Details of the Target
General Information of Target
| Target ID | LDTP10958 | |||||
|---|---|---|---|---|---|---|
| Target Name | Sigma non-opioid intracellular receptor 1 (SIGMAR1) | |||||
| Gene Name | SIGMAR1 | |||||
| Gene ID | 10280 | |||||
| Synonyms |
OPRS1; SRBP; Sigma non-opioid intracellular receptor 1; Aging-associated gene 8 protein; SR31747-binding protein; SR-BP; Sigma 1-type opioid receptor; SIG-1R; Sigma1-receptor; Sigma1R; hSigmaR1 |
|||||
| 3D Structure | ||||||
| Sequence |
MSTGLRYKSKLATPEDKQDIDKQYVGFATLPNQVHRKSVKKGFDFTLMVAGESGLGKSTL
VHSLFLTDLYKDRKLLSAEERISQTVEILKHTVDIEEKGVKLKLTIVDTPGFGDAVNNTE CWKPITDYVDQQFEQYFRDESGLNRKNIQDNRVHCCLYFISPFGHGLRPVDVGFMKALHE KVNIVPLIAKADCLVPSEIRKLKERIREEIDKFGIHVYQFPECDSDEDEDFKQQDRELKE SAPFAVIGSNTVVEAKGQRVRGRLYPWGIVEVENQAHCDFVKLRNMLIRTHMHDLKDVTC DVHYENYRAHCIQQMTSKLTQDSRMESPIPILPLPTPDAETEKLIRMKDEELRRMQEMLQ RMKQQMQDQ |
|||||
| Target Type |
Successful
|
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
ERG2 family
|
|||||
| Subcellular location |
Nucleus inner membrane
|
|||||
| Function |
Functions in lipid transport from the endoplasmic reticulum and is involved in a wide array of cellular functions probably through regulation of the biogenesis of lipid microdomains at the plasma membrane. Involved in the regulation of different receptors it plays a role in BDNF signaling and EGF signaling. Also regulates ion channels like the potassium channel and could modulate neurotransmitter release. Plays a role in calcium signaling through modulation together with ANK2 of the ITP3R-dependent calcium efflux at the endoplasmic reticulum. Plays a role in several other cell functions including proliferation, survival and death. Originally identified for its ability to bind various psychoactive drugs it is involved in learning processes, memory and mood alteration. Necessary for proper mitochondrial axonal transport in motor neurons, in particular the retrograde movement of mitochondria. Plays a role in protecting cells against oxidative stress-induced cell death via its interaction with RNF112.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
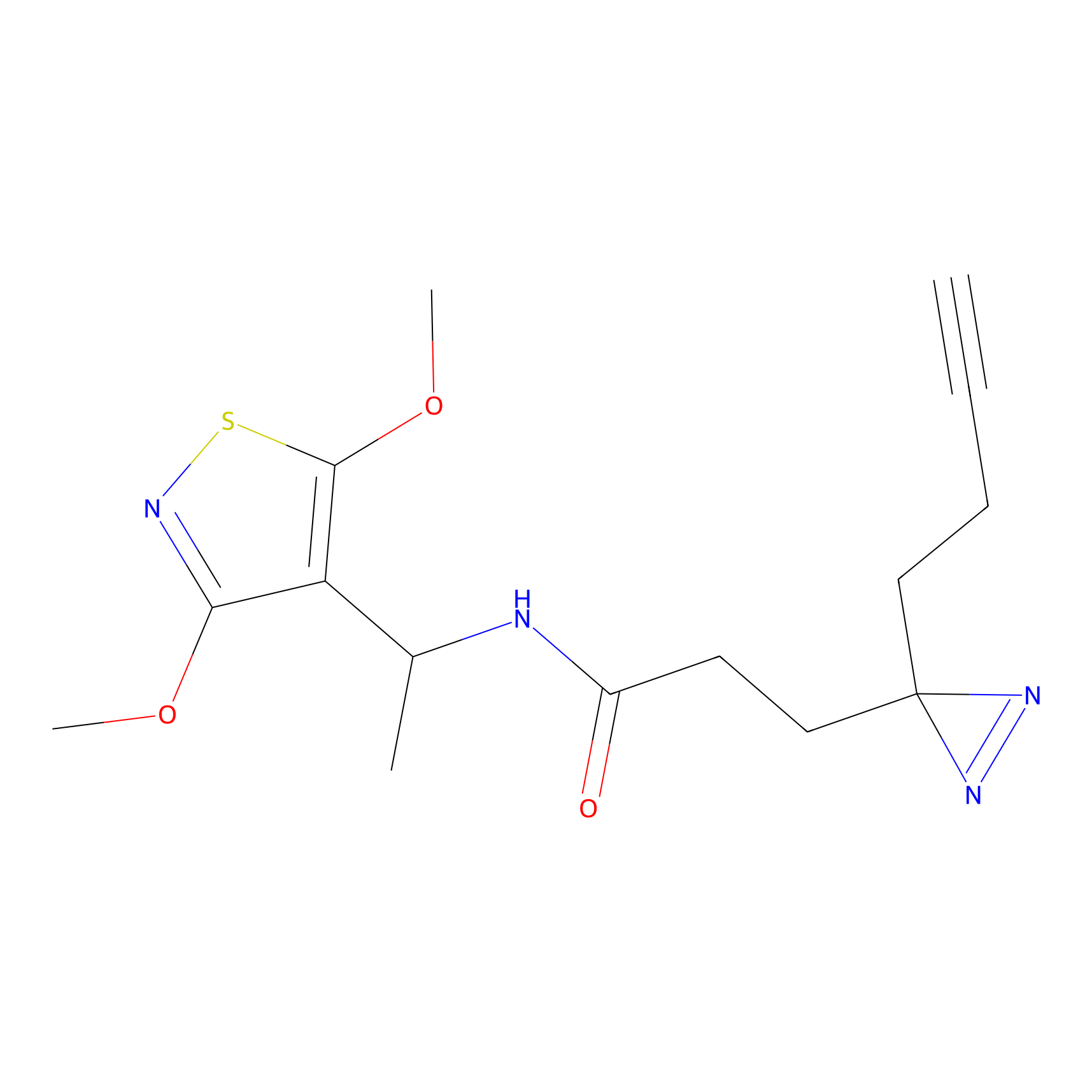
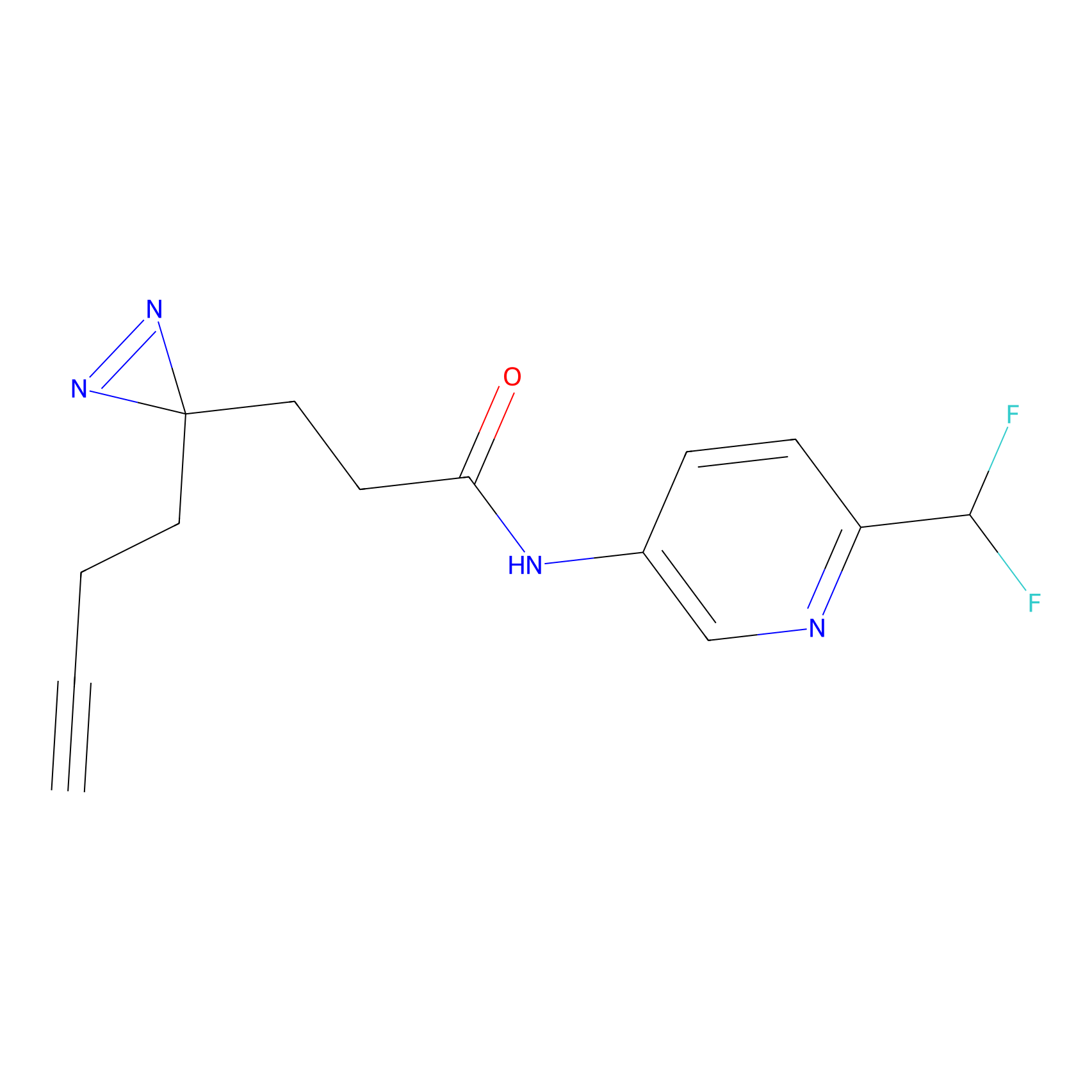
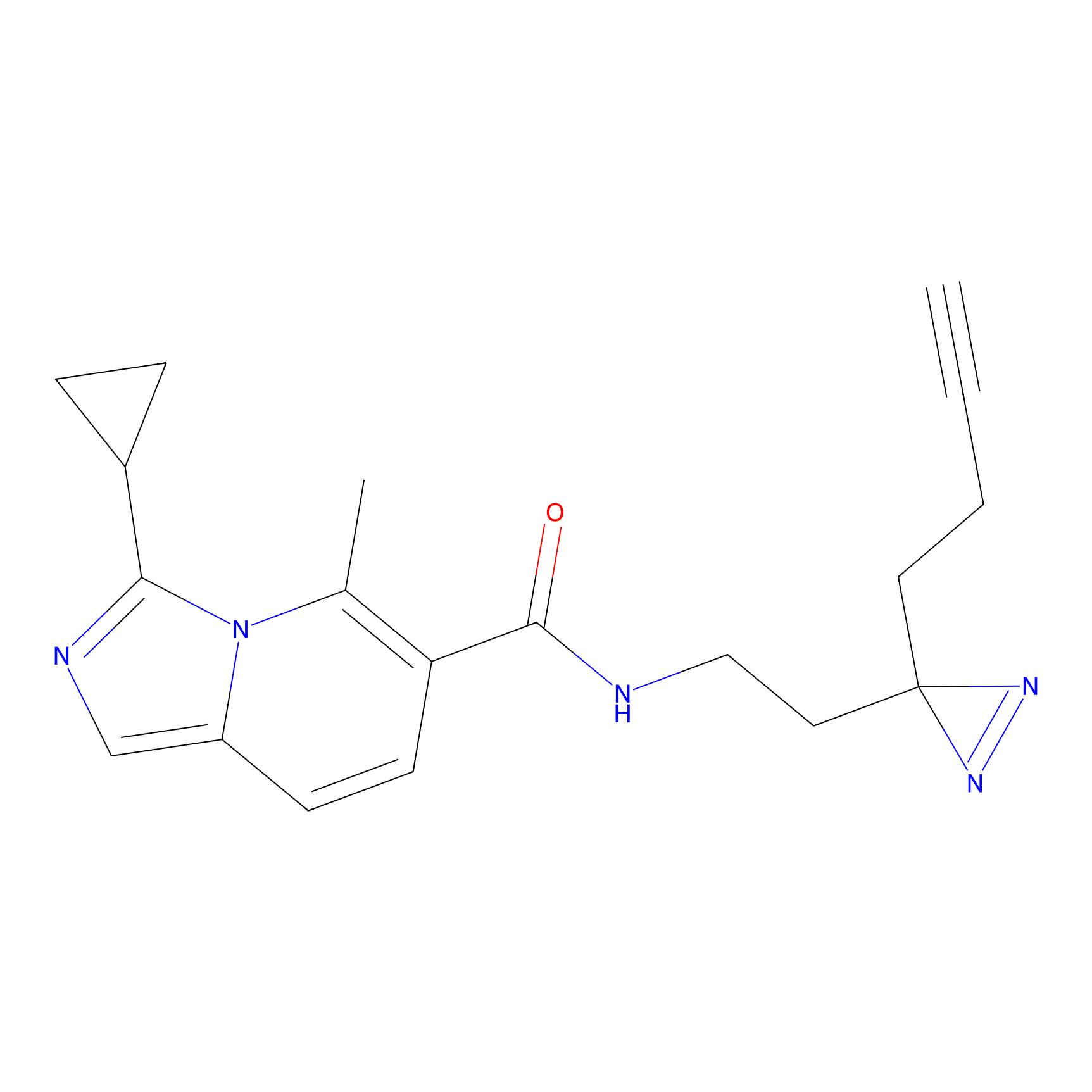
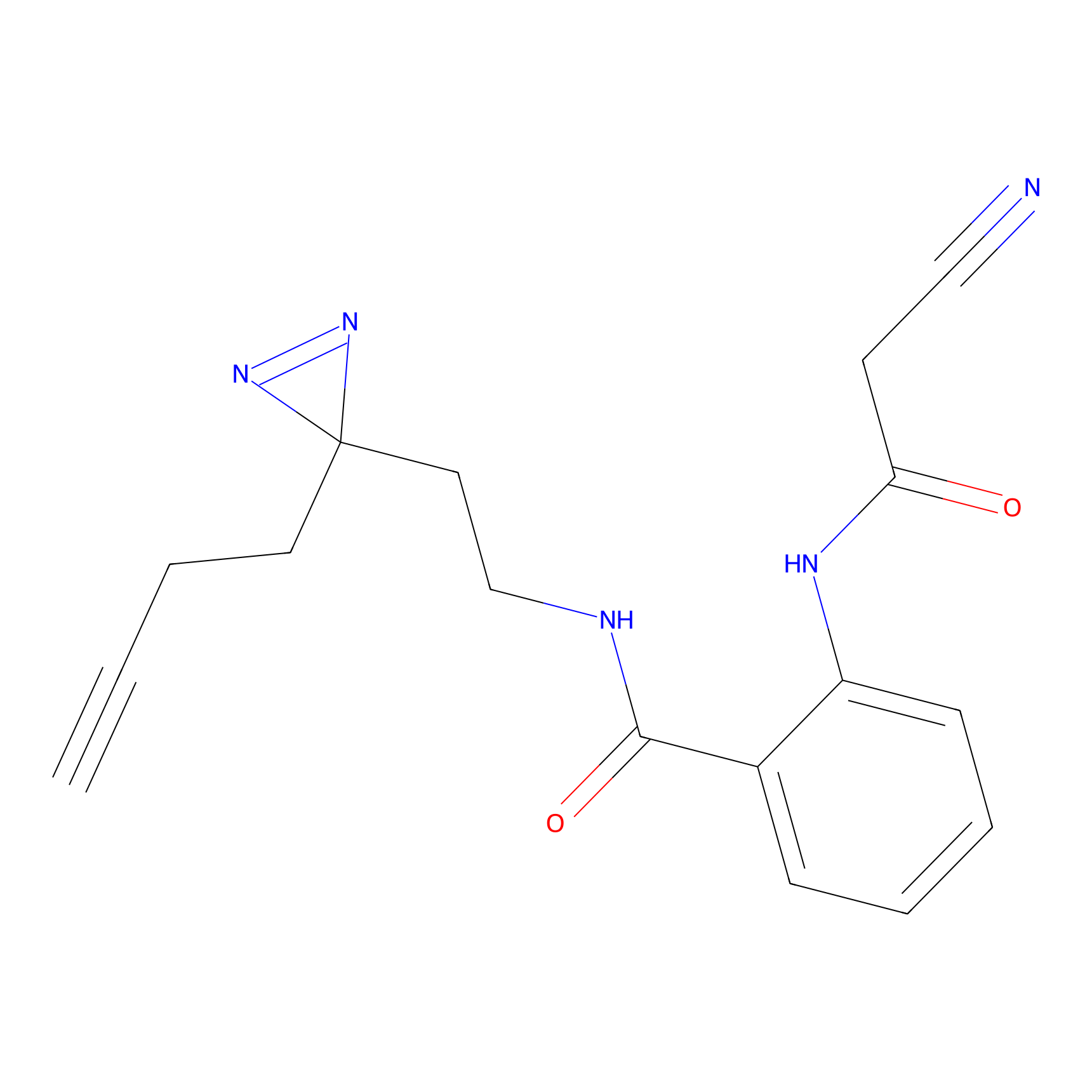
Jackson_1 Probe Info |
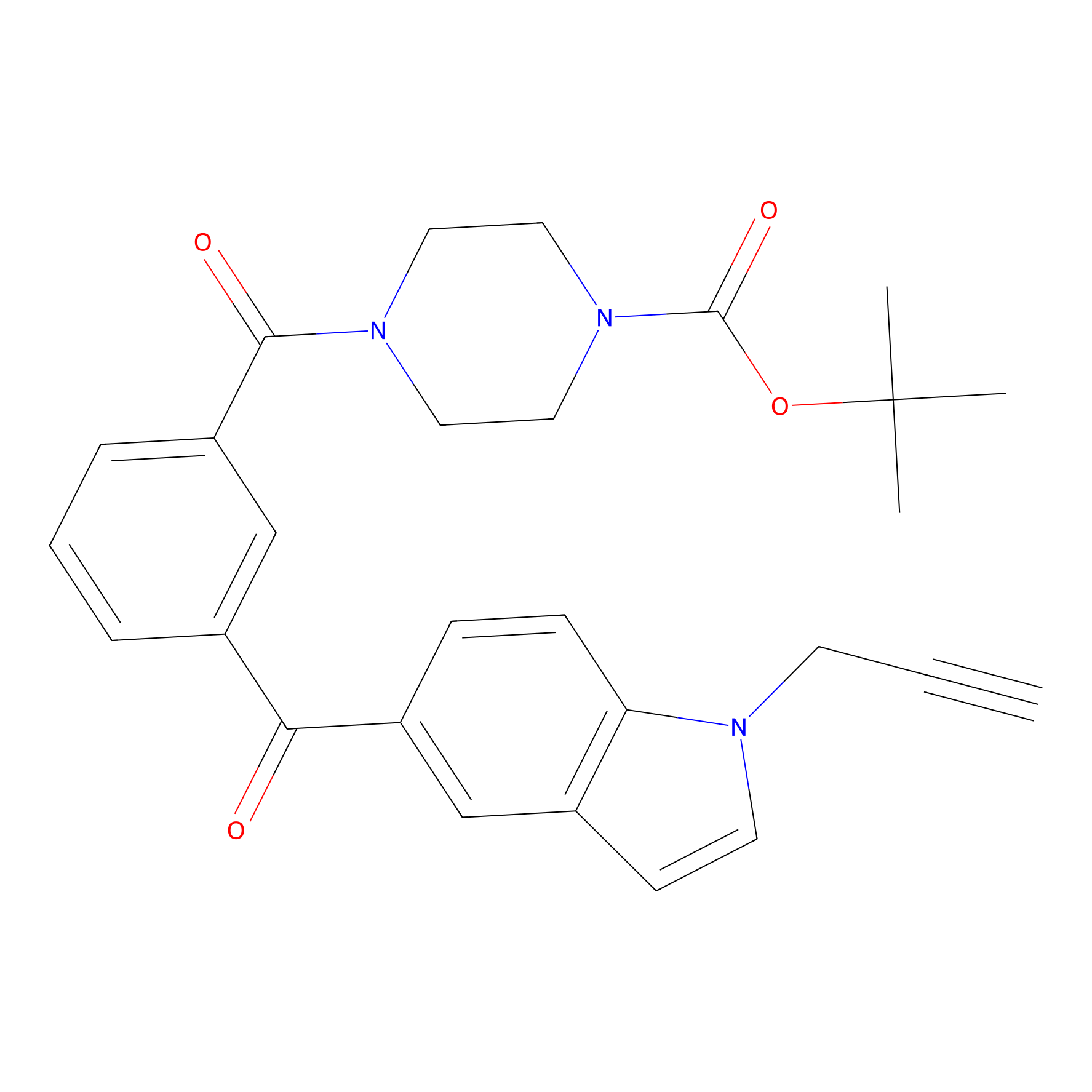 |
2.46 | LDD0120 | [1] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Serpin H1 (SERPINH1) | Serpin family | P50454 | |||
The Drug(s) Related To This Target
Approved
Phase 3
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Adx N05 | . | D0GW6K | |||
| Avp-786 | . | D0Z6IB | |||
| Cm-2395 | . | D0X6XH | |||
Phase 2
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Igmesine | Small molecular drug | D0N5LJ | |||
| Opc-14523 | Small molecular drug | D0R8GR | |||
Phase 1
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Ssr-125047 | Small molecular drug | D0Q7QY | |||
| Anavex 2-73 | . | D09AOO | |||
| Sa-5845 | . | D0G9SH | |||
| Ssr-125329a | . | D0T6ES | |||
Preclinical
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Cutamesine | Small molecular drug | D00KVE | |||
| Anavex 1-41 | . | D0T0UC | |||
| Anavex 1007 | . | D0M6OW | |||
Investigative
Patented
Discontinued
References