Details of the Target
General Information of Target
| Target ID | LDTP01039 | |||||
|---|---|---|---|---|---|---|
| Target Name | Protein-S-isoprenylcysteine O-methyltransferase (ICMT) | |||||
| Gene Name | ICMT | |||||
| Gene ID | 23463 | |||||
| Synonyms |
PCCMT; Protein-S-isoprenylcysteine O-methyltransferase; EC 2.1.1.100; Isoprenylcysteine carboxylmethyltransferase; Prenylated protein carboxyl methyltransferase; PPMT; Prenylcysteine carboxyl methyltransferase; pcCMT
|
|||||
| 3D Structure | ||||||
| Sequence |
MAGCAARAPPGSEARLSLATFLLGASVLALPLLTRAGLQGRTGLALYVAGLNALLLLLYR
PPRYQIAIRACFLGFVFGCGTLLSFSQSSWSHFGWYMCSLSLFHYSEYLVTAVNNPKSLS LDSFLLNHSLEYTVAALSSWLEFTLENIFWPELKQITWLSVTGLLMVVFGECLRKAAMFT AGSNFNHVVQNEKSDTHTLVTSGVYAWFRHPSYVGWFYWSIGTQVMLCNPICGVSYALTV WRFFRDRTEEEEISLIHFFGEEYLEYKKRVPTGLPFIKGVKVDL |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Class VI-like SAM-binding methyltransferase superfamily, Isoprenylcysteine carboxyl methyltransferase family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function | Catalyzes the post-translational methylation of isoprenylated C-terminal cysteine residues. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
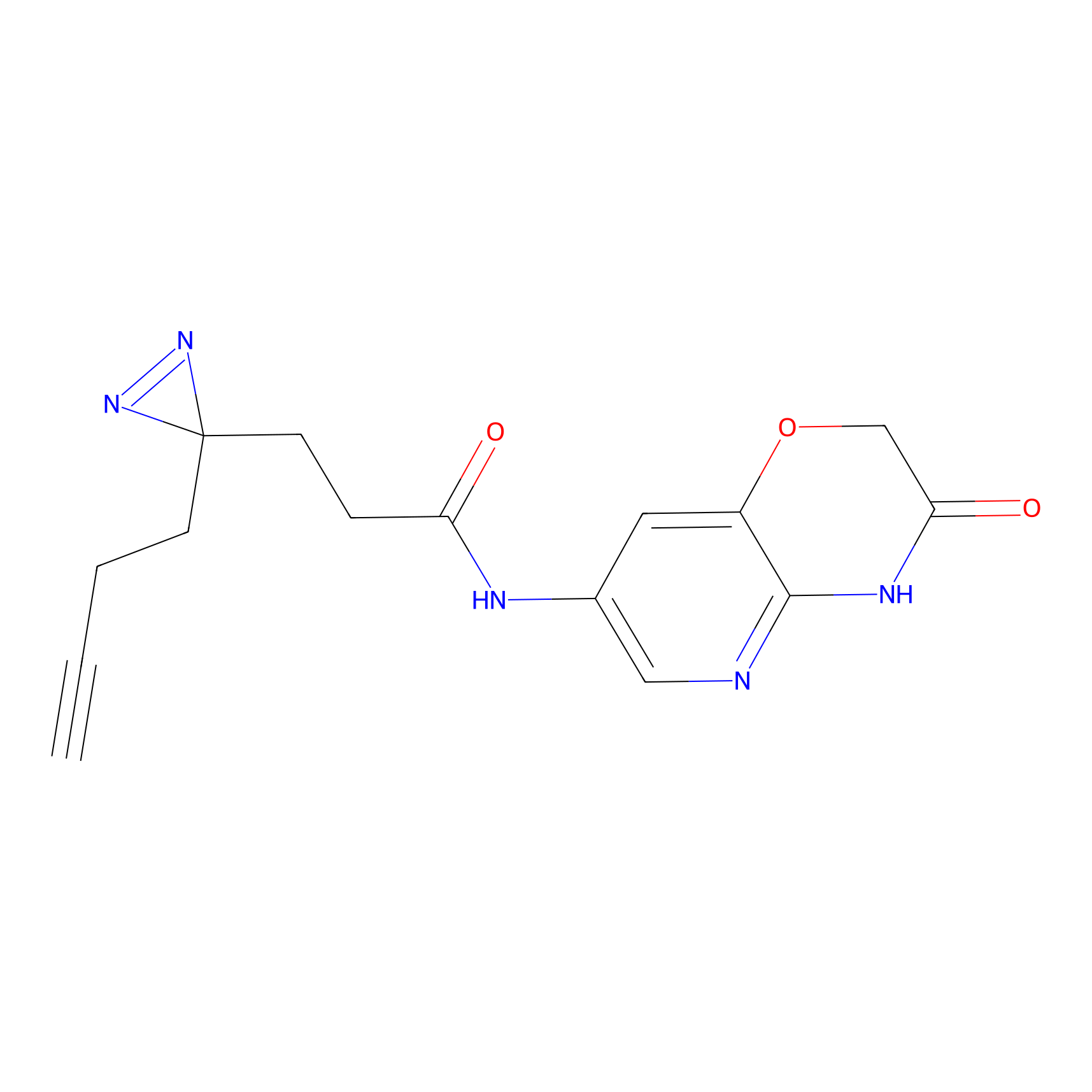
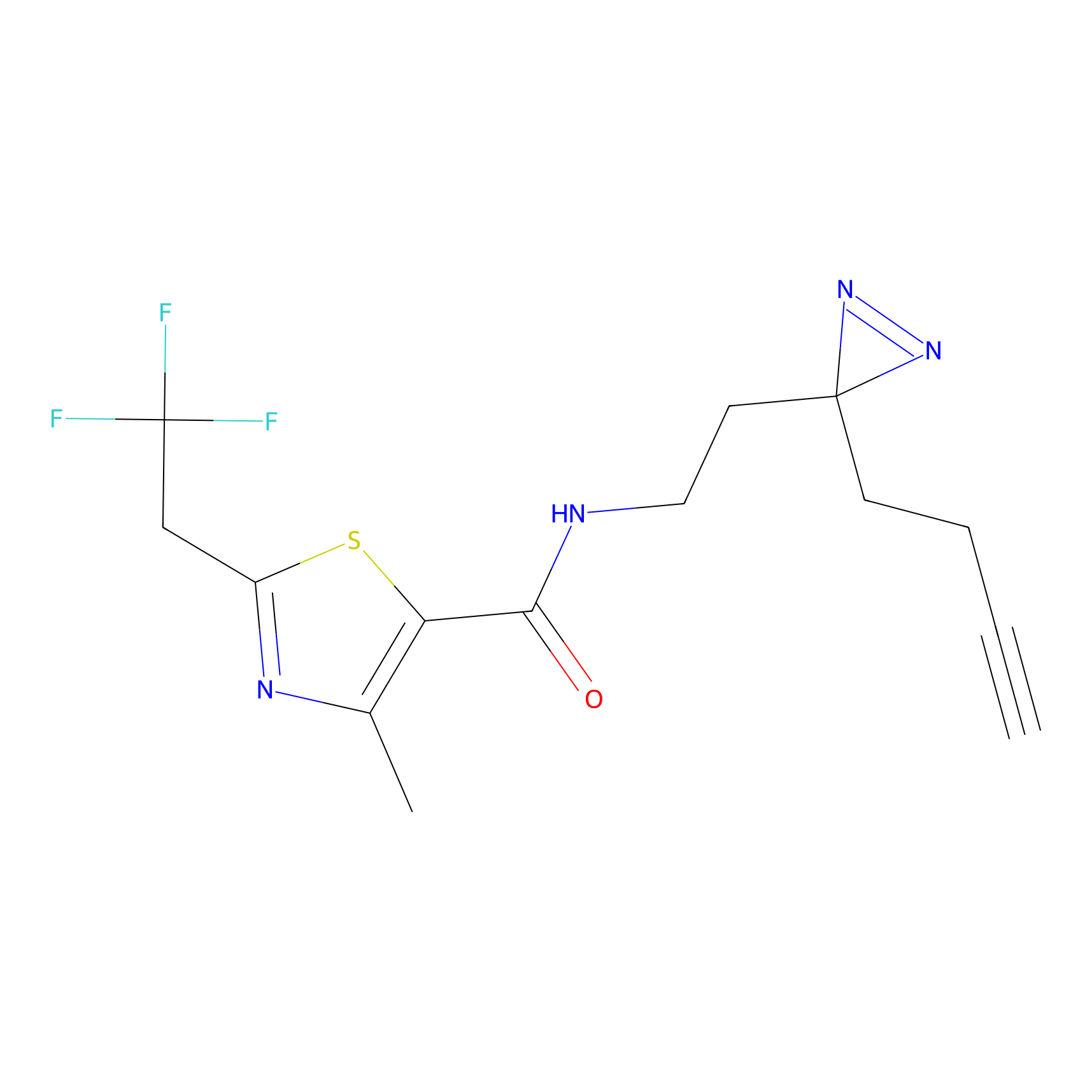
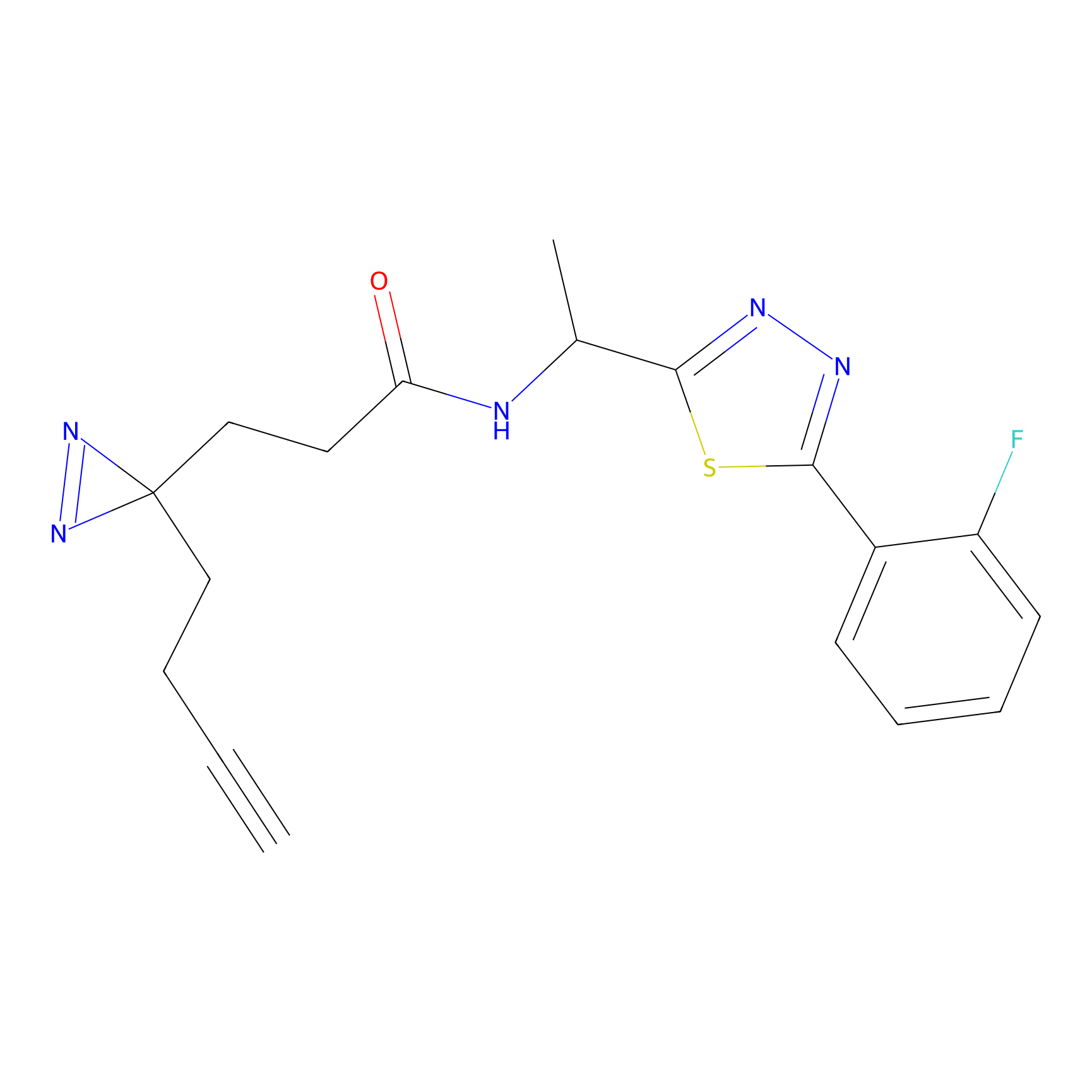
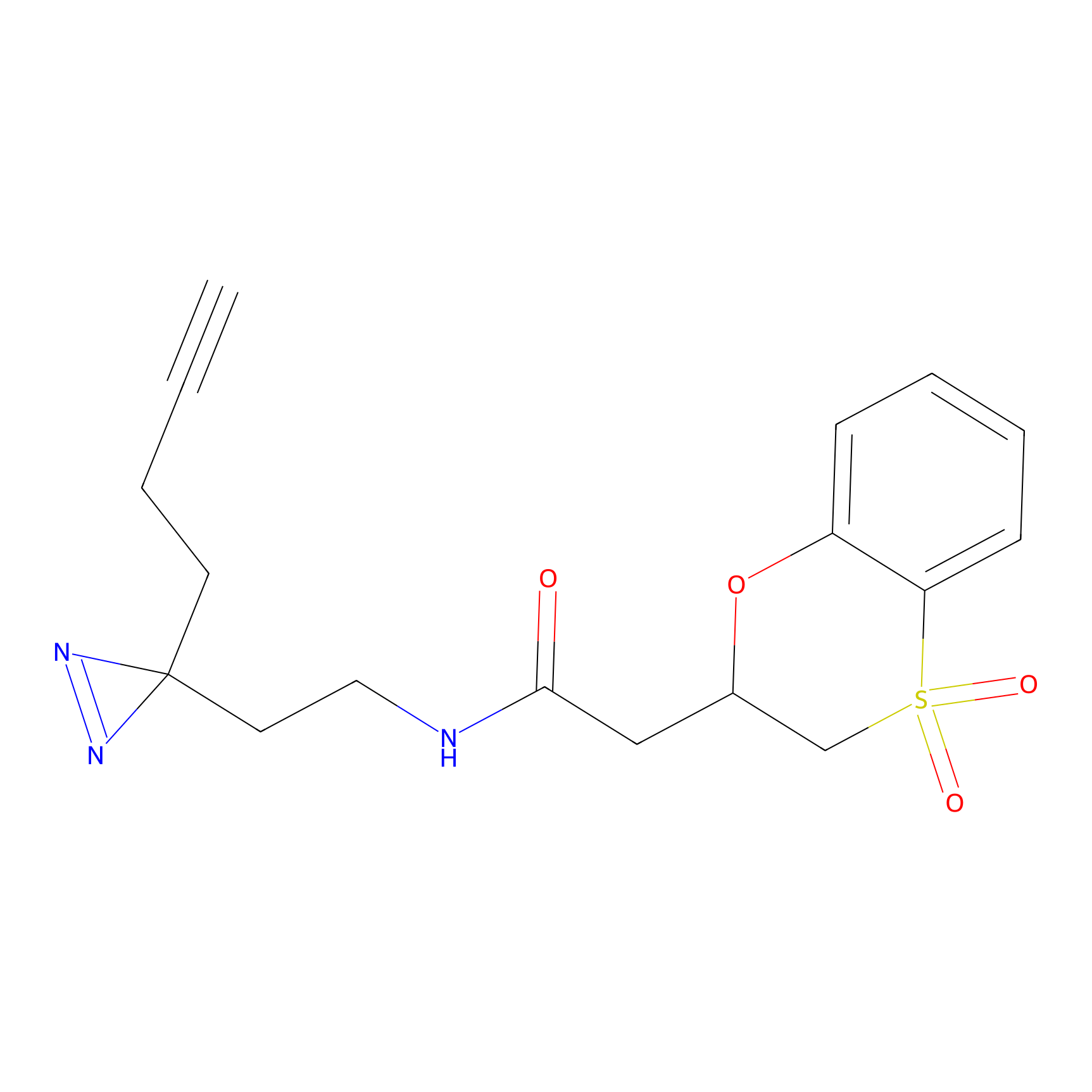
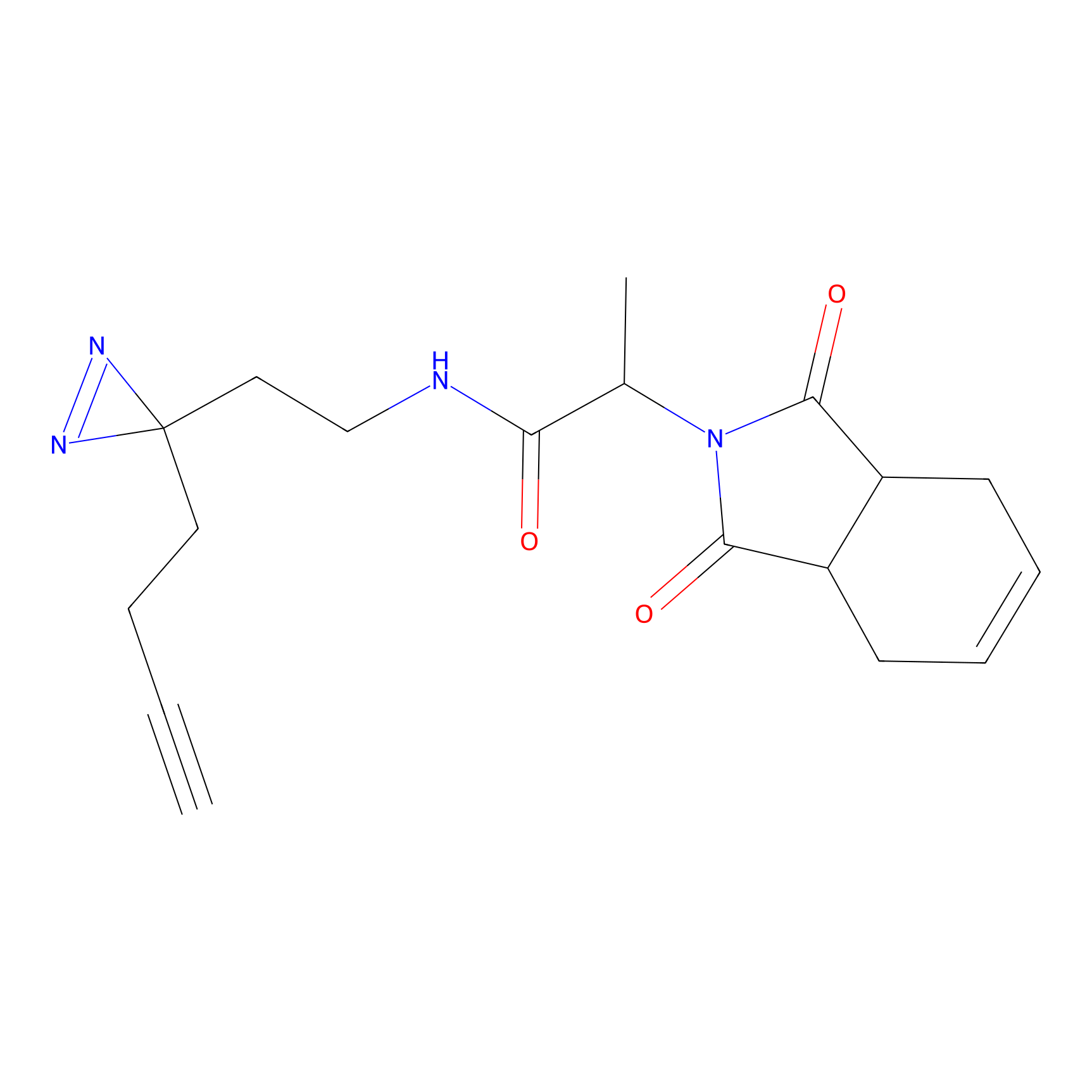
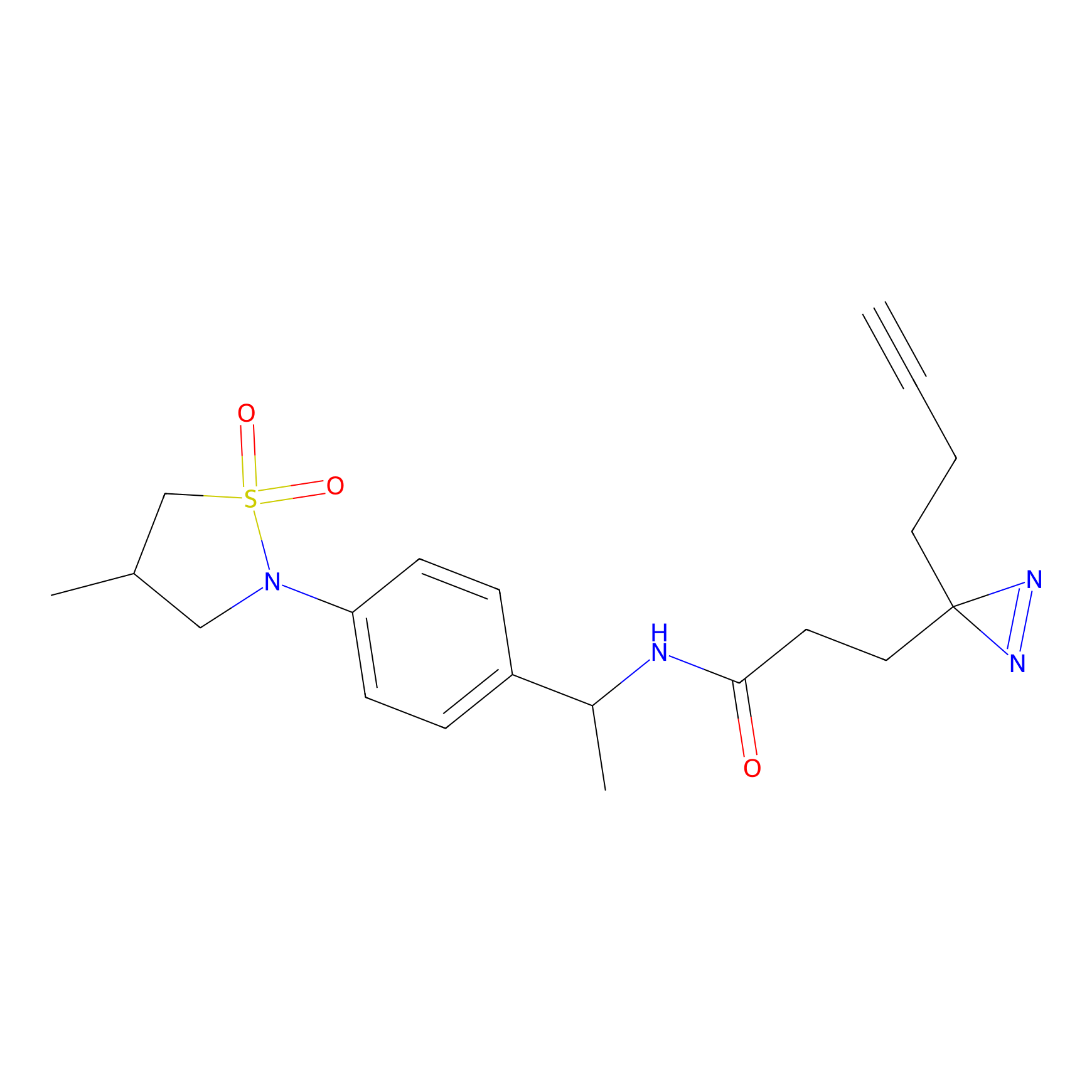
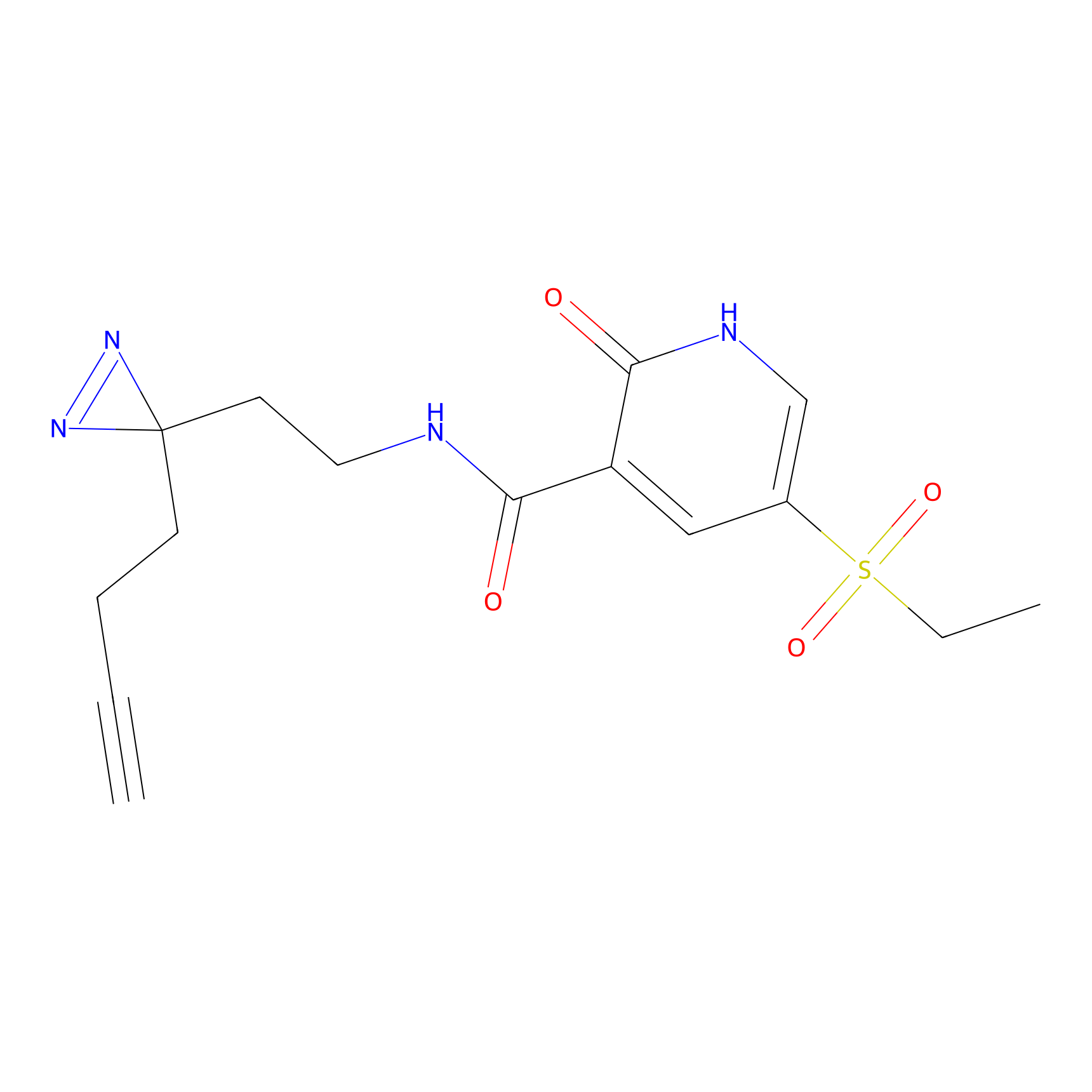
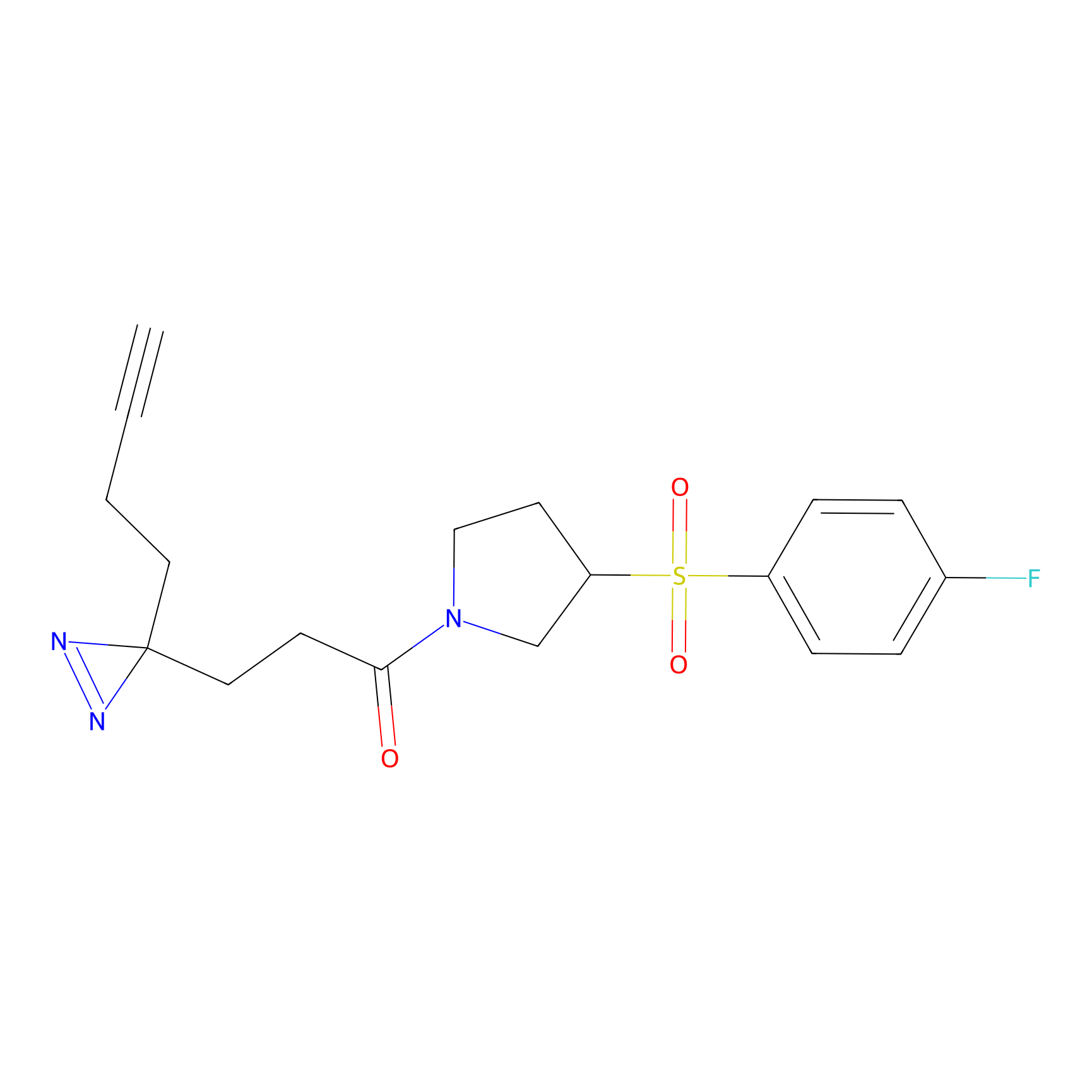
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
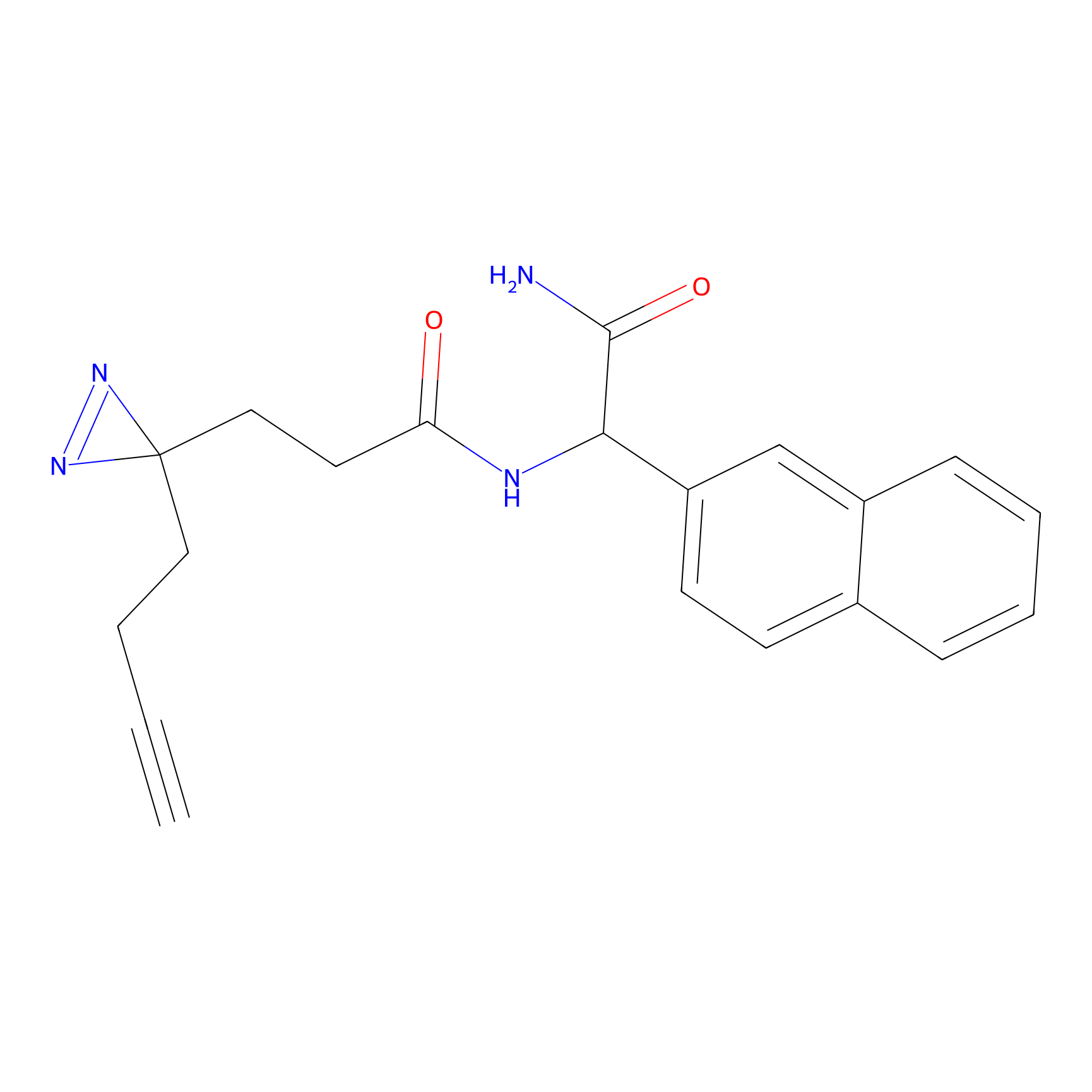
DA-P2 Probe Info |
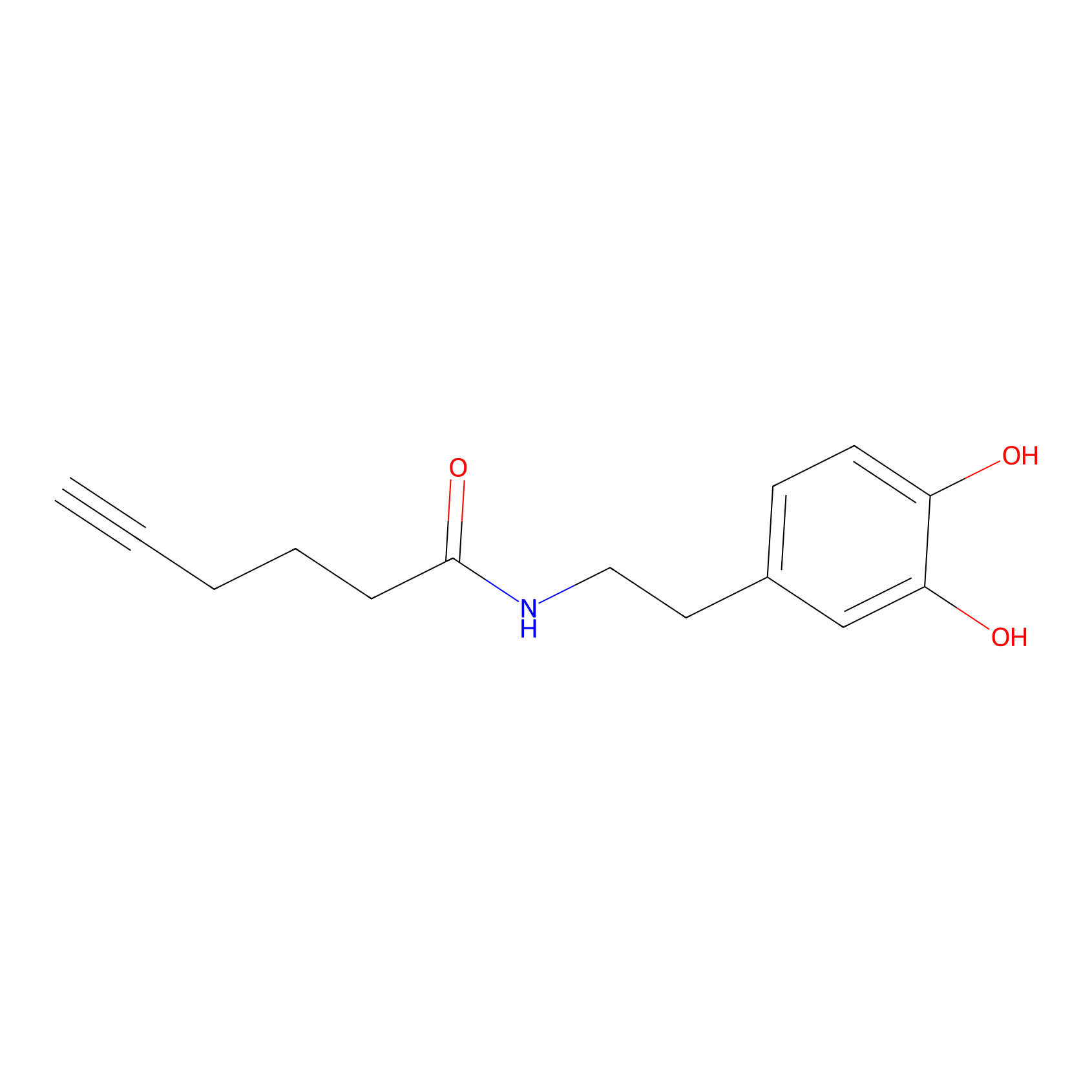 |
5.57 | LDD0348 | [1] | |
|
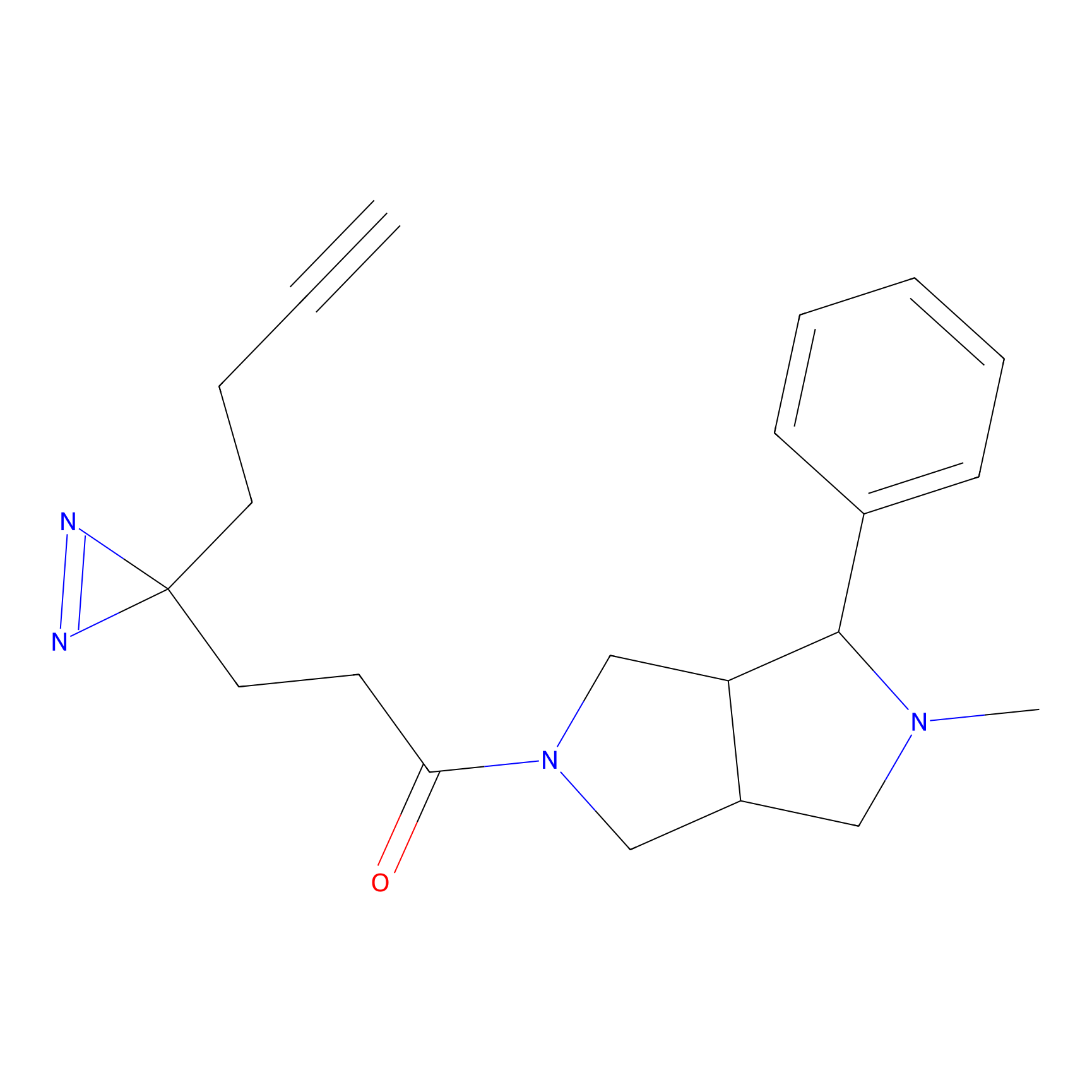
CHEMBL5175495 Probe Info |
 |
9.10 | LDD0196 | [2] | |
|
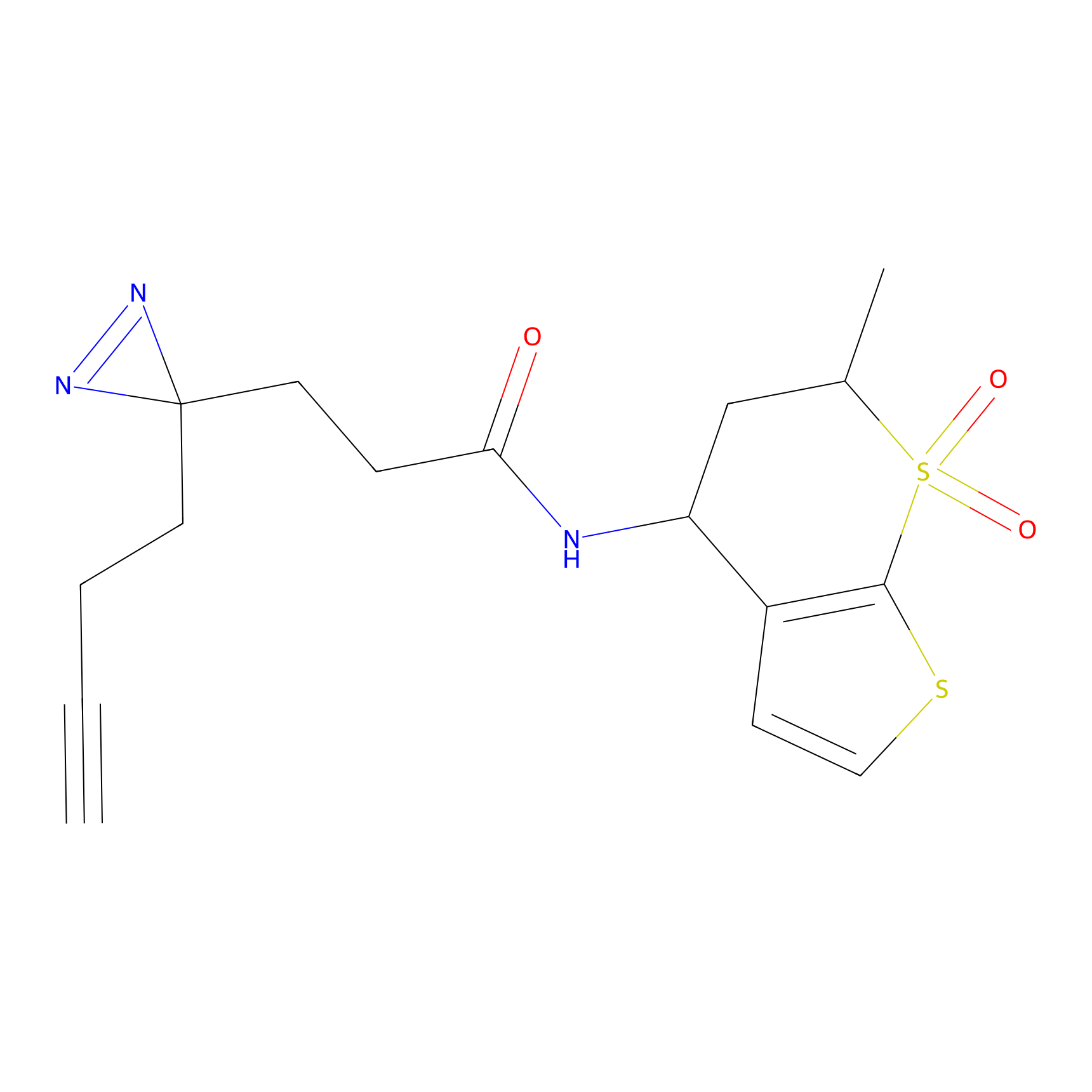
STPyne Probe Info |
 |
K278(7.57); K281(6.11) | LDD0277 | [3] | |
|
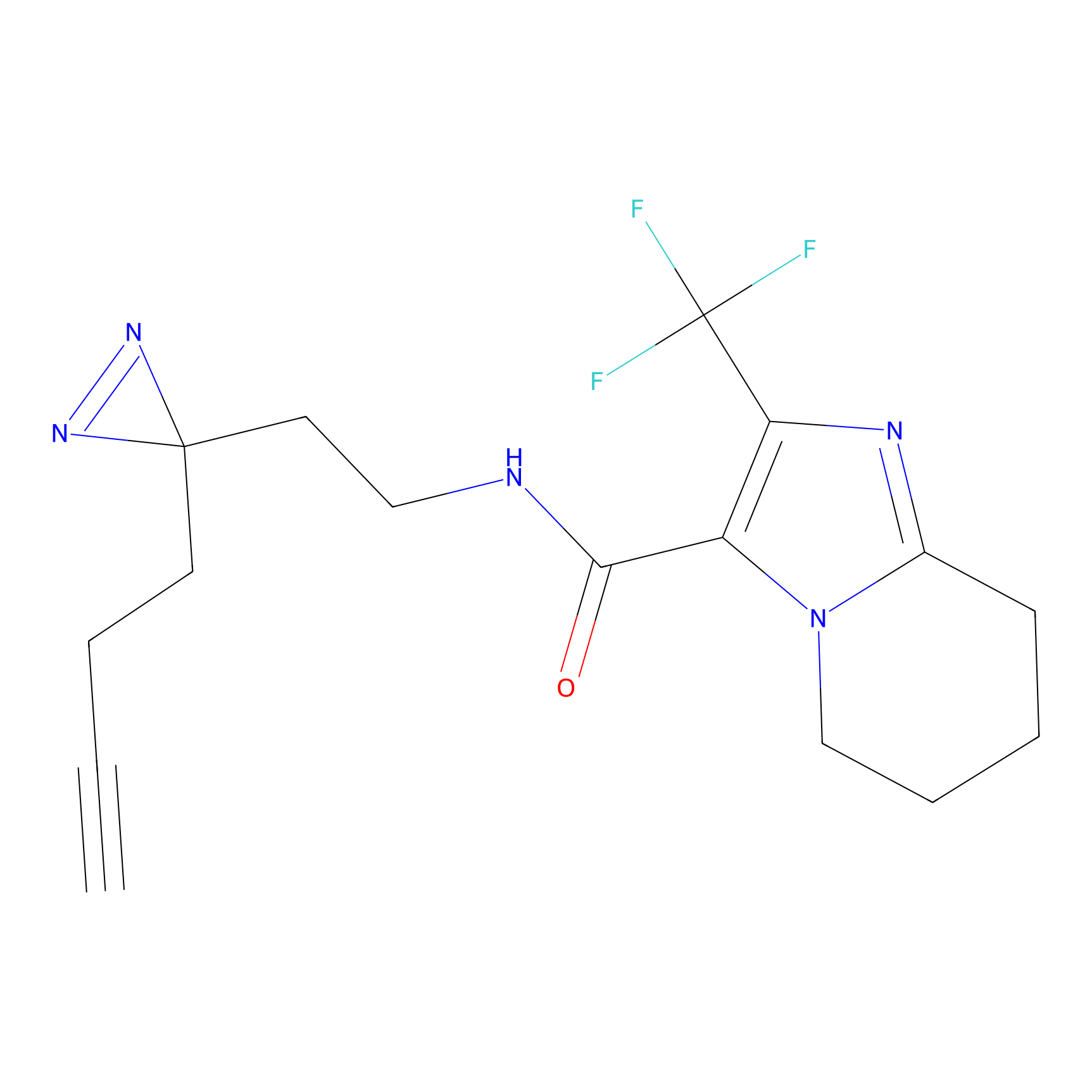
Johansson_61 Probe Info |
 |
_(11.53) | LDD1489 | [4] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [5] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [6] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [6] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [7] | |
|
AOyne Probe Info |
 |
12.20 | LDD0443 | [8] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Protein jagunal homolog 1 (JAGN1) | Jagunal family | Q8N5M9 | |||
| PDZK1-interacting protein 1 (PDZK1IP1) | . | Q13113 | |||
References