Details of the Target
General Information of Target
| Target ID | LDTP06426 | |||||
|---|---|---|---|---|---|---|
| Target Name | Translocating chain-associated membrane protein 1 (TRAM1) | |||||
| Gene Name | TRAM1 | |||||
| Gene ID | 23471 | |||||
| Synonyms |
TRAM; Translocating chain-associated membrane protein 1; Protein TRAM1 |
|||||
| 3D Structure | ||||||
| Sequence |
MAIRKKSTKSPPVLSHEFVLQNHADIVSCVAMVFLLGLMFEITAKASIIFVTLQYNVTLP
ATEEQATESVSLYYYGIKDLATVFFYMLVAIIIHAVIQEYMLDKINRRMHFSKTKHSKFN ESGQLSAFYLFACVWGTFILISENYISDPTILWRAYPHNLMTFQMKFFYISQLAYWLHAF PELYFQKTKKEDIPRQLVYIGLYLFHIAGAYLLNLNHLGLVLLVLHYFVEFLFHISRLFY FSNEKYQKGFSLWAVLFVLGRLLTLILSVLTVGFGLARAENQKLDFSTGNFNVLAVRIAV LASICVTQAFMMWKFINFQLRRWREHSAFQAPAVKKKPTVTKGRSSKKGTENGVNGTLTS NVADSPRNKKEKSS |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
TRAM family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Involved in the translocation of nascent protein chains into or through the endoplasmic reticulum (ER) membrane by facilitating the proper chain positioning at the SEC61 channel. Regulates the exposure of nascent secretory protein chain to the cytosol during translocation into the ER. May affect the phospholipid bilayer in the vicinity of the lateral gate of the SEC61 channel, thereby facilitating ER protein transport. Intimately associates with transmembrane (TM) domain of nascent membrane proteins during the entire integration process into the ER membrane. Associates with the second TM domain of G-protein-coupled receptor opsin/OPSD nascent chain in the ER membrane, which may facilitate its integration into the membrane. Under conditions of ER stress, participates in the disposal of misfolded ER membrane proteins during the unfolded protein response (UPR), an integrated stress response (ISR) pathway, by selectively retrotranslocating misfolded ER-membrane proteins from the ER into the cytosol where they are ubiquitinated and degraded by the proteasome.; (Microbial infection) In case of cytomegalovirus infection, participates in US2- and US11-mediated ER-to-cytosol retrotranslocation and subsequent degradation of major histocompatibility complex (MHC) class I heavy chains, thereby decreasing the immune detection by cytotoxic T-cells.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
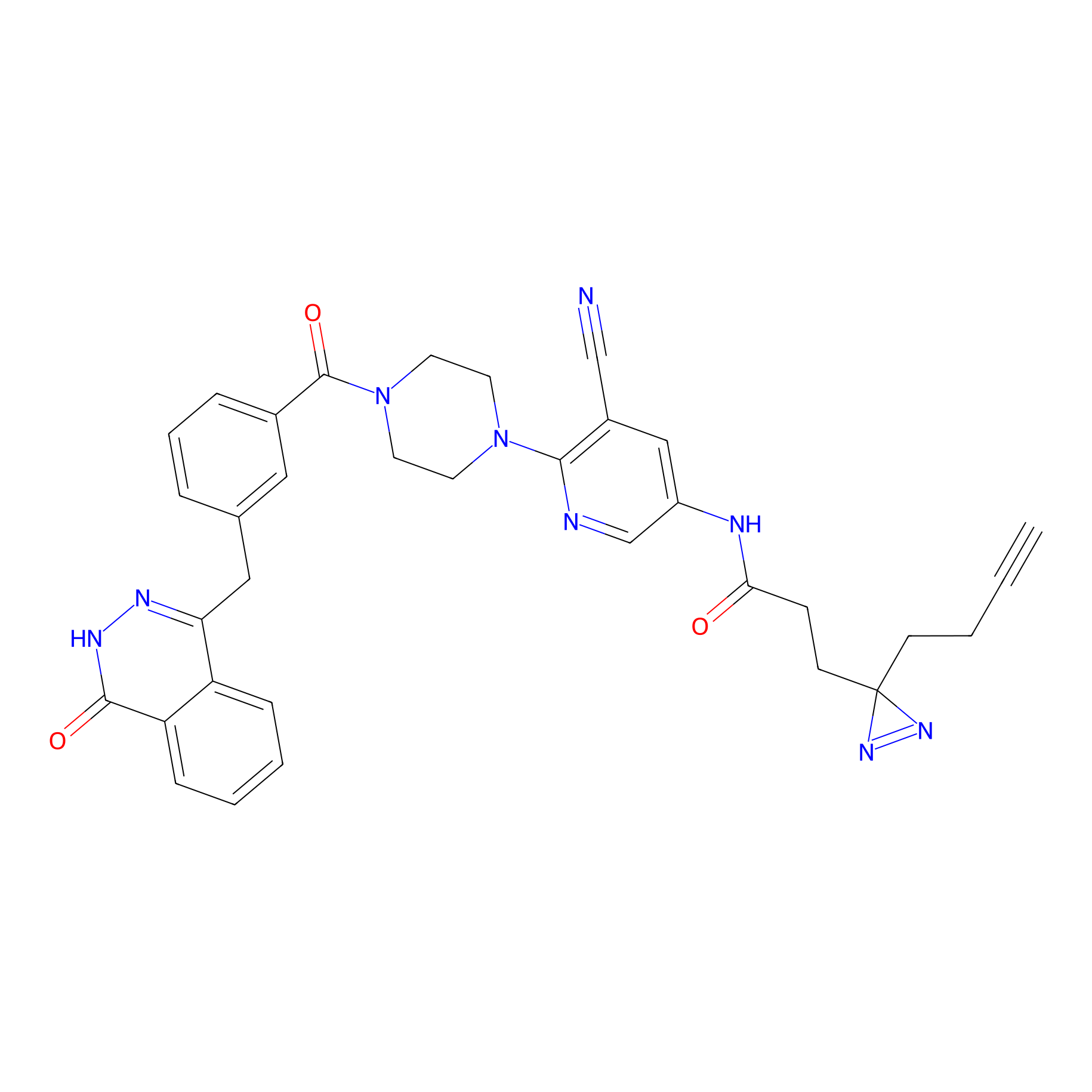
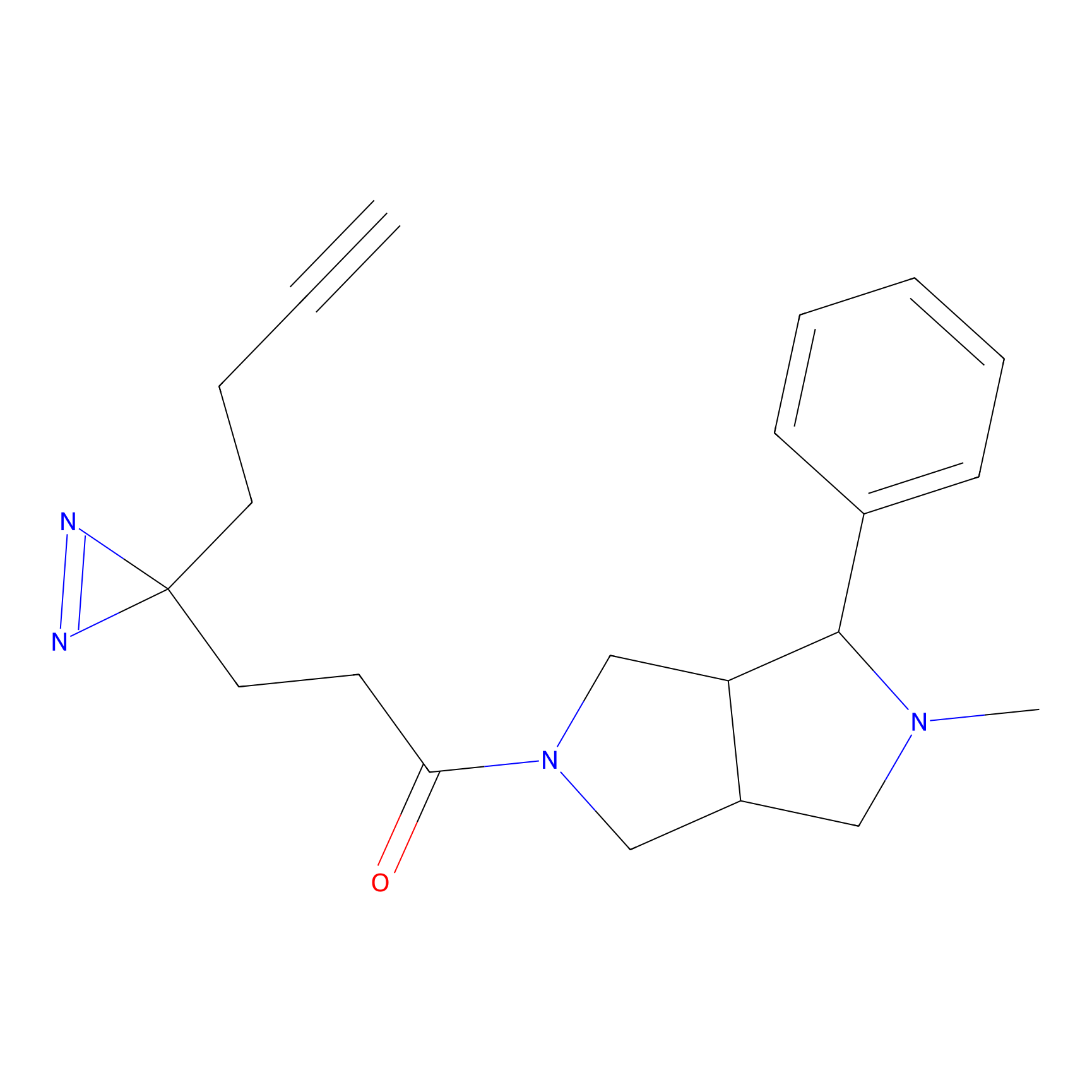
|
FBPP2 Probe Info |
 |
4.54 | LDD0318 | [1] | |
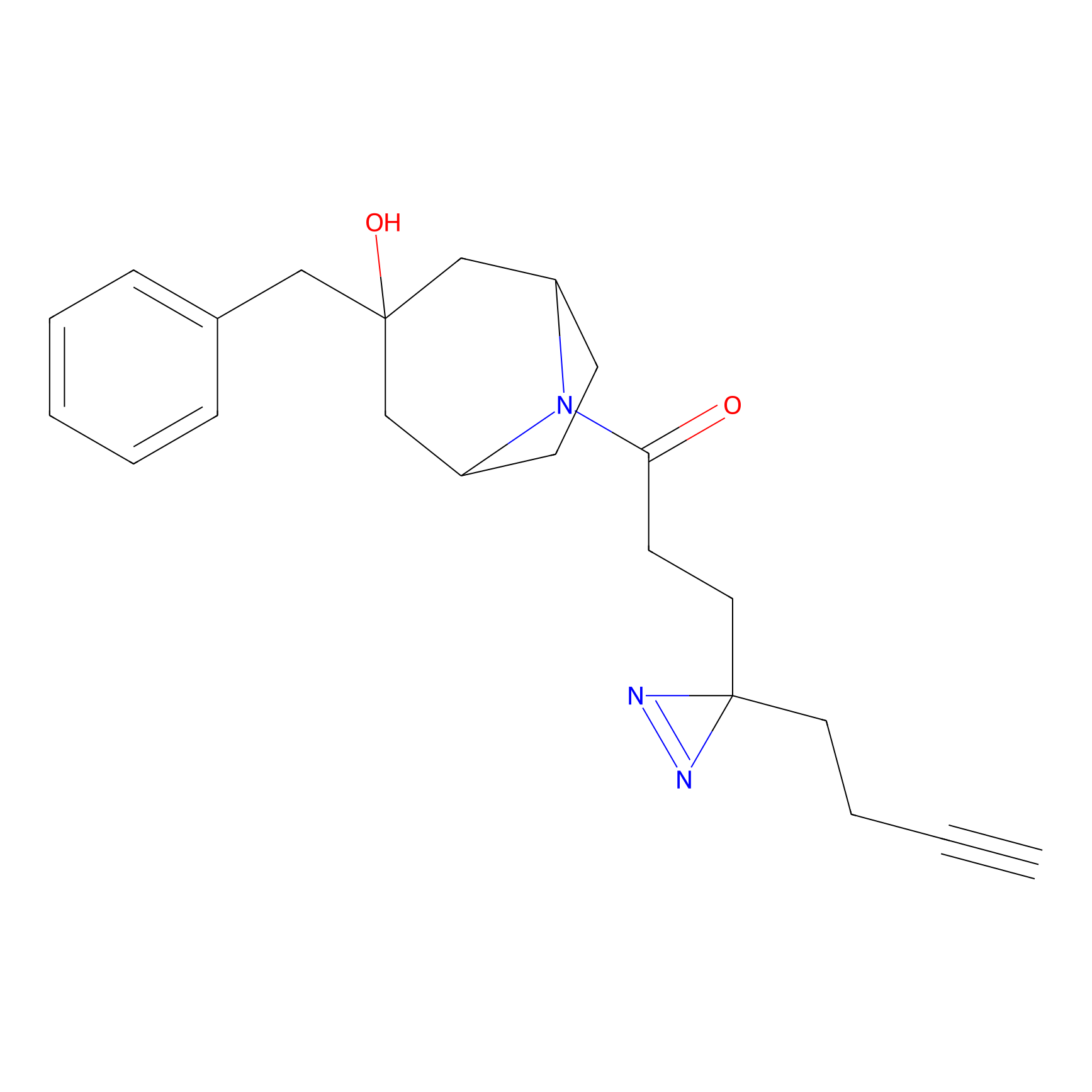
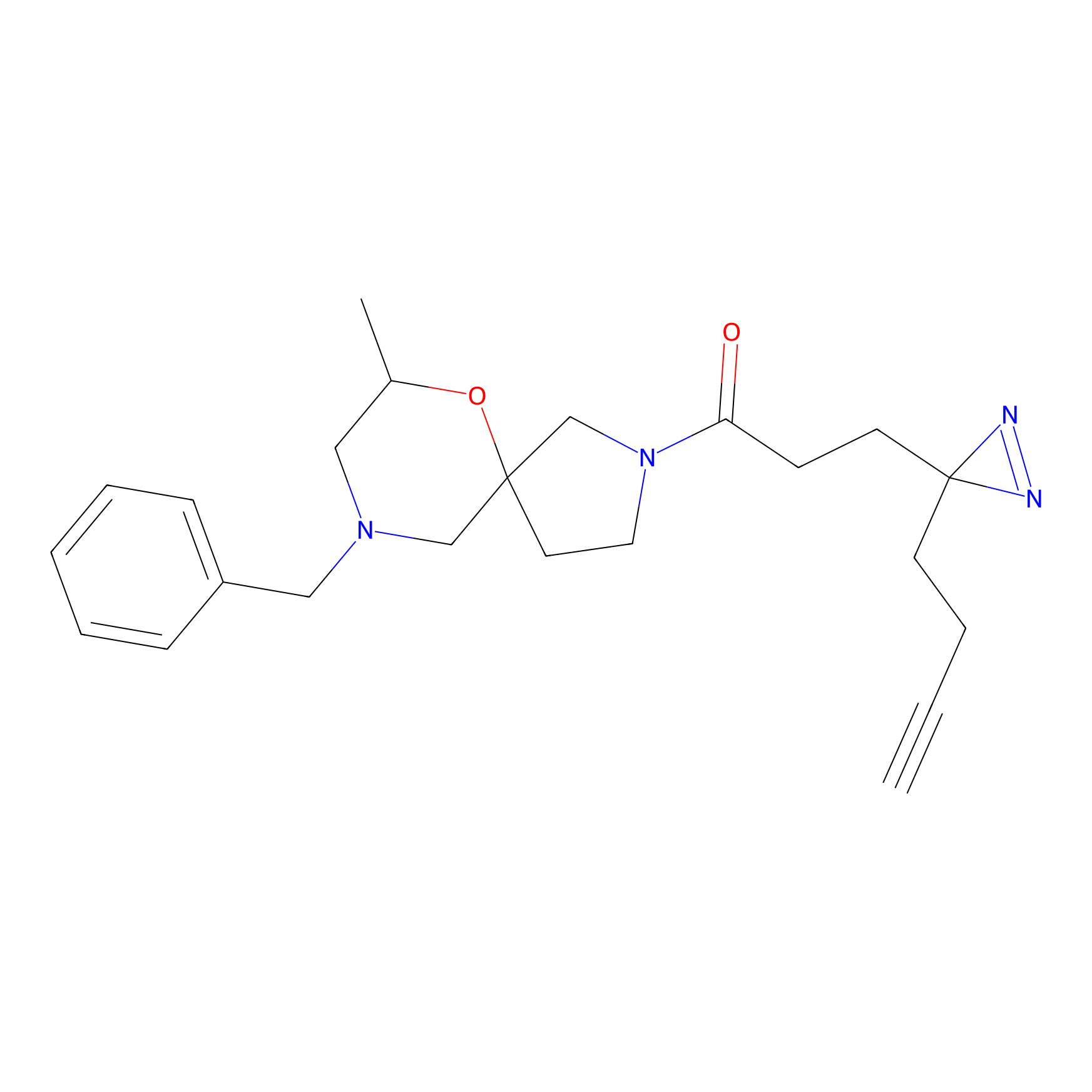
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [2] | |
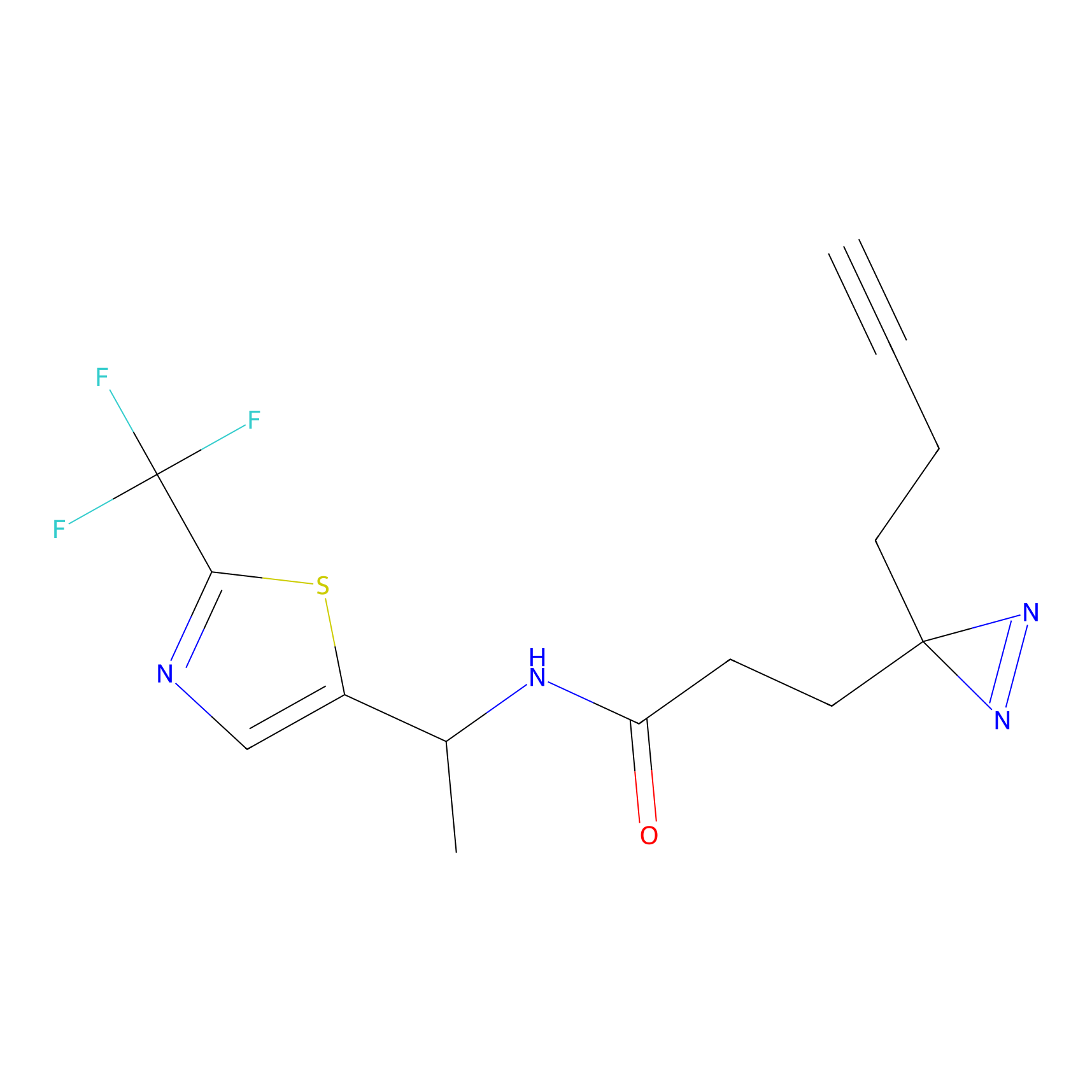
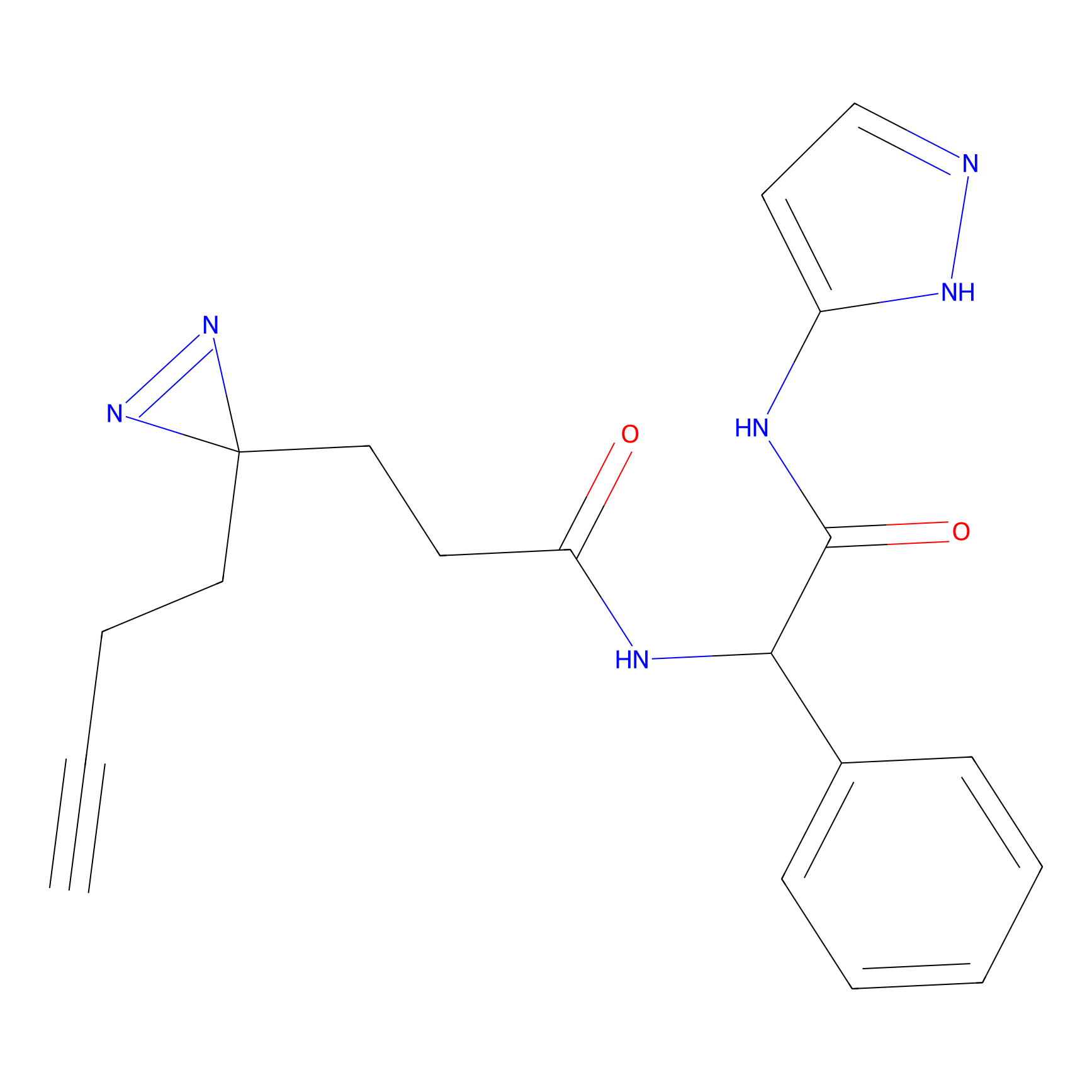
|
STPyne Probe Info |
 |
K113(4.39); K342(8.25) | LDD0277 | [3] | |
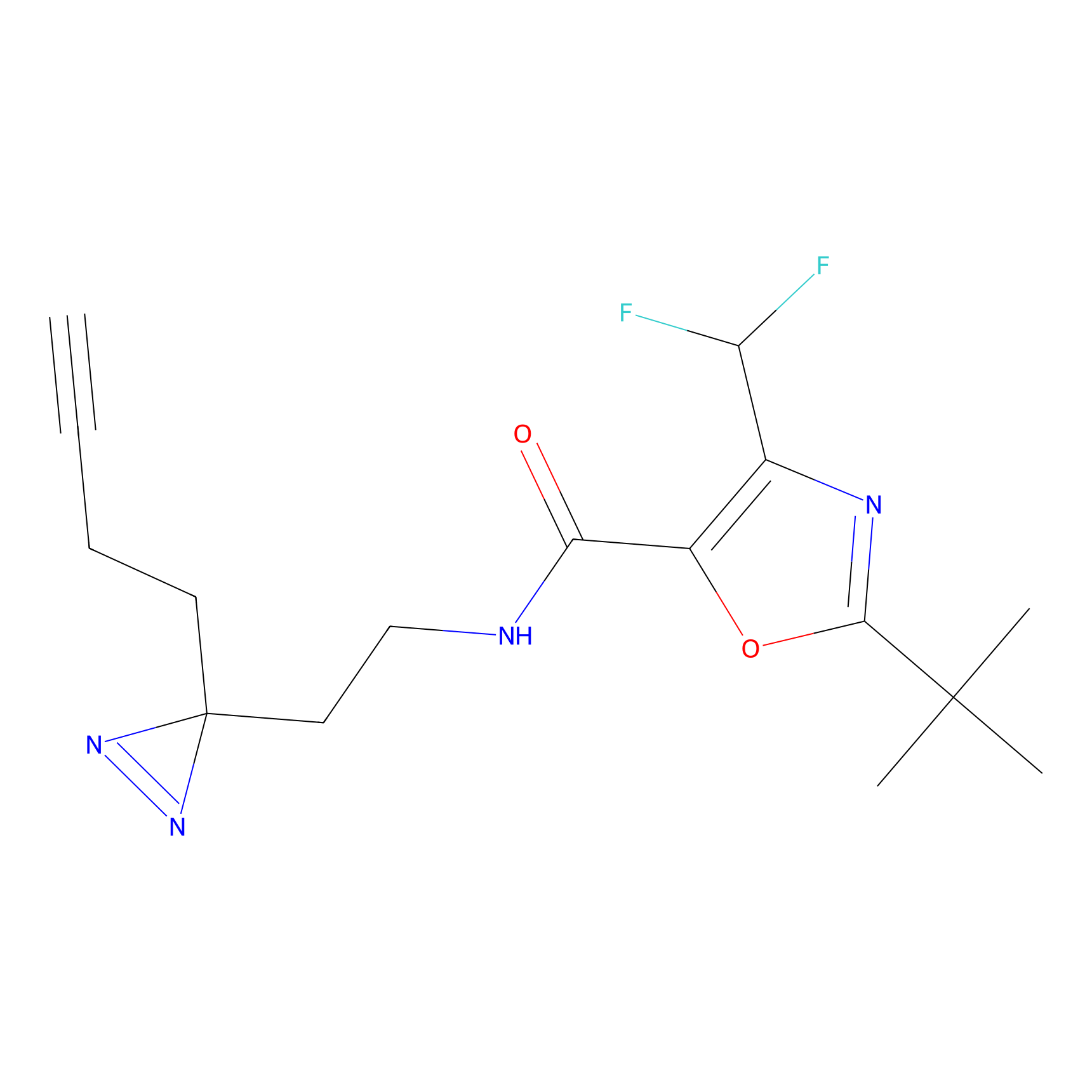
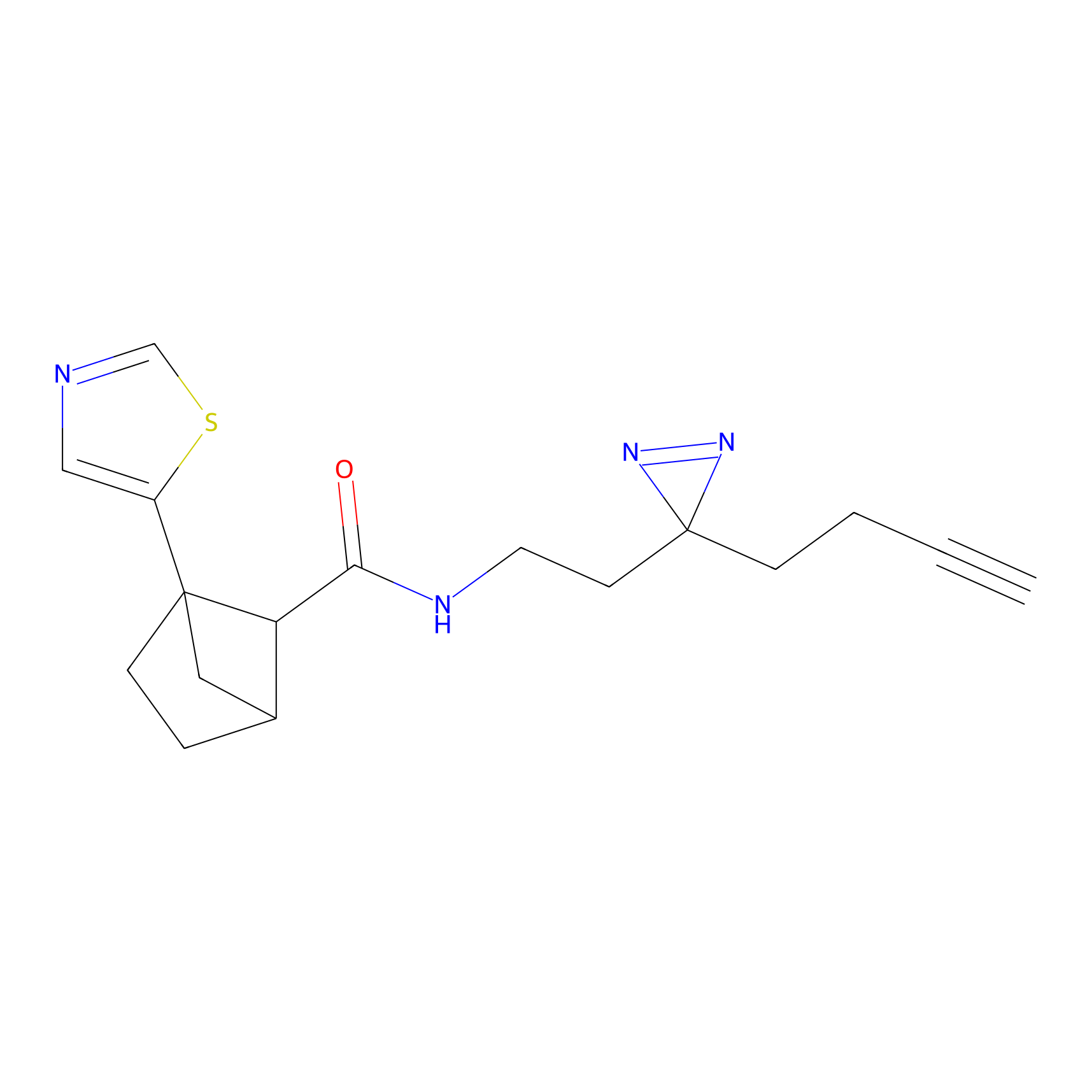
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [4] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [4] |
| LDCM0185 | Compound 17 | HEK-293T | 4.09 | LDD0512 | [7] |
| LDCM0191 | Compound 21 | HEK-293T | 8.20 | LDD0508 | [7] |
| LDCM0190 | Compound 34 | HEK-293T | 13.70 | LDD0510 | [7] |
| LDCM0192 | Compound 35 | HEK-293T | 9.71 | LDD0509 | [7] |
| LDCM0193 | Compound 36 | HEK-293T | 10.26 | LDD0511 | [7] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [4] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [4] |
| LDCM0084 | Ro 48-8071 | A-549 | 5.25 | LDD0145 | [8] |
The Interaction Atlas With This Target
References