Details of the Target
General Information of Target
| Target ID | LDTP05905 | |||||
|---|---|---|---|---|---|---|
| Target Name | Acid ceramidase (ASAH1) | |||||
| Gene Name | ASAH1 | |||||
| Gene ID | 427 | |||||
| Synonyms |
ASAH; Acid ceramidase; AC; ACDase; Acid CDase; EC 3.5.1.23; Acylsphingosine deacylase; N-acylethanolamine hydrolase ASAH1; EC 3.5.1.-; N-acylsphingosine amidohydrolase; Putative 32 kDa heart protein; PHP32) [Cleaved into: Acid ceramidase subunit alpha; Acid ceramidase subunit beta]
|
|||||
| 3D Structure | ||||||
| Sequence |
MPGRSCVALVLLAAAVSCAVAQHAPPWTEDCRKSTYPPSGPTYRGAVPWYTINLDLPPYK
RWHELMLDKAPVLKVIVNSLKNMINTFVPSGKIMQVVDEKLPGLLGNFPGPFEEEMKGIA AVTDIPLGEIISFNIFYELFTICTSIVAEDKKGHLIHGRNMDFGVFLGWNINNDTWVITE QLKPLTVNLDFQRNNKTVFKASSFAGYVGMLTGFKPGLFSLTLNERFSINGGYLGILEWI LGKKDVMWIGFLTRTVLENSTSYEEAKNLLTKTKILAPAYFILGGNQSGEGCVITRDRKE SLDVYELDAKQGRWYVVQTNYDRWKHPFFLDDRRTPAKMCLNRTSQENISFETMYDVLST KPVLNKLTVYTTLIDVTKGQFETYLRDCPDPCIGW |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Acid ceramidase family
|
|||||
| Subcellular location |
Lysosome; Nucleus
|
|||||
| Function |
Lysosomal ceramidase that hydrolyzes sphingolipid ceramides into sphingosine and free fatty acids at acidic pH. Ceramides, sphingosine, and its phosphorylated form sphingosine-1-phosphate are bioactive lipids that mediate cellular signaling pathways regulating several biological processes including cell proliferation, apoptosis and differentiation. Has a higher catalytic efficiency towards C12-ceramides versus other ceramides. Also catalyzes the reverse reaction allowing the synthesis of ceramides from fatty acids and sphingosine. For the reverse synthetic reaction, the natural sphingosine D-erythro isomer is more efficiently utilized as a substrate compared to D-erythro-dihydrosphingosine and D-erythro-phytosphingosine, while the fatty acids with chain lengths of 12 or 14 carbons are the most efficiently used. Has also an N-acylethanolamine hydrolase activity. By regulating the levels of ceramides, sphingosine and sphingosine-1-phosphate in the epidermis, mediates the calcium-induced differentiation of epidermal keratinocytes. Also indirectly regulates tumor necrosis factor/TNF-induced apoptosis. By regulating the intracellular balance between ceramides and sphingosine, in adrenocortical cells, probably also acts as a regulator of steroidogenesis.; [Isoform 2]: May directly regulate steroidogenesis by binding the nuclear receptor NR5A1 and negatively regulating its transcriptional activity.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
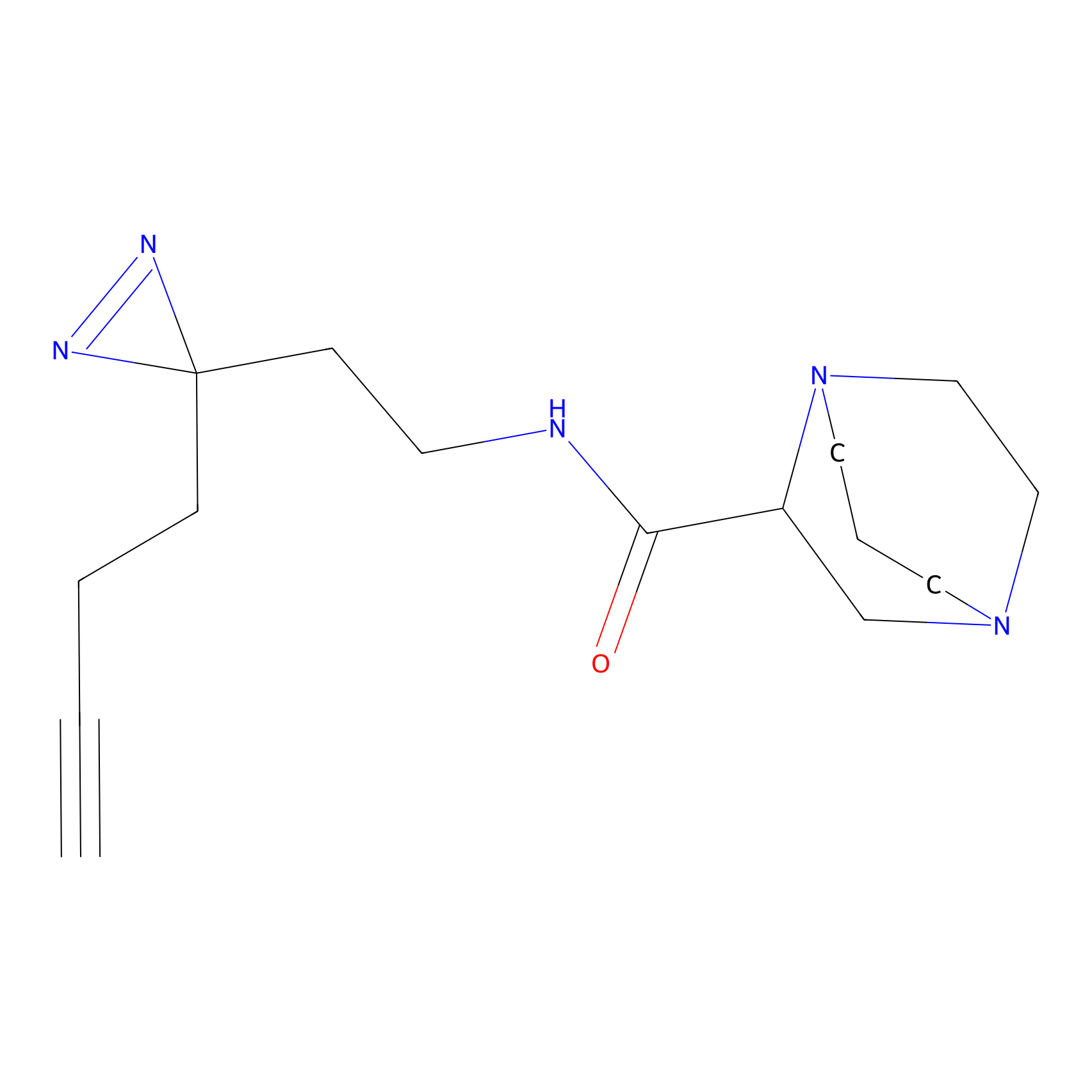
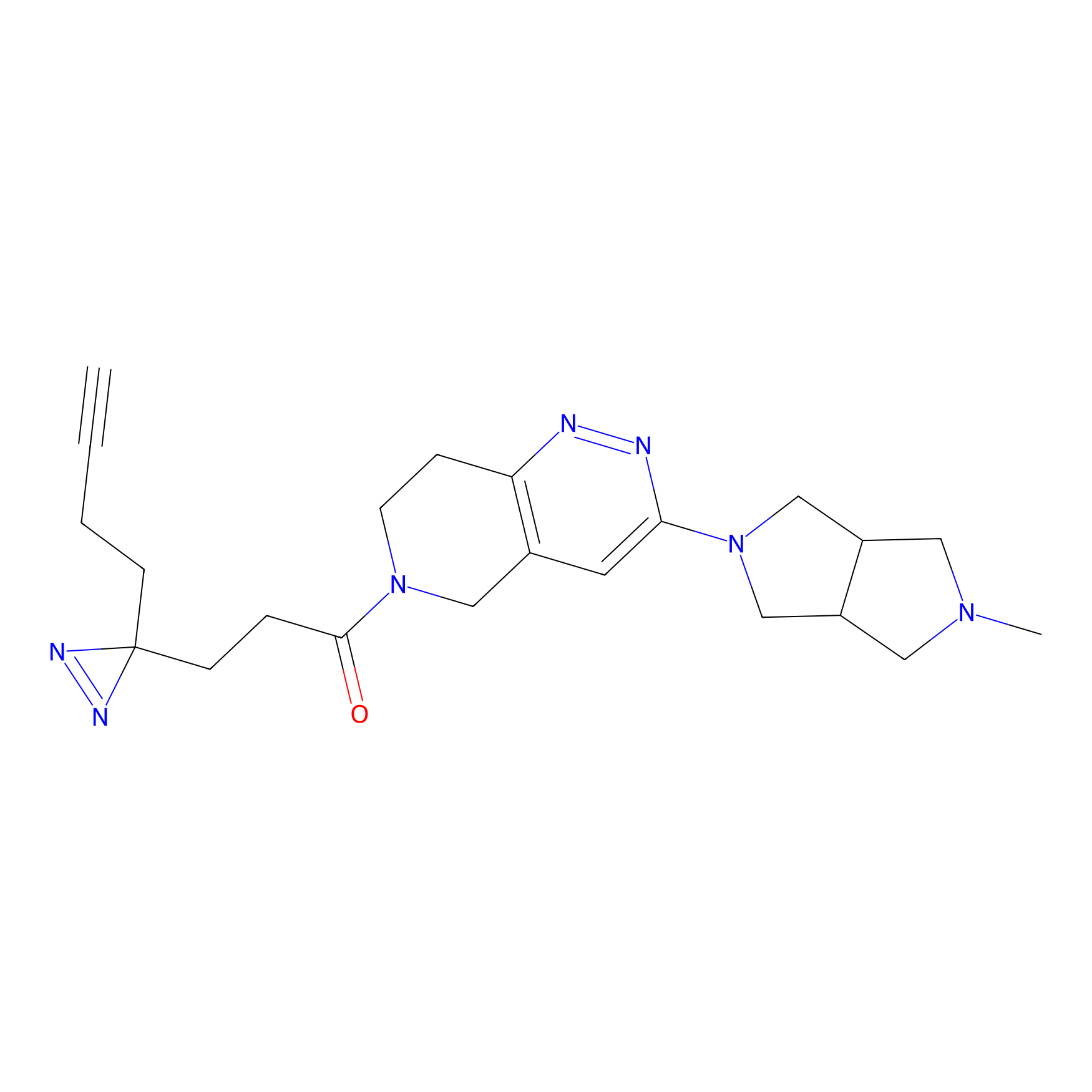
Alkylaryl probe 1 Probe Info |
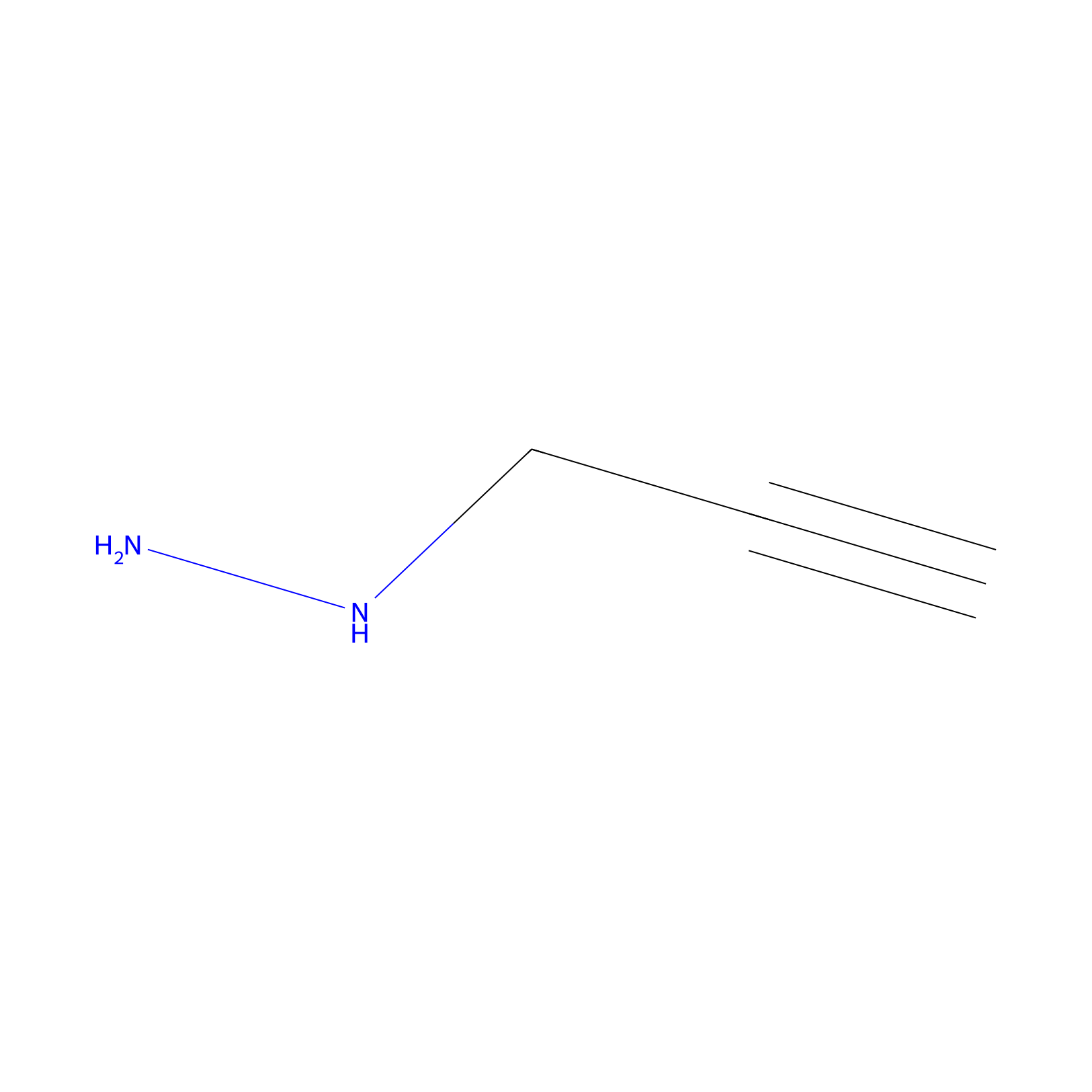 |
20.00 | LDD0387 | [1] | |
|
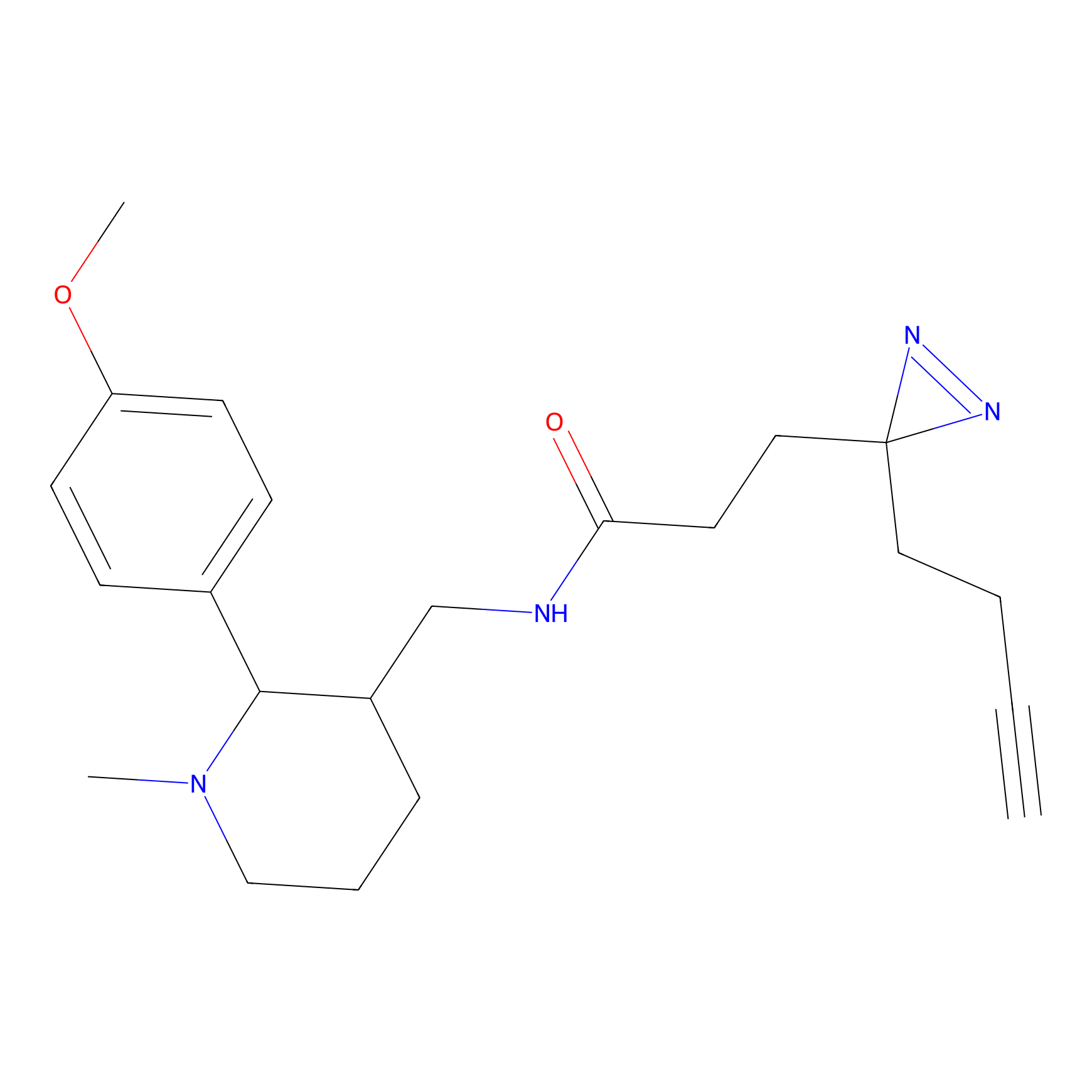
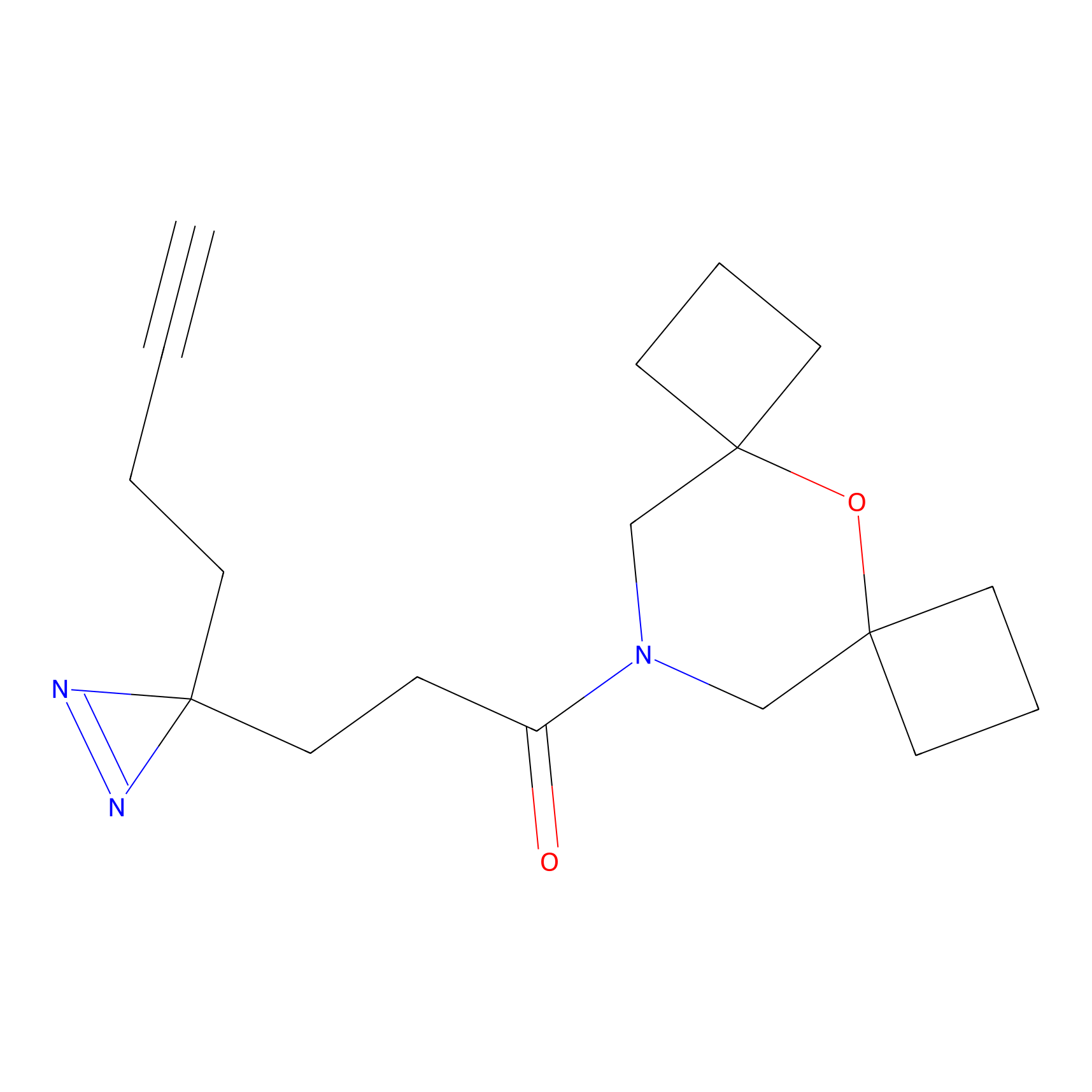
Alkylaryl probe 2 Probe Info |
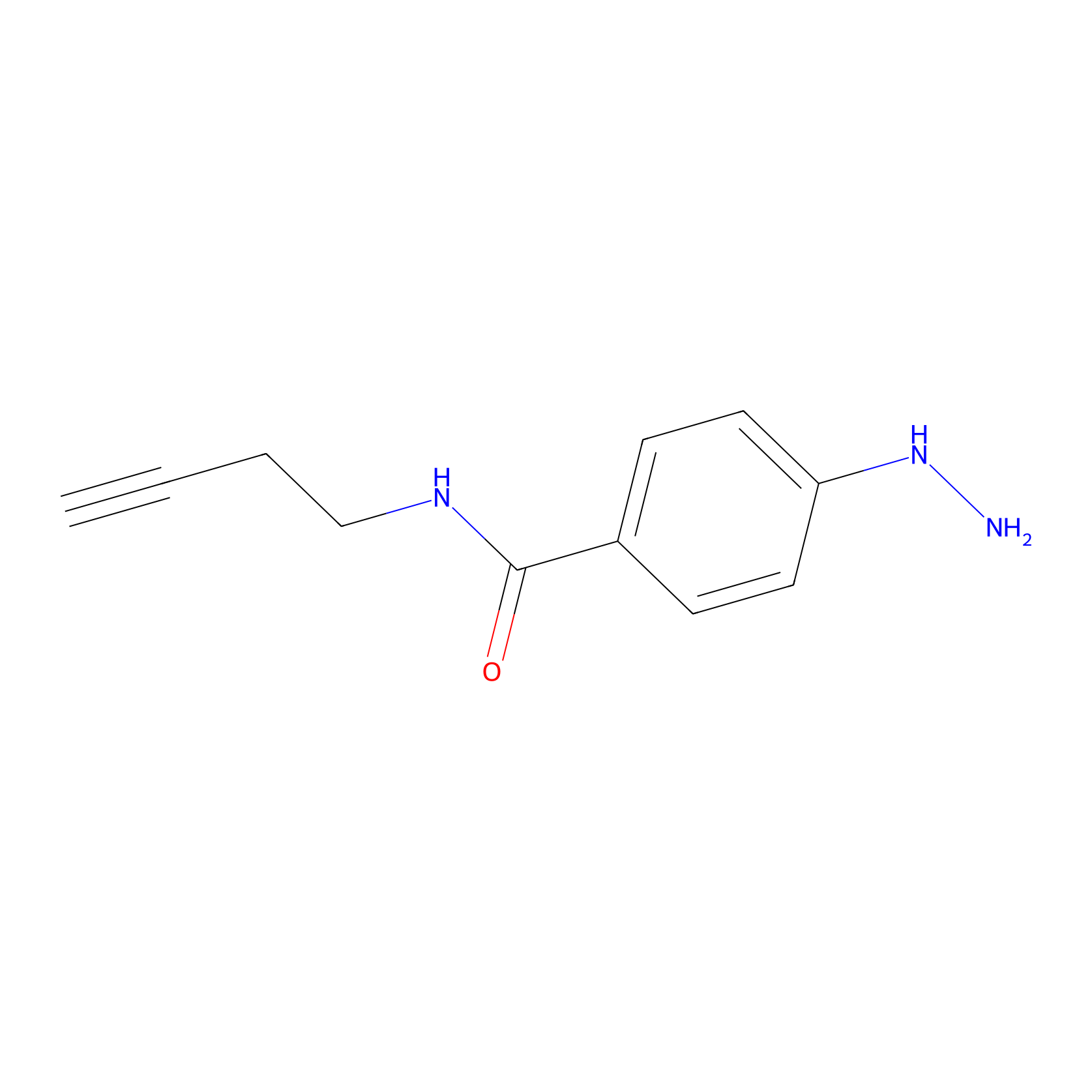 |
20.00 | LDD0391 | [1] | |
|
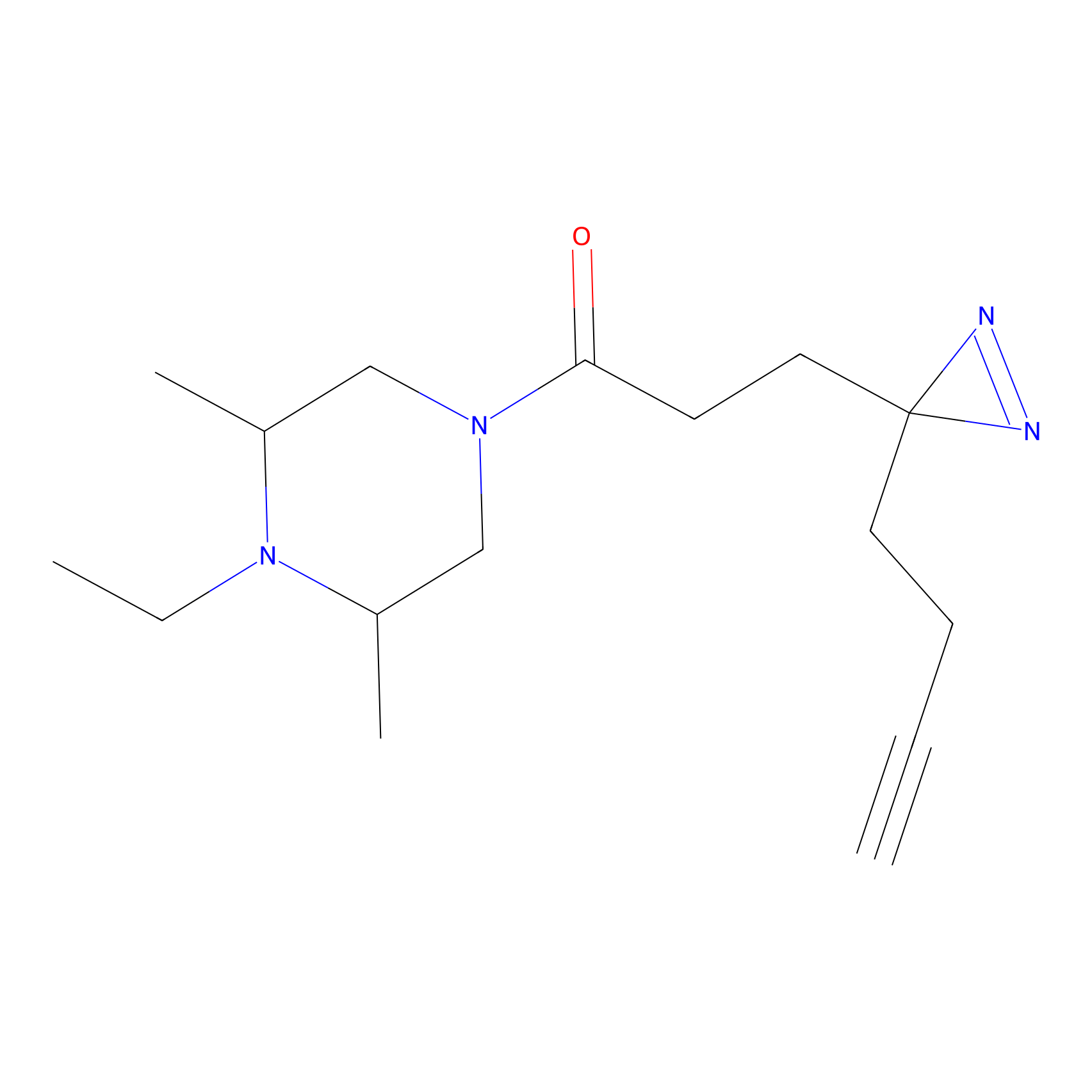
FBPP2 Probe Info |
 |
152.19 | LDD0318 | [2] | |
|
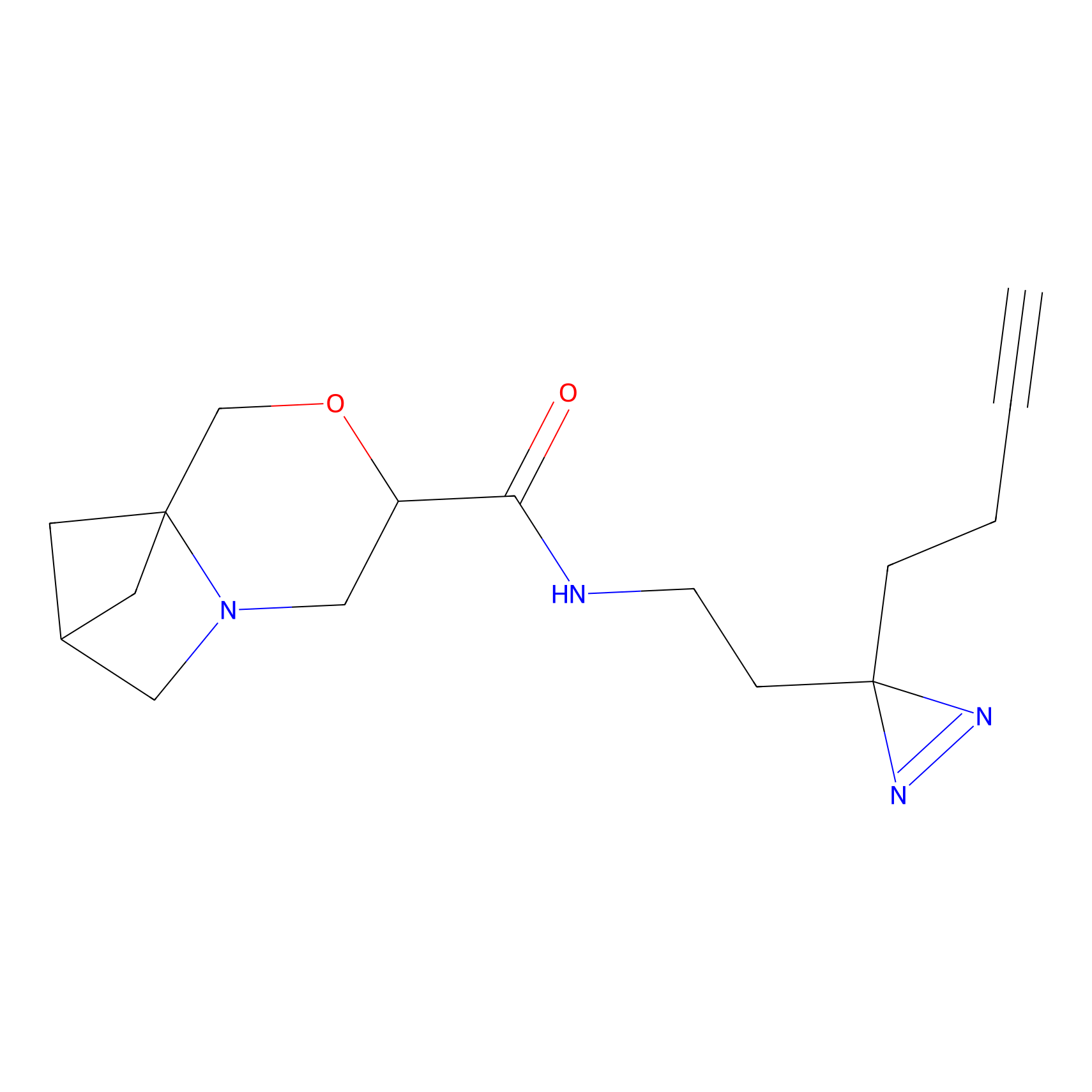
FBP2 Probe Info |
 |
2.63 | LDD0317 | [2] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [3] | |
|
STPyne Probe Info |
 |
K152(5.56); K325(9.07) | LDD0277 | [4] | |
|
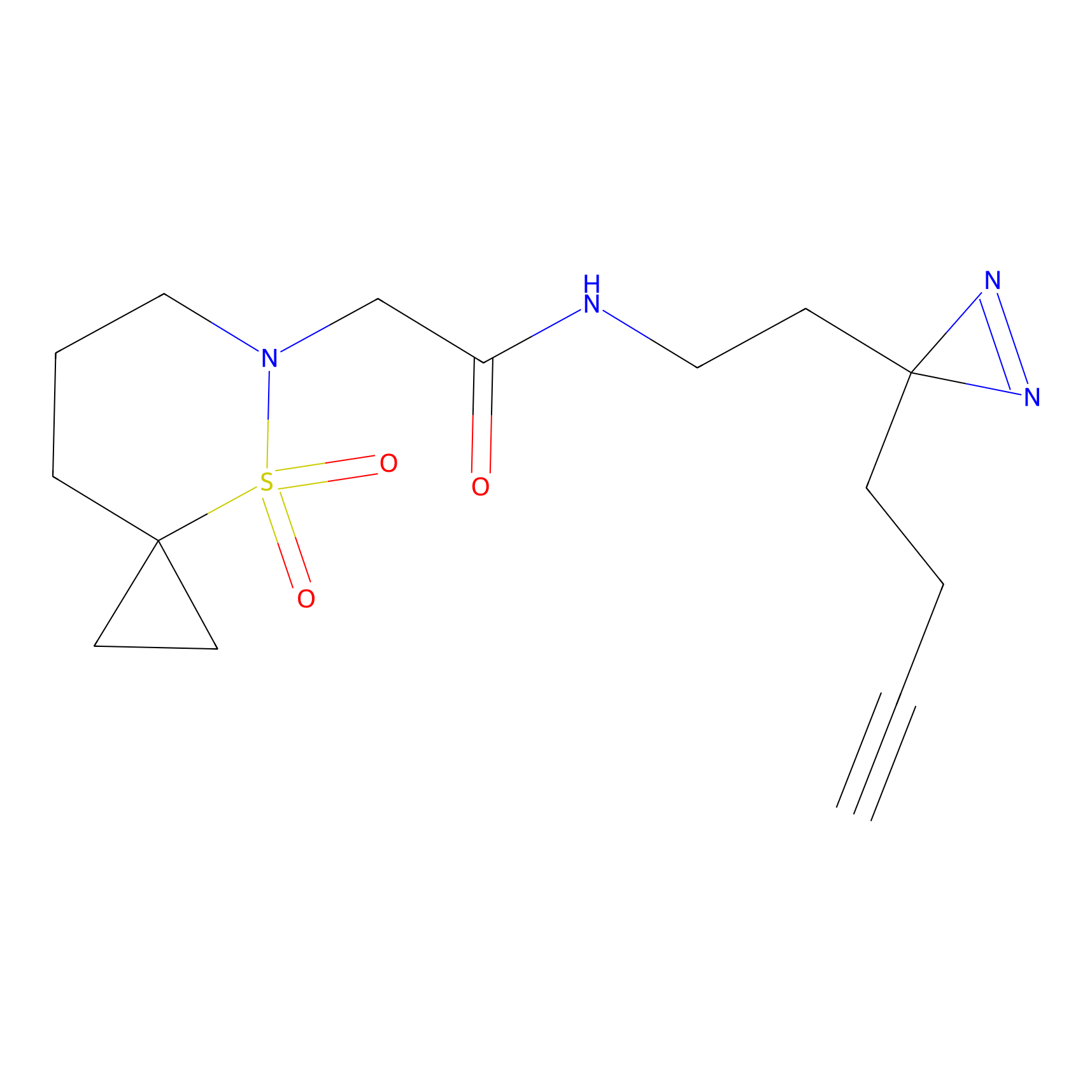
Alkylaryl probe 3 Probe Info |
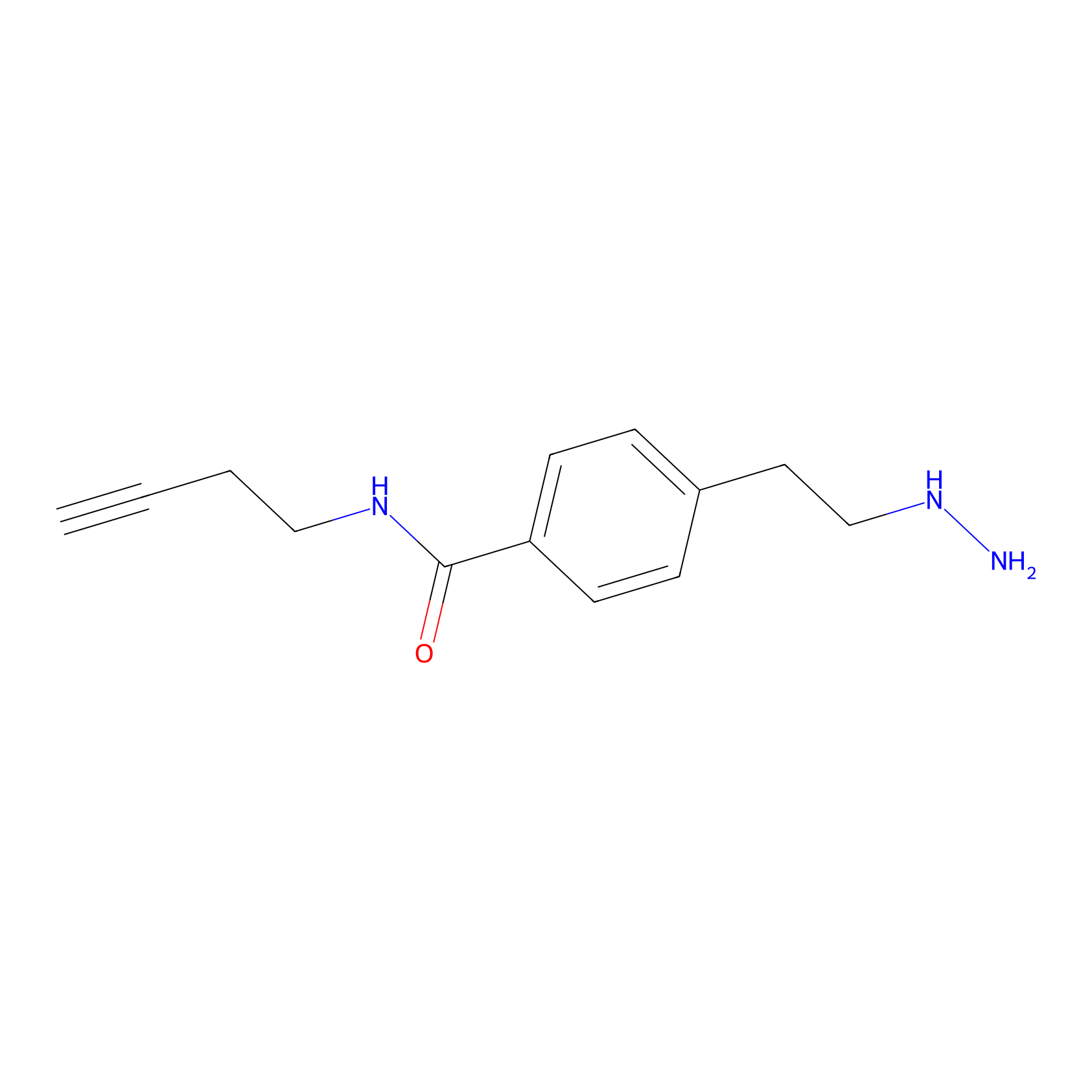 |
3.82 | LDD0384 | [1] | |
|
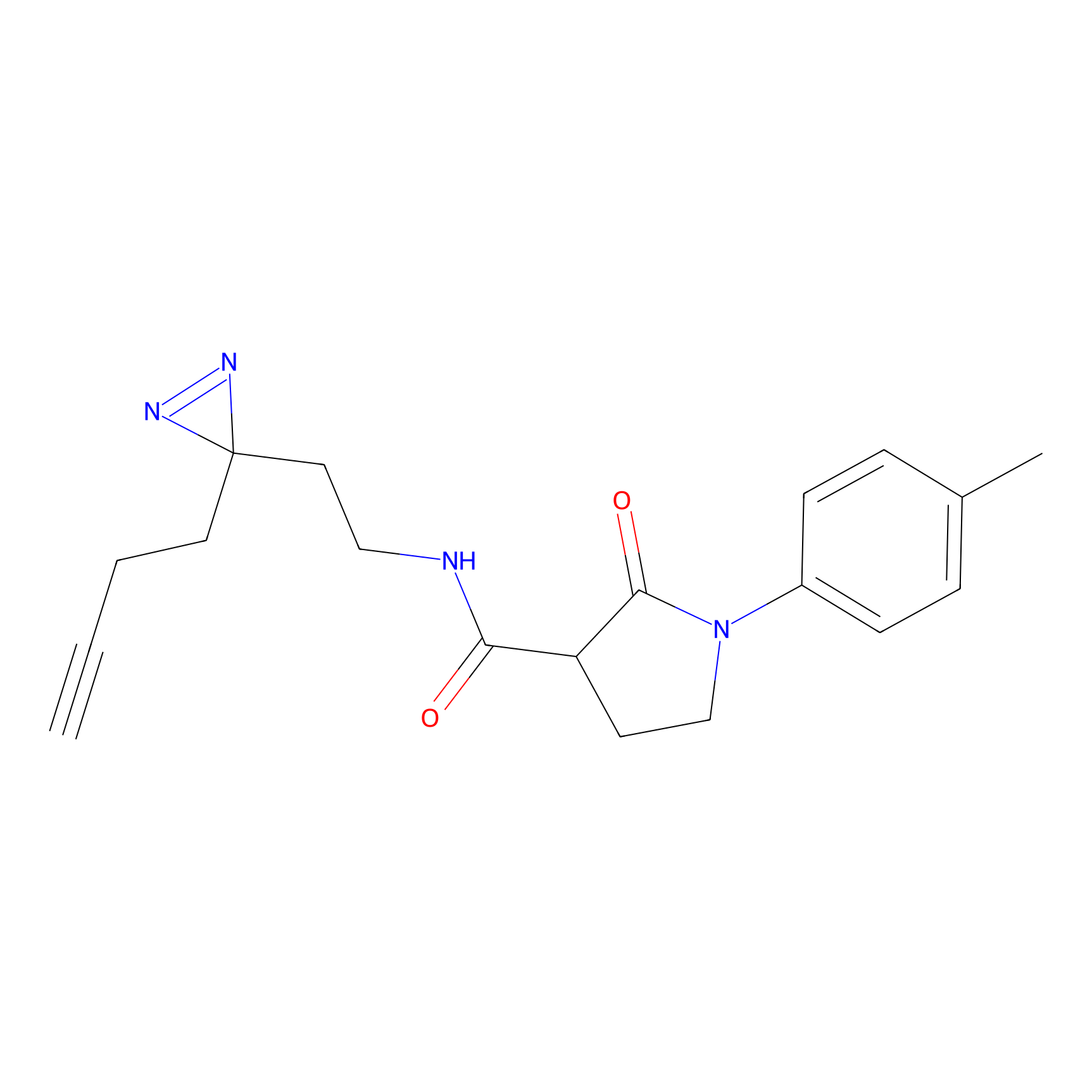
Jackson_14 Probe Info |
 |
2.38 | LDD0123 | [5] | |
|
IPM Probe Info |
 |
N.A. | LDD2156 | [6] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References