Details of the Target
General Information of Target
| Target ID | LDTP05066 | |||||
|---|---|---|---|---|---|---|
| Target Name | Signal peptidase complex catalytic subunit SEC11A (SEC11A) | |||||
| Gene Name | SEC11A | |||||
| Gene ID | 23478 | |||||
| Synonyms |
SEC11L1; SPC18; SPCS4A; Signal peptidase complex catalytic subunit SEC11A; EC 3.4.21.89; Endopeptidase SP18; Microsomal signal peptidase 18 kDa subunit; SPase 18 kDa subunit; SEC11 homolog A; SEC11-like protein 1; SPC18
|
|||||
| 3D Structure | ||||||
| Sequence |
MLSLDFLDDVRRMNKRQLYYQVLNFGMIVSSALMIWKGLMVITGSESPIVVVLSGSMEPA
FHRGDLLFLTNRVEDPIRVGEIVVFRIEGREIPIVHRVLKIHEKQNGHIKFLTKGDNNAV DDRGLYKQGQHWLEKKDVVGRARGFVPYIGIVTILMNDYPKFKYAVLFLLGLFVLVHRE |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Peptidase S26B family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Catalytic component of the signal peptidase complex (SPC) which catalyzes the cleavage of N-terminal signal sequences from nascent proteins as they are translocated into the lumen of the endoplasmic reticulum. Specifically cleaves N-terminal signal peptides that contain a hydrophobic alpha-helix (h-region) shorter than 18-20 amino acids.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
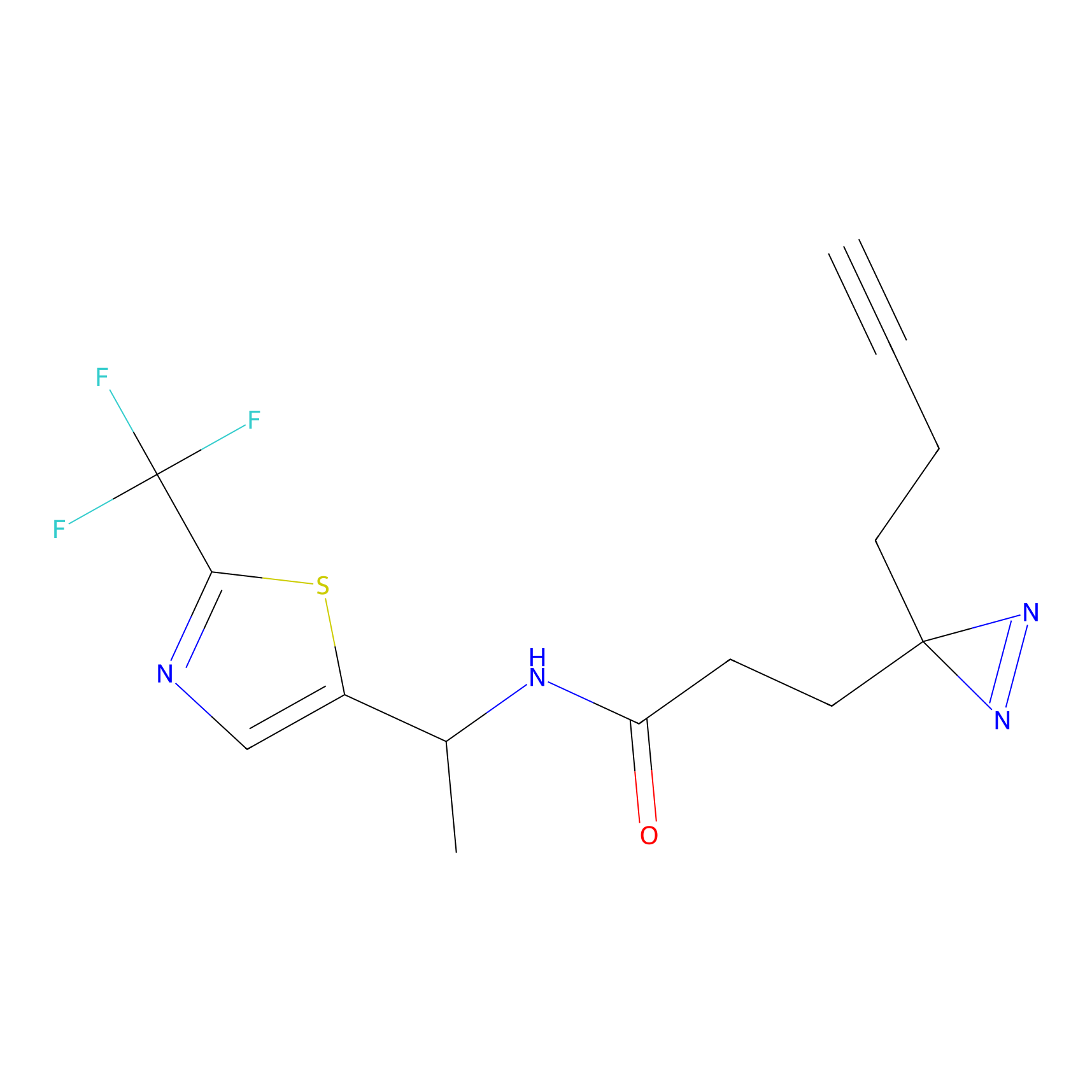
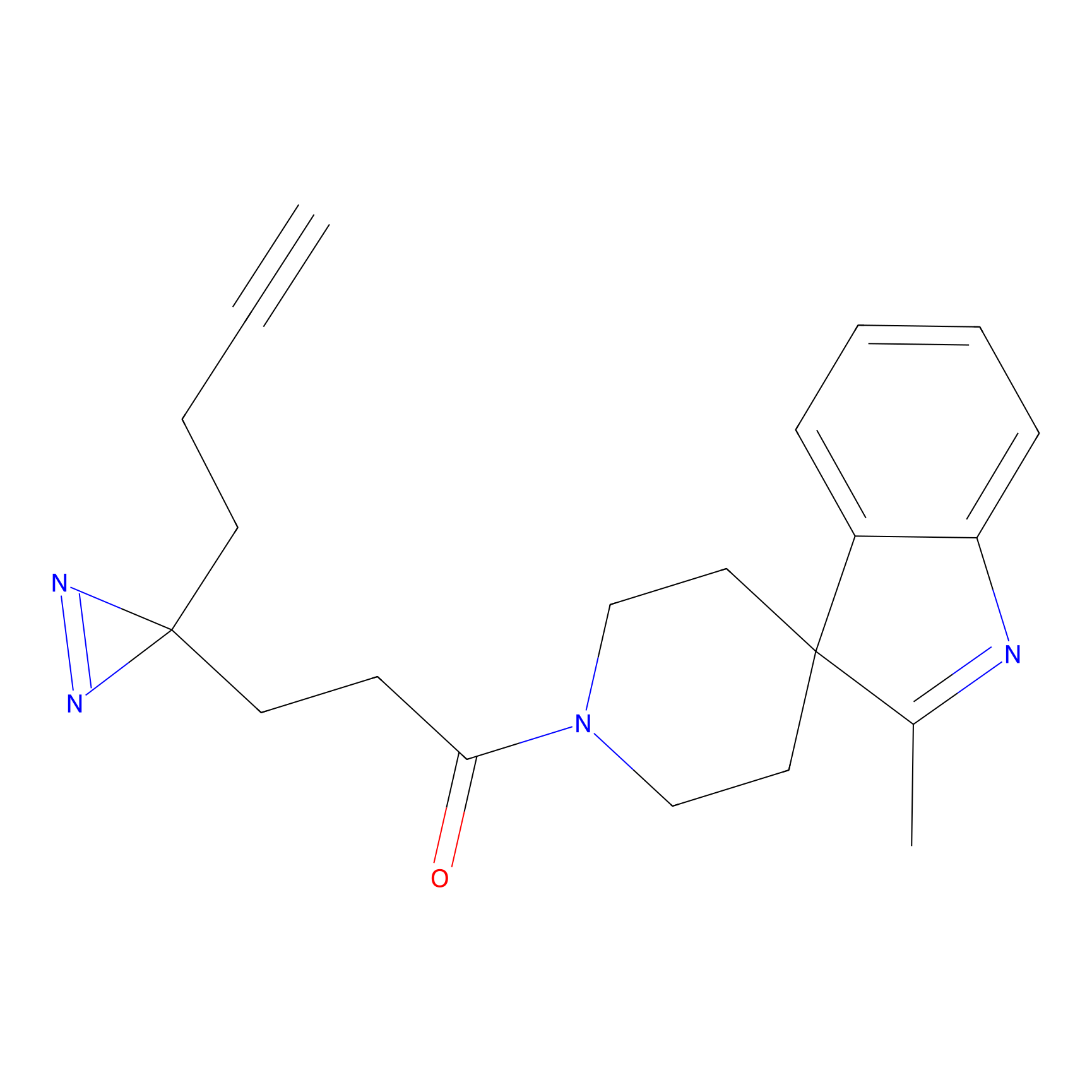
|
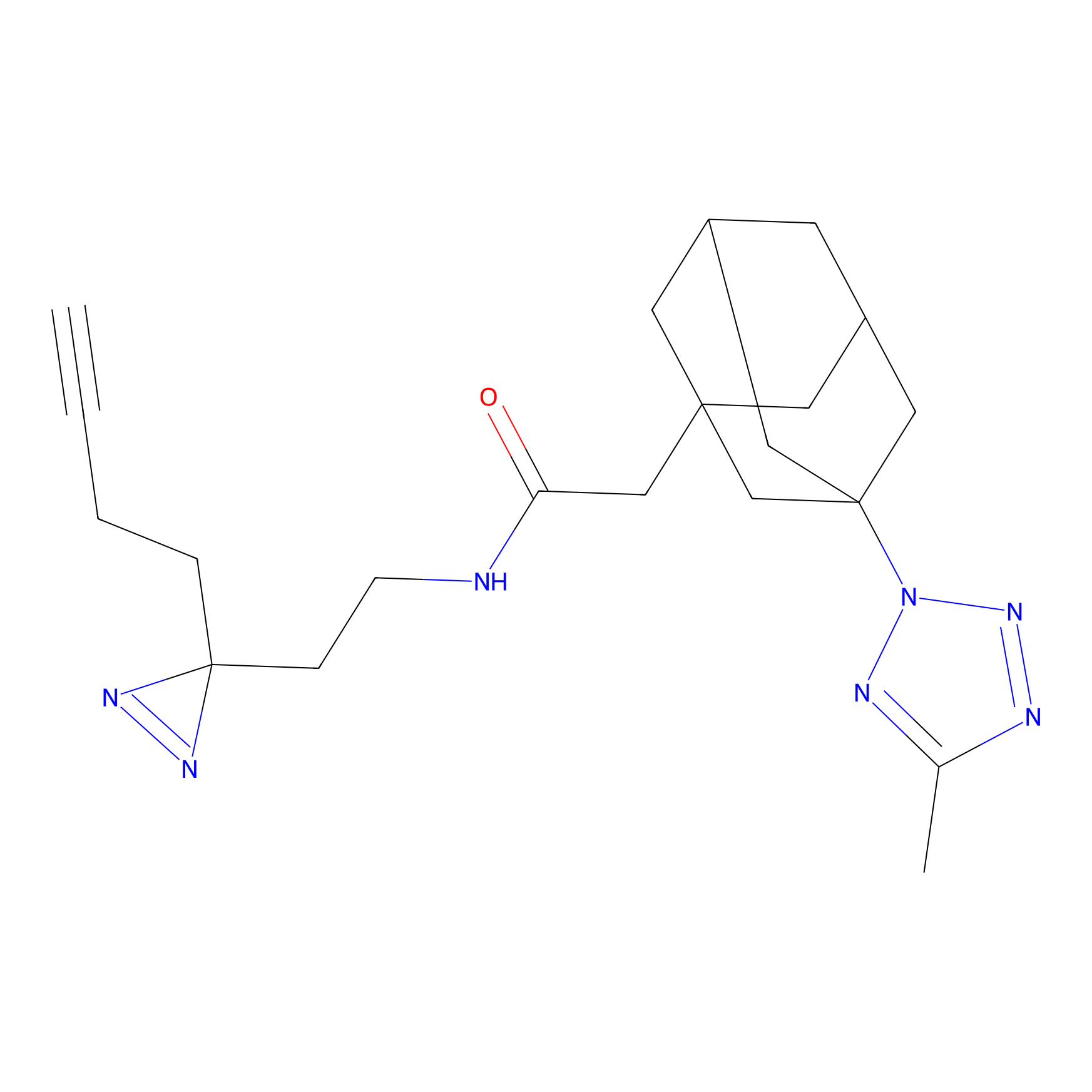
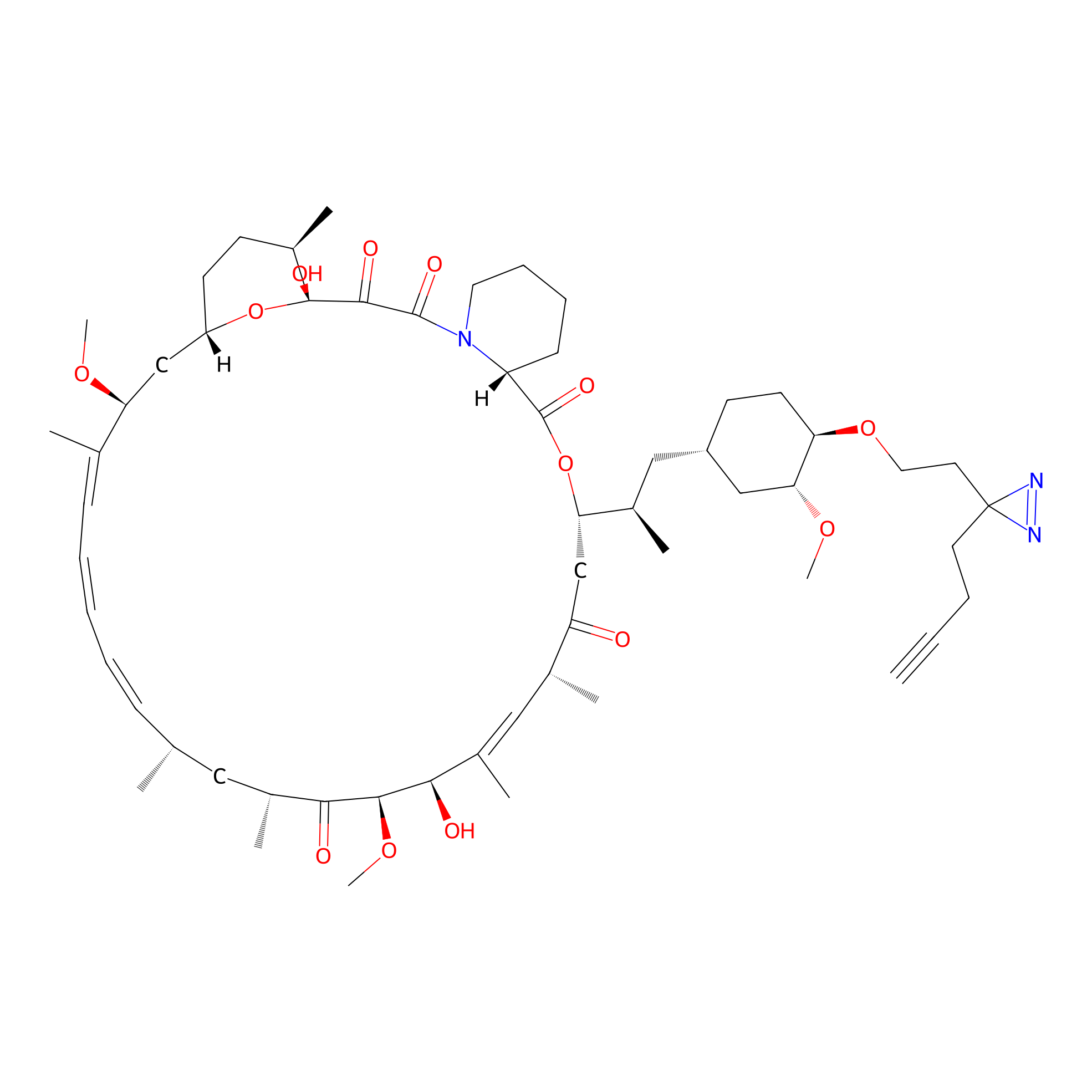
CHEMBL5175495 Probe Info |
 |
14.60 | LDD0196 | [1] | |
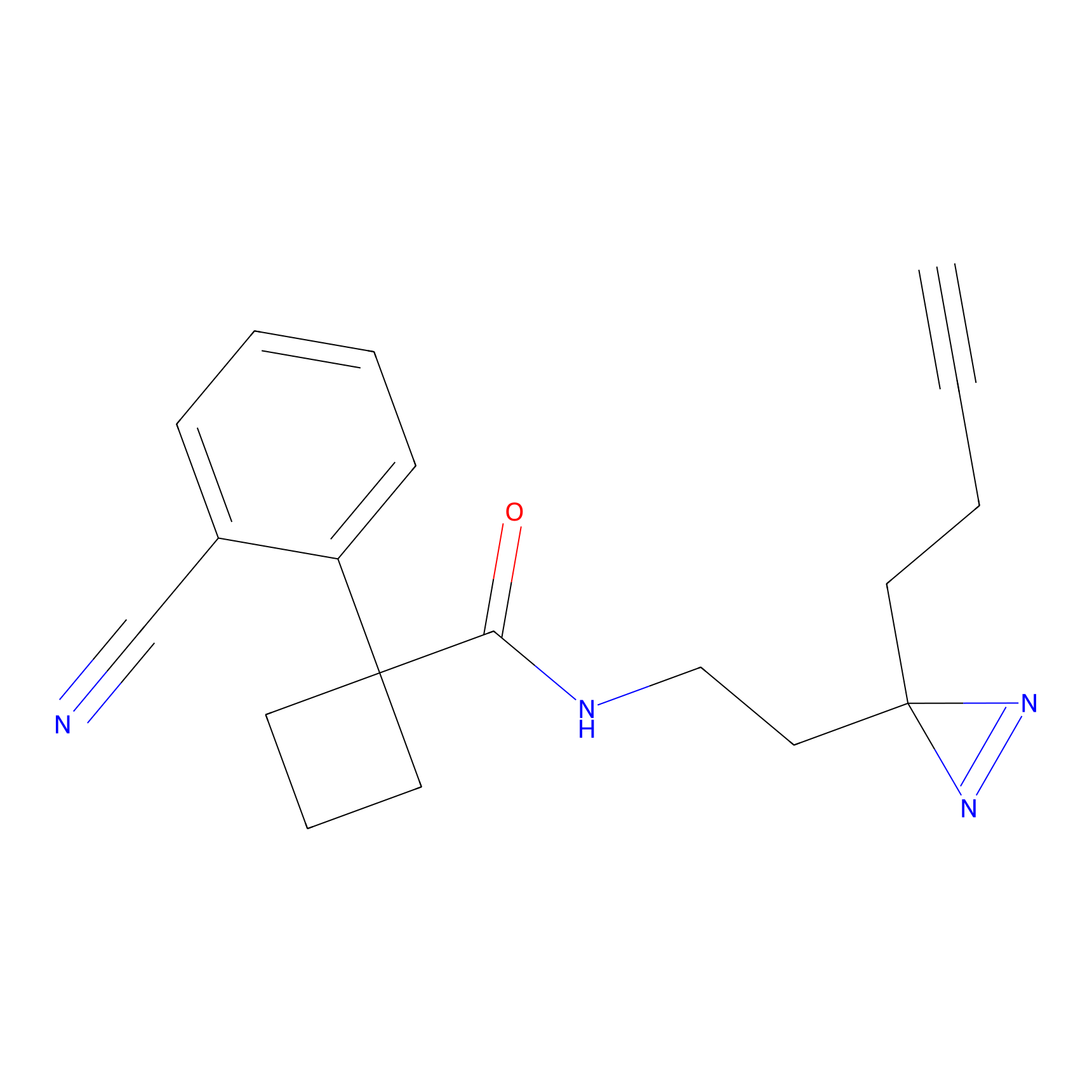
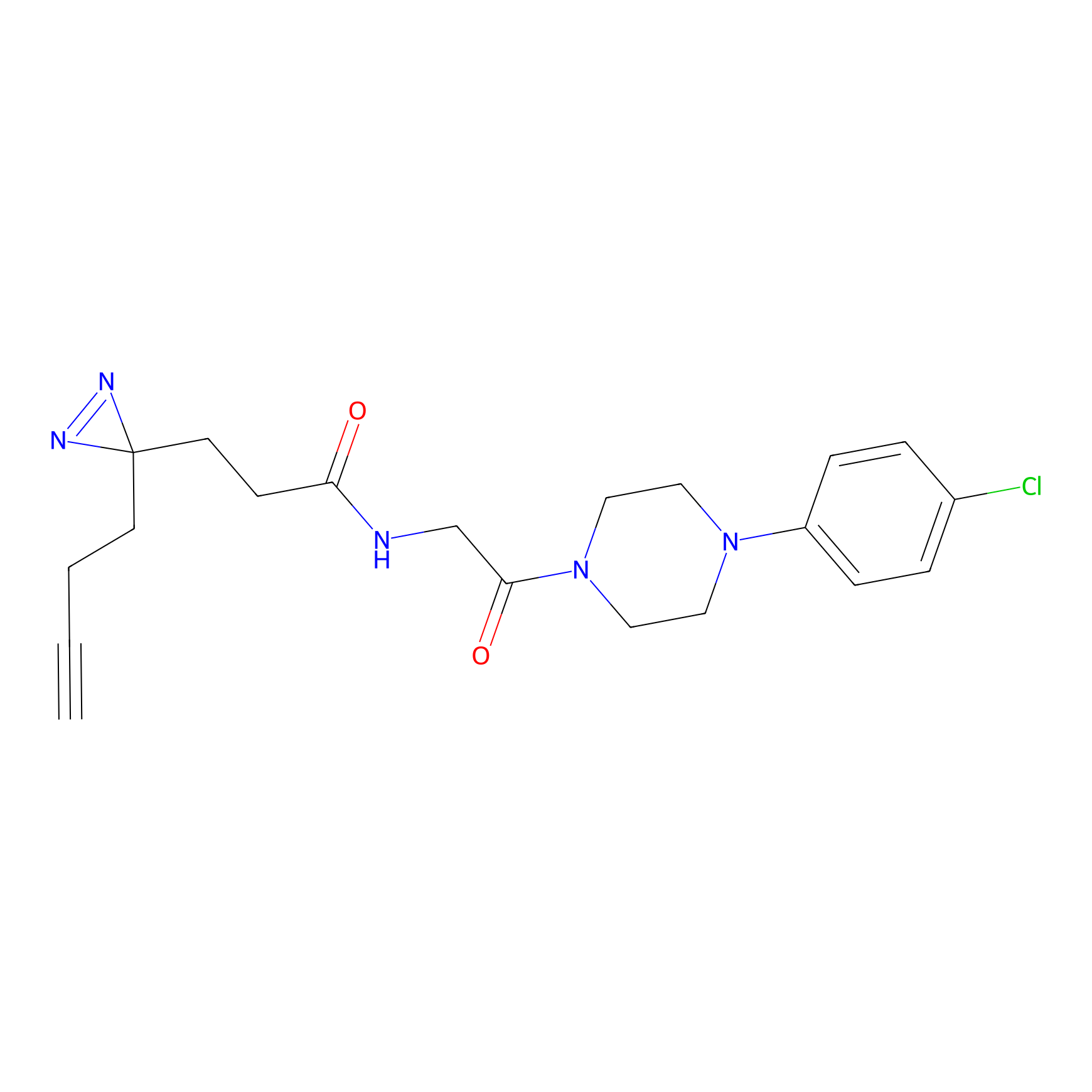
|
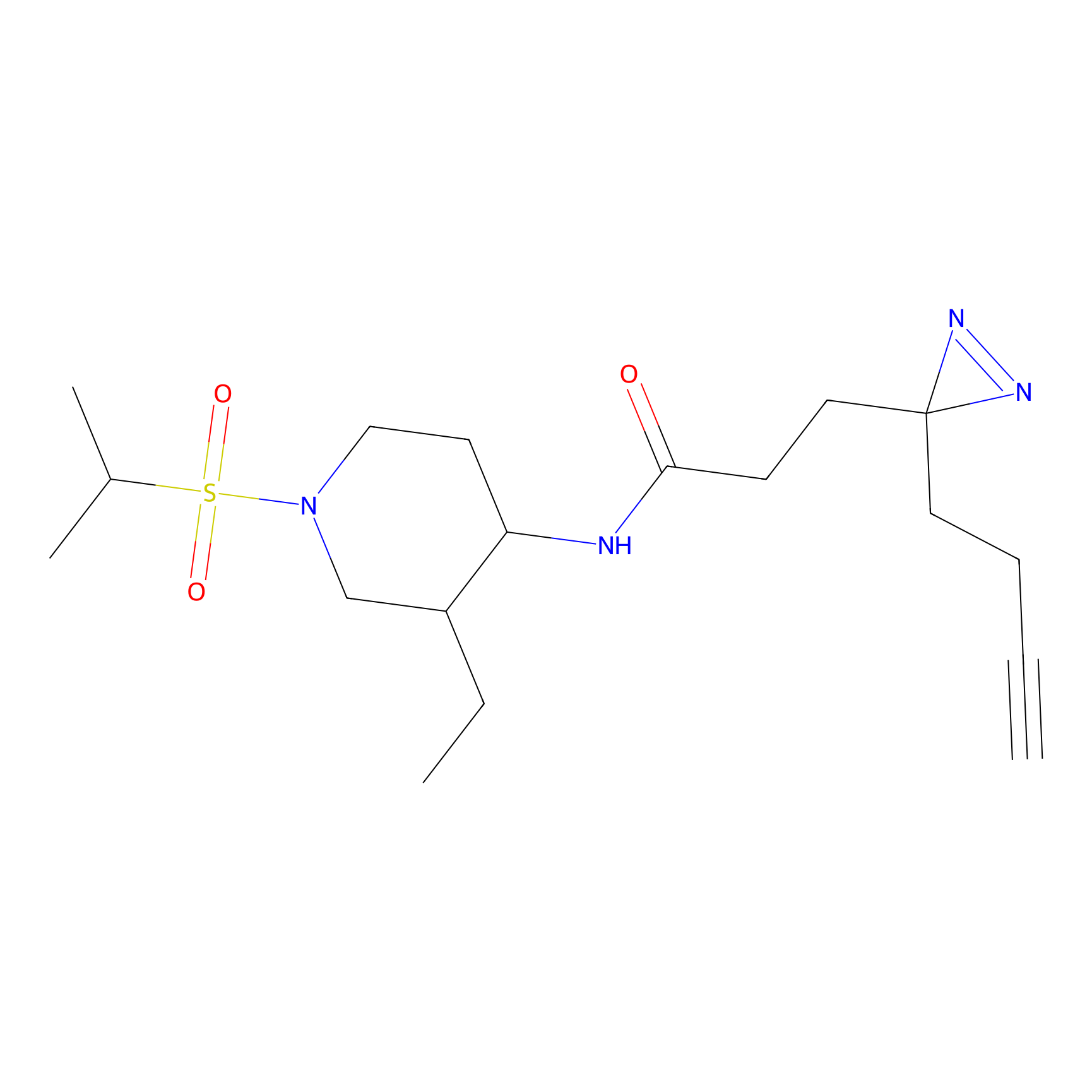
C-Sul Probe Info |
 |
2.89 | LDD0066 | [2] | |
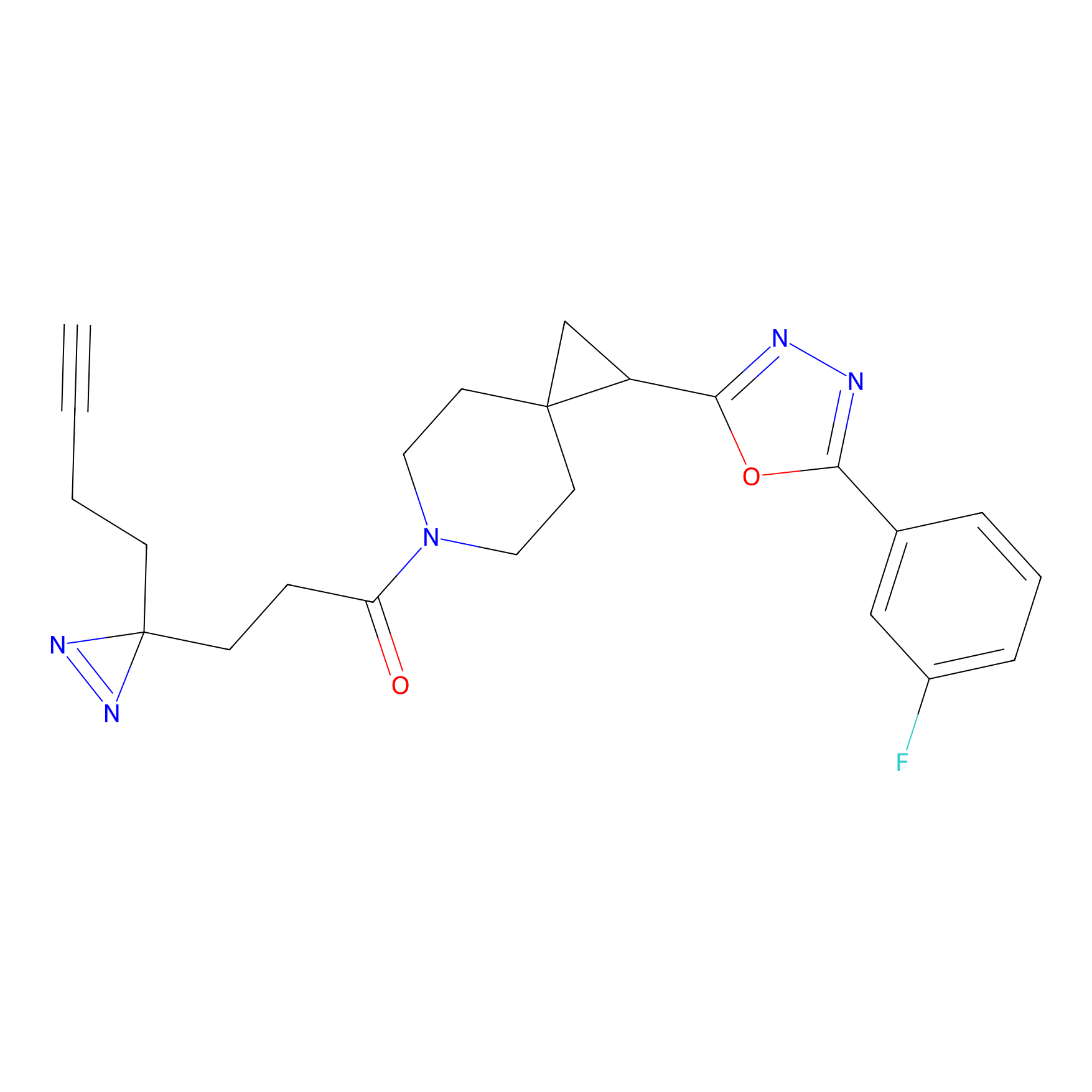
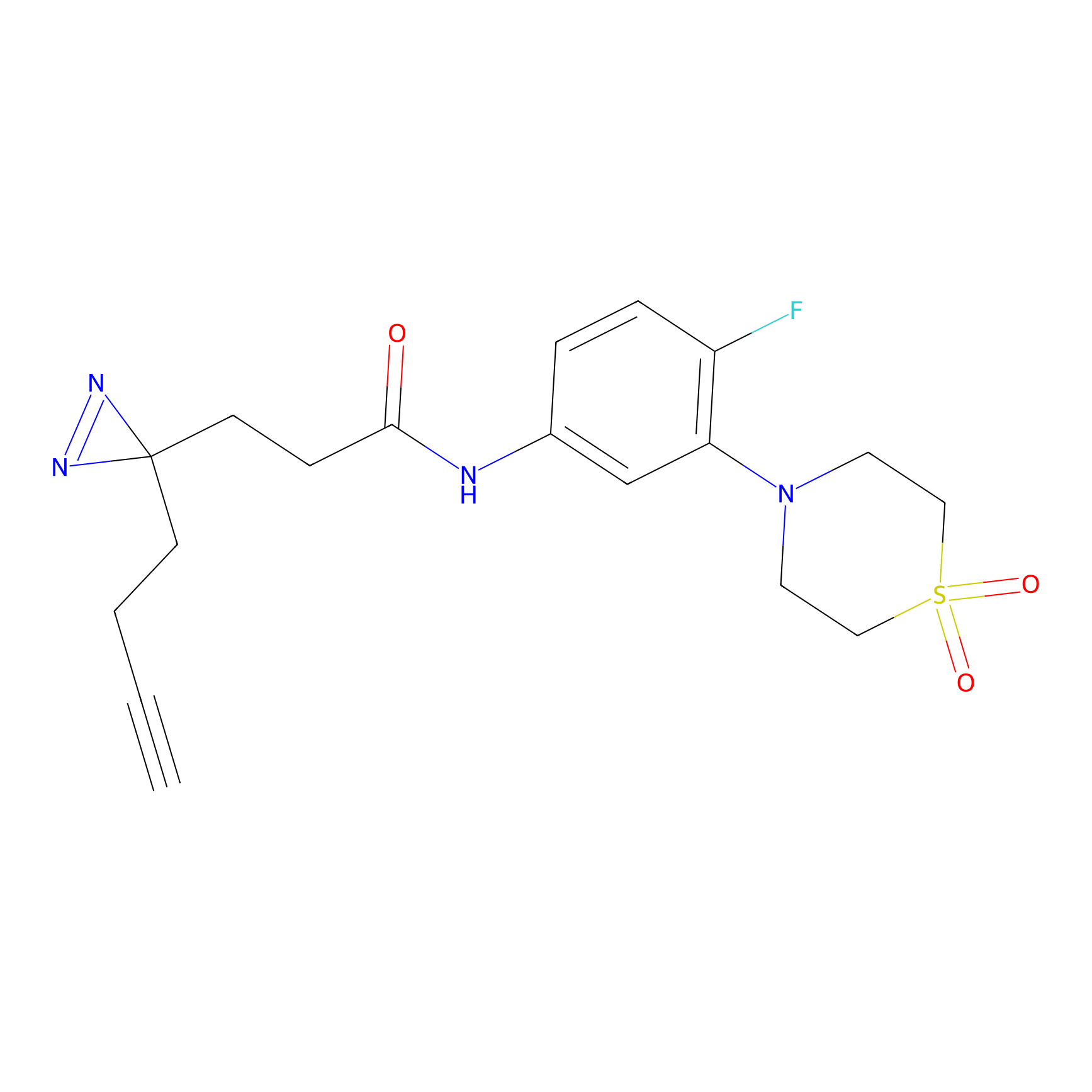
|
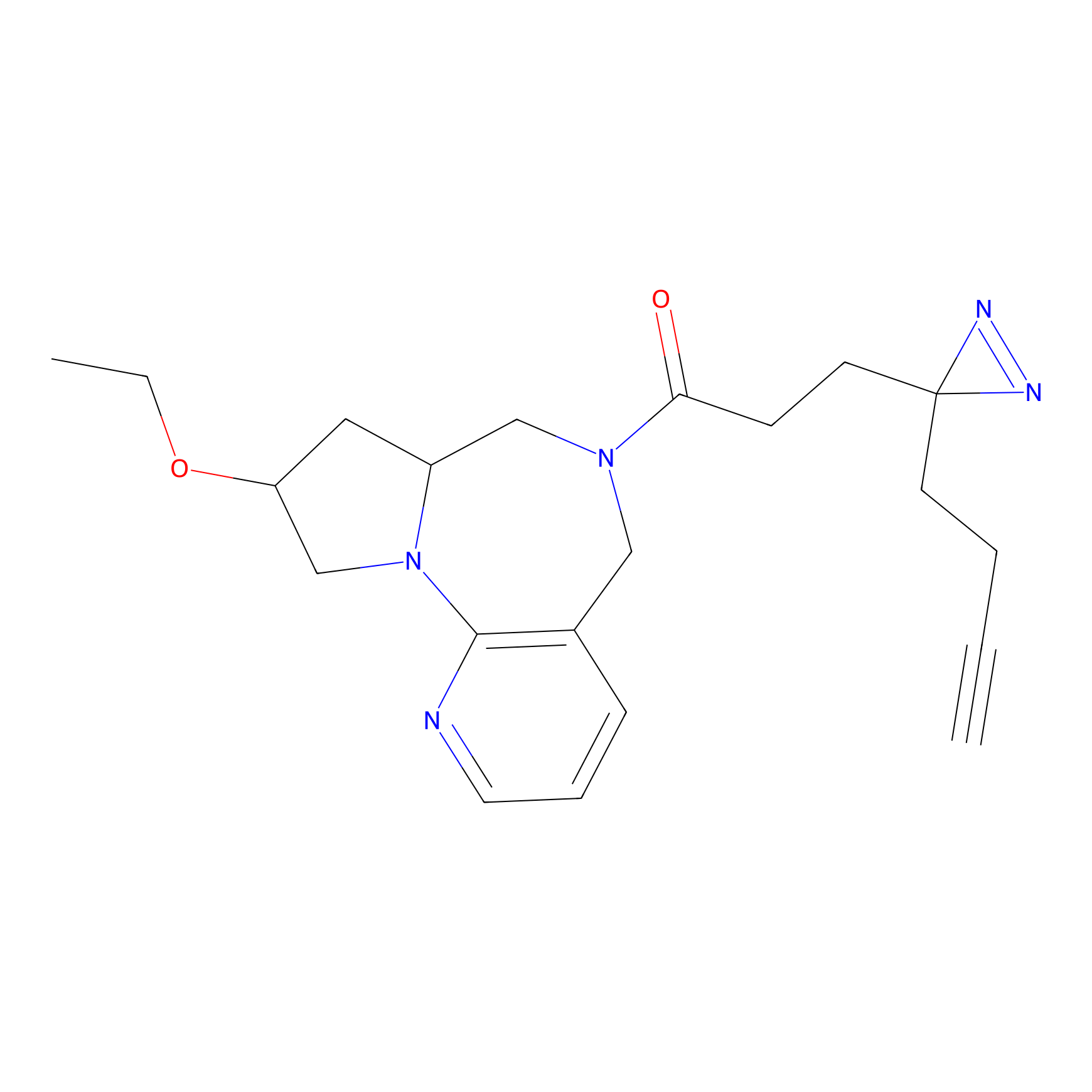
ONAyne Probe Info |
 |
K100(1.37); K136(0.92) | LDD0274 | [3] | |
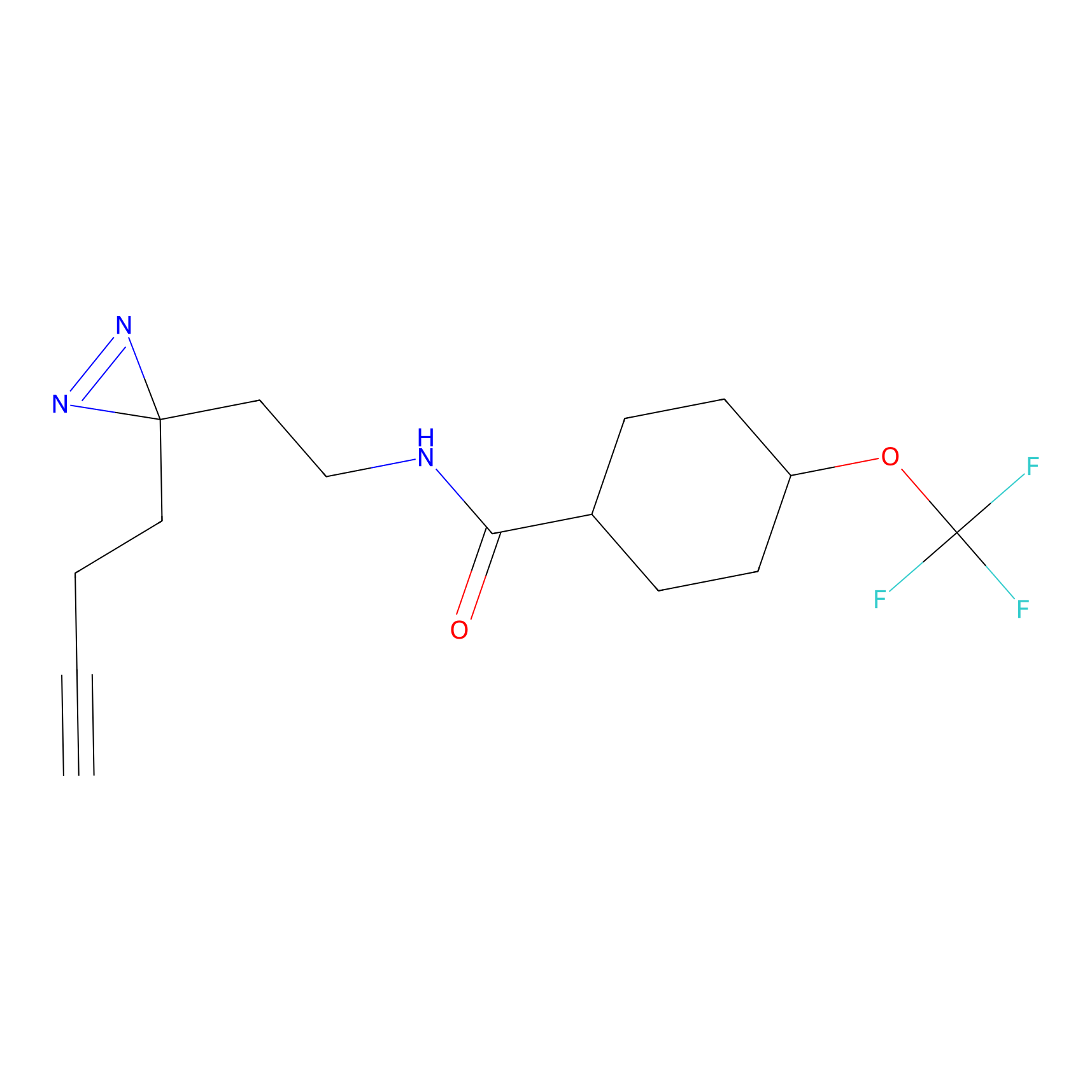
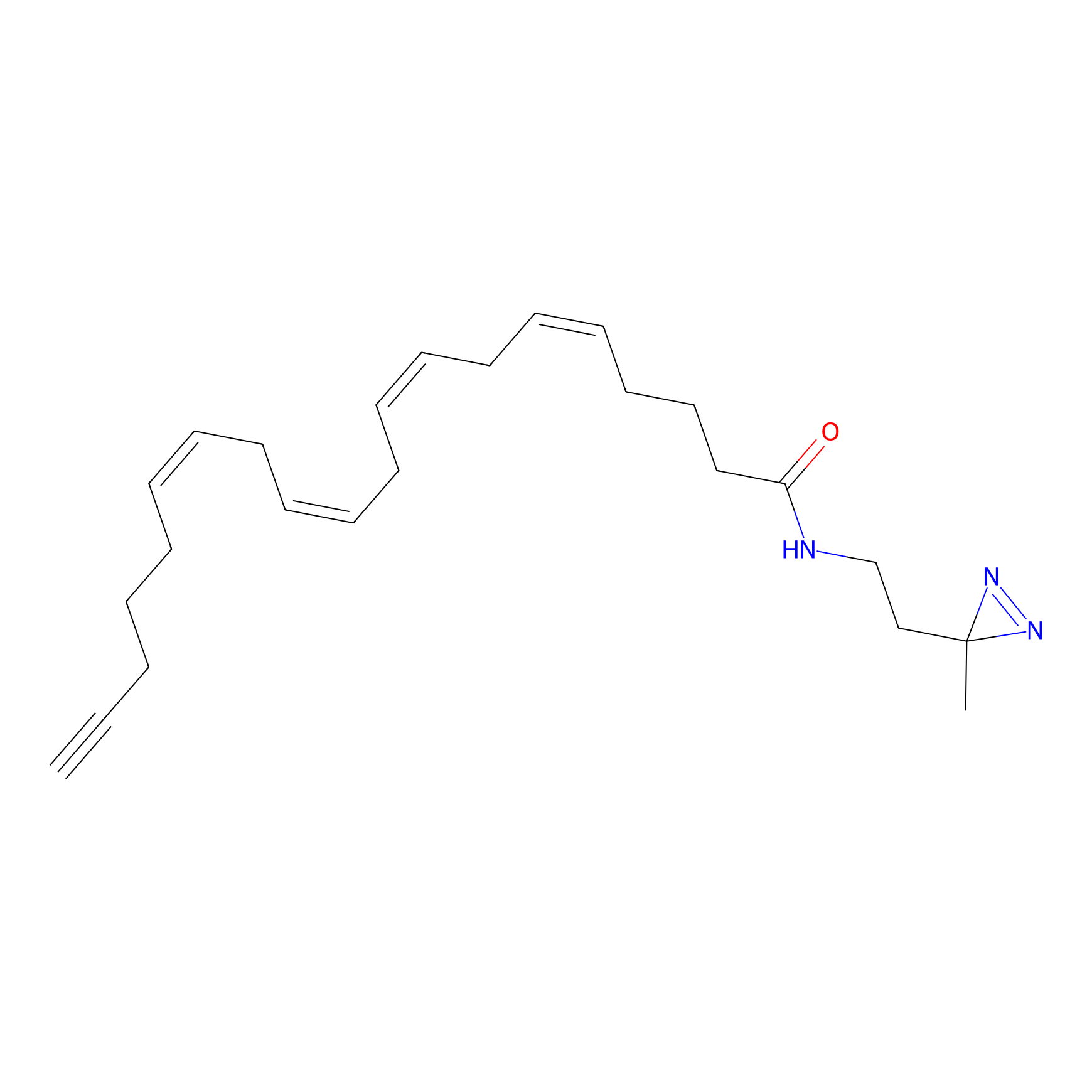
|
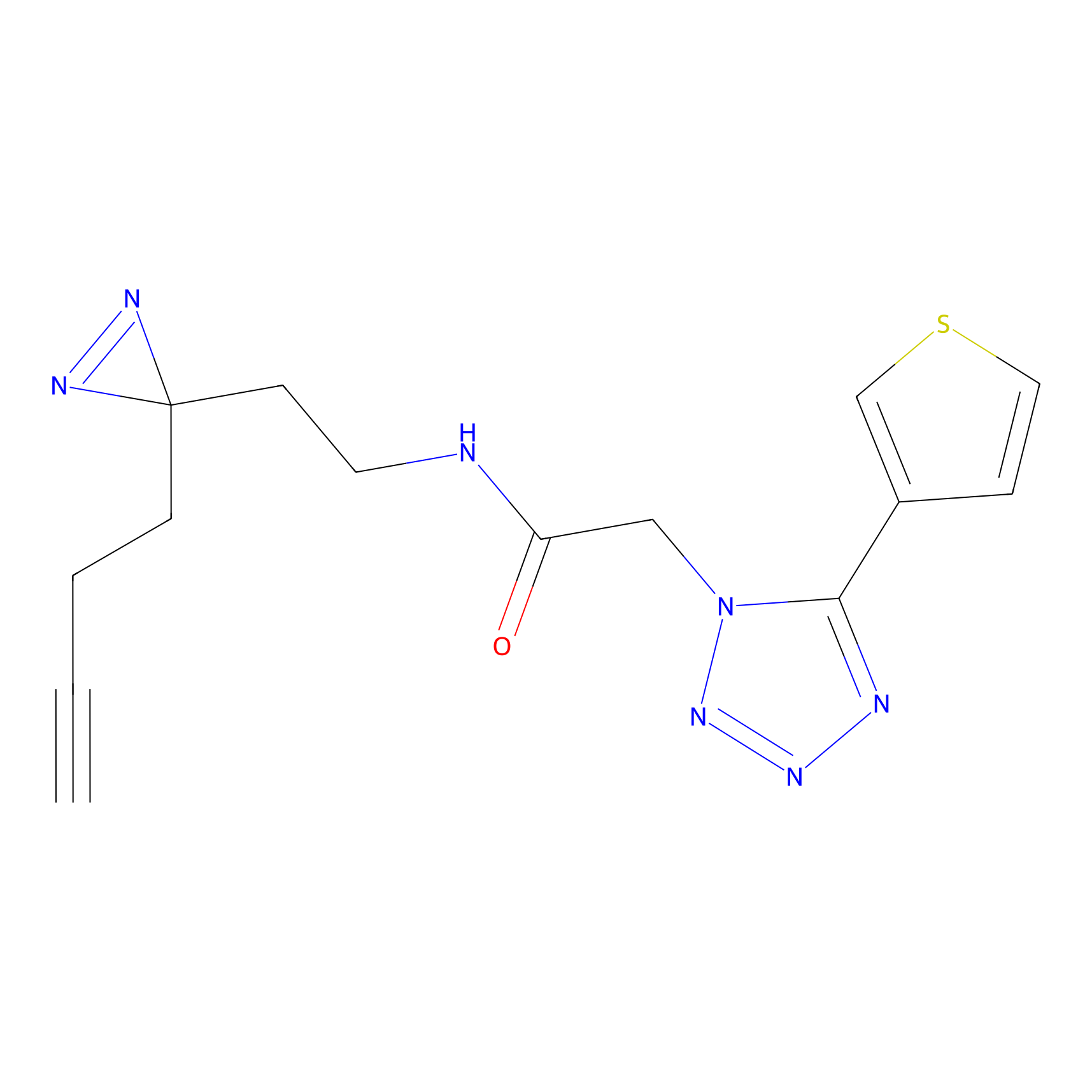
STPyne Probe Info |
 |
K100(7.34); K104(9.37); K110(4.22); K114(10.00) | LDD0277 | [3] | |
|
Jackson_14 Probe Info |
 |
4.00 | LDD0123 | [4] | |
|
HPAP Probe Info |
 |
4.30 | LDD0062 | [5] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0233 | [6] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References