Details of the Target
General Information of Target
| Target ID | LDTP11076 | |||||
|---|---|---|---|---|---|---|
| Target Name | Apolipoprotein L2 (APOL2) | |||||
| Gene Name | APOL2 | |||||
| Gene ID | 23780 | |||||
| Synonyms |
Apolipoprotein L2; Apolipoprotein L-II; ApoL-II |
|||||
| 3D Structure | ||||||
| Sequence |
MERQEESLSARPALETEGLRFLHTTVGSLLATYGWYIVFSCILLYVVFQKLSARLRALRQ
RQLDRAAAAVEPDVVVKRQEALAAARLKMQEELNAQVEKHKEKLKQLEEEKRRQKIEMWD SMQEGKSYKGNAKKPQEEDSPGPSTSSVLKRKSDRKPLRGGGYNPLSGEGGGACSWRPGR RGPSSGGUG |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Apolipoprotein L family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function | May affect the movement of lipids in the cytoplasm or allow the binding of lipids to organelles. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
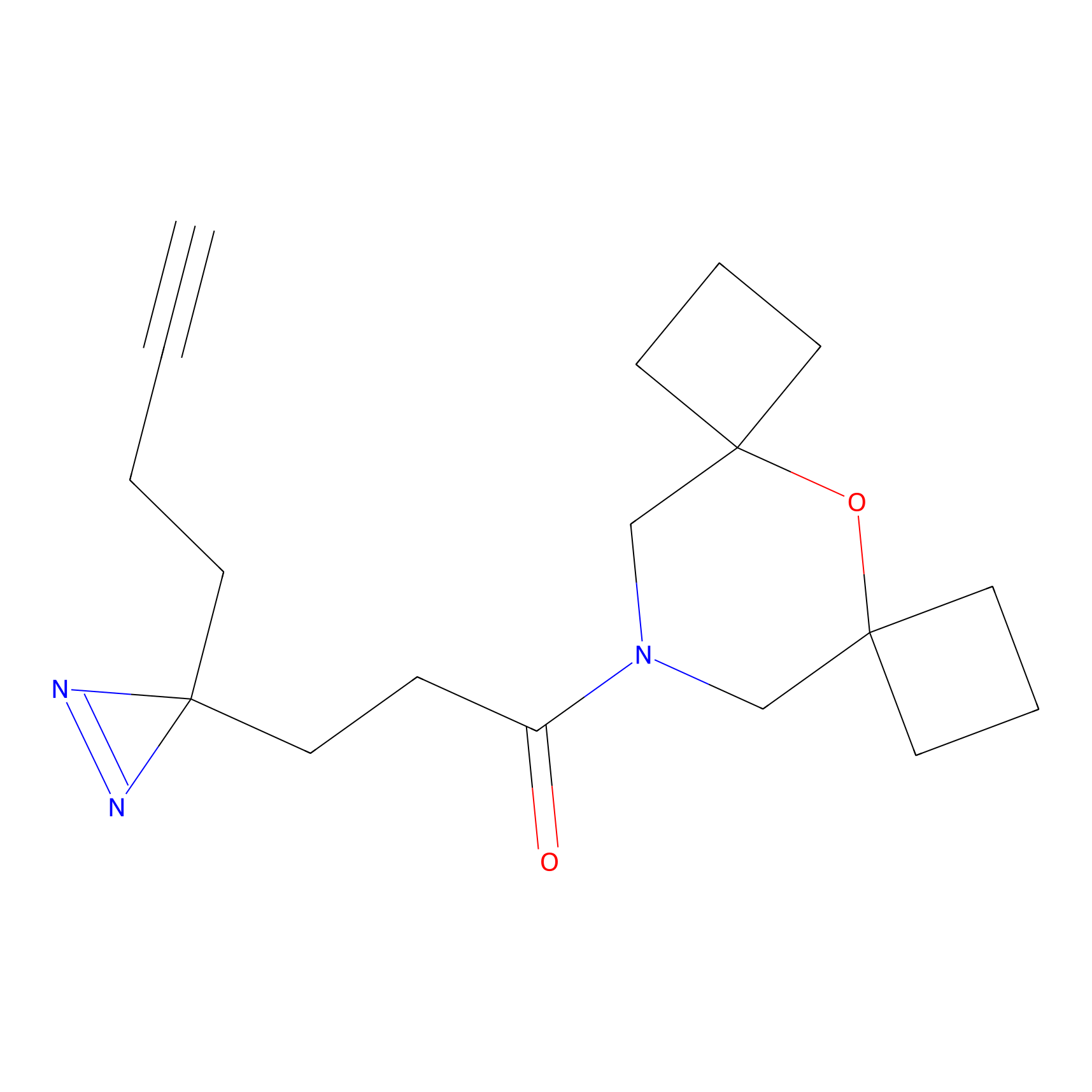
|
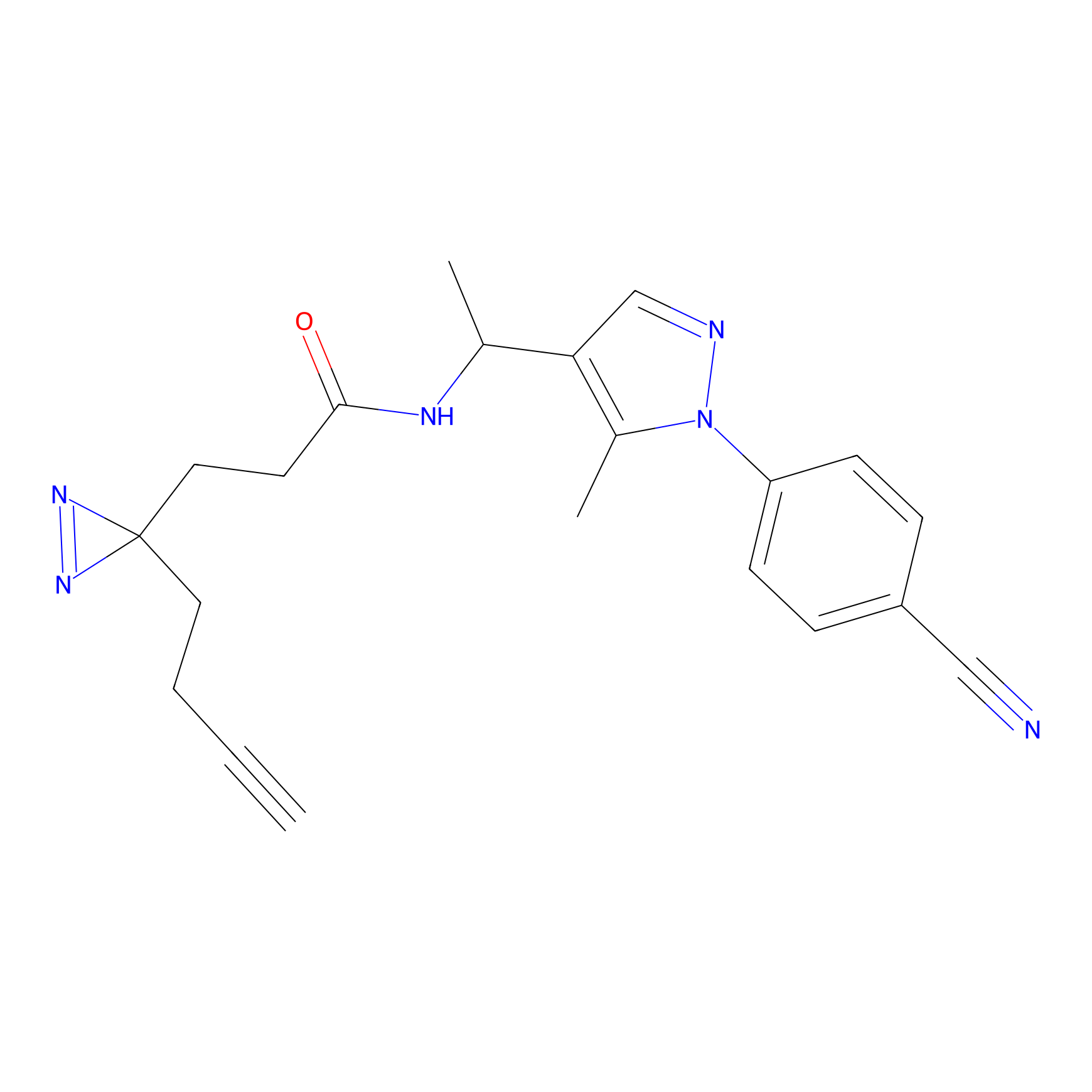
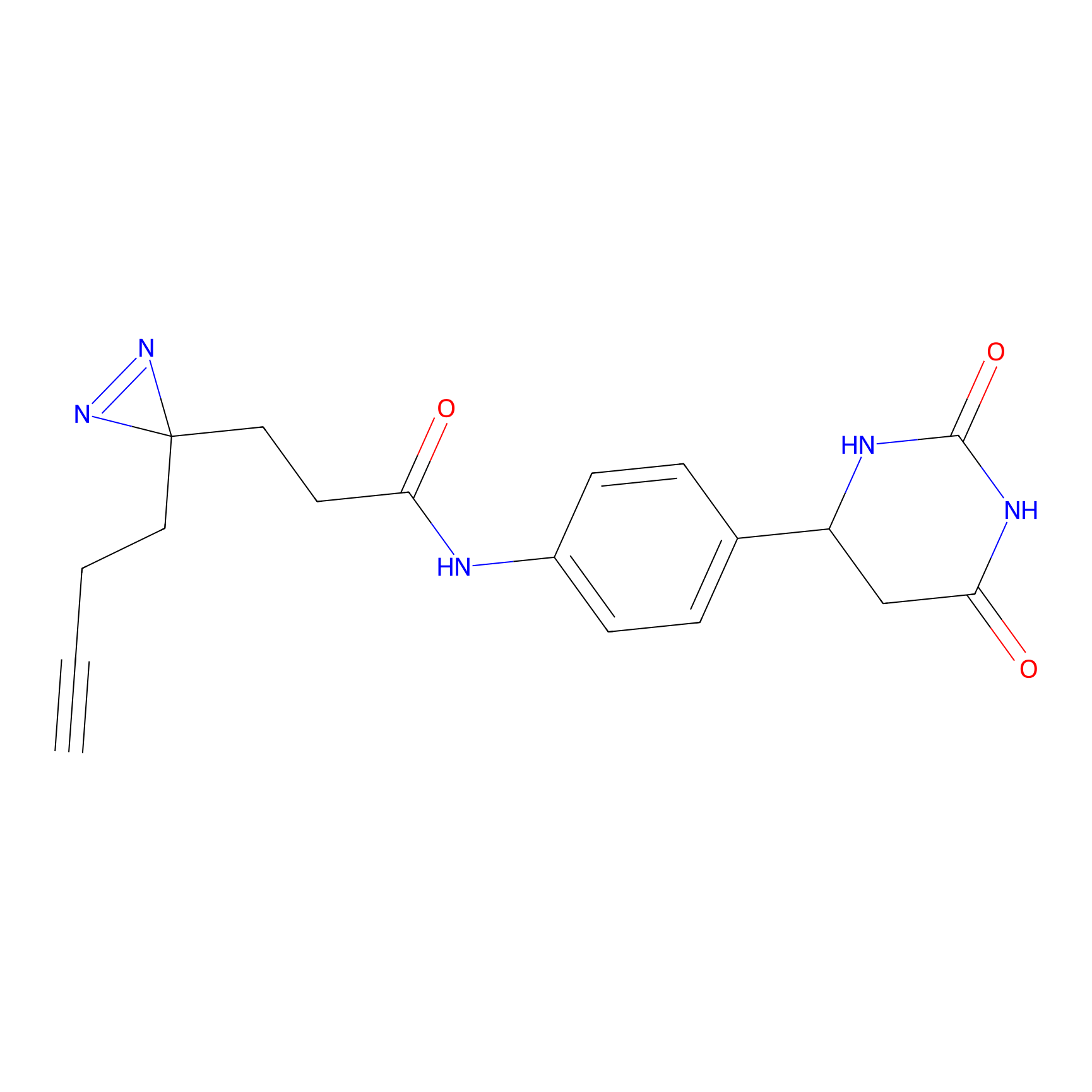
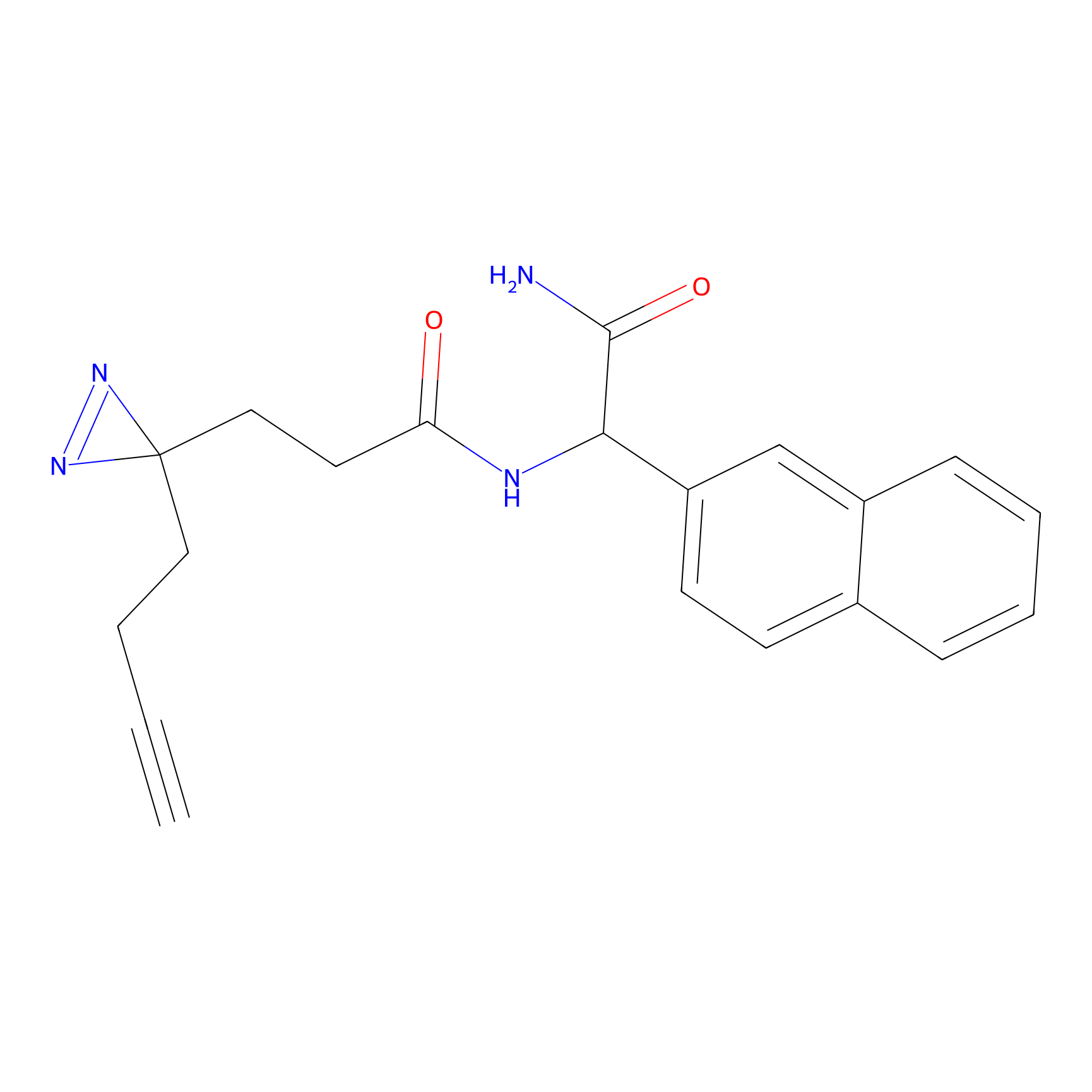
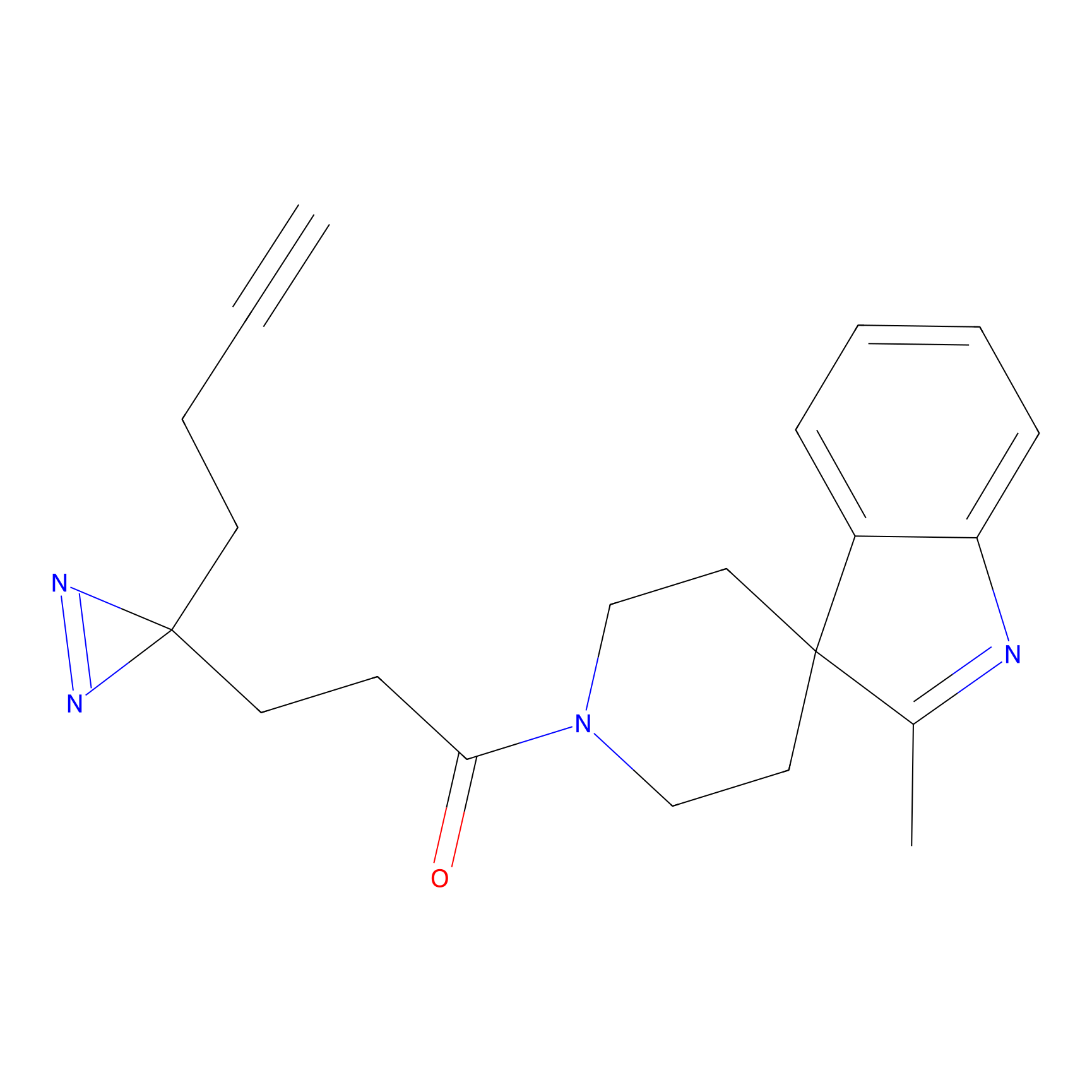
FBPP2 Probe Info |
 |
3.62 | LDD0318 | [1] | |
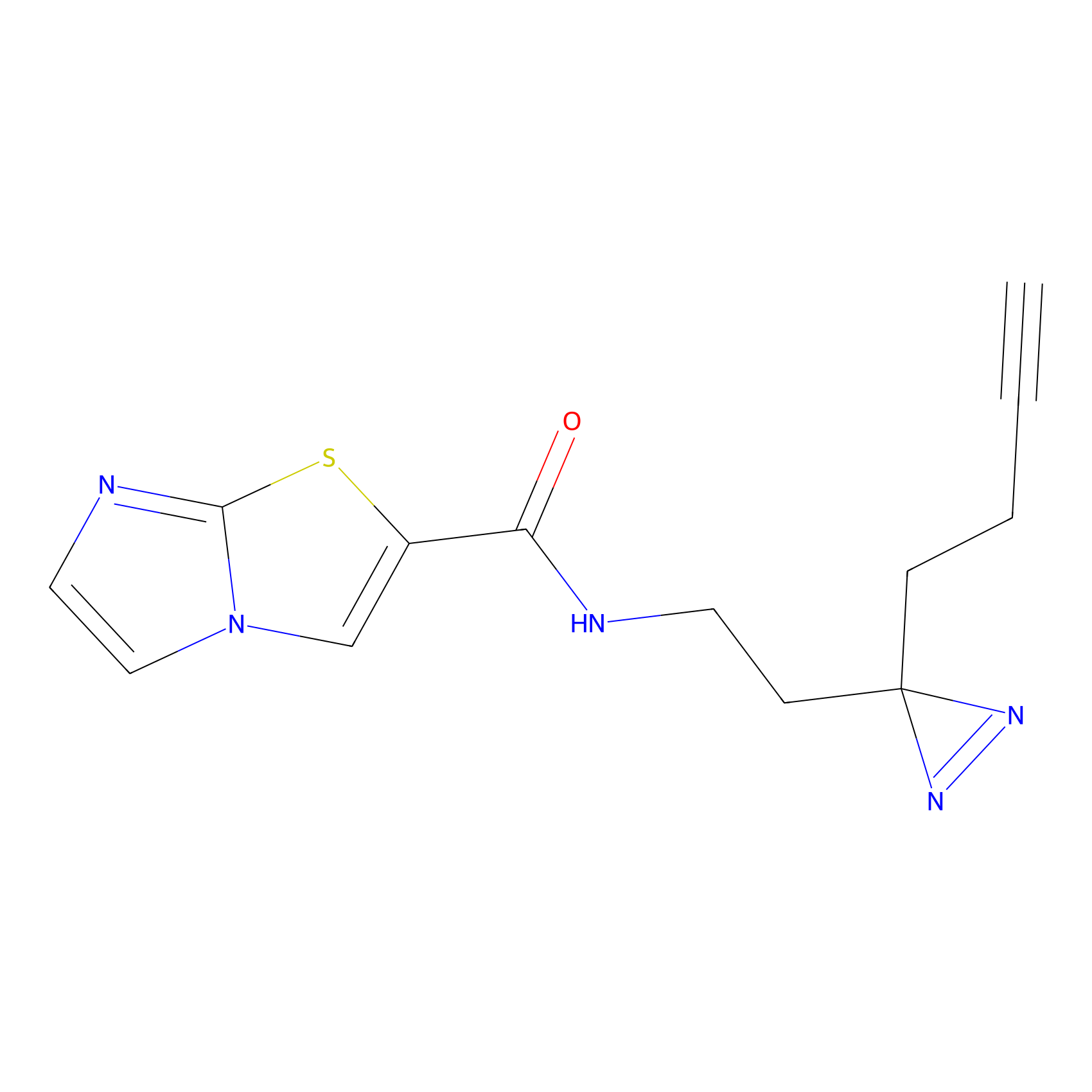
|
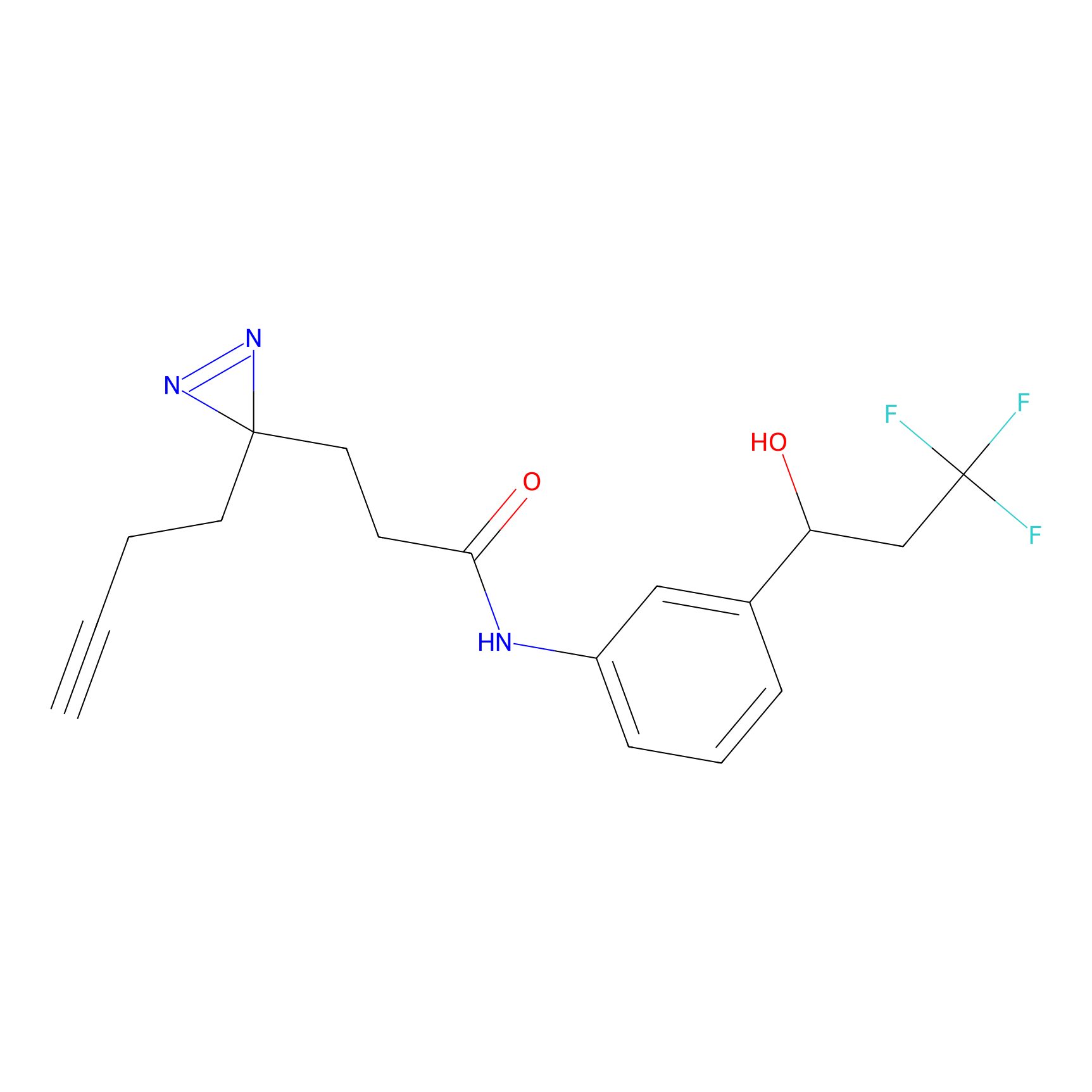
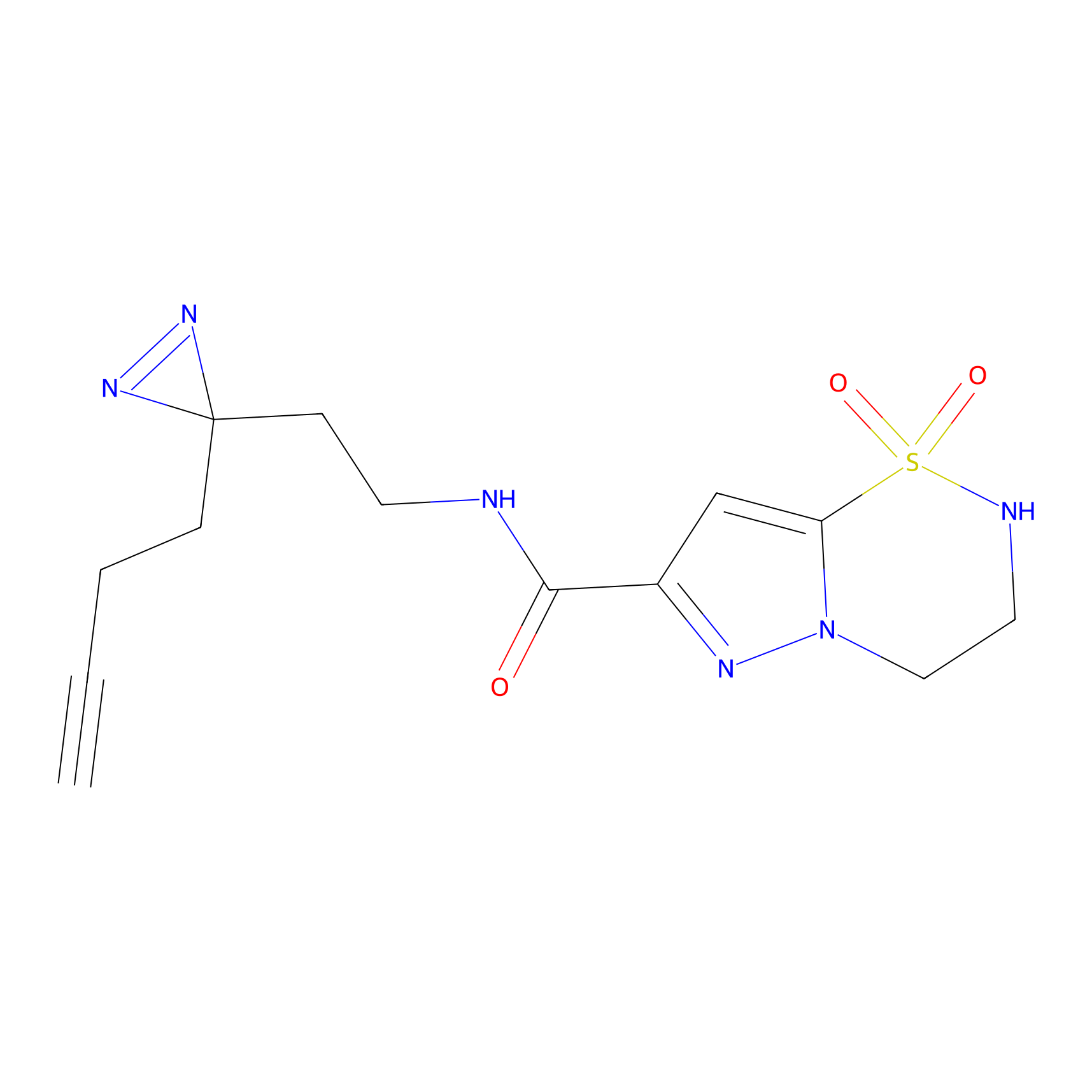
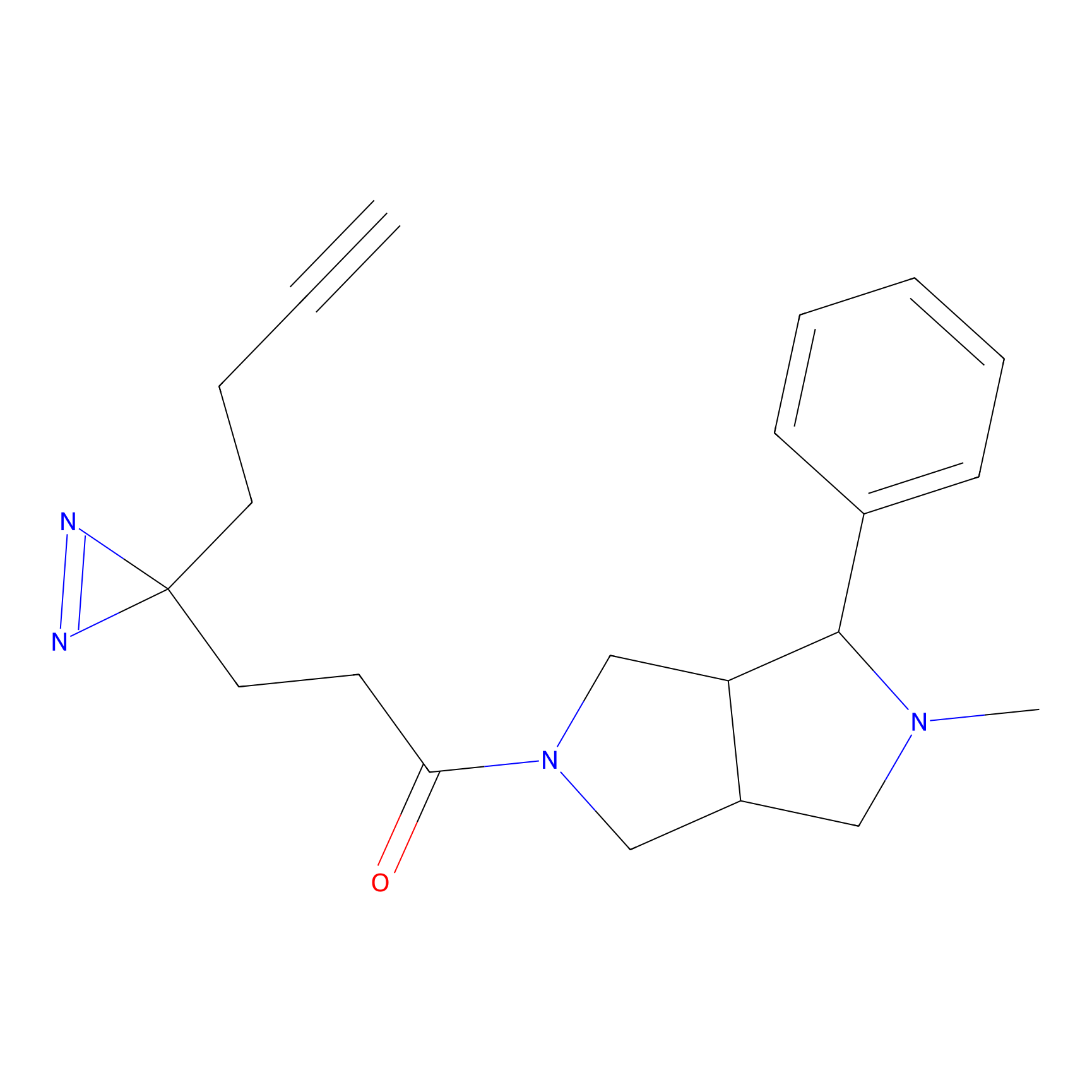
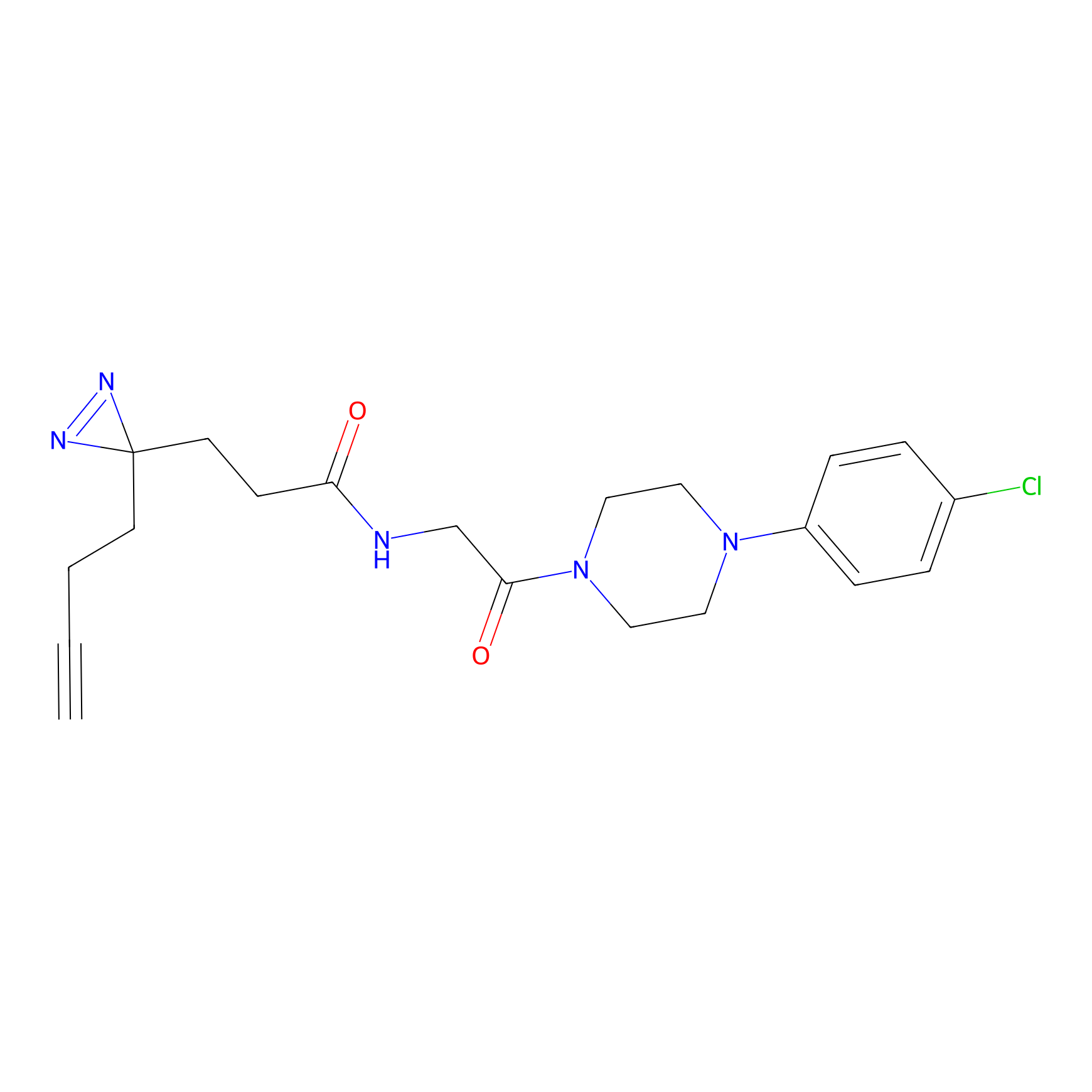
TG42 Probe Info |
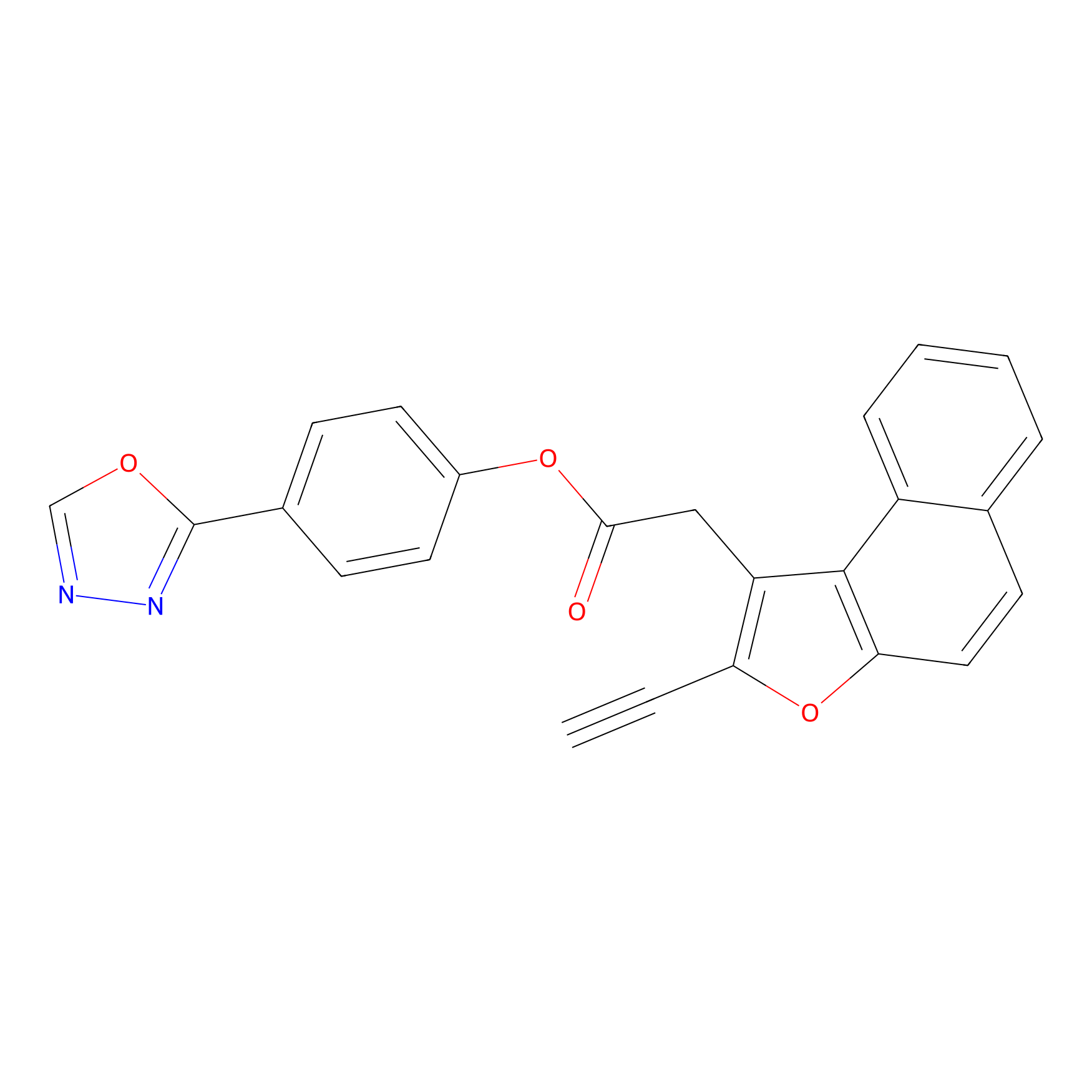 |
5.80 | LDD0326 | [2] | |
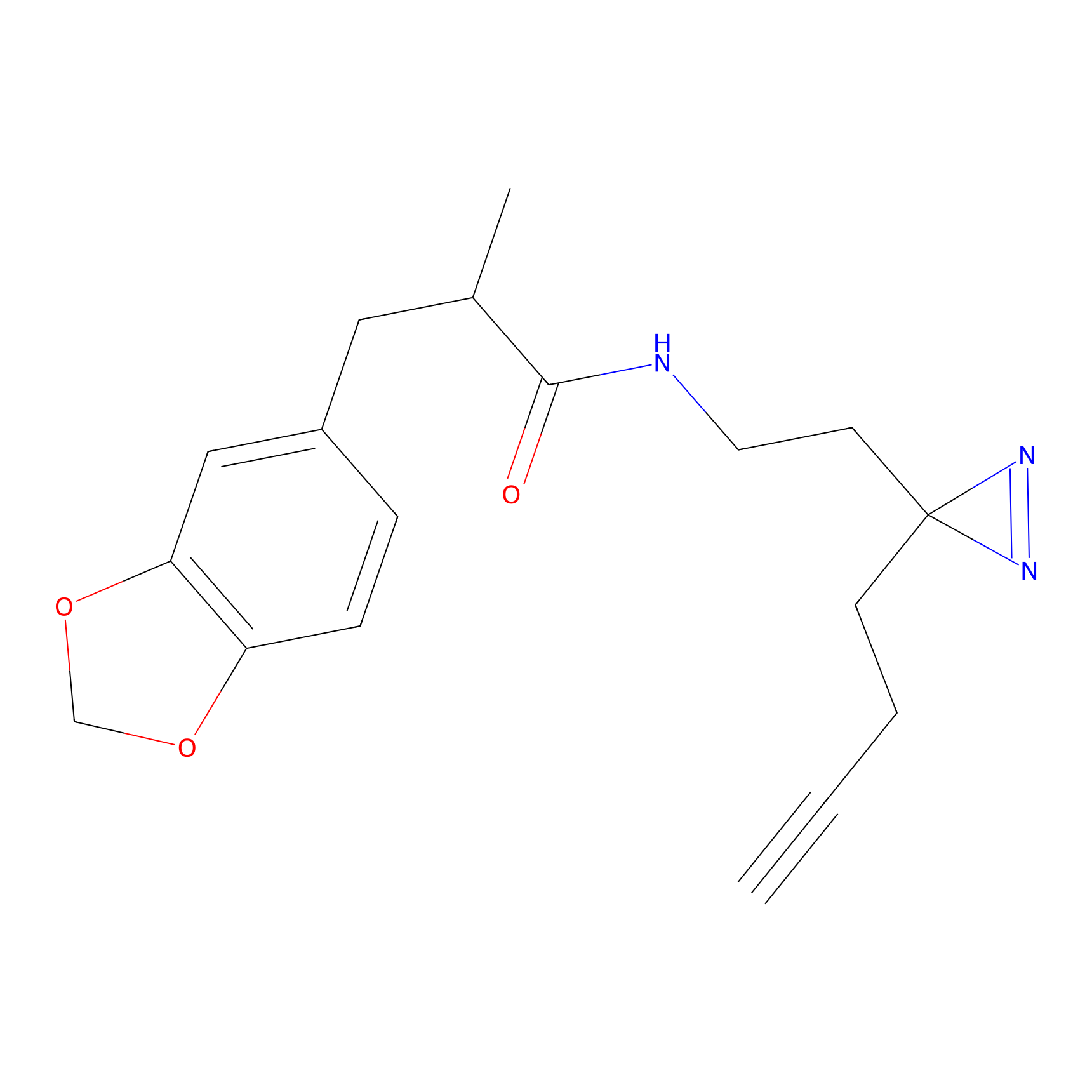
|
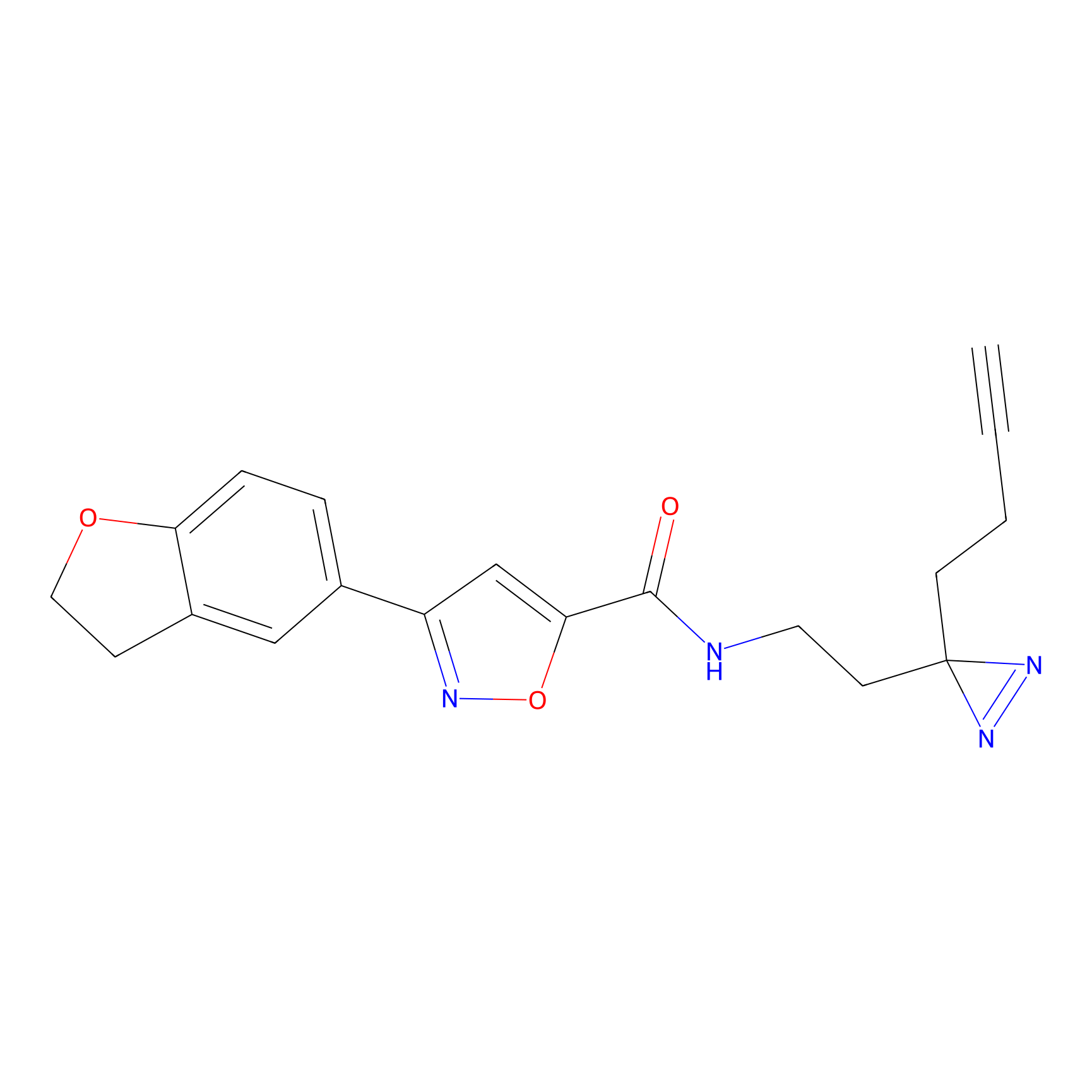
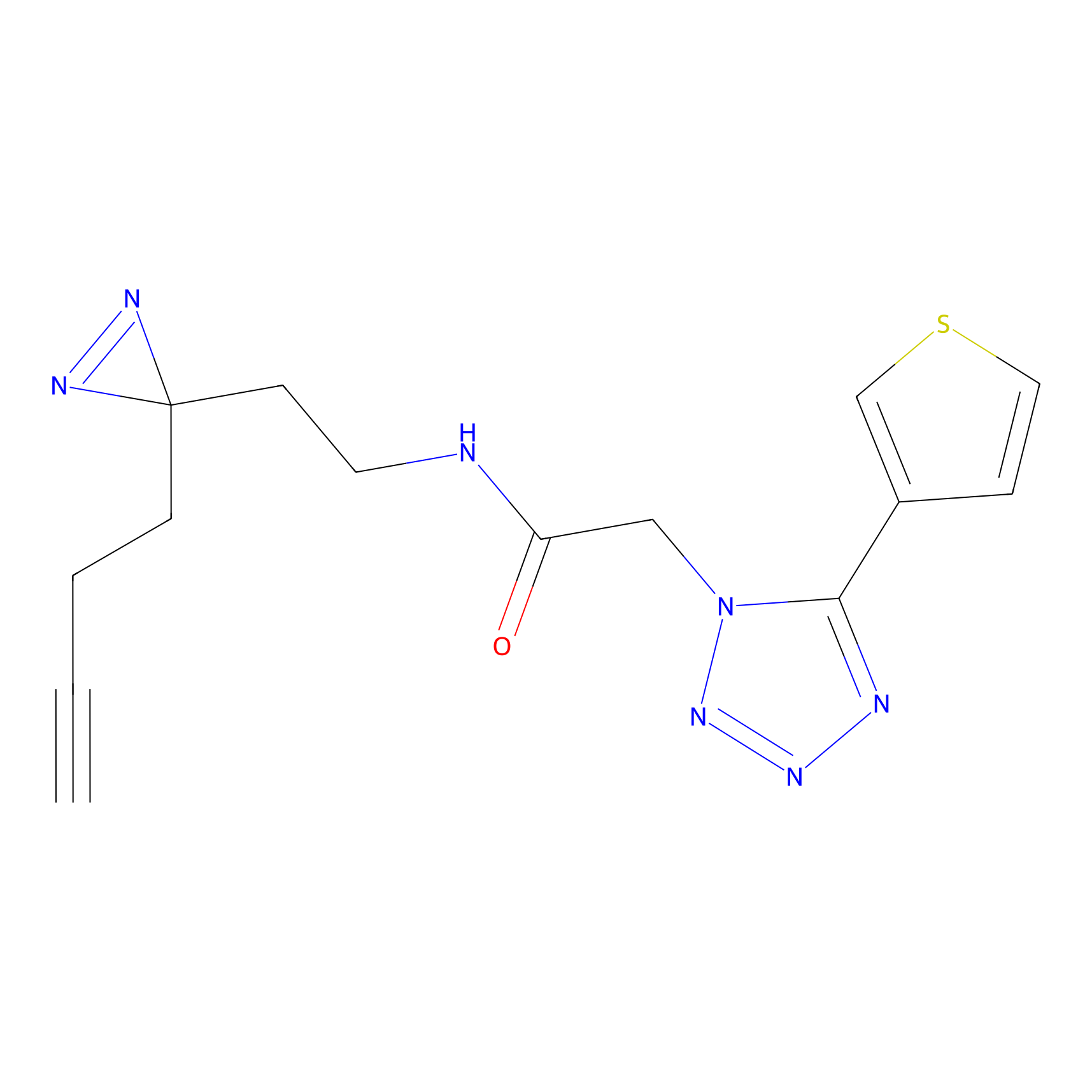
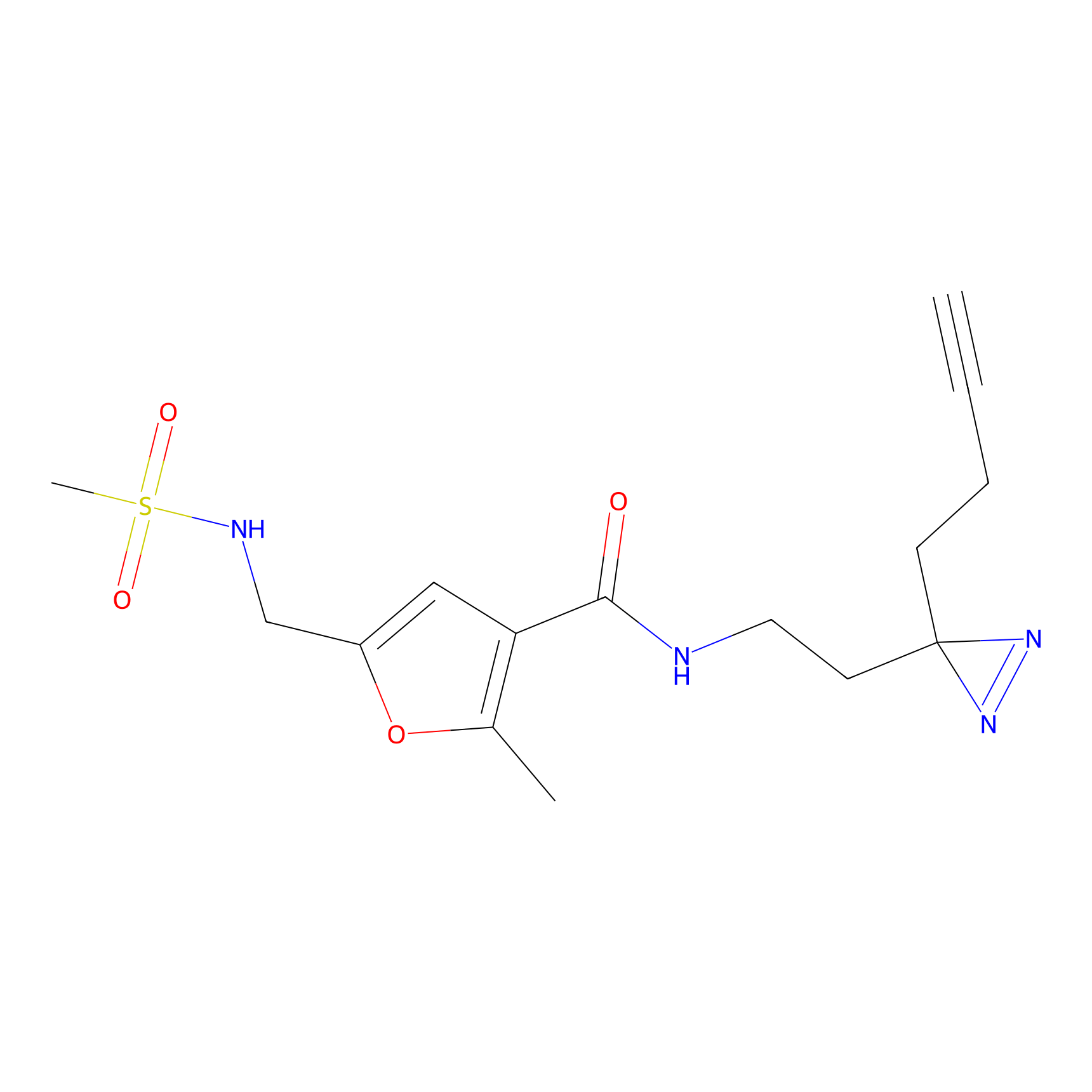
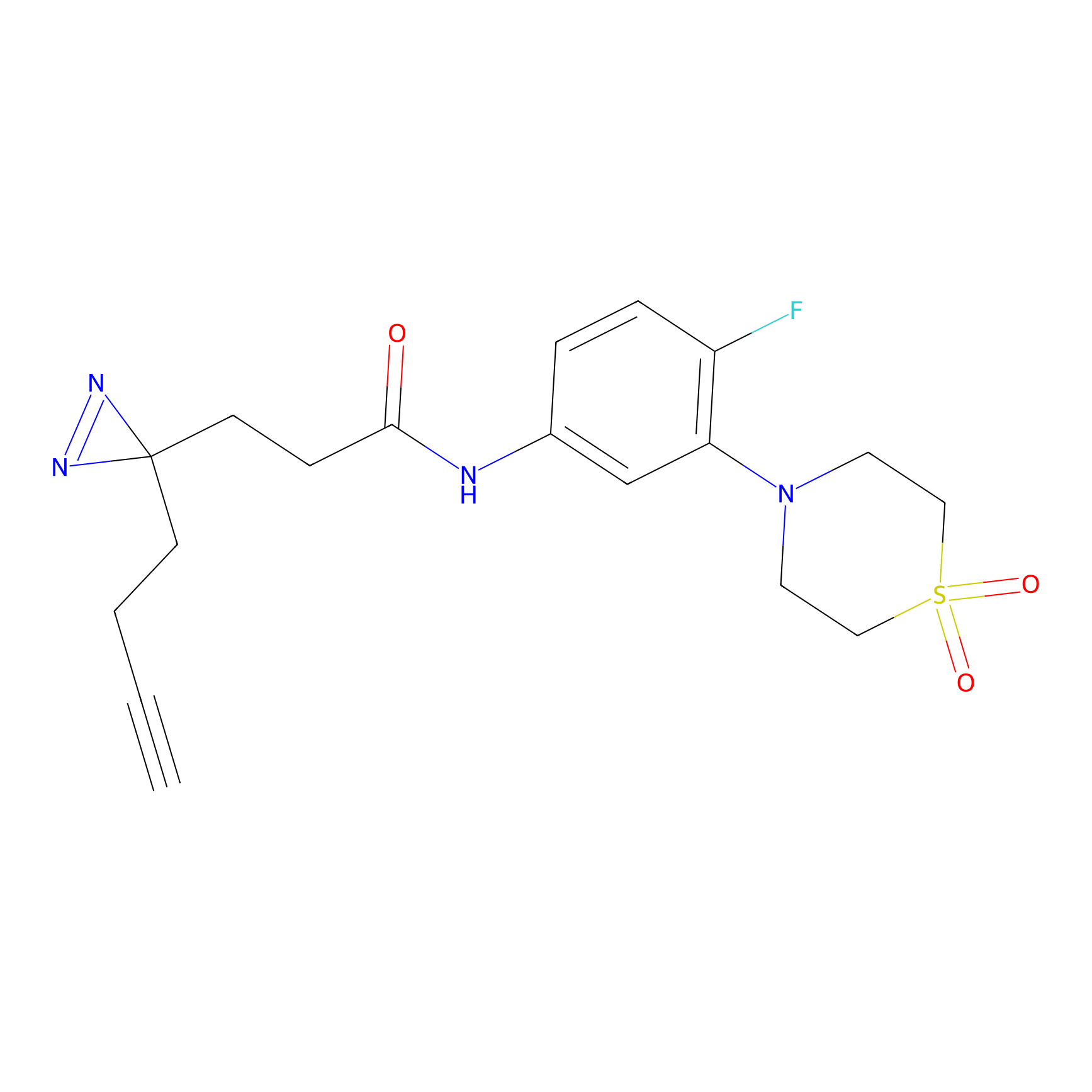
STPyne Probe Info |
 |
K320(10.00); K73(10.00) | LDD0277 | [3] | |
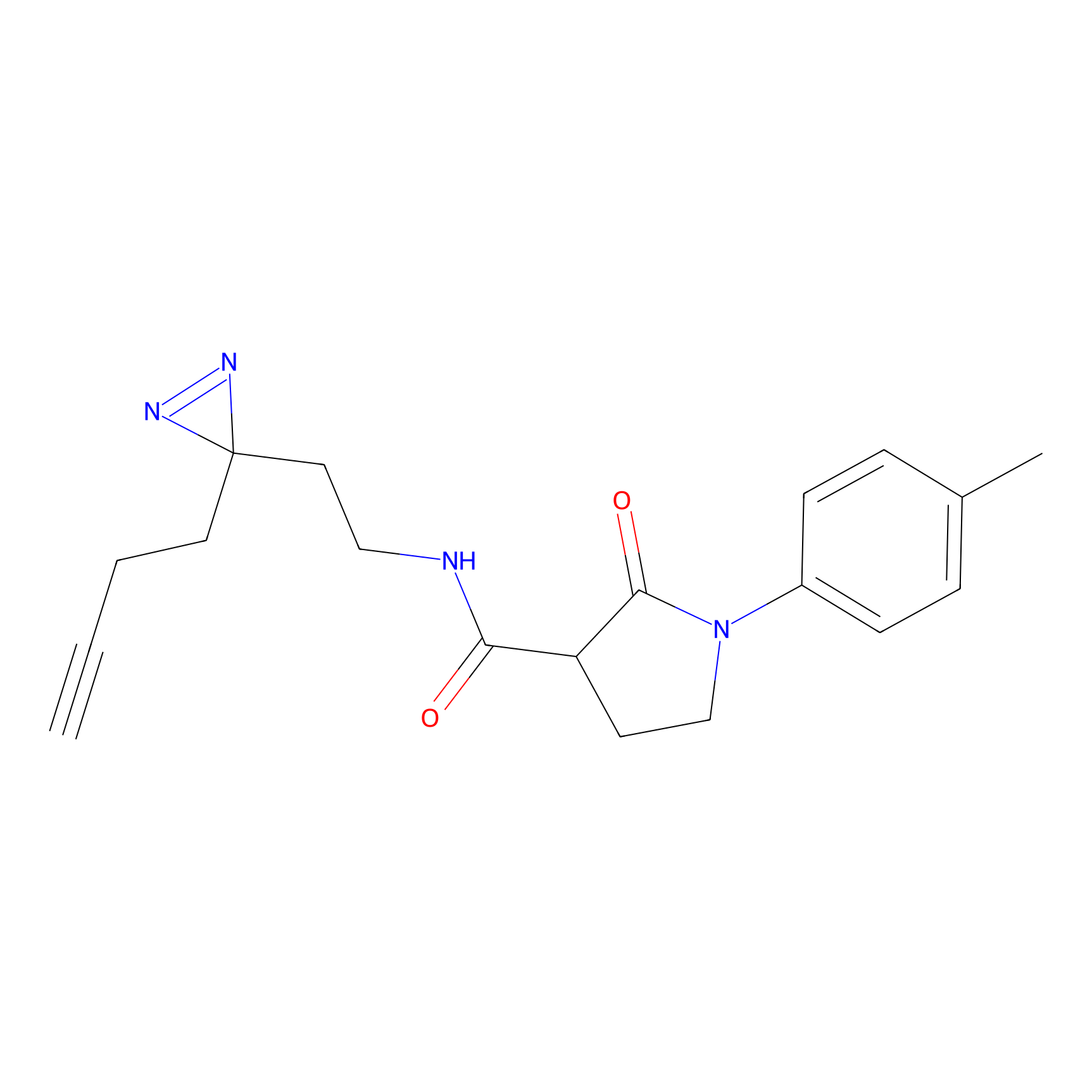
|
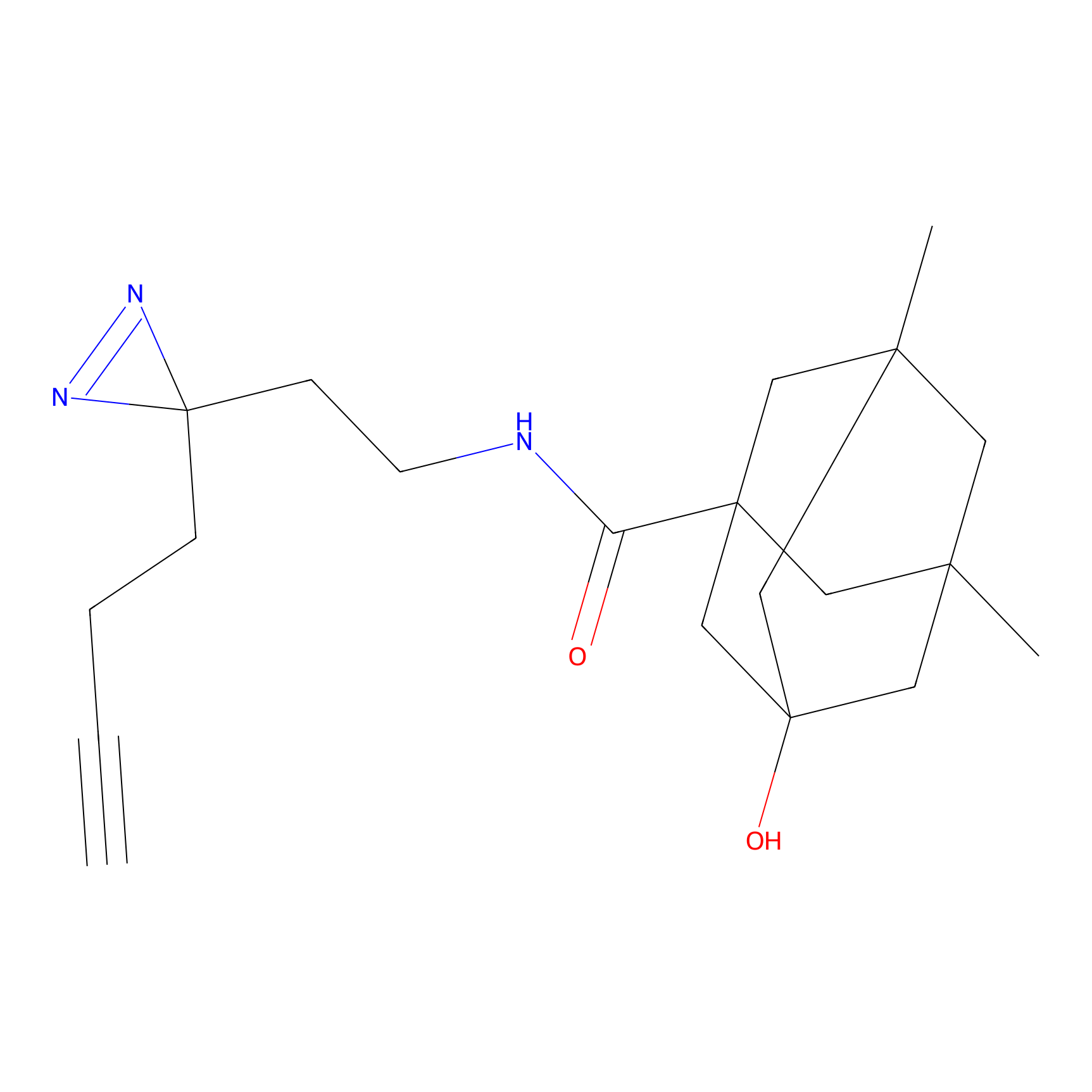
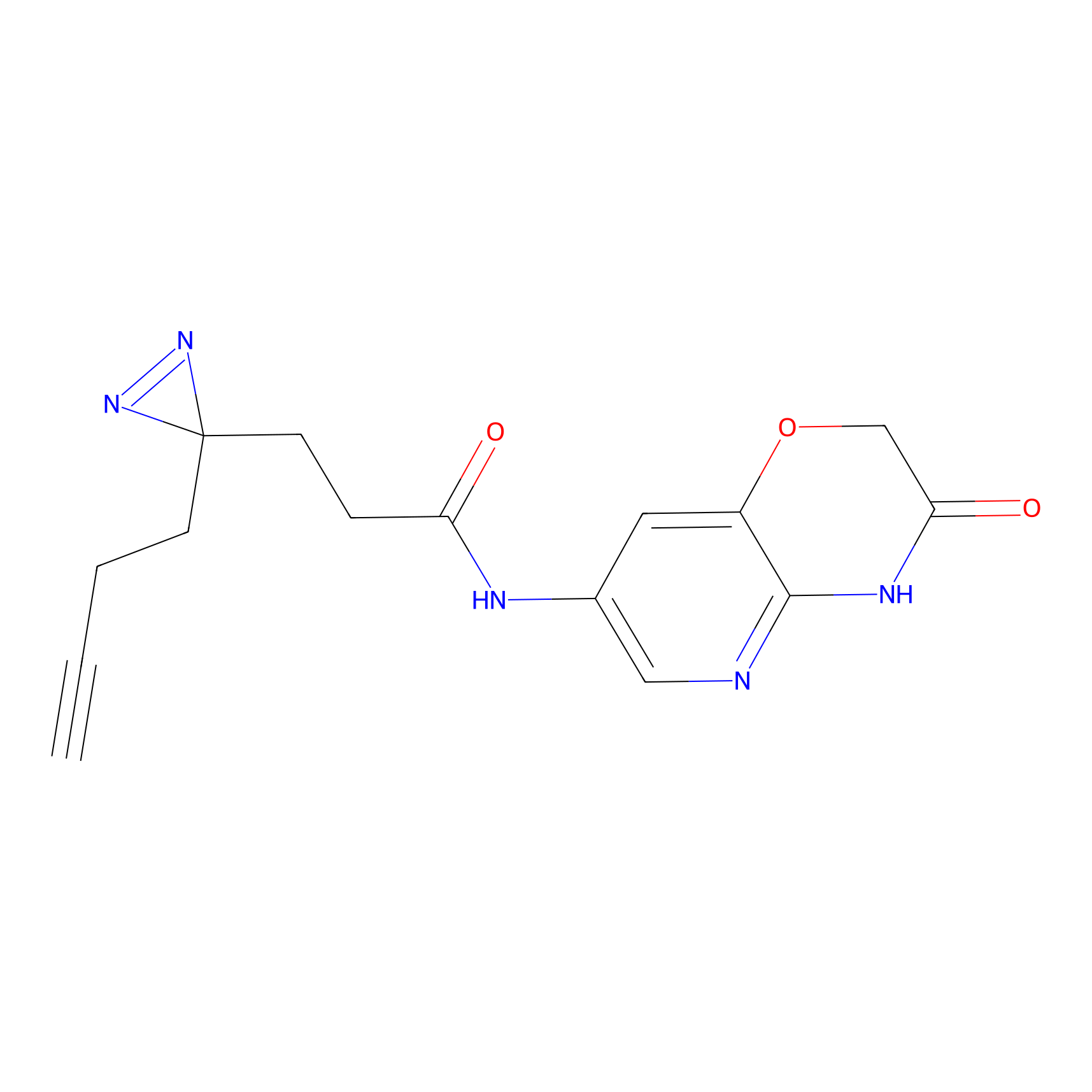
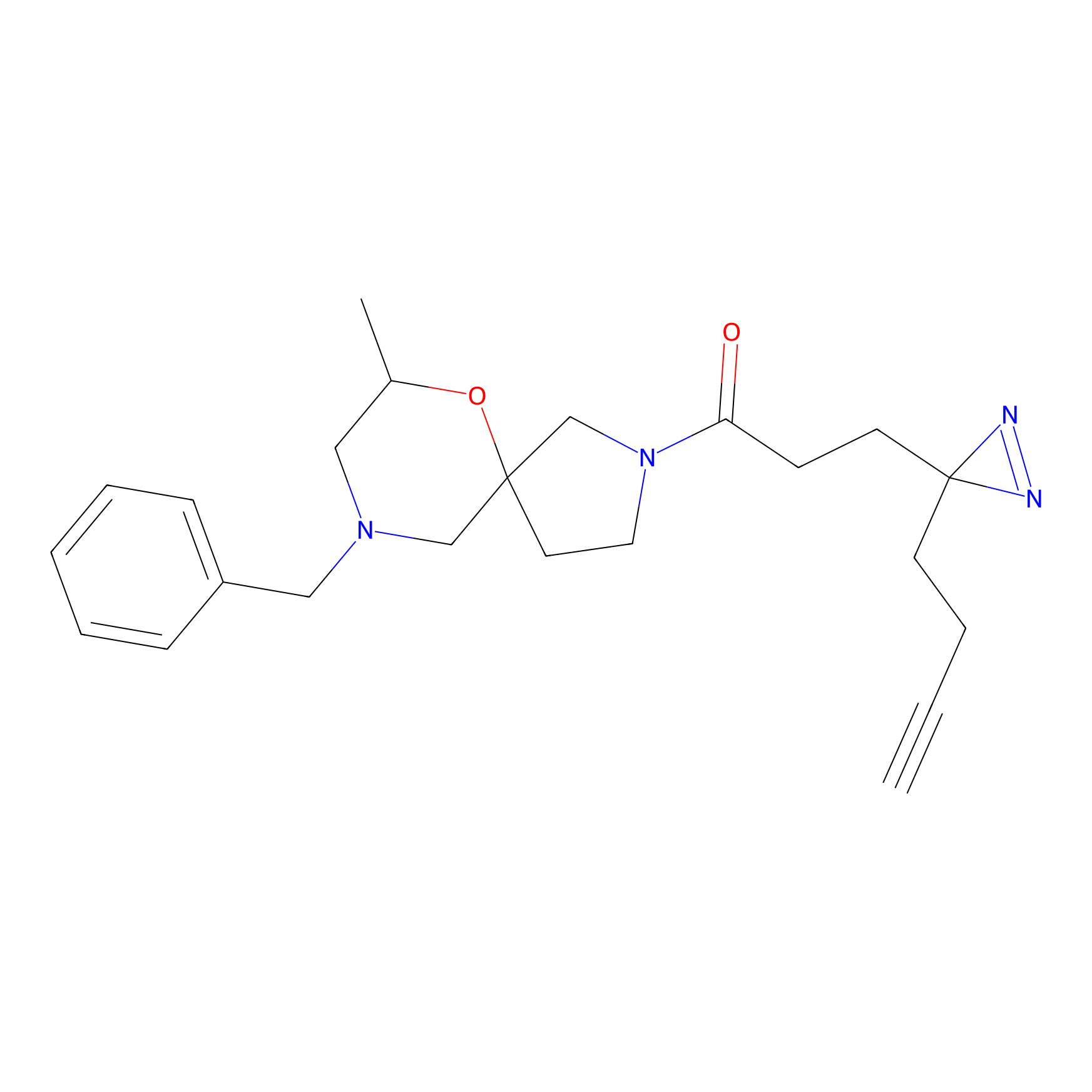
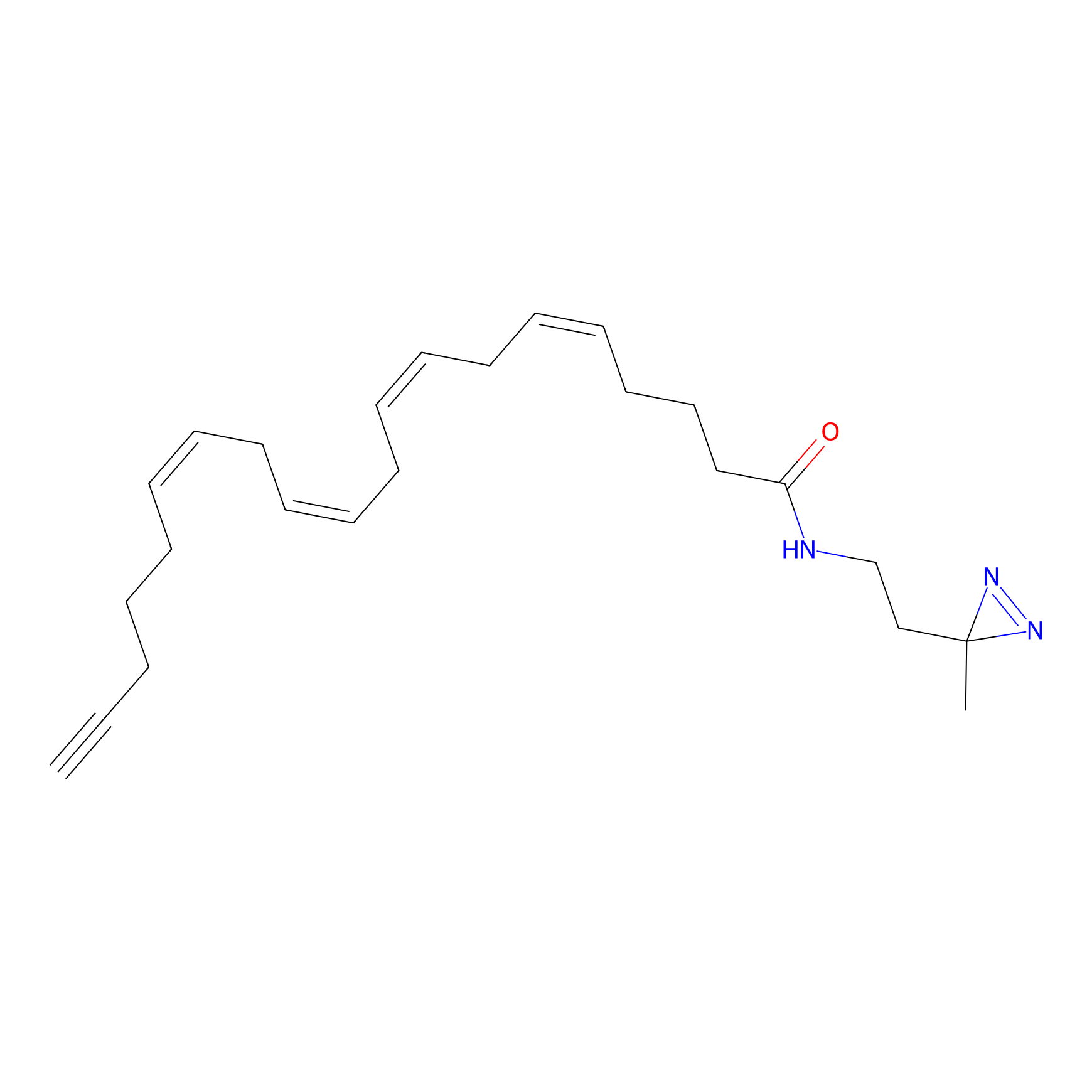
m-APA Probe Info |
 |
12.15 | LDD0403 | [4] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [5] | |
|
AOyne Probe Info |
 |
13.40 | LDD0443 | [6] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Glutathione S-transferase 3, mitochondrial (MGST3) | MAPEG family | O14880 | |||
| E3 ubiquitin-protein ligase MARCHF3 (MARCHF3) | . | Q86UD3 | |||
Transporter and channel
Other
References