Details of the Target
General Information of Target
| Target ID | LDTP13902 | |||||
|---|---|---|---|---|---|---|
| Target Name | Cytochrome c oxidase assembly factor 3 homolog, mitochondrial (COA3) | |||||
| Gene Name | COA3 | |||||
| Gene ID | 28958 | |||||
| Synonyms |
CCDC56; MITRAC12; Cytochrome c oxidase assembly factor 3 homolog, mitochondrial; Coiled-coil domain-containing protein 56; Mitochondrial translation regulation assembly intermediate of cytochrome c oxidase protein of 12 kDa
|
|||||
| 3D Structure | ||||||
| Sequence |
MLRPKALTQVLSQANTGGVQSTLLLNNEGSLLAYSGYGDTDARVTAAIASNIWAAYDRNG
NQAFNEDNLKFILMDCMEGRVAITRVANLLLCMYAKETVGFGMLKAKAQALVQYLEEPLT QVAAS |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
COA3 family
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Core component of the MITRAC (mitochondrial translation regulation assembly intermediate of cytochrome c oxidase complex) complex, that regulates cytochrome c oxidase assembly. MITRAC complexes regulate both translation of mitochondrial encoded components and assembly of nuclear-encoded components imported in mitochondrion. Required for efficient translation of MT-CO1 and mitochondrial respiratory chain complex IV assembly.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
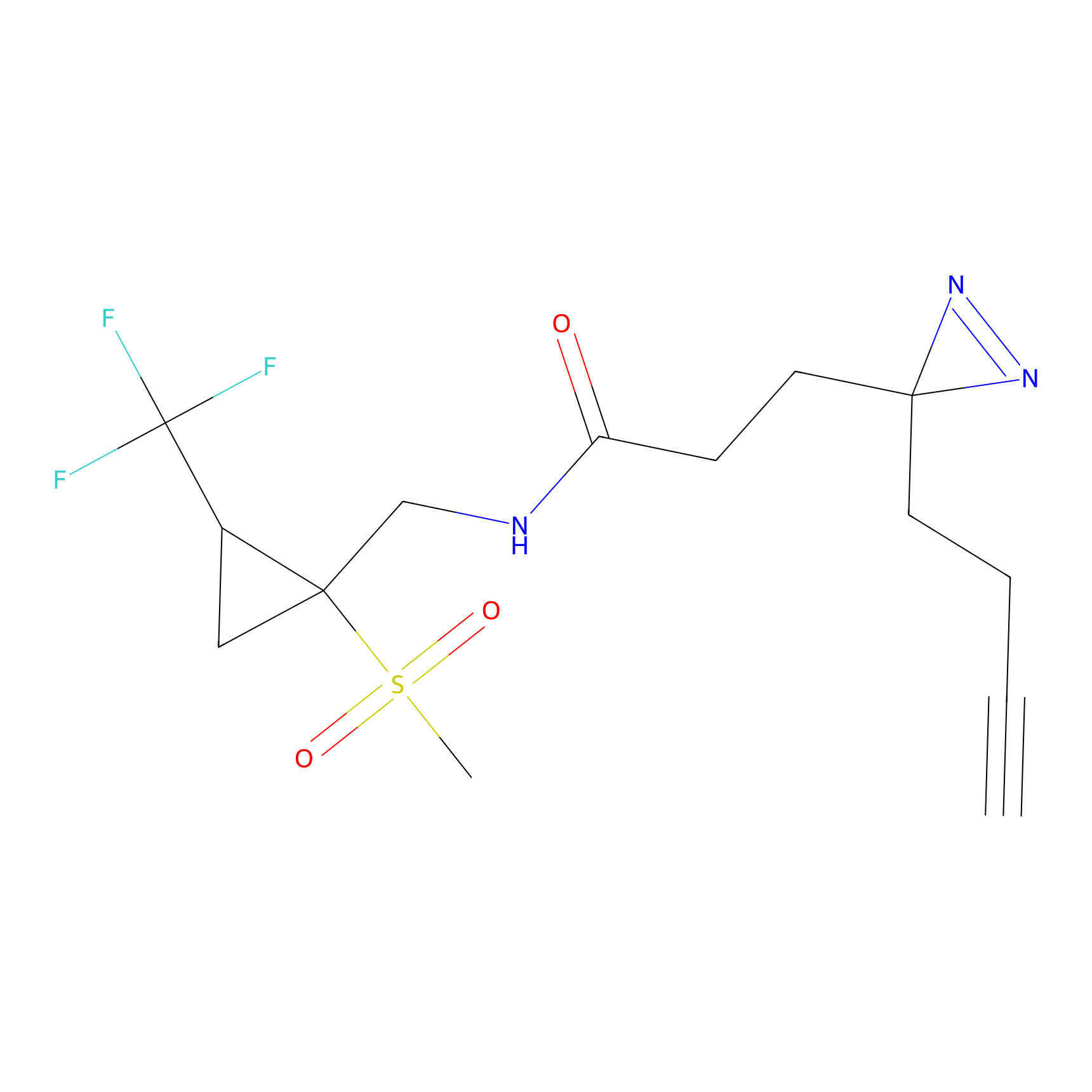
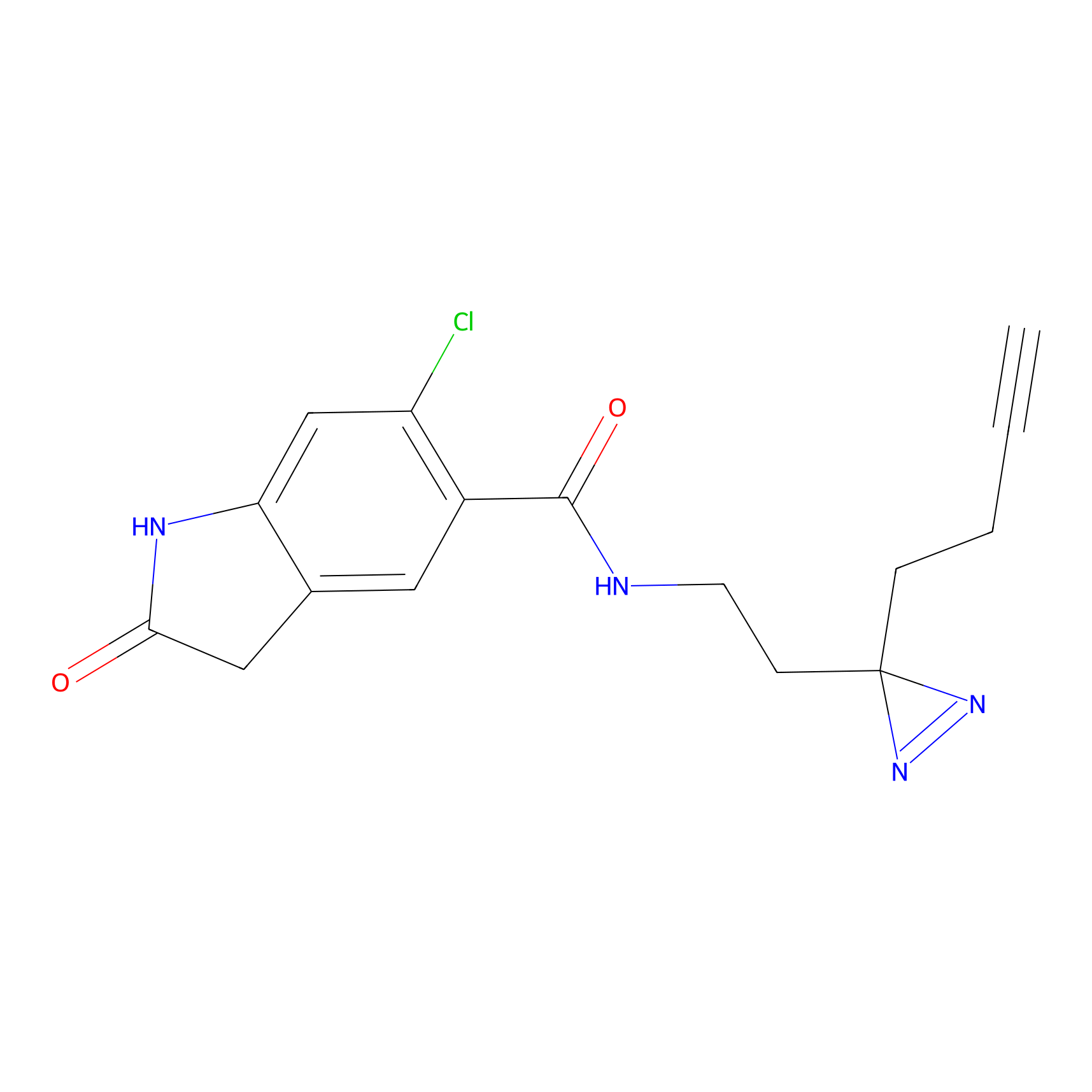
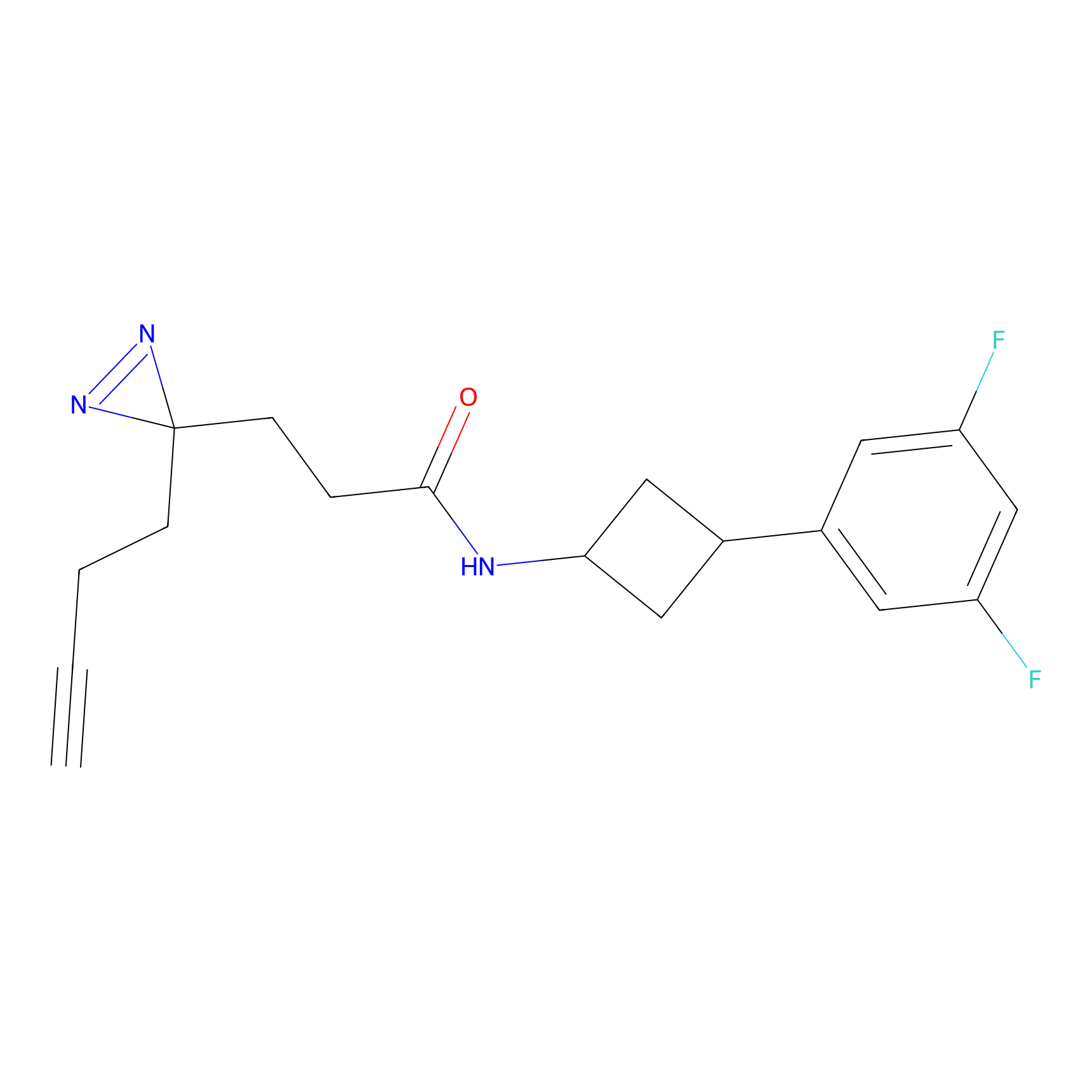
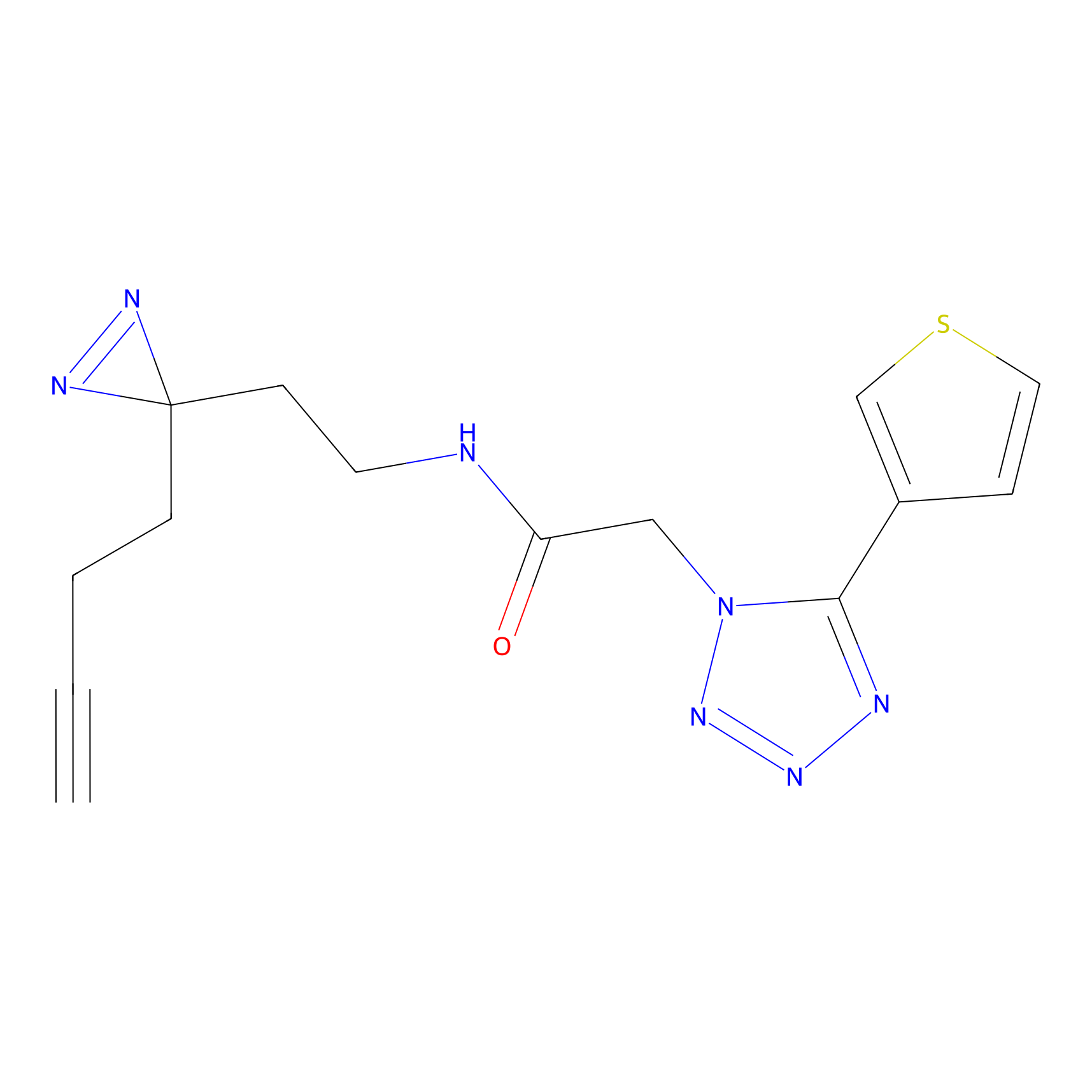
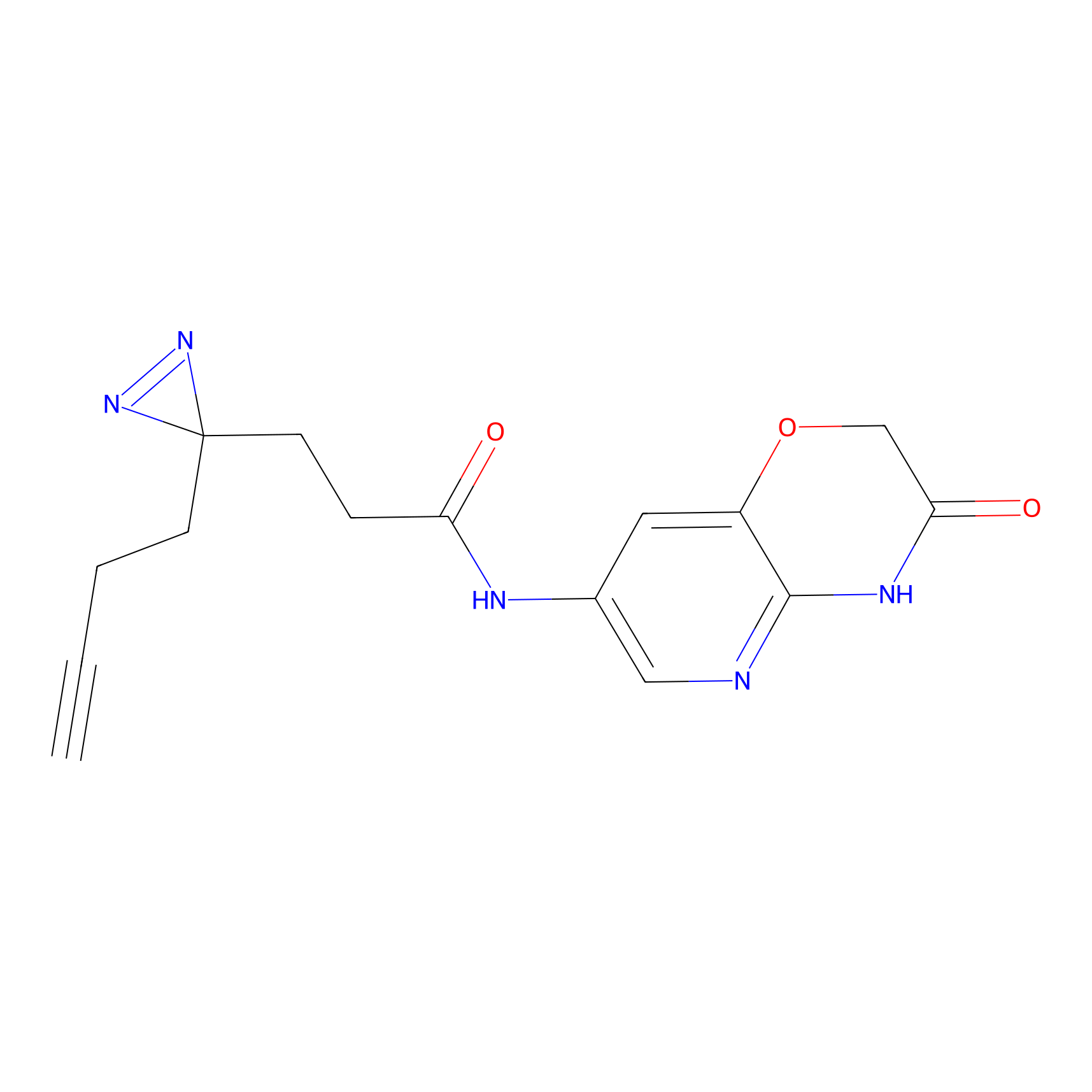
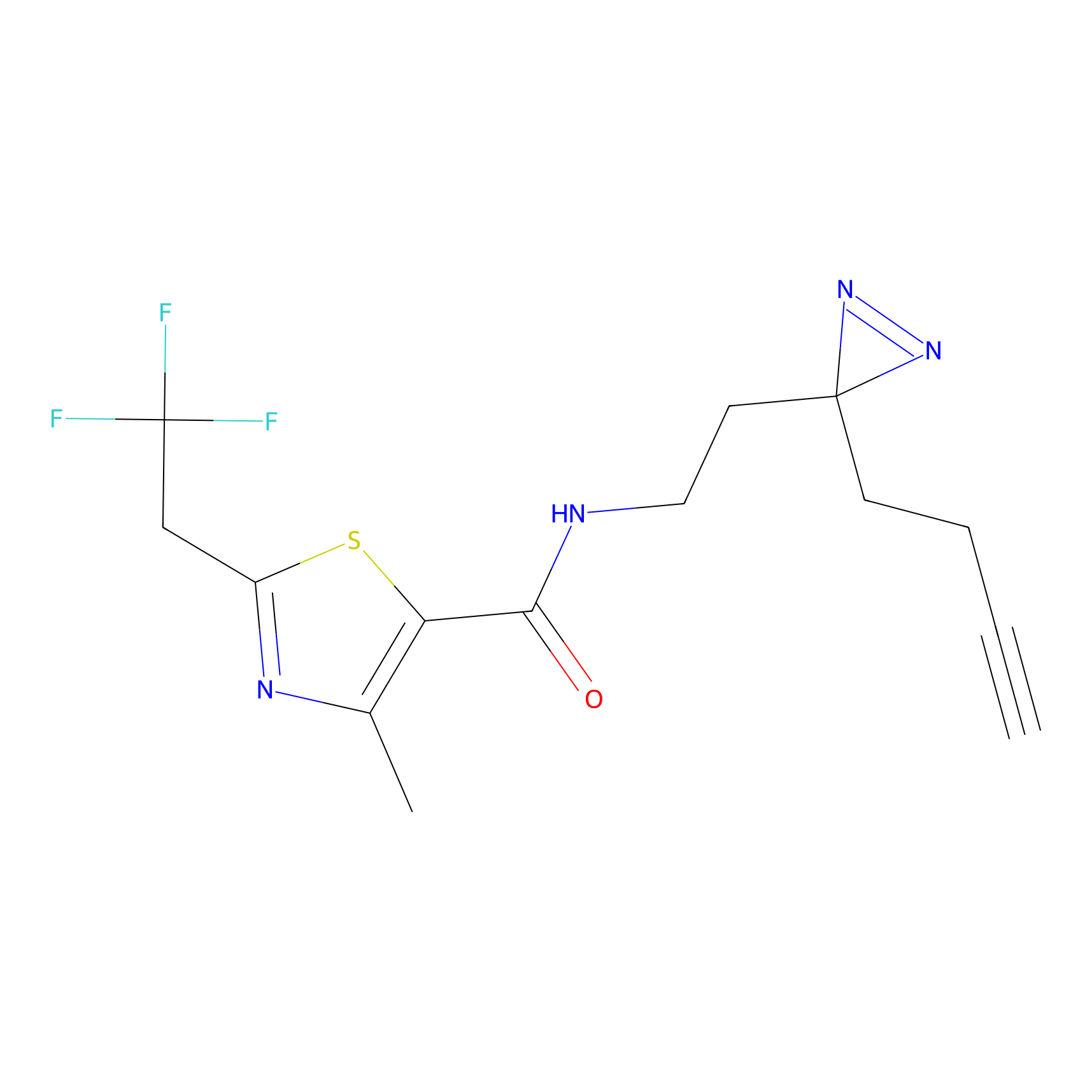
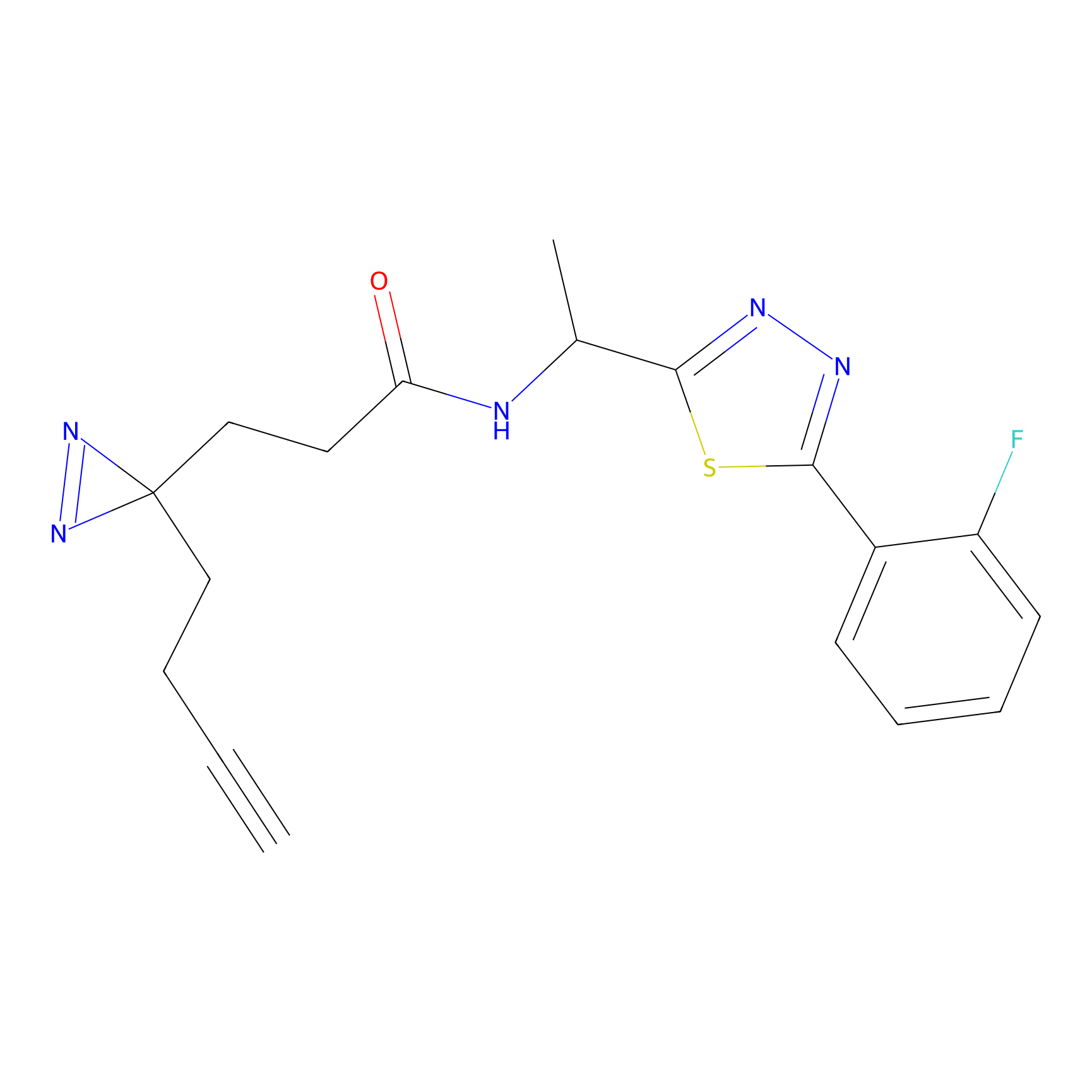
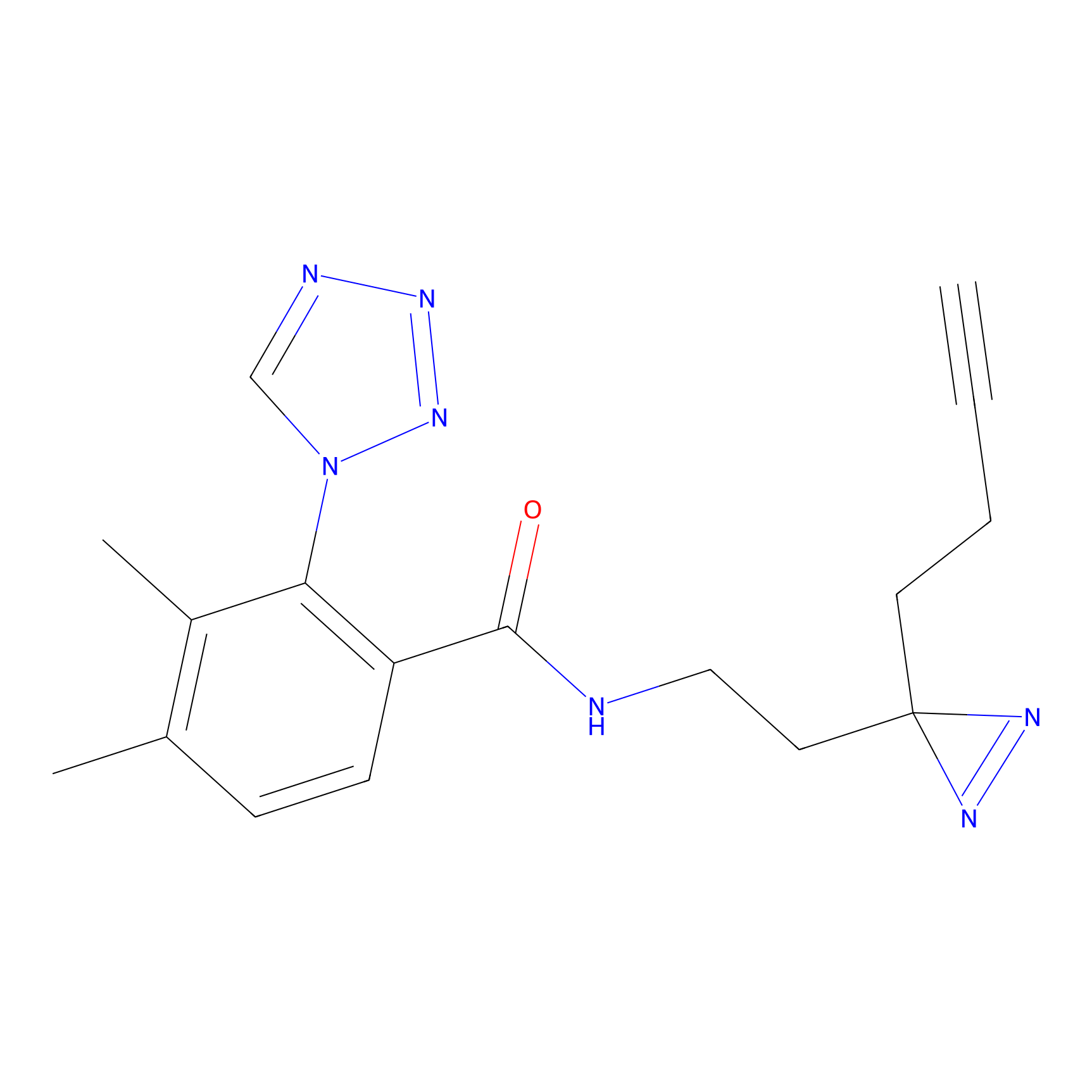
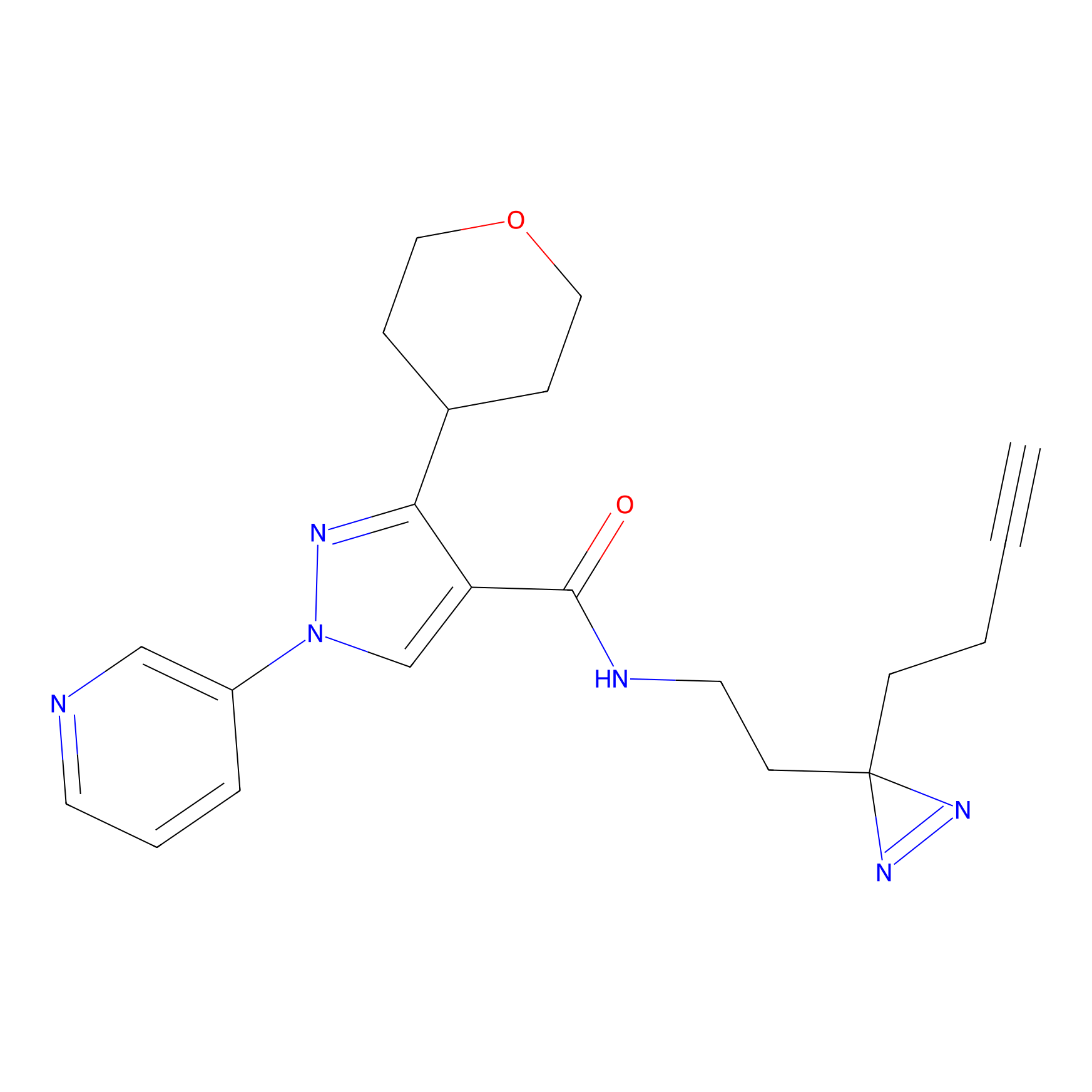
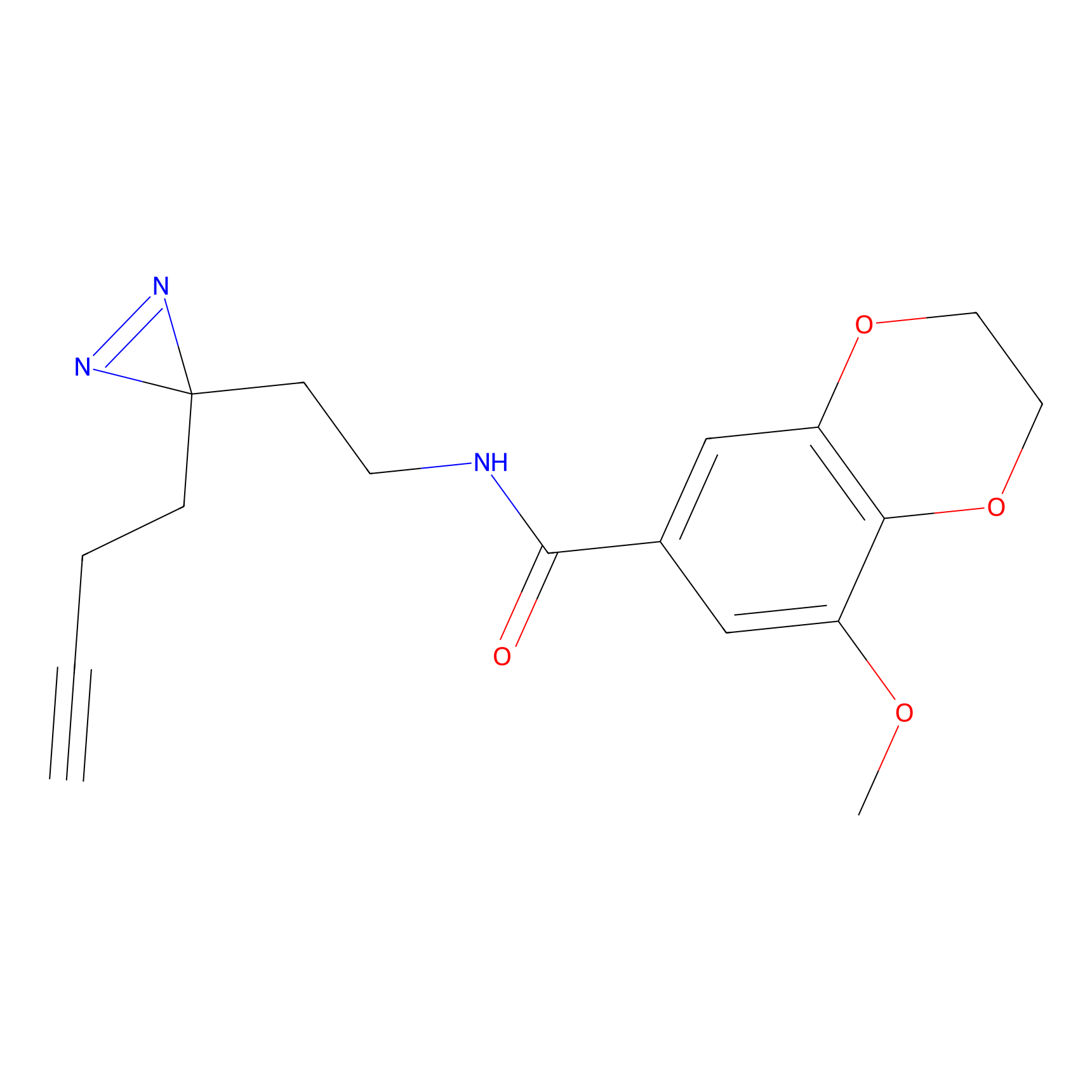
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
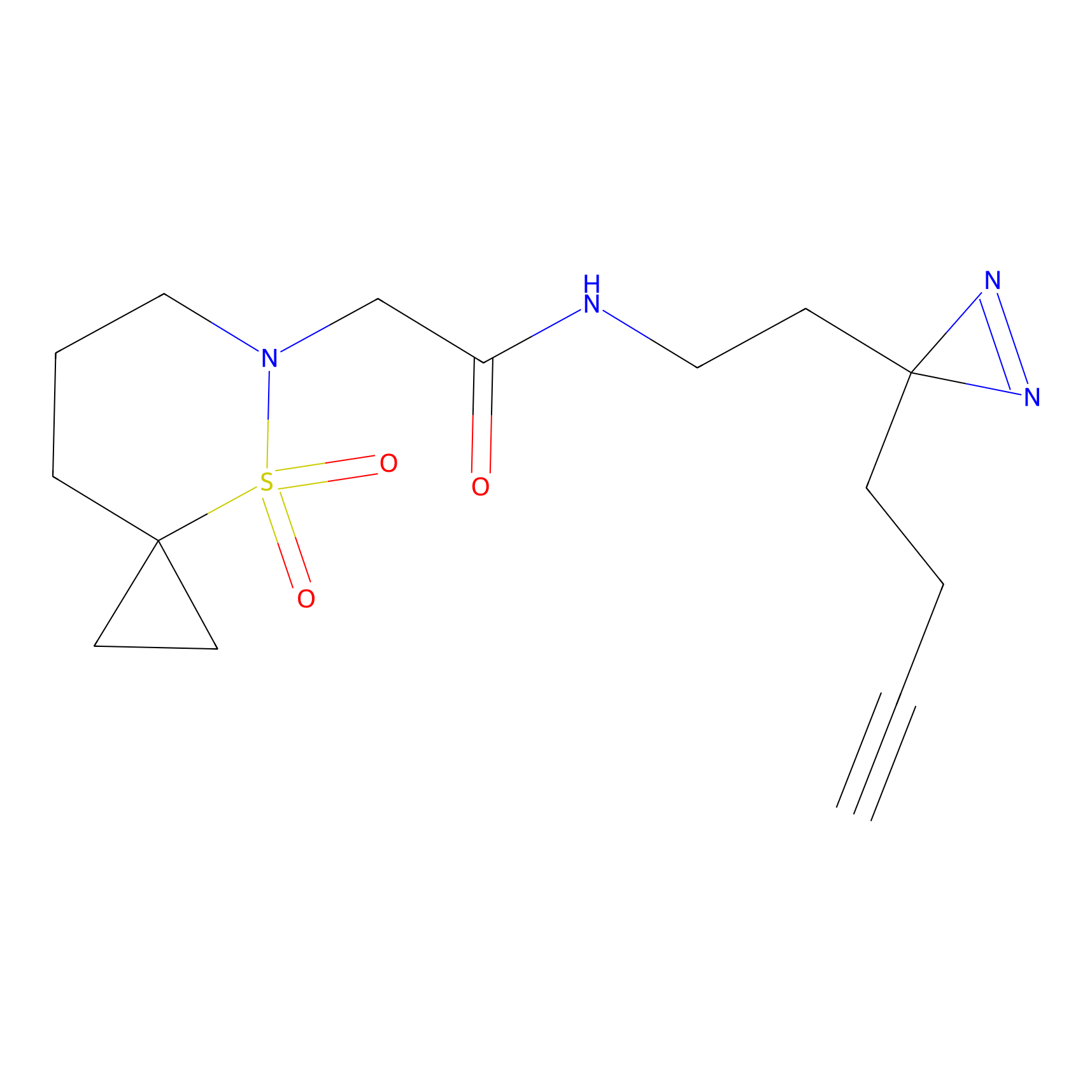
STPyne Probe Info |
 |
K48(6.75) | LDD0277 | [1] | |
|
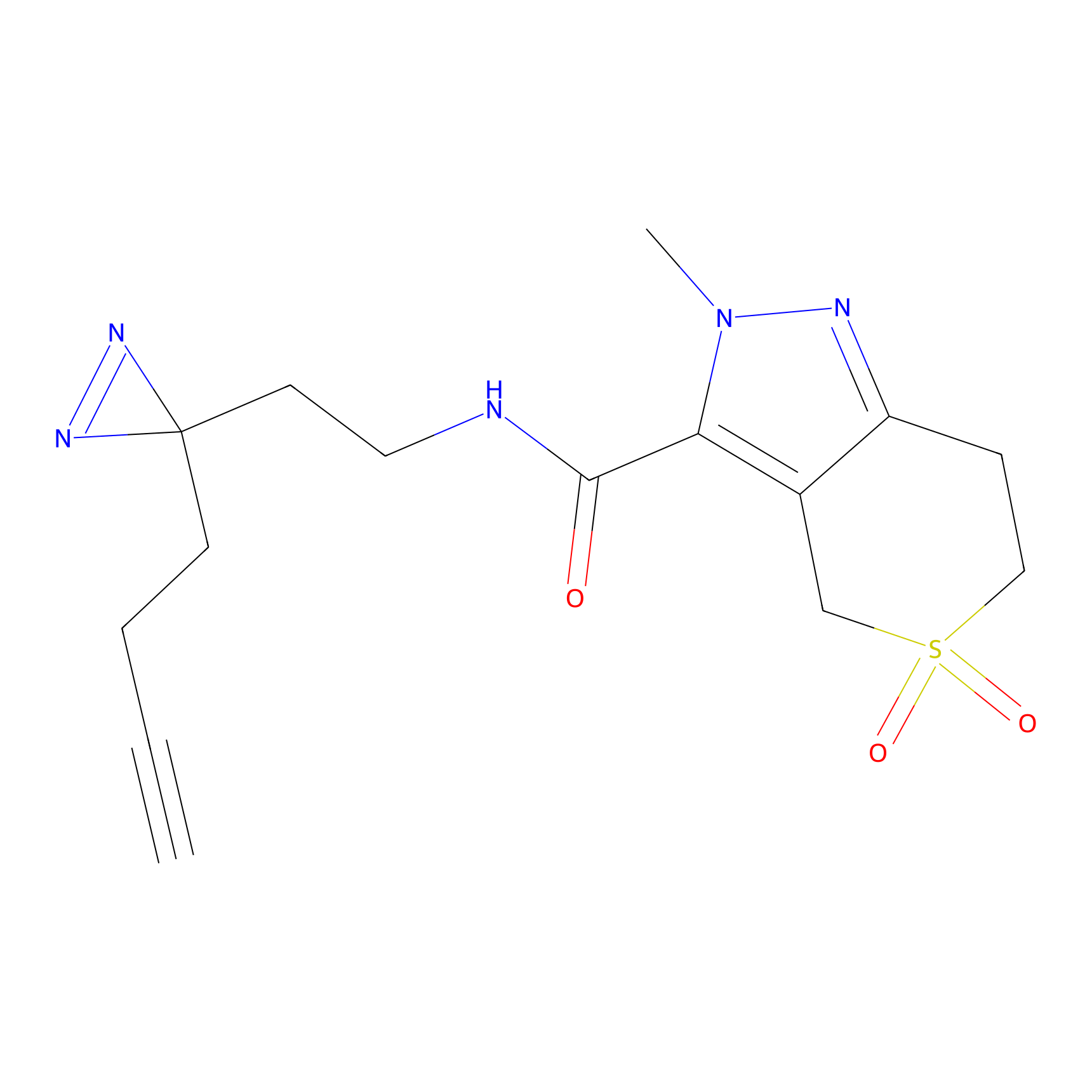
AZ-9 Probe Info |
 |
D86(10.00) | LDD2209 | [2] | |
|
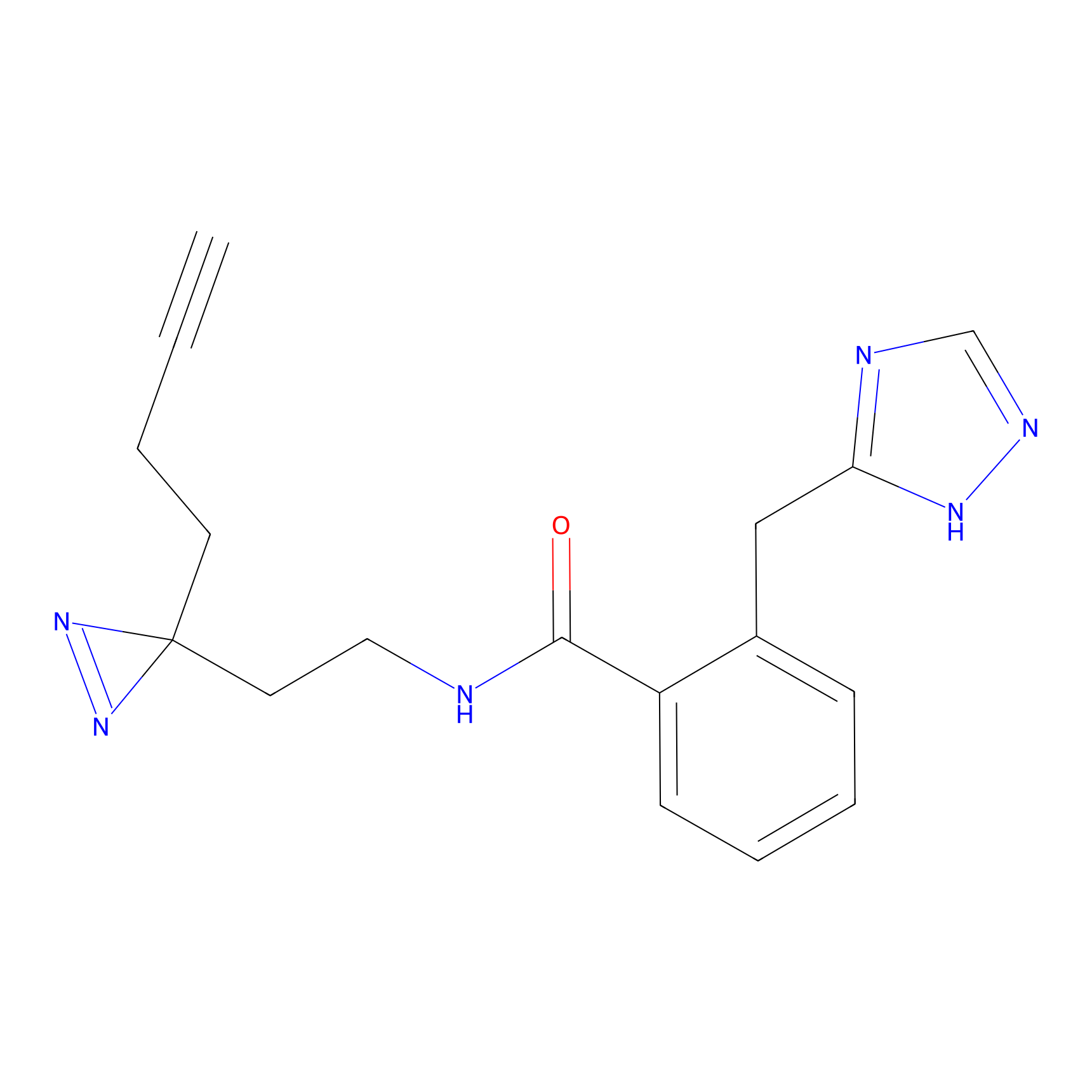
Acrolein Probe Info |
 |
N.A. | LDD0221 | [3] | |
|
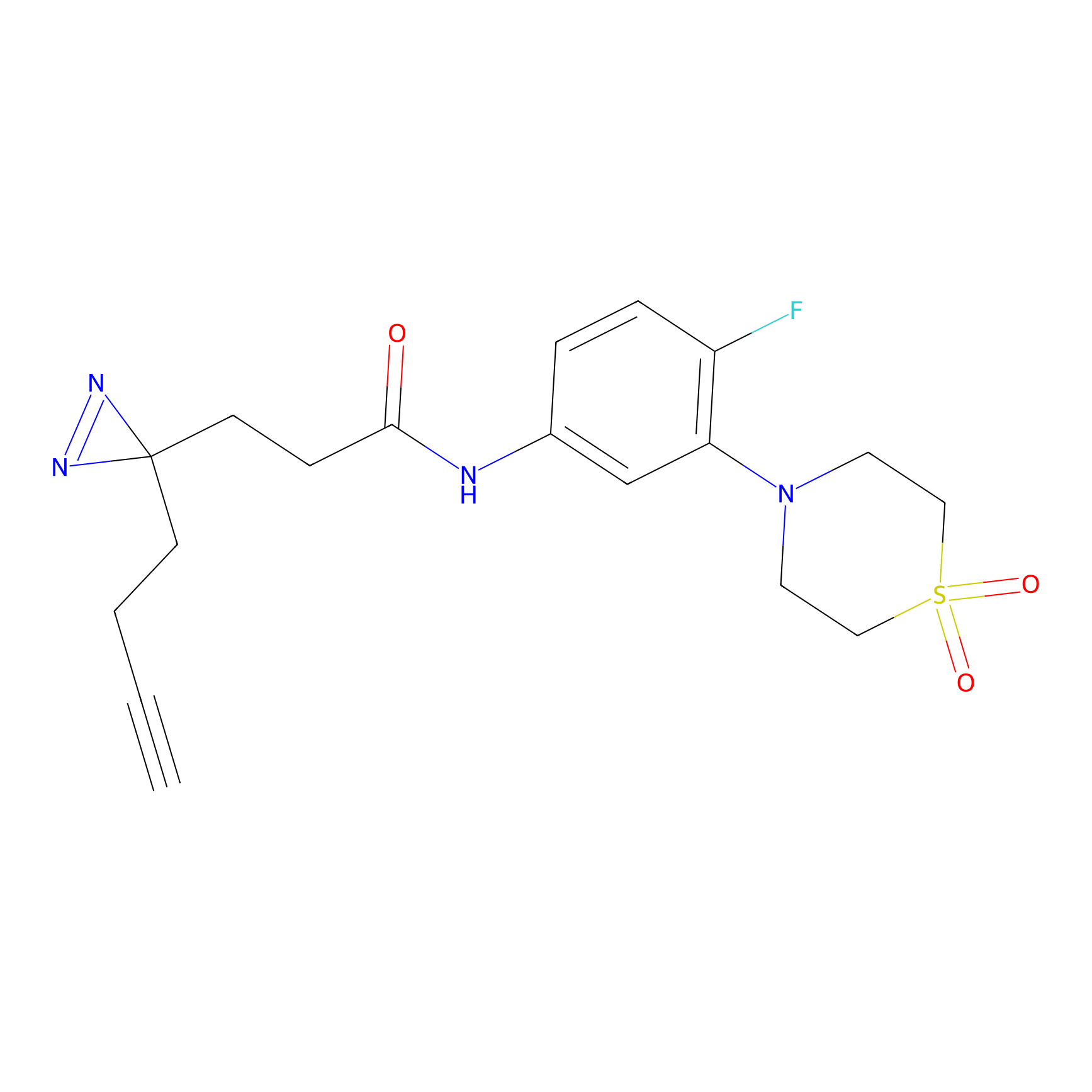
ATP probe Probe Info |
 |
N.A. | LDD0199 | [4] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [5] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References