Details of the Target
General Information of Target
| Target ID | LDTP06945 | |||||
|---|---|---|---|---|---|---|
| Target Name | Sigma intracellular receptor 2 (TMEM97) | |||||
| Gene Name | TMEM97 | |||||
| Gene ID | 27346 | |||||
| Synonyms |
MAC30; S2R; Sigma intracellular receptor 2; S2R; Sigma-2 receptor; Sigma2 receptor; Meningioma-associated protein 30; MAC30; Transmembrane protein 97 |
|||||
| 3D Structure | ||||||
| Sequence |
MGAPATRRCVEWLLGLYFLSHIPITLFMDLQAVLPRELYPVEFRNLLKWYAKEFKDPLLQ
EPPAWFKSFLFCELVFQLPFFPIATYAFLKGSCKWIRTPAIIYSVHTMTTLIPILSTFLF EDFSKASGFKGQRPETLHERLTLVSVYAPYLLIPFILLIFMLRSPYYKYEEKRKKK |
|||||
| Target Type |
Clinical trial
|
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
TMEM97/sigma-2 receptor family
|
|||||
| Subcellular location |
Nucleus membrane
|
|||||
| Function |
Sigma-2 receptor which contributes to ameliorate dysfunctional cellular processes and slow degenerative progression by regulating cell functions including cholesterol biosynthesis/trafficking, membrane trafficking, autophagy, lipid membrane-bound protein trafficking, and receptor stabilization at the cell surface (Probable). Forms a ternary complex with PGRMC1 receptor and low density lipoprotein receptor/LDLR at the plasma membrane, which increases LDLR-mediated LDL cholesterol internalization. Decreases lysosomal sterol transporter NPC1 availability to the cell, probably through NPC1-binding, hence controlling lipid transport, including cholesterol and LBPA, outside of late endosome/lysosome. Binds regio- and stereoselective ligand 20(S)-hydroxycholesterol (20(S)-OHC) which enhances TMEM97-NPC1 interaction and decreases TMEM97-PGRMC1 and TMEM97-TSPO interactions, thereby linking OHC binding to cholesterol homeostasis. Also able to bind cholesterol. Binds histatin 1 (Hst 1)/HN1 salivary peptide at the ER membrane, which is critical for increasing mitochondria-ER contacts and stimulating Hst1 wound healing properties. May alter the activity of some cytochrome P450 proteins. Although shows homologies with sterol isomerases (EXPERA domain), not able to catalyze sterol isomerization (Probable). However, may act as sensors of these molecules (Probable). Acts as a quality control factor in the ER, promoting the proteolytic degradation of nonproductive and extramitochondrial precursor proteins in the ER membrane thus removing them from the ER surface.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
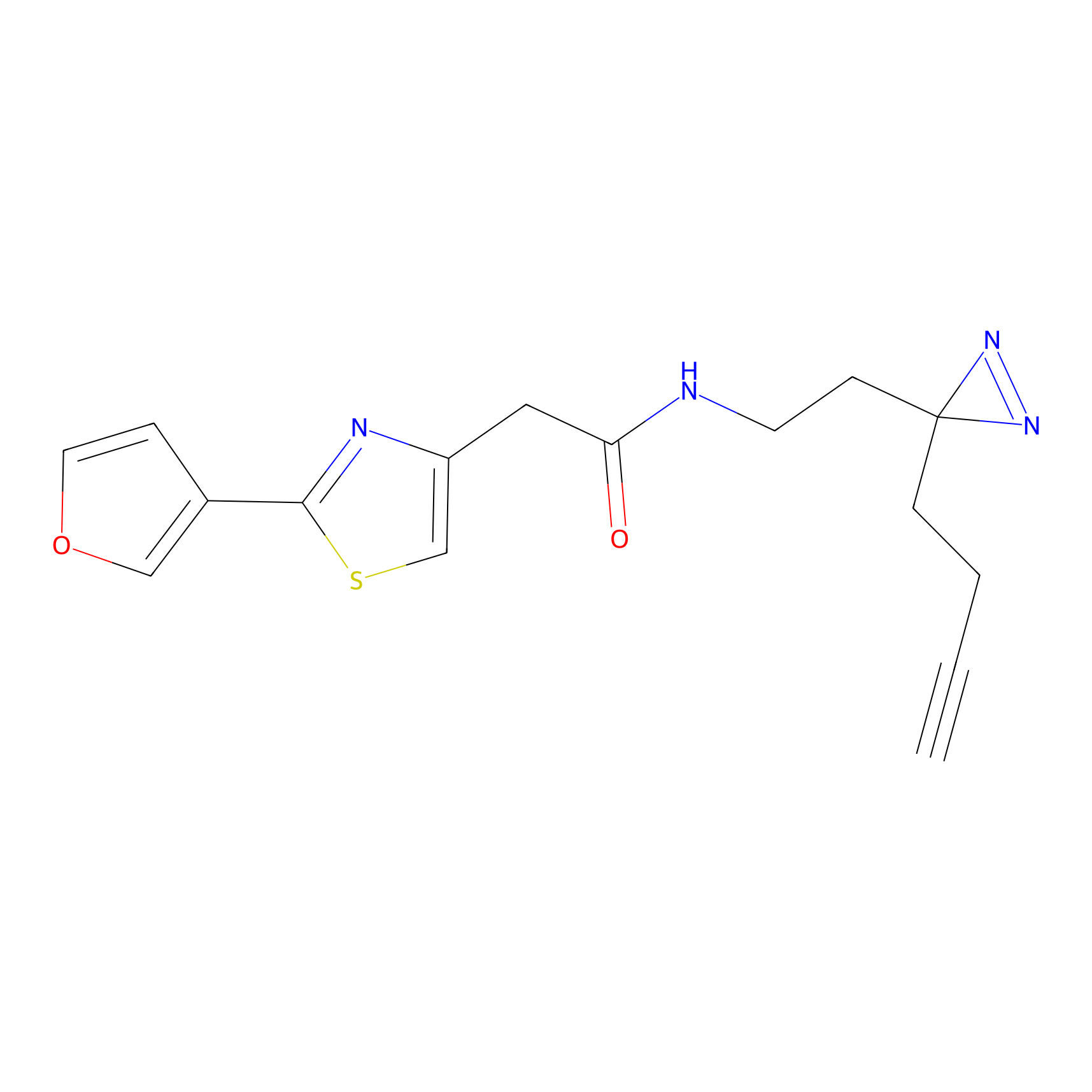
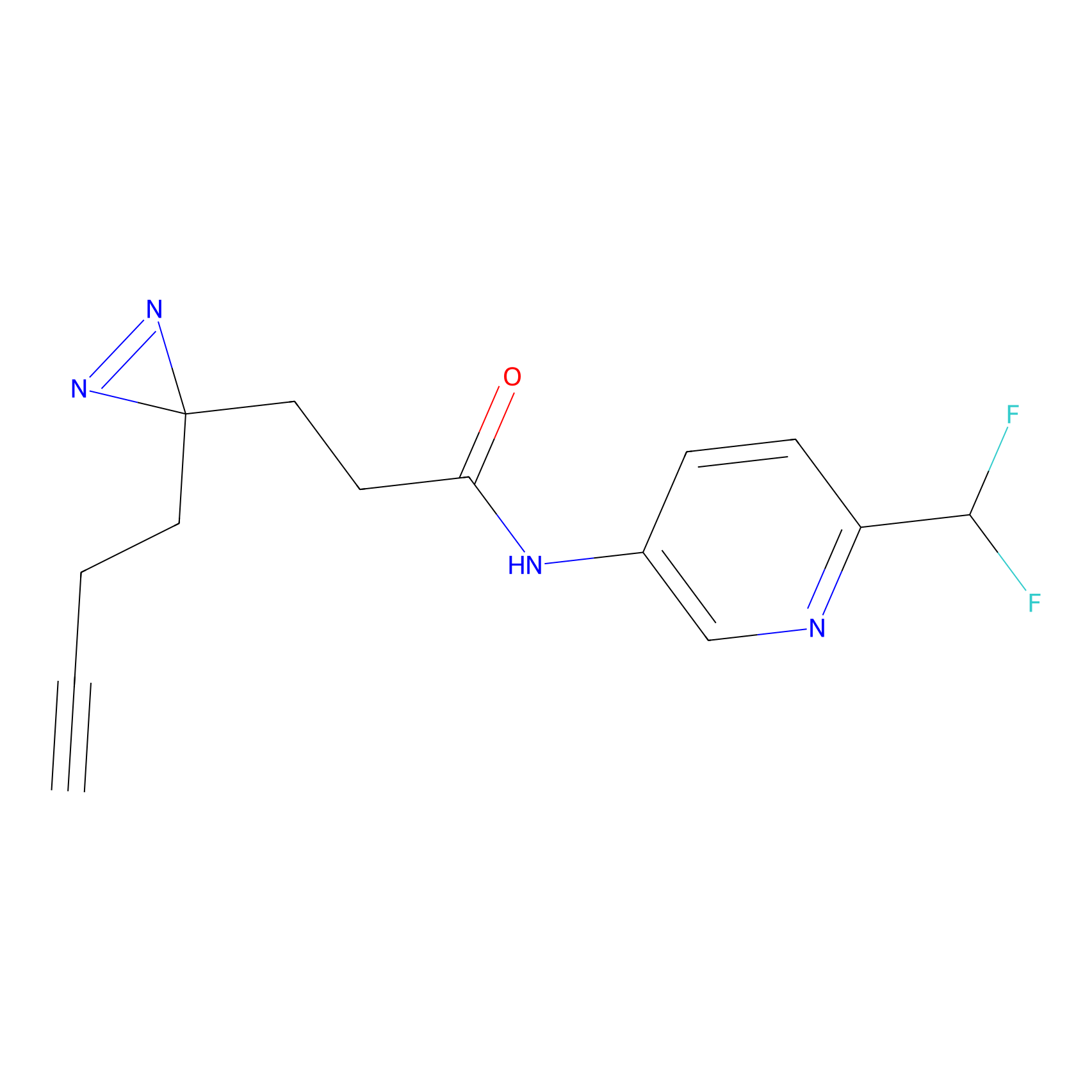
YN-1 Probe Info |
 |
100.00 | LDD0444 | [1] | |
|
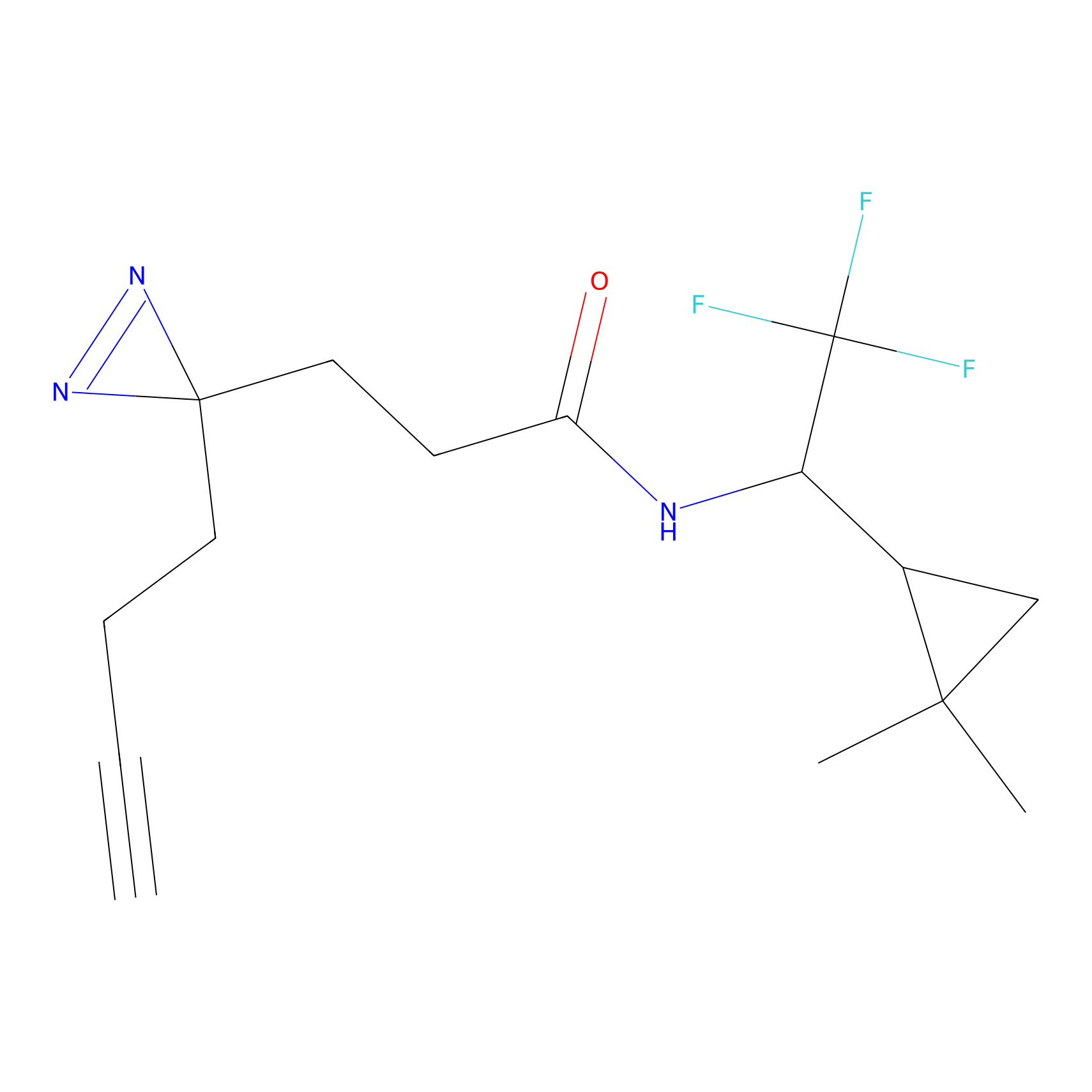
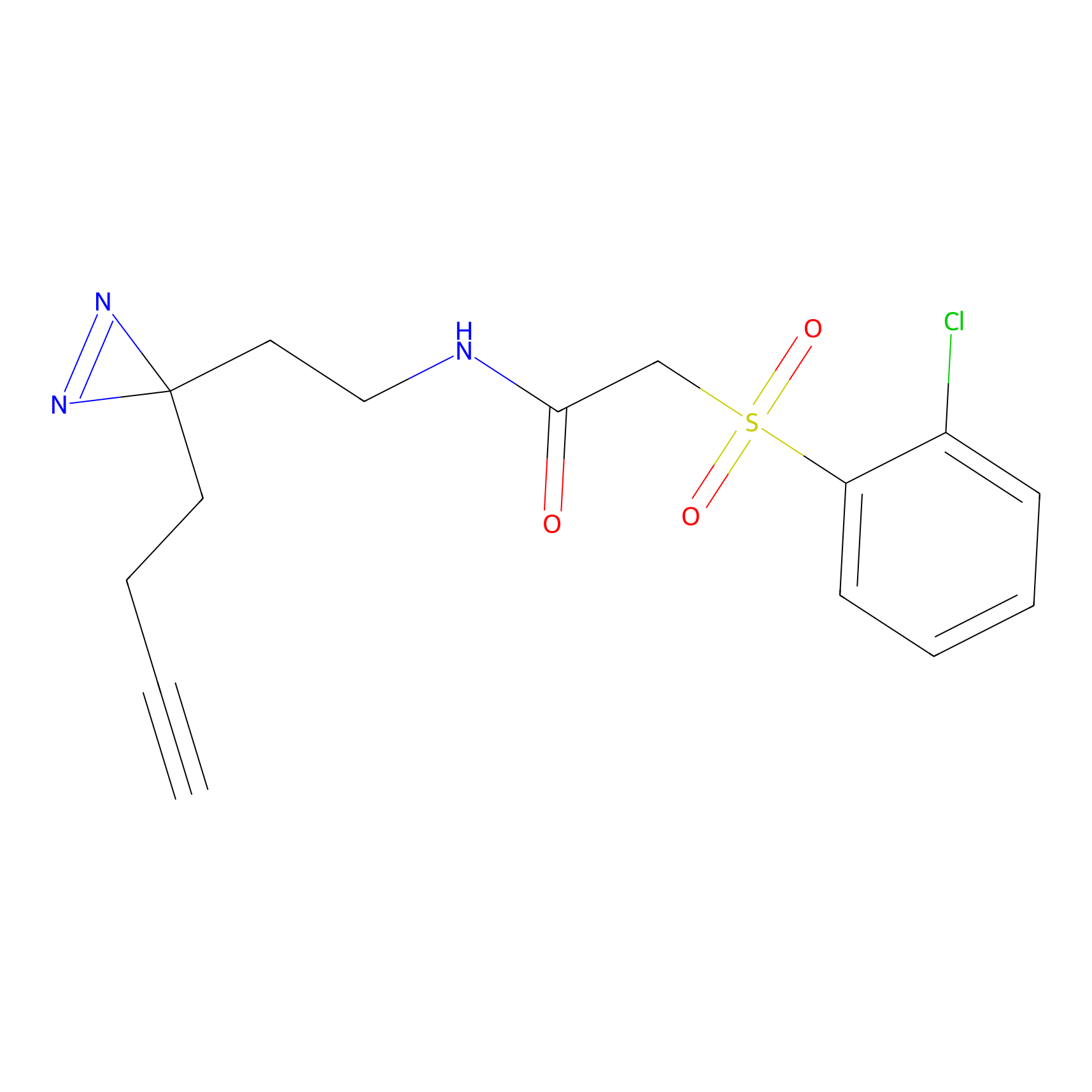
YN-4 Probe Info |
 |
100.00 | LDD0445 | [1] | |
|
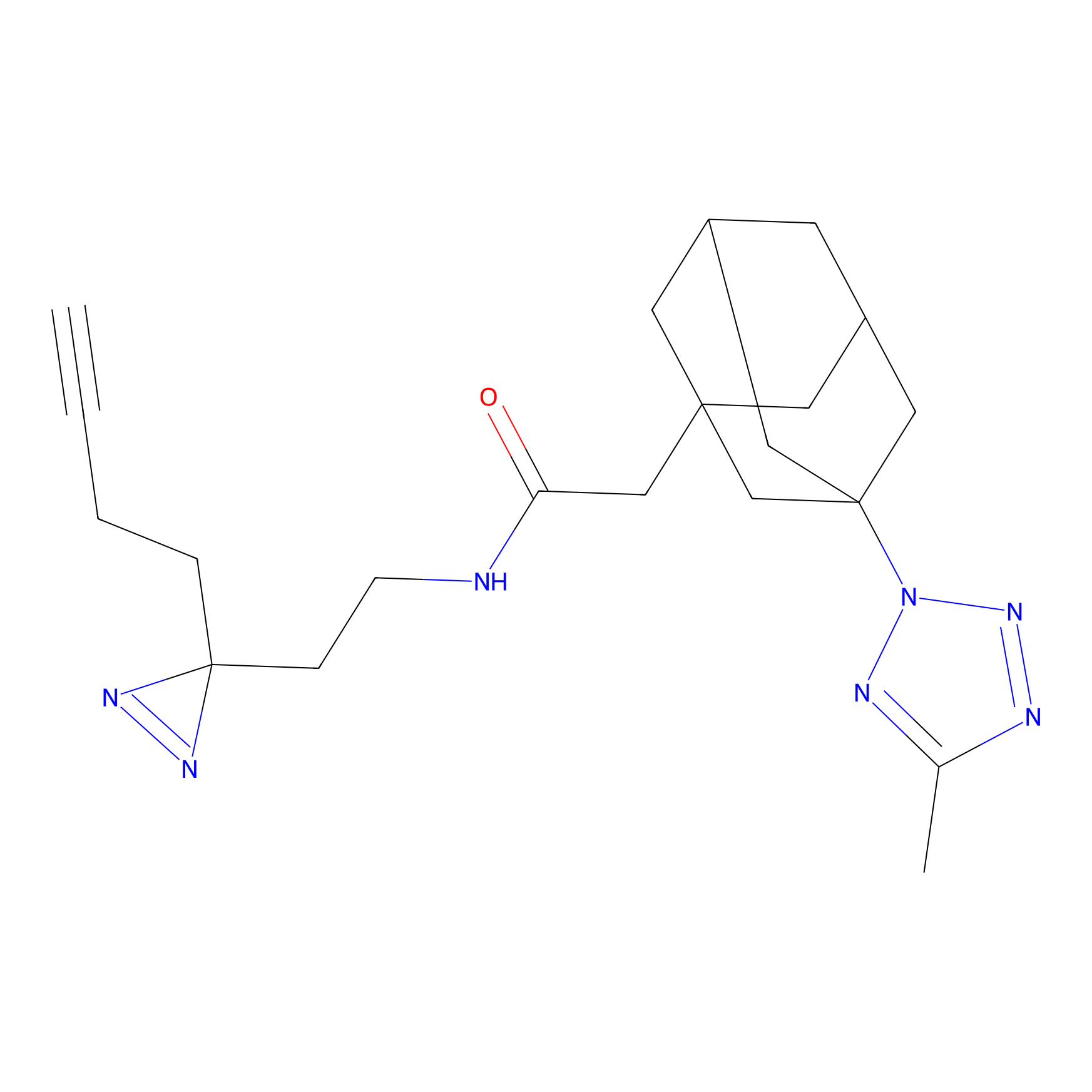
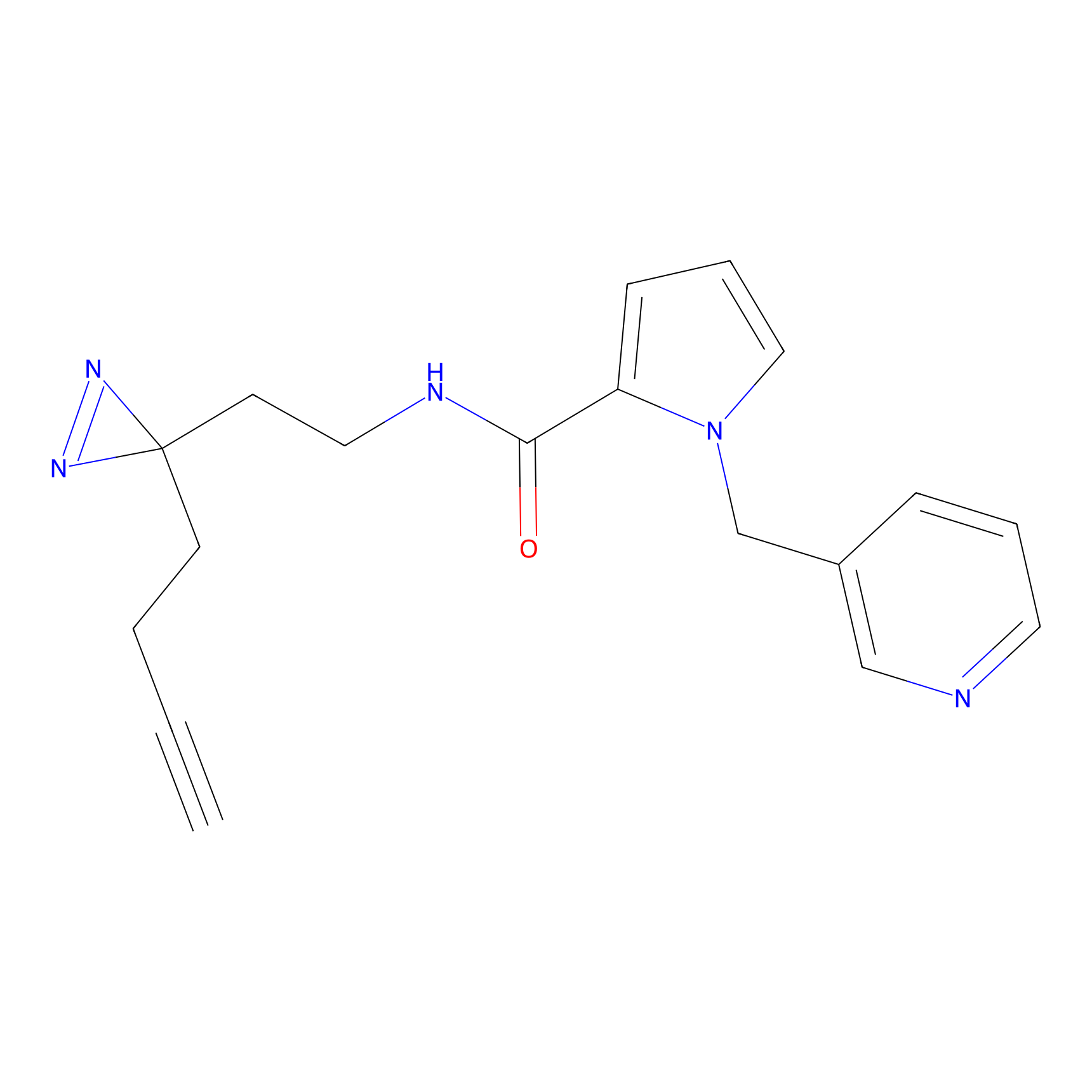
STPyne Probe Info |
 |
K48(7.17) | LDD0277 | [2] | |
|
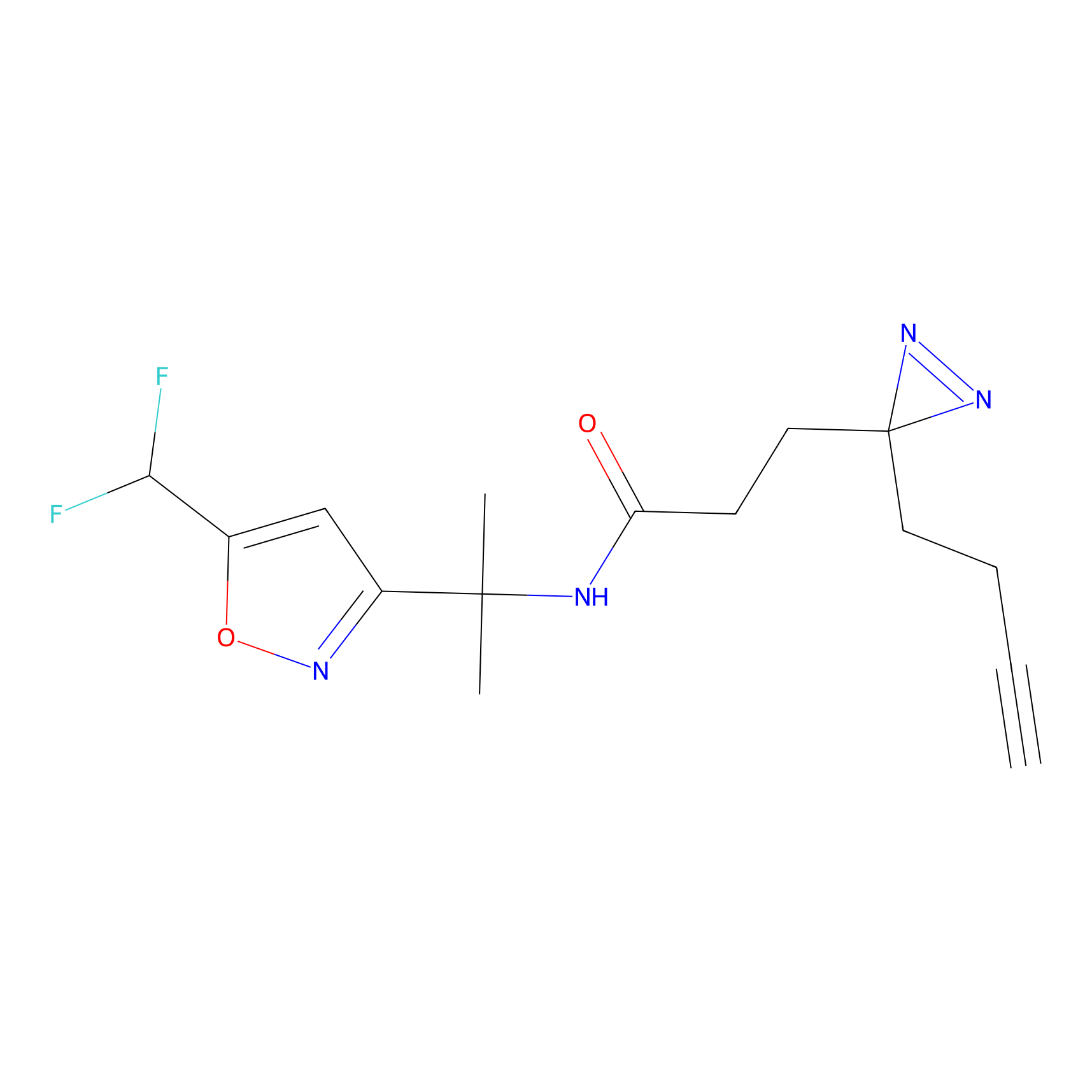
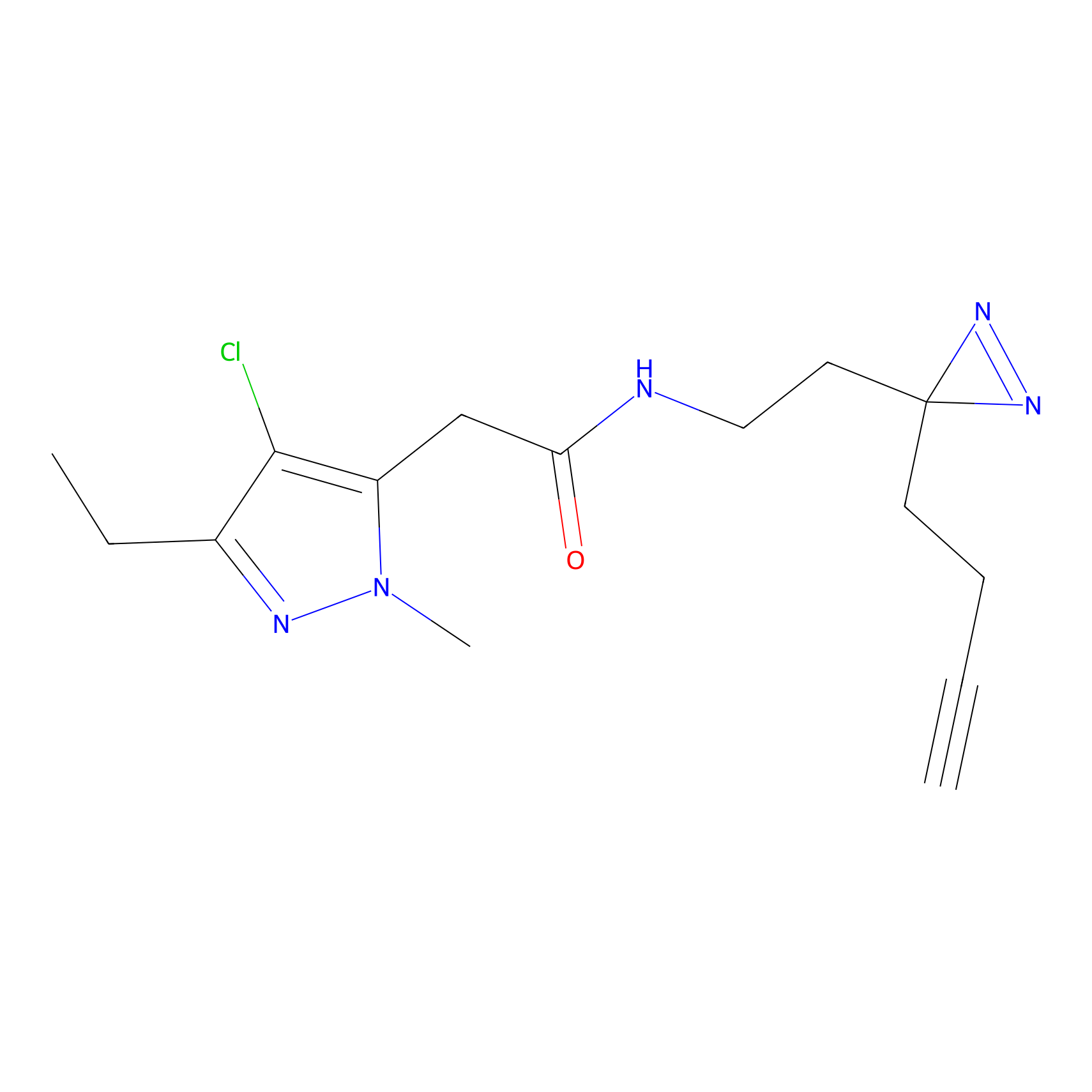
OPA-S-S-alkyne Probe Info |
 |
K130(3.33) | LDD3494 | [3] | |
|
FBPP2 Probe Info |
 |
2.86 | LDD0054 | [4] | |
|
Jackson_1 Probe Info |
 |
2.04 | LDD0120 | [5] | |
|
HPAP Probe Info |
 |
5.03 | LDD0062 | [6] | |
|
Alkyne-RA190 Probe Info |
 |
9.91 | LDD0299 | [7] | |
|
DBIA Probe Info |
 |
C72(0.99) | LDD1507 | [8] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [9] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [10] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [10] | |
|
W1 Probe Info |
 |
E135(0.00); E139(0.00); Q132(0.00); H138(0.00) | LDD0236 | [11] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [12] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C72(0.99) | LDD1507 | [8] |
| LDCM0215 | AC10 | HEK-293T | C72(0.83); C93(1.07) | LDD1508 | [8] |
| LDCM0226 | AC11 | HEK-293T | C72(0.72) | LDD1509 | [8] |
| LDCM0237 | AC12 | HEK-293T | C72(0.91) | LDD1510 | [8] |
| LDCM0259 | AC14 | HEK-293T | C72(0.94) | LDD1512 | [8] |
| LDCM0270 | AC15 | HEK-293T | C72(0.95) | LDD1513 | [8] |
| LDCM0276 | AC17 | HEK-293T | C72(0.94) | LDD1515 | [8] |
| LDCM0277 | AC18 | HEK-293T | C72(0.78); C93(0.99) | LDD1516 | [8] |
| LDCM0278 | AC19 | HEK-293T | C72(0.68) | LDD1517 | [8] |
| LDCM0279 | AC2 | HEK-293T | C72(0.73); C93(0.93) | LDD1518 | [8] |
| LDCM0280 | AC20 | HEK-293T | C72(0.92) | LDD1519 | [8] |
| LDCM0281 | AC21 | HEK-293T | C72(0.79) | LDD1520 | [8] |
| LDCM0282 | AC22 | HEK-293T | C72(0.96) | LDD1521 | [8] |
| LDCM0283 | AC23 | HEK-293T | C72(0.99) | LDD1522 | [8] |
| LDCM0284 | AC24 | HEK-293T | C72(1.03) | LDD1523 | [8] |
| LDCM0285 | AC25 | HEK-293T | C72(0.93) | LDD1524 | [8] |
| LDCM0286 | AC26 | HEK-293T | C72(0.68); C93(0.91) | LDD1525 | [8] |
| LDCM0287 | AC27 | HEK-293T | C72(0.70) | LDD1526 | [8] |
| LDCM0288 | AC28 | HEK-293T | C72(0.91) | LDD1527 | [8] |
| LDCM0289 | AC29 | HEK-293T | C72(0.75) | LDD1528 | [8] |
| LDCM0290 | AC3 | HEK-293T | C72(0.64) | LDD1529 | [8] |
| LDCM0291 | AC30 | HEK-293T | C72(0.93) | LDD1530 | [8] |
| LDCM0292 | AC31 | HEK-293T | C72(0.89) | LDD1531 | [8] |
| LDCM0293 | AC32 | HEK-293T | C72(1.03) | LDD1532 | [8] |
| LDCM0294 | AC33 | HEK-293T | C72(1.02) | LDD1533 | [8] |
| LDCM0295 | AC34 | HEK-293T | C72(0.79); C93(1.12) | LDD1534 | [8] |
| LDCM0296 | AC35 | HEK-293T | C72(0.68) | LDD1535 | [8] |
| LDCM0297 | AC36 | HEK-293T | C72(0.95) | LDD1536 | [8] |
| LDCM0298 | AC37 | HEK-293T | C72(0.78) | LDD1537 | [8] |
| LDCM0299 | AC38 | HEK-293T | C72(1.03) | LDD1538 | [8] |
| LDCM0300 | AC39 | HEK-293T | C72(0.99) | LDD1539 | [8] |
| LDCM0301 | AC4 | HEK-293T | C72(0.96) | LDD1540 | [8] |
| LDCM0302 | AC40 | HEK-293T | C72(1.09) | LDD1541 | [8] |
| LDCM0303 | AC41 | HEK-293T | C72(0.92) | LDD1542 | [8] |
| LDCM0304 | AC42 | HEK-293T | C72(0.69); C93(1.04) | LDD1543 | [8] |
| LDCM0305 | AC43 | HEK-293T | C72(0.68) | LDD1544 | [8] |
| LDCM0306 | AC44 | HEK-293T | C72(0.94) | LDD1545 | [8] |
| LDCM0307 | AC45 | HEK-293T | C72(0.80) | LDD1546 | [8] |
| LDCM0308 | AC46 | HEK-293T | C72(0.99) | LDD1547 | [8] |
| LDCM0309 | AC47 | HEK-293T | C72(1.02) | LDD1548 | [8] |
| LDCM0310 | AC48 | HEK-293T | C72(1.01) | LDD1549 | [8] |
| LDCM0311 | AC49 | HEK-293T | C72(1.02) | LDD1550 | [8] |
| LDCM0312 | AC5 | HEK-293T | C72(0.80) | LDD1551 | [8] |
| LDCM0313 | AC50 | HEK-293T | C72(0.80); C93(1.00) | LDD1552 | [8] |
| LDCM0314 | AC51 | HEK-293T | C72(0.65) | LDD1553 | [8] |
| LDCM0315 | AC52 | HEK-293T | C72(0.95) | LDD1554 | [8] |
| LDCM0316 | AC53 | HEK-293T | C72(0.81) | LDD1555 | [8] |
| LDCM0317 | AC54 | HEK-293T | C72(0.96) | LDD1556 | [8] |
| LDCM0318 | AC55 | HEK-293T | C72(0.96) | LDD1557 | [8] |
| LDCM0319 | AC56 | HEK-293T | C72(1.04) | LDD1558 | [8] |
| LDCM0320 | AC57 | HEK-293T | C72(1.03) | LDD1559 | [8] |
| LDCM0321 | AC58 | HEK-293T | C72(0.90); C93(0.89) | LDD1560 | [8] |
| LDCM0322 | AC59 | HEK-293T | C72(0.89) | LDD1561 | [8] |
| LDCM0323 | AC6 | HEK-293T | C72(0.96) | LDD1562 | [8] |
| LDCM0324 | AC60 | HEK-293T | C72(1.06) | LDD1563 | [8] |
| LDCM0325 | AC61 | HEK-293T | C72(0.93) | LDD1564 | [8] |
| LDCM0326 | AC62 | HEK-293T | C72(0.93) | LDD1565 | [8] |
| LDCM0327 | AC63 | HEK-293T | C72(1.01) | LDD1566 | [8] |
| LDCM0328 | AC64 | HEK-293T | C72(0.97) | LDD1567 | [8] |
| LDCM0334 | AC7 | HEK-293T | C72(0.88) | LDD1568 | [8] |
| LDCM0345 | AC8 | HEK-293T | C72(1.09) | LDD1569 | [8] |
| LDCM0248 | AKOS034007472 | HEK-293T | C72(0.72) | LDD1511 | [8] |
| LDCM0356 | AKOS034007680 | HEK-293T | C72(0.99) | LDD1570 | [8] |
| LDCM0275 | AKOS034007705 | HEK-293T | C72(1.13) | LDD1514 | [8] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [10] |
| LDCM0070 | Cisar_cp37 | MDA-MB-231 | 2.04 | LDD0120 | [5] |
| LDCM0367 | CL1 | HEK-293T | C72(1.21) | LDD1571 | [8] |
| LDCM0368 | CL10 | HEK-293T | C72(1.91) | LDD1572 | [8] |
| LDCM0369 | CL100 | HEK-293T | C72(1.17) | LDD1573 | [8] |
| LDCM0370 | CL101 | HEK-293T | C72(1.09) | LDD1574 | [8] |
| LDCM0371 | CL102 | HEK-293T | C72(1.12) | LDD1575 | [8] |
| LDCM0372 | CL103 | HEK-293T | C72(0.93) | LDD1576 | [8] |
| LDCM0373 | CL104 | HEK-293T | C72(0.99) | LDD1577 | [8] |
| LDCM0374 | CL105 | HEK-293T | C72(0.90) | LDD1578 | [8] |
| LDCM0375 | CL106 | HEK-293T | C72(0.99) | LDD1579 | [8] |
| LDCM0376 | CL107 | HEK-293T | C72(1.01) | LDD1580 | [8] |
| LDCM0377 | CL108 | HEK-293T | C72(0.95) | LDD1581 | [8] |
| LDCM0378 | CL109 | HEK-293T | C72(1.13) | LDD1582 | [8] |
| LDCM0379 | CL11 | HEK-293T | C72(1.34) | LDD1583 | [8] |
| LDCM0380 | CL110 | HEK-293T | C72(1.09) | LDD1584 | [8] |
| LDCM0381 | CL111 | HEK-293T | C72(0.96) | LDD1585 | [8] |
| LDCM0382 | CL112 | HEK-293T | C72(0.87) | LDD1586 | [8] |
| LDCM0383 | CL113 | HEK-293T | C72(1.07) | LDD1587 | [8] |
| LDCM0384 | CL114 | HEK-293T | C72(1.10) | LDD1588 | [8] |
| LDCM0385 | CL115 | HEK-293T | C72(0.87) | LDD1589 | [8] |
| LDCM0386 | CL116 | HEK-293T | C72(1.04) | LDD1590 | [8] |
| LDCM0387 | CL117 | HEK-293T | C72(0.92) | LDD1591 | [8] |
| LDCM0388 | CL118 | HEK-293T | C72(0.93) | LDD1592 | [8] |
| LDCM0389 | CL119 | HEK-293T | C72(0.84) | LDD1593 | [8] |
| LDCM0390 | CL12 | HEK-293T | C72(1.53) | LDD1594 | [8] |
| LDCM0391 | CL120 | HEK-293T | C72(1.01) | LDD1595 | [8] |
| LDCM0392 | CL121 | HEK-293T | C72(1.01) | LDD1596 | [8] |
| LDCM0393 | CL122 | HEK-293T | C72(1.12) | LDD1597 | [8] |
| LDCM0394 | CL123 | HEK-293T | C72(1.26) | LDD1598 | [8] |
| LDCM0395 | CL124 | HEK-293T | C72(1.08) | LDD1599 | [8] |
| LDCM0396 | CL125 | HEK-293T | C72(1.02) | LDD1600 | [8] |
| LDCM0397 | CL126 | HEK-293T | C72(1.12) | LDD1601 | [8] |
| LDCM0398 | CL127 | HEK-293T | C72(0.98) | LDD1602 | [8] |
| LDCM0399 | CL128 | HEK-293T | C72(1.05) | LDD1603 | [8] |
| LDCM0400 | CL13 | HEK-293T | C72(1.15) | LDD1604 | [8] |
| LDCM0401 | CL14 | HEK-293T | C72(0.99) | LDD1605 | [8] |
| LDCM0402 | CL15 | HEK-293T | C72(1.32) | LDD1606 | [8] |
| LDCM0403 | CL16 | HEK-293T | C72(1.03) | LDD1607 | [8] |
| LDCM0404 | CL17 | HEK-293T | C72(1.46) | LDD1608 | [8] |
| LDCM0405 | CL18 | HEK-293T | C72(0.84); C93(1.02) | LDD1609 | [8] |
| LDCM0406 | CL19 | HEK-293T | C72(0.76) | LDD1610 | [8] |
| LDCM0407 | CL2 | HEK-293T | C72(1.05) | LDD1611 | [8] |
| LDCM0408 | CL20 | HEK-293T | C72(1.16) | LDD1612 | [8] |
| LDCM0409 | CL21 | HEK-293T | C72(1.24) | LDD1613 | [8] |
| LDCM0410 | CL22 | HEK-293T | C72(1.17) | LDD1614 | [8] |
| LDCM0411 | CL23 | HEK-293T | C72(1.15) | LDD1615 | [8] |
| LDCM0412 | CL24 | HEK-293T | C72(1.45) | LDD1616 | [8] |
| LDCM0413 | CL25 | HEK-293T | C72(1.44) | LDD1617 | [8] |
| LDCM0414 | CL26 | HEK-293T | C72(0.91) | LDD1618 | [8] |
| LDCM0415 | CL27 | HEK-293T | C72(0.93) | LDD1619 | [8] |
| LDCM0416 | CL28 | HEK-293T | C72(1.06) | LDD1620 | [8] |
| LDCM0417 | CL29 | HEK-293T | C72(1.04) | LDD1621 | [8] |
| LDCM0418 | CL3 | HEK-293T | C72(1.22) | LDD1622 | [8] |
| LDCM0419 | CL30 | HEK-293T | C72(0.89); C93(0.90) | LDD1623 | [8] |
| LDCM0420 | CL31 | HEK-293T | C72(0.71) | LDD1624 | [8] |
| LDCM0421 | CL32 | HEK-293T | C72(1.03) | LDD1625 | [8] |
| LDCM0422 | CL33 | HEK-293T | C72(1.67) | LDD1626 | [8] |
| LDCM0423 | CL34 | HEK-293T | C72(1.20) | LDD1627 | [8] |
| LDCM0424 | CL35 | HEK-293T | C72(1.15) | LDD1628 | [8] |
| LDCM0425 | CL36 | HEK-293T | C72(1.48) | LDD1629 | [8] |
| LDCM0426 | CL37 | HEK-293T | C72(1.42) | LDD1630 | [8] |
| LDCM0428 | CL39 | HEK-293T | C72(1.11) | LDD1632 | [8] |
| LDCM0429 | CL4 | HEK-293T | C72(1.16) | LDD1633 | [8] |
| LDCM0430 | CL40 | HEK-293T | C72(1.00) | LDD1634 | [8] |
| LDCM0431 | CL41 | HEK-293T | C72(1.12) | LDD1635 | [8] |
| LDCM0432 | CL42 | HEK-293T | C72(0.80); C93(0.97) | LDD1636 | [8] |
| LDCM0433 | CL43 | HEK-293T | C72(0.75) | LDD1637 | [8] |
| LDCM0434 | CL44 | HEK-293T | C72(0.96) | LDD1638 | [8] |
| LDCM0435 | CL45 | HEK-293T | C72(0.99) | LDD1639 | [8] |
| LDCM0436 | CL46 | HEK-293T | C72(1.04) | LDD1640 | [8] |
| LDCM0437 | CL47 | HEK-293T | C72(1.11) | LDD1641 | [8] |
| LDCM0438 | CL48 | HEK-293T | C72(1.41) | LDD1642 | [8] |
| LDCM0439 | CL49 | HEK-293T | C72(1.51) | LDD1643 | [8] |
| LDCM0440 | CL5 | HEK-293T | C72(1.20) | LDD1644 | [8] |
| LDCM0441 | CL50 | HEK-293T | C72(1.05) | LDD1645 | [8] |
| LDCM0443 | CL52 | HEK-293T | C72(1.13) | LDD1646 | [8] |
| LDCM0444 | CL53 | HEK-293T | C72(1.26) | LDD1647 | [8] |
| LDCM0445 | CL54 | HEK-293T | C72(1.34); C93(3.01) | LDD1648 | [8] |
| LDCM0446 | CL55 | HEK-293T | C72(0.80) | LDD1649 | [8] |
| LDCM0447 | CL56 | HEK-293T | C72(1.10) | LDD1650 | [8] |
| LDCM0448 | CL57 | HEK-293T | C72(1.25) | LDD1651 | [8] |
| LDCM0449 | CL58 | HEK-293T | C72(1.20) | LDD1652 | [8] |
| LDCM0450 | CL59 | HEK-293T | C72(1.19) | LDD1653 | [8] |
| LDCM0451 | CL6 | HEK-293T | C72(1.11); C93(1.66) | LDD1654 | [8] |
| LDCM0452 | CL60 | HEK-293T | C72(1.52) | LDD1655 | [8] |
| LDCM0453 | CL61 | HEK-293T | C72(1.12) | LDD1656 | [8] |
| LDCM0454 | CL62 | HEK-293T | C72(0.88) | LDD1657 | [8] |
| LDCM0455 | CL63 | HEK-293T | C72(0.91) | LDD1658 | [8] |
| LDCM0456 | CL64 | HEK-293T | C72(1.21) | LDD1659 | [8] |
| LDCM0457 | CL65 | HEK-293T | C72(1.04) | LDD1660 | [8] |
| LDCM0458 | CL66 | HEK-293T | C72(0.80); C93(1.02) | LDD1661 | [8] |
| LDCM0459 | CL67 | HEK-293T | C72(0.74) | LDD1662 | [8] |
| LDCM0460 | CL68 | HEK-293T | C72(1.14) | LDD1663 | [8] |
| LDCM0461 | CL69 | HEK-293T | C72(1.01) | LDD1664 | [8] |
| LDCM0462 | CL7 | HEK-293T | C72(0.76) | LDD1665 | [8] |
| LDCM0463 | CL70 | HEK-293T | C72(1.24) | LDD1666 | [8] |
| LDCM0464 | CL71 | HEK-293T | C72(1.08) | LDD1667 | [8] |
| LDCM0465 | CL72 | HEK-293T | C72(1.30) | LDD1668 | [8] |
| LDCM0466 | CL73 | HEK-293T | C72(1.27) | LDD1669 | [8] |
| LDCM0467 | CL74 | HEK-293T | C72(0.95) | LDD1670 | [8] |
| LDCM0469 | CL76 | HEK-293T | C72(0.95) | LDD1672 | [8] |
| LDCM0470 | CL77 | HEK-293T | C72(1.32) | LDD1673 | [8] |
| LDCM0471 | CL78 | HEK-293T | C72(0.78); C93(0.99) | LDD1674 | [8] |
| LDCM0472 | CL79 | HEK-293T | C72(0.73) | LDD1675 | [8] |
| LDCM0473 | CL8 | HEK-293T | C72(4.56) | LDD1676 | [8] |
| LDCM0474 | CL80 | HEK-293T | C72(1.02) | LDD1677 | [8] |
| LDCM0475 | CL81 | HEK-293T | C72(0.86) | LDD1678 | [8] |
| LDCM0476 | CL82 | HEK-293T | C72(1.06) | LDD1679 | [8] |
| LDCM0477 | CL83 | HEK-293T | C72(1.18) | LDD1680 | [8] |
| LDCM0478 | CL84 | HEK-293T | C72(1.50) | LDD1681 | [8] |
| LDCM0479 | CL85 | HEK-293T | C72(1.15) | LDD1682 | [8] |
| LDCM0480 | CL86 | HEK-293T | C72(1.10) | LDD1683 | [8] |
| LDCM0481 | CL87 | HEK-293T | C72(1.02) | LDD1684 | [8] |
| LDCM0482 | CL88 | HEK-293T | C72(1.02) | LDD1685 | [8] |
| LDCM0483 | CL89 | HEK-293T | C72(1.10) | LDD1686 | [8] |
| LDCM0484 | CL9 | HEK-293T | C72(1.06) | LDD1687 | [8] |
| LDCM0485 | CL90 | HEK-293T | C72(1.58); C93(2.58) | LDD1688 | [8] |
| LDCM0486 | CL91 | HEK-293T | C72(0.81) | LDD1689 | [8] |
| LDCM0487 | CL92 | HEK-293T | C72(1.25) | LDD1690 | [8] |
| LDCM0488 | CL93 | HEK-293T | C72(1.19) | LDD1691 | [8] |
| LDCM0489 | CL94 | HEK-293T | C72(1.10) | LDD1692 | [8] |
| LDCM0490 | CL95 | HEK-293T | C72(1.63) | LDD1693 | [8] |
| LDCM0491 | CL96 | HEK-293T | C72(1.55) | LDD1694 | [8] |
| LDCM0492 | CL97 | HEK-293T | C72(1.26) | LDD1695 | [8] |
| LDCM0493 | CL98 | HEK-293T | C72(1.04) | LDD1696 | [8] |
| LDCM0494 | CL99 | HEK-293T | C72(0.91) | LDD1697 | [8] |
| LDCM0189 | Compound 16 | HEK-293T | 10.00 | LDD0492 | [14] |
| LDCM0185 | Compound 17 | HEK-293T | 9.19 | LDD0504 | [14] |
| LDCM0184 | Compound 20 | HEK-293T | 13.91 | LDD0503 | [14] |
| LDCM0186 | Compound 31 | HEK-293T | 12.12 | LDD0506 | [14] |
| LDCM0187 | Compound 32 | HEK-293T | 45.93 | LDD0507 | [14] |
| LDCM0190 | Compound 34 | HEK-293T | 14.81 | LDD0497 | [14] |
| LDCM0194 | Compound 37 | HEK-293T | 6.51 | LDD0498 | [14] |
| LDCM0495 | E2913 | HEK-293T | C72(0.98) | LDD1698 | [8] |
| LDCM0468 | Fragment33 | HEK-293T | C72(0.90) | LDD1671 | [8] |
| LDCM0427 | Fragment51 | HEK-293T | C72(0.96) | LDD1631 | [8] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [10] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [10] |
| LDCM0014 | Panhematin | HEK-293T | 5.03 | LDD0062 | [6] |
| LDCM0131 | RA190 | MM1.R | 9.91 | LDD0299 | [7] |
| LDCM0084 | Ro 48-8071 | A-549 | 20.00 | LDD0144 | [15] |
| LDCM0008 | Tranylcypromine | SH-SY5Y | 2.86 | LDD0054 | [4] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Paired-like homeodomain transcription factor LEUTX (LEUTX) | Paired homeobox family | A8MZ59 | |||
GPCR
Immunoglobulin
Cytokine and receptor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Interferon gamma receptor 2 (IFNGR2) | Type II cytokine receptor family | P38484 | |||
Other
The Drug(s) Related To This Target
Phase 3
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Min-101 | . | D0L7TA | |||
Phase 2
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Ct1812 | . | D0FX8N | |||
Preclinical
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Anavex 1007 | . | D0M6OW | |||
Investigative
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| 1,3-ditolylguanidine | Small molecular drug | D02WJT | |||
| Sm 21 | Small molecular drug | D0X7BG | |||
Patented
References