Details of the Target
General Information of Target
| Target ID | LDTP09088 | |||||
|---|---|---|---|---|---|---|
| Target Name | Phospholipase A2 group XV (PLA2G15) | |||||
| Gene Name | PLA2G15 | |||||
| Gene ID | 23659 | |||||
| Synonyms |
LYPLA3; Phospholipase A2 group XV; 1-O-acylceramide synthase; ACS; LCAT-like lysophospholipase; LLPL; EC 3.1.1.5; Lysophospholipase 3; Lysosomal phospholipase A and acyltransferase; EC 2.3.1.-, EC 3.1.1.32, EC 3.1.1.4; Lysosomal phospholipase A2; LPLA2
|
|||||
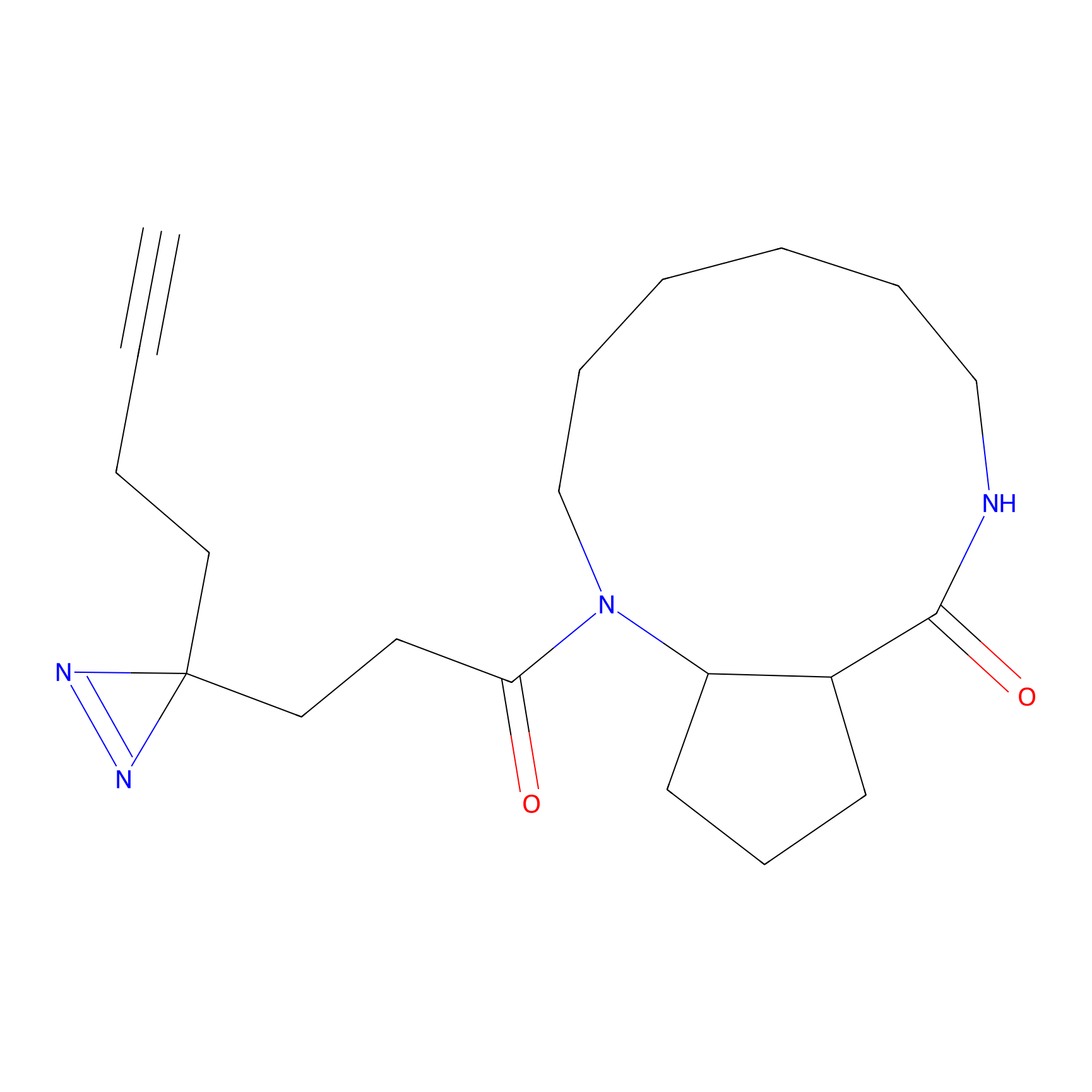
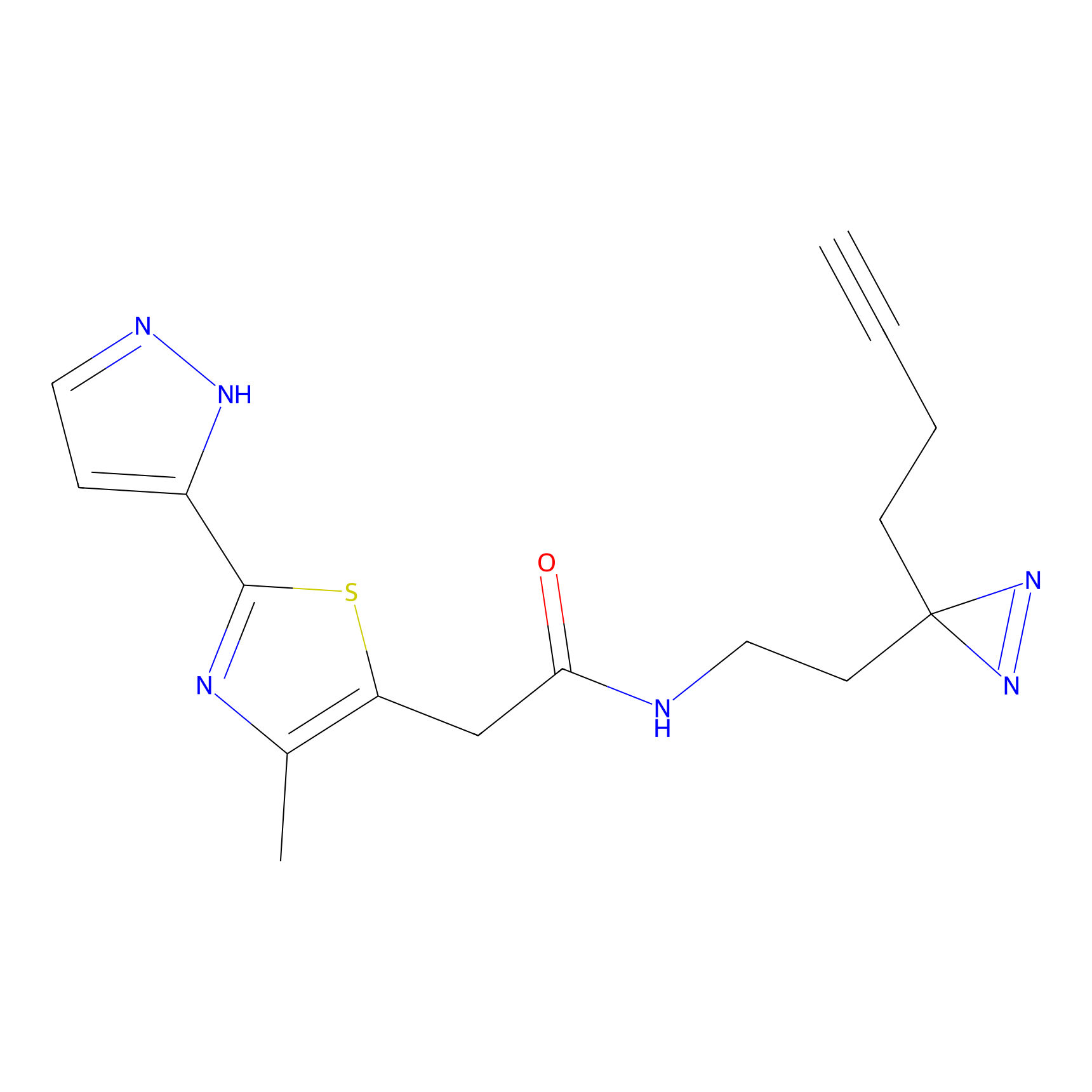
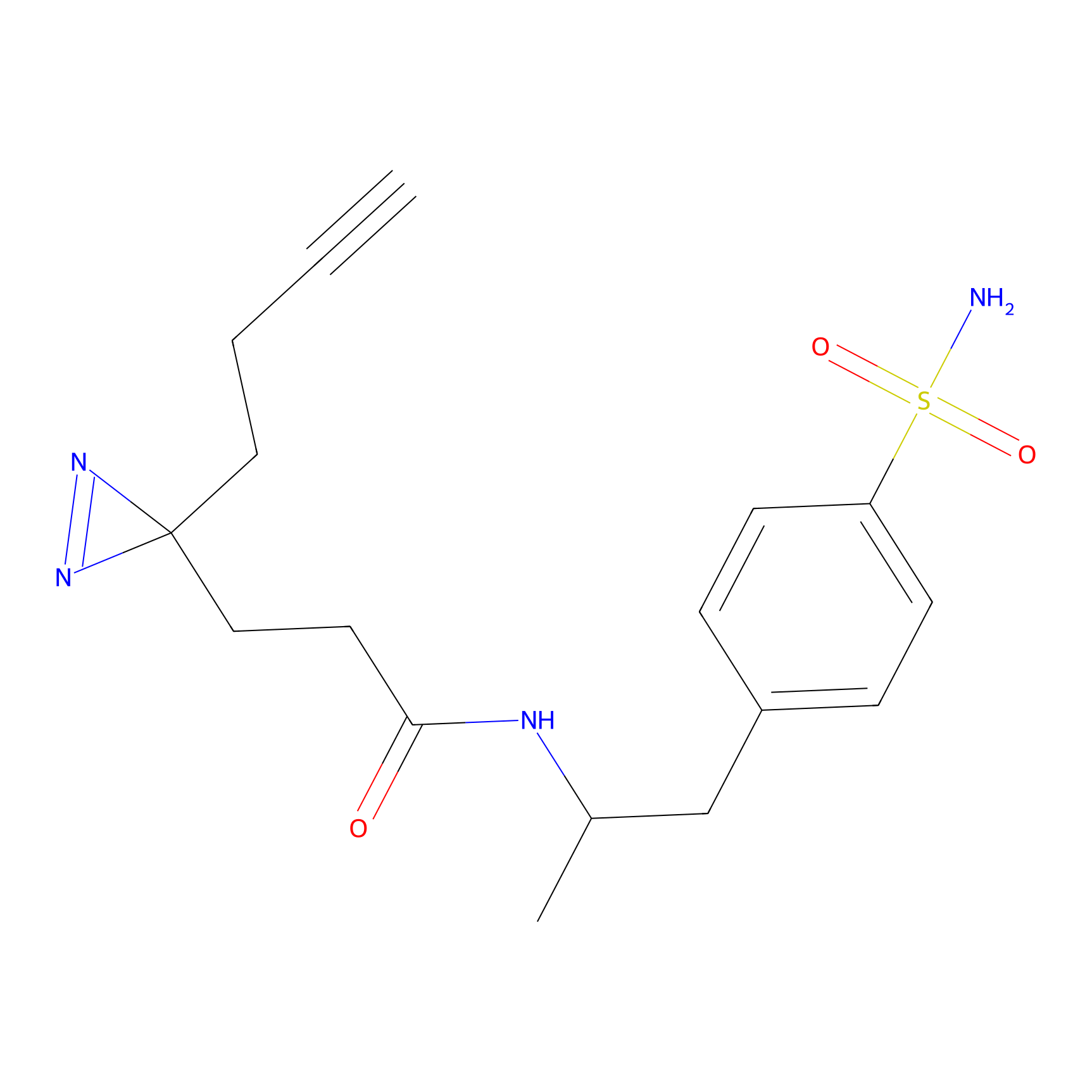
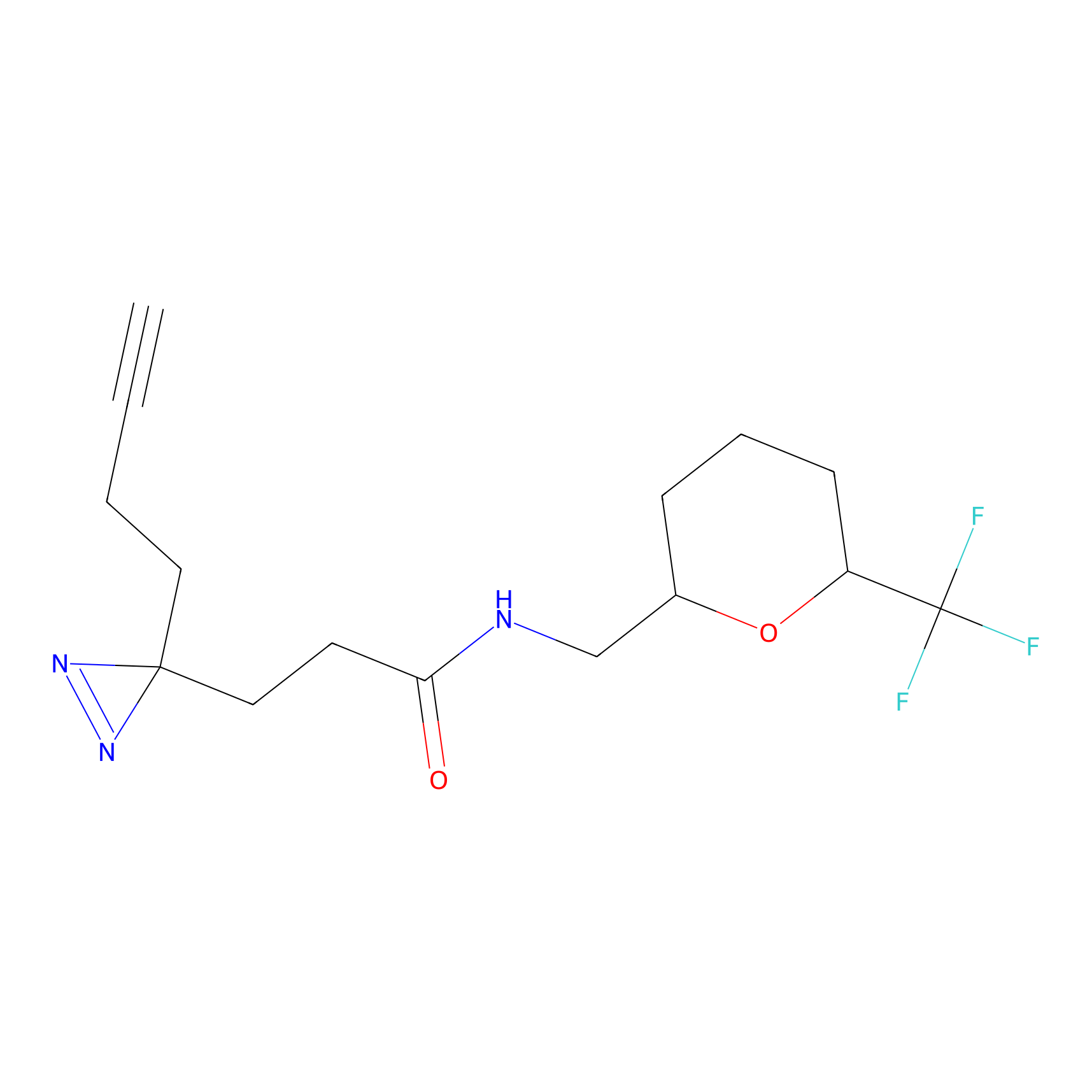
| 3D Structure | ||||||
| Sequence |
MGLHLRPYRVGLLPDGLLFLLLLLMLLADPALPAGRHPPVVLVPGDLGNQLEAKLDKPTV
VHYLCSKKTESYFTIWLNLELLLPVIIDCWIDNIRLVYNKTSRATQFPDGVDVRVPGFGK TFSLEFLDPSKSSVGSYFHTMVESLVGWGYTRGEDVRGAPYDWRRAPNENGPYFLALREM IEEMYQLYGGPVVLVAHSMGNMYTLYFLQRQPQAWKDKYIRAFVSLGAPWGGVAKTLRVL ASGDNNRIPVIGPLKIREQQRSAVSTSWLLPYNYTWSPEKVFVQTPTINYTLRDYRKFFQ DIGFEDGWLMRQDTEGLVEATMPPGVQLHCLYGTGVPTPDSFYYESFPDRDPKICFGDGD GTVNLKSALQCQAWQSRQEHQVLLQELPGSEHIEMLANATTLAYLKRVLLGP |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
AB hydrolase superfamily, Lipase family
|
|||||
| Subcellular location |
Lysosome
|
|||||
| Function |
Has dual calcium-independent phospholipase and O-acyltransferase activities with a potential role in glycerophospholipid homeostasis and remodeling of acyl groups of lipophilic alcohols present in acidic cellular compartments . Catalyzes hydrolysis of the ester bond of the fatty acyl group attached at sn-1 or sn-2 position of phospholipids (phospholipase A1 or A2 activity) and transfer it to the hydroxyl group at the first carbon of lipophilic alcohols (O-acyltransferase activity) . Among preferred fatty acyl donors are phosphatidylcholines, phosphatidylethanolamines, phosphatidylglycerols and phosphatidylserines. Favors sn-2 over sn-1 deacylation of unsaturated fatty acyl groups of phosphatidylcholines and phosphatidylethanolamines. Among preferred fatty acyl acceptors are natural lipophilic alcohols including short-chain ceramide N-acetyl-sphingosine (C2 ceramide), alkylacylglycerols, monoacylglycerols, and acylethanolamides such as anandamide and oleoylethanolamide. Selectively hydrolyzes the sn-1 fatty acyl group of truncated oxidized phospholipids and may play a role in detoxification of reactive oxidized phospholipids during oxidative stress. Required for normal phospholipid degradation in alveolar macrophages with potential implications in pulmonary surfactant clearance. At neutral pH, hydrolyzes the sn-1 fatty acyl group of the lysophosphatidylcholines.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| JURKAT | SNV: p.S198N | . | |||
| OE33 | SNV: p.R6H | DBIA Probe Info | |||
| PC9 | SNV: p.M1? | . | |||
| SNU1 | SNV: p.E391K | DBIA Probe Info | |||
| WM2664 | Insertion: p.I91DfsTer2 | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
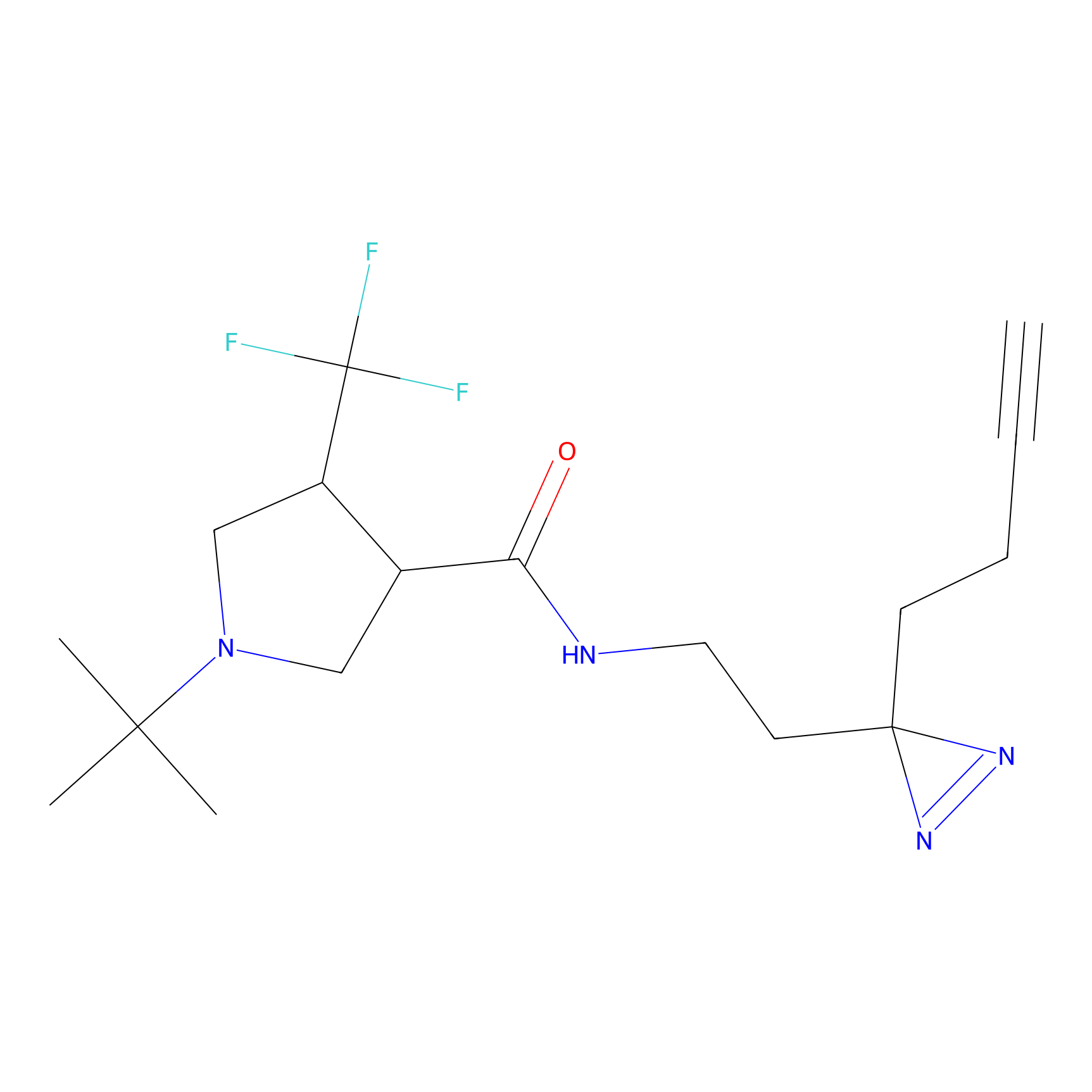
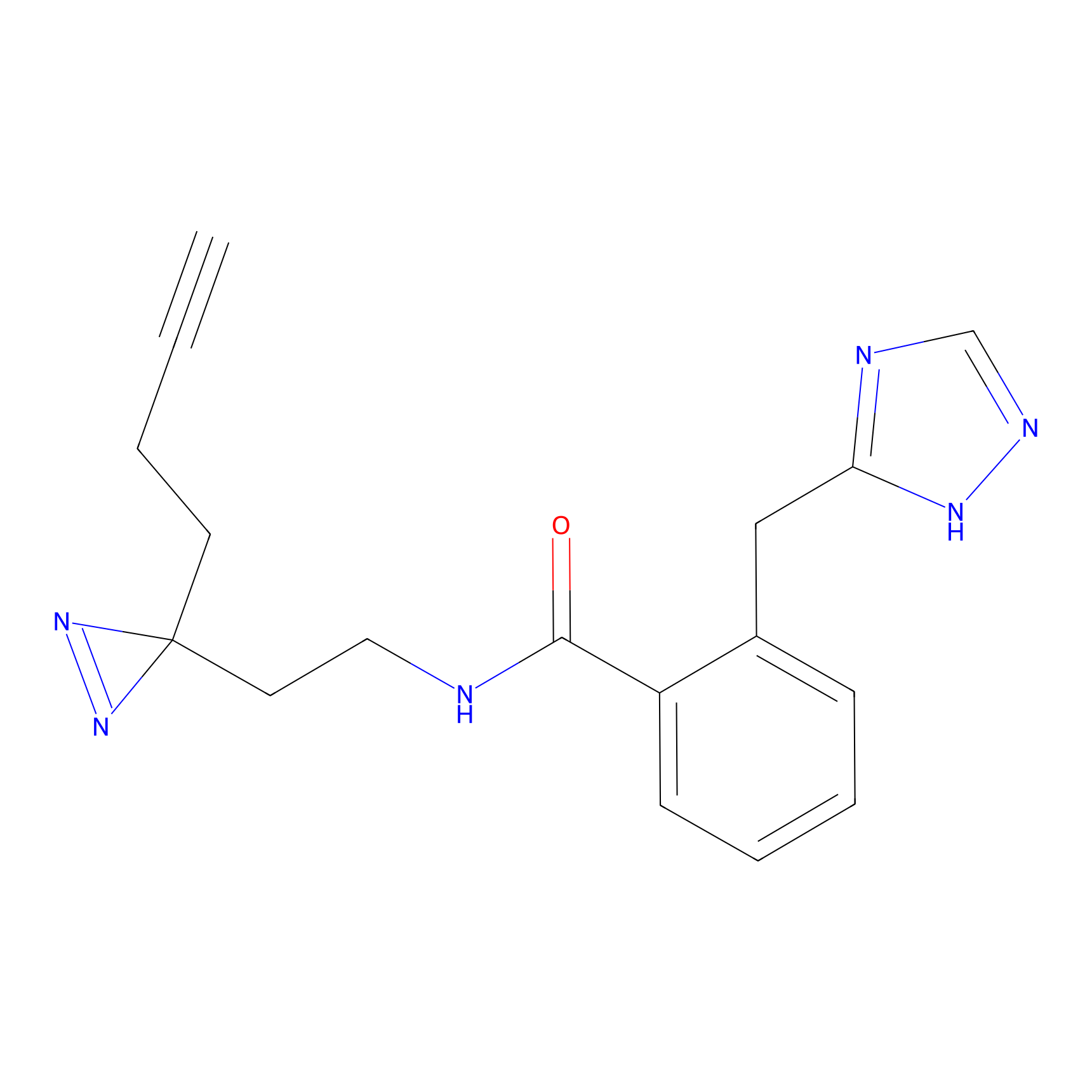
|
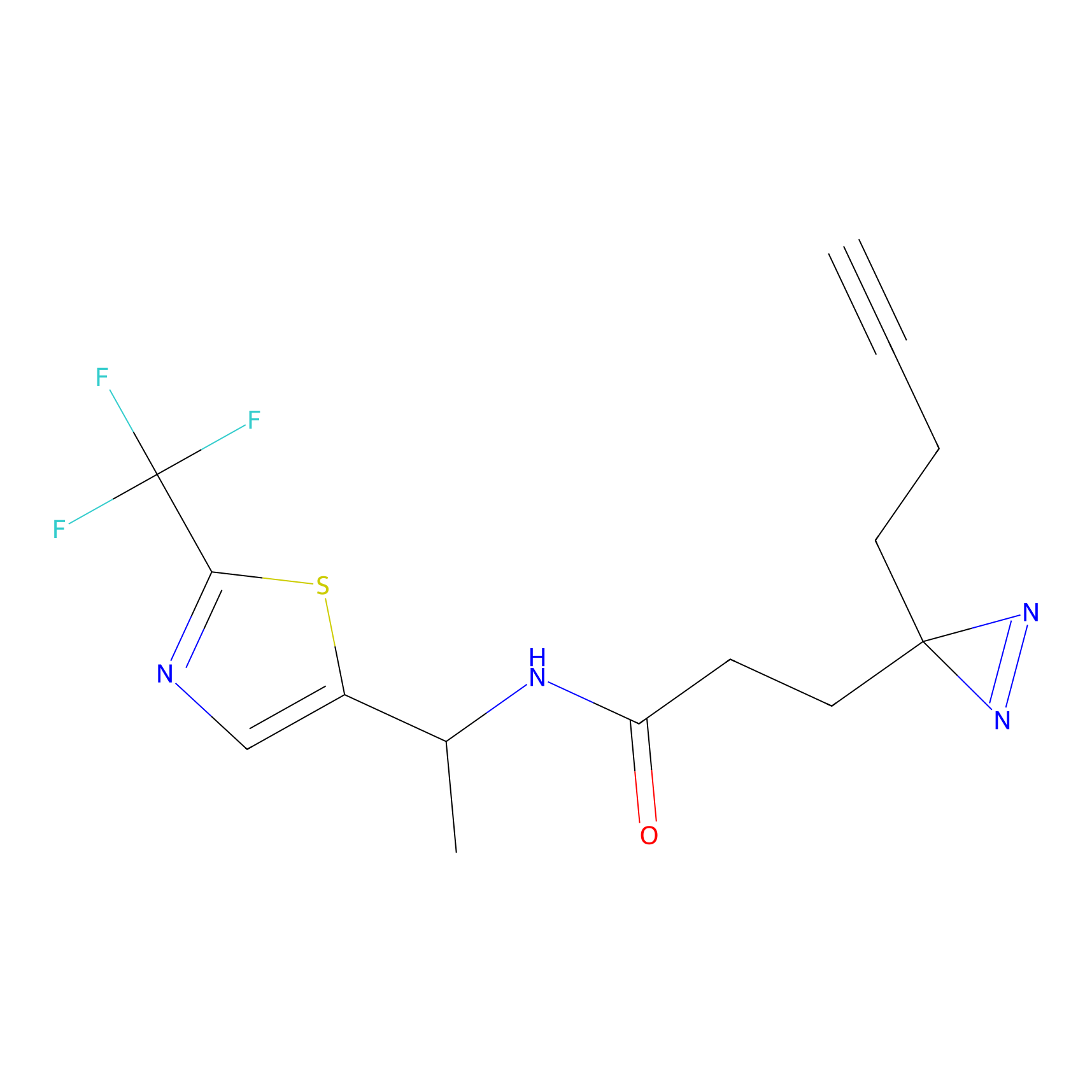
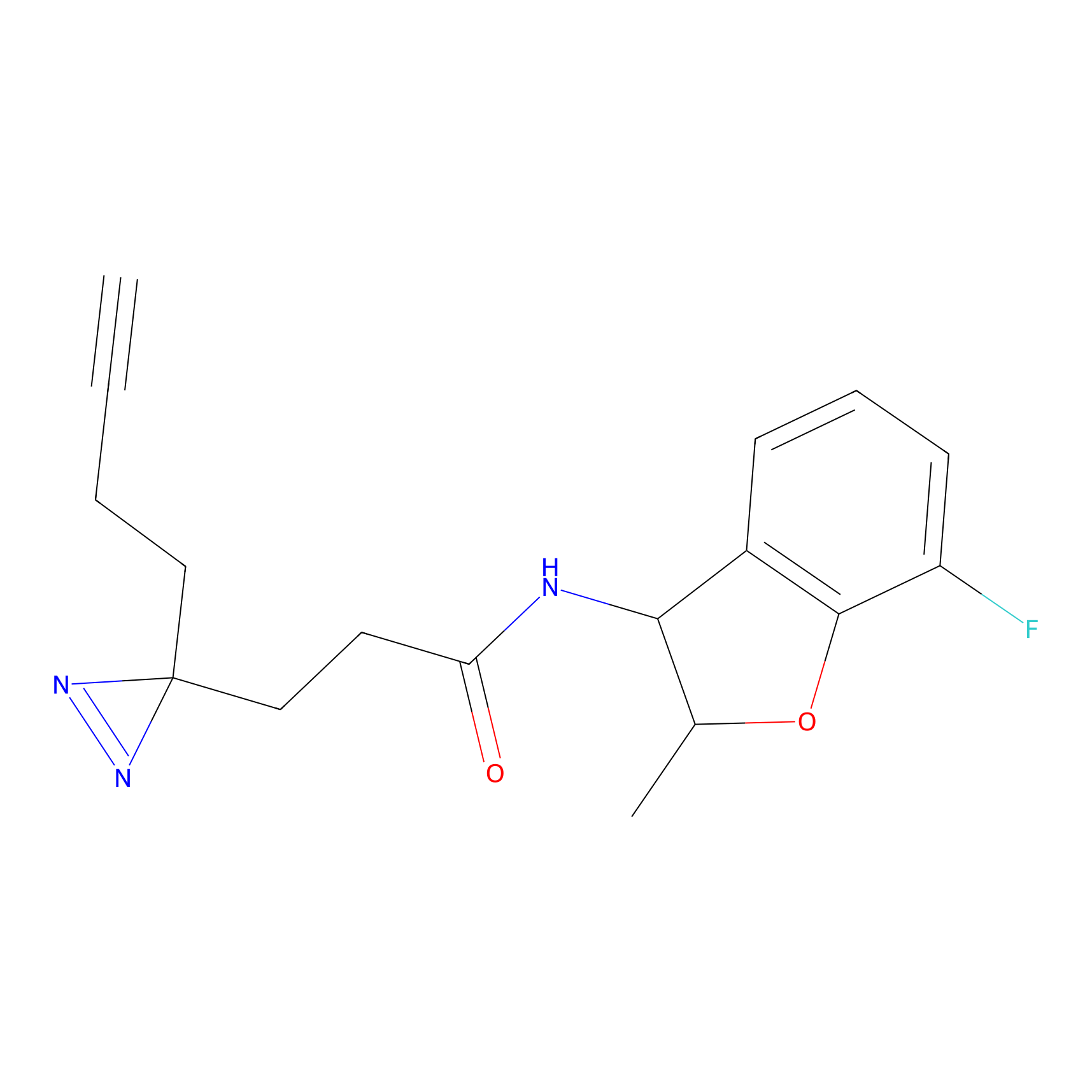
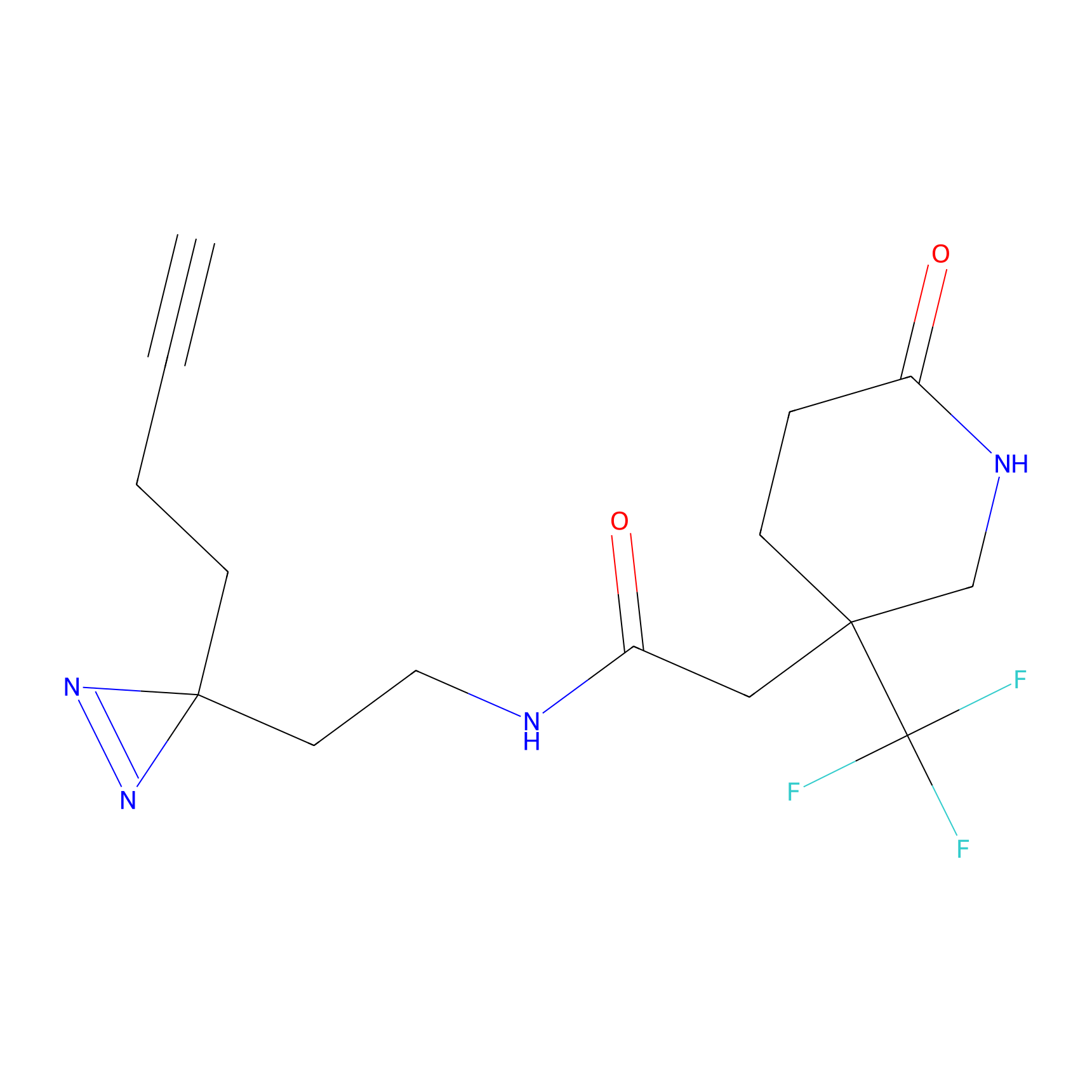
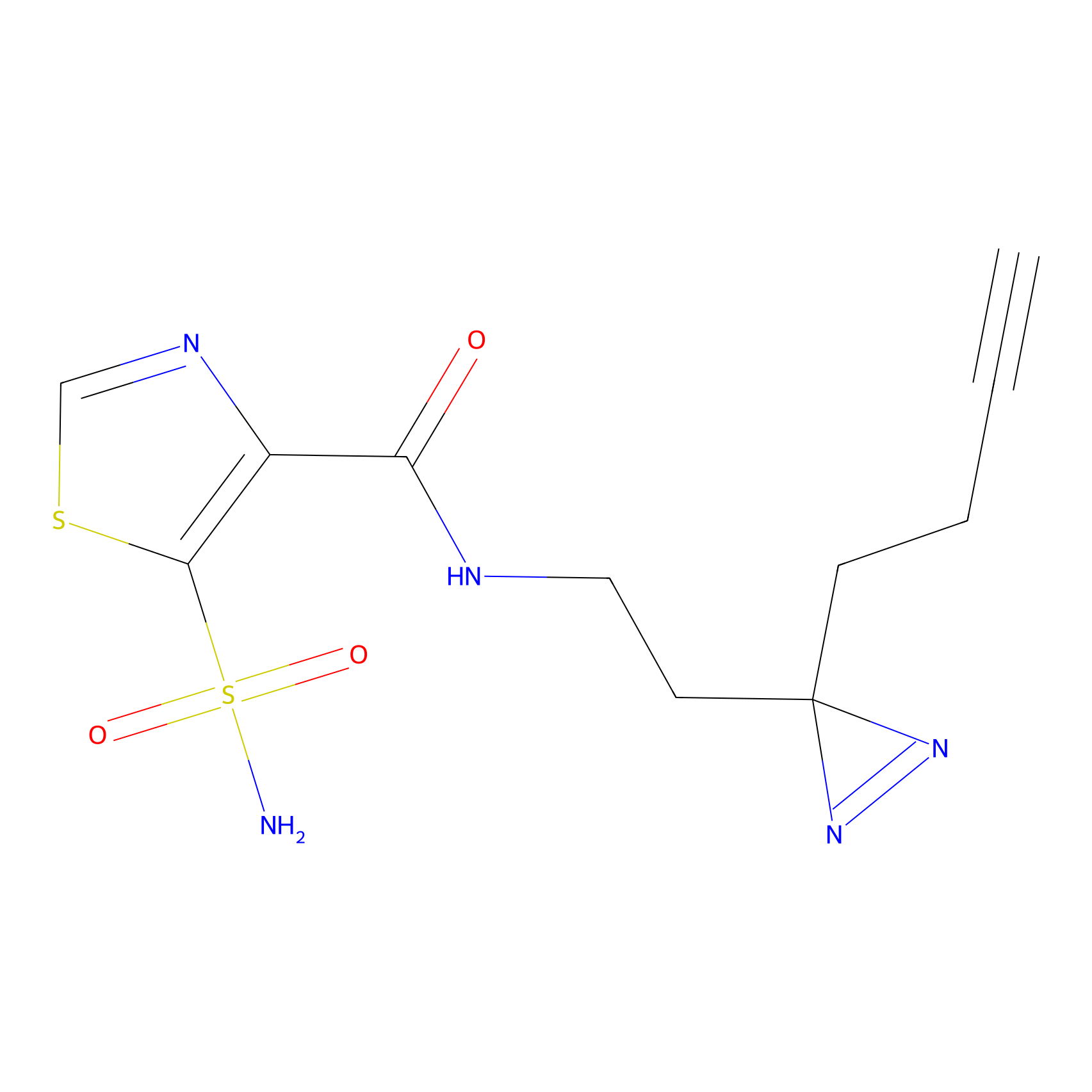
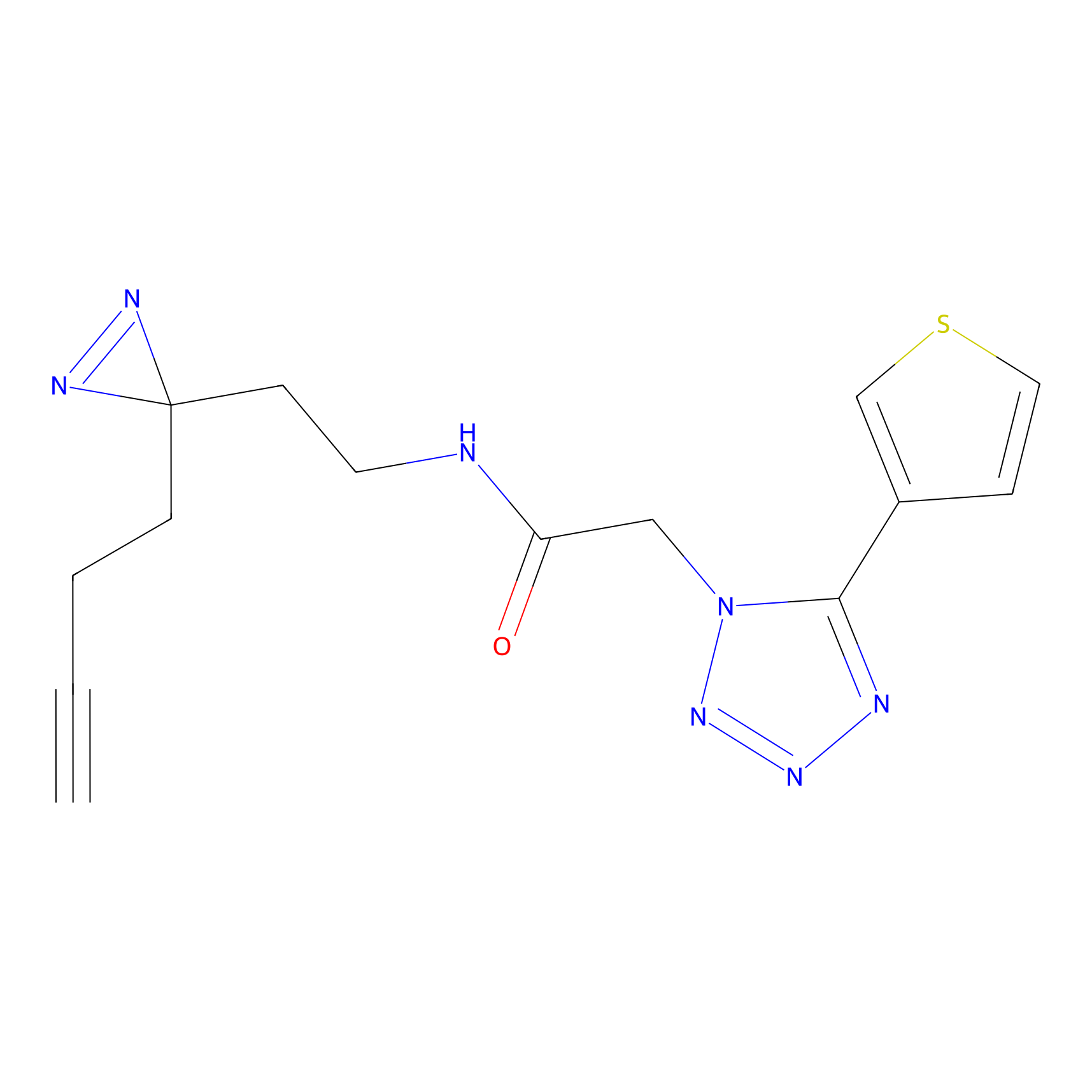
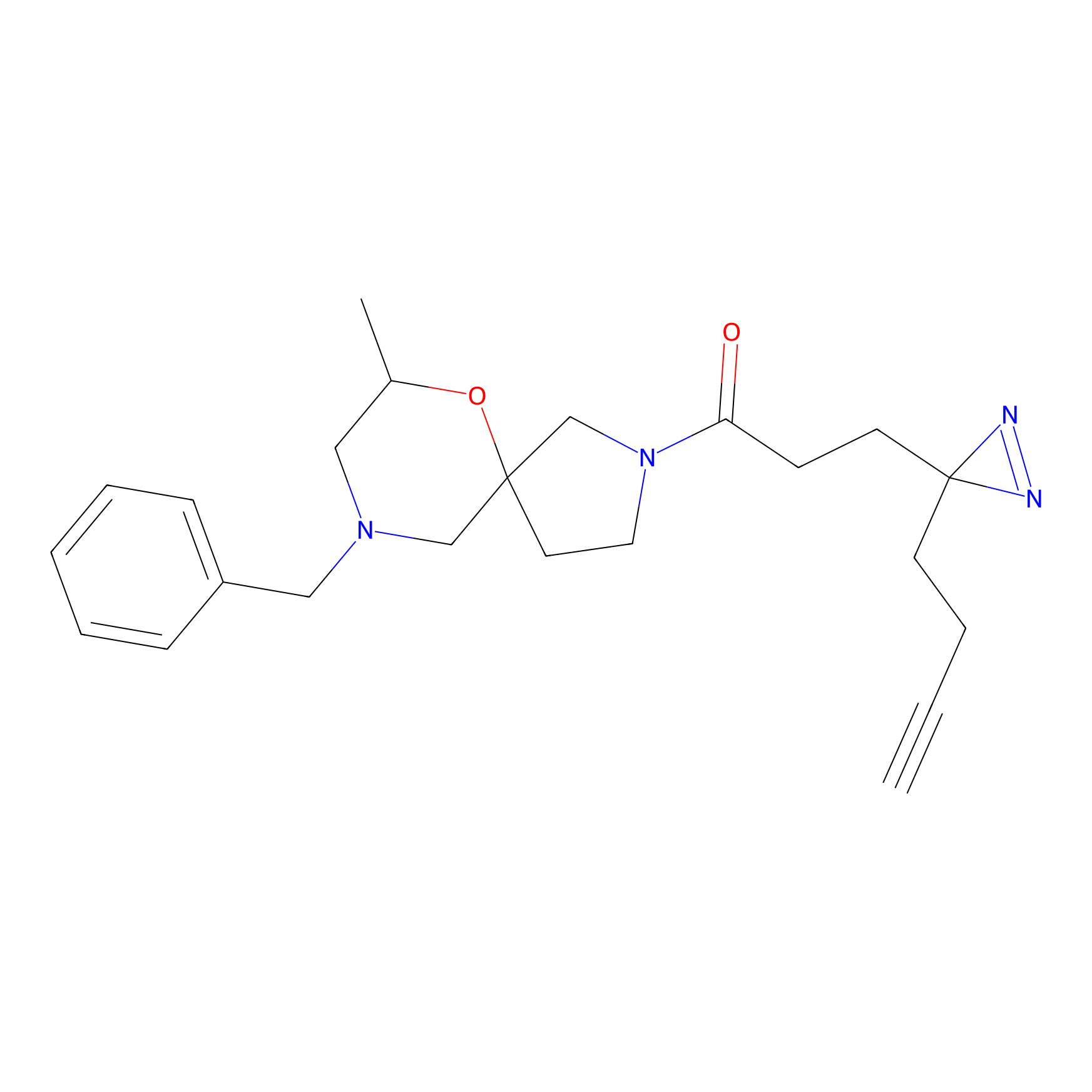
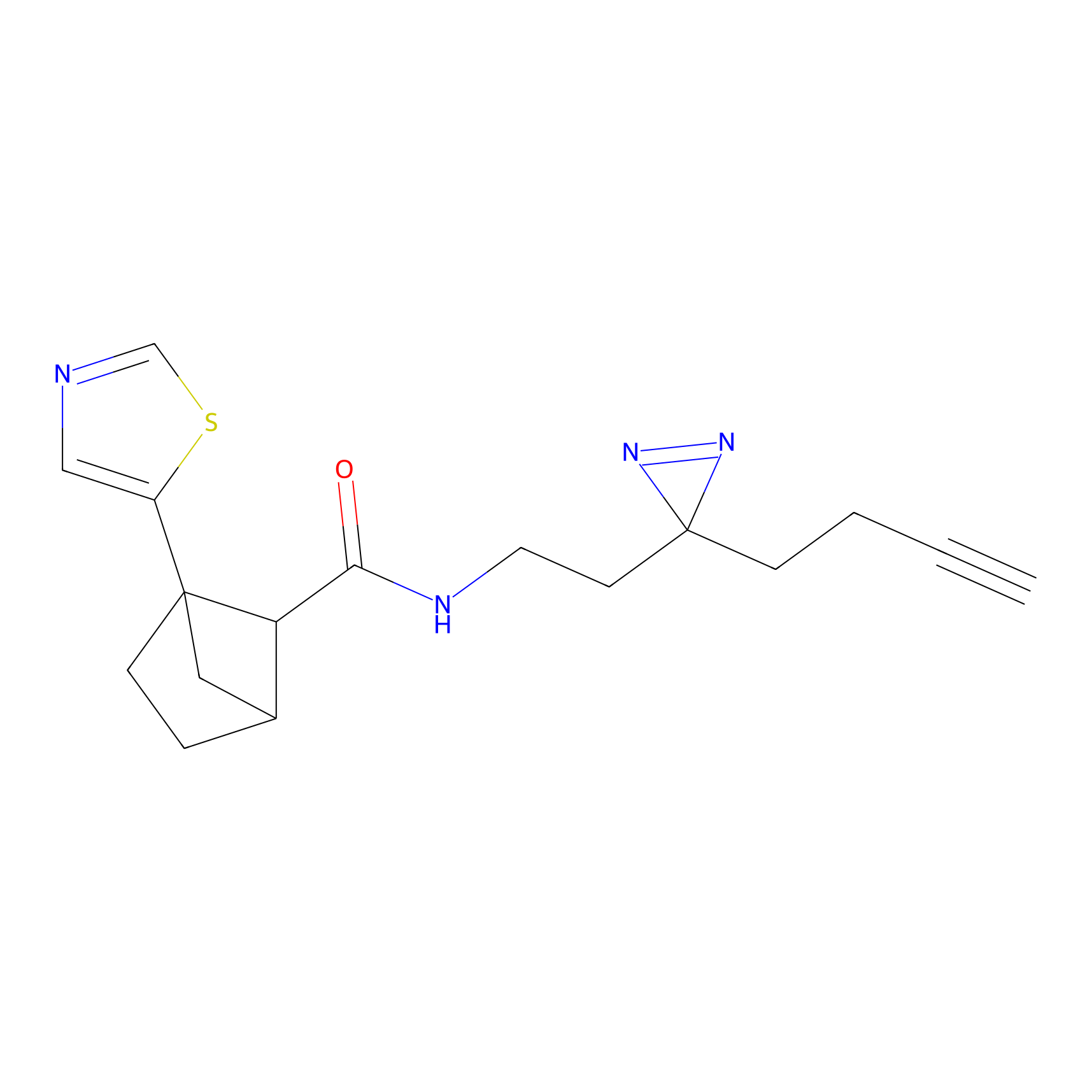
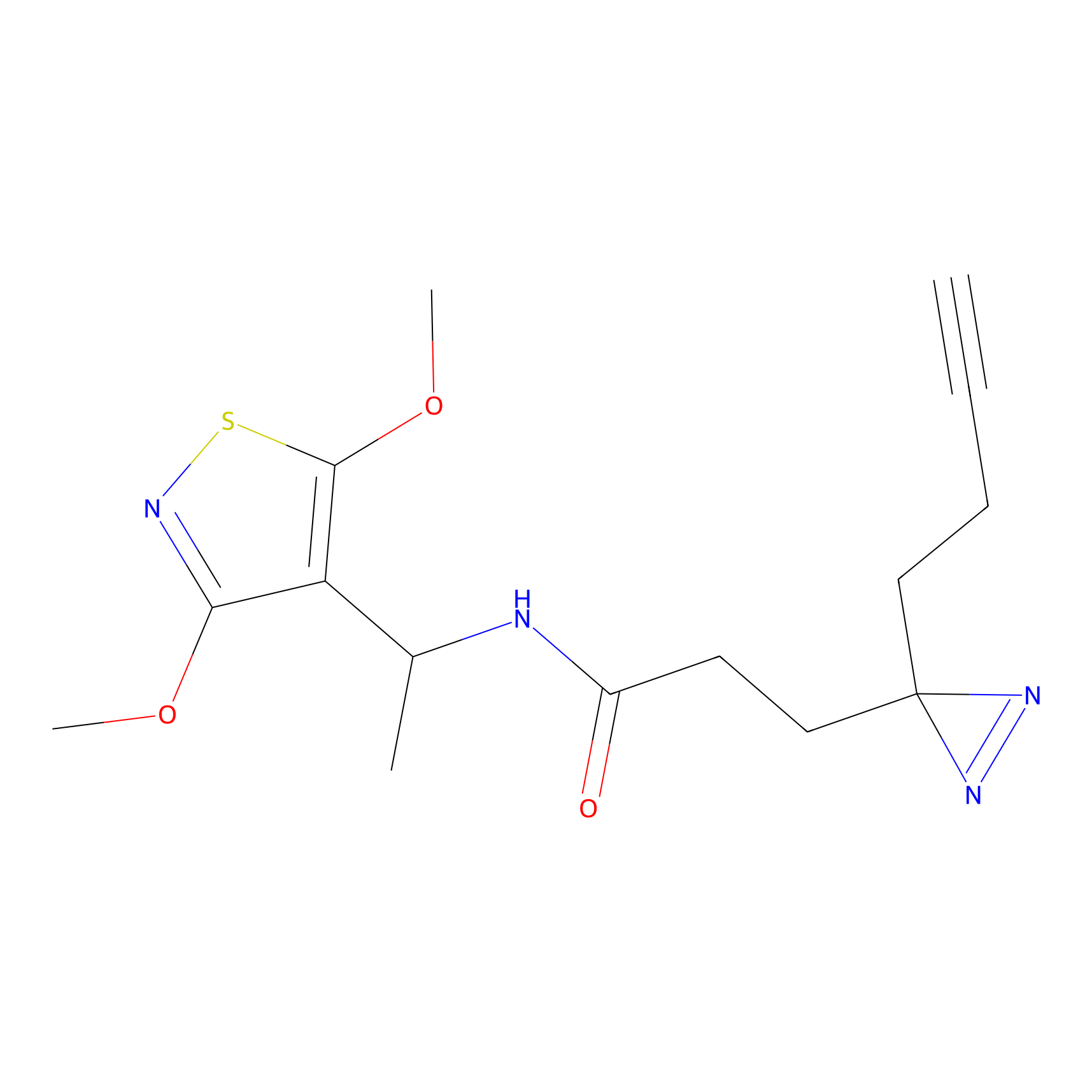
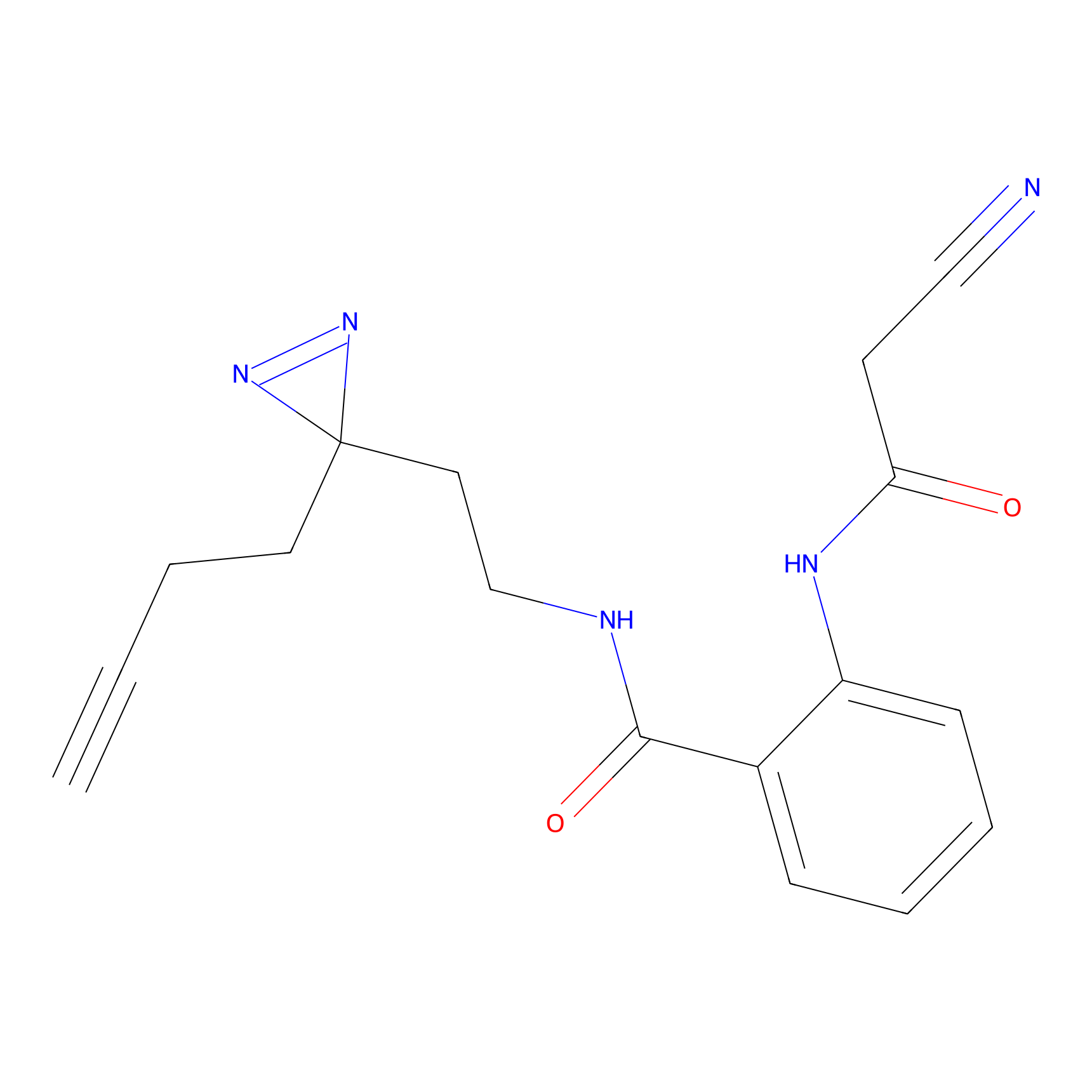
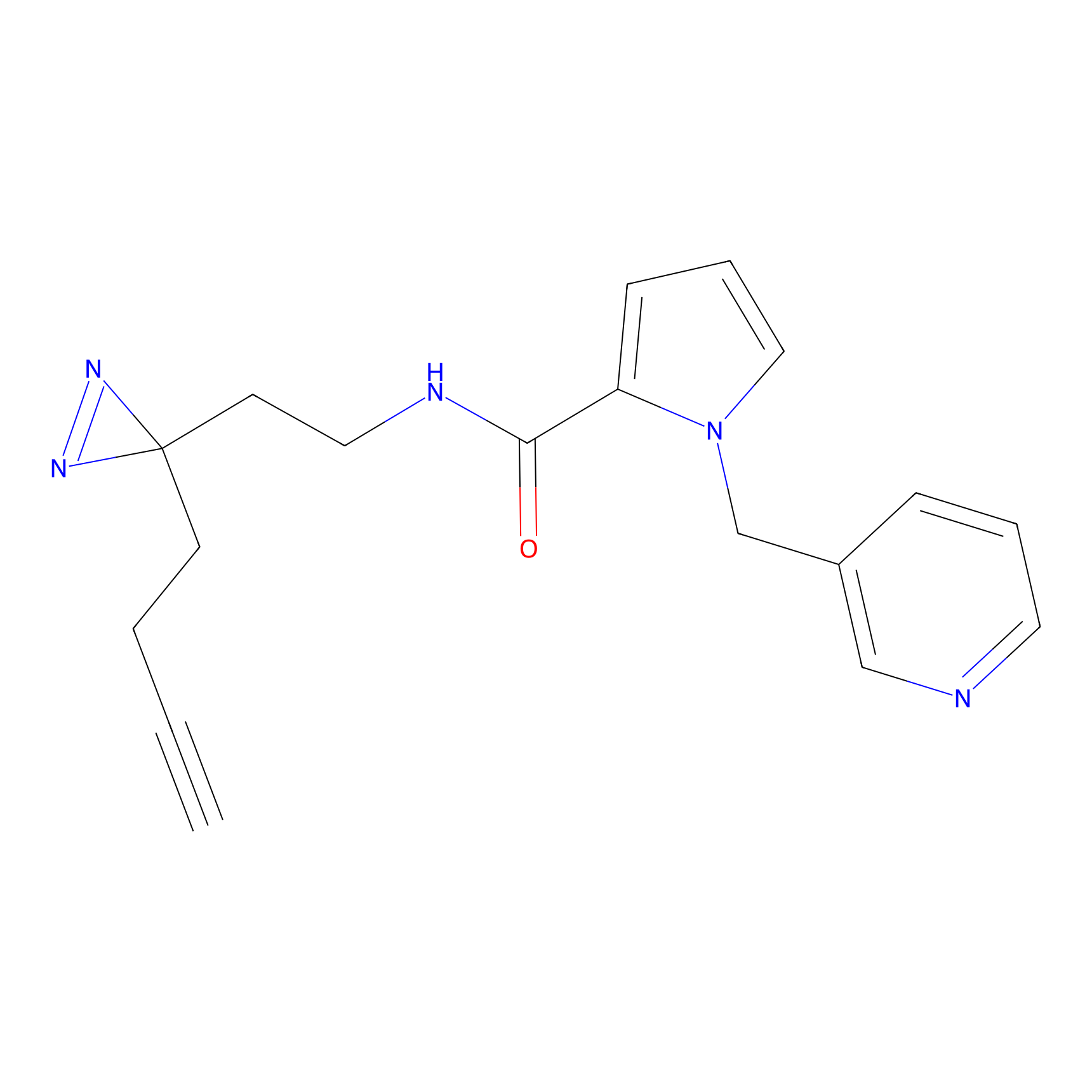
HDSF-alk Probe Info |
 |
5.32 | LDD0197 | [1] | |
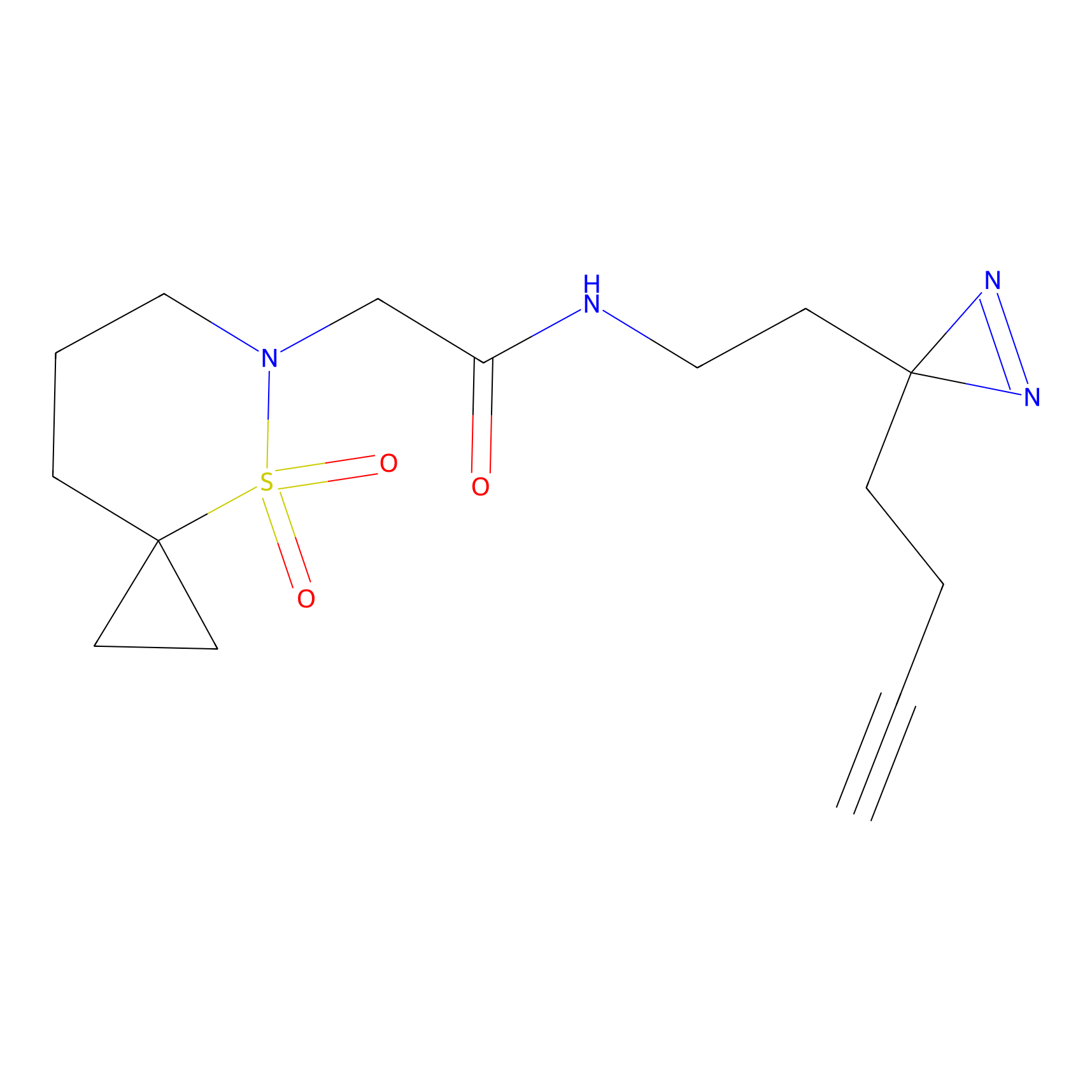
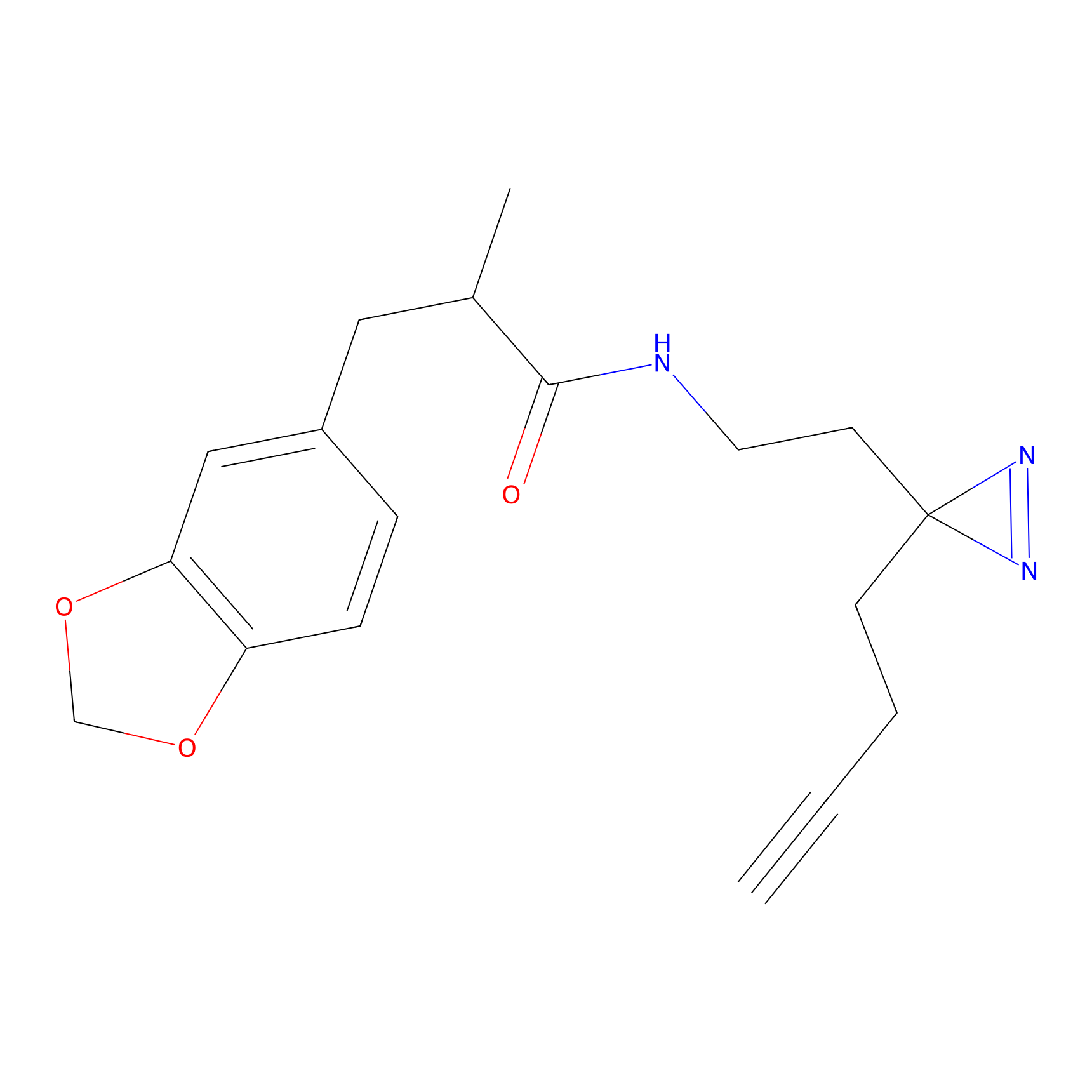
|
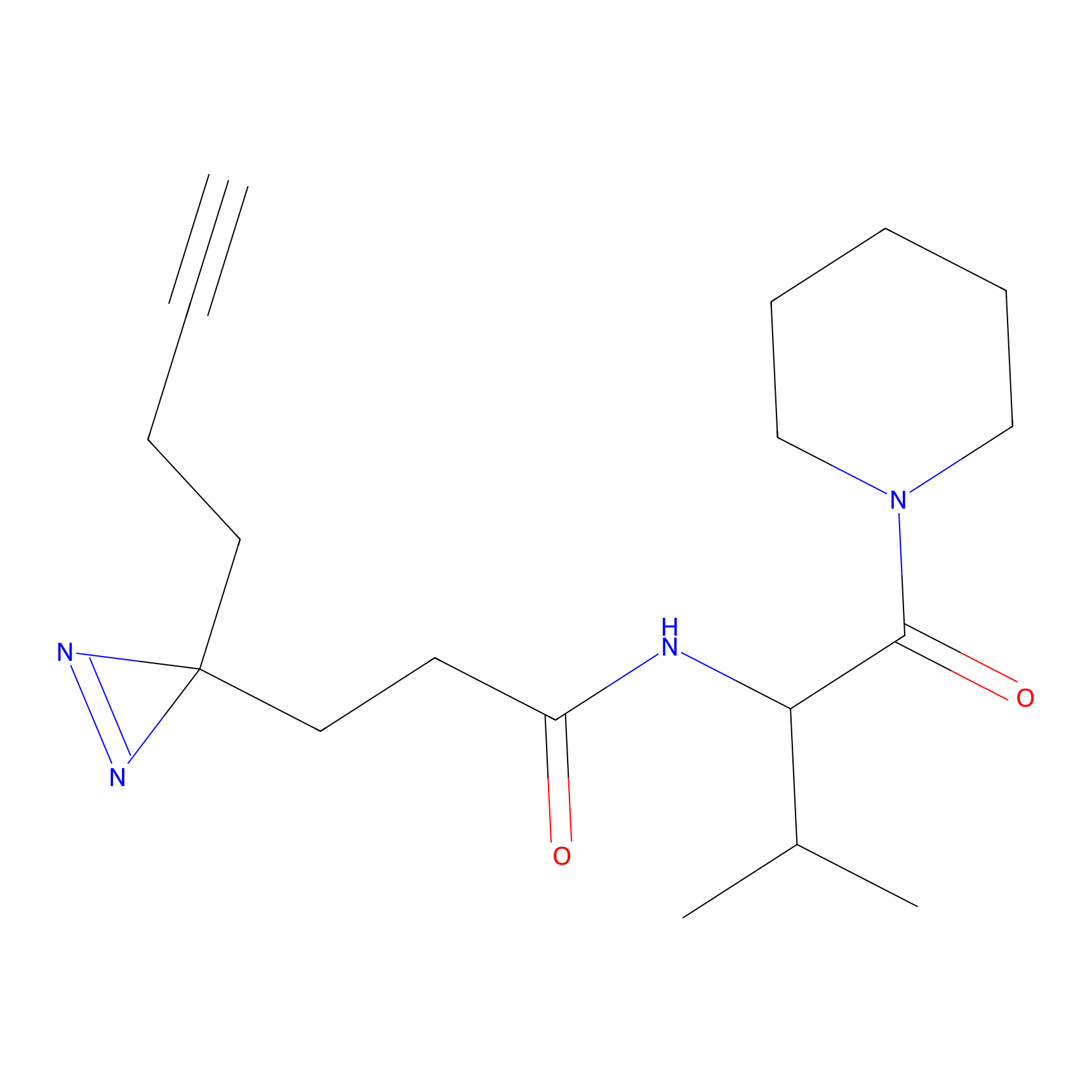
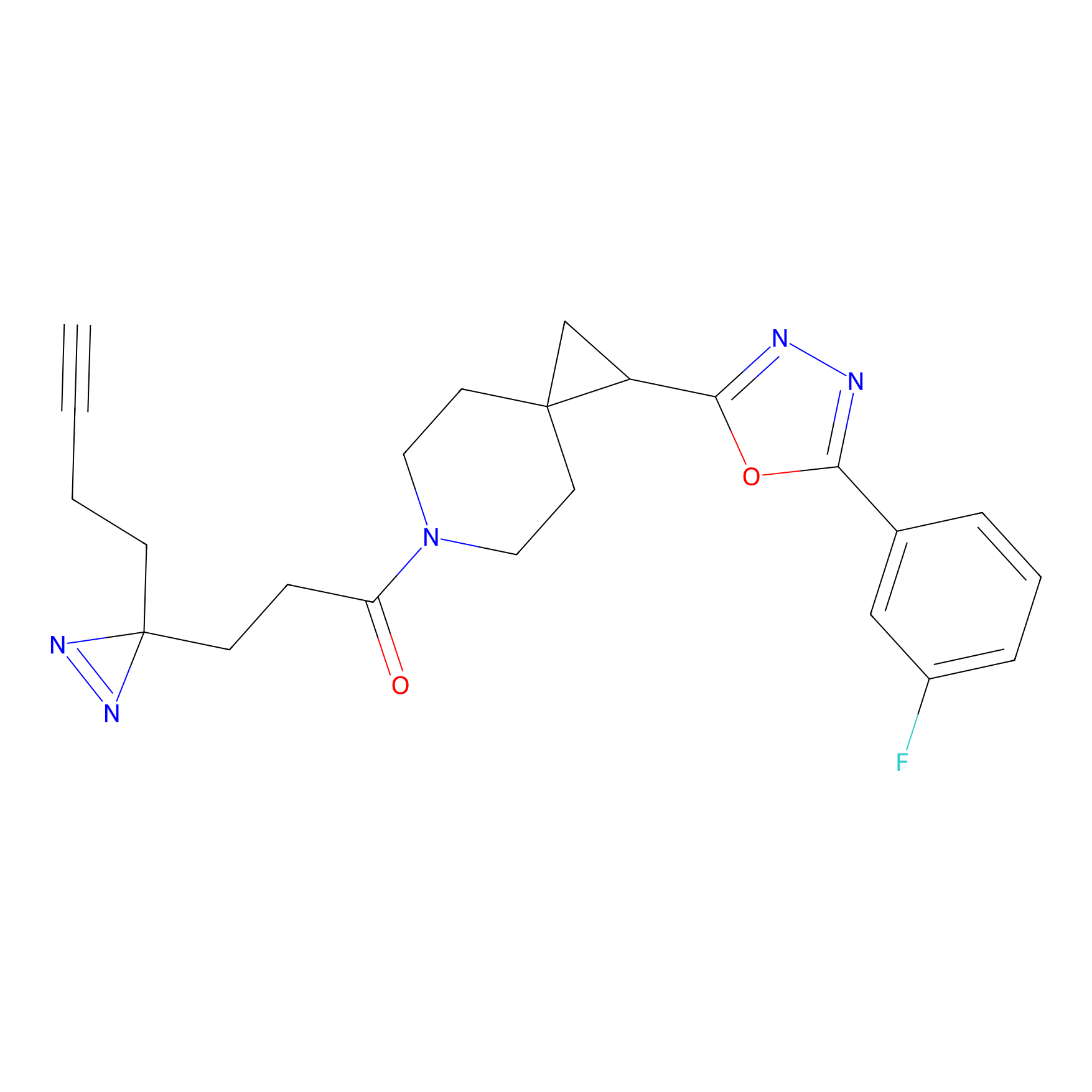
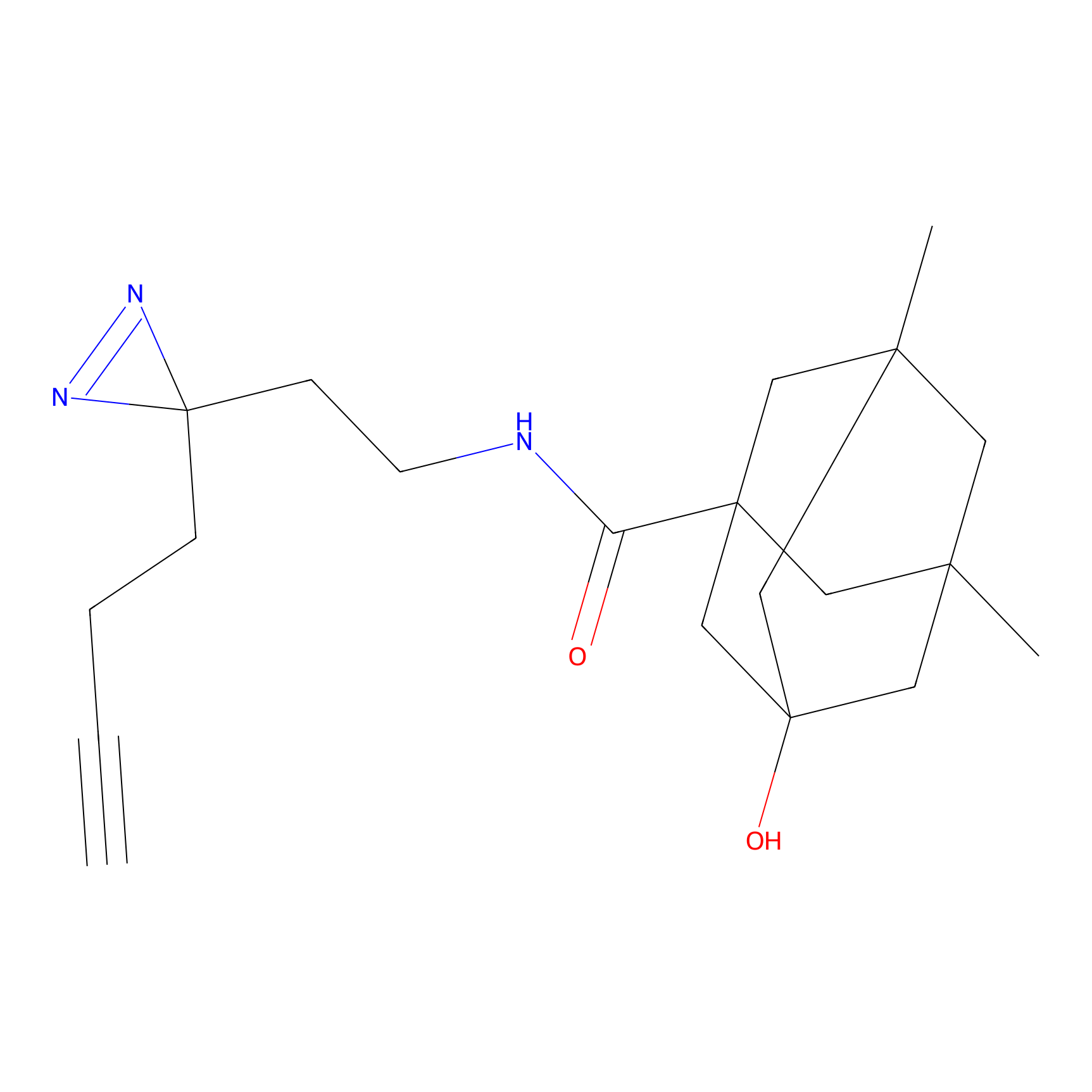
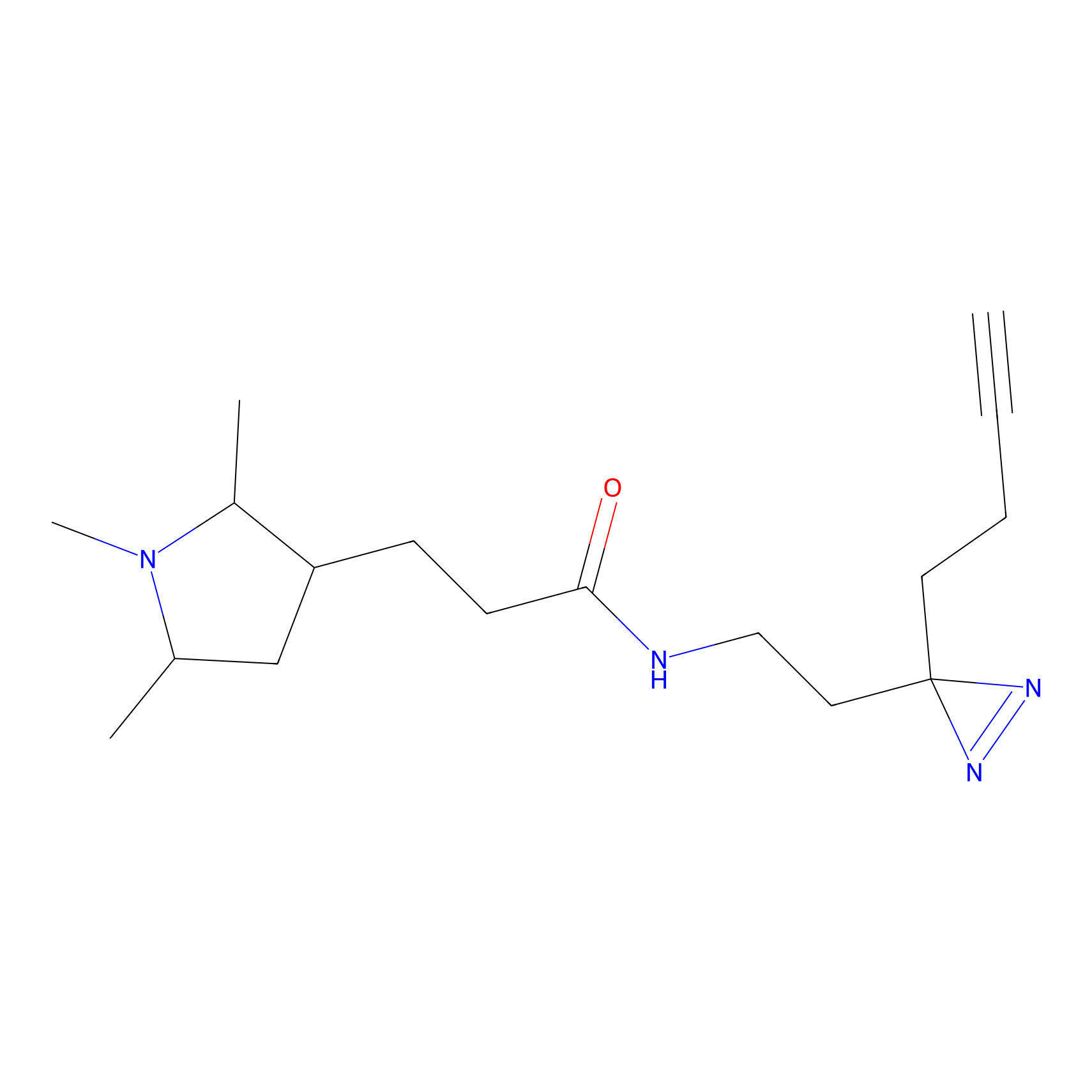
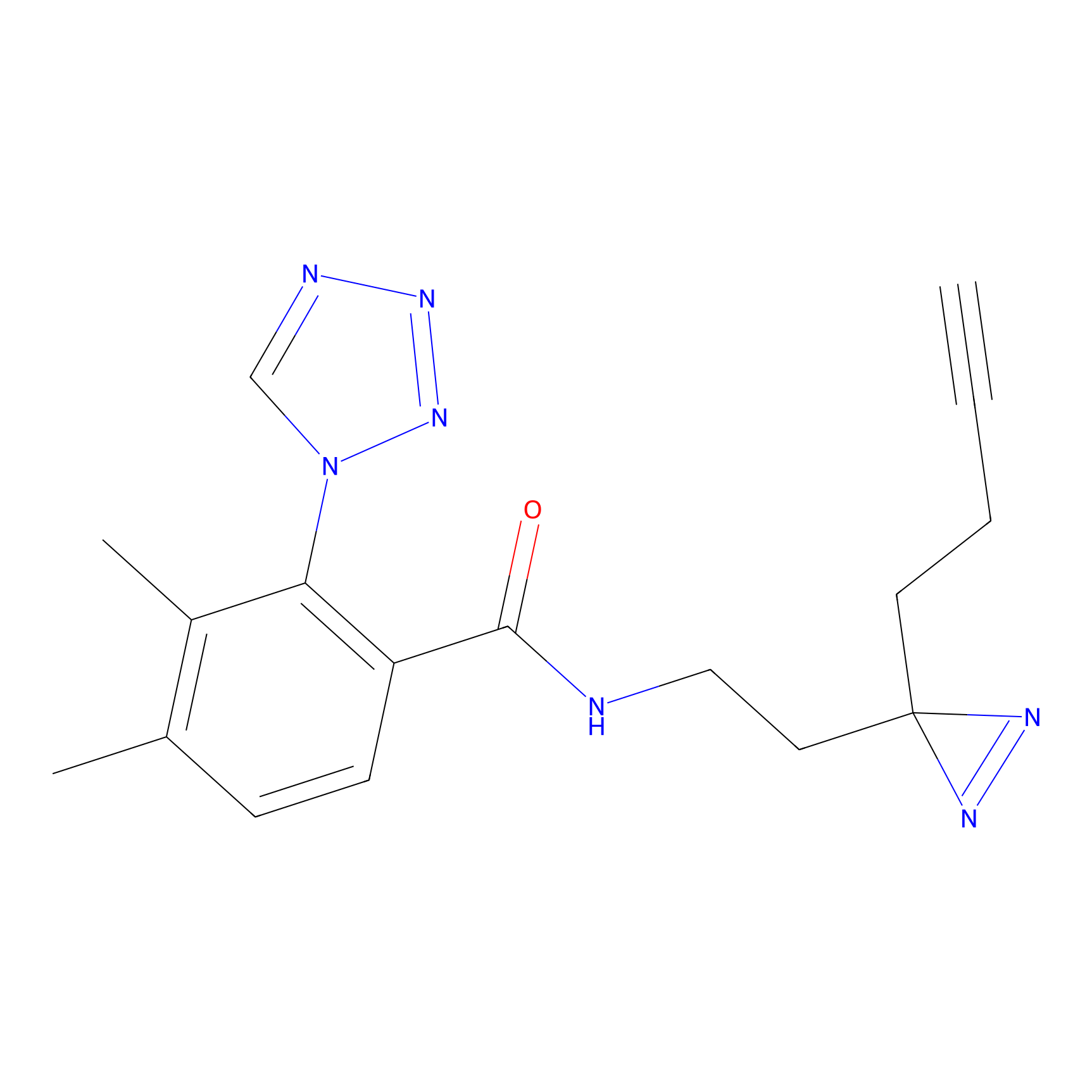
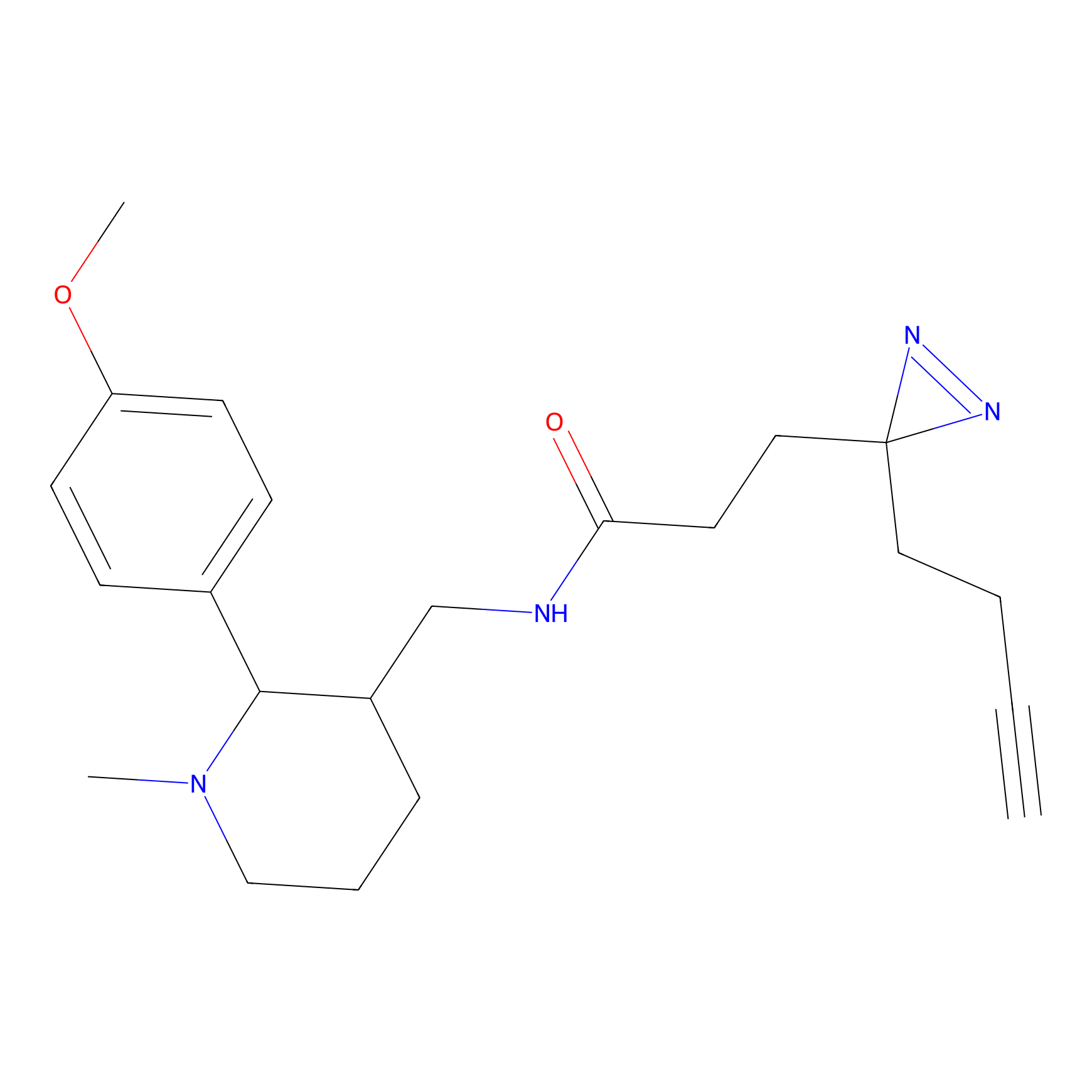
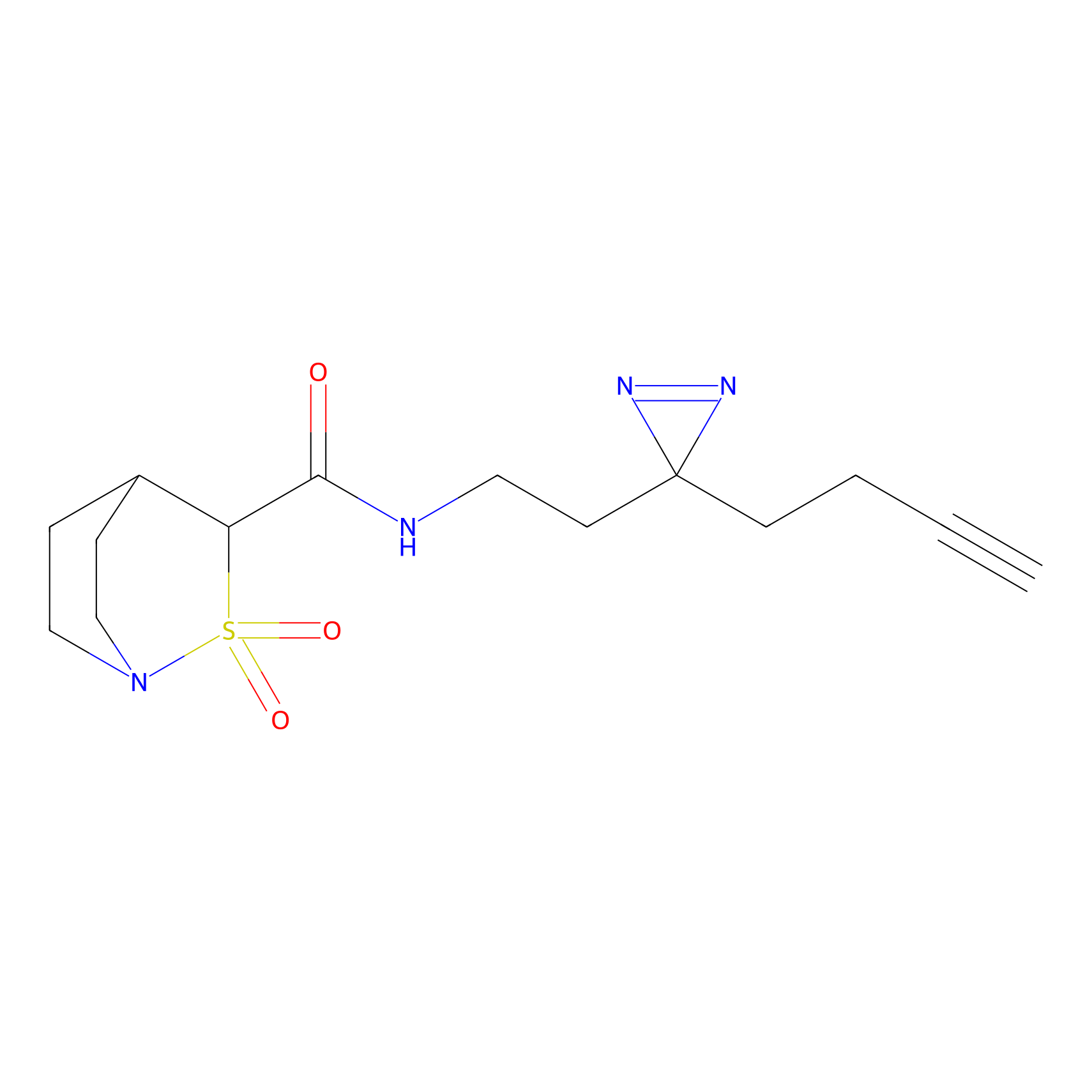
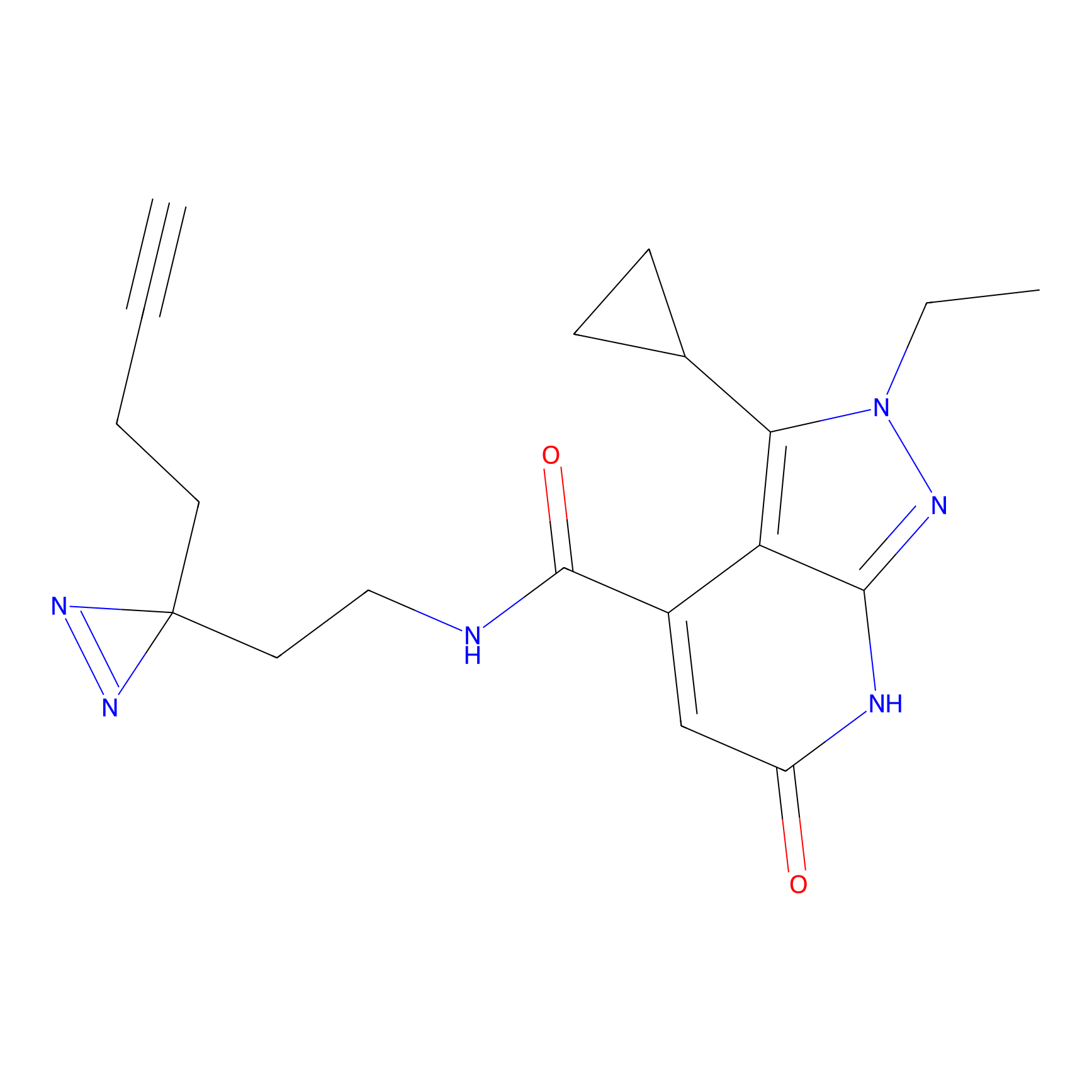
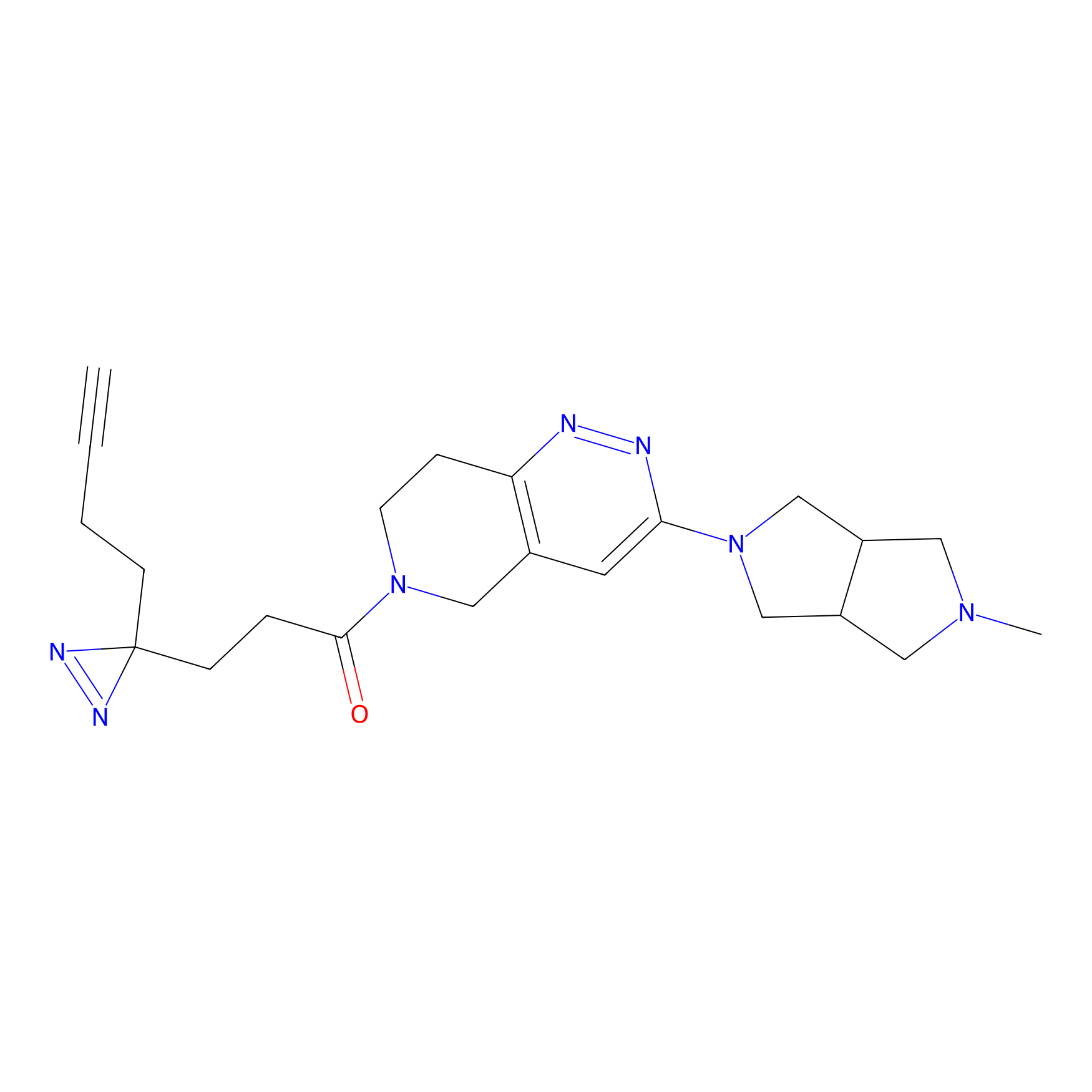
IPM Probe Info |
 |
N.A. | LDD0241 | [2] | |
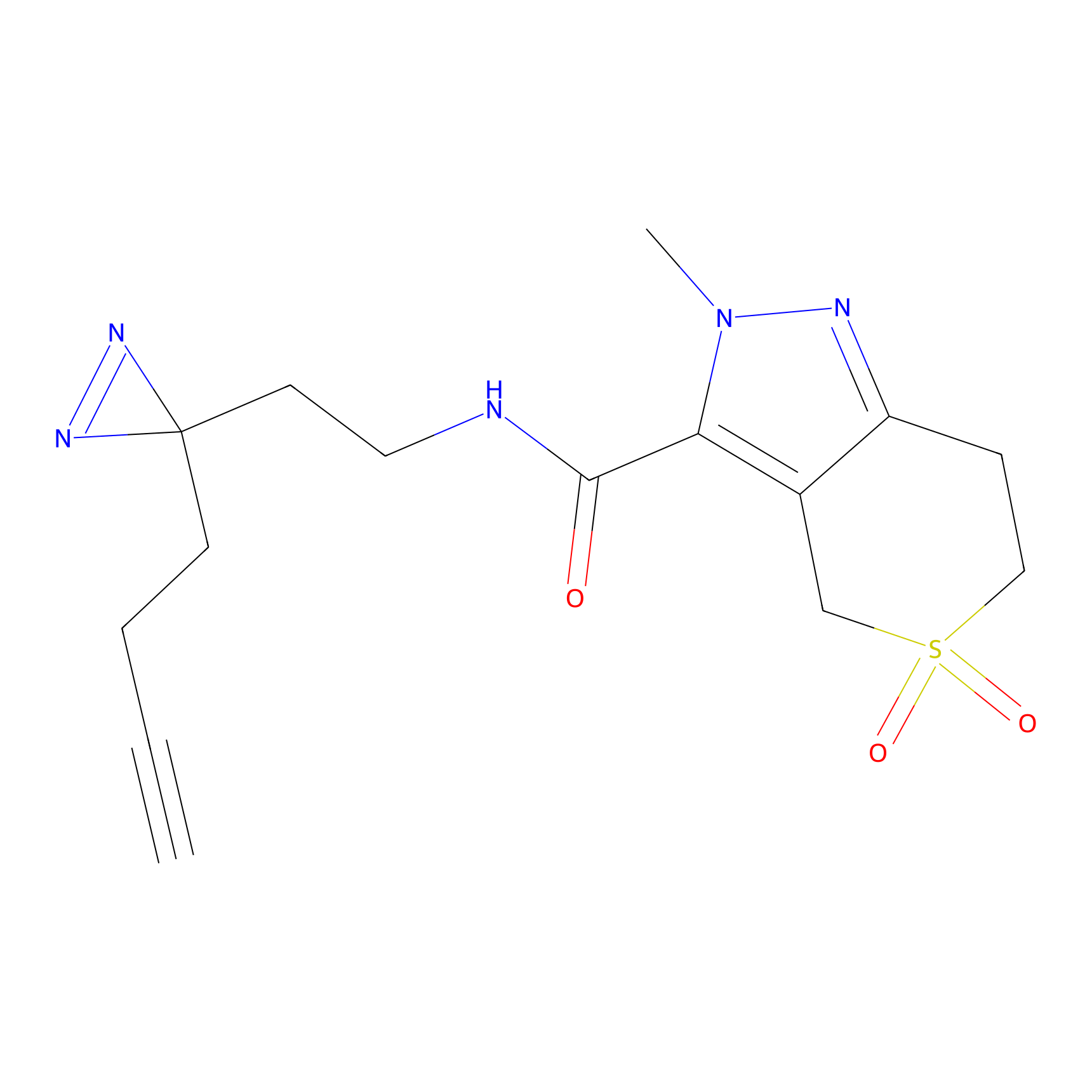
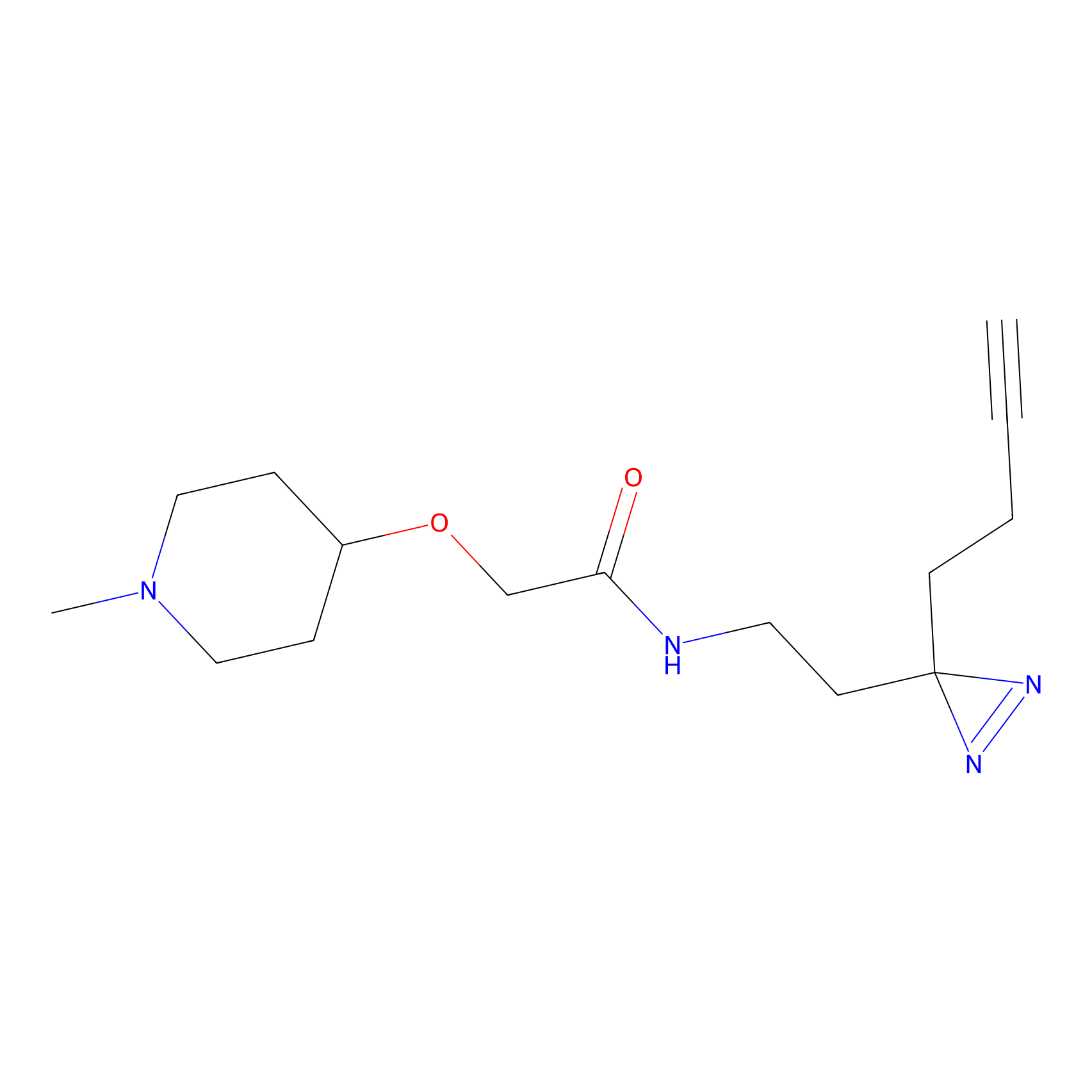
|
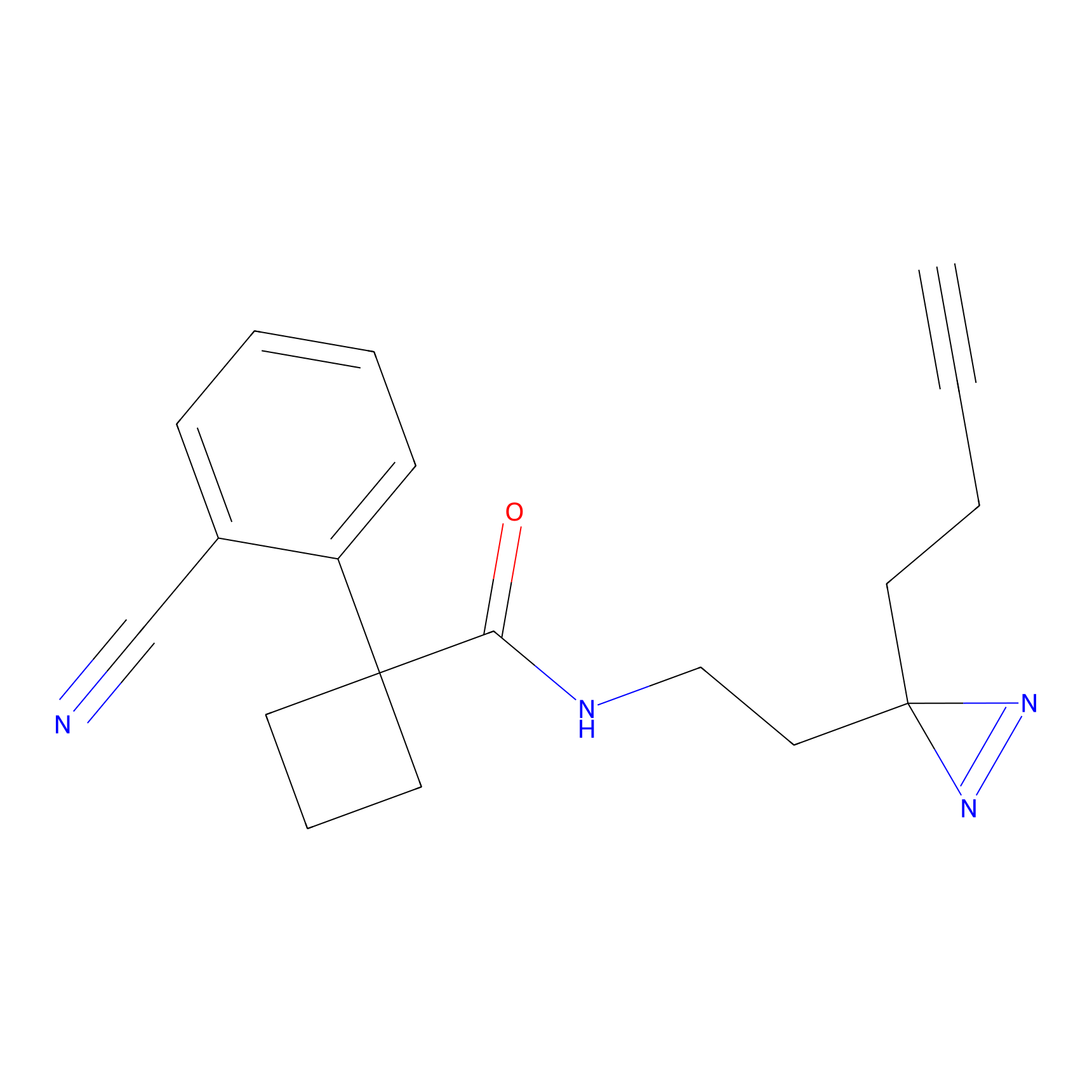
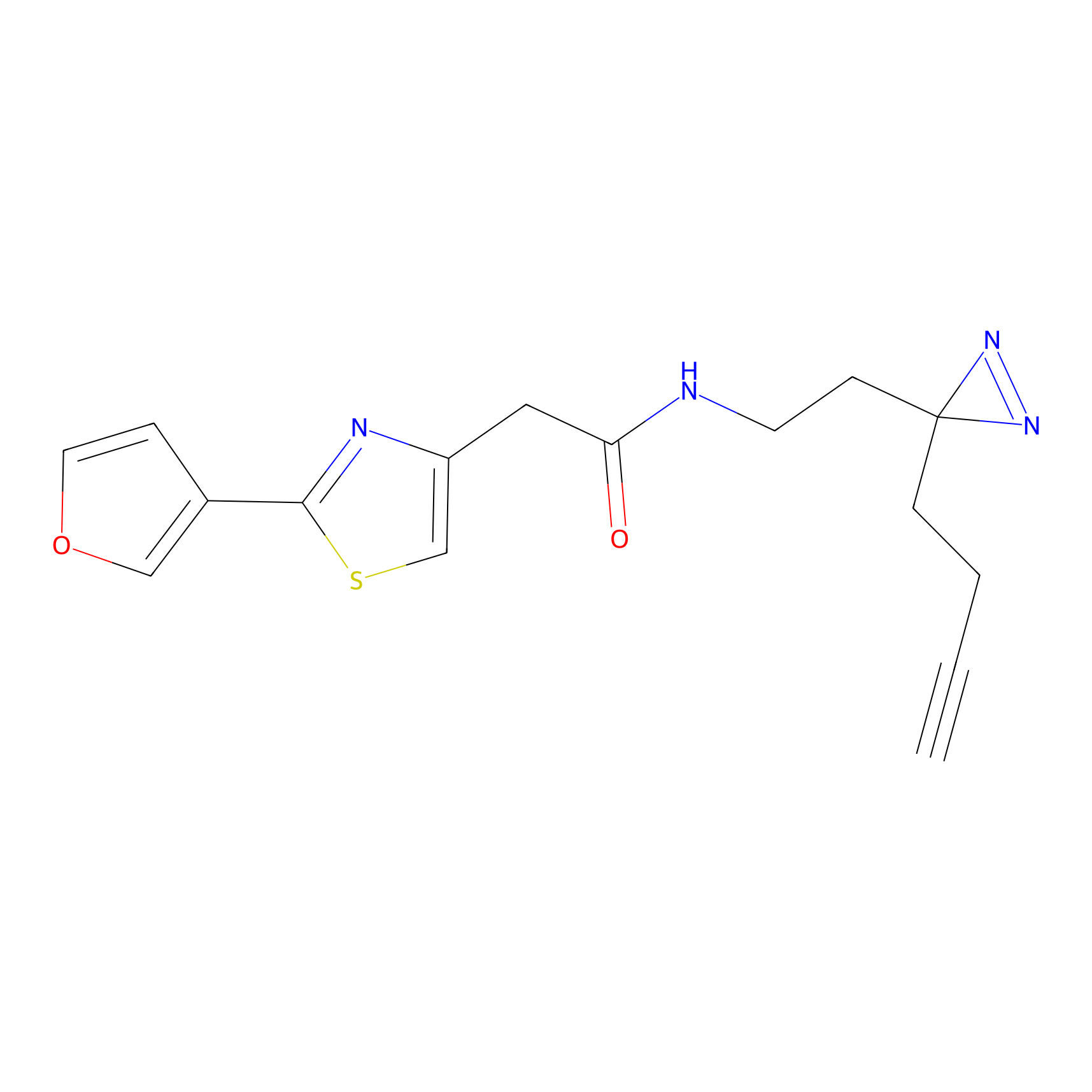
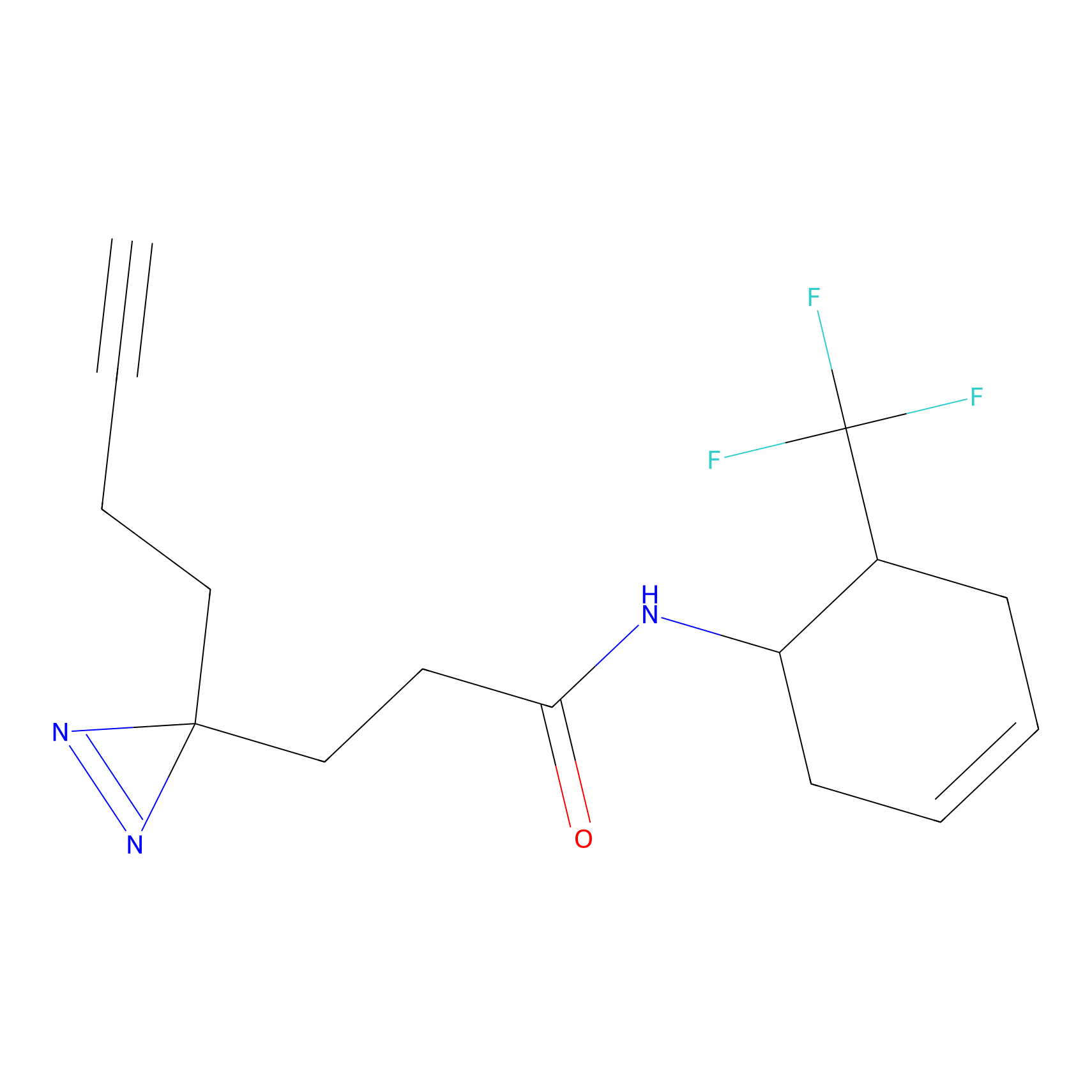
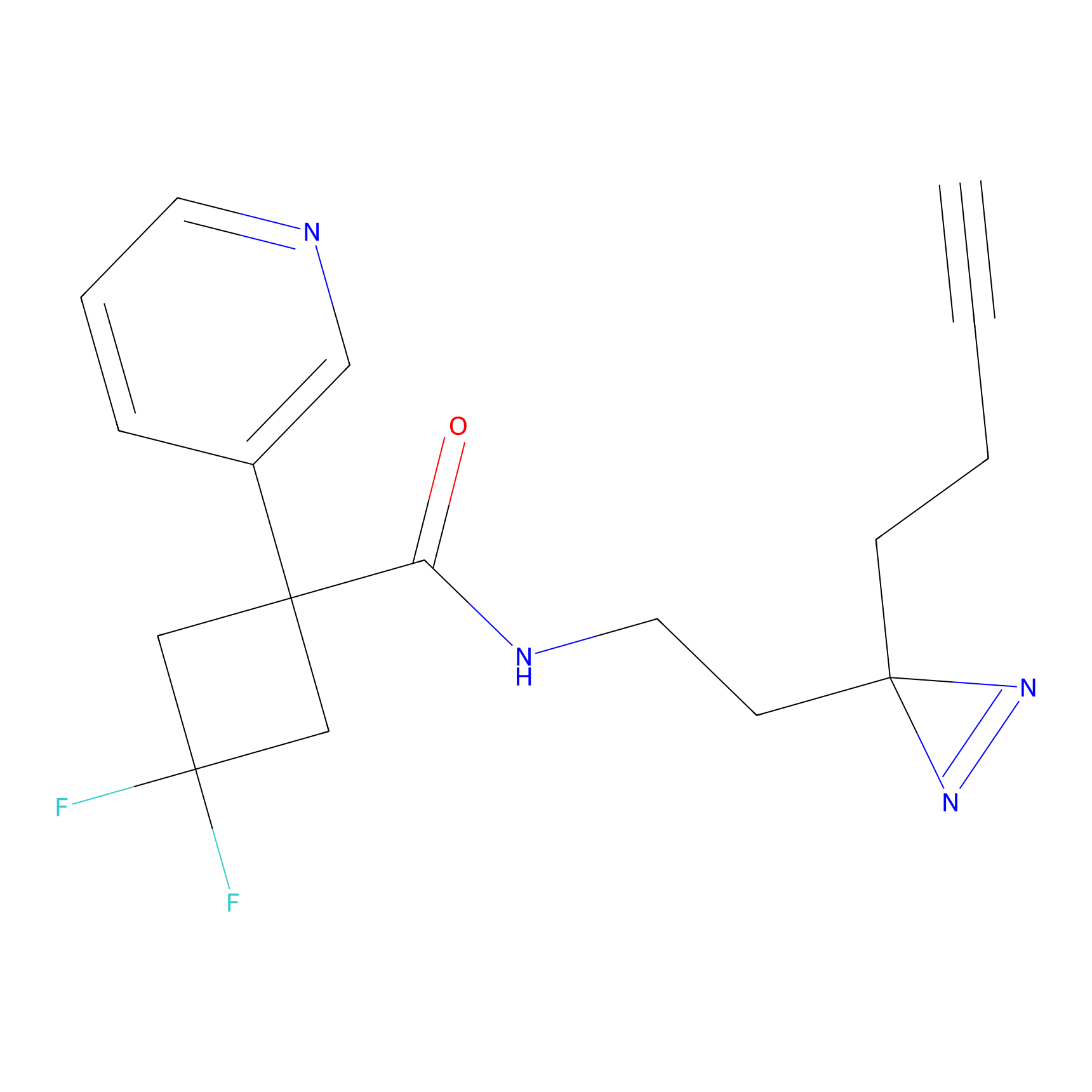
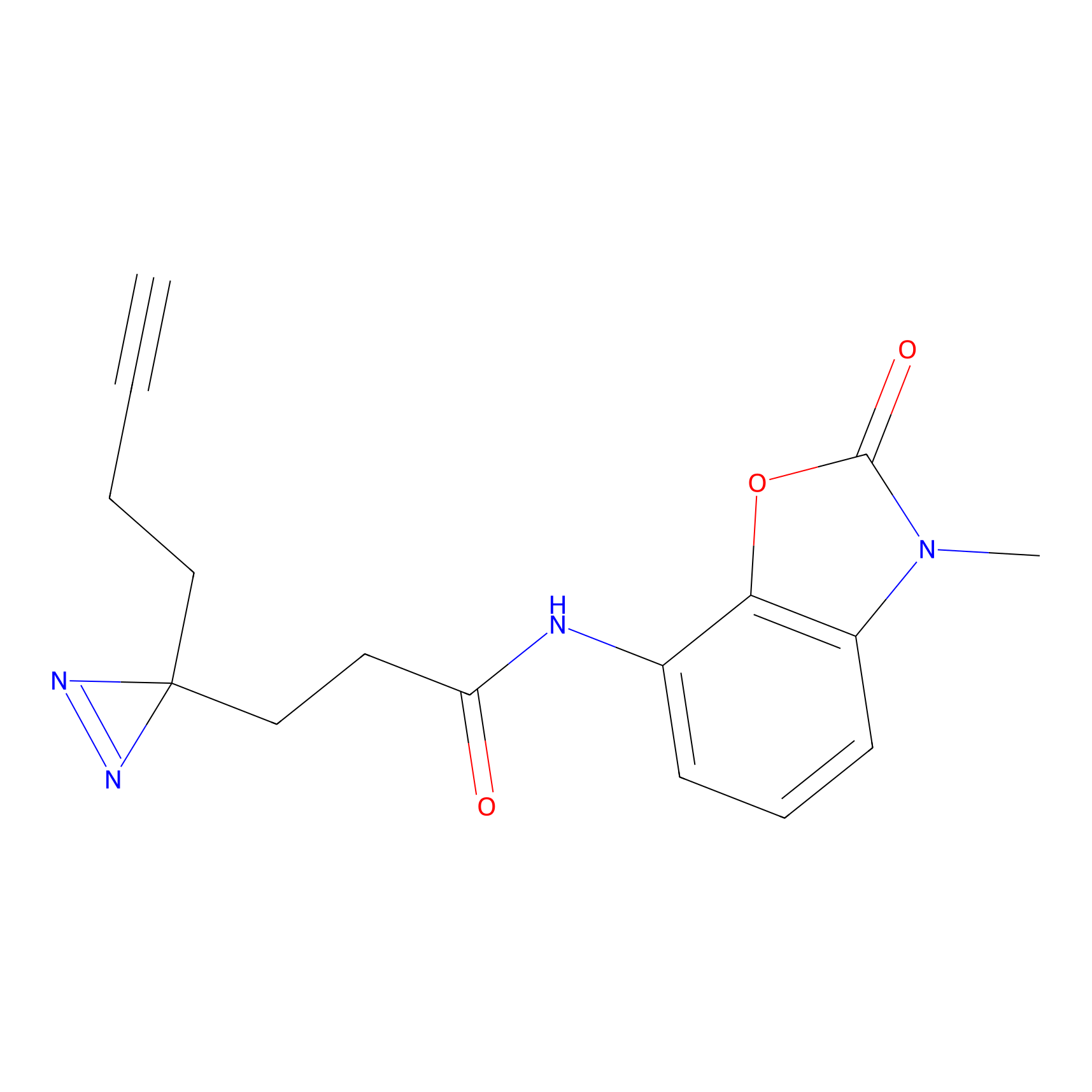
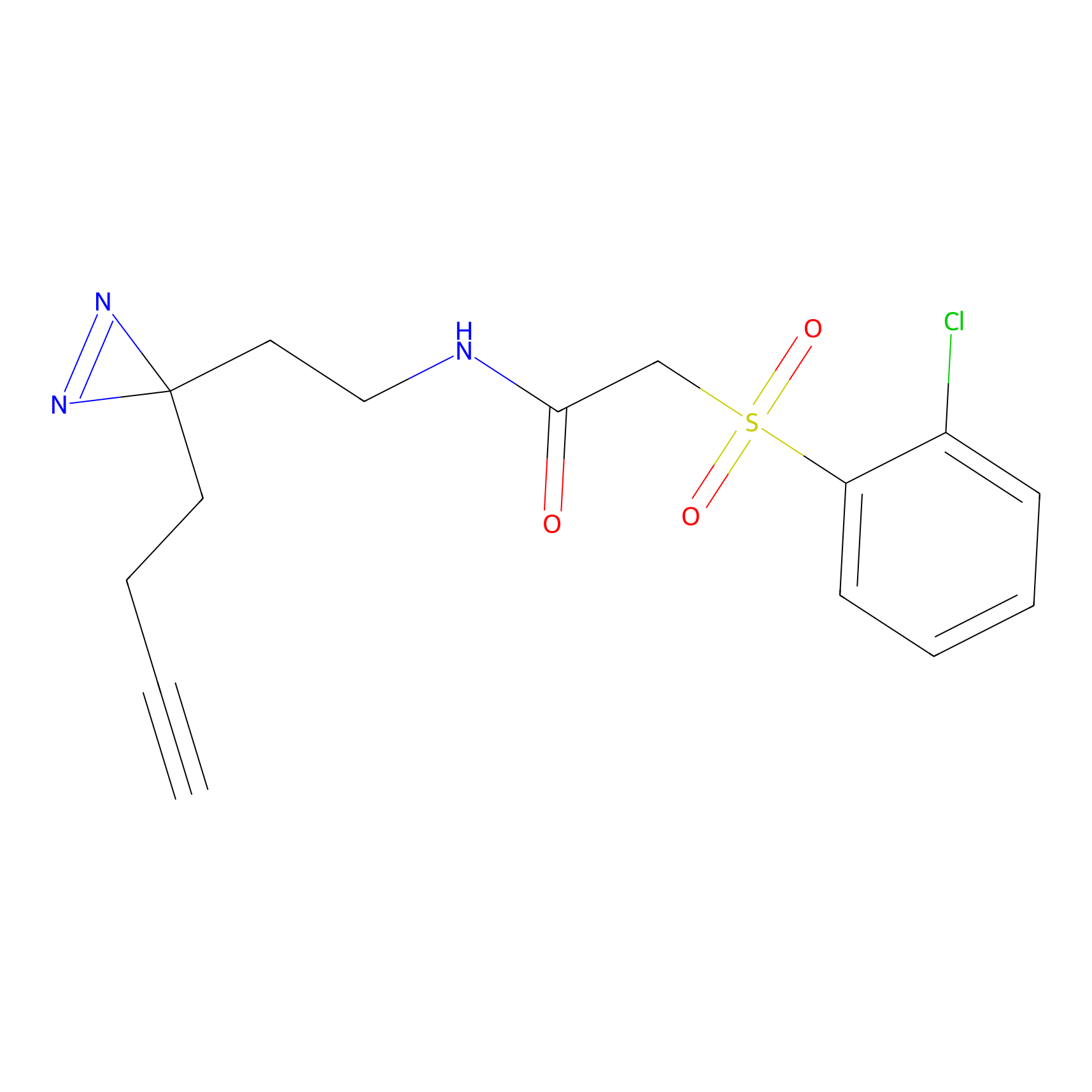
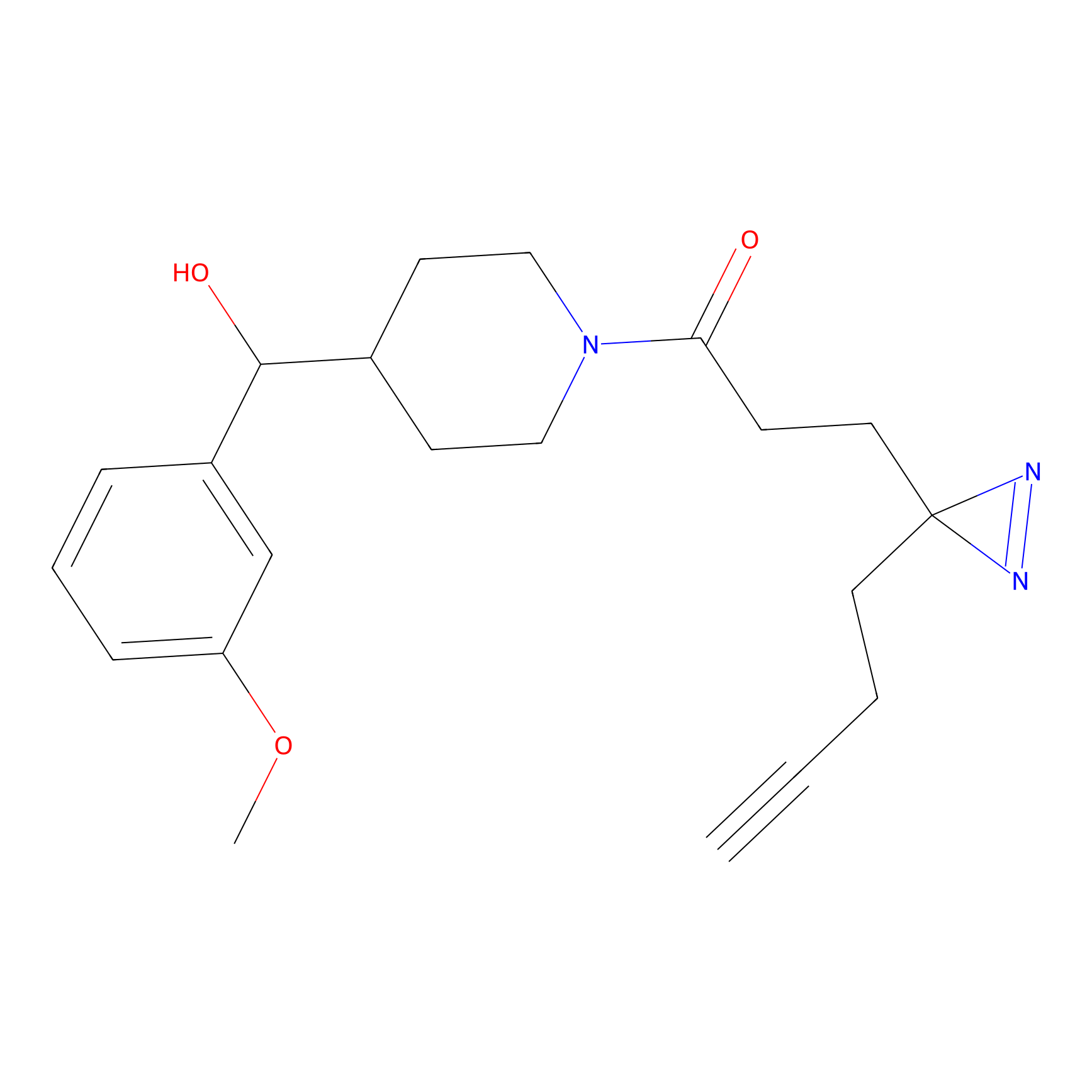
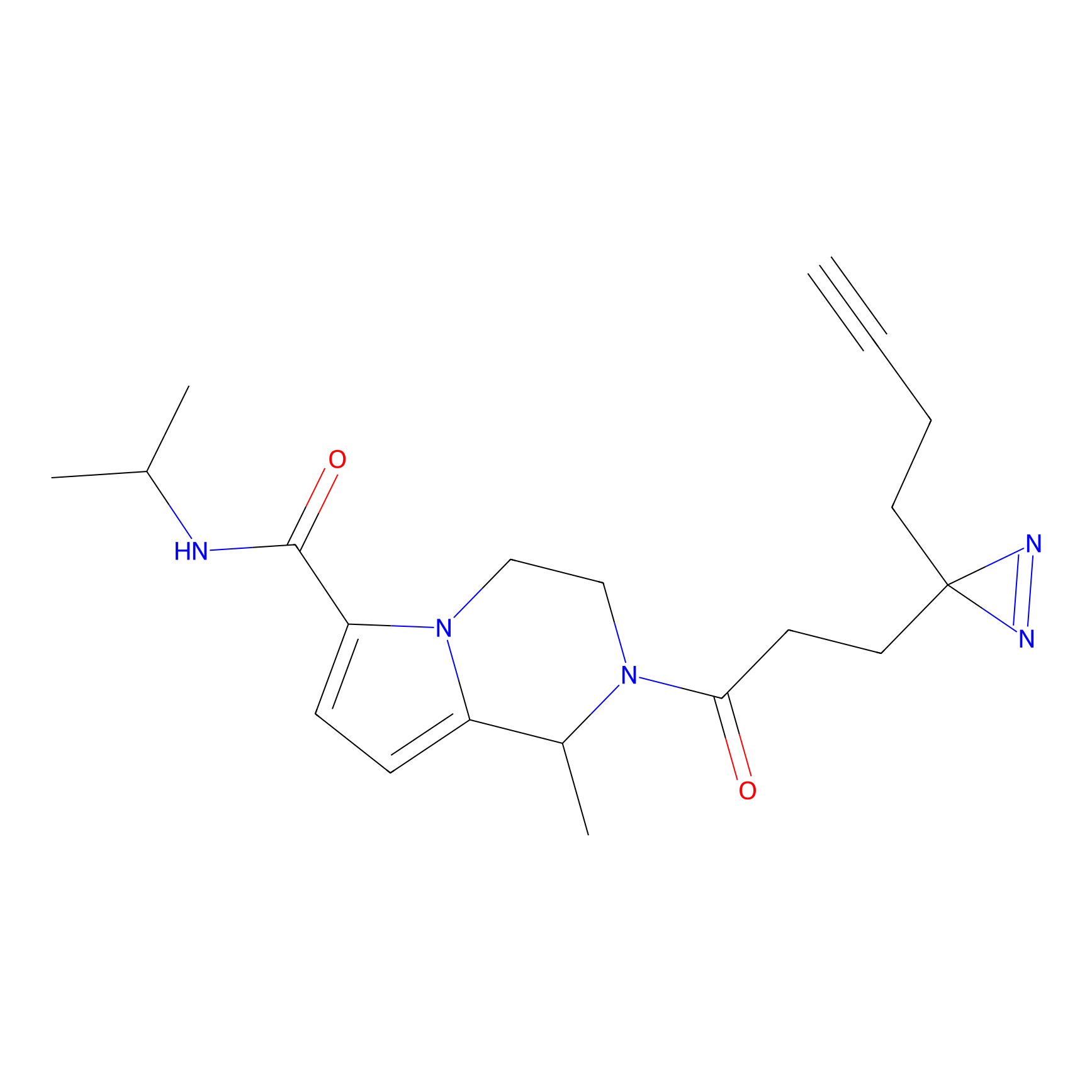
DBIA Probe Info |
 |
C355(3.37) | LDD3310 | [3] | |
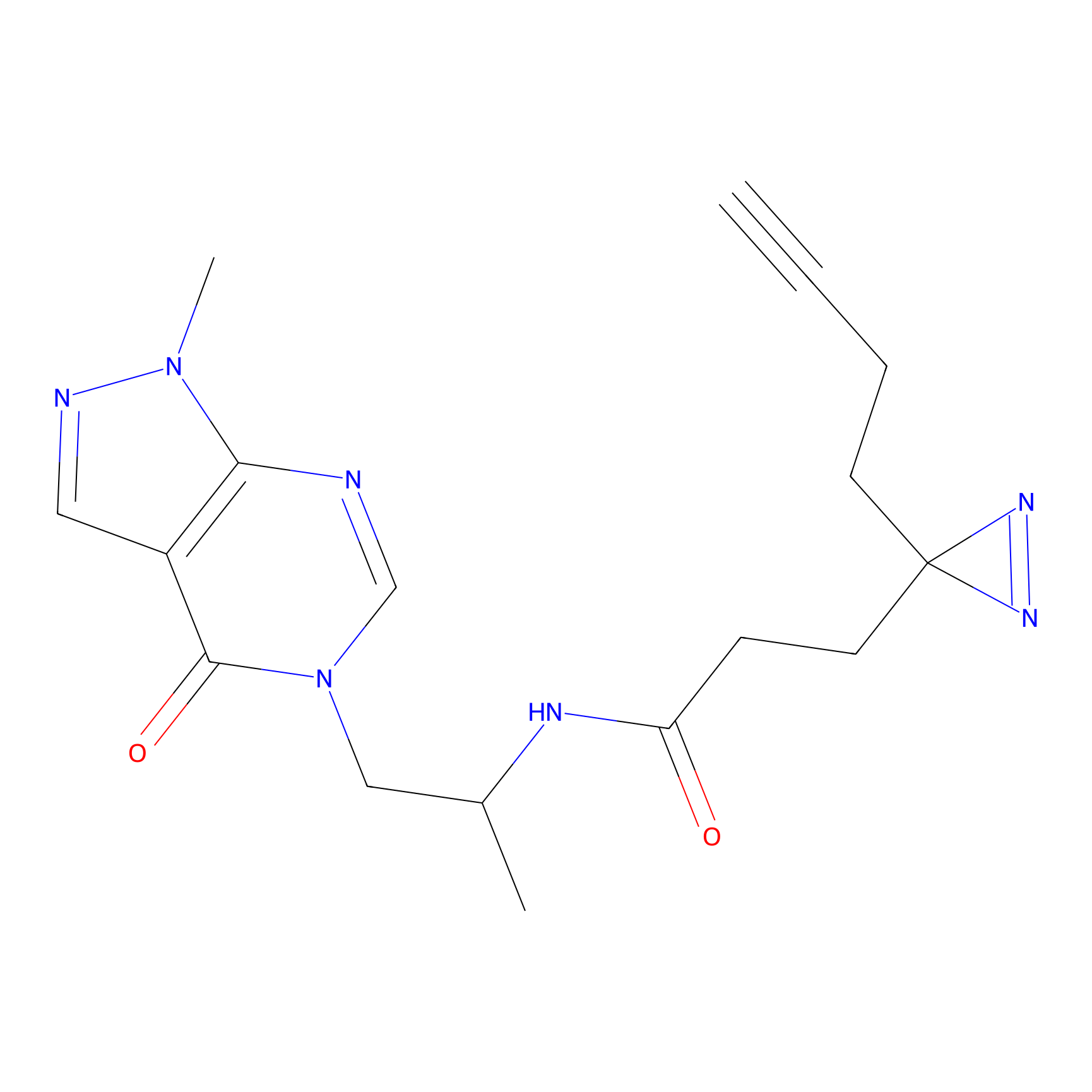
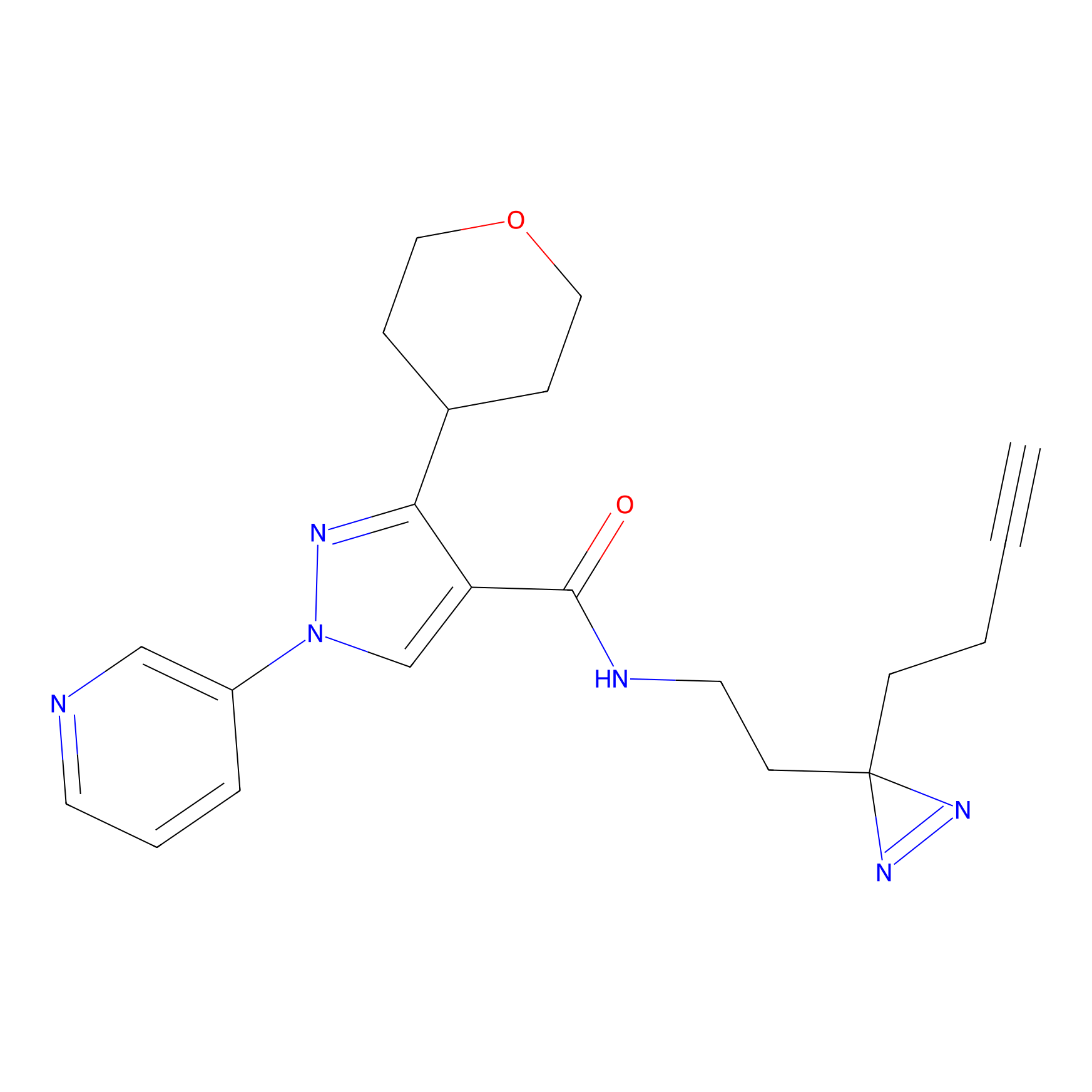
|
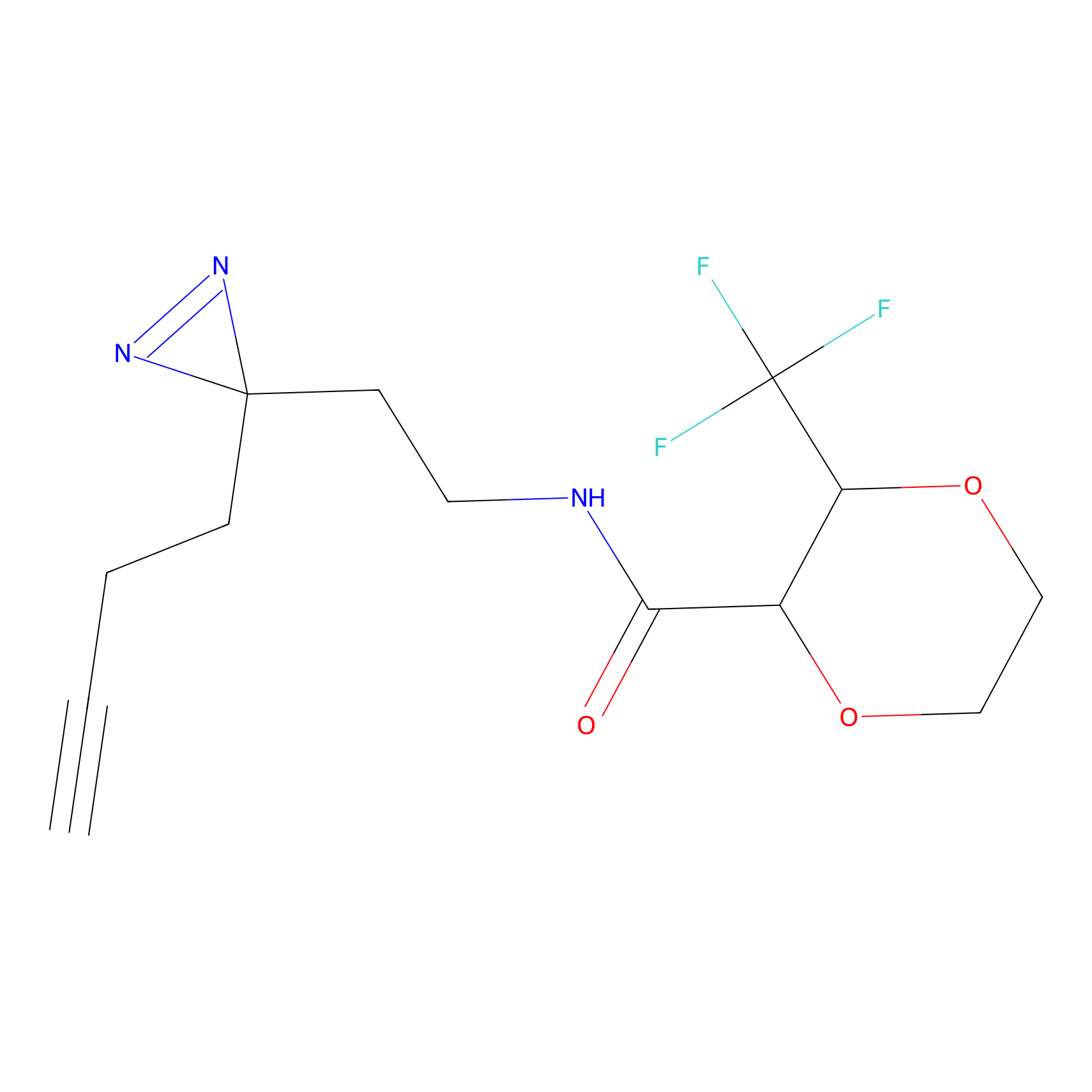
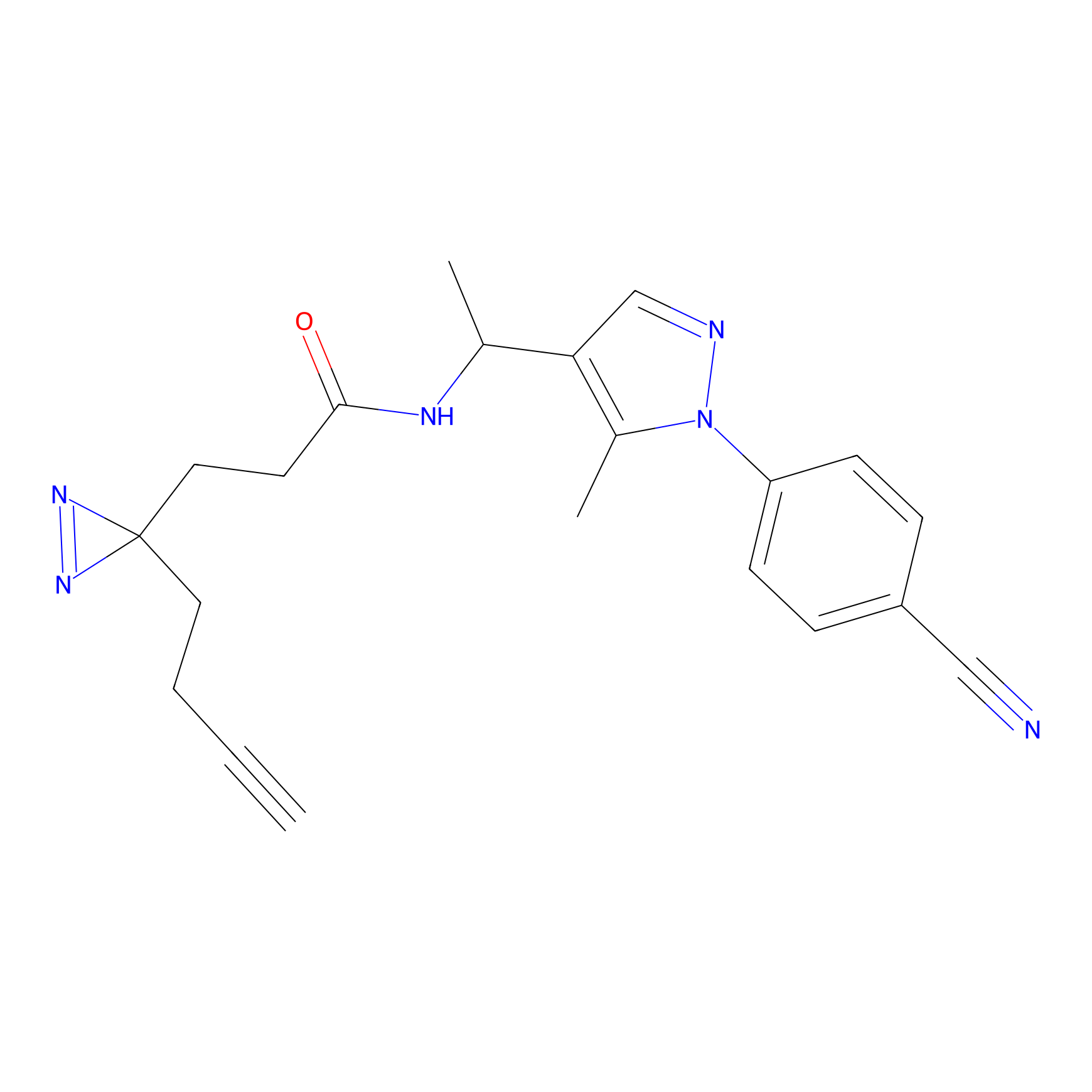
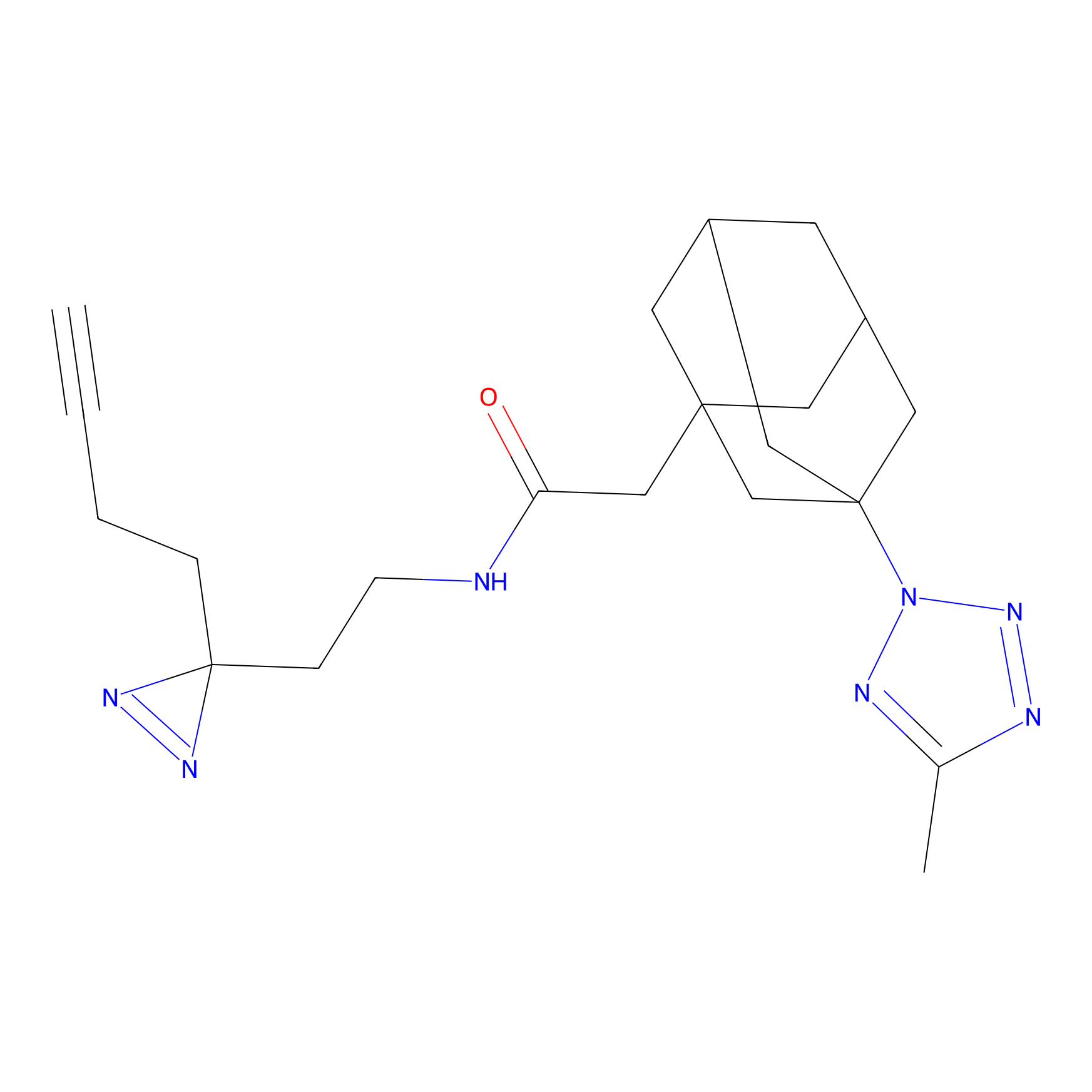
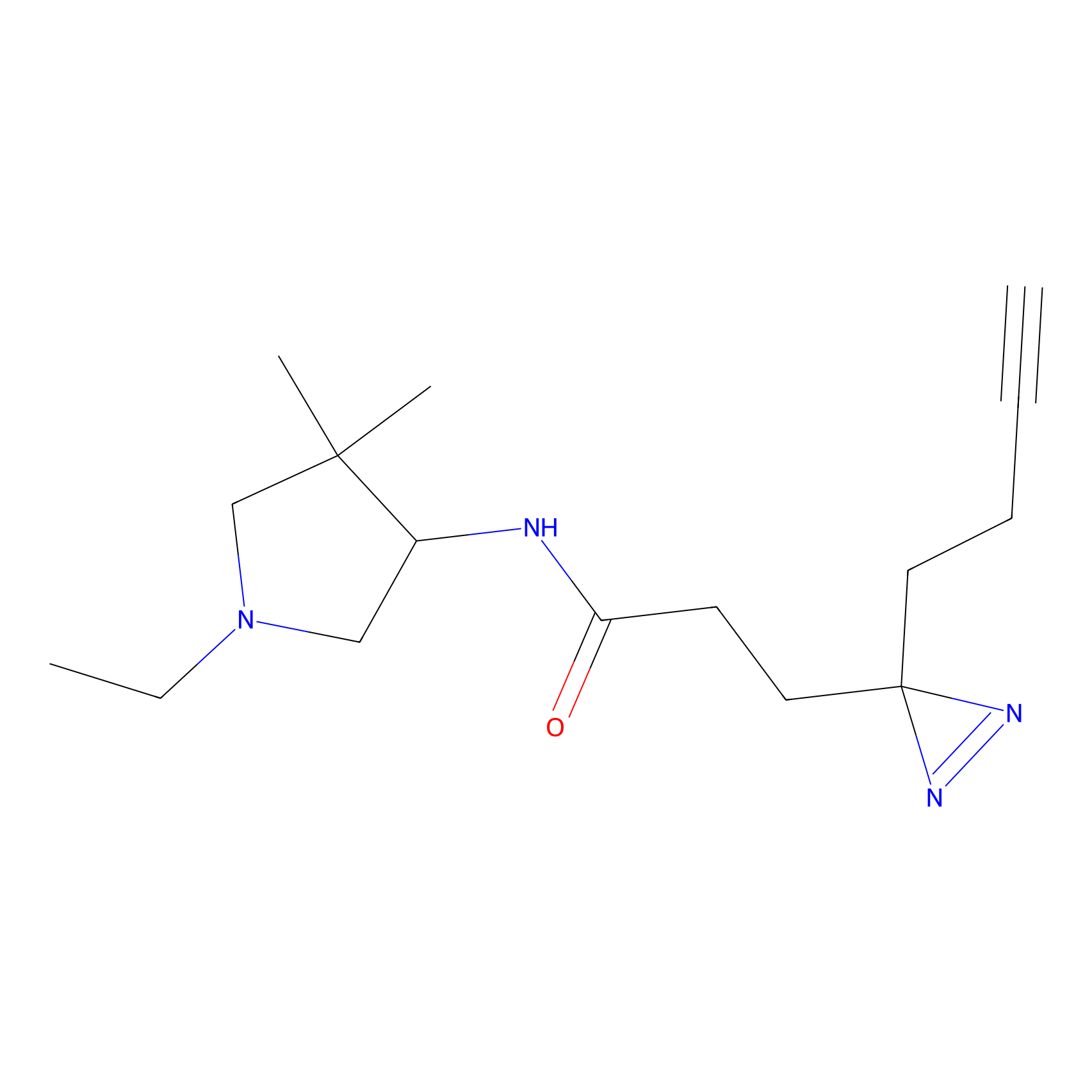
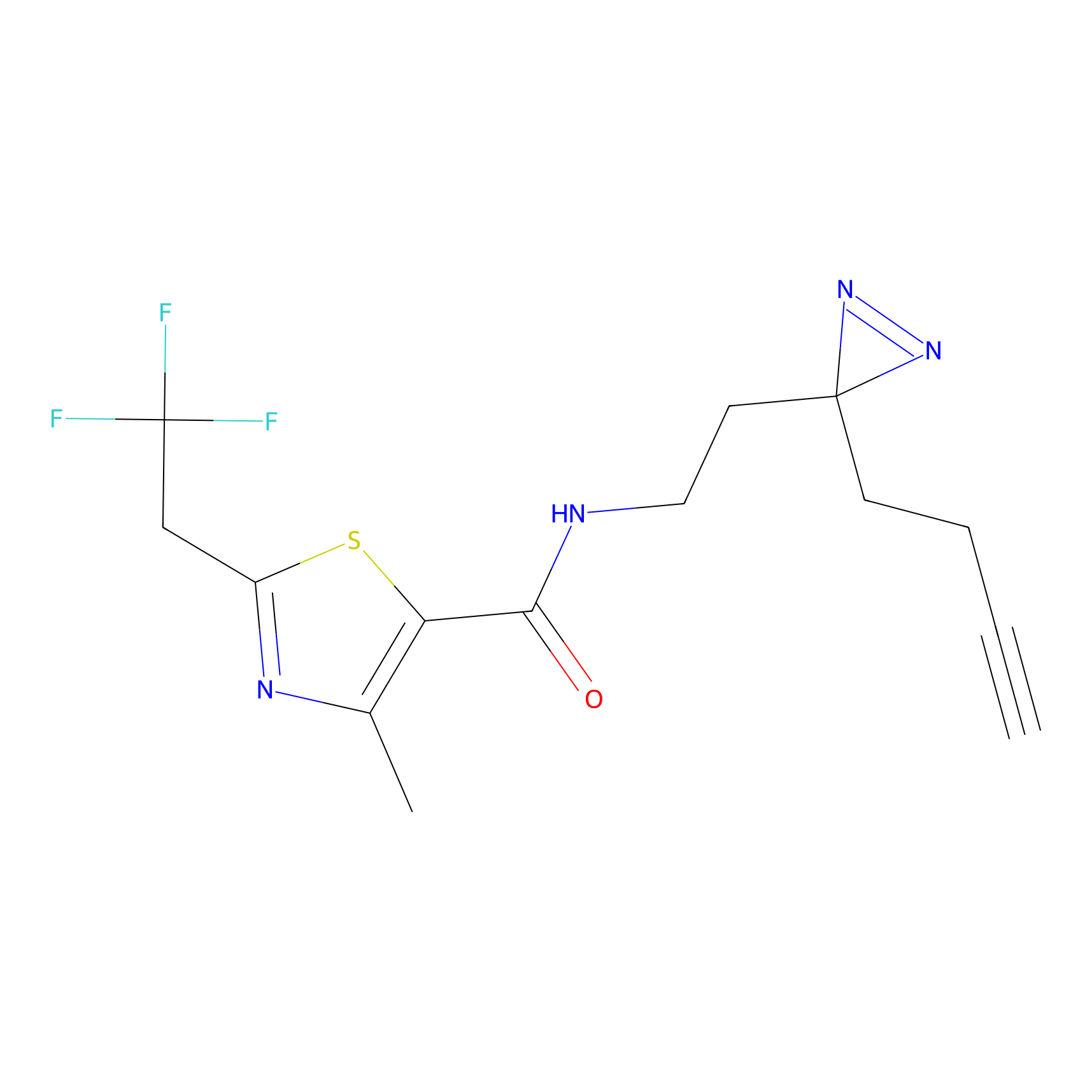
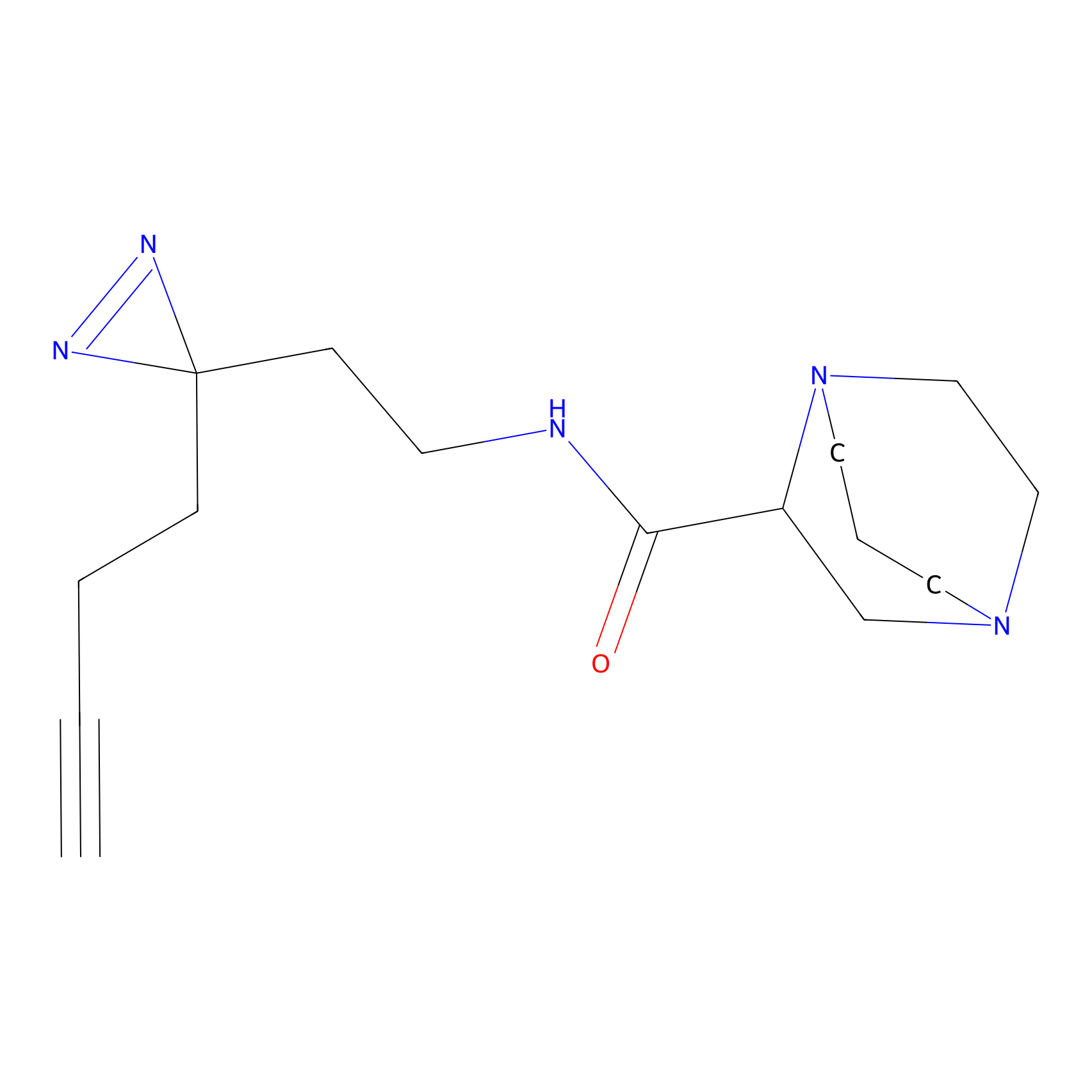
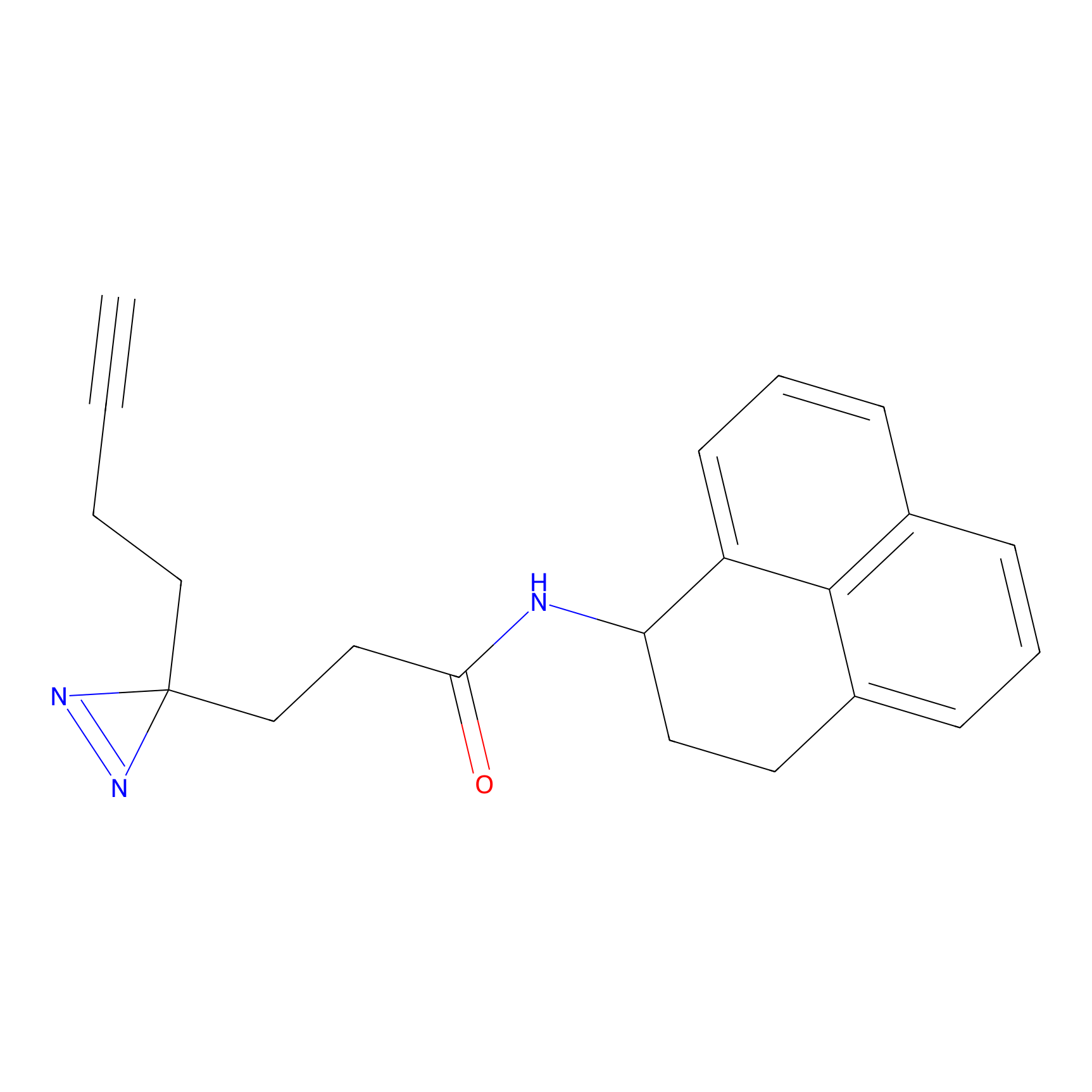
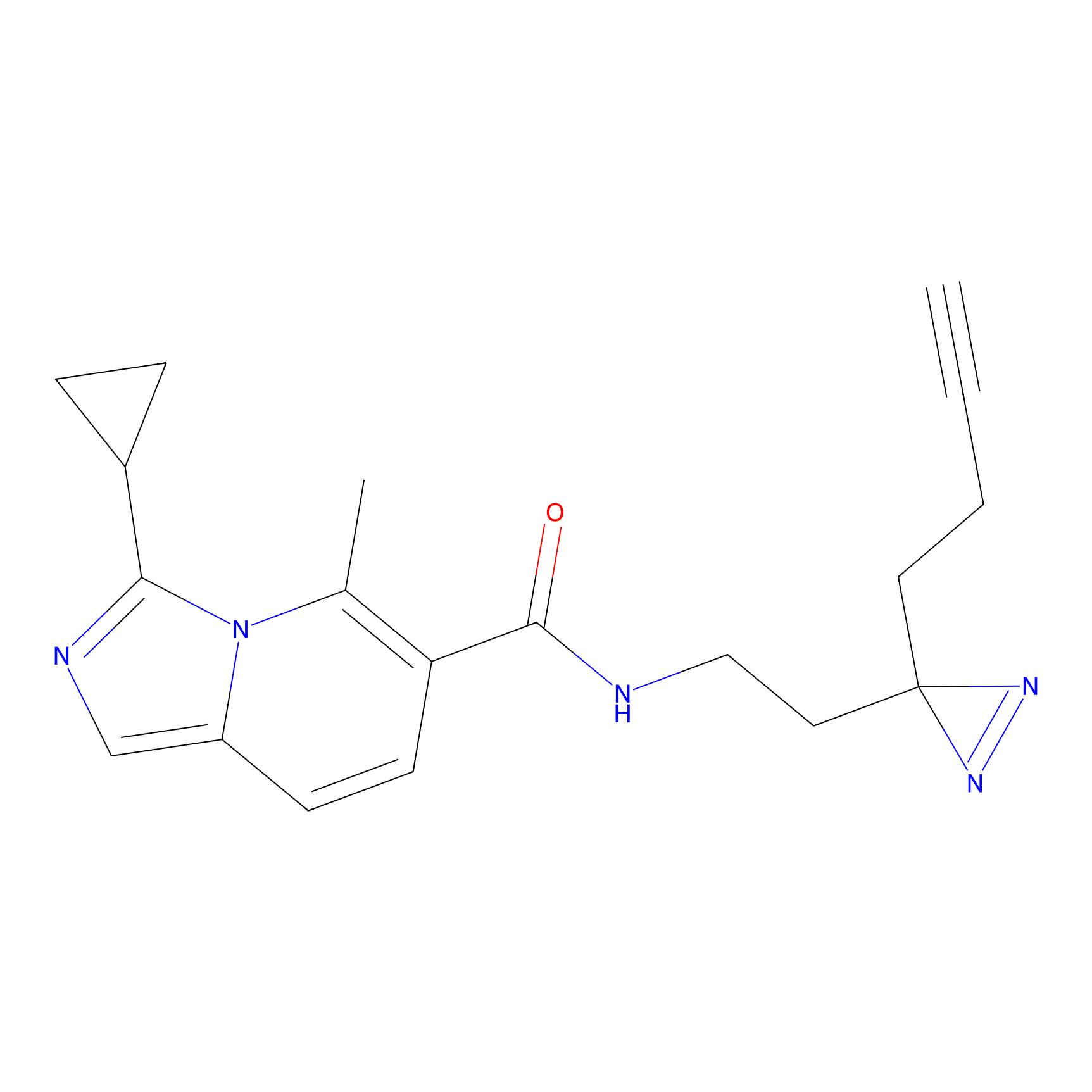
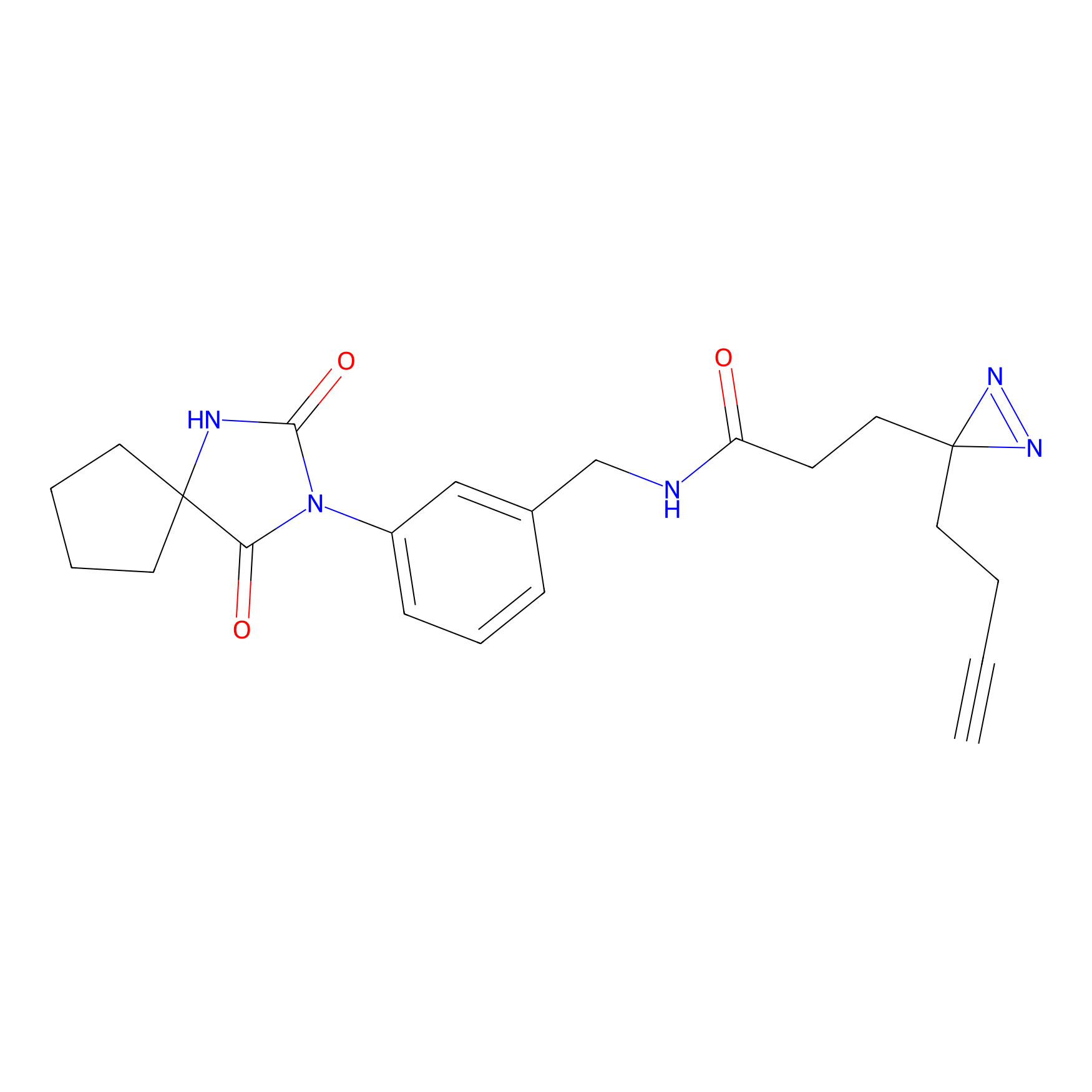
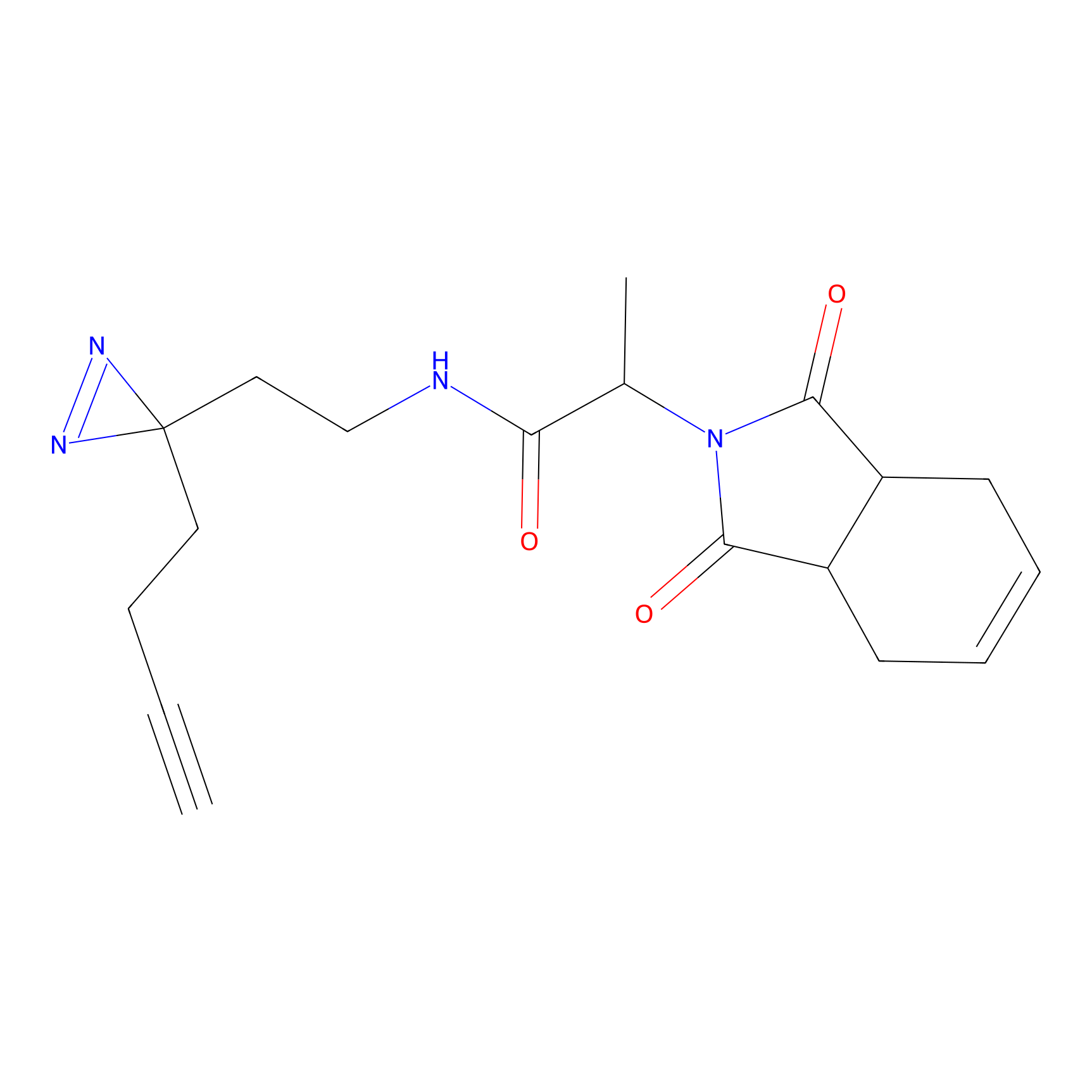
FBPP2 Probe Info |
 |
2.80 | LDD0054 | [4] | |
|
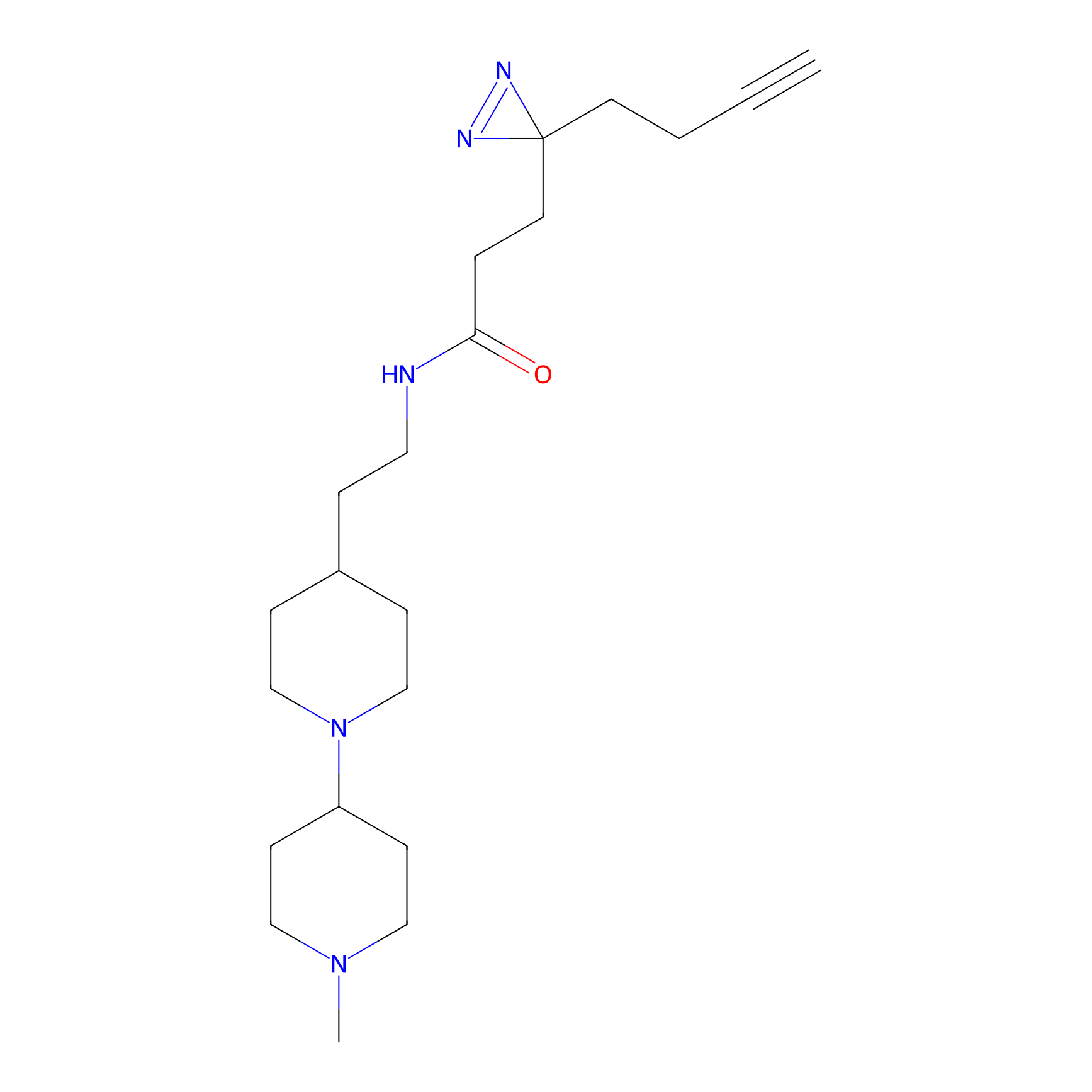
4-Iodoacetamidophenylacetylene Probe Info |
 |
C355(0.00); C371(0.00) | LDD0038 | [5] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [5] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [5] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [6] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C355(0.99) | LDD1507 | [9] |
| LDCM0215 | AC10 | HEK-293T | C355(0.97) | LDD1508 | [9] |
| LDCM0226 | AC11 | HEK-293T | C355(1.05) | LDD1509 | [9] |
| LDCM0237 | AC12 | HEK-293T | C355(1.03) | LDD1510 | [9] |
| LDCM0259 | AC14 | HEK-293T | C355(1.08) | LDD1512 | [9] |
| LDCM0270 | AC15 | HEK-293T | C355(0.98) | LDD1513 | [9] |
| LDCM0276 | AC17 | HEK-293T | C355(0.96) | LDD1515 | [9] |
| LDCM0277 | AC18 | HEK-293T | C355(1.00) | LDD1516 | [9] |
| LDCM0278 | AC19 | HEK-293T | C355(0.92) | LDD1517 | [9] |
| LDCM0279 | AC2 | HEK-293T | C355(1.03) | LDD1518 | [9] |
| LDCM0280 | AC20 | HEK-293T | C355(1.03) | LDD1519 | [9] |
| LDCM0281 | AC21 | HEK-293T | C355(1.05) | LDD1520 | [9] |
| LDCM0282 | AC22 | HEK-293T | C355(1.04) | LDD1521 | [9] |
| LDCM0283 | AC23 | HEK-293T | C355(1.04) | LDD1522 | [9] |
| LDCM0284 | AC24 | HEK-293T | C355(0.95) | LDD1523 | [9] |
| LDCM0285 | AC25 | HEK-293T | C355(1.01) | LDD1524 | [9] |
| LDCM0286 | AC26 | HEK-293T | C355(0.95) | LDD1525 | [9] |
| LDCM0287 | AC27 | HEK-293T | C355(1.03) | LDD1526 | [9] |
| LDCM0288 | AC28 | HEK-293T | C355(1.04) | LDD1527 | [9] |
| LDCM0289 | AC29 | HEK-293T | C355(1.02) | LDD1528 | [9] |
| LDCM0290 | AC3 | HEK-293T | C355(1.11) | LDD1529 | [9] |
| LDCM0291 | AC30 | HEK-293T | C355(1.04) | LDD1530 | [9] |
| LDCM0292 | AC31 | HEK-293T | C355(1.03) | LDD1531 | [9] |
| LDCM0293 | AC32 | HEK-293T | C355(1.01) | LDD1532 | [9] |
| LDCM0294 | AC33 | HEK-293T | C355(0.99) | LDD1533 | [9] |
| LDCM0295 | AC34 | HEK-293T | C355(0.98) | LDD1534 | [9] |
| LDCM0296 | AC35 | HEK-293T | C355(1.08) | LDD1535 | [9] |
| LDCM0297 | AC36 | HEK-293T | C355(1.05) | LDD1536 | [9] |
| LDCM0298 | AC37 | HEK-293T | C355(1.00) | LDD1537 | [9] |
| LDCM0299 | AC38 | HEK-293T | C355(1.03) | LDD1538 | [9] |
| LDCM0300 | AC39 | HEK-293T | C355(1.02) | LDD1539 | [9] |
| LDCM0301 | AC4 | HEK-293T | C355(1.09) | LDD1540 | [9] |
| LDCM0302 | AC40 | HEK-293T | C355(0.96) | LDD1541 | [9] |
| LDCM0303 | AC41 | HEK-293T | C355(1.04) | LDD1542 | [9] |
| LDCM0304 | AC42 | HEK-293T | C355(1.10) | LDD1543 | [9] |
| LDCM0305 | AC43 | HEK-293T | C355(1.03) | LDD1544 | [9] |
| LDCM0306 | AC44 | HEK-293T | C355(1.02) | LDD1545 | [9] |
| LDCM0307 | AC45 | HEK-293T | C355(0.99) | LDD1546 | [9] |
| LDCM0308 | AC46 | HEK-293T | C355(1.06) | LDD1547 | [9] |
| LDCM0309 | AC47 | HEK-293T | C355(1.09) | LDD1548 | [9] |
| LDCM0310 | AC48 | HEK-293T | C355(1.04) | LDD1549 | [9] |
| LDCM0311 | AC49 | HEK-293T | C355(0.97) | LDD1550 | [9] |
| LDCM0312 | AC5 | HEK-293T | C355(1.05) | LDD1551 | [9] |
| LDCM0313 | AC50 | HEK-293T | C355(0.98) | LDD1552 | [9] |
| LDCM0314 | AC51 | HEK-293T | C355(1.05) | LDD1553 | [9] |
| LDCM0315 | AC52 | HEK-293T | C355(1.08) | LDD1554 | [9] |
| LDCM0316 | AC53 | HEK-293T | C355(1.02) | LDD1555 | [9] |
| LDCM0317 | AC54 | HEK-293T | C355(1.10) | LDD1556 | [9] |
| LDCM0318 | AC55 | HEK-293T | C355(1.05) | LDD1557 | [9] |
| LDCM0319 | AC56 | HEK-293T | C355(1.00) | LDD1558 | [9] |
| LDCM0320 | AC57 | HEK-293T | C355(1.07) | LDD1559 | [9] |
| LDCM0321 | AC58 | HEK-293T | C355(0.96) | LDD1560 | [9] |
| LDCM0322 | AC59 | HEK-293T | C355(1.01) | LDD1561 | [9] |
| LDCM0323 | AC6 | HEK-293T | C355(1.07) | LDD1562 | [9] |
| LDCM0324 | AC60 | HEK-293T | C355(1.10) | LDD1563 | [9] |
| LDCM0325 | AC61 | HEK-293T | C355(1.09) | LDD1564 | [9] |
| LDCM0326 | AC62 | HEK-293T | C355(1.09) | LDD1565 | [9] |
| LDCM0327 | AC63 | HEK-293T | C355(1.04) | LDD1566 | [9] |
| LDCM0328 | AC64 | HEK-293T | C355(0.98) | LDD1567 | [9] |
| LDCM0334 | AC7 | HEK-293T | C355(1.16) | LDD1568 | [9] |
| LDCM0345 | AC8 | HEK-293T | C355(1.03) | LDD1569 | [9] |
| LDCM0248 | AKOS034007472 | HEK-293T | C355(1.00) | LDD1511 | [9] |
| LDCM0356 | AKOS034007680 | HEK-293T | C355(1.12) | LDD1570 | [9] |
| LDCM0275 | AKOS034007705 | HEK-293T | C355(0.97) | LDD1514 | [9] |
| LDCM0367 | CL1 | HEK-293T | C355(0.98) | LDD1571 | [9] |
| LDCM0368 | CL10 | HEK-293T | C355(0.92) | LDD1572 | [9] |
| LDCM0369 | CL100 | HEK-293T | C355(0.95) | LDD1573 | [9] |
| LDCM0370 | CL101 | HEK-293T | C355(1.07) | LDD1574 | [9] |
| LDCM0371 | CL102 | HEK-293T | C355(0.97) | LDD1575 | [9] |
| LDCM0372 | CL103 | HEK-293T | C355(0.93) | LDD1576 | [9] |
| LDCM0373 | CL104 | HEK-293T | C355(0.99) | LDD1577 | [9] |
| LDCM0374 | CL105 | HEK-293T | C355(1.05) | LDD1578 | [9] |
| LDCM0375 | CL106 | HEK-293T | C355(0.95) | LDD1579 | [9] |
| LDCM0376 | CL107 | HEK-293T | C355(0.97) | LDD1580 | [9] |
| LDCM0377 | CL108 | HEK-293T | C355(0.99) | LDD1581 | [9] |
| LDCM0378 | CL109 | HEK-293T | C355(0.92) | LDD1582 | [9] |
| LDCM0379 | CL11 | HEK-293T | C355(1.06) | LDD1583 | [9] |
| LDCM0380 | CL110 | HEK-293T | C355(0.97) | LDD1584 | [9] |
| LDCM0381 | CL111 | HEK-293T | C355(1.03) | LDD1585 | [9] |
| LDCM0382 | CL112 | HEK-293T | C355(0.92) | LDD1586 | [9] |
| LDCM0383 | CL113 | HEK-293T | C355(0.98) | LDD1587 | [9] |
| LDCM0384 | CL114 | HEK-293T | C355(0.97) | LDD1588 | [9] |
| LDCM0385 | CL115 | HEK-293T | C355(0.96) | LDD1589 | [9] |
| LDCM0386 | CL116 | HEK-293T | C355(0.95) | LDD1590 | [9] |
| LDCM0387 | CL117 | HEK-293T | C355(0.97) | LDD1591 | [9] |
| LDCM0388 | CL118 | HEK-293T | C355(1.04) | LDD1592 | [9] |
| LDCM0389 | CL119 | HEK-293T | C355(1.03) | LDD1593 | [9] |
| LDCM0390 | CL12 | HEK-293T | C355(1.06) | LDD1594 | [9] |
| LDCM0391 | CL120 | HEK-293T | C355(1.06) | LDD1595 | [9] |
| LDCM0392 | CL121 | HEK-293T | C355(1.04) | LDD1596 | [9] |
| LDCM0393 | CL122 | HEK-293T | C355(0.97) | LDD1597 | [9] |
| LDCM0394 | CL123 | HEK-293T | C355(0.91) | LDD1598 | [9] |
| LDCM0395 | CL124 | HEK-293T | C355(1.00) | LDD1599 | [9] |
| LDCM0396 | CL125 | HEK-293T | C355(1.00) | LDD1600 | [9] |
| LDCM0397 | CL126 | HEK-293T | C355(1.07) | LDD1601 | [9] |
| LDCM0398 | CL127 | HEK-293T | C355(1.03) | LDD1602 | [9] |
| LDCM0399 | CL128 | HEK-293T | C355(0.97) | LDD1603 | [9] |
| LDCM0400 | CL13 | HEK-293T | C355(1.05) | LDD1604 | [9] |
| LDCM0401 | CL14 | HEK-293T | C355(1.00) | LDD1605 | [9] |
| LDCM0402 | CL15 | HEK-293T | C355(0.92) | LDD1606 | [9] |
| LDCM0403 | CL16 | HEK-293T | C355(1.04) | LDD1607 | [9] |
| LDCM0404 | CL17 | HEK-293T | C355(0.89) | LDD1608 | [9] |
| LDCM0405 | CL18 | HEK-293T | C355(1.03) | LDD1609 | [9] |
| LDCM0406 | CL19 | HEK-293T | C355(1.05) | LDD1610 | [9] |
| LDCM0407 | CL2 | HEK-293T | C355(1.06) | LDD1611 | [9] |
| LDCM0408 | CL20 | HEK-293T | C355(1.06) | LDD1612 | [9] |
| LDCM0409 | CL21 | HEK-293T | C355(0.99) | LDD1613 | [9] |
| LDCM0410 | CL22 | HEK-293T | C355(1.04) | LDD1614 | [9] |
| LDCM0411 | CL23 | HEK-293T | C355(1.00) | LDD1615 | [9] |
| LDCM0412 | CL24 | HEK-293T | C355(1.02) | LDD1616 | [9] |
| LDCM0413 | CL25 | HEK-293T | C355(0.88) | LDD1617 | [9] |
| LDCM0414 | CL26 | HEK-293T | C355(1.05) | LDD1618 | [9] |
| LDCM0415 | CL27 | HEK-293T | C355(1.01) | LDD1619 | [9] |
| LDCM0416 | CL28 | HEK-293T | C355(0.97) | LDD1620 | [9] |
| LDCM0417 | CL29 | HEK-293T | C355(1.03) | LDD1621 | [9] |
| LDCM0418 | CL3 | HEK-293T | C355(1.03) | LDD1622 | [9] |
| LDCM0419 | CL30 | HEK-293T | C355(1.04) | LDD1623 | [9] |
| LDCM0420 | CL31 | HEK-293T | C355(1.07) | LDD1624 | [9] |
| LDCM0421 | CL32 | HEK-293T | C355(0.95) | LDD1625 | [9] |
| LDCM0422 | CL33 | HEK-293T | C355(0.97) | LDD1626 | [9] |
| LDCM0423 | CL34 | HEK-293T | C355(1.08) | LDD1627 | [9] |
| LDCM0424 | CL35 | HEK-293T | C355(0.99) | LDD1628 | [9] |
| LDCM0425 | CL36 | HEK-293T | C355(1.12) | LDD1629 | [9] |
| LDCM0426 | CL37 | HEK-293T | C355(0.95) | LDD1630 | [9] |
| LDCM0428 | CL39 | HEK-293T | C355(0.90) | LDD1632 | [9] |
| LDCM0429 | CL4 | HEK-293T | C355(0.98) | LDD1633 | [9] |
| LDCM0430 | CL40 | HEK-293T | C355(1.03) | LDD1634 | [9] |
| LDCM0431 | CL41 | HEK-293T | C355(0.99) | LDD1635 | [9] |
| LDCM0432 | CL42 | HEK-293T | C355(1.04) | LDD1636 | [9] |
| LDCM0433 | CL43 | HEK-293T | C355(1.12) | LDD1637 | [9] |
| LDCM0434 | CL44 | HEK-293T | C355(1.00) | LDD1638 | [9] |
| LDCM0435 | CL45 | HEK-293T | C355(0.99) | LDD1639 | [9] |
| LDCM0436 | CL46 | HEK-293T | C355(1.03) | LDD1640 | [9] |
| LDCM0437 | CL47 | HEK-293T | C355(1.00) | LDD1641 | [9] |
| LDCM0438 | CL48 | HEK-293T | C355(1.38) | LDD1642 | [9] |
| LDCM0439 | CL49 | HEK-293T | C355(1.01) | LDD1643 | [9] |
| LDCM0440 | CL5 | HEK-293T | C355(1.06) | LDD1644 | [9] |
| LDCM0441 | CL50 | HEK-293T | C355(1.02) | LDD1645 | [9] |
| LDCM0443 | CL52 | HEK-293T | C355(0.98) | LDD1646 | [9] |
| LDCM0444 | CL53 | HEK-293T | C355(0.93) | LDD1647 | [9] |
| LDCM0445 | CL54 | HEK-293T | C355(0.97) | LDD1648 | [9] |
| LDCM0446 | CL55 | HEK-293T | C355(1.03) | LDD1649 | [9] |
| LDCM0447 | CL56 | HEK-293T | C355(1.07) | LDD1650 | [9] |
| LDCM0448 | CL57 | HEK-293T | C355(0.90) | LDD1651 | [9] |
| LDCM0449 | CL58 | HEK-293T | C355(1.03) | LDD1652 | [9] |
| LDCM0450 | CL59 | HEK-293T | C355(1.08) | LDD1653 | [9] |
| LDCM0451 | CL6 | HEK-293T | C355(0.99) | LDD1654 | [9] |
| LDCM0452 | CL60 | HEK-293T | C355(0.99) | LDD1655 | [9] |
| LDCM0453 | CL61 | HEK-293T | C355(1.00) | LDD1656 | [9] |
| LDCM0454 | CL62 | HEK-293T | C355(1.05) | LDD1657 | [9] |
| LDCM0455 | CL63 | HEK-293T | C355(1.01) | LDD1658 | [9] |
| LDCM0456 | CL64 | HEK-293T | C355(0.88) | LDD1659 | [9] |
| LDCM0457 | CL65 | HEK-293T | C355(1.02) | LDD1660 | [9] |
| LDCM0458 | CL66 | HEK-293T | C355(0.99) | LDD1661 | [9] |
| LDCM0459 | CL67 | HEK-293T | C355(1.04) | LDD1662 | [9] |
| LDCM0460 | CL68 | HEK-293T | C355(0.98) | LDD1663 | [9] |
| LDCM0461 | CL69 | HEK-293T | C355(1.06) | LDD1664 | [9] |
| LDCM0462 | CL7 | HEK-293T | C355(1.06) | LDD1665 | [9] |
| LDCM0463 | CL70 | HEK-293T | C355(1.06) | LDD1666 | [9] |
| LDCM0464 | CL71 | HEK-293T | C355(1.02) | LDD1667 | [9] |
| LDCM0465 | CL72 | HEK-293T | C355(1.14) | LDD1668 | [9] |
| LDCM0466 | CL73 | HEK-293T | C355(1.03) | LDD1669 | [9] |
| LDCM0467 | CL74 | HEK-293T | C355(1.10) | LDD1670 | [9] |
| LDCM0469 | CL76 | HEK-293T | C355(0.96) | LDD1672 | [9] |
| LDCM0470 | CL77 | HEK-293T | C355(0.97) | LDD1673 | [9] |
| LDCM0471 | CL78 | HEK-293T | C355(0.89) | LDD1674 | [9] |
| LDCM0472 | CL79 | HEK-293T | C355(0.97) | LDD1675 | [9] |
| LDCM0473 | CL8 | HEK-293T | C355(0.92) | LDD1676 | [9] |
| LDCM0474 | CL80 | HEK-293T | C355(1.10) | LDD1677 | [9] |
| LDCM0475 | CL81 | HEK-293T | C355(1.09) | LDD1678 | [9] |
| LDCM0476 | CL82 | HEK-293T | C355(1.05) | LDD1679 | [9] |
| LDCM0477 | CL83 | HEK-293T | C355(1.00) | LDD1680 | [9] |
| LDCM0478 | CL84 | HEK-293T | C355(1.02) | LDD1681 | [9] |
| LDCM0479 | CL85 | HEK-293T | C355(1.01) | LDD1682 | [9] |
| LDCM0480 | CL86 | HEK-293T | C355(1.07) | LDD1683 | [9] |
| LDCM0481 | CL87 | HEK-293T | C355(1.00) | LDD1684 | [9] |
| LDCM0482 | CL88 | HEK-293T | C355(0.97) | LDD1685 | [9] |
| LDCM0483 | CL89 | HEK-293T | C355(1.09) | LDD1686 | [9] |
| LDCM0484 | CL9 | HEK-293T | C355(1.05) | LDD1687 | [9] |
| LDCM0485 | CL90 | HEK-293T | C355(0.82) | LDD1688 | [9] |
| LDCM0486 | CL91 | HEK-293T | C355(1.08) | LDD1689 | [9] |
| LDCM0487 | CL92 | HEK-293T | C355(1.09) | LDD1690 | [9] |
| LDCM0488 | CL93 | HEK-293T | C355(1.00) | LDD1691 | [9] |
| LDCM0489 | CL94 | HEK-293T | C355(1.06) | LDD1692 | [9] |
| LDCM0490 | CL95 | HEK-293T | C355(0.95) | LDD1693 | [9] |
| LDCM0491 | CL96 | HEK-293T | C355(1.06) | LDD1694 | [9] |
| LDCM0492 | CL97 | HEK-293T | C355(0.98) | LDD1695 | [9] |
| LDCM0493 | CL98 | HEK-293T | C355(0.98) | LDD1696 | [9] |
| LDCM0494 | CL99 | HEK-293T | C355(0.99) | LDD1697 | [9] |
| LDCM0495 | E2913 | HEK-293T | C355(0.96) | LDD1698 | [9] |
| LDCM0468 | Fragment33 | HEK-293T | C355(0.99) | LDD1671 | [9] |
| LDCM0427 | Fragment51 | HEK-293T | C355(0.95) | LDD1631 | [9] |
| LDCM0022 | KB02 | 22RV1 | C355(1.42) | LDD2243 | [3] |
| LDCM0023 | KB03 | 22RV1 | C355(1.38) | LDD2660 | [3] |
| LDCM0024 | KB05 | COLO792 | C355(3.37) | LDD3310 | [3] |
| LDCM0131 | RA190 | MM1.R | C355(1.45) | LDD0304 | [10] |
| LDCM0008 | Tranylcypromine | SH-SY5Y | 2.80 | LDD0054 | [4] |
References