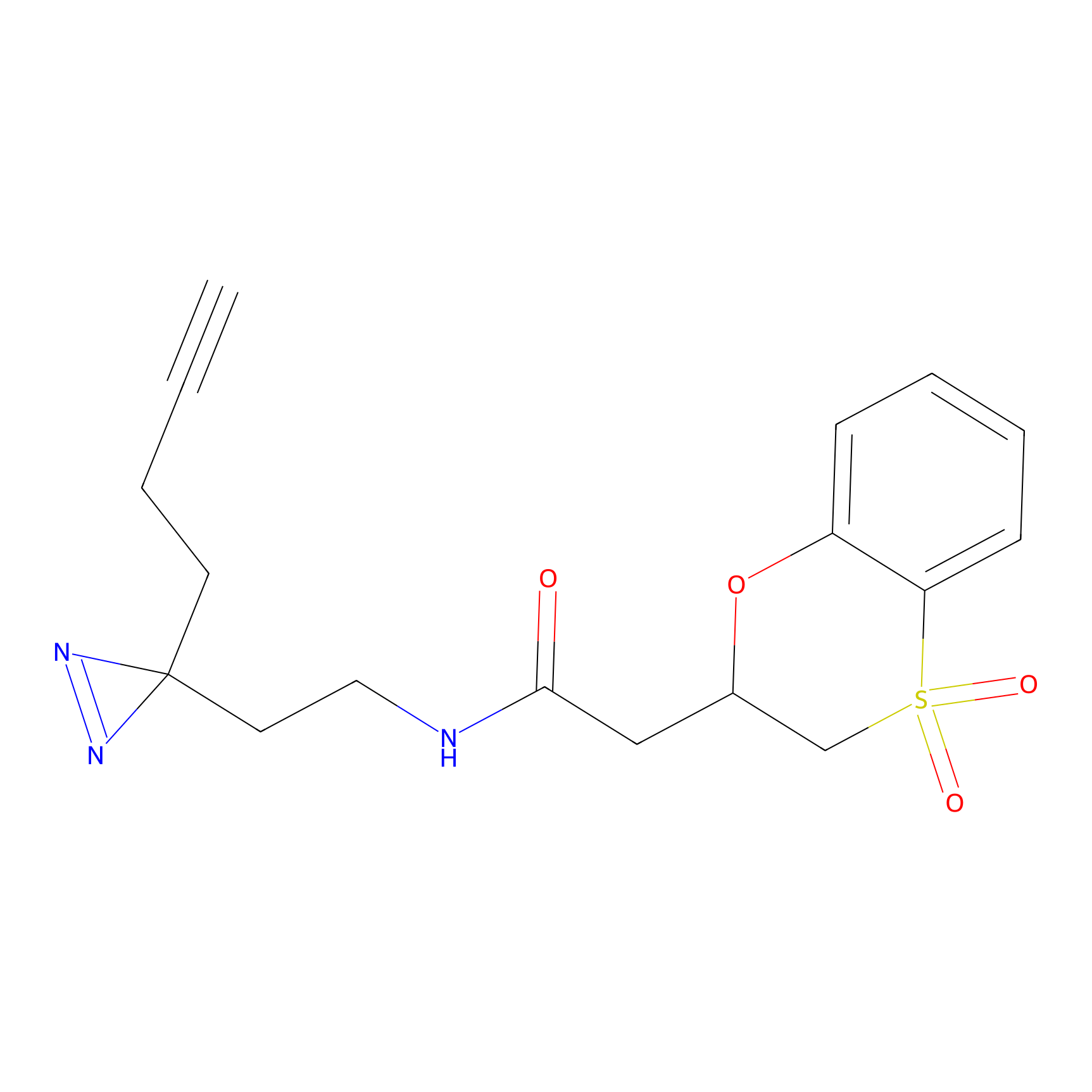
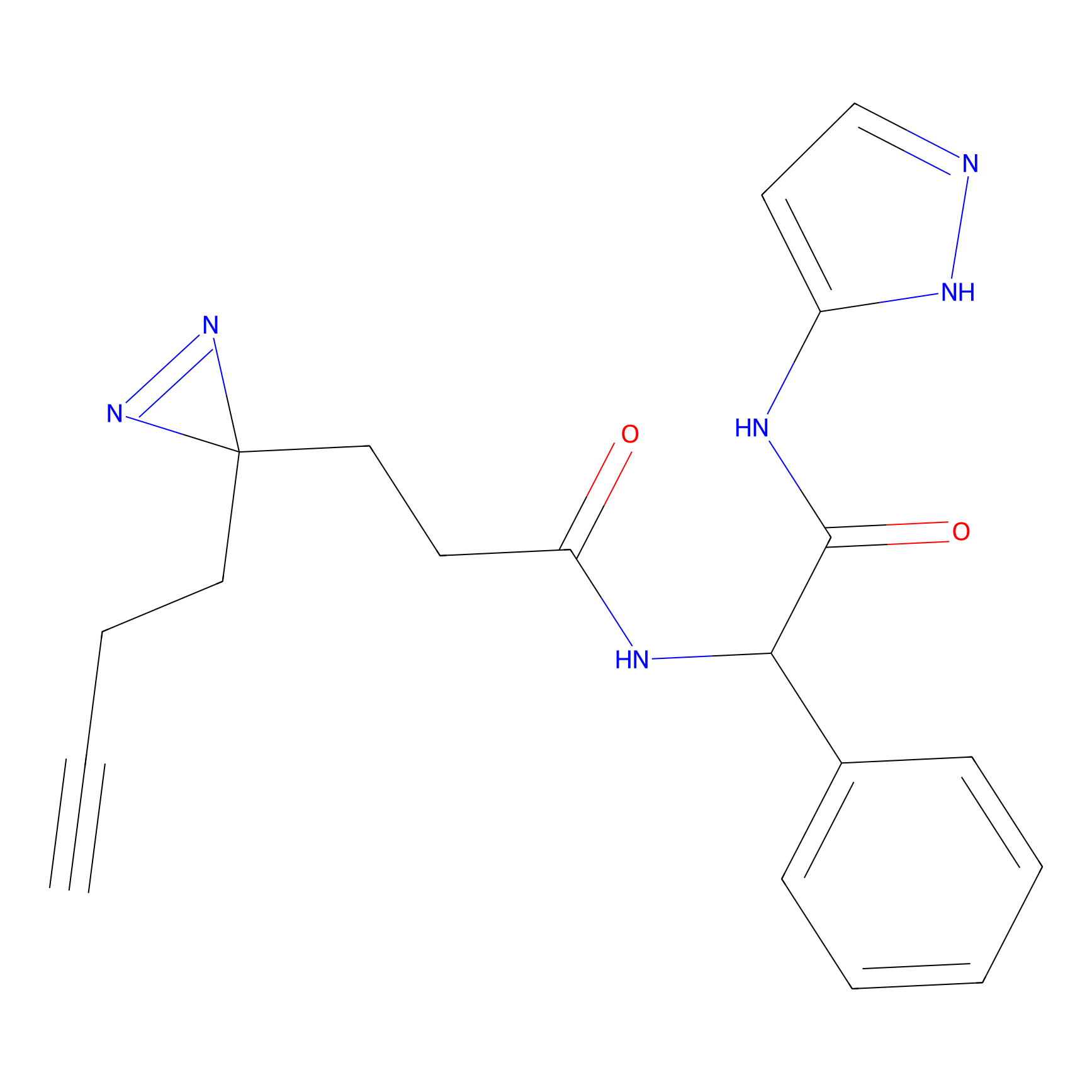
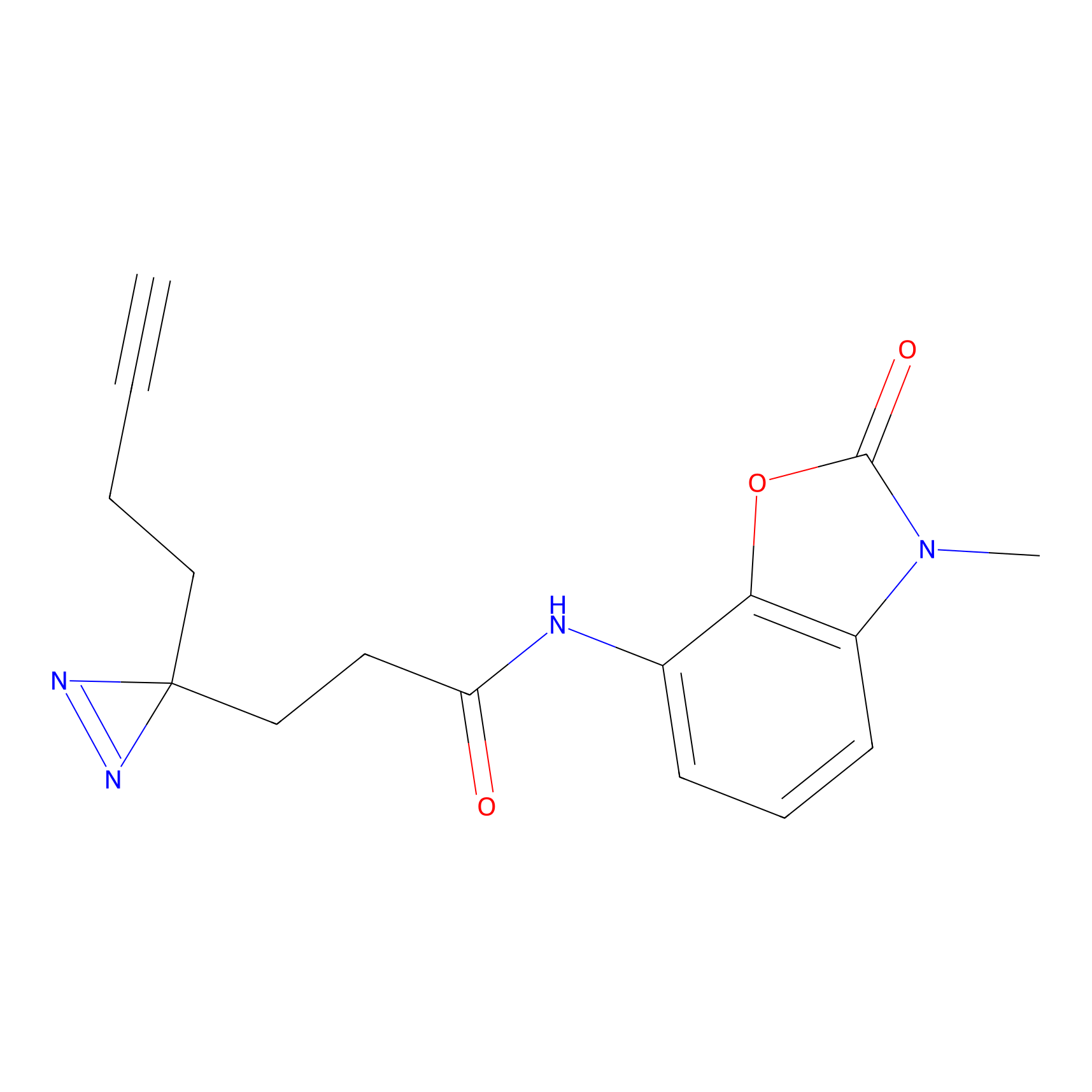
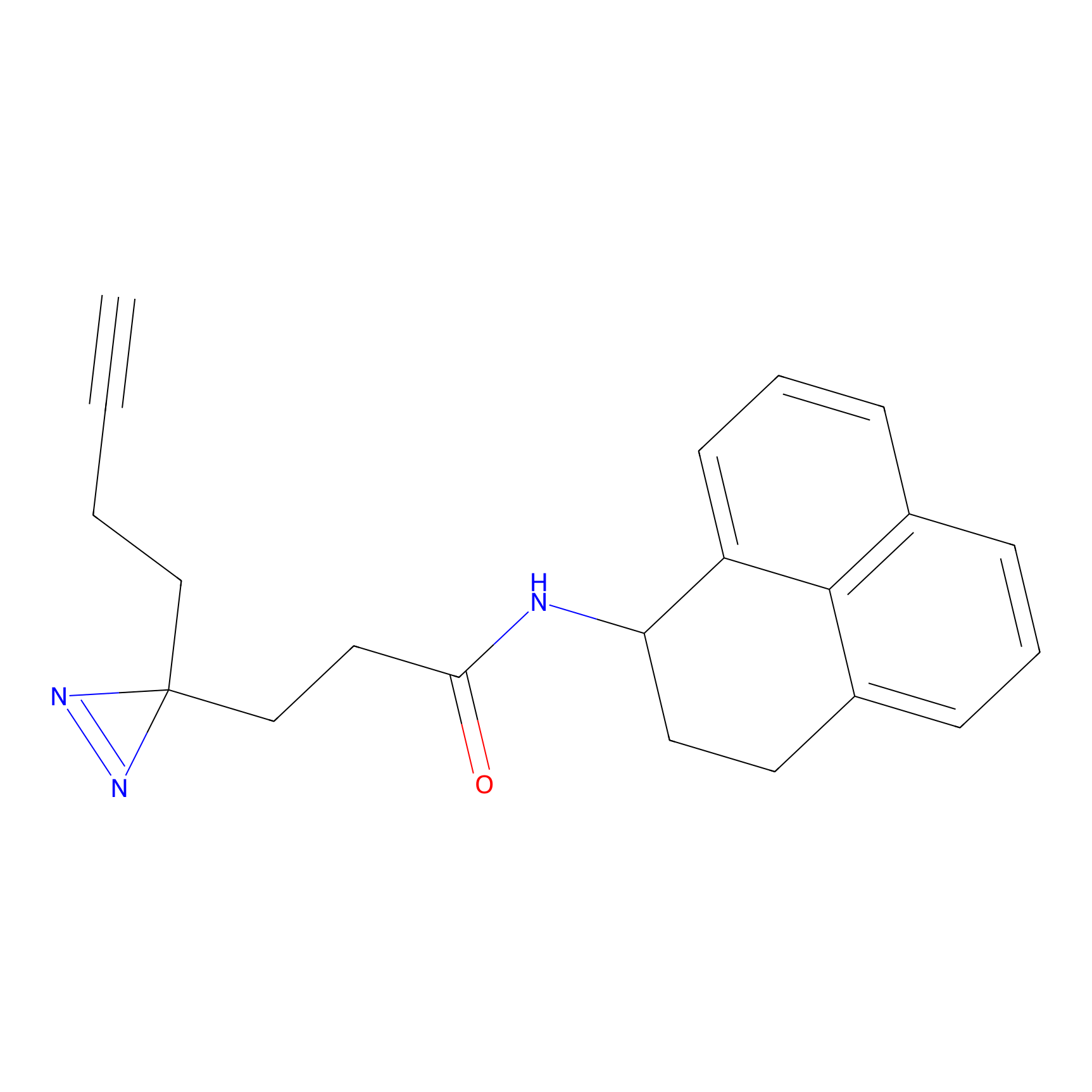
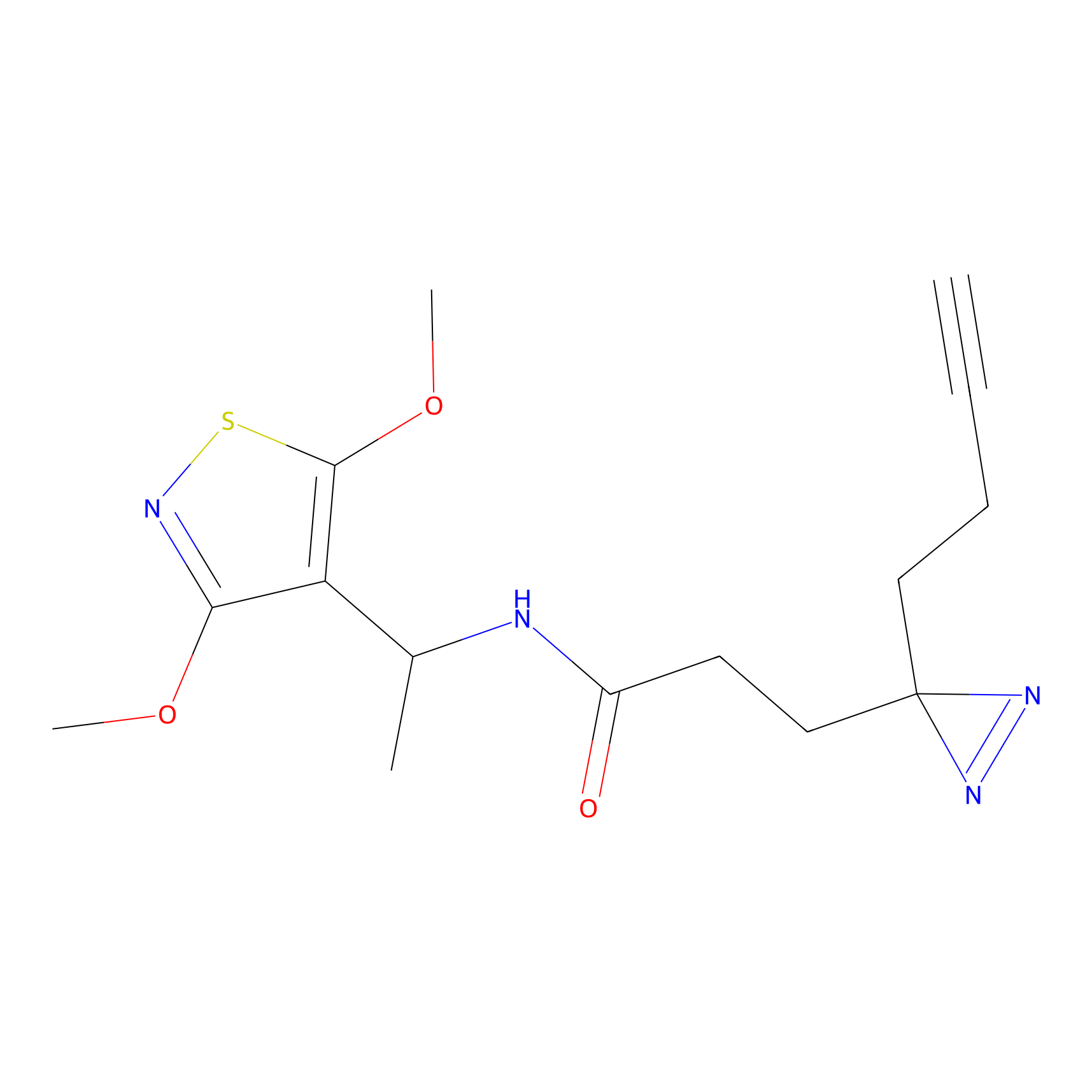
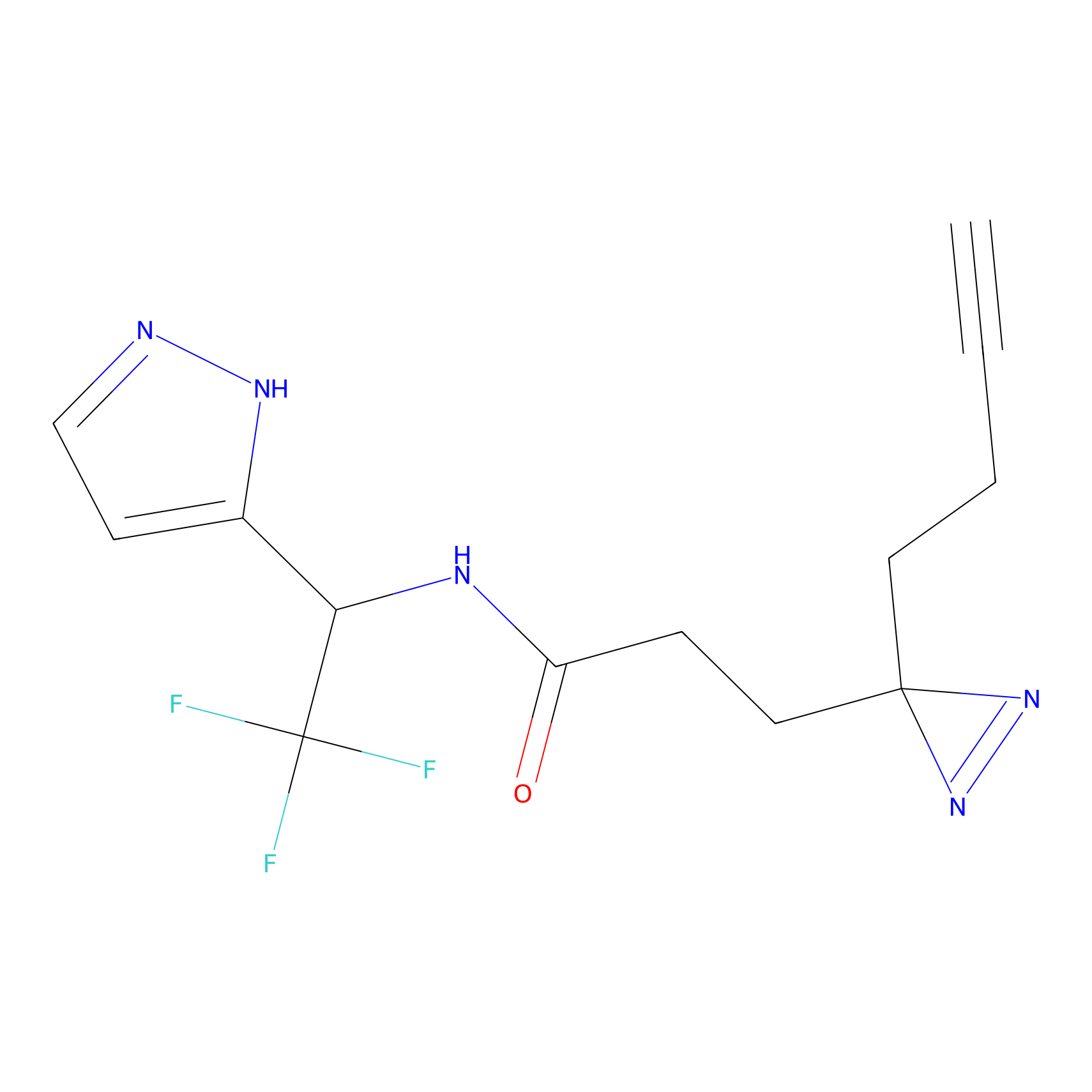
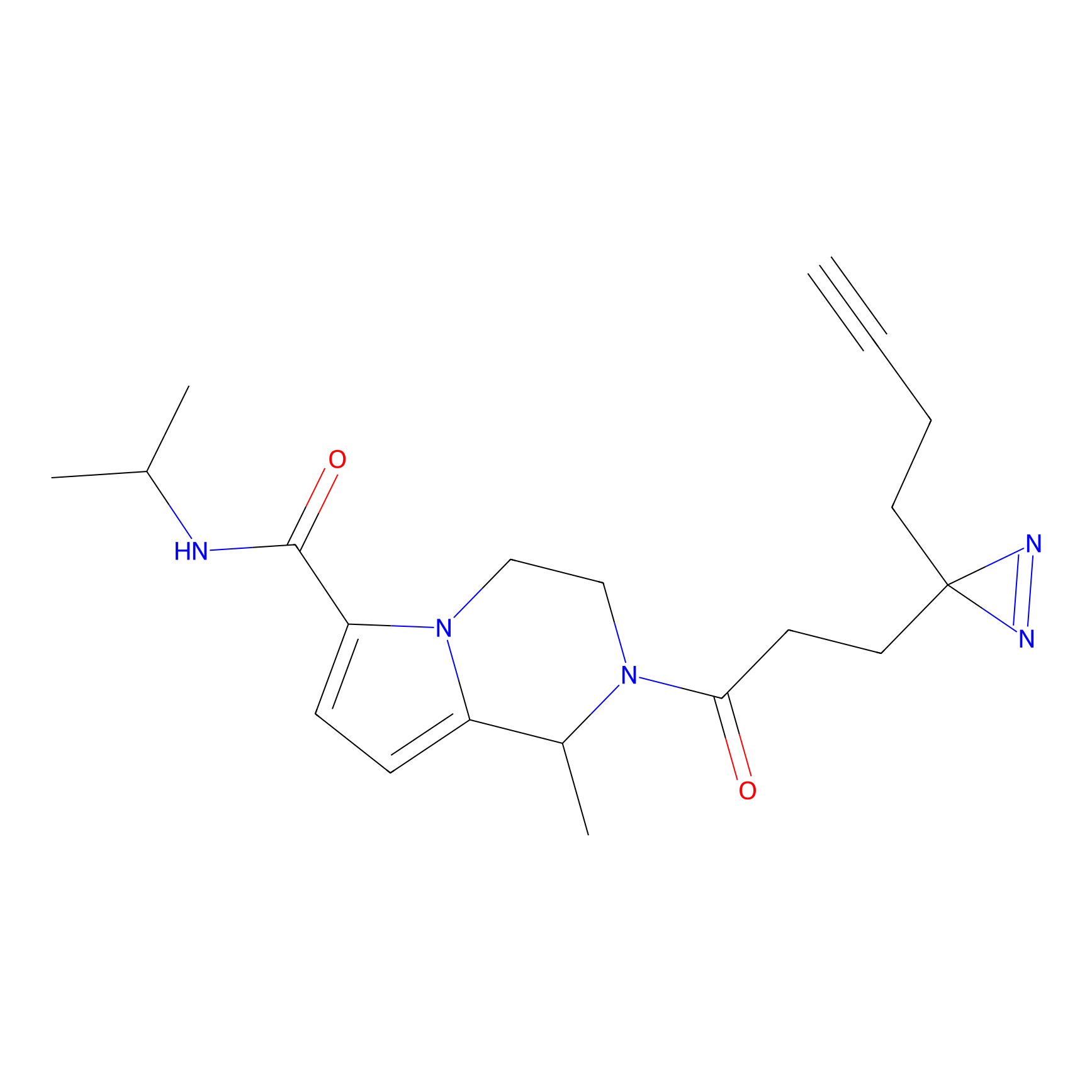
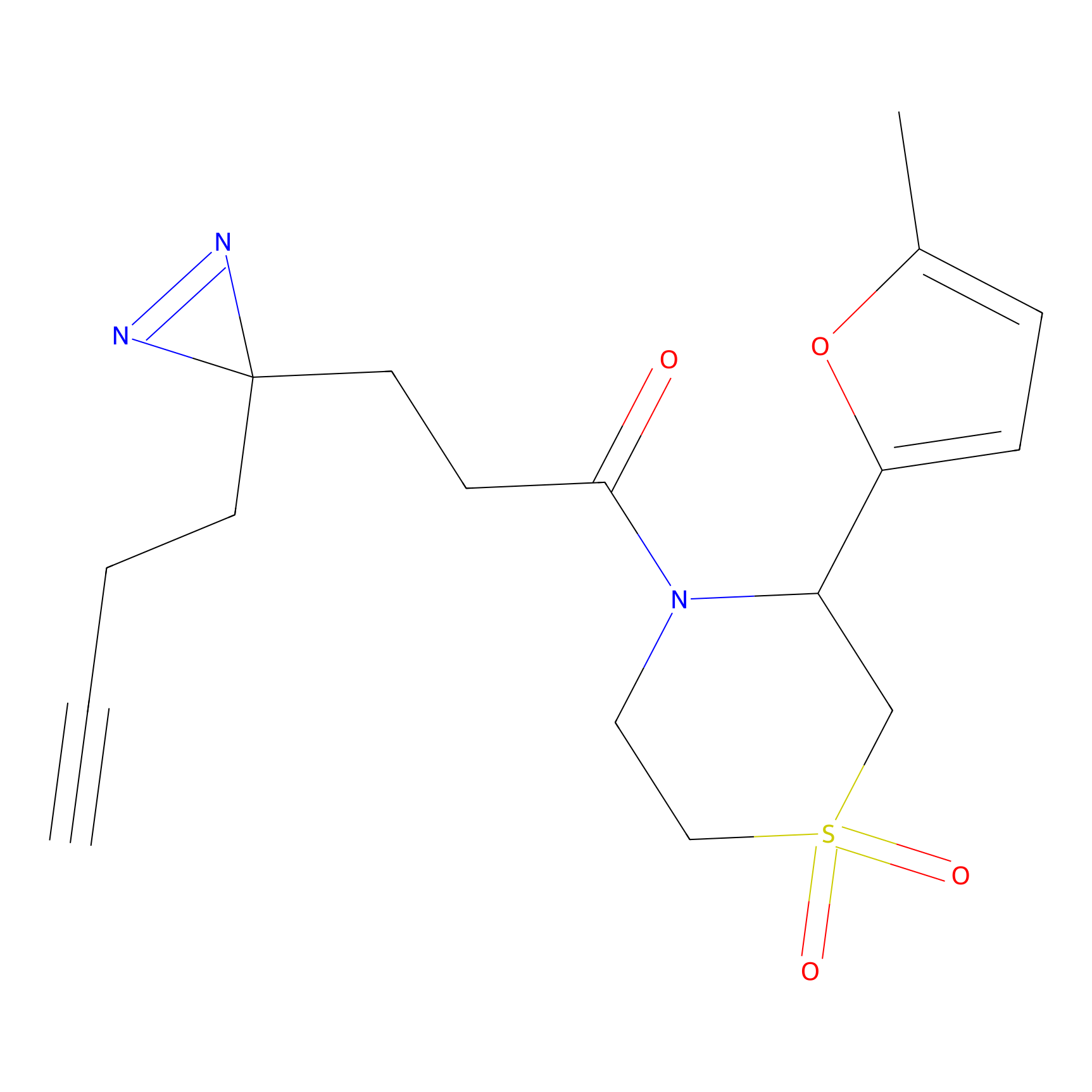
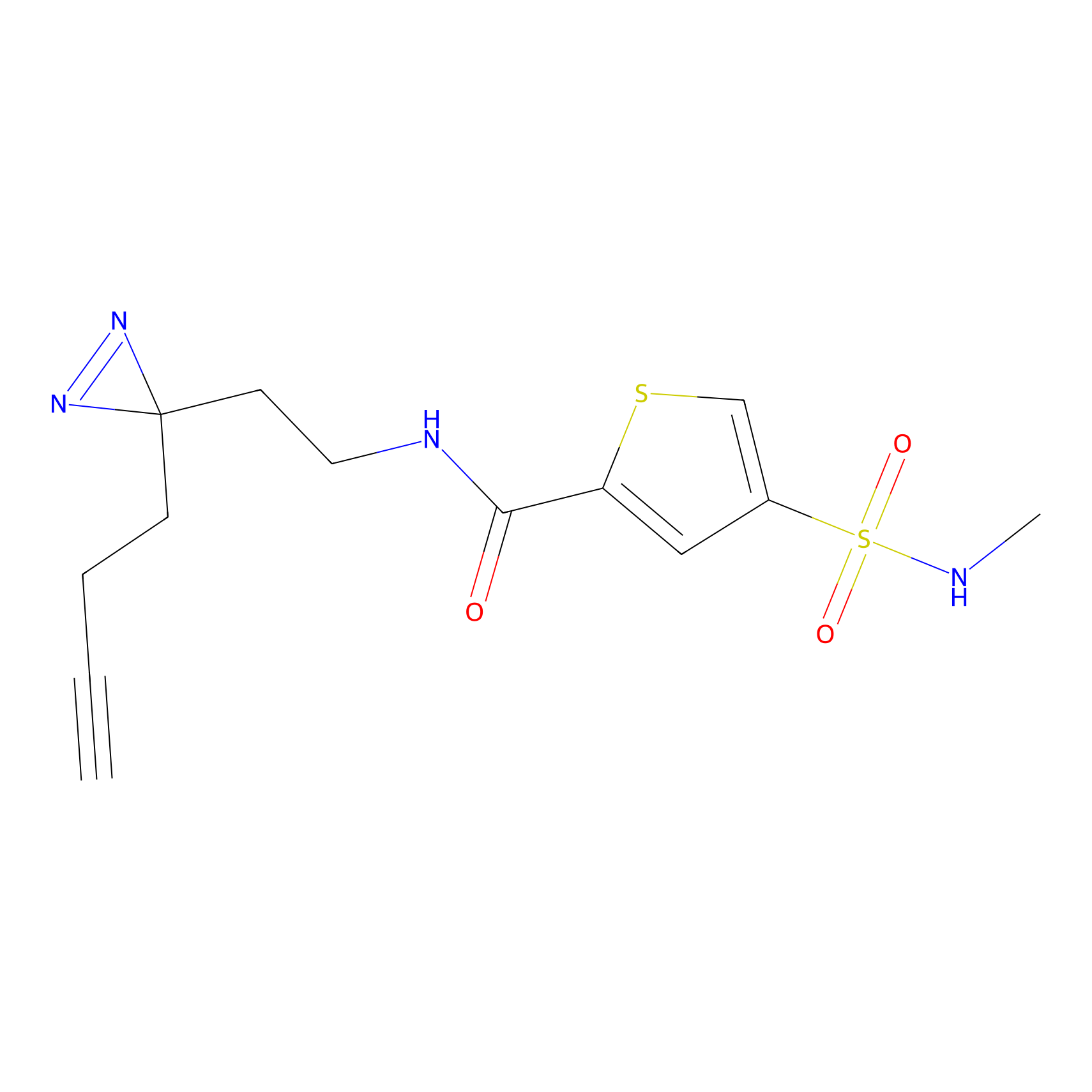
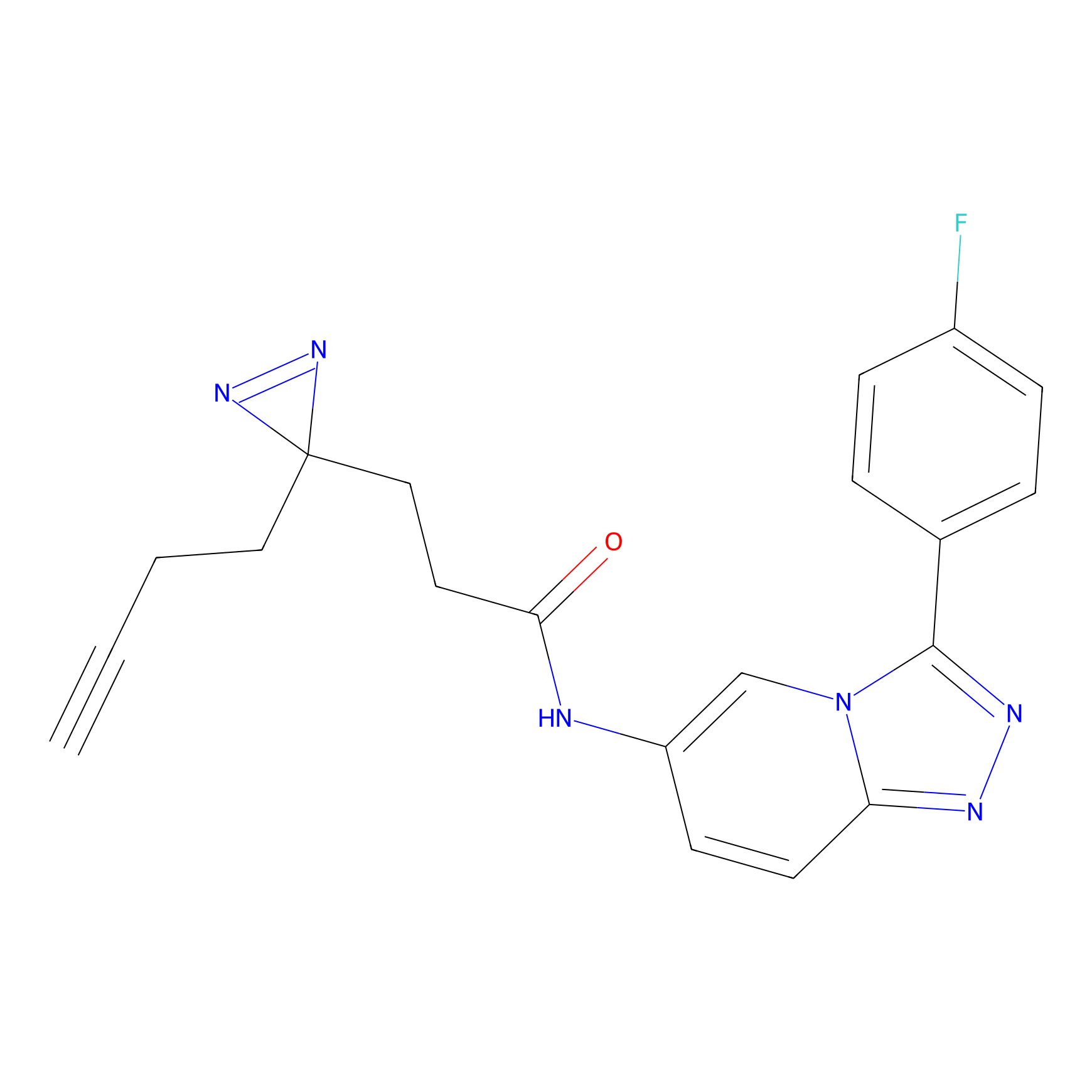
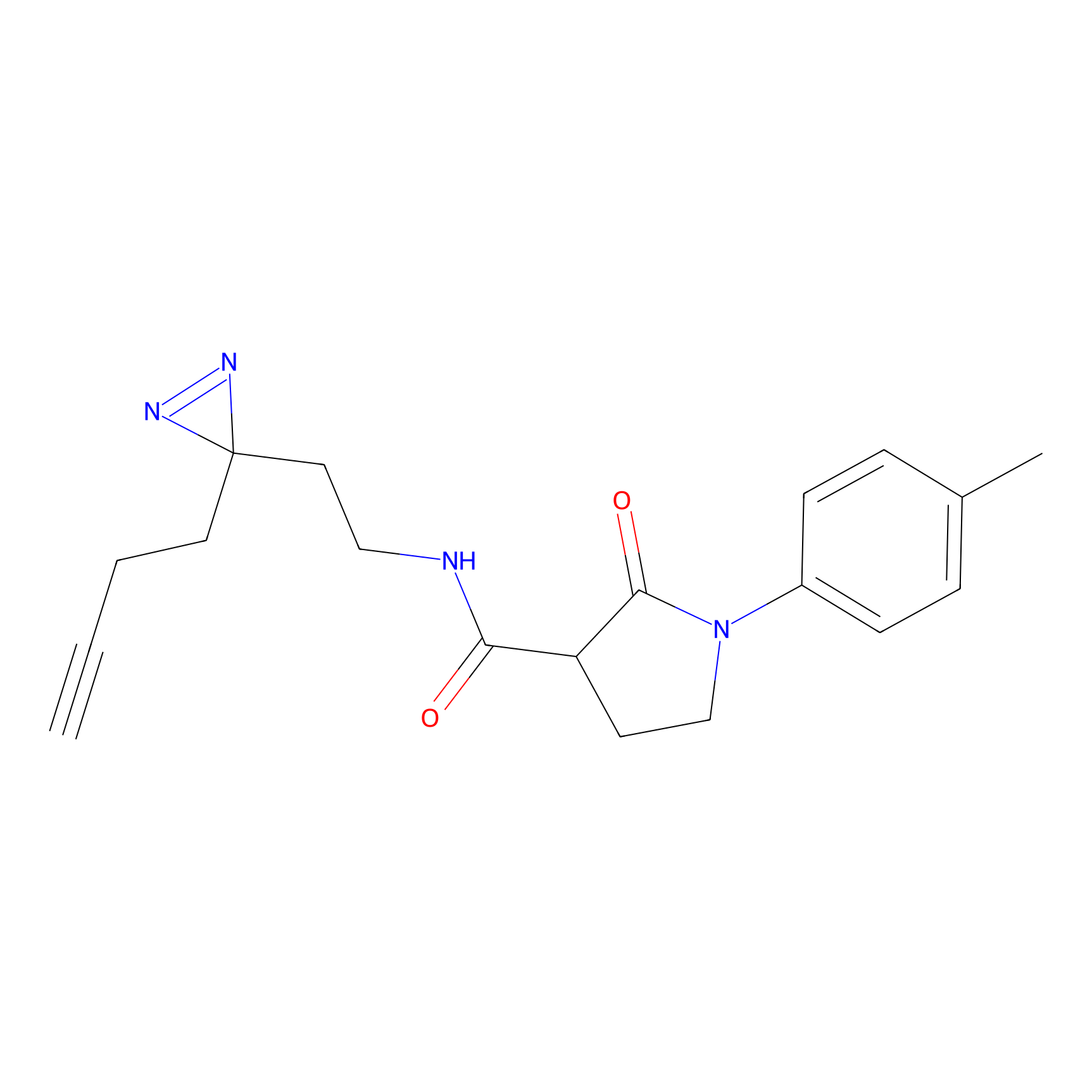
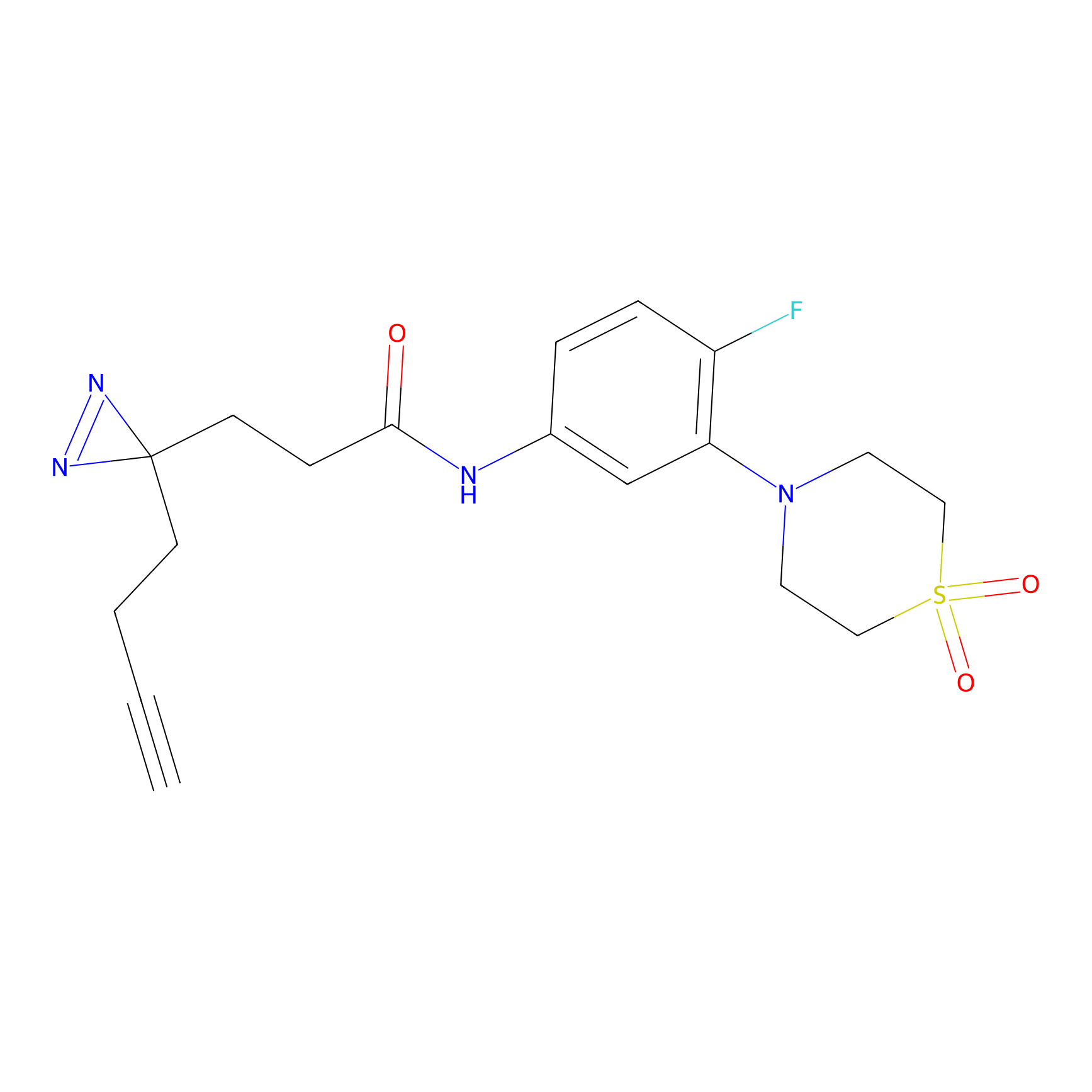
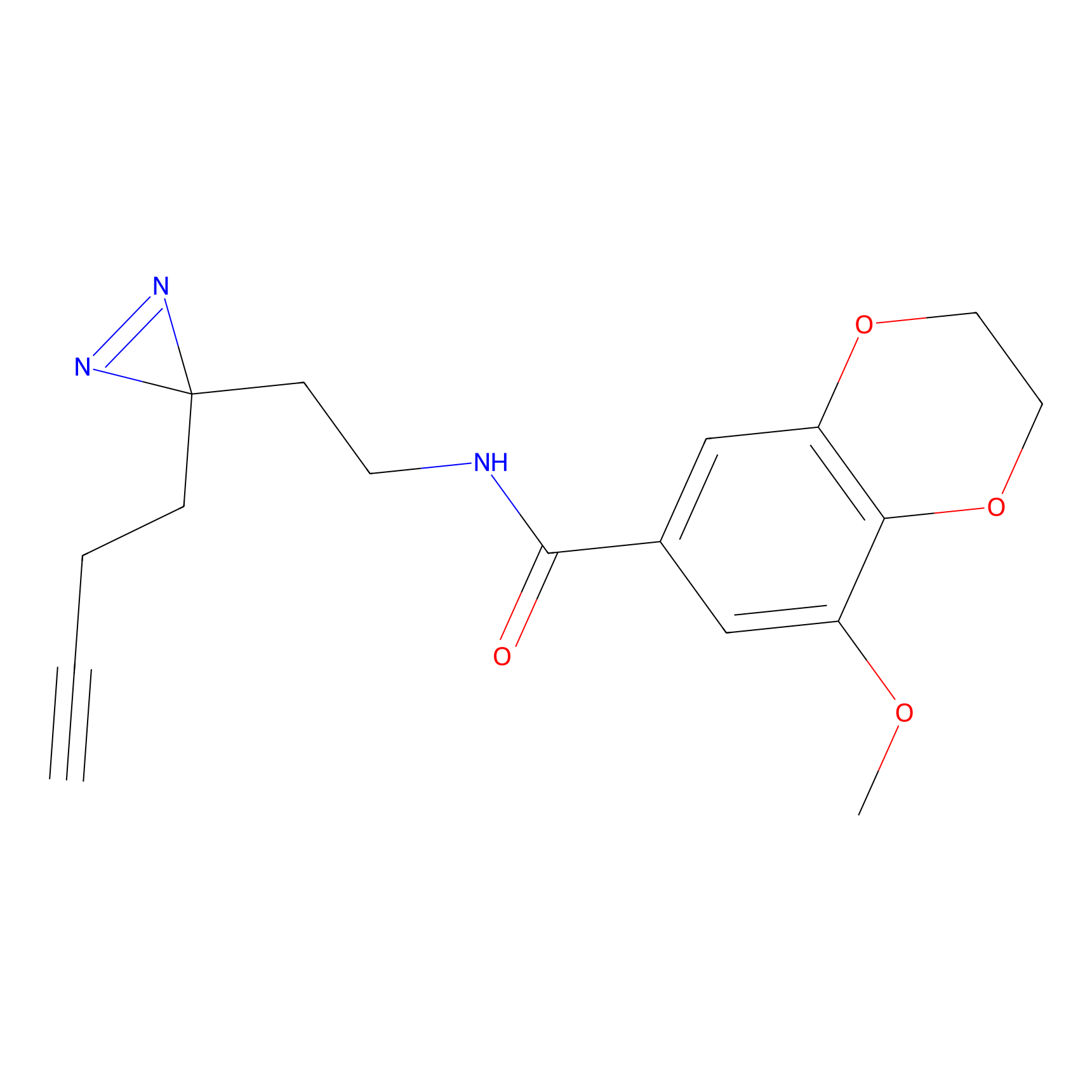
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
ILS-1 Probe Info |
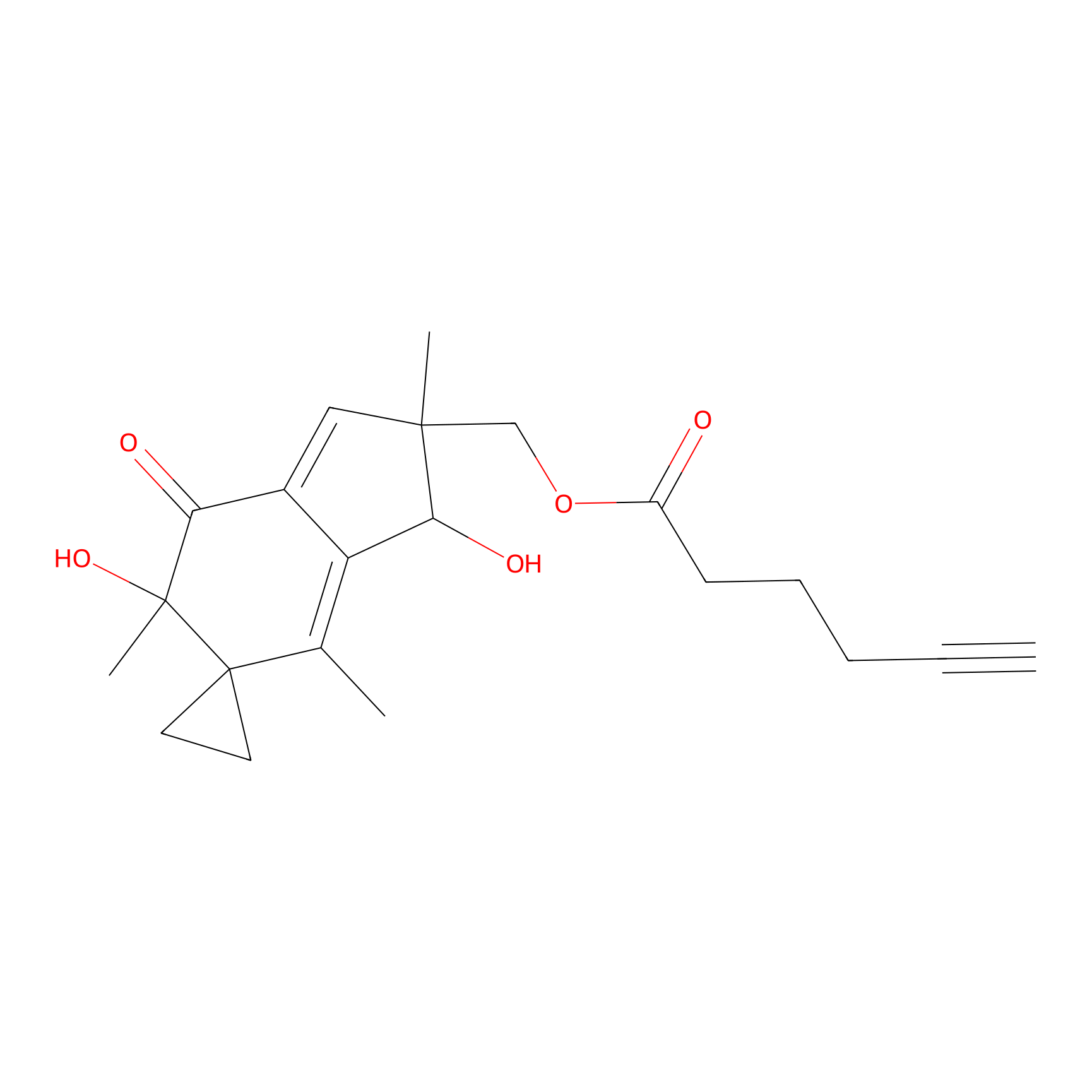 |
2.58 | LDD0415 | [1] | |
|
TH211 Probe Info |
 |
Y134(12.06) | LDD0257 | [2] | |
|
TH214 Probe Info |
 |
Y134(10.13) | LDD0258 | [2] | |
|
TH216 Probe Info |
 |
Y134(20.00) | LDD0259 | [2] | |
|
OPA-S-S-alkyne Probe Info |
 |
K243(1.07); K39(1.45) | LDD3494 | [3] | |
|
HPAP Probe Info |
 |
3.37 | LDD0062 | [4] | |
|
HHS-475 Probe Info |
 |
Y134(1.08) | LDD0264 | [5] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [6] | |
|
OSF Probe Info |
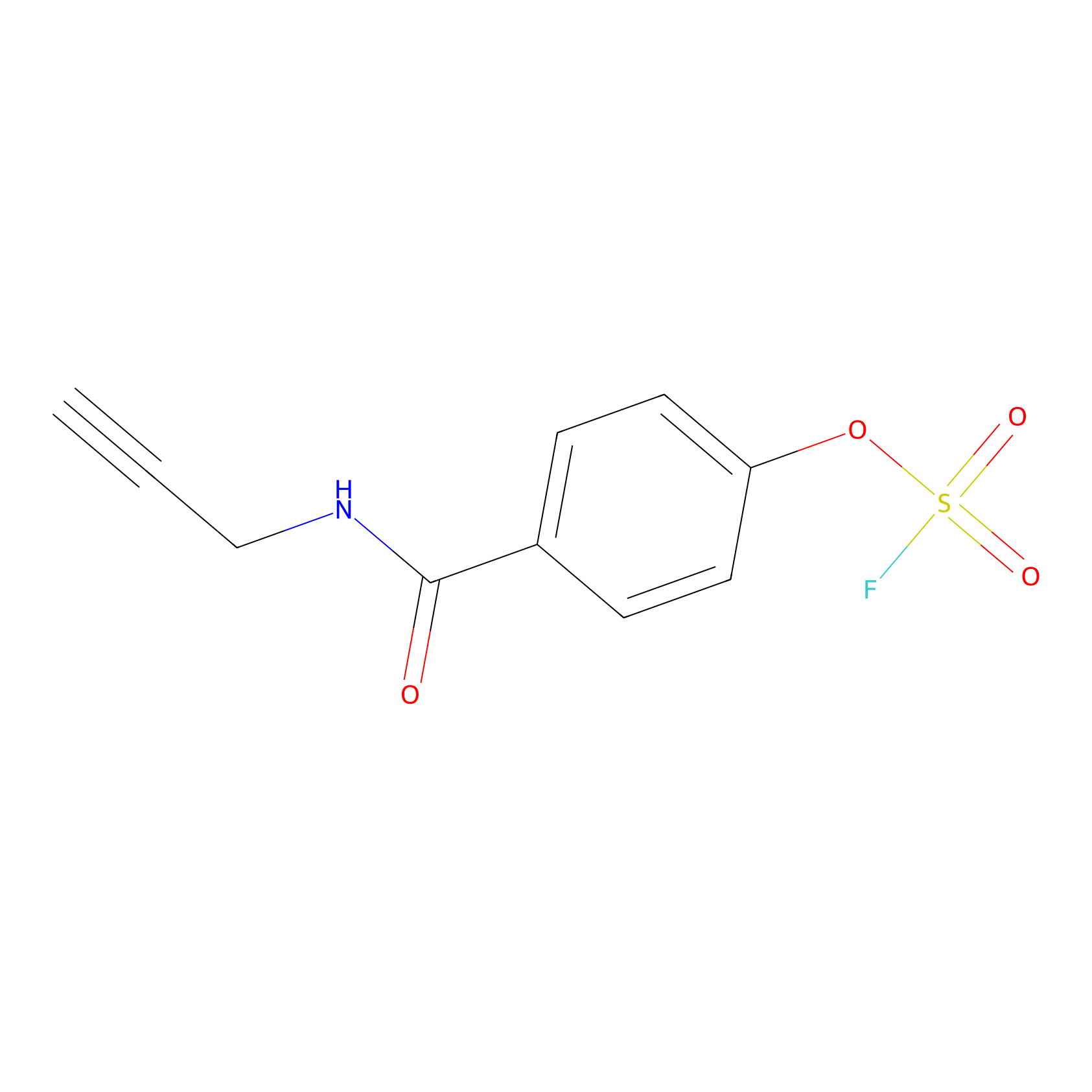 |
N.A. | LDD0029 | [7] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [7] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [8] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [9] | |
|
Acrolein Probe Info |
 |
H84(0.00); H25(0.00) | LDD0217 | [10] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [11] | |
|
HHS-482 Probe Info |
 |
Y134(1.74) | LDD2239 | [12] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [10] |
| LDCM0116 | HHS-0101 | DM93 | Y134(1.08) | LDD0264 | [5] |
| LDCM0117 | HHS-0201 | DM93 | Y134(1.79) | LDD0265 | [5] |
| LDCM0118 | HHS-0301 | DM93 | Y134(1.18) | LDD0266 | [5] |
| LDCM0119 | HHS-0401 | DM93 | Y134(1.30) | LDD0267 | [5] |
| LDCM0120 | HHS-0701 | DM93 | Y134(1.13) | LDD0268 | [5] |
| LDCM0107 | IAA | HeLa | H84(0.00); H25(0.00) | LDD0221 | [10] |
| LDCM0109 | NEM | HeLa | H25(0.00); H84(0.00); H119(0.00) | LDD0223 | [10] |
| LDCM0014 | Panhematin | HEK-293T | 3.37 | LDD0062 | [4] |
| LDCM0081 | Rofecoxib | A-549 | 2.51 | LDD0141 | [15] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
GPCR
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Probable G-protein coupled receptor 152 (GPR152) | G-protein coupled receptor 1 family | Q8TDT2 | |||
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| B-cell antigen receptor complex-associated protein alpha chain (CD79A) | . | P11912 | |||
Other
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Nadh | Small molecular drug | DB00157 | |||
| Vitamin E | Small molecular drug | DB00163 | |||
| Alpha-tocopherol Succinate | . | DB14001 | |||
| D-alpha-tocopherol Acetate | . | DB14002 | |||
Phase 2
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Stannsoporfin | Small molecular drug | D02JAY | |||
Investigative
References