Details of the Target
General Information of Target
| Target ID | LDTP10329 | |||||
|---|---|---|---|---|---|---|
| Target Name | Ceramide synthase 2 (CERS2) | |||||
| Gene Name | CERS2 | |||||
| Gene ID | 29956 | |||||
| Synonyms |
LASS2; TMSG1; Ceramide synthase 2; CerS2; LAG1 longevity assurance homolog 2; SP260; Sphingosine N-acyltransferase CERS2; EC 2.3.1.24; Tumor metastasis-suppressor gene 1 protein; Very-long-chain ceramide synthase CERS2; EC 2.3.1.297
|
|||||
| 3D Structure | ||||||
| Sequence |
MLRREARLRREYLYRKAREEAQRSAQERKERLRRALEENRLIPTELRREALALQGSLEFD
DAGGEGVTSHVDDEYRWAGVEDPKVMITTSRDPSSRLKMFAKELKLVFPGAQRMNRGRHE VGALVRACKANGVTDLLVVHEHRGTPVGLIVSHLPFGPTAYFTLCNVVMRHDIPDLGTMS EAKPHLITHGFSSRLGKRVSDILRYLFPVPKDDSHRVITFANQDDYISFRHHVYKKTDHR NVELTEVGPRFELKLYMIRLGTLEQEATADVEWRWHPYTNTARKRVFLSTE |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Ceramide synthase that catalyzes the transfer of the acyl chain from acyl-CoA to a sphingoid base, with high selectivity toward very-long-chain fatty acyl-CoA (chain length C22-C27). N-acylates sphinganine and sphingosine bases to form dihydroceramides and ceramides in de novo synthesis and salvage pathways, respectively. Plays a non-redundant role in the synthesis of ceramides with very-long-chain fatty acids in kidney, liver and brain. Regulates the abundance of myelin-specific sphingolipids galactosylceramide and sulfatide that affects myelin sheath architecture and motor neuron functions.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
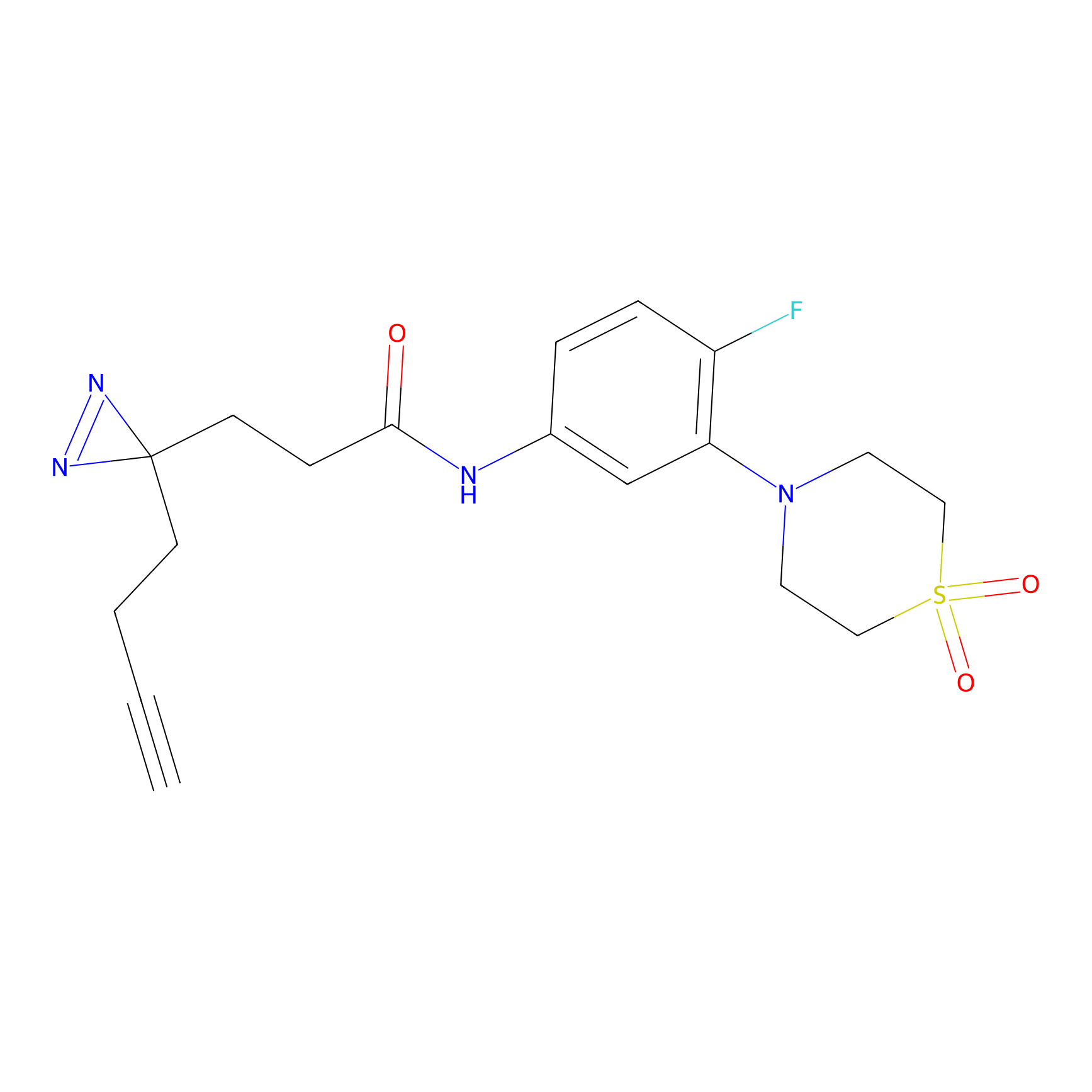
CHEMBL5175495 Probe Info |
 |
5.81 | LDD0196 | [1] | |
|
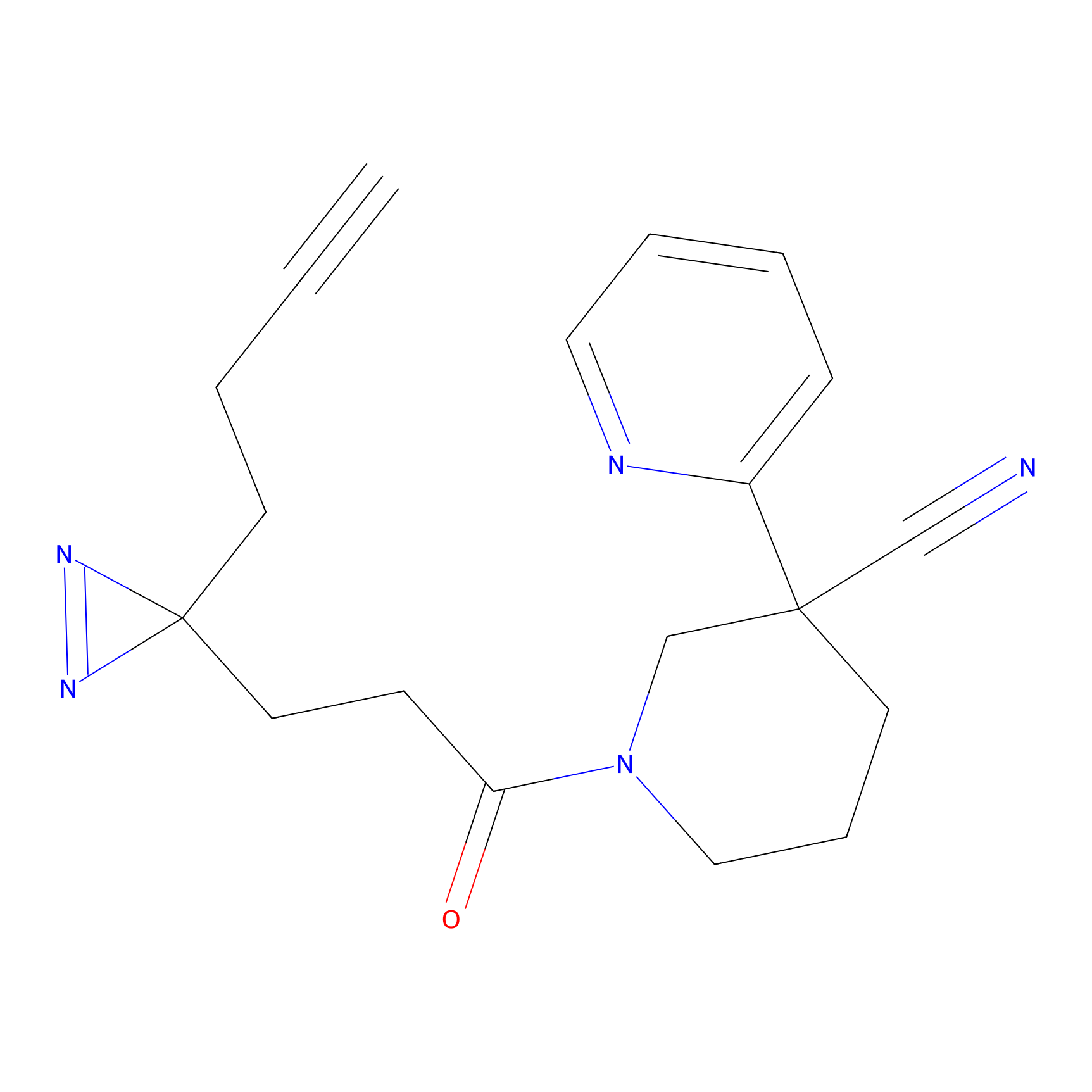
CY4 Probe Info |
 |
100.00 | LDD0244 | [2] | |
|
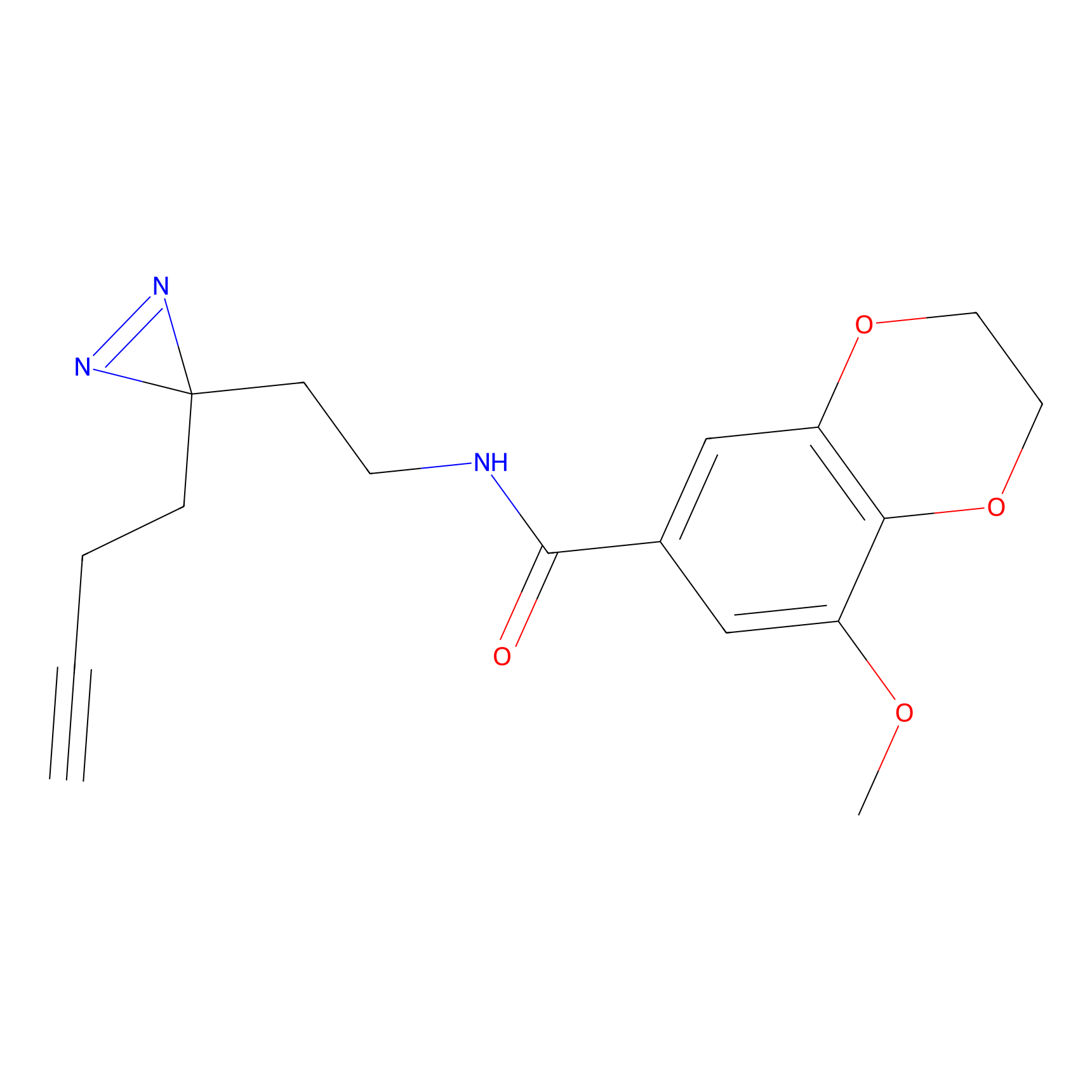
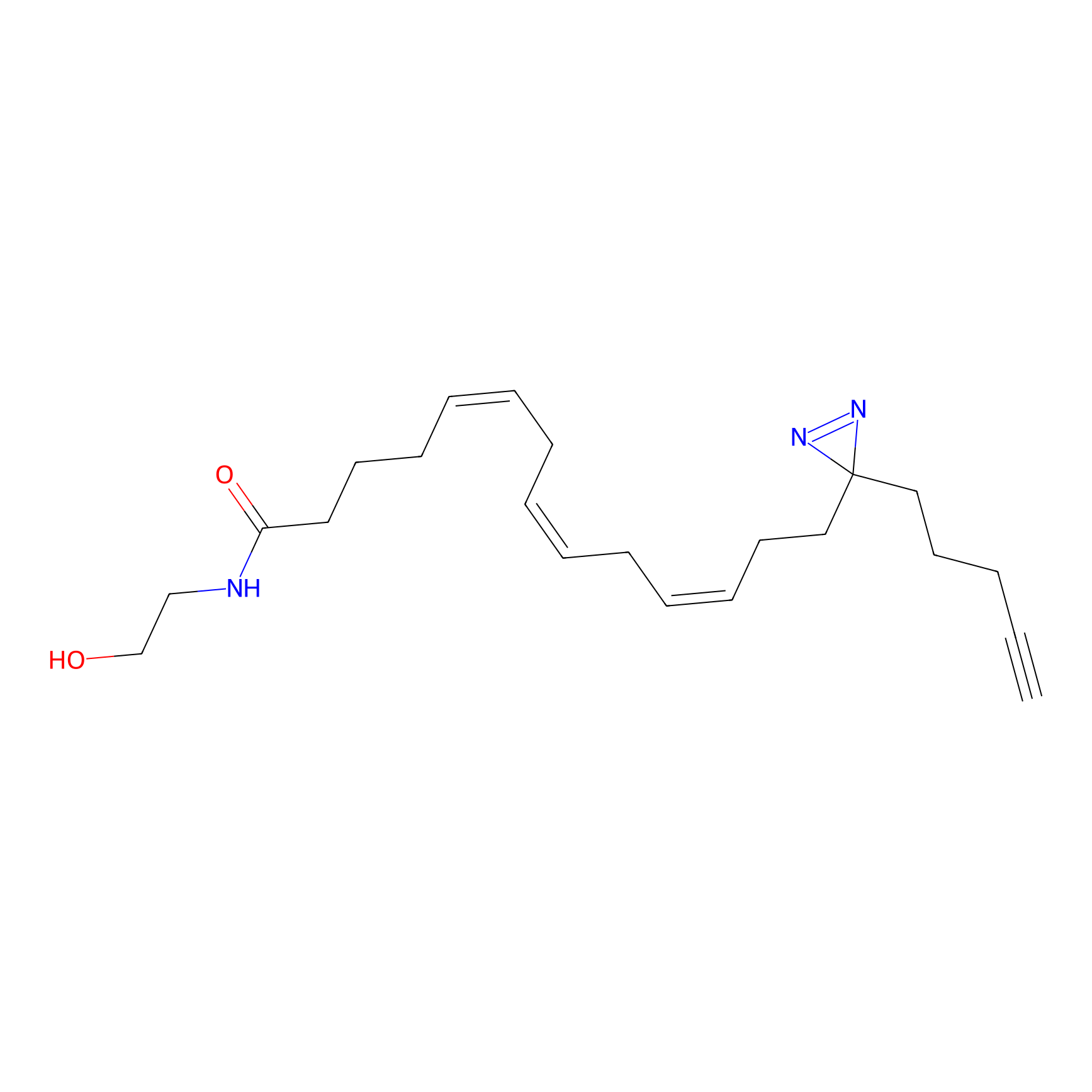
TH211 Probe Info |
 |
Y88(9.41) | LDD0260 | [3] | |
|
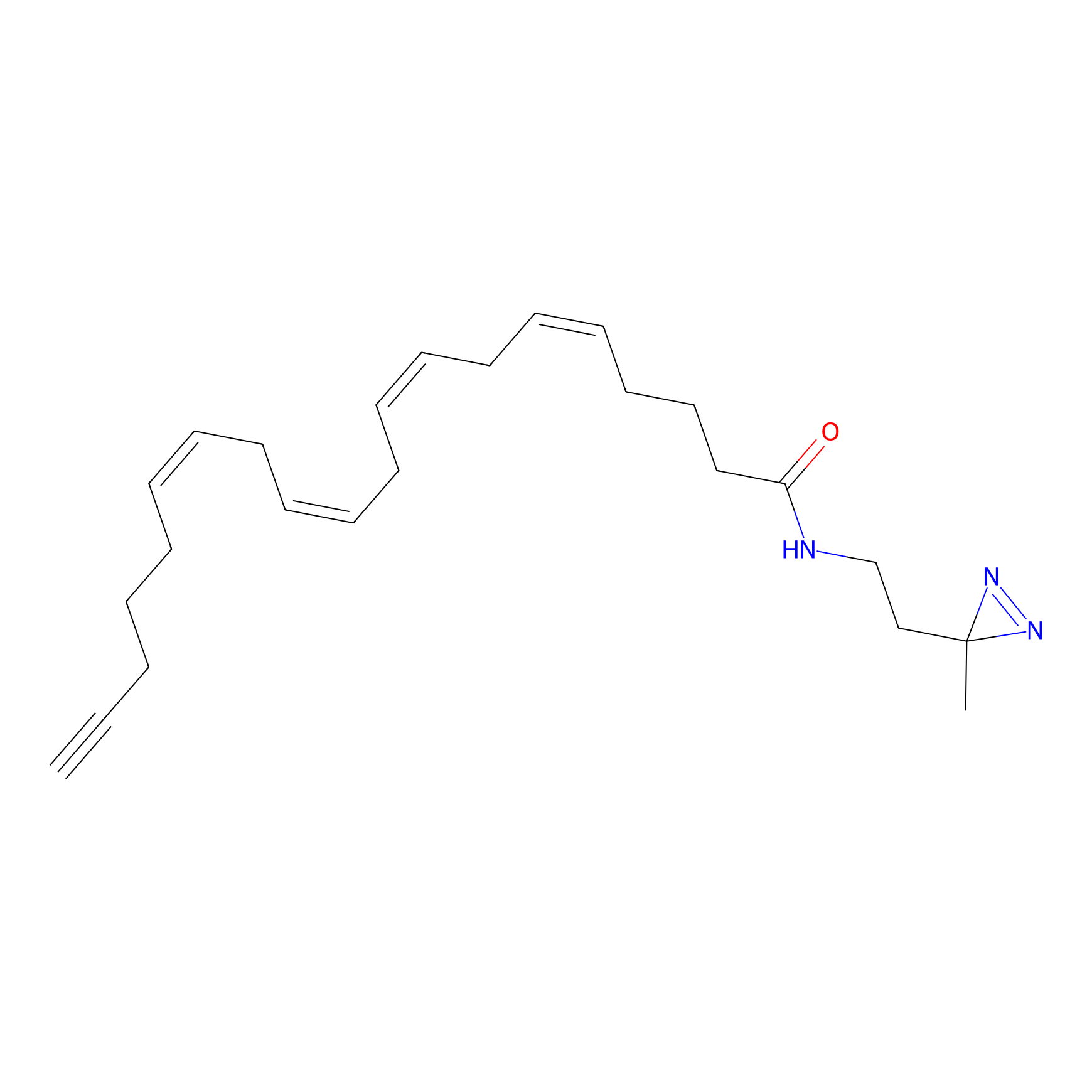
STPyne Probe Info |
 |
K131(10.00); K93(1.98) | LDD0277 | [4] | |
|
Jackson_14 Probe Info |
 |
2.33 | LDD0123 | [5] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [6] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [7] | |
|
AOyne Probe Info |
 |
14.50 | LDD0443 | [8] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [7] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [7] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [7] |
| LDCM0016 | Ranjitkar_cp1 | MDA-MB-231 | 2.33 | LDD0123 | [5] |
| LDCM0084 | Ro 48-8071 | A-549 | 2.52 | LDD0145 | [12] |
| LDCM0136 | SR-4559 | HEK-293T | 4.99 | LDD0333 | [13] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Solute carrier family 22 member 1 (SLC22A1) | Organic cation transporter (TC 2.A.1.19) family | O15245 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nuclear factor erythroid 2-related factor 2 (NFE2L2) | BZIP family | Q16236 | |||
Other
References