Details of the Target
General Information of Target
| Target ID | LDTP14511 | |||||
|---|---|---|---|---|---|---|
| Target Name | ATP synthase subunit g, mitochondrial (ATP5MG) | |||||
| Gene Name | ATP5MG | |||||
| Gene ID | 10632 | |||||
| Synonyms |
ATP5L; ATP synthase subunit g, mitochondrial; ATPase subunit g; ATP synthase membrane subunit g |
|||||
| 3D Structure | ||||||
| Sequence |
MGKDYYKILGIPSGANEDEIKKAYRKMALKYHPDKNKEPNAEEKFKEIAEAYDVLSDPKK
RGLYDQYGEEGLKTGGGTSGGSSGSFHYTFHGDPHATFASFFGGSNPFDIFFASSRSTRP FSGFDPDDMDVDEDEDPFGAFGRFGFNGLSRGPRRAPEPLYPRRKVQDPPVVHELRVSLE EIYHGSTKRMKITRRRLNPDGRTVRTEDKILHIVIKRGWKEGTKITFPKEGDATPDNIPA DIVFVLKDKPHAHFRRDGTNVLYSALISLKEALCGCTVNIPTIDGRVIPLPCNDVIKPGT VKRLRGEGLPFPKVPTQRGDLIVEFKVRFPDRLTPQTRQILKQHLPCS |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
ATPase g subunit family
|
|||||
| Subcellular location |
Mitochondrion
|
|||||
| Function |
Mitochondrial membrane ATP synthase (F(1)F(0) ATP synthase or Complex V) produces ATP from ADP in the presence of a proton gradient across the membrane which is generated by electron transport complexes of the respiratory chain. F-type ATPases consist of two structural domains, F(1) - containing the extramembraneous catalytic core, and F(0) - containing the membrane proton channel, linked together by a central stalk and a peripheral stalk. During catalysis, ATP synthesis in the catalytic domain of F(1) is coupled via a rotary mechanism of the central stalk subunits to proton translocation. Part of the complex F(0) domain. Minor subunit located with subunit a in the membrane.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
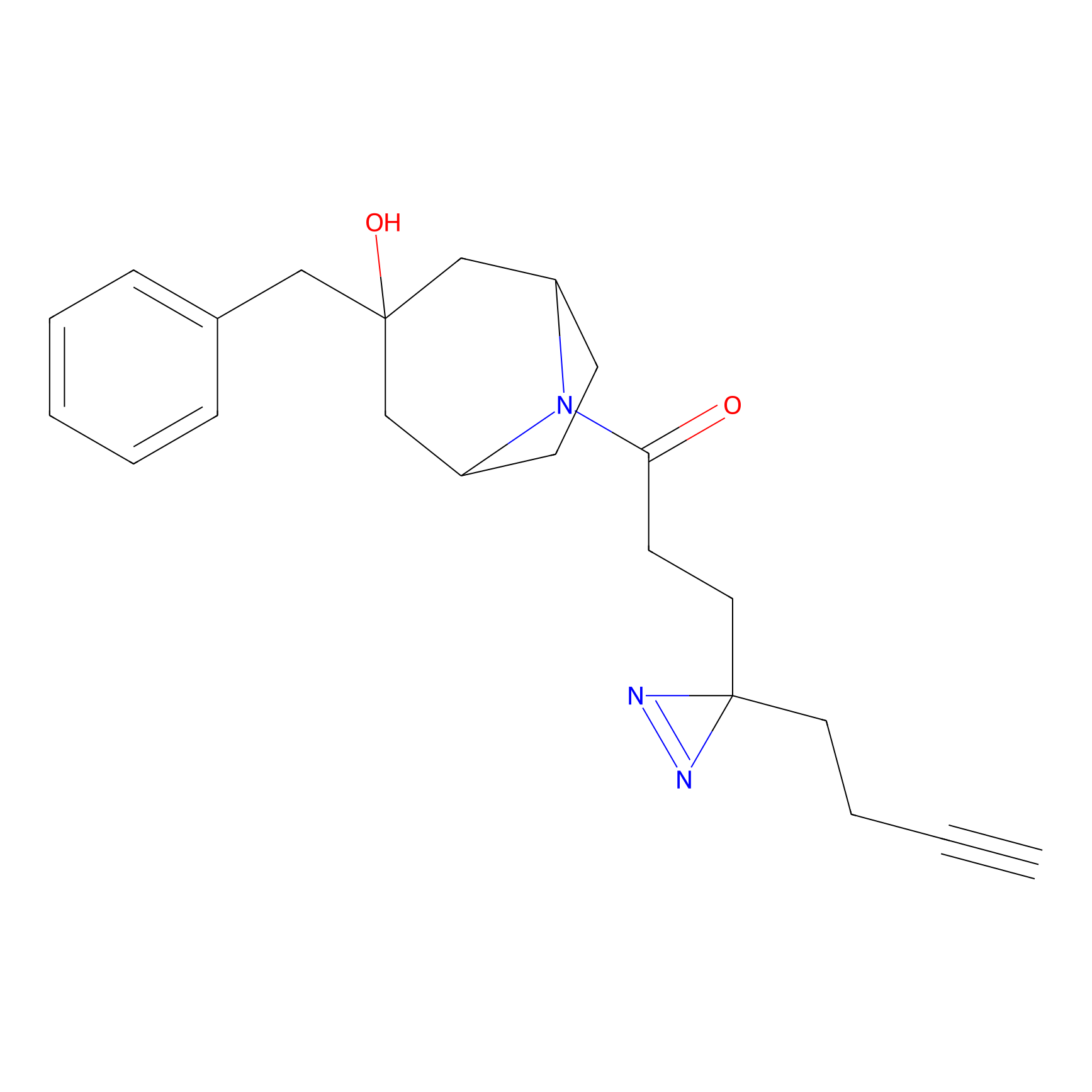
TH211 Probe Info |
 |
Y22(16.42) | LDD0260 | [1] | |
|
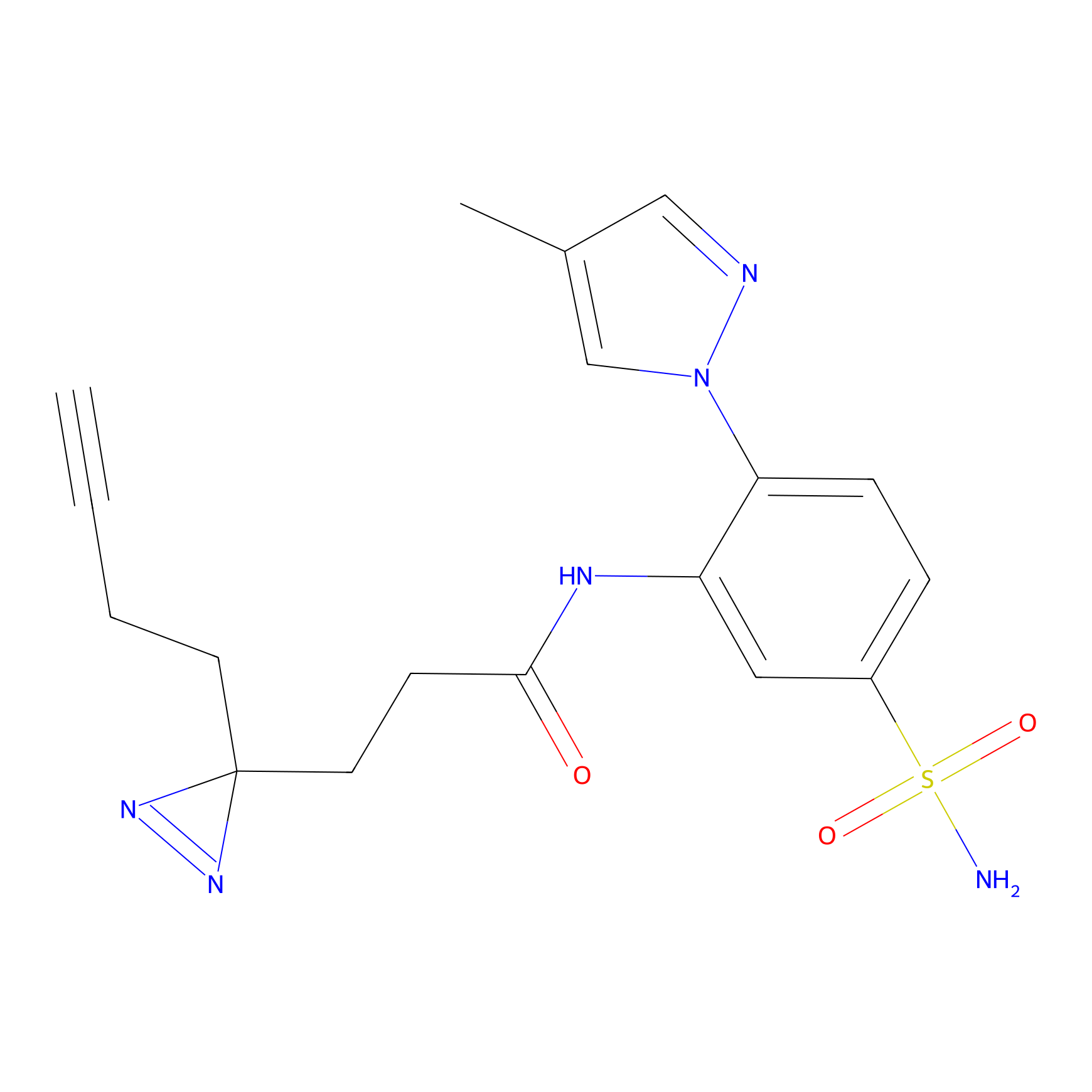
ATP probe Probe Info |
 |
N.A. | LDD0199 | [2] | |
|
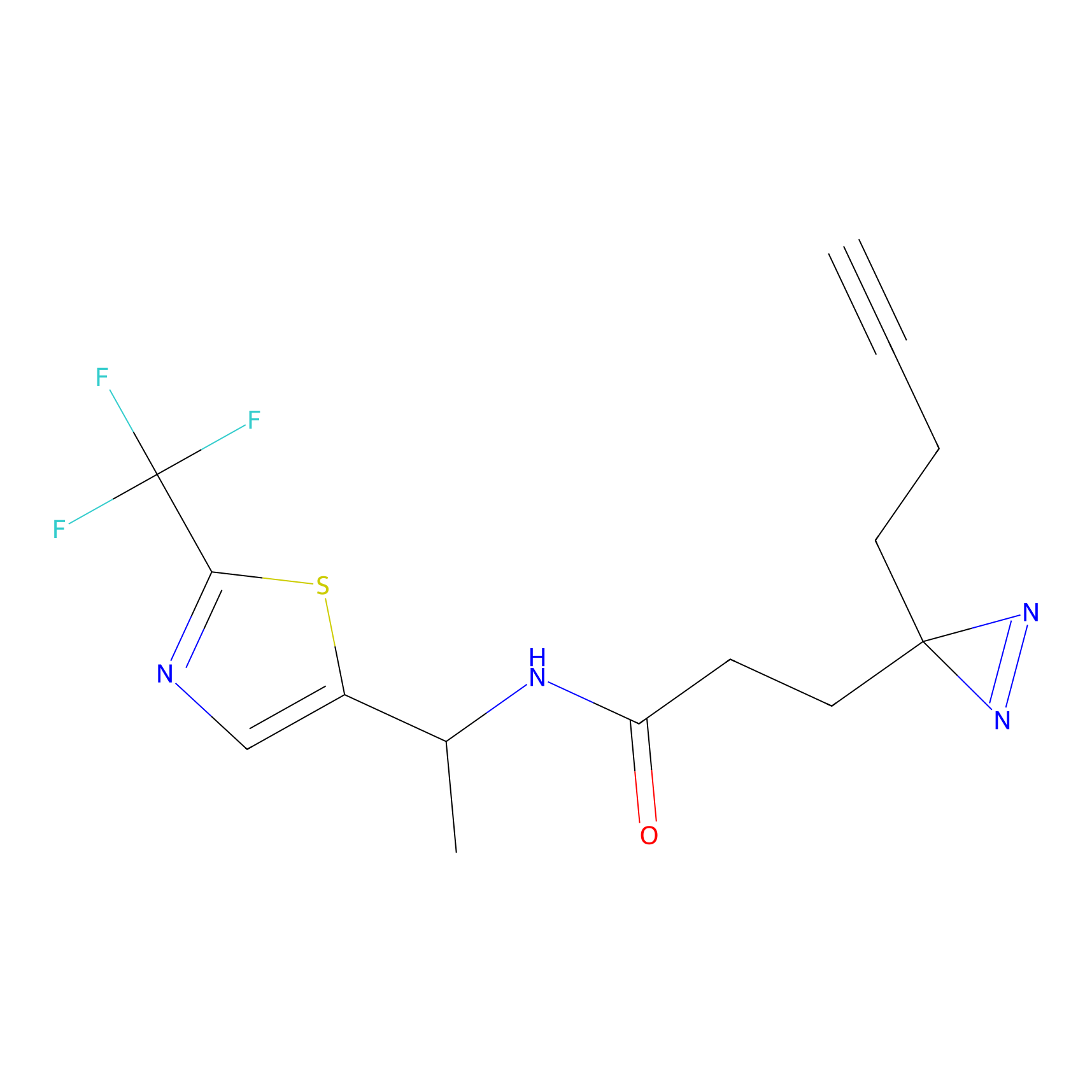
1d-yne Probe Info |
 |
N.A. | LDD0356 | [3] | |
|
NHS Probe Info |
 |
K11(0.00); K55(0.00); K24(0.00); K66(0.00) | LDD0010 | [4] | |
|
SF Probe Info |
 |
Y22(0.00); Y32(0.00) | LDD0028 | [5] | |
|
STPyne Probe Info |
 |
K66(0.00); K24(0.00) | LDD0009 | [4] | |
|
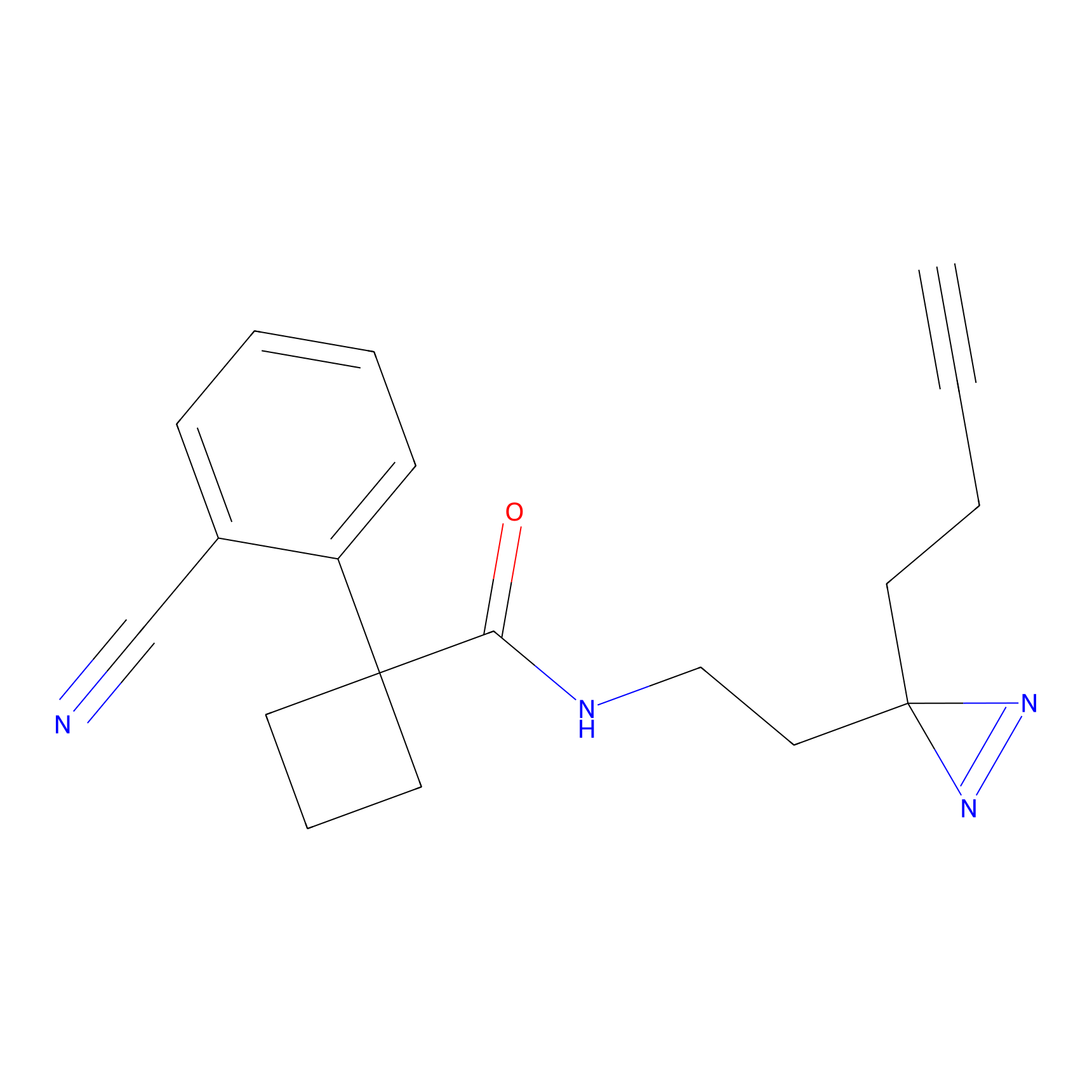
1c-yne Probe Info |
 |
K66(0.00); K11(0.00) | LDD0228 | [3] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References