Details of the Target
General Information of Target
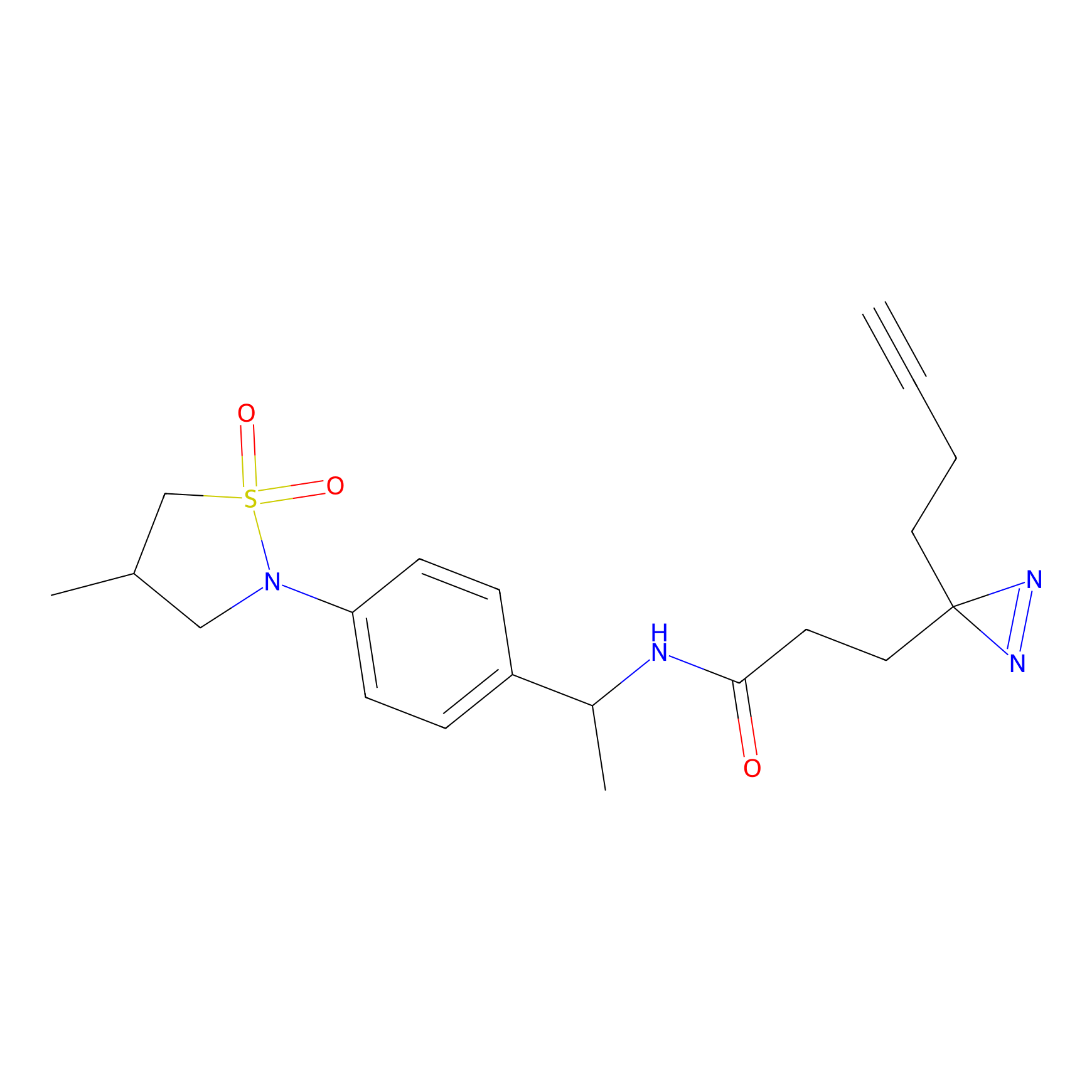
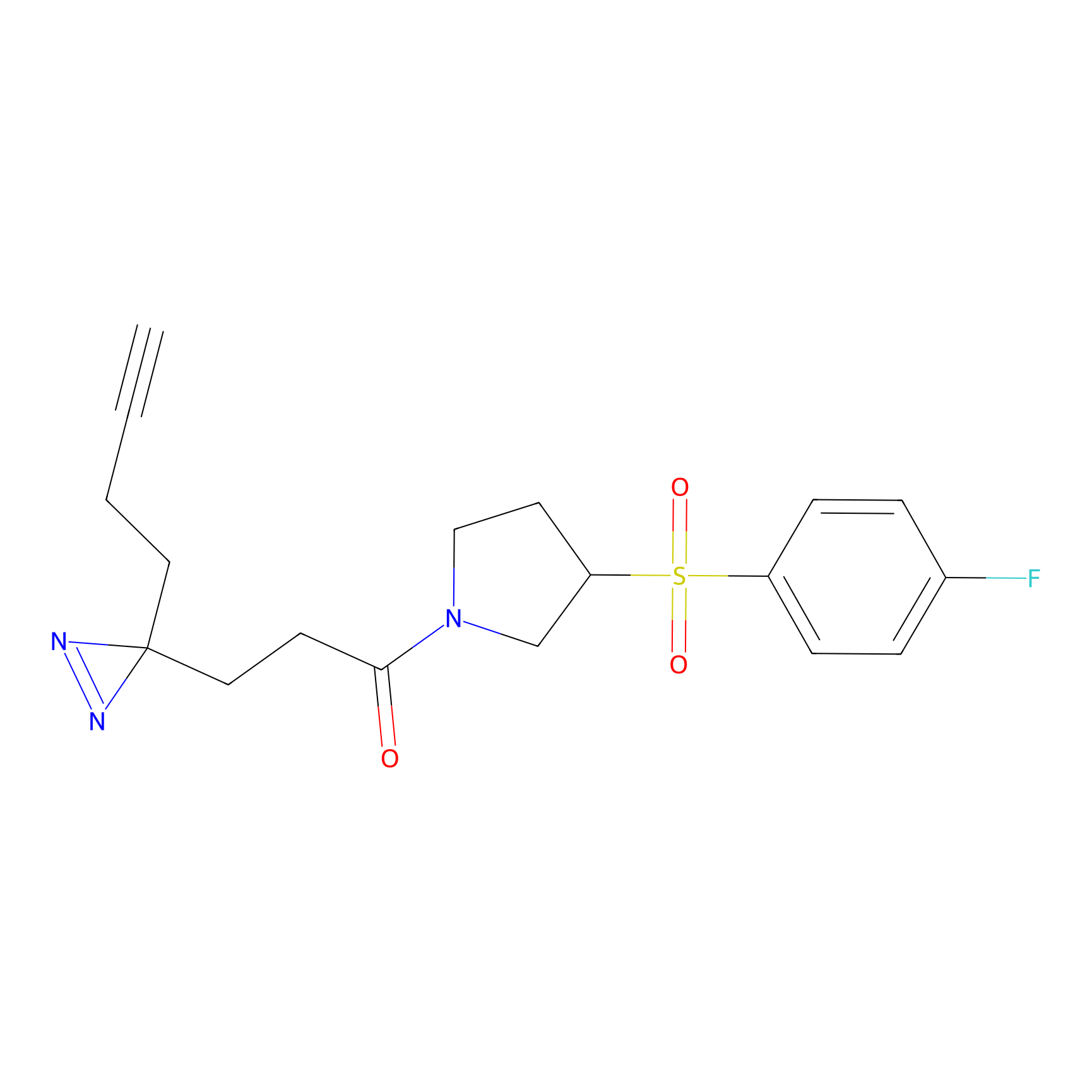
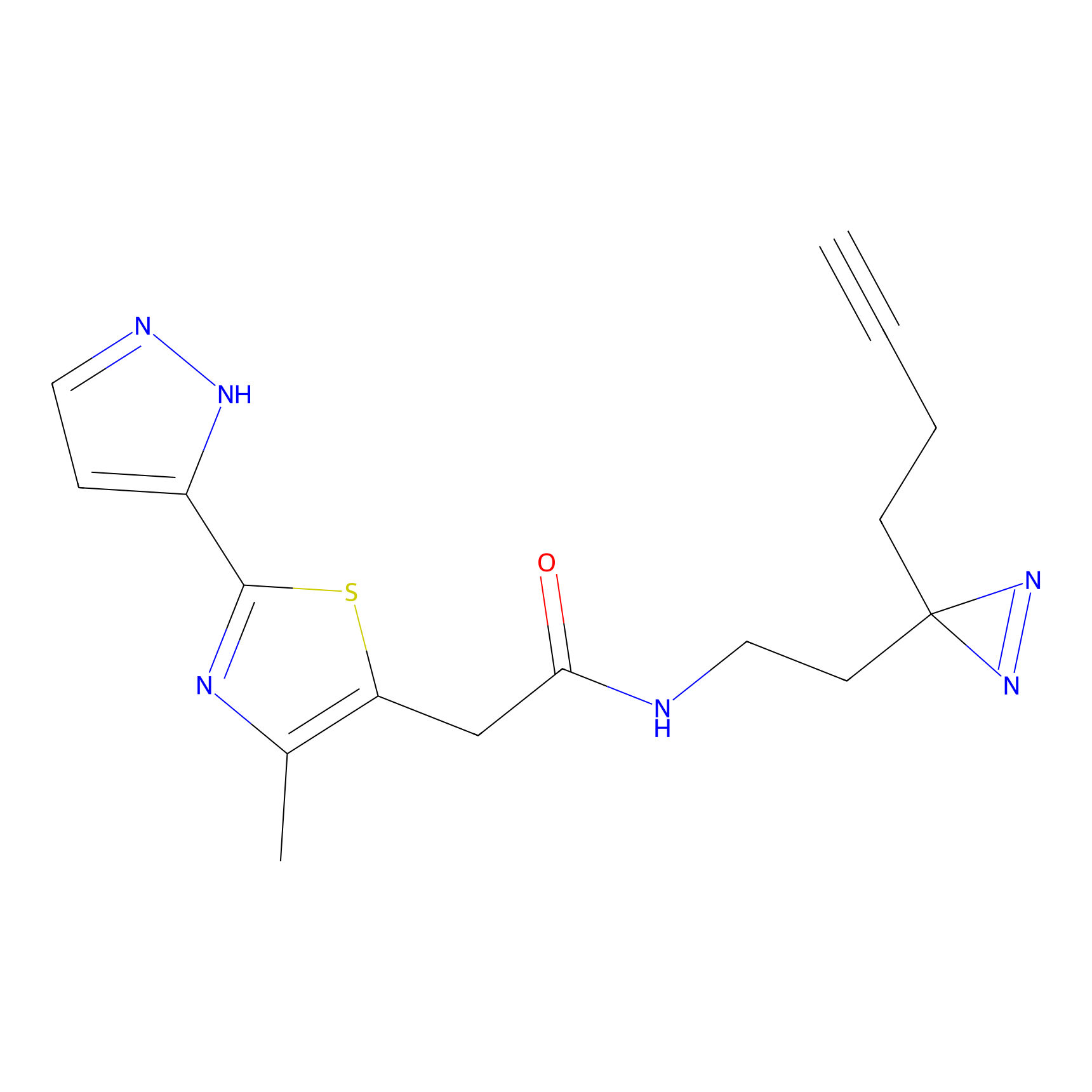
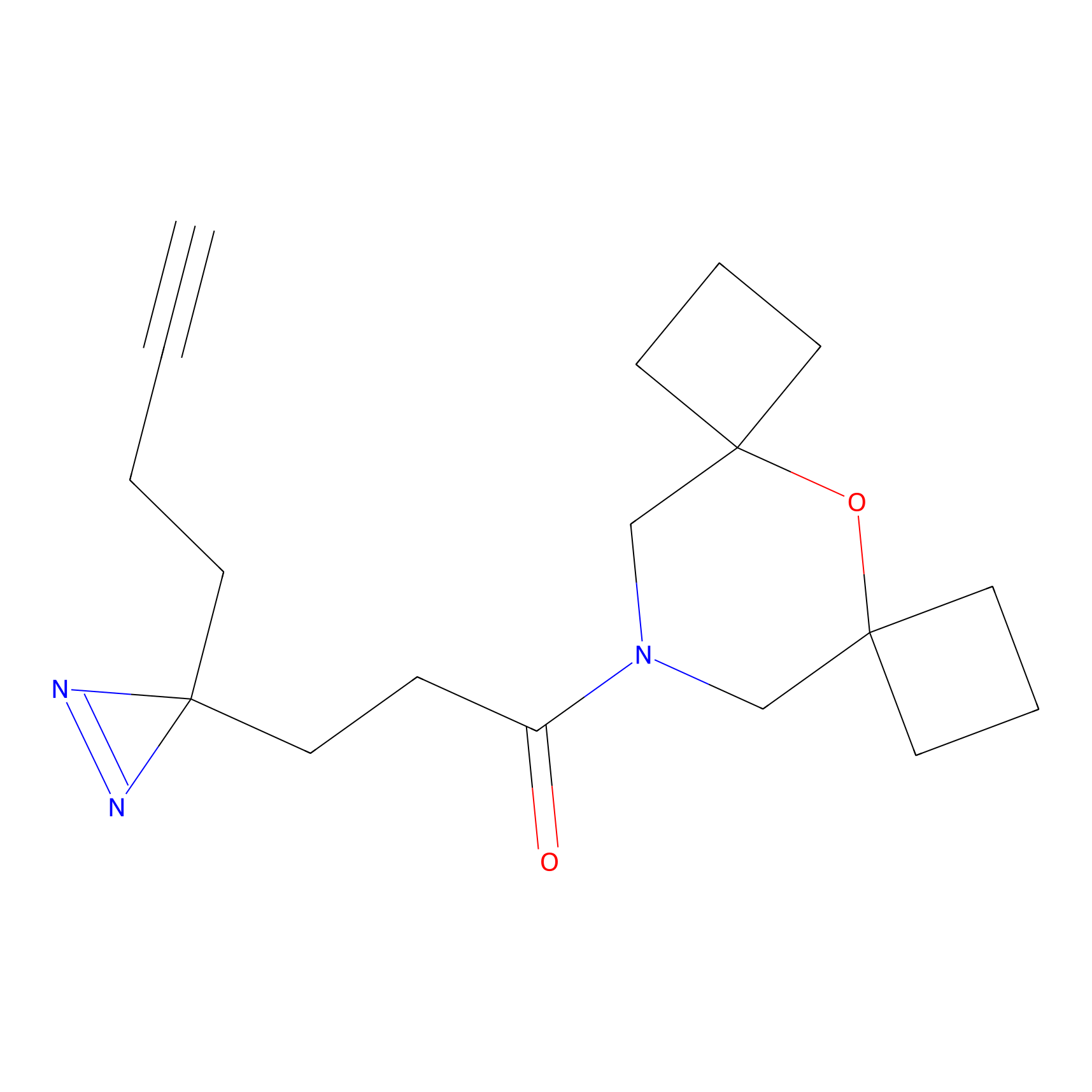
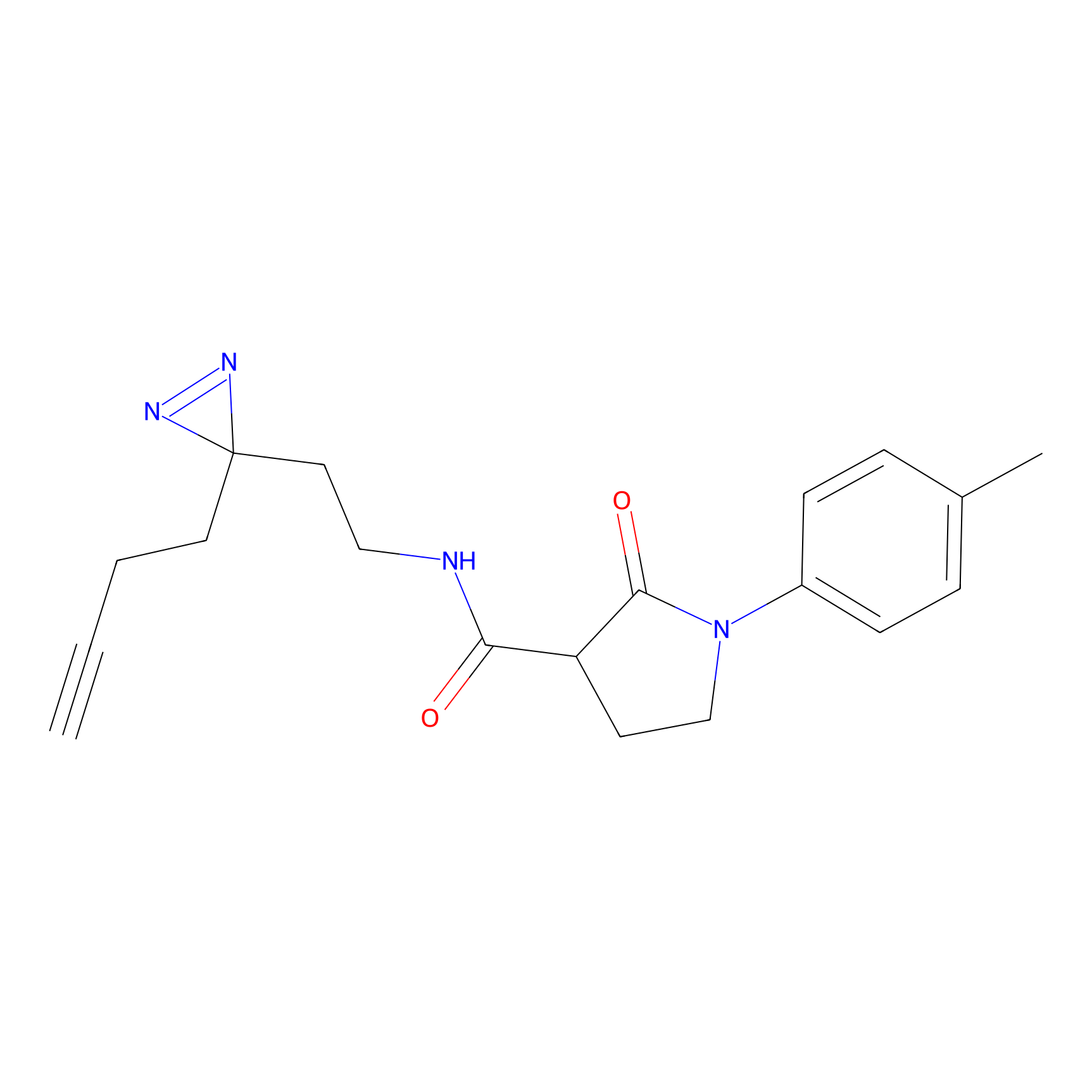
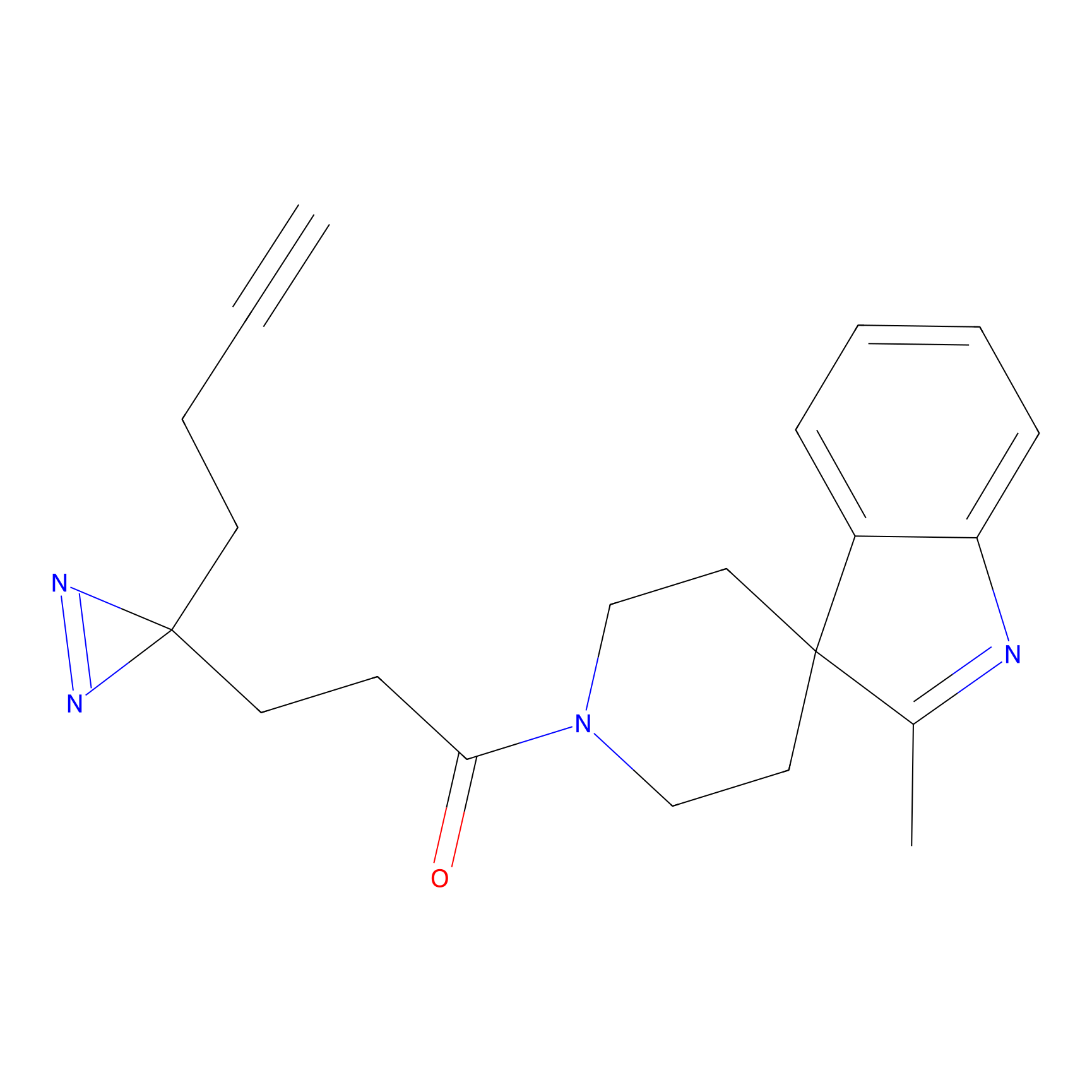
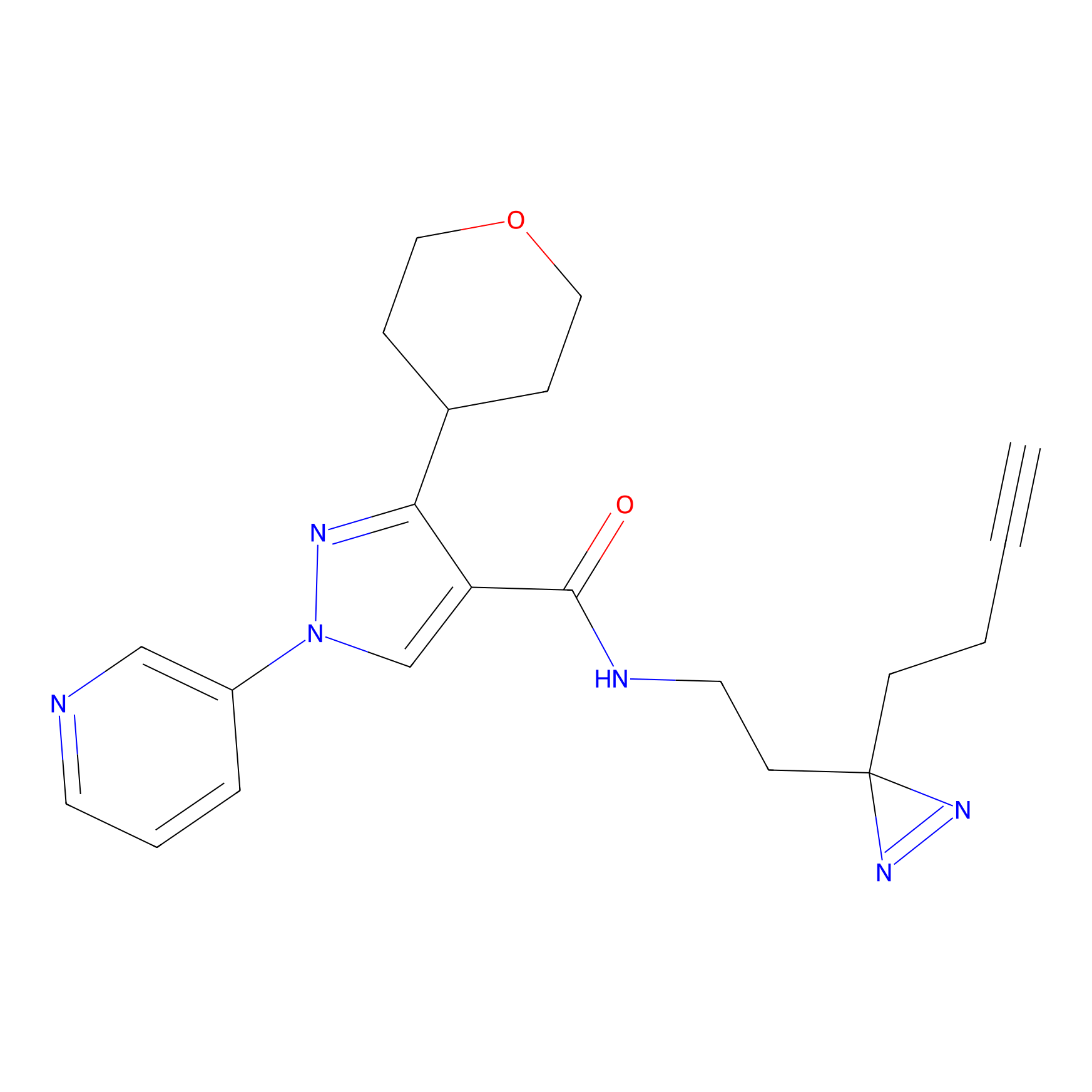
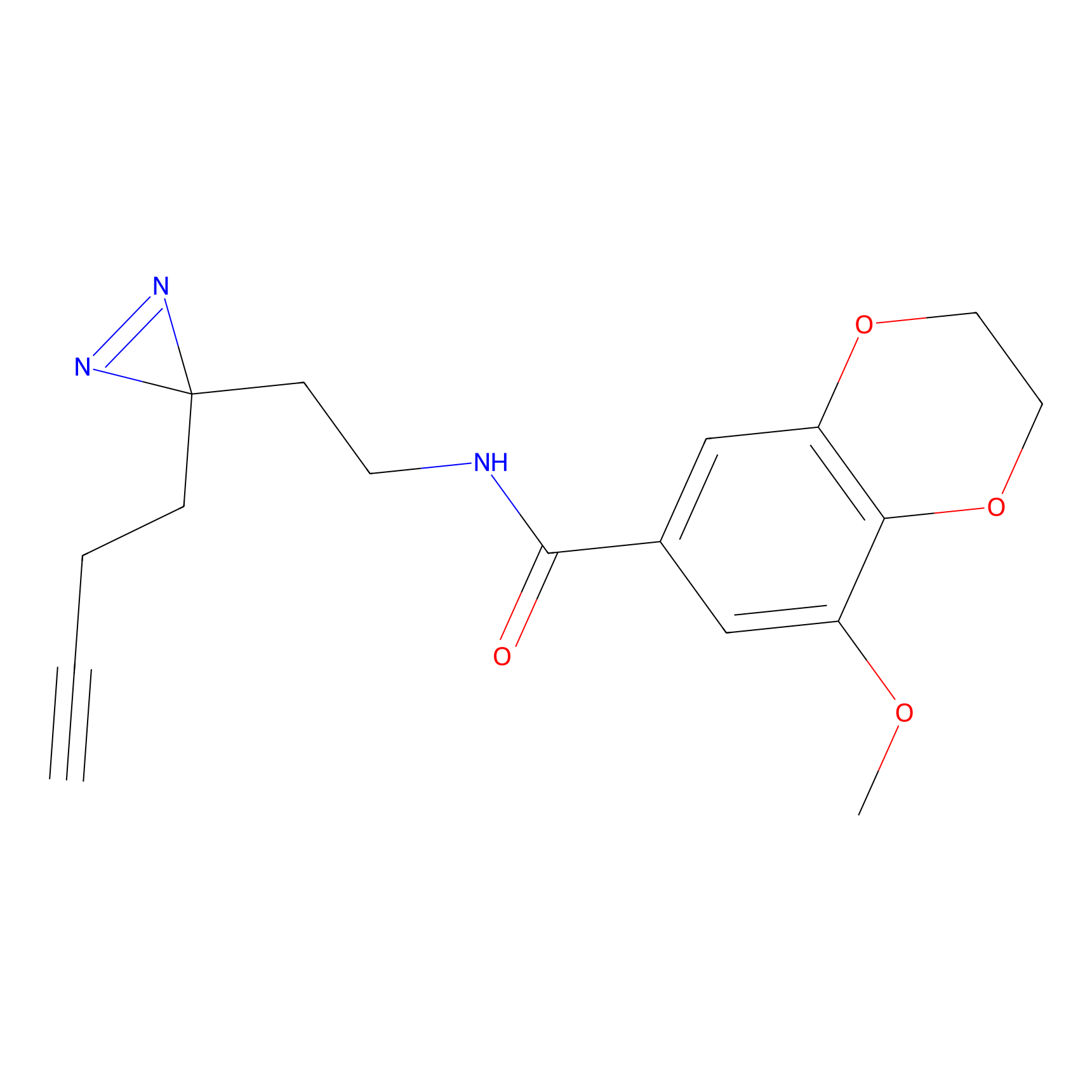
Probe(s) Labeling This Target
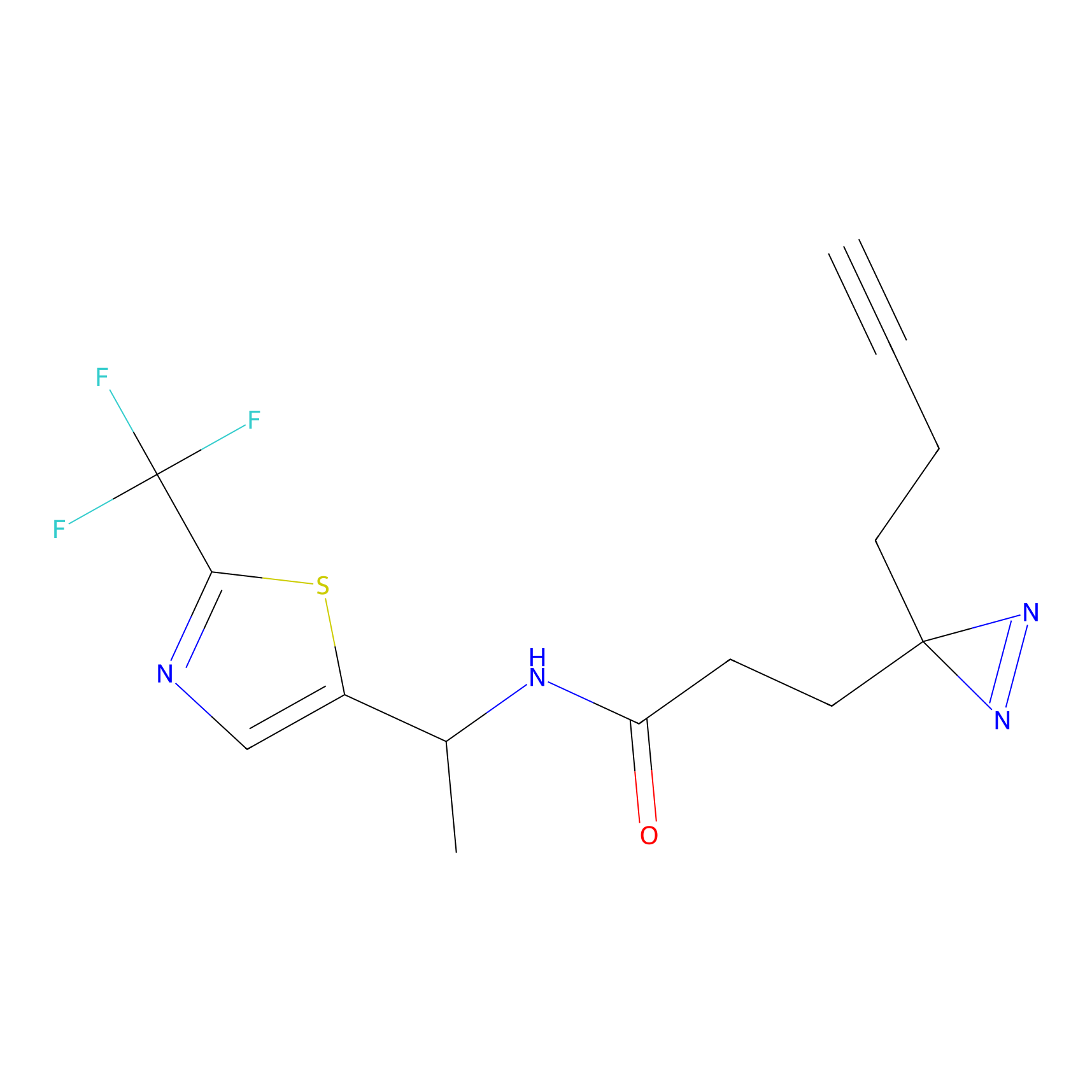
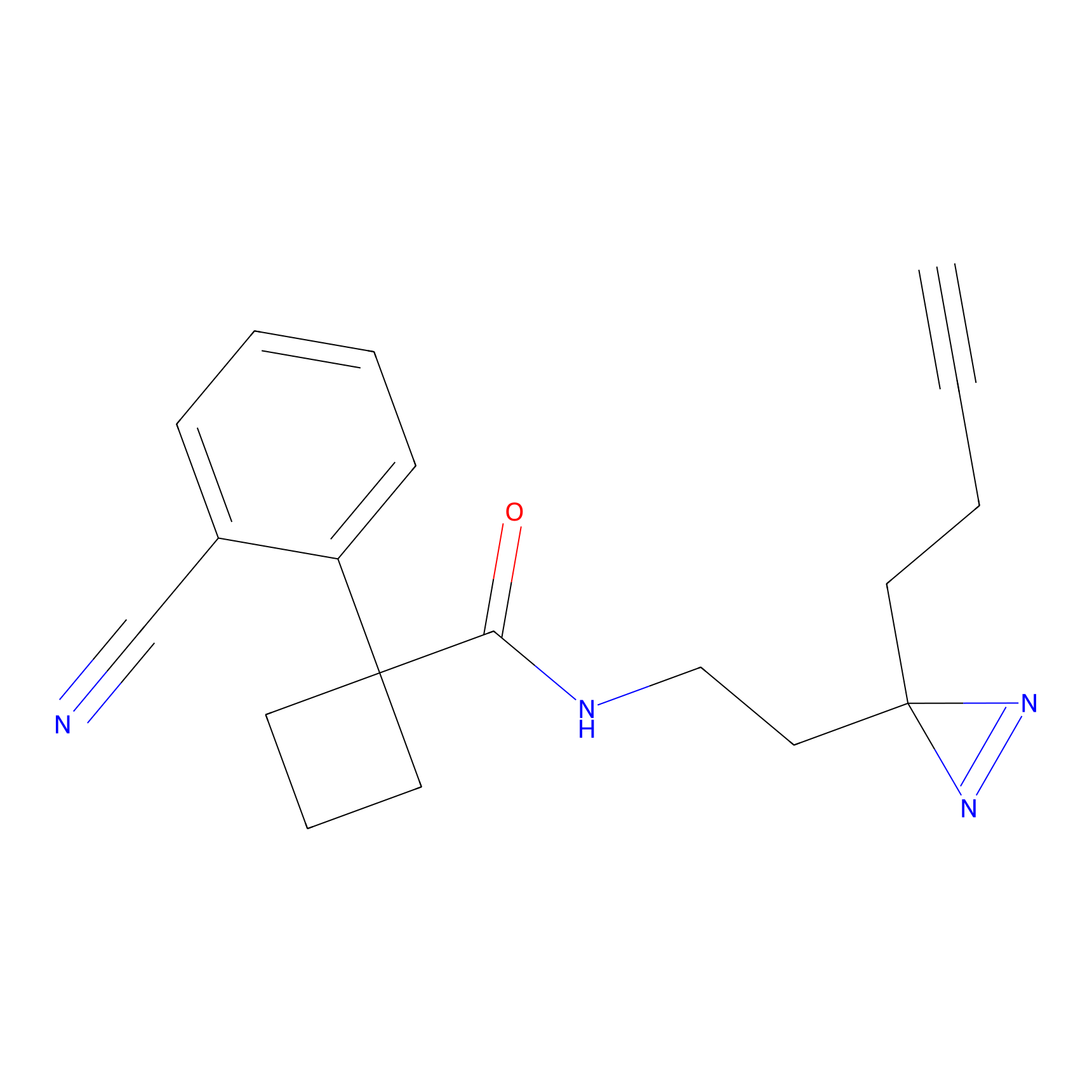
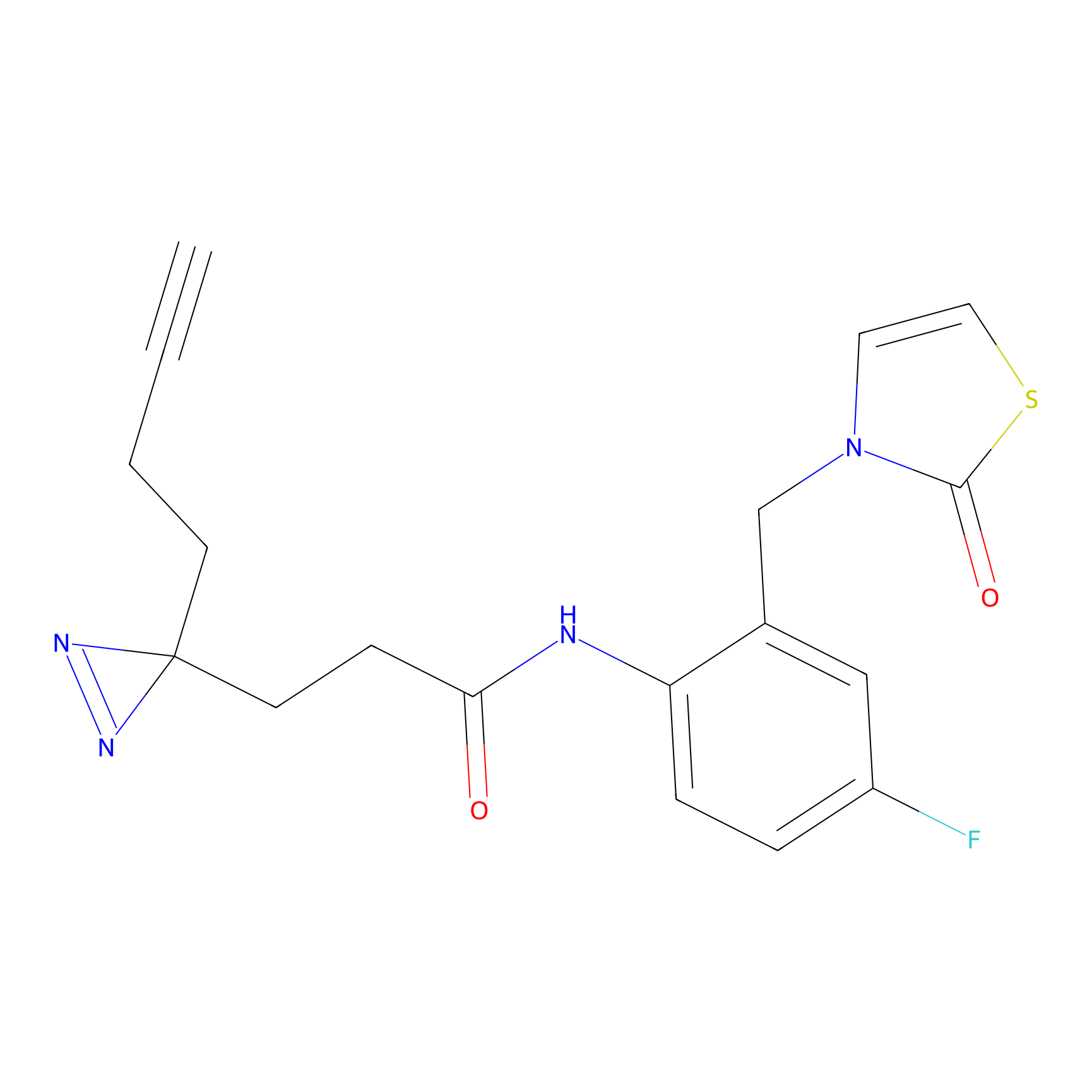
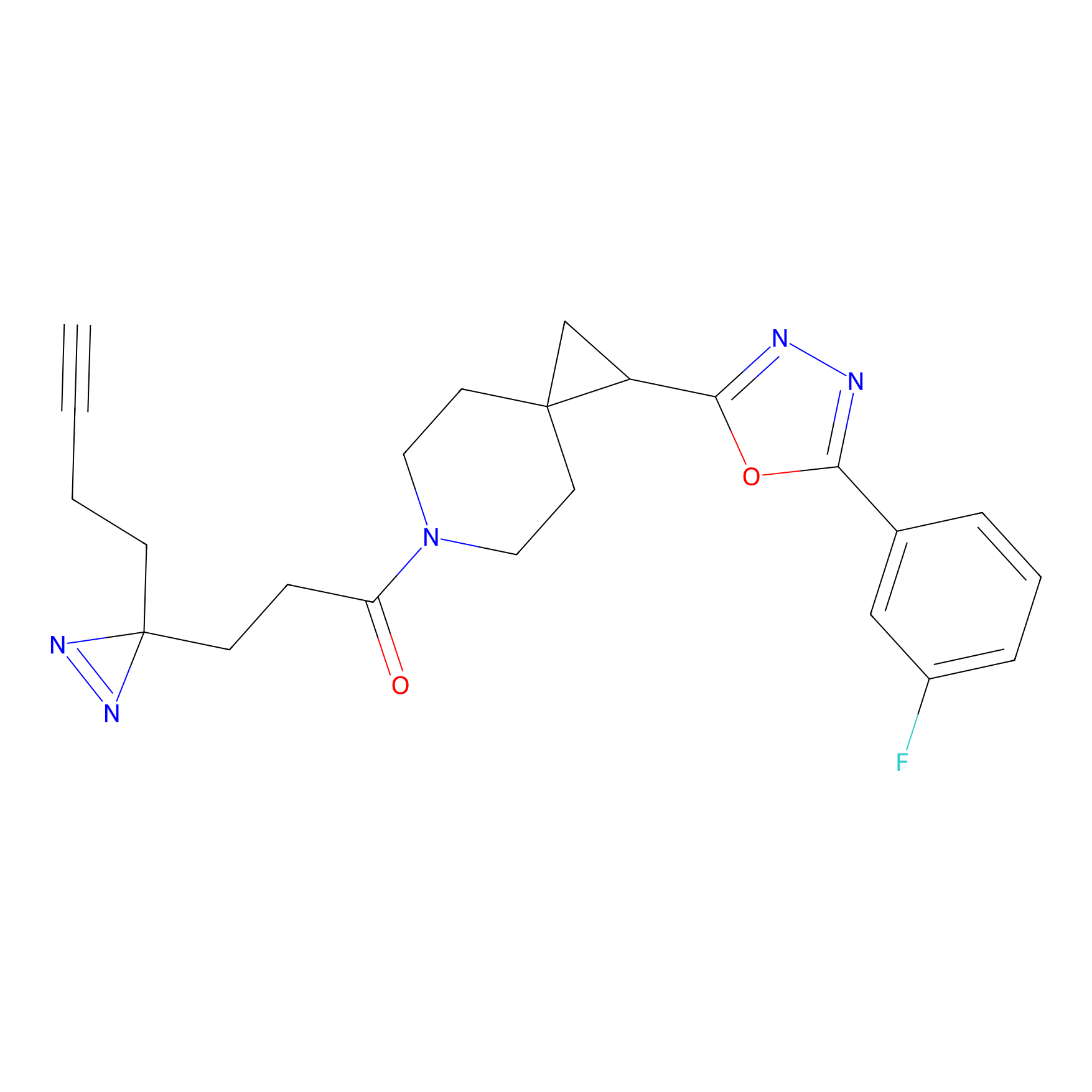
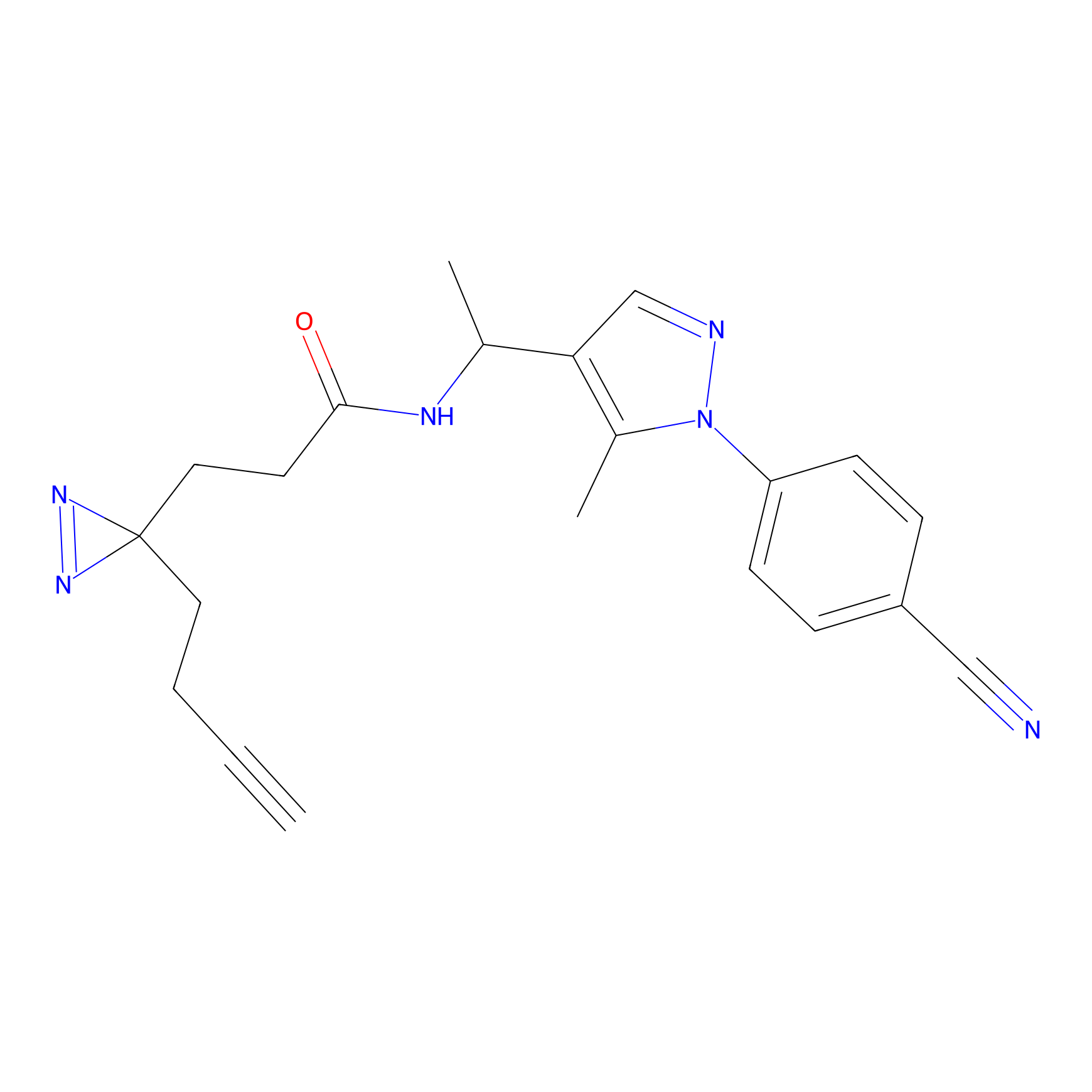
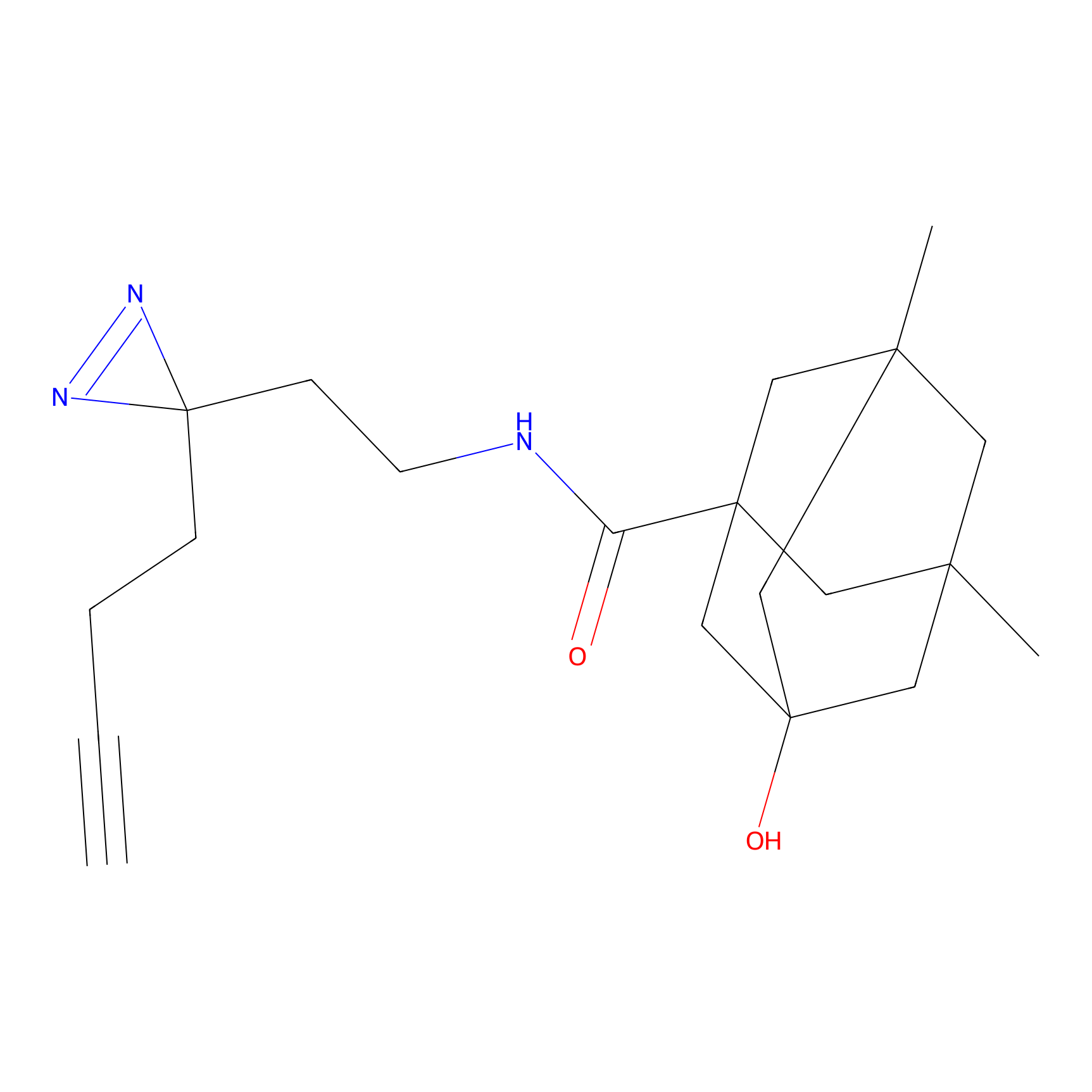
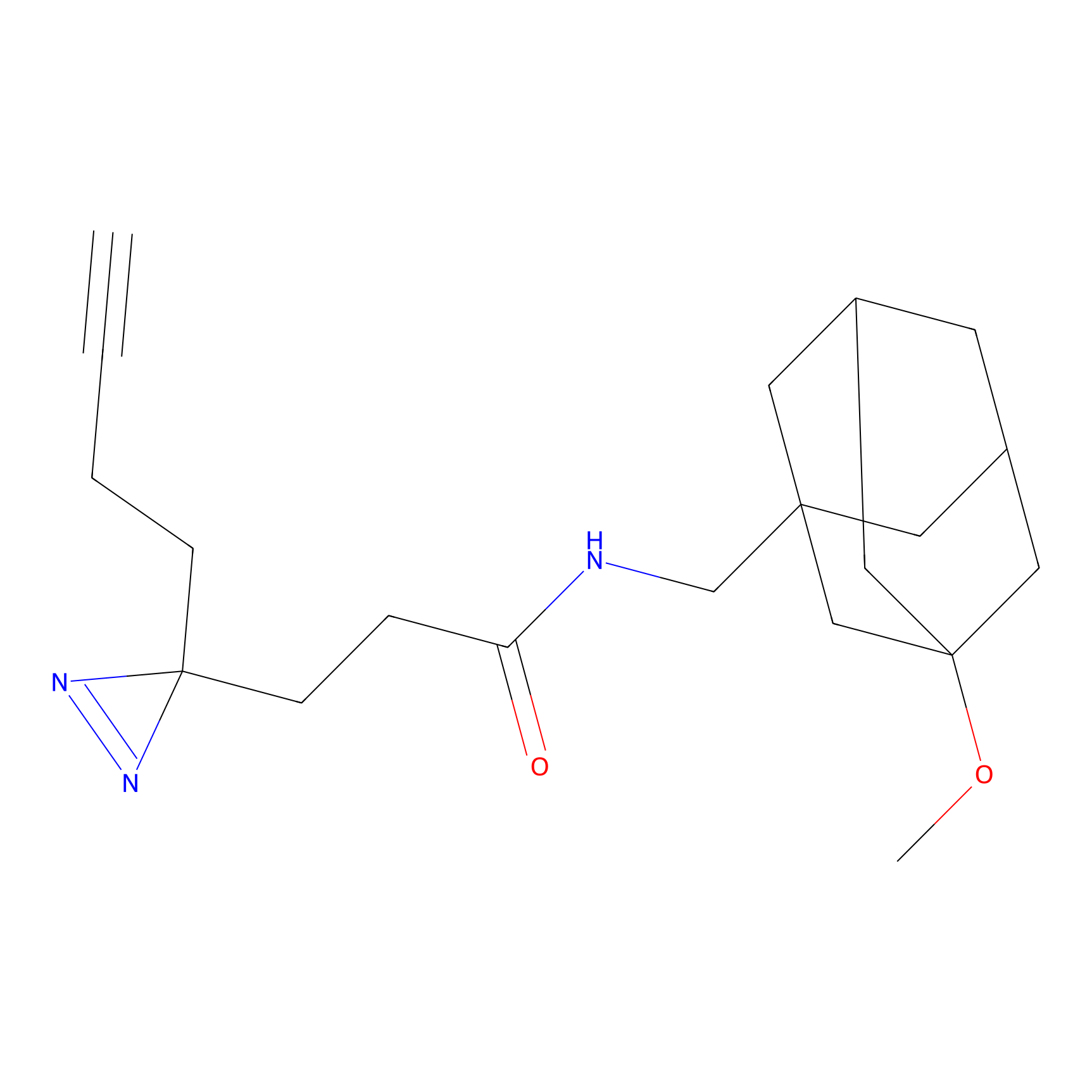
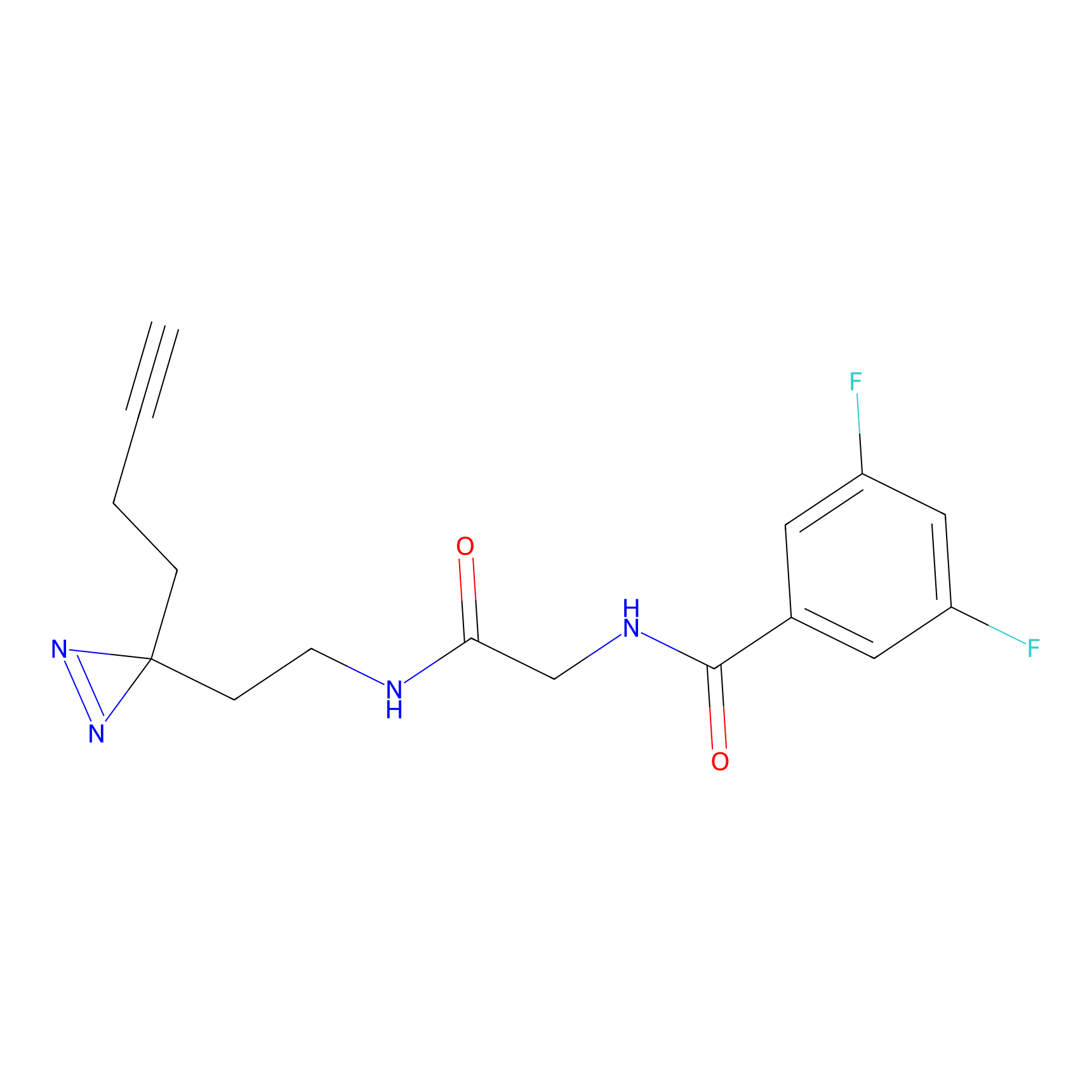
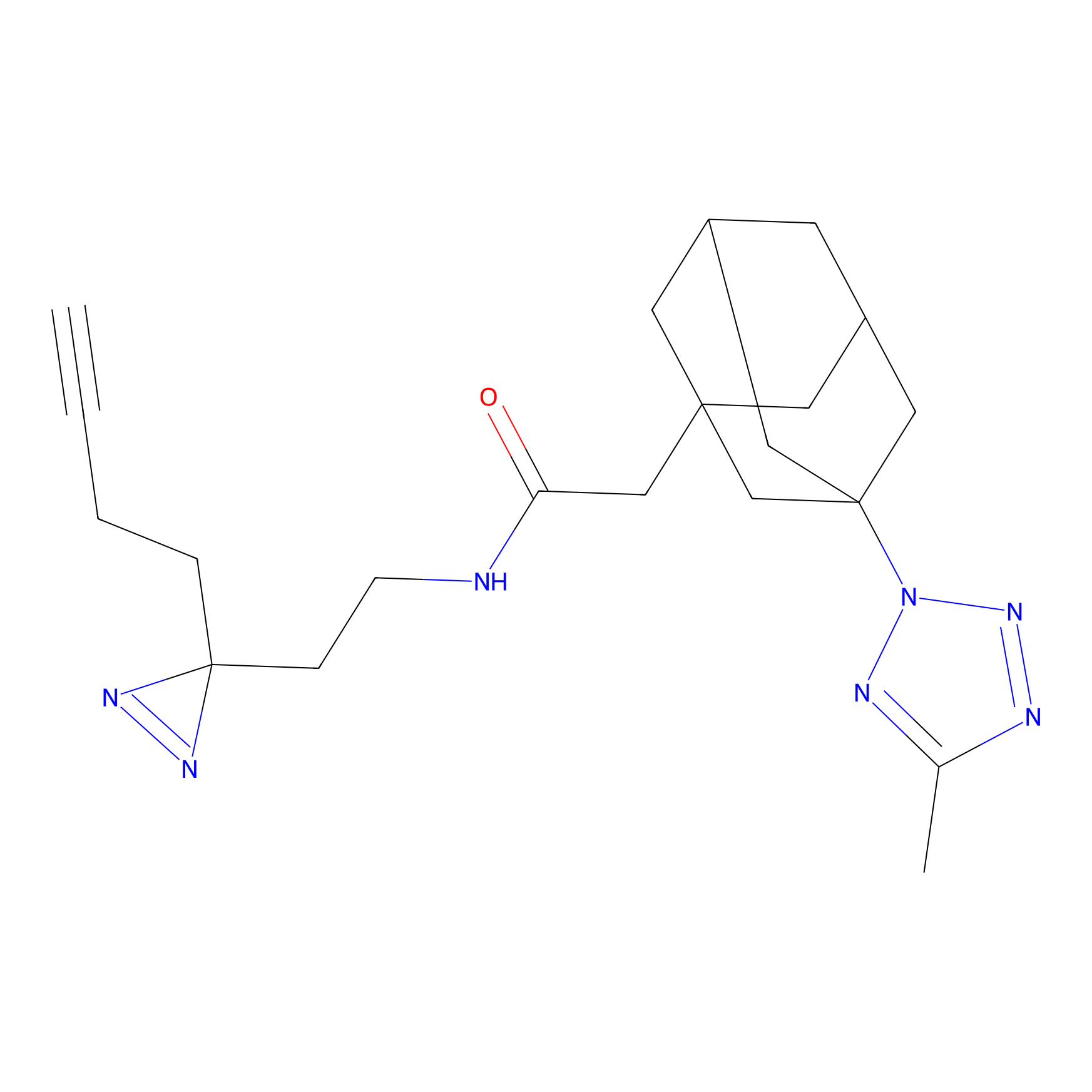
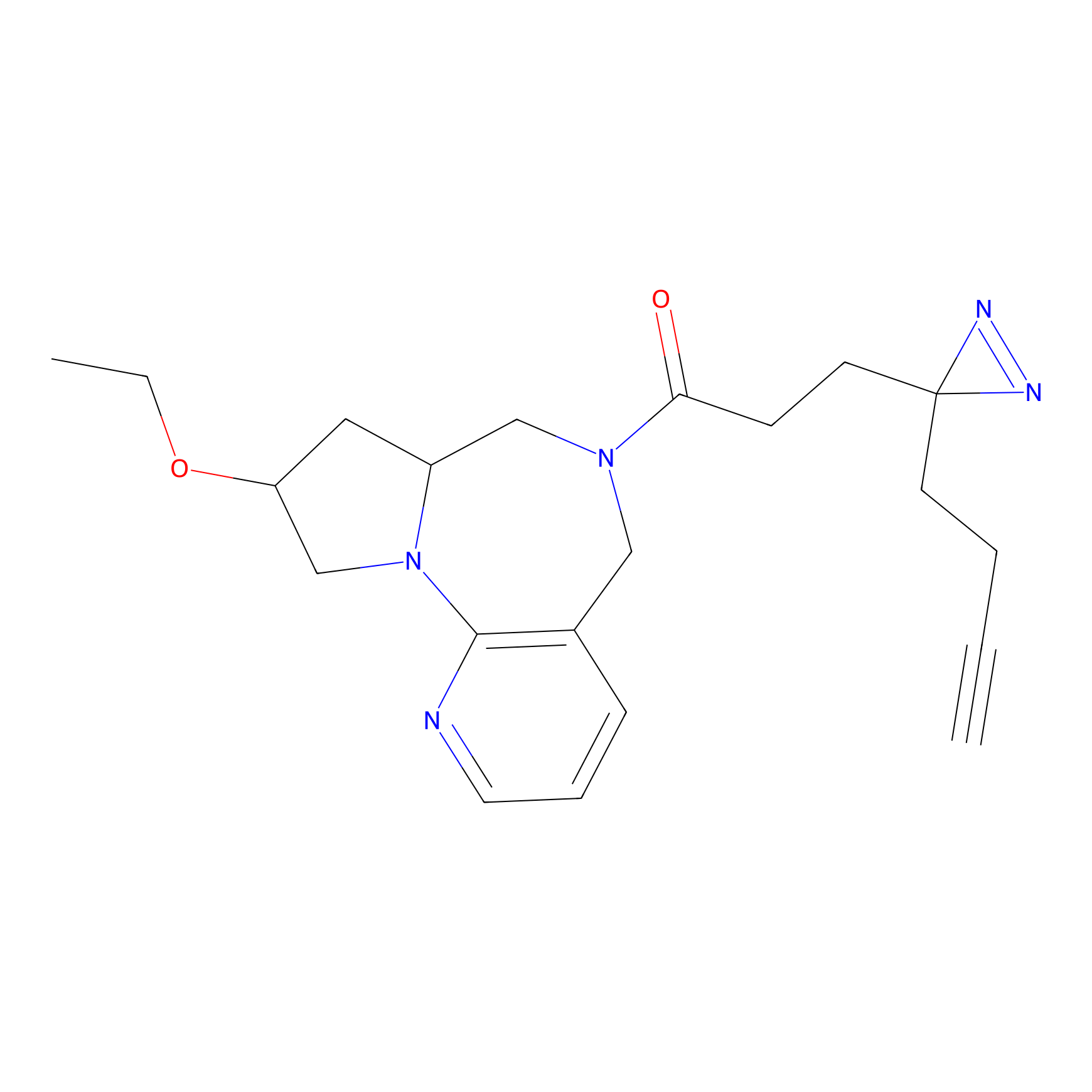
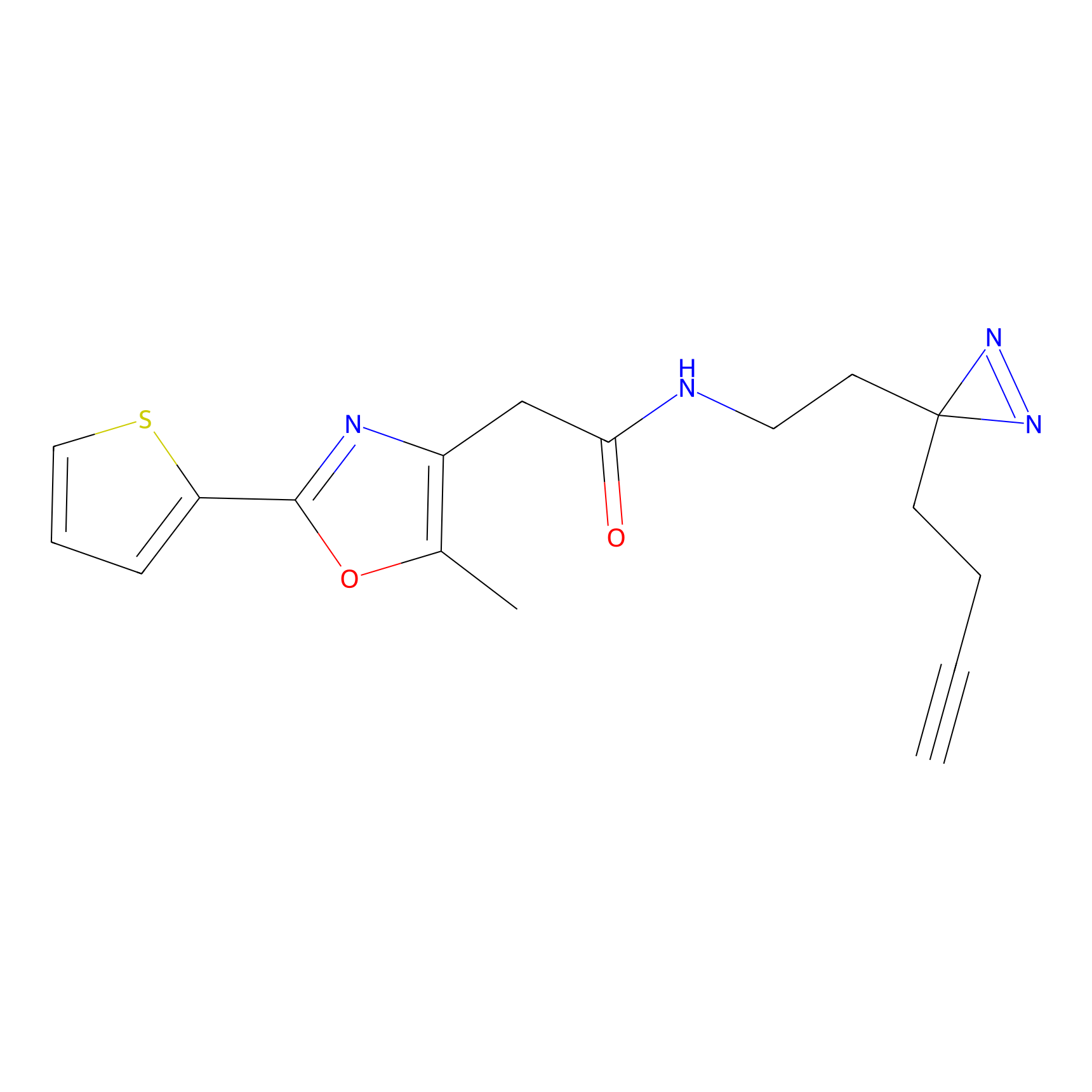
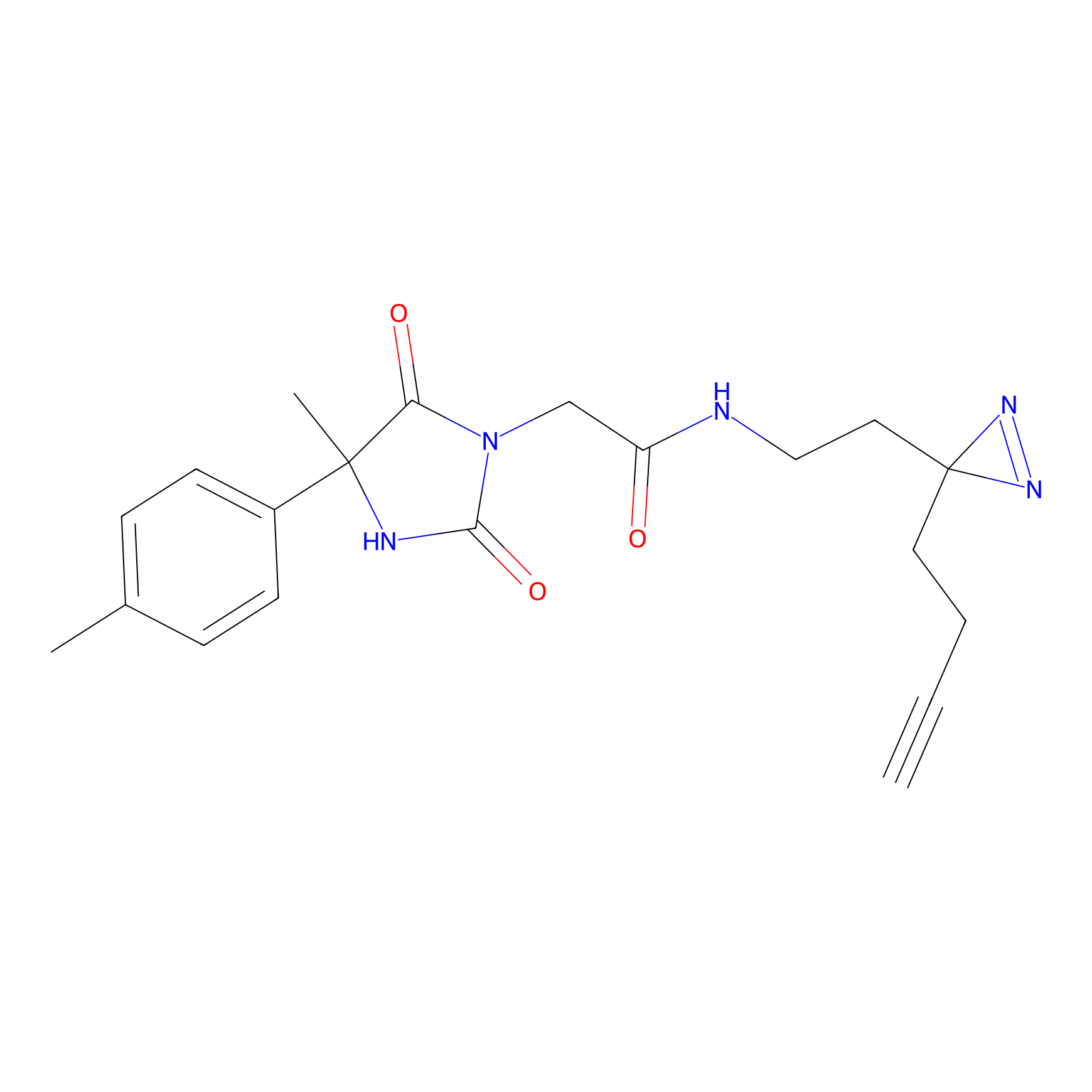
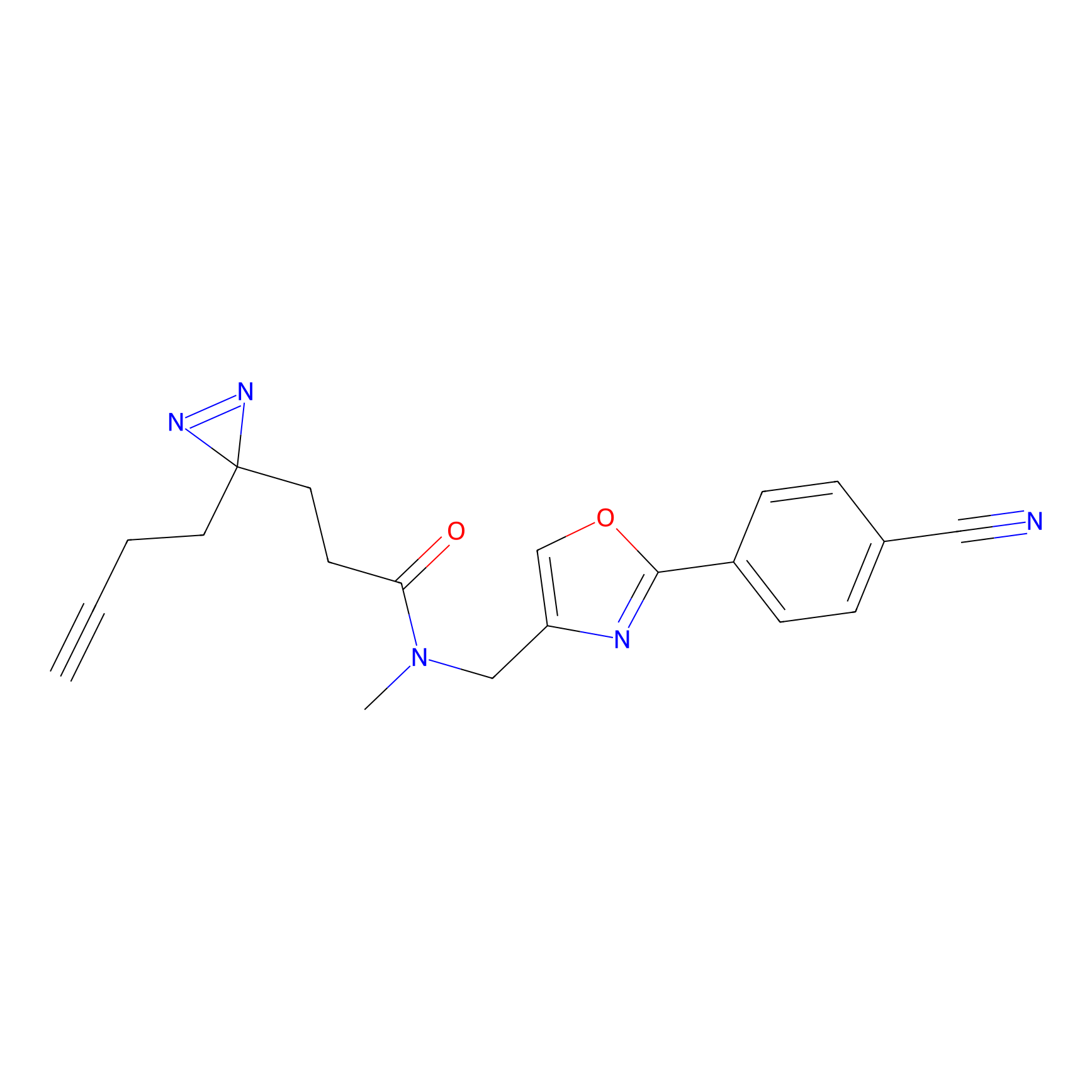
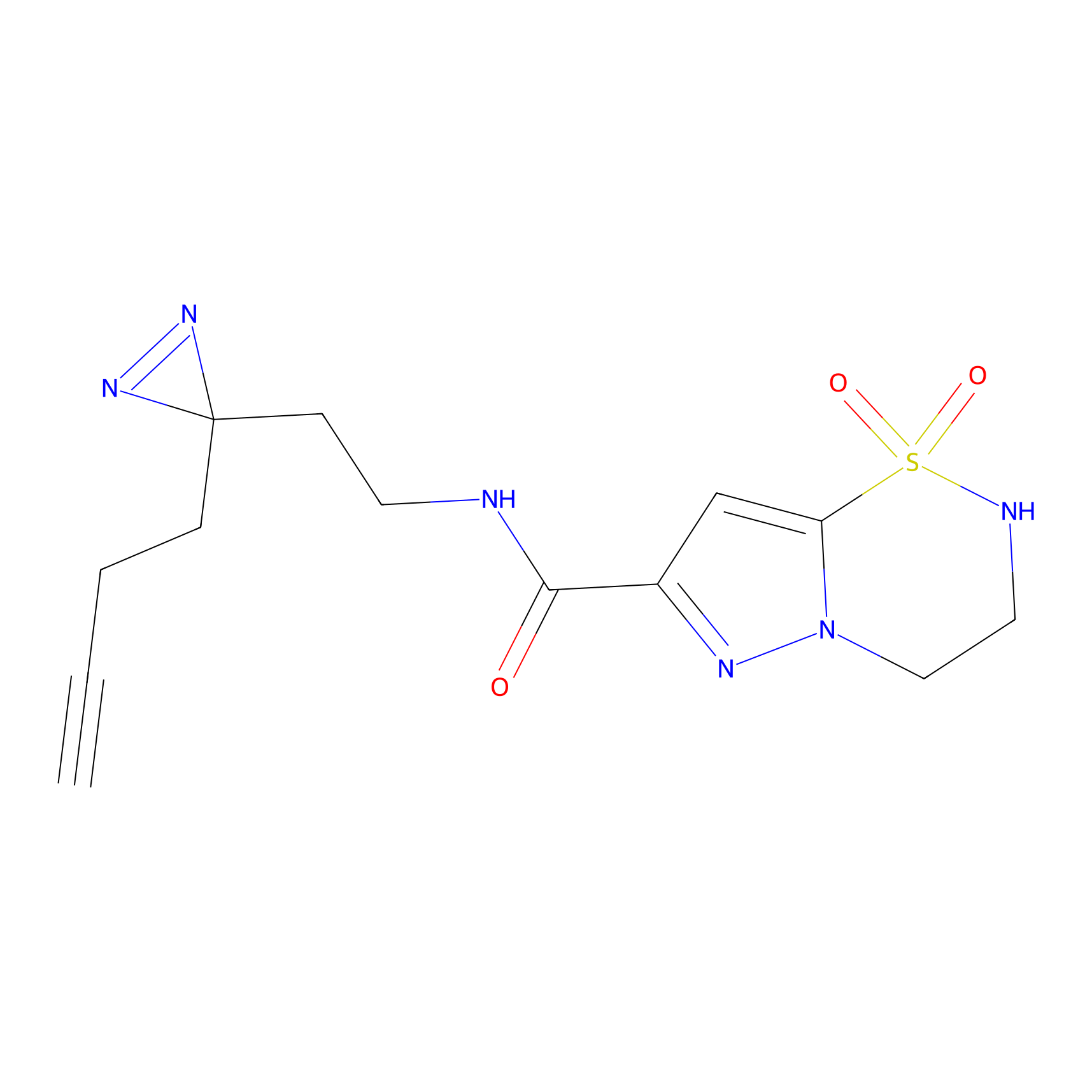
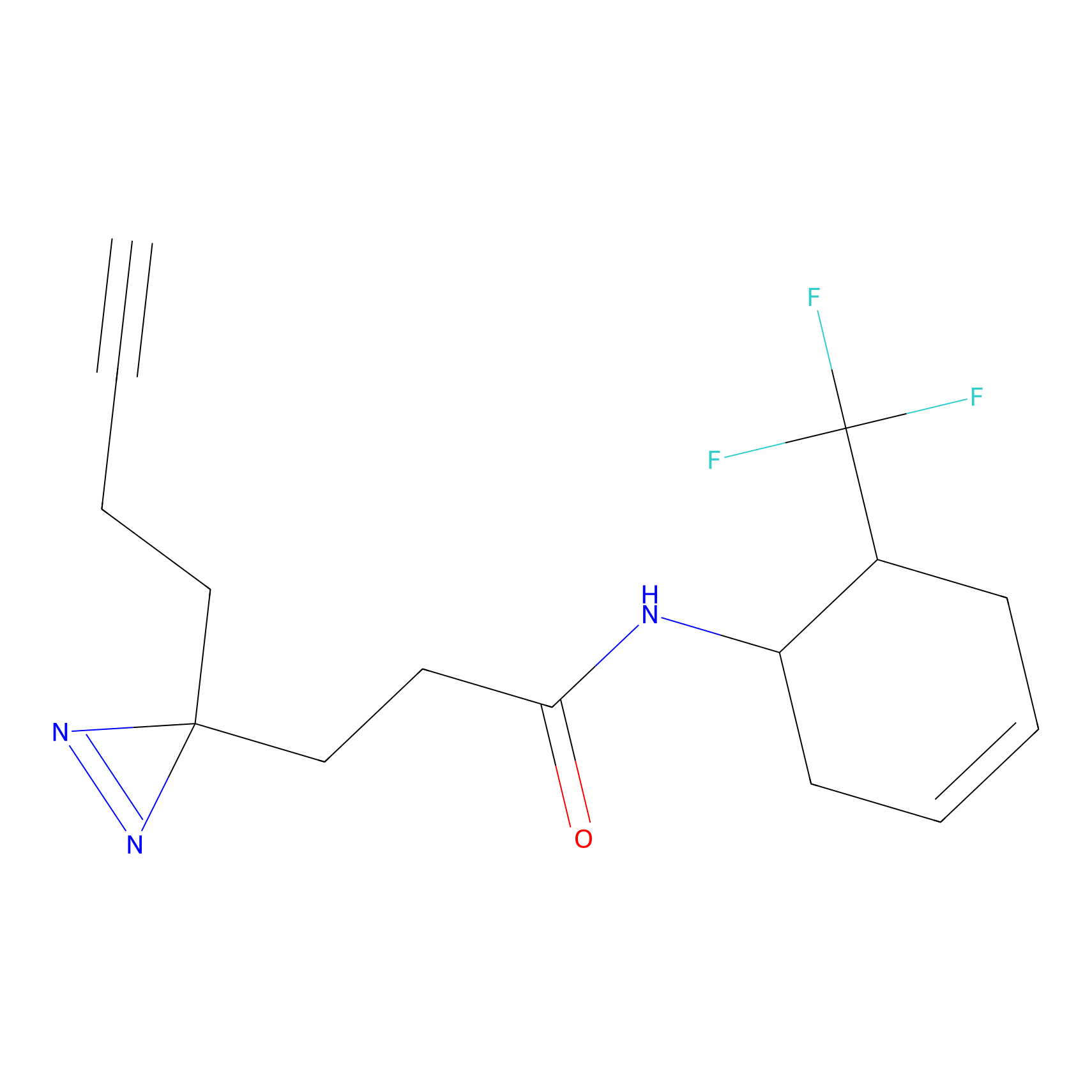
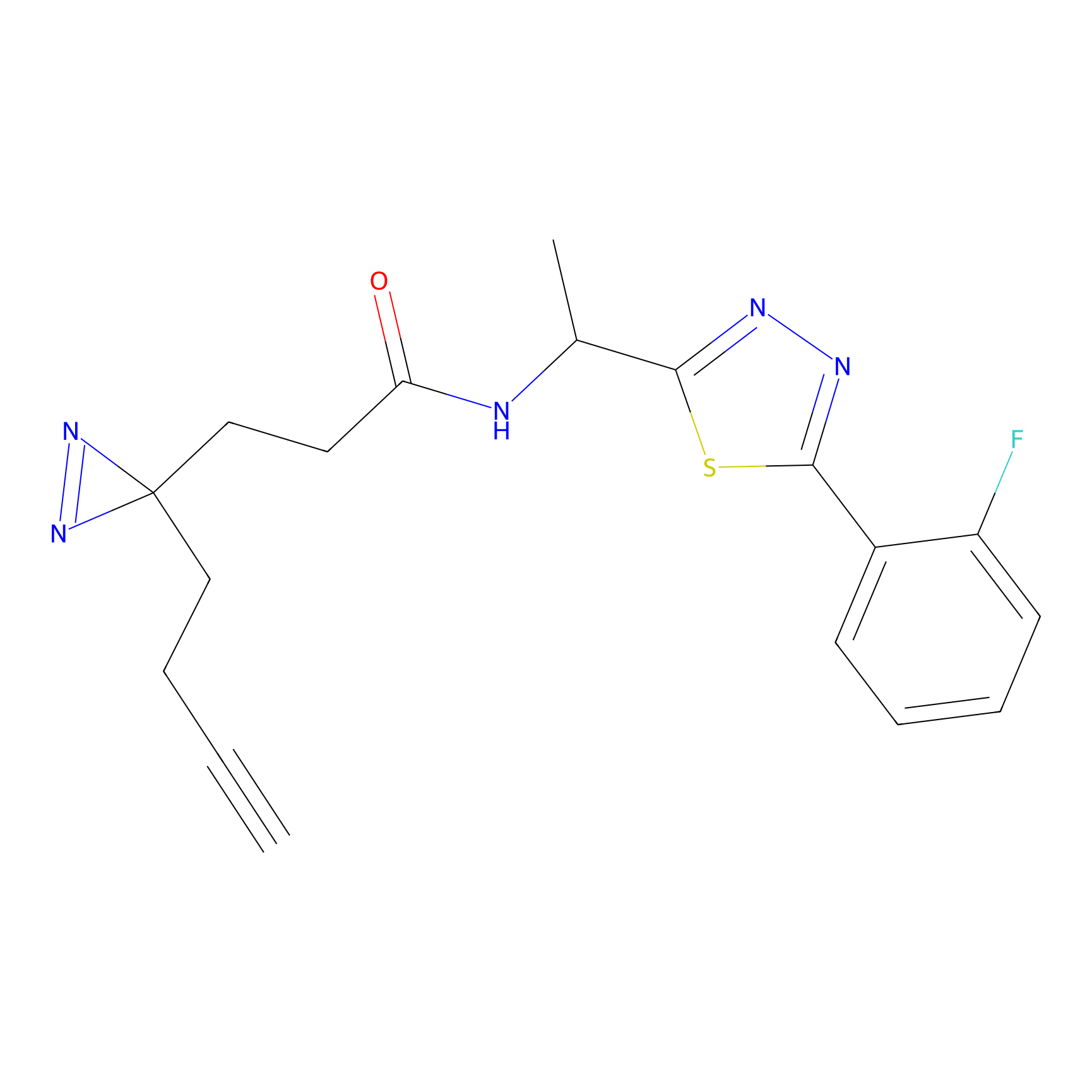
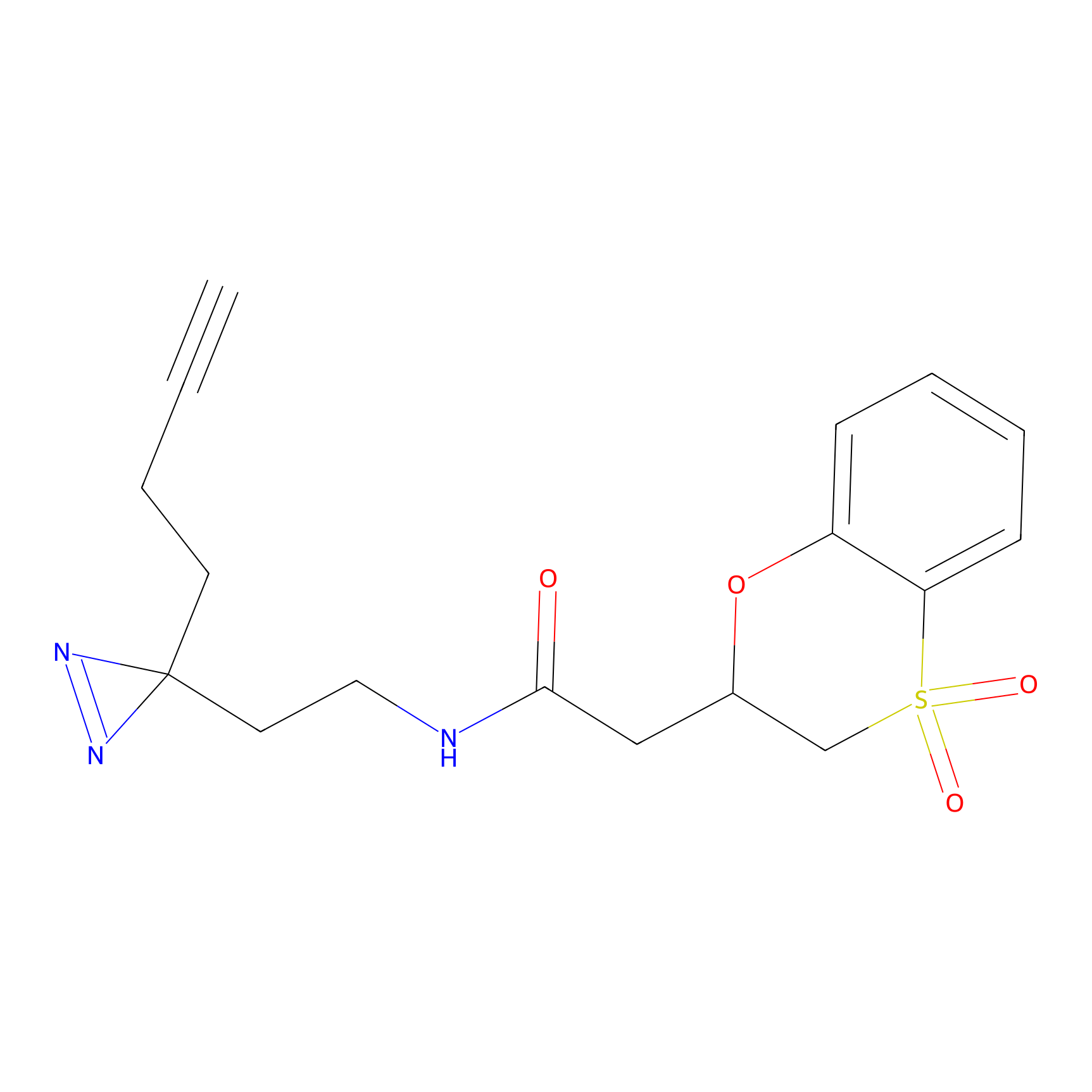
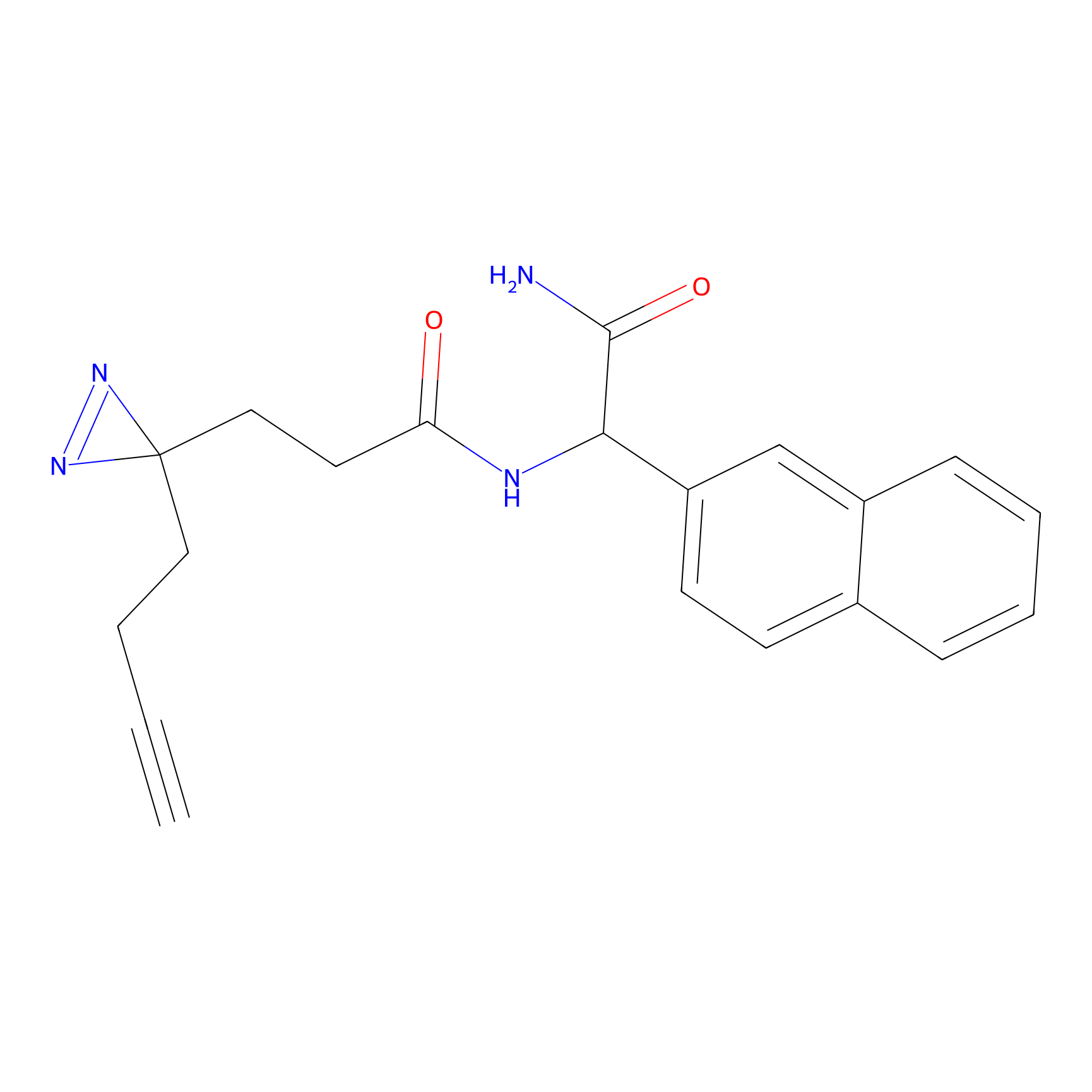
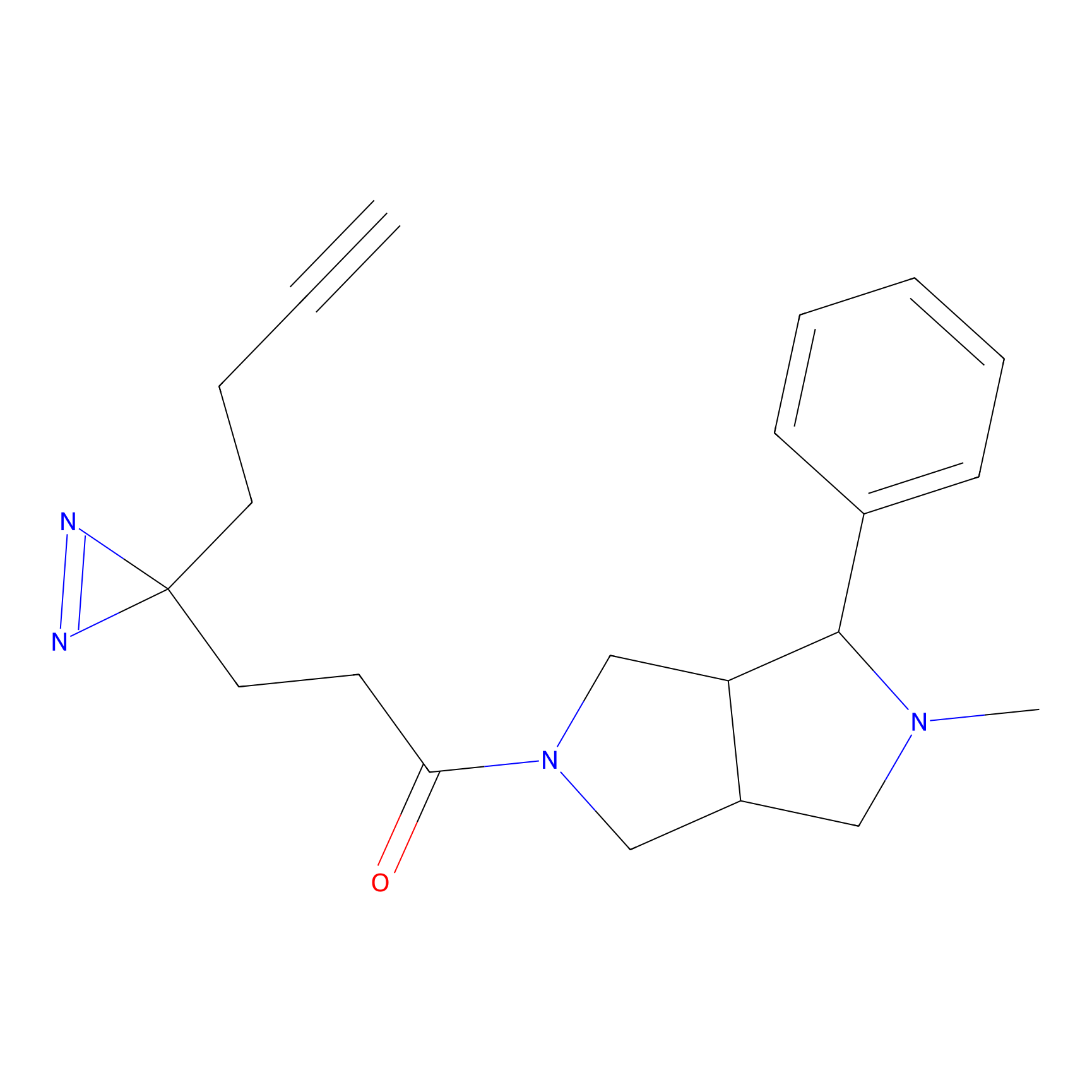
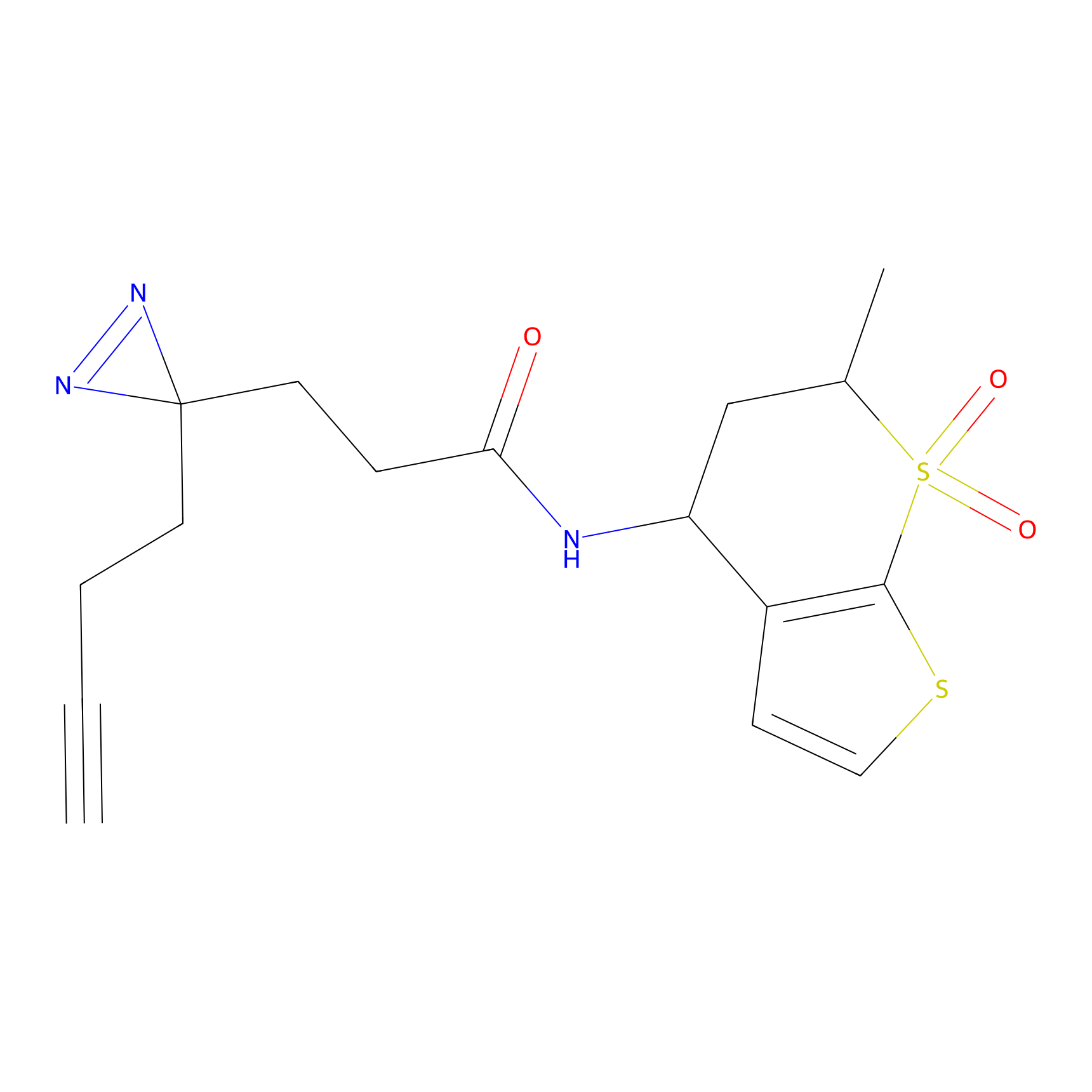
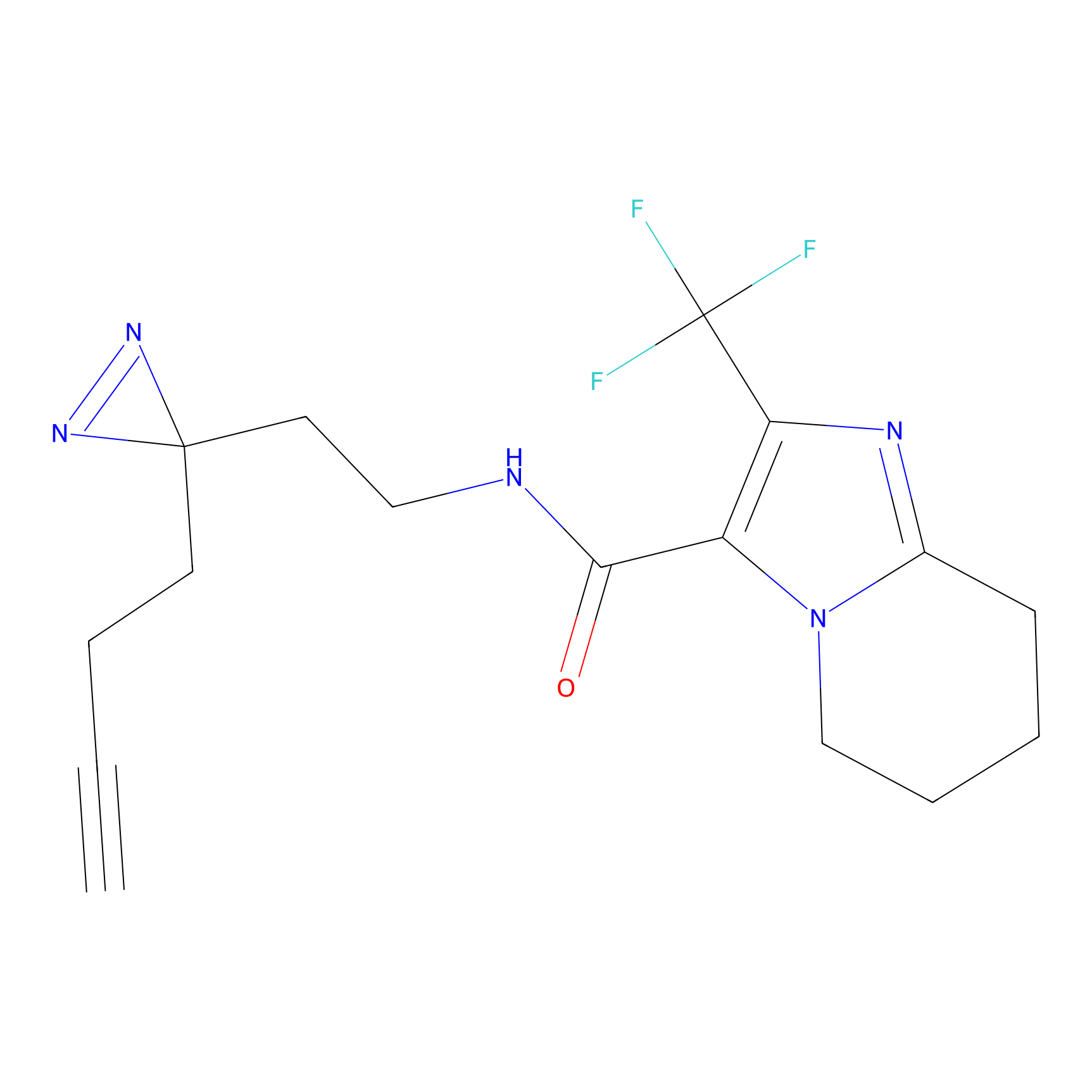
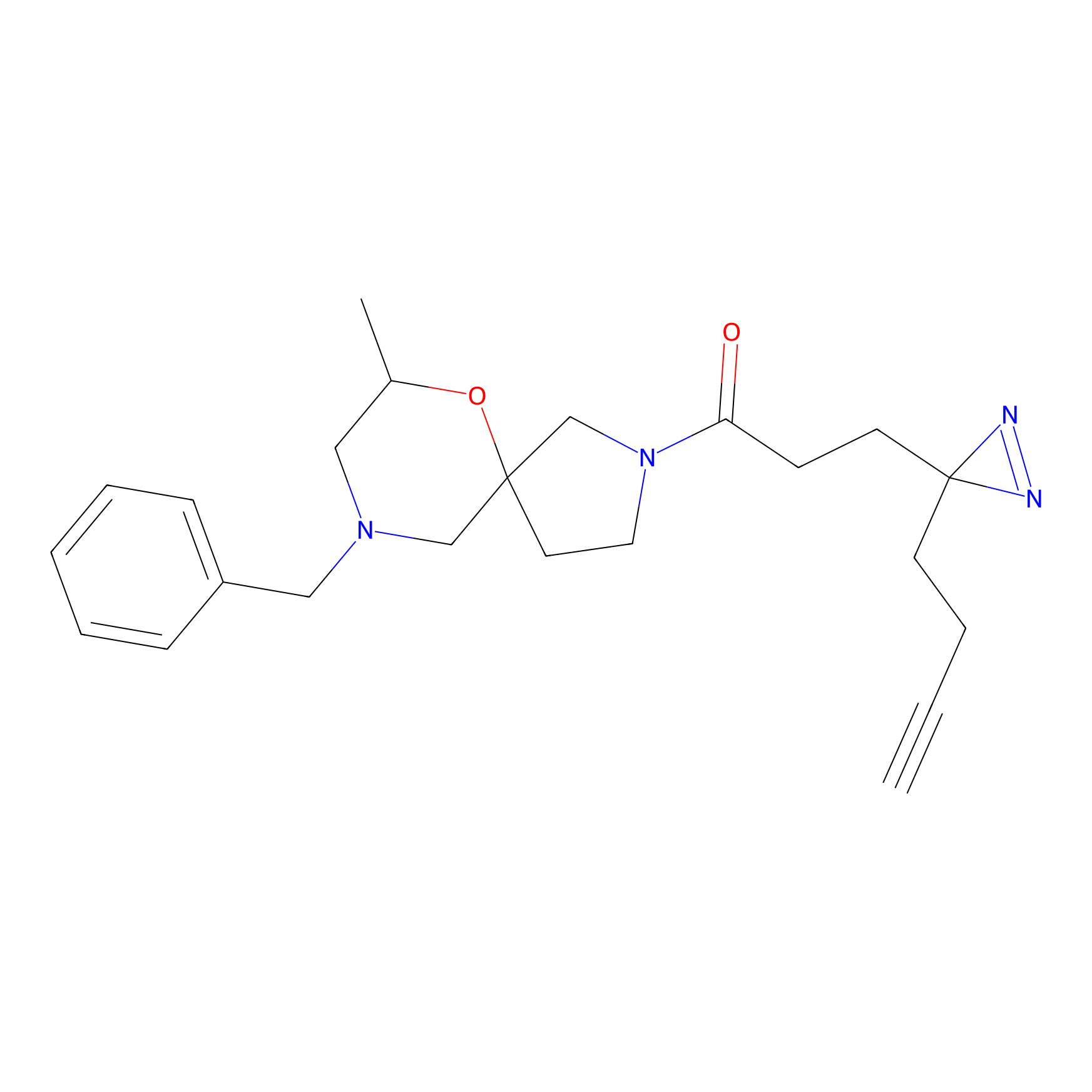
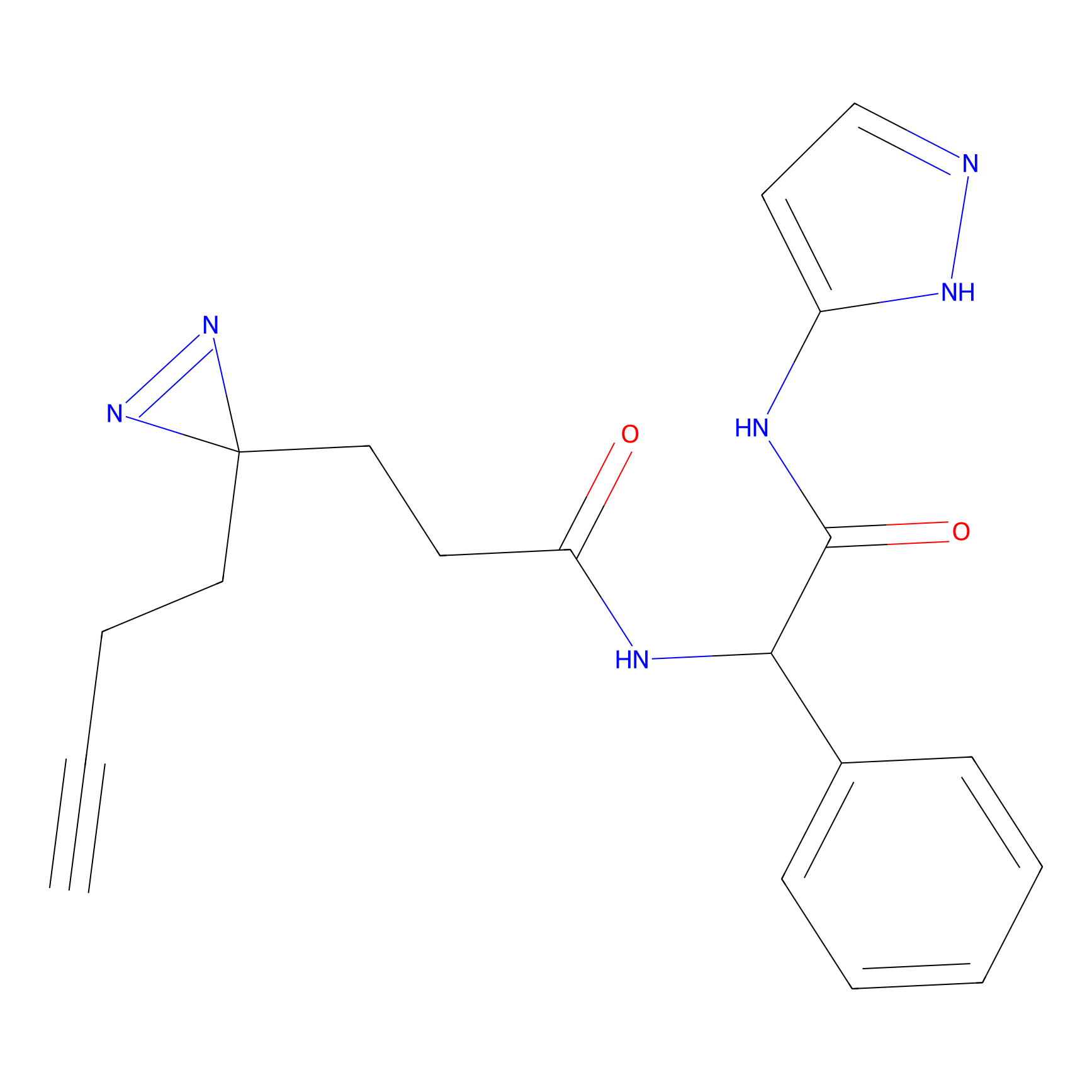
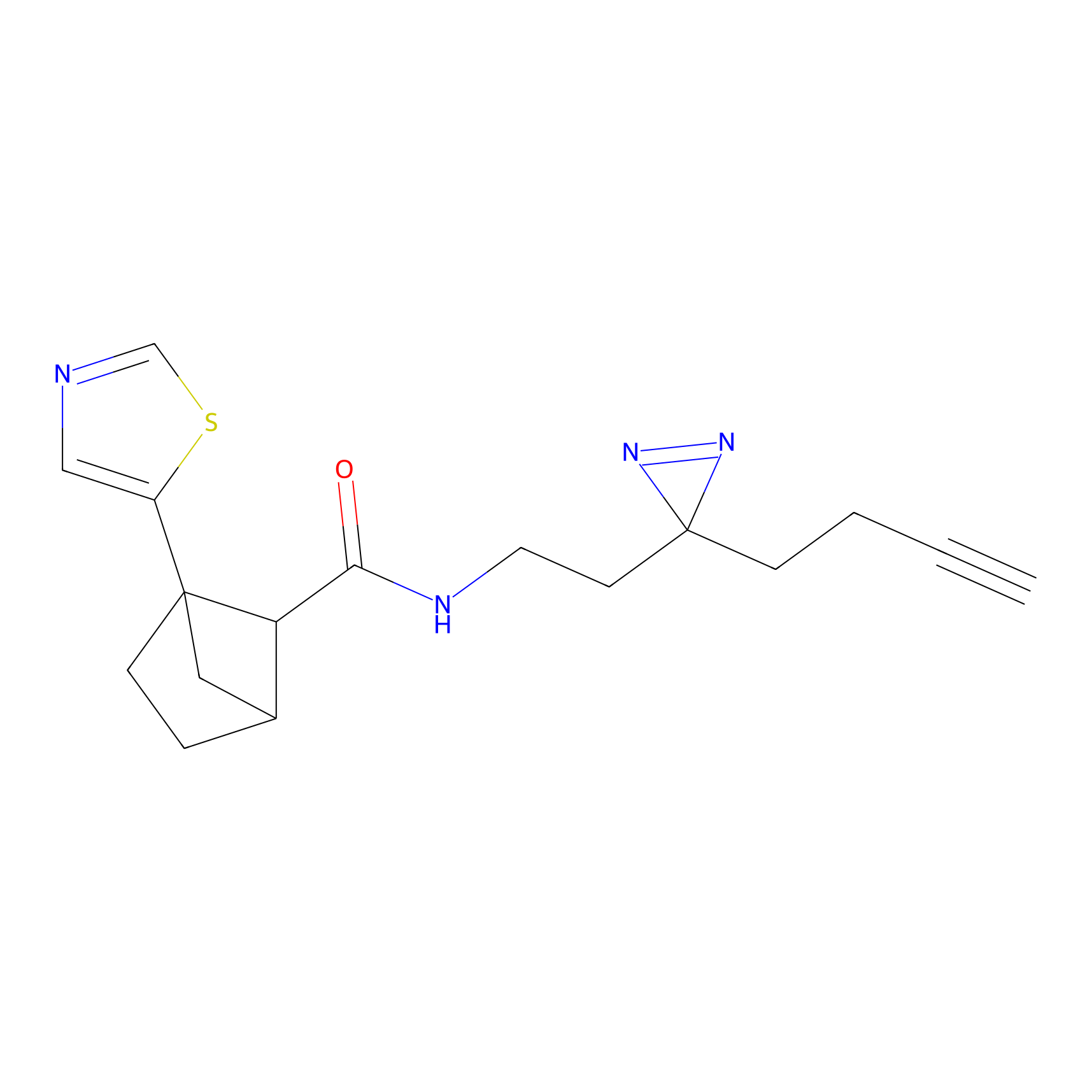
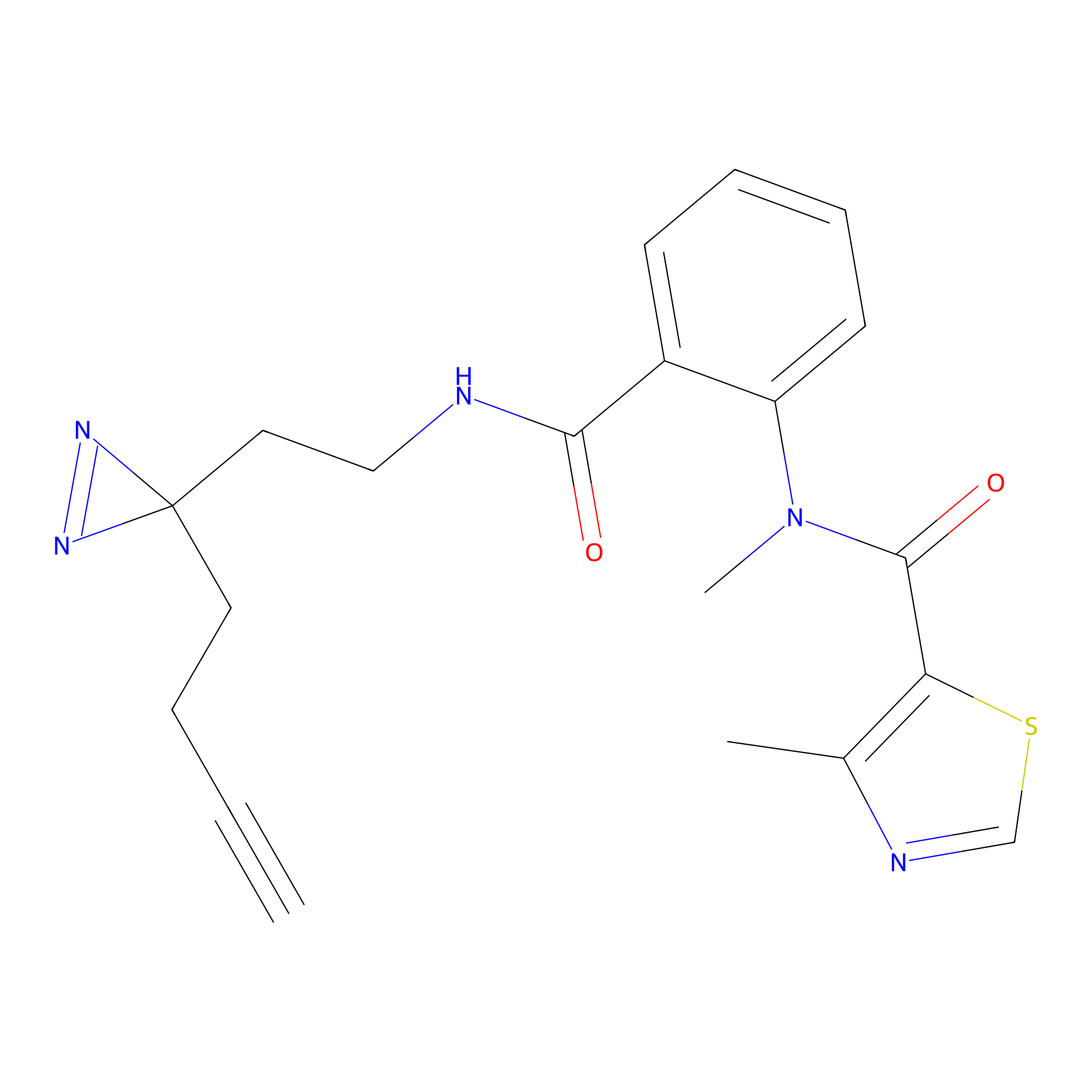
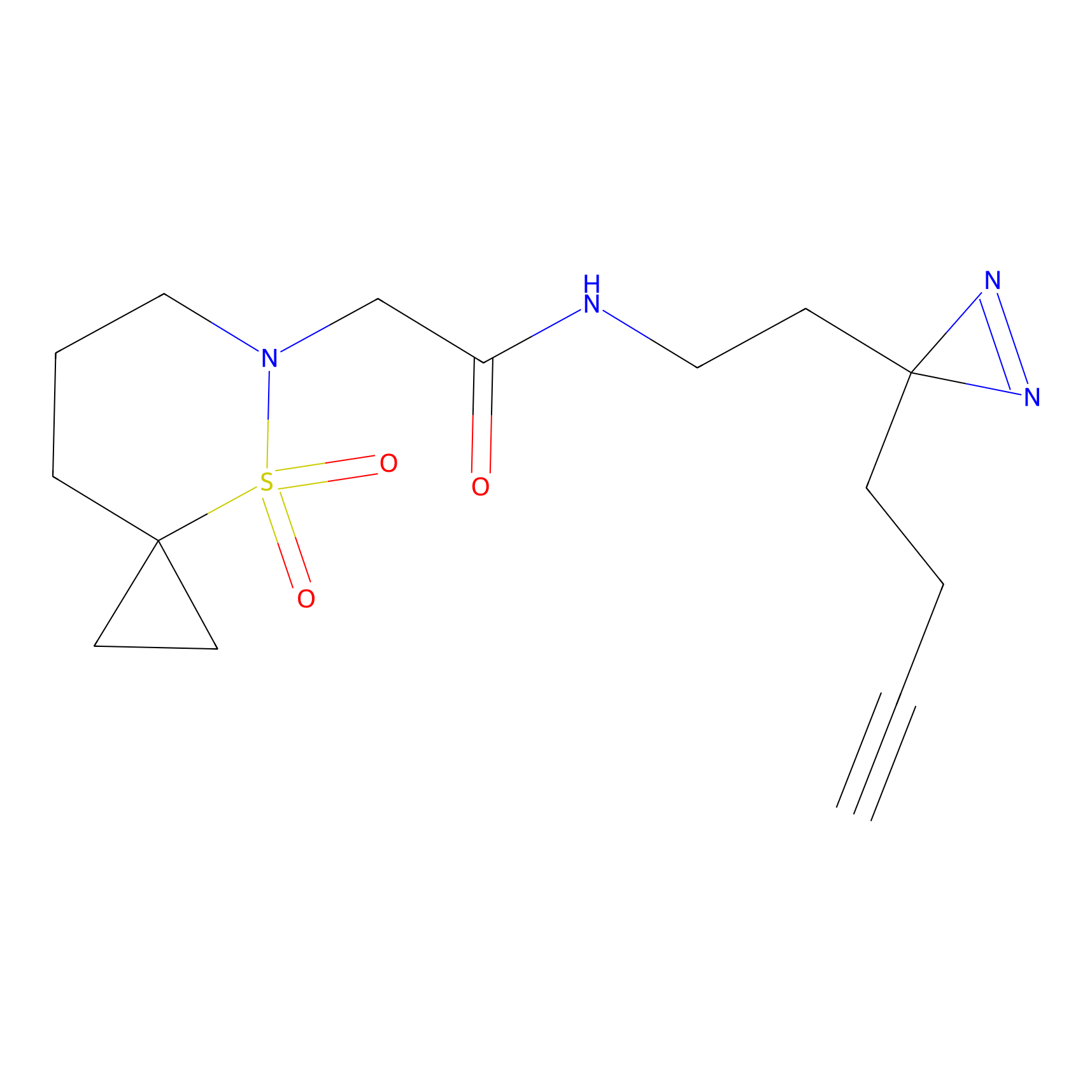
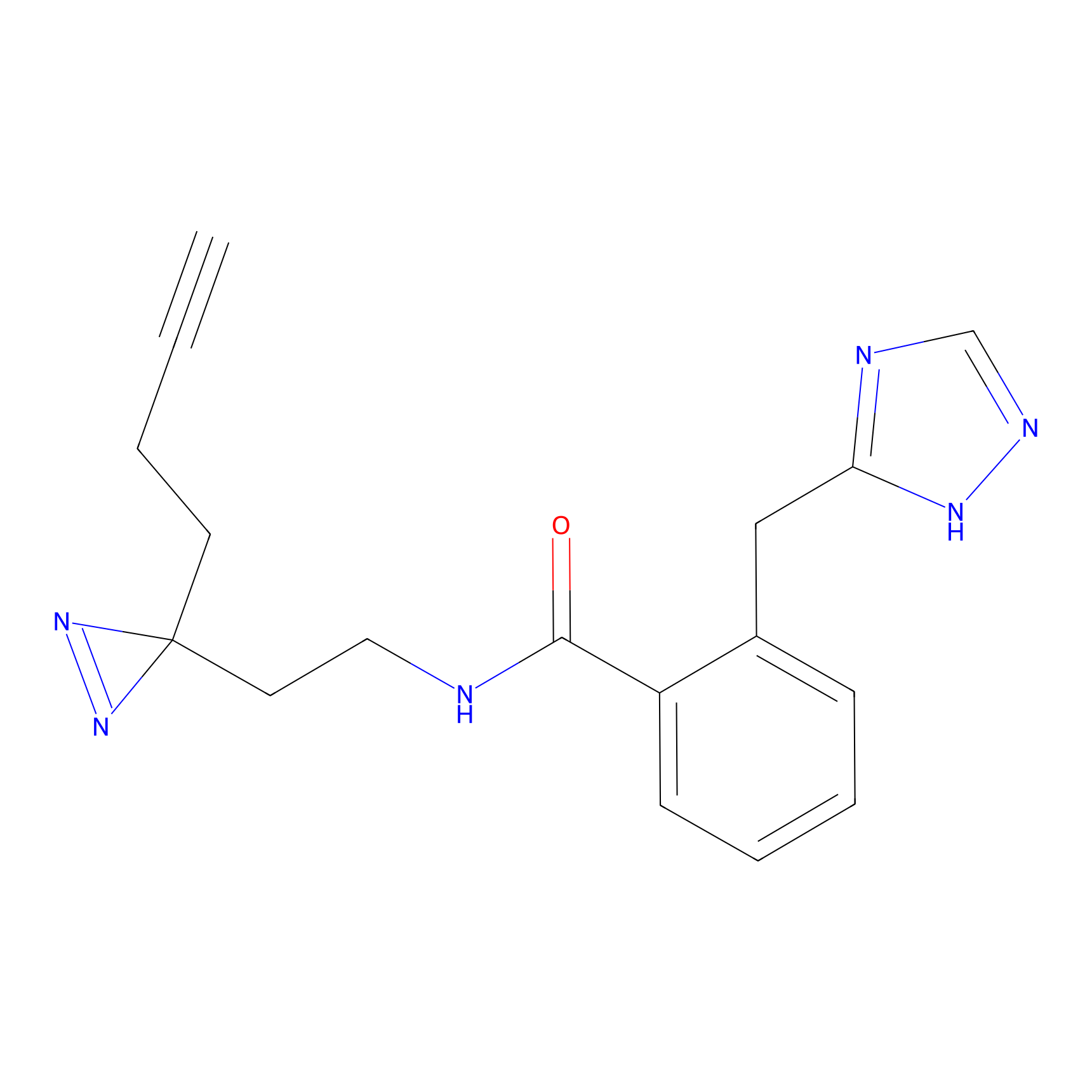
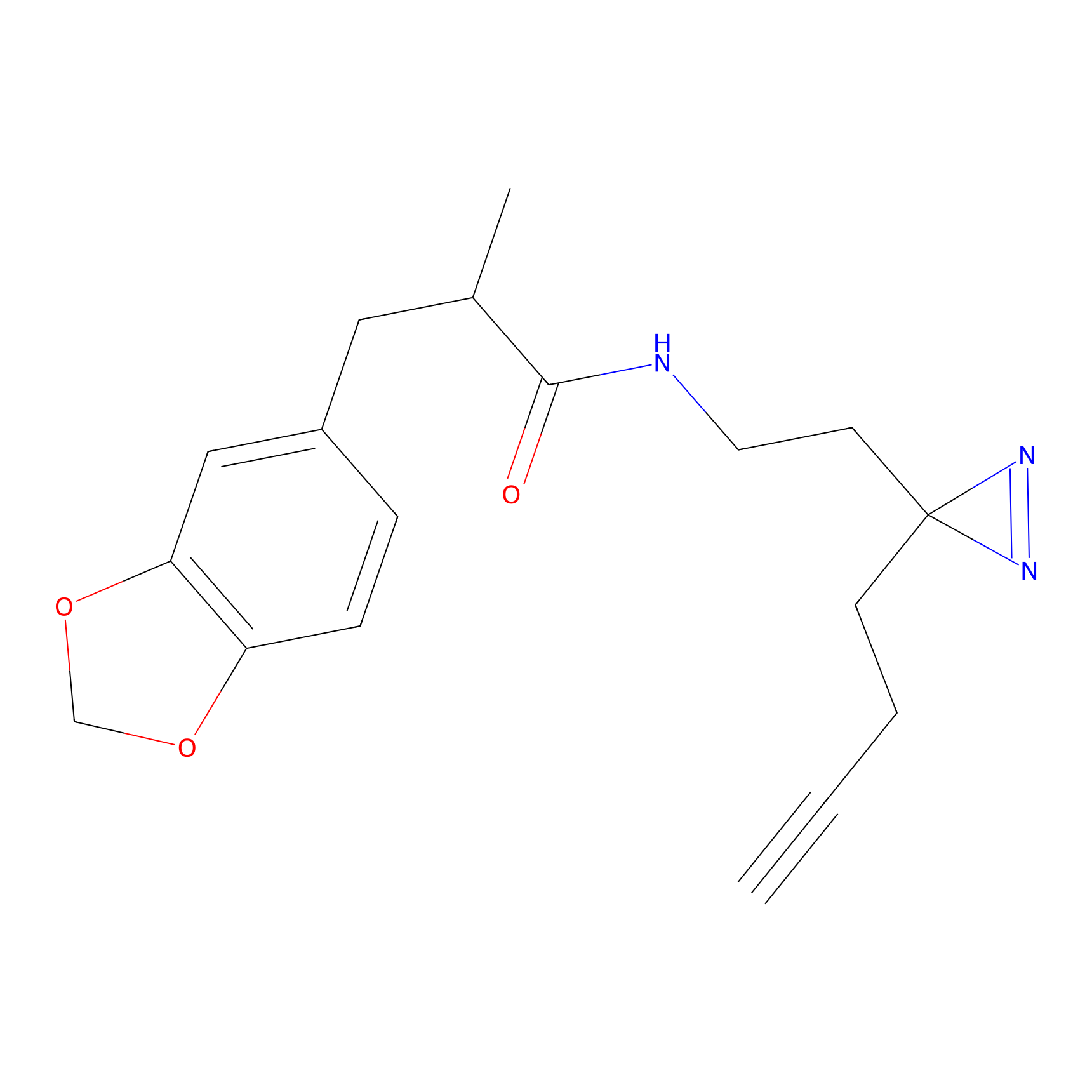
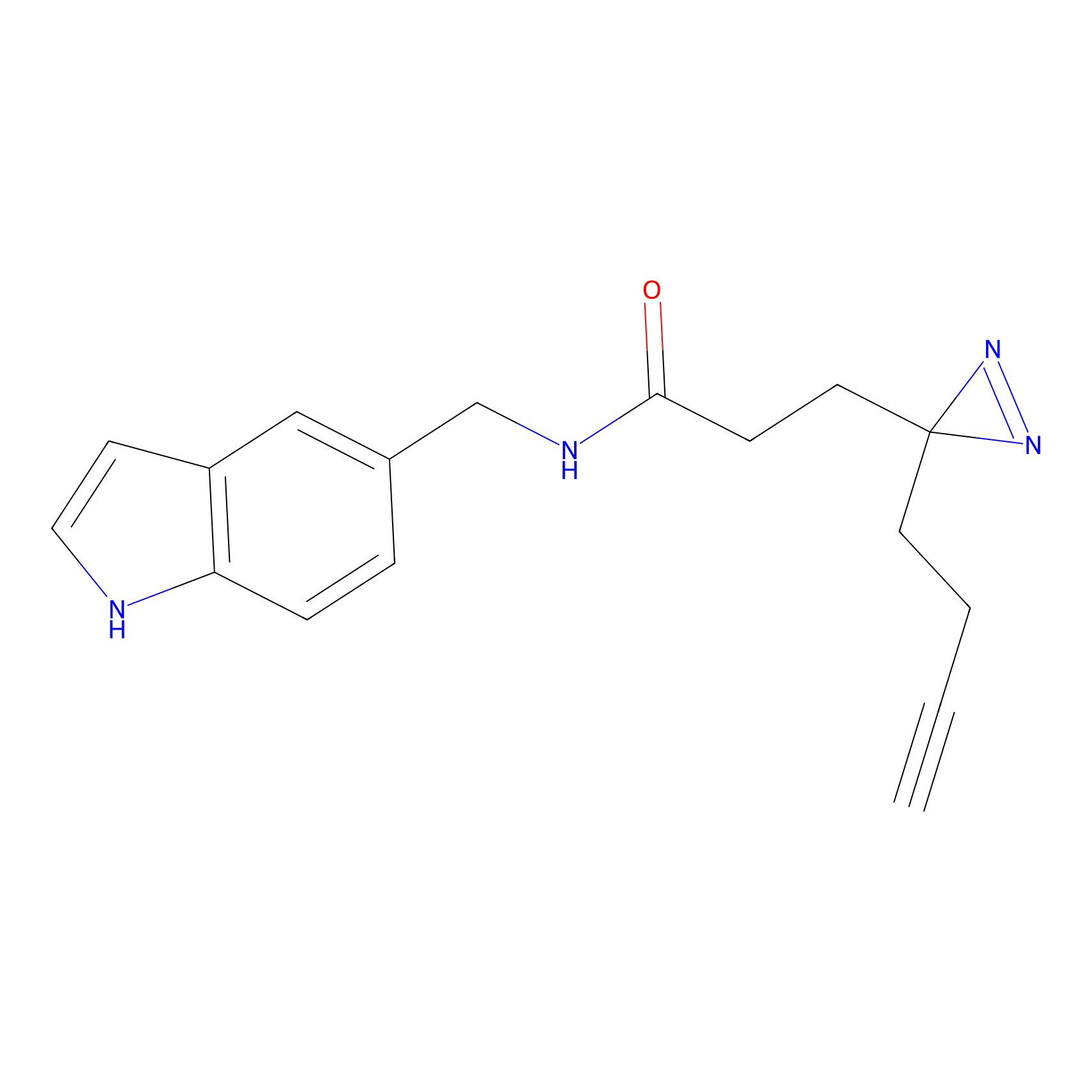
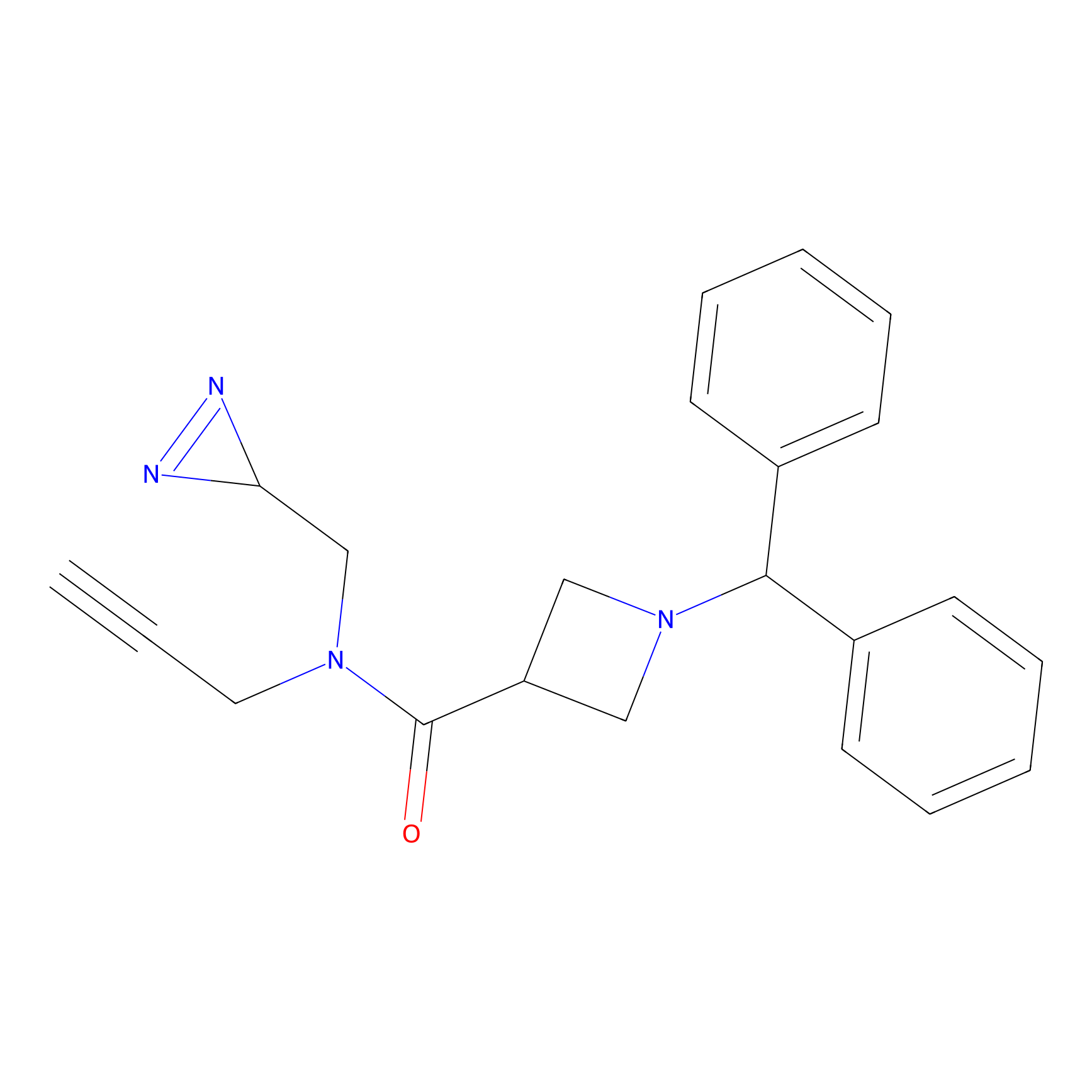
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
13.01 | LDD0318 | [1] | |
|
C-Sul Probe Info |
 |
3.01 | LDD0066 | [2] | |
|
ONAyne Probe Info |
 |
K177(9.64) | LDD0274 | [3] | |
|
STPyne Probe Info |
 |
K118(10.00); K177(10.00); K364(2.09); K400(7.69) | LDD0277 | [3] | |
|
OPA-S-S-alkyne Probe Info |
 |
K177(1.71); K302(2.22); K364(2.75); K183(3.16) | LDD3494 | [4] | |
|
Probe 1 Probe Info |
 |
Y179(55.31) | LDD3495 | [5] | |
|
5E-2FA Probe Info |
 |
H131(0.00); H415(0.00); H172(0.00); H395(0.00) | LDD2235 | [6] | |
|
ATP probe Probe Info |
 |
K421(0.00); K71(0.00); K218(0.00) | LDD0199 | [7] | |
|
m-APA Probe Info |
 |
H131(0.00); H415(0.00); H172(0.00); H395(0.00) | LDD2231 | [6] | |
|
Acrolein Probe Info |
 |
H460(0.00); H415(0.00); H423(0.00); H67(0.00) | LDD0217 | [8] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H415(0.00); H460(0.00); H423(0.00) | LDD0222 | [8] |
| LDCM0185 | Compound 17 | HEK-293T | 3.98 | LDD0512 | [10] |
| LDCM0191 | Compound 21 | HEK-293T | 9.95 | LDD0508 | [10] |
| LDCM0190 | Compound 34 | HEK-293T | 9.76 | LDD0510 | [10] |
| LDCM0192 | Compound 35 | HEK-293T | 9.43 | LDD0509 | [10] |
| LDCM0193 | Compound 36 | HEK-293T | 10.05 | LDD0511 | [10] |
| LDCM0107 | IAA | HeLa | H460(0.00); H423(0.00) | LDD0221 | [8] |
| LDCM0109 | NEM | HeLa | H172(0.00); H67(0.00); H460(0.00); H415(0.00) | LDD0223 | [8] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Amyloid-beta precursor protein (APP) | APP family | P05067 | |||
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Calcium Phosphate | Small molecular drug | DB11348 | |||
| Calcium Phosphate Dihydrate | Small molecular drug | DB14481 | |||
| Calcium Citrate | . | DB11093 | |||
Investigative
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Kml110 | Small molecular drug | D0S9AT | |||
| Mjn228 | Small molecular drug | D01XLZ | |||
References