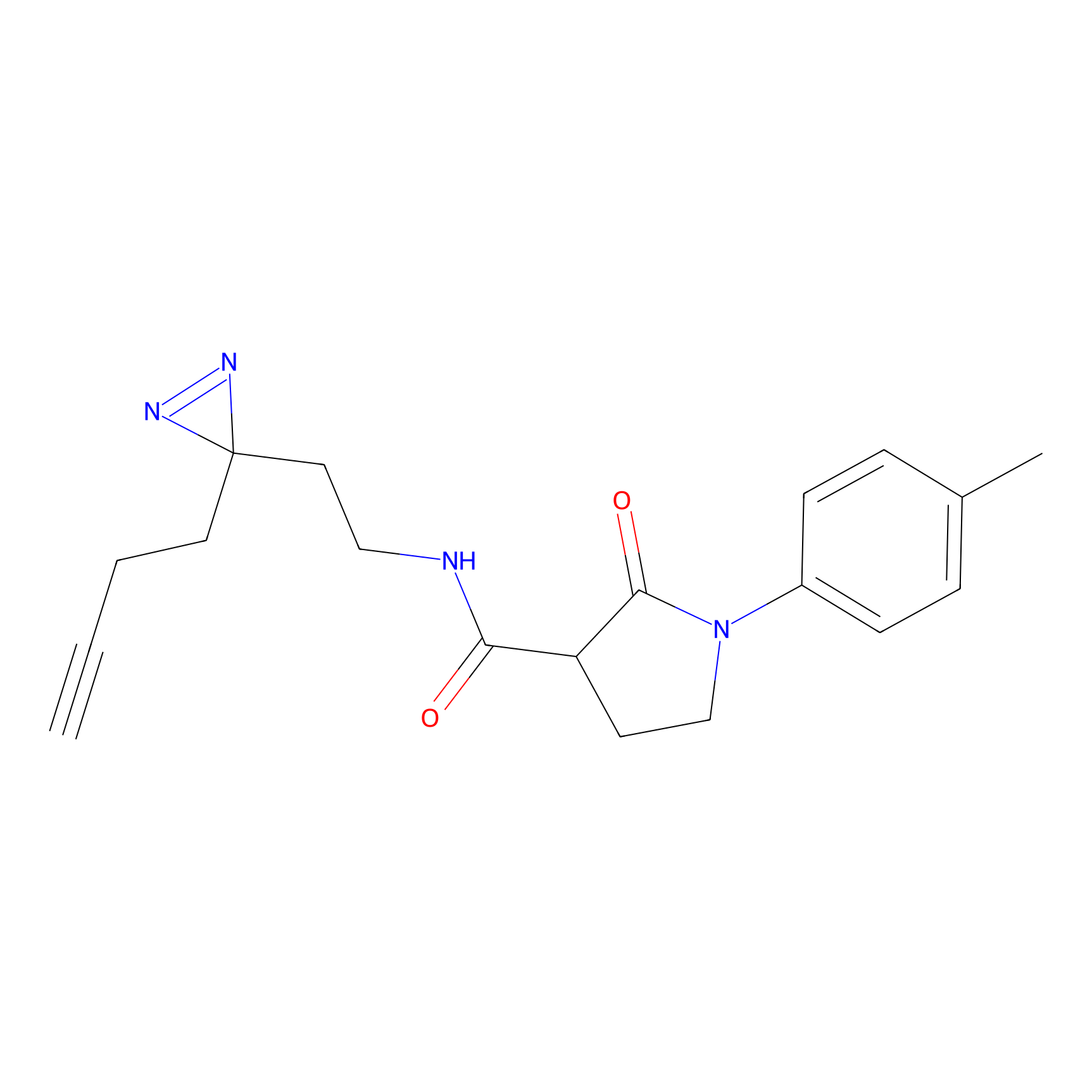
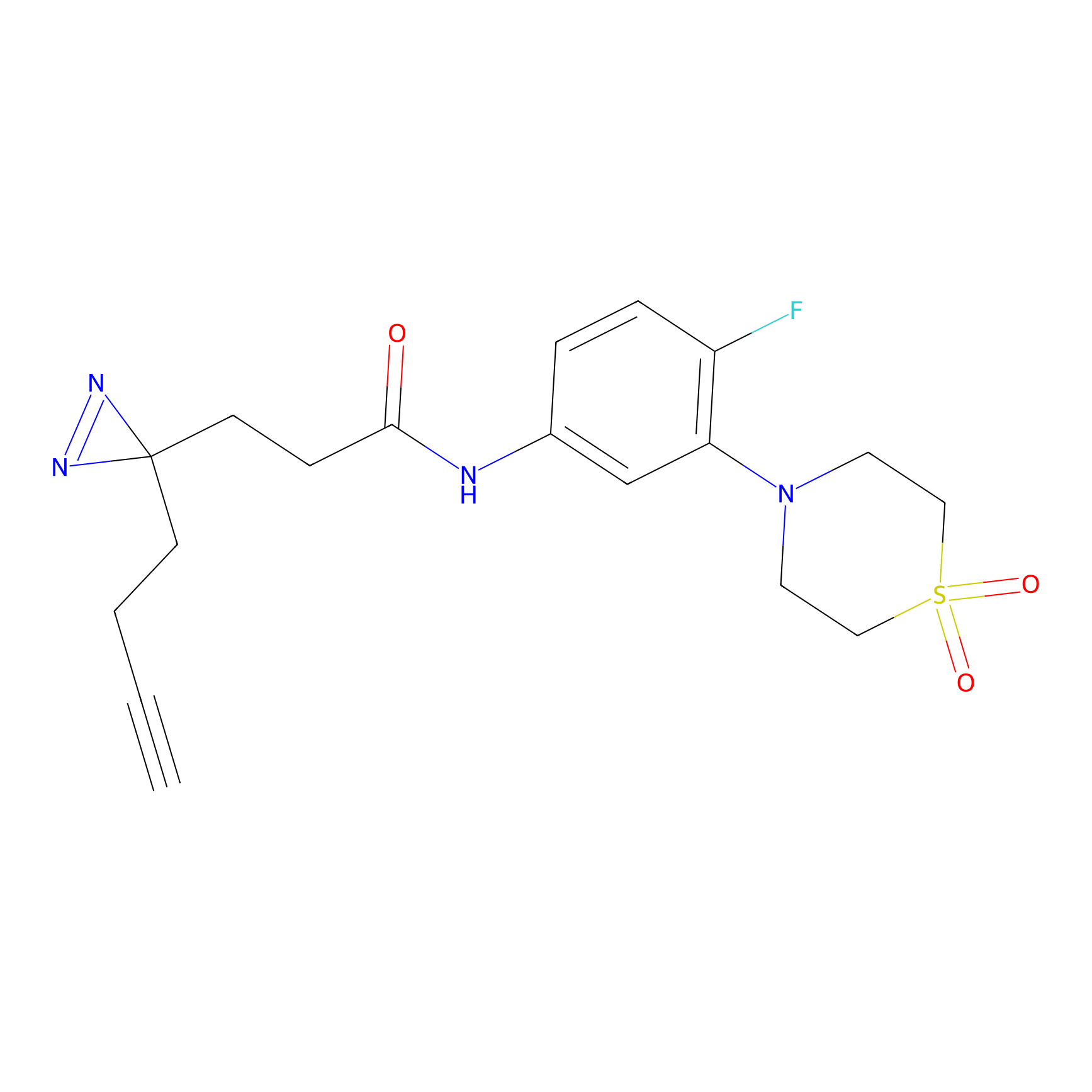
Details of the Target
General Information of Target
Probe(s) Labeling This Target
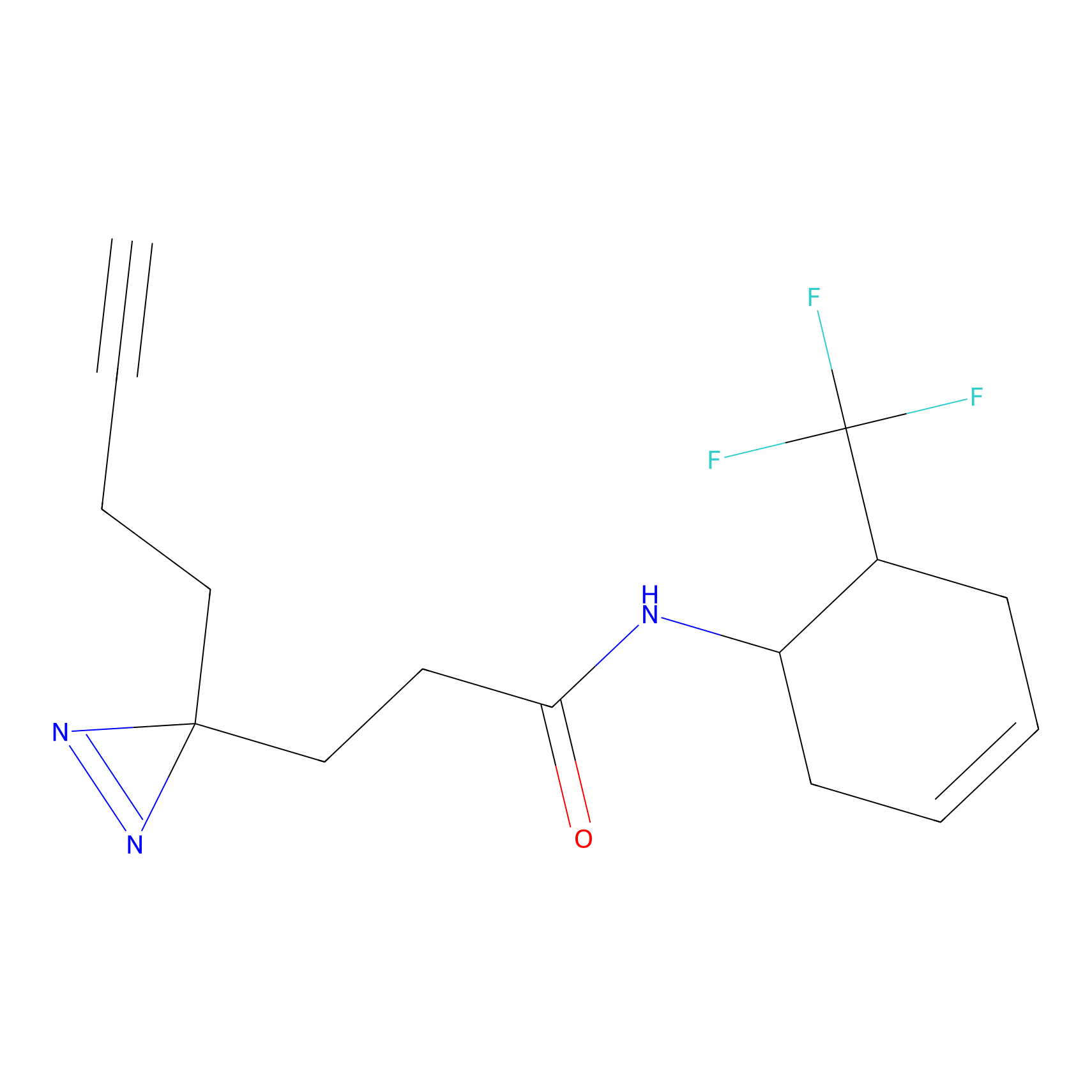
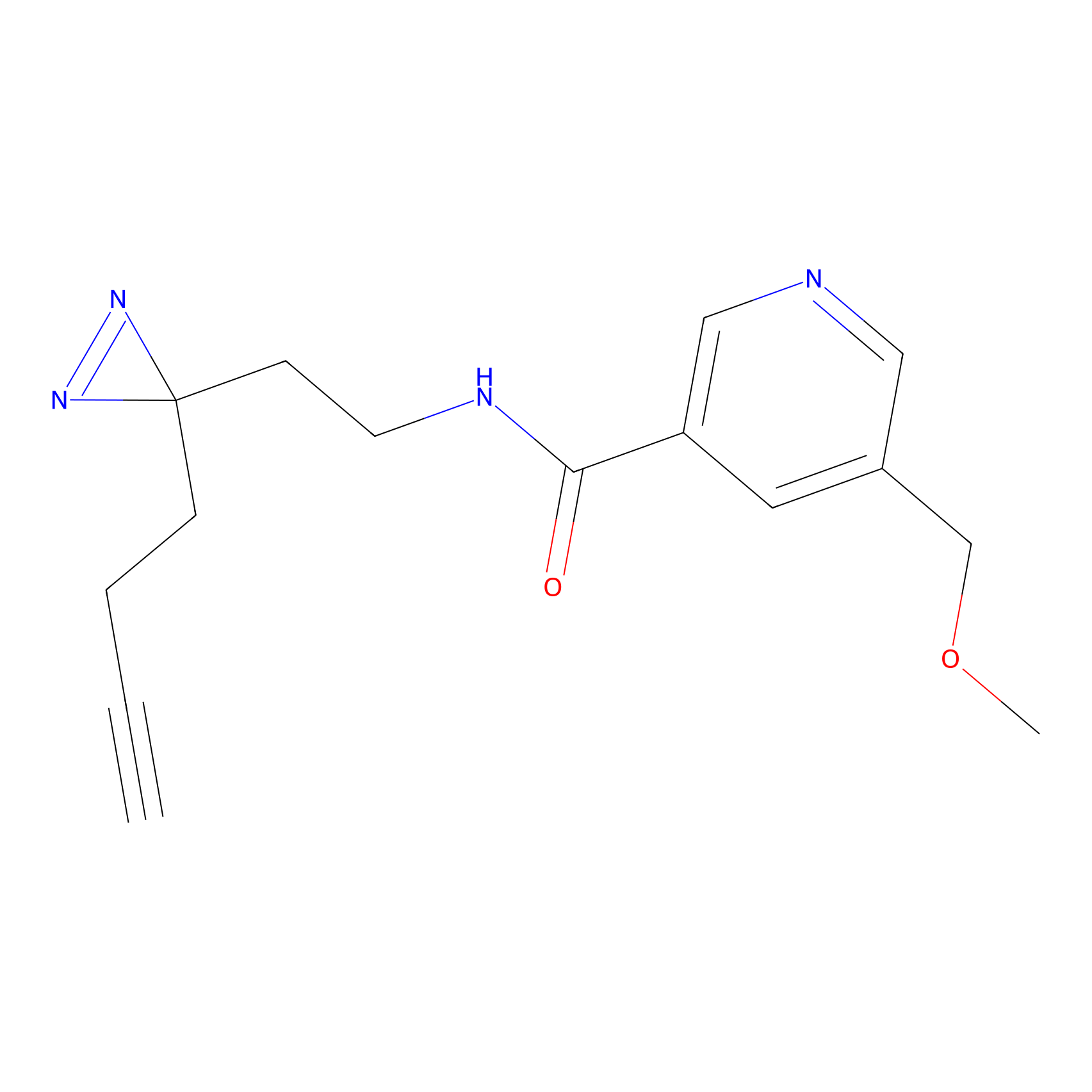
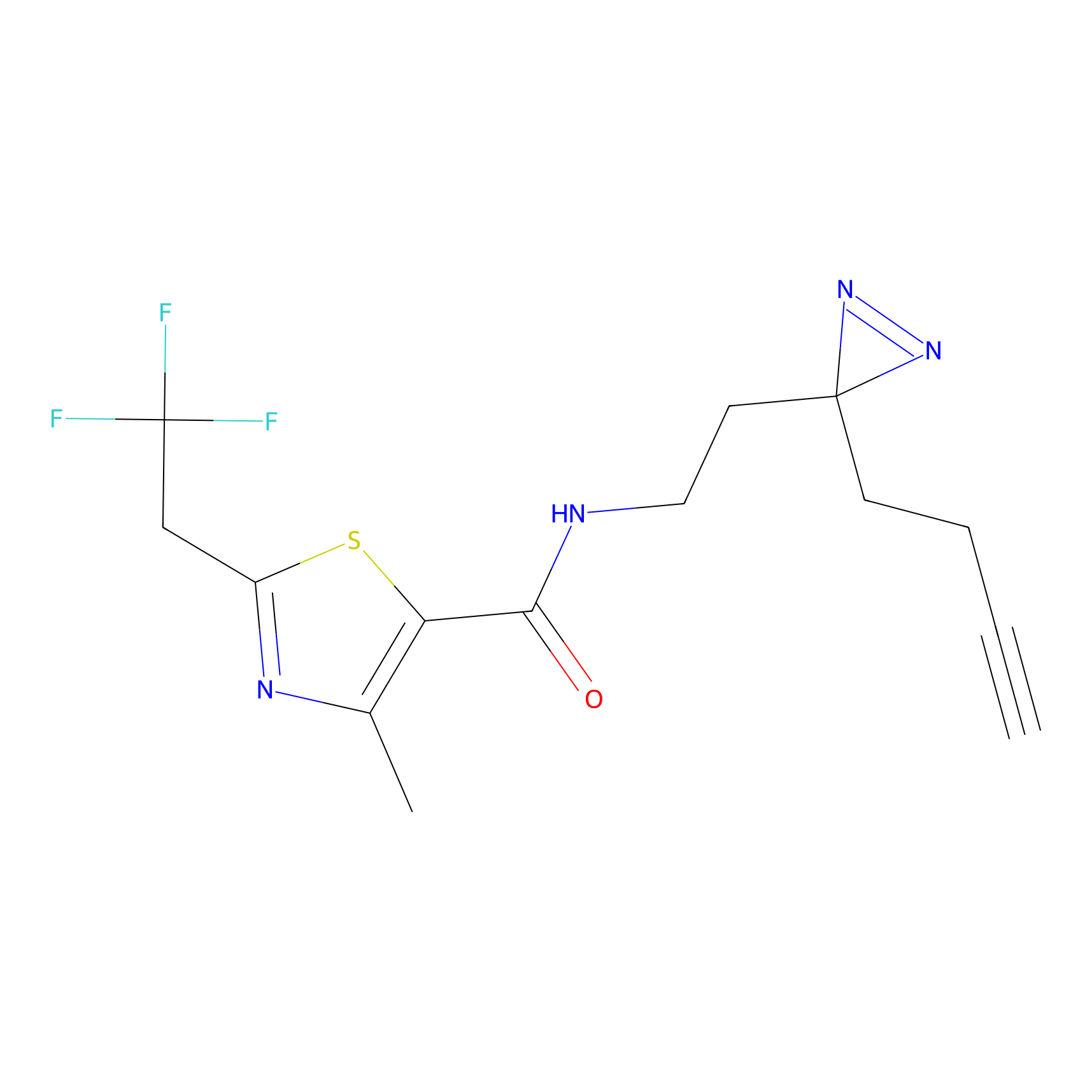
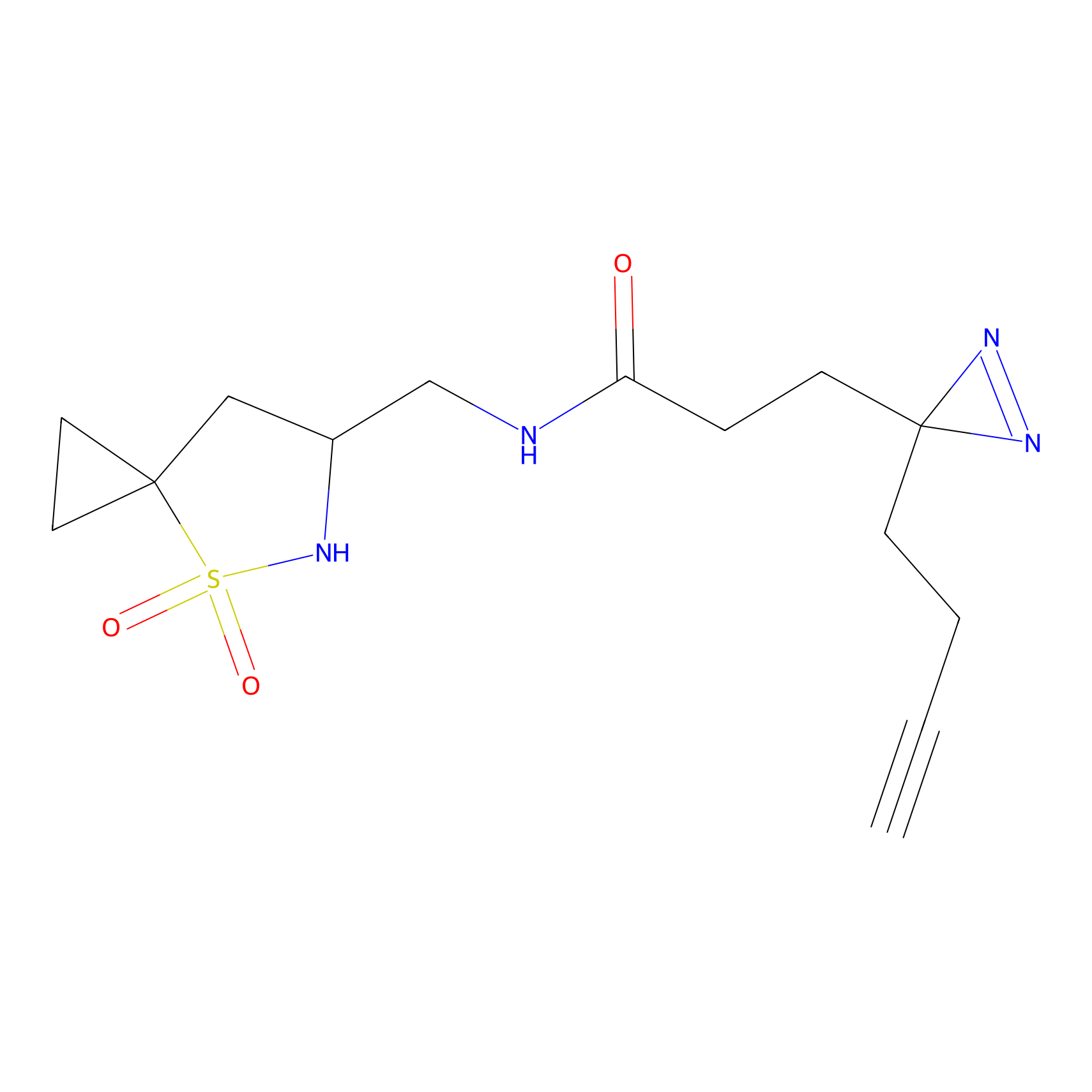
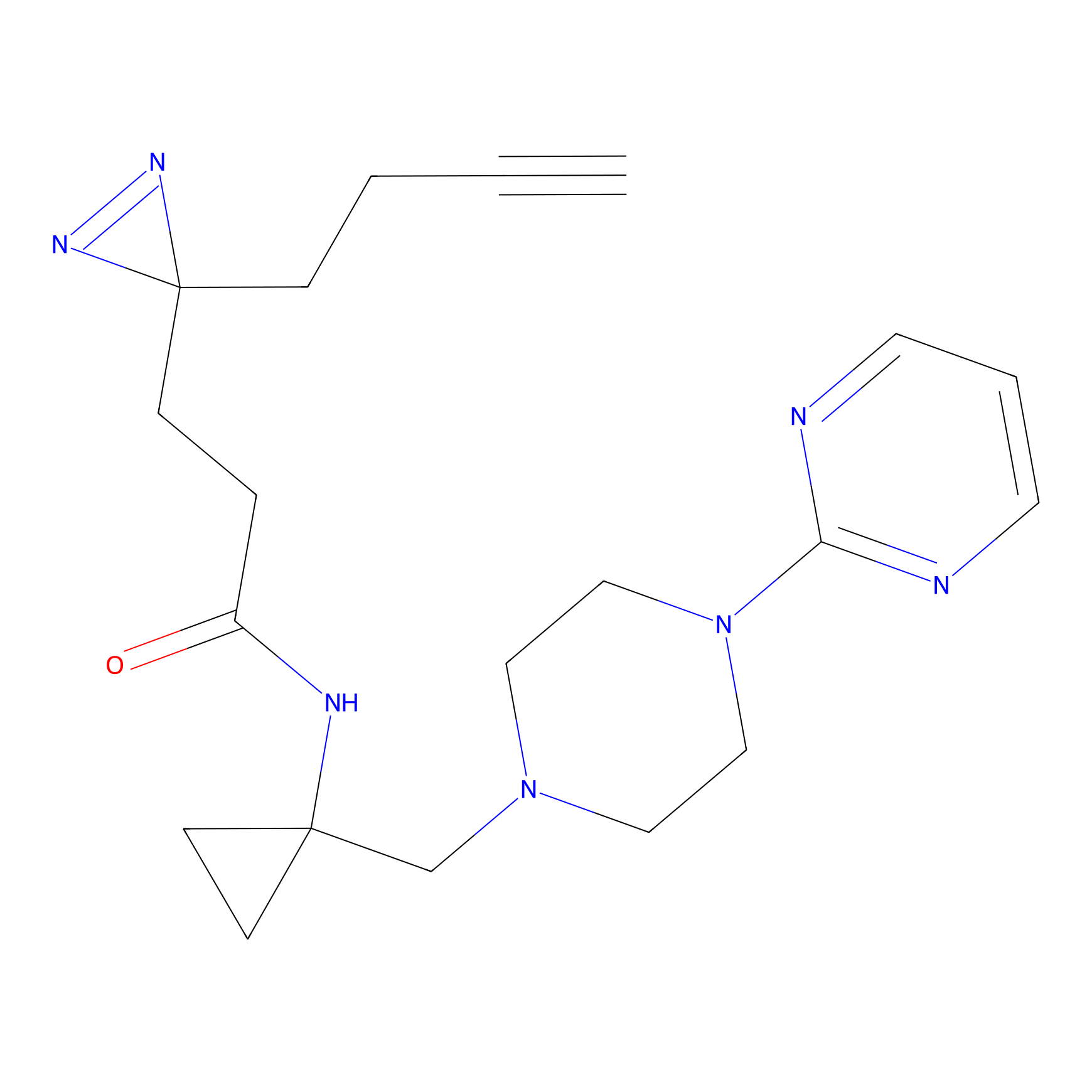
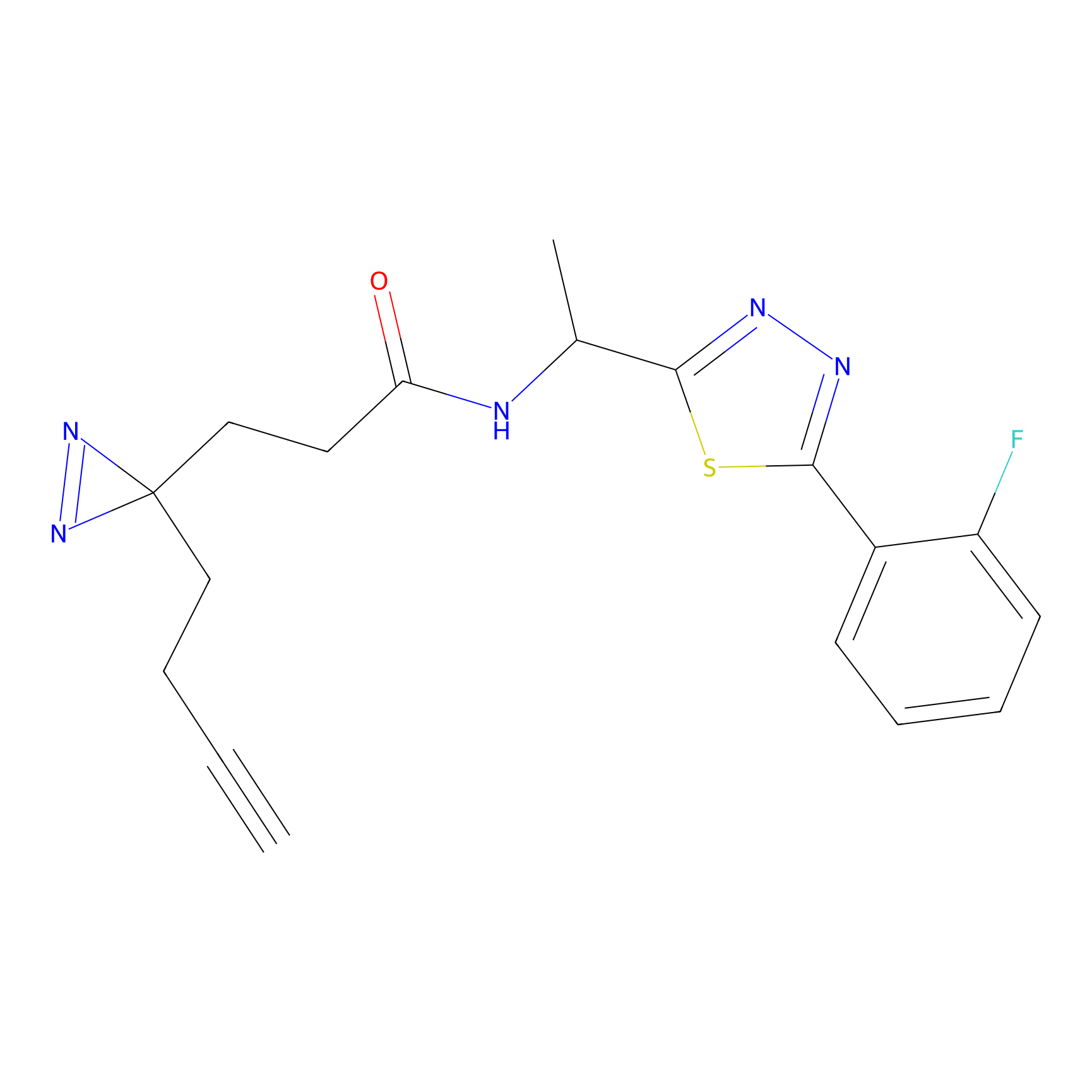
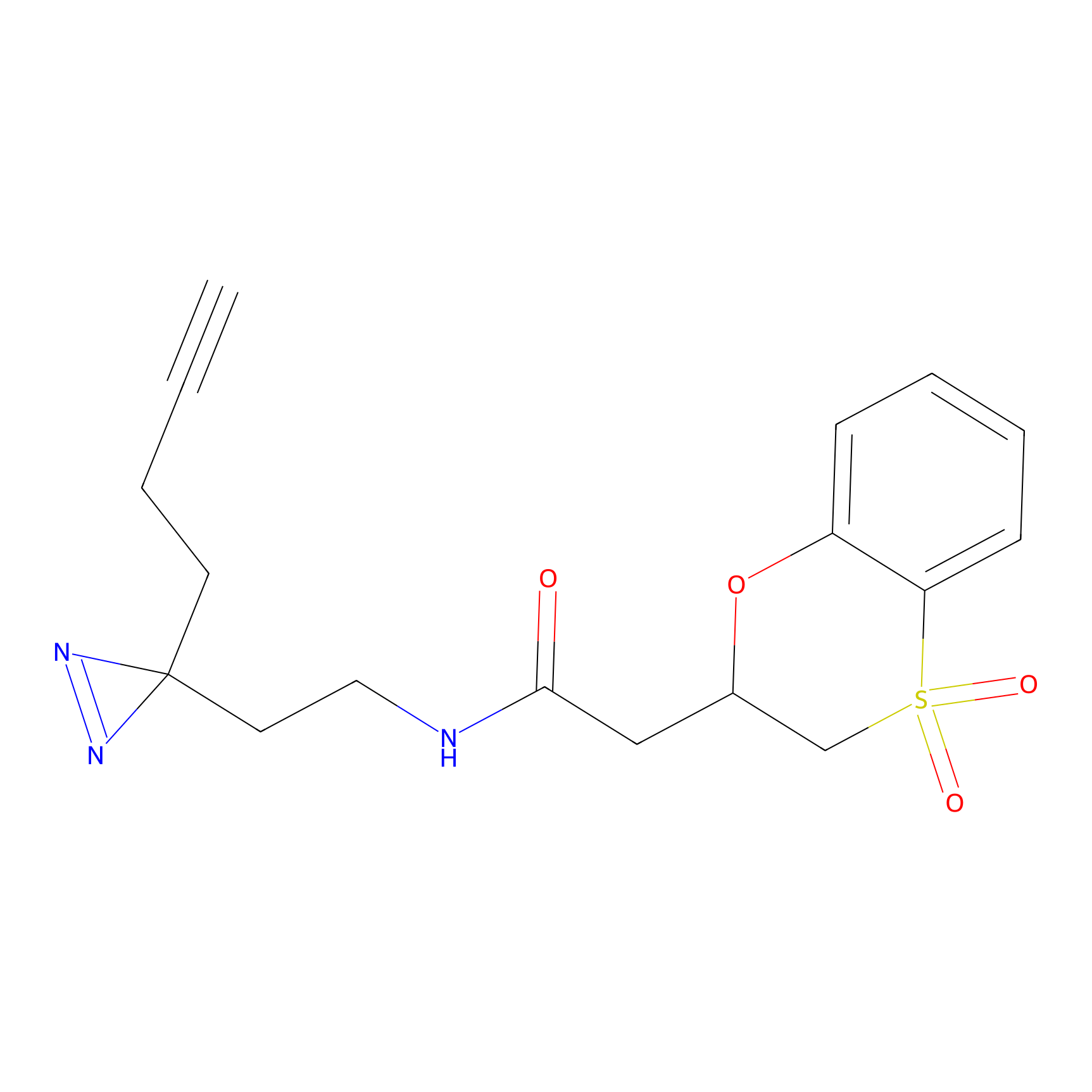
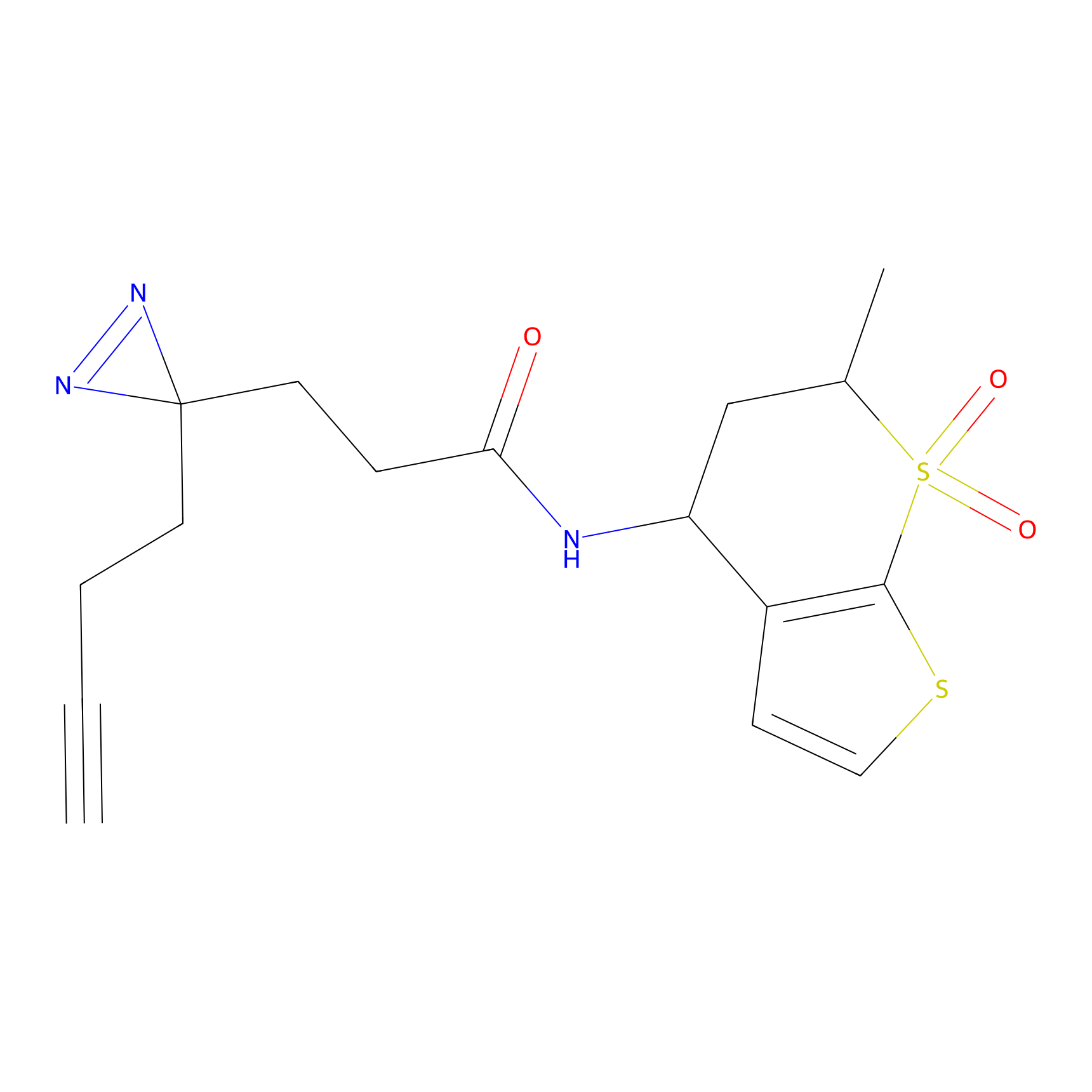
ABPP Probe
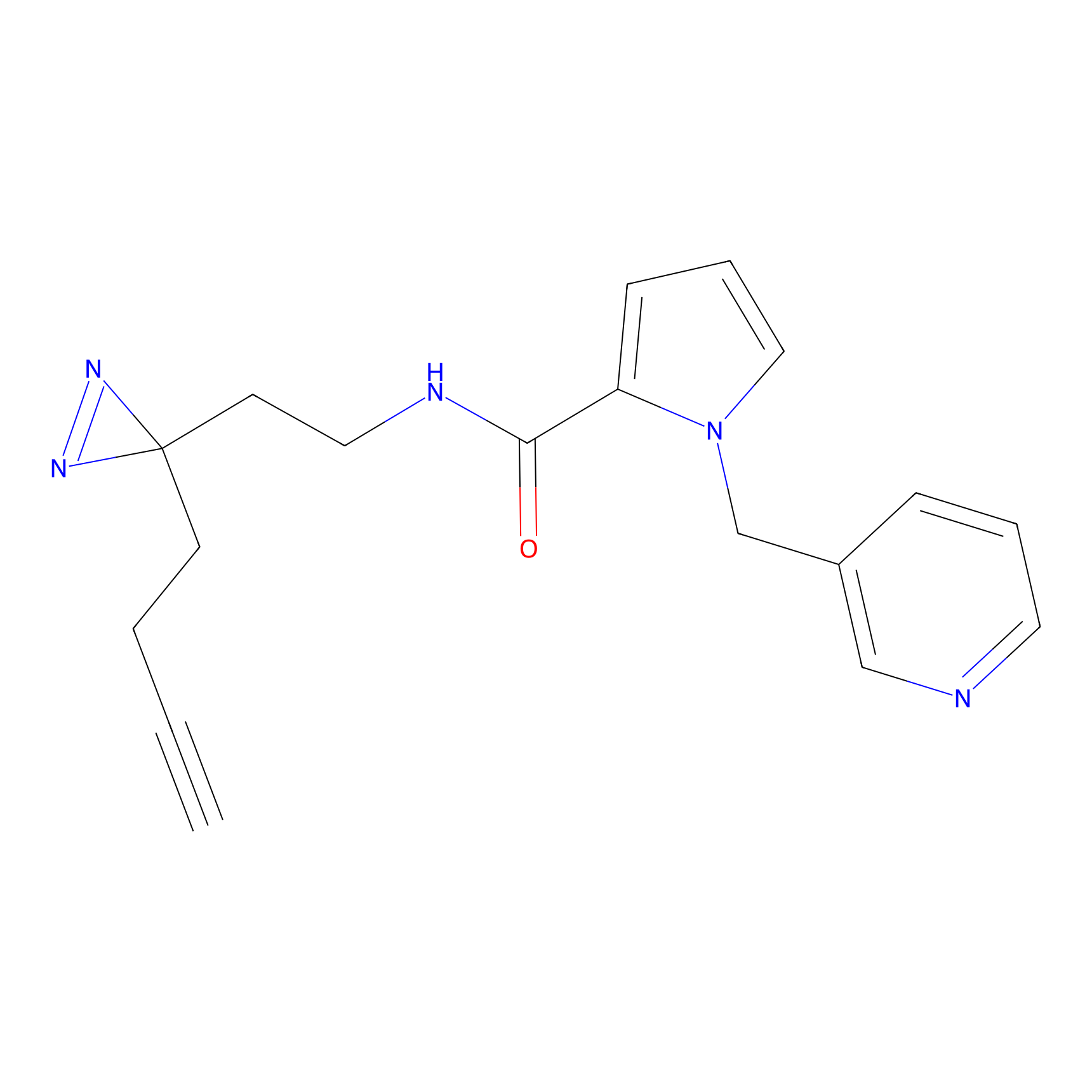
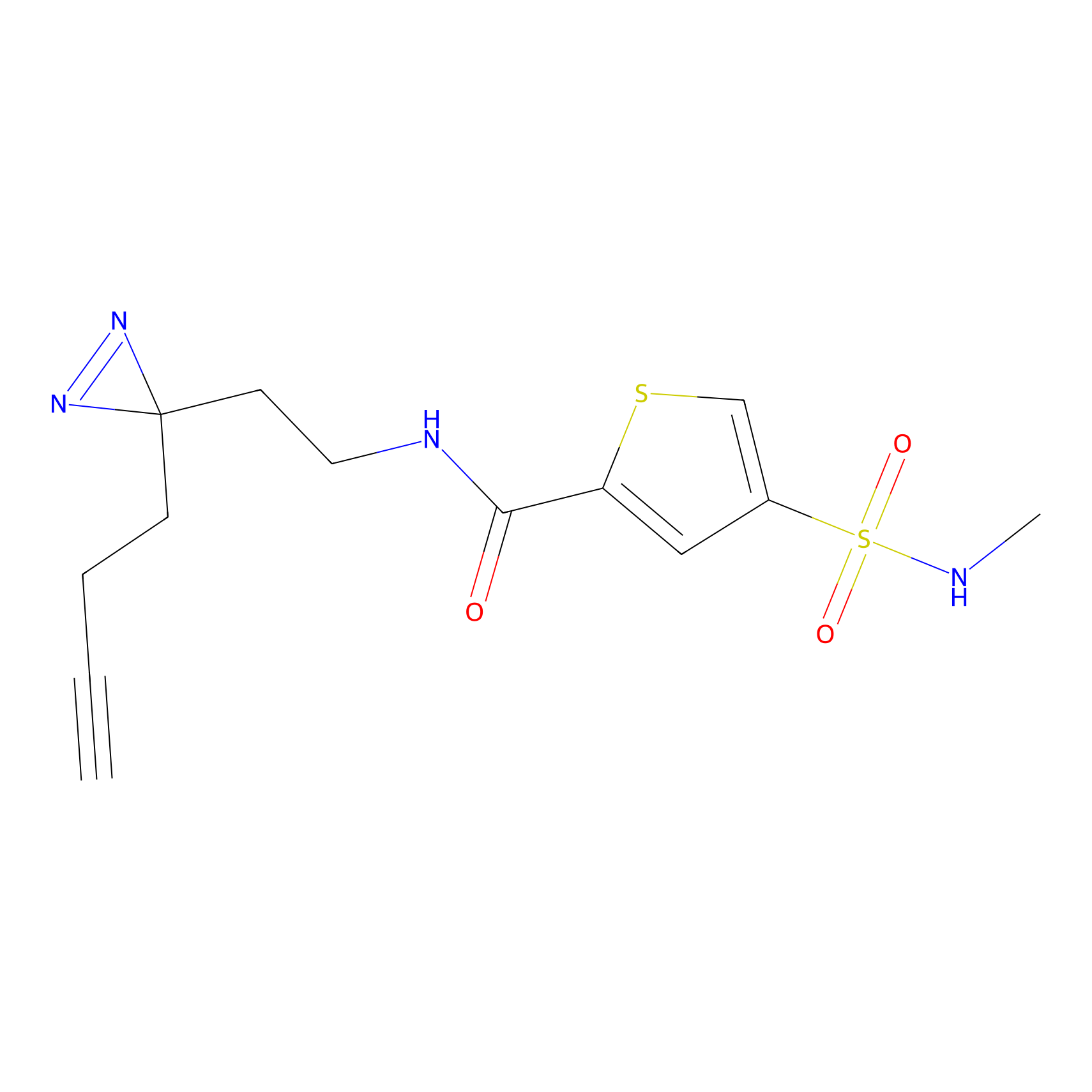
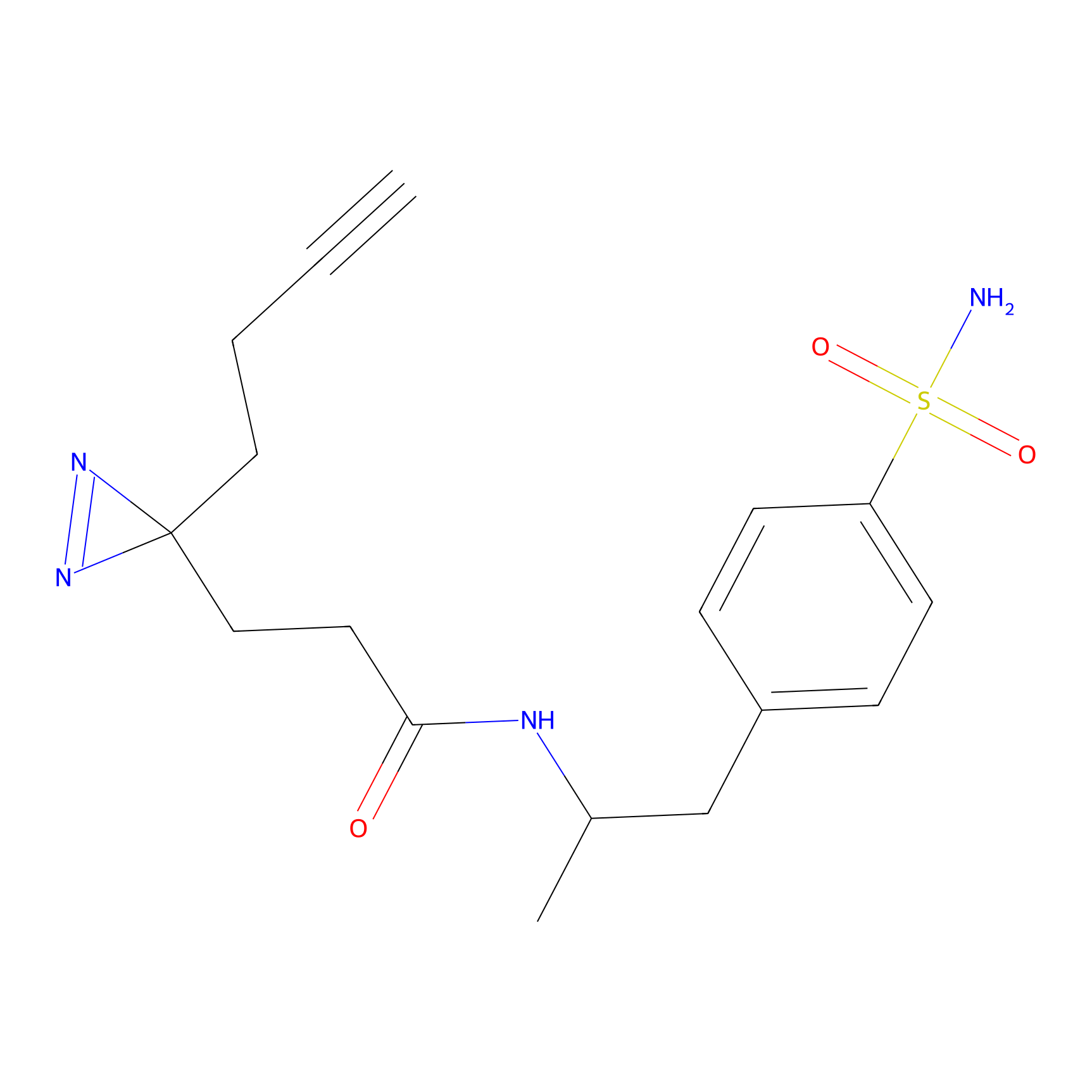
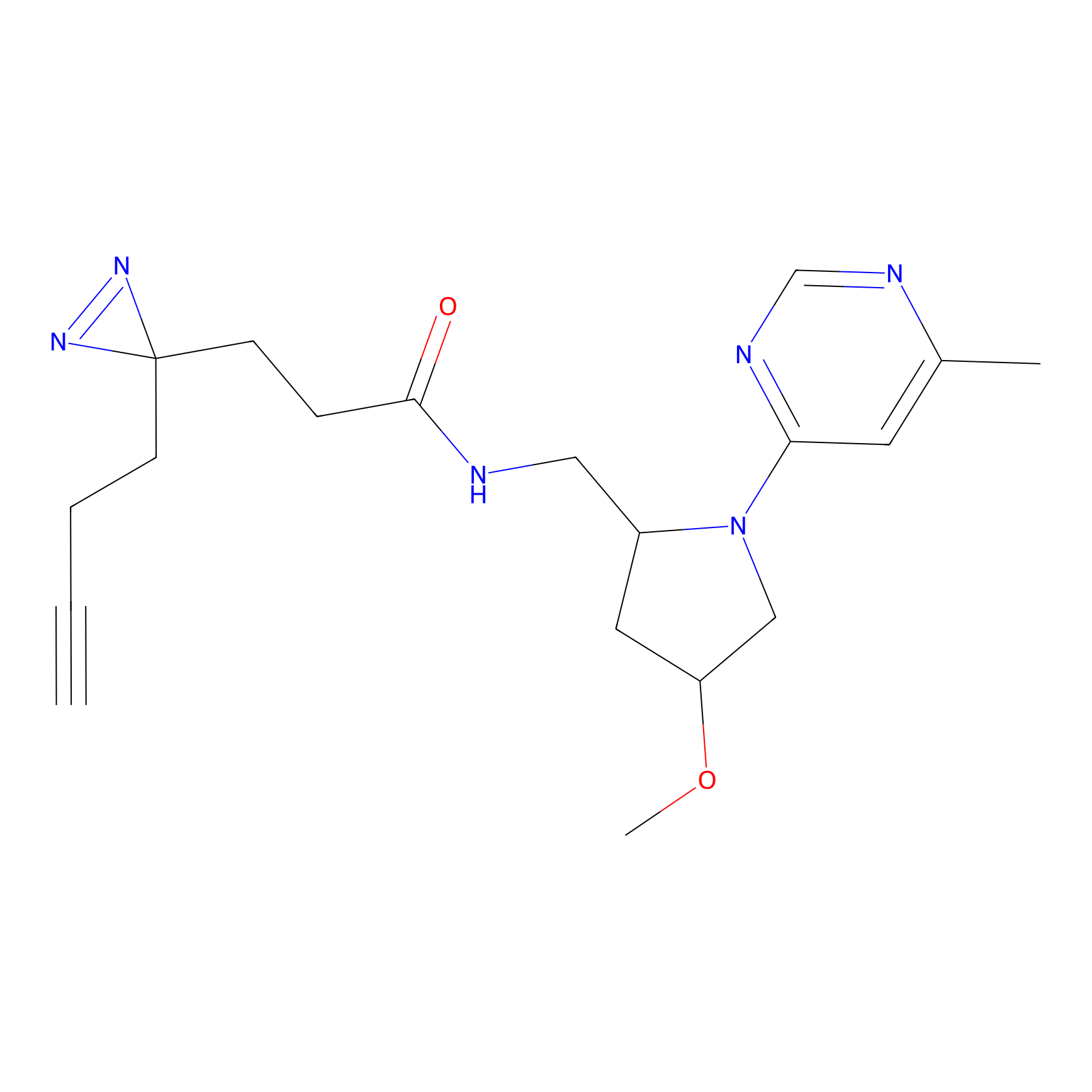
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
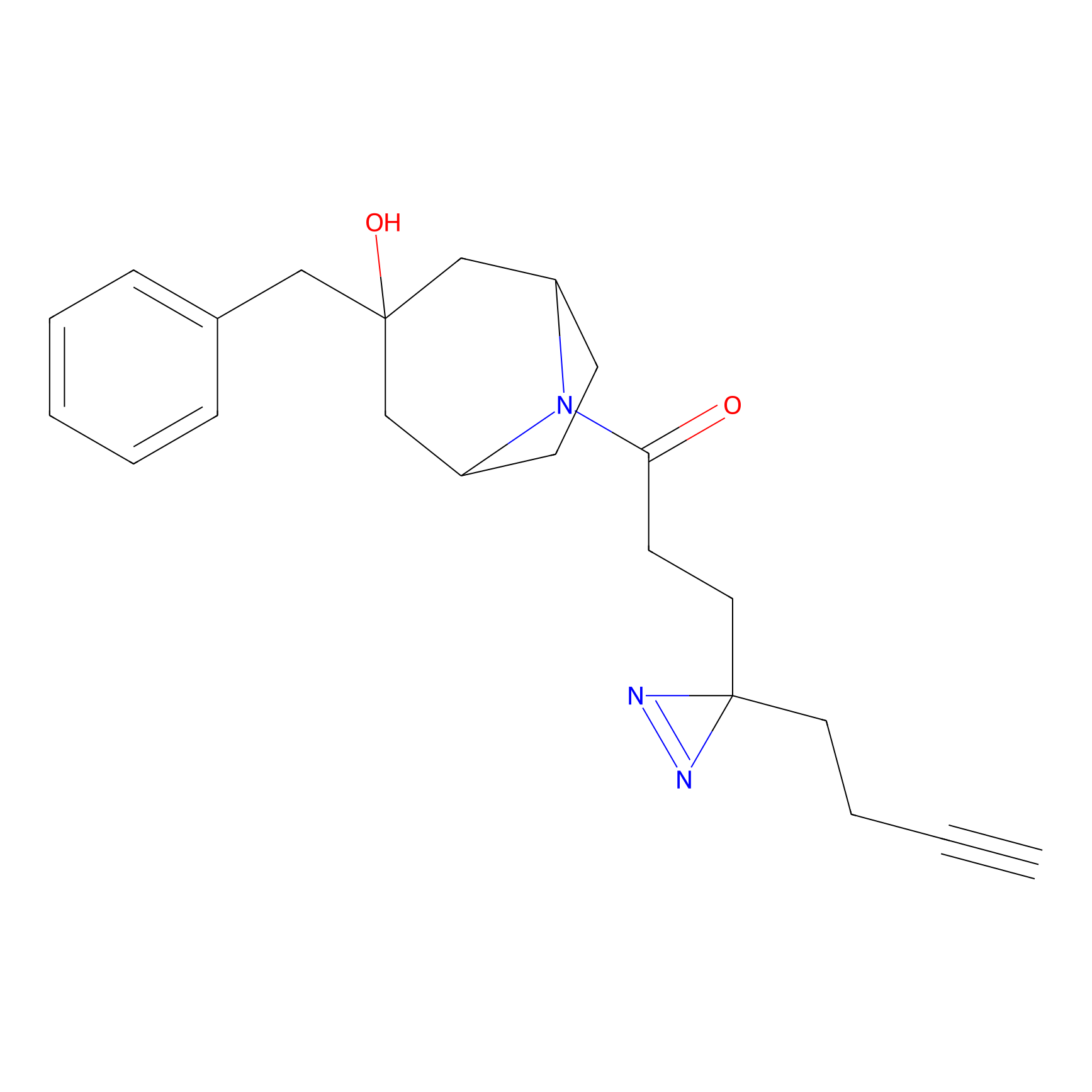
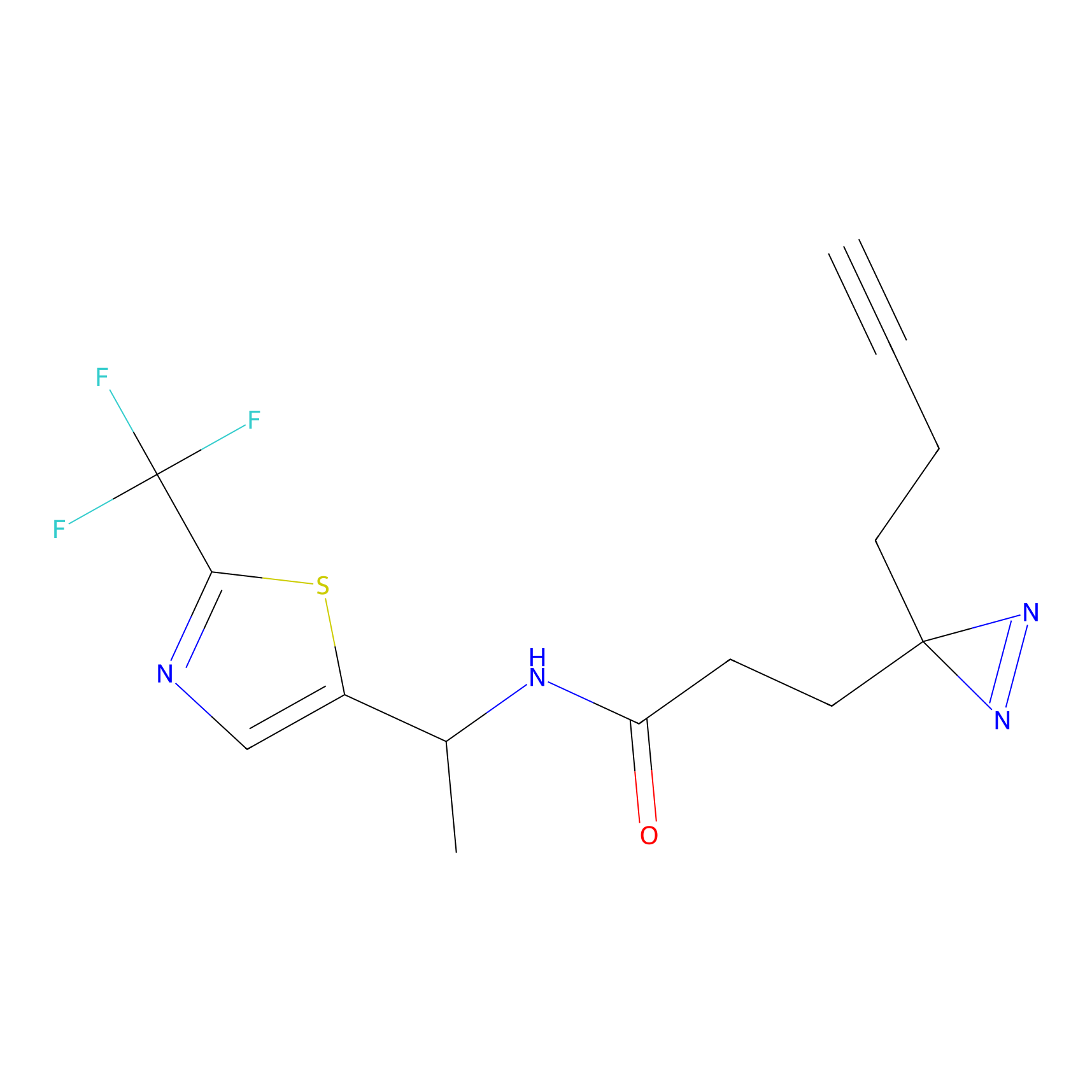
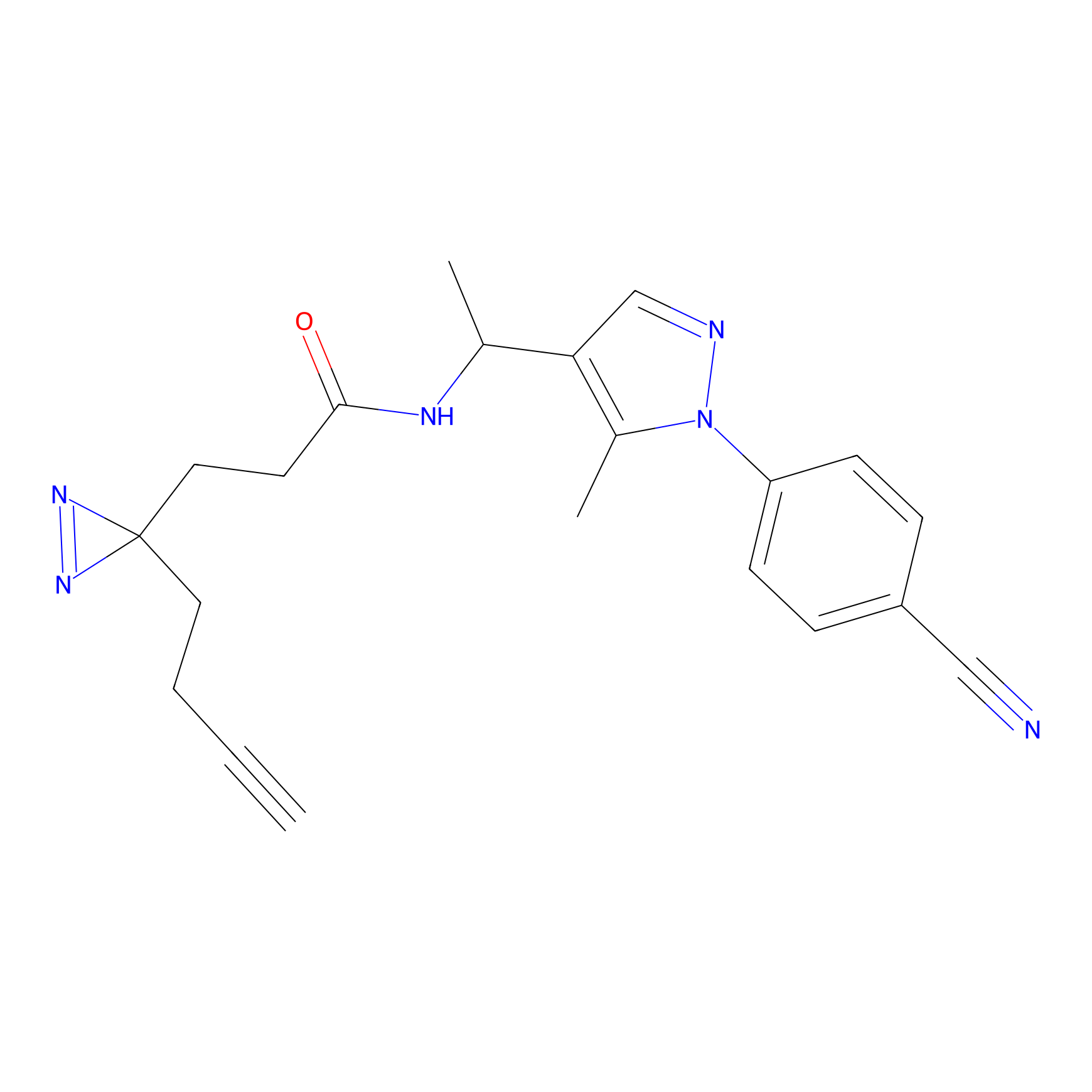
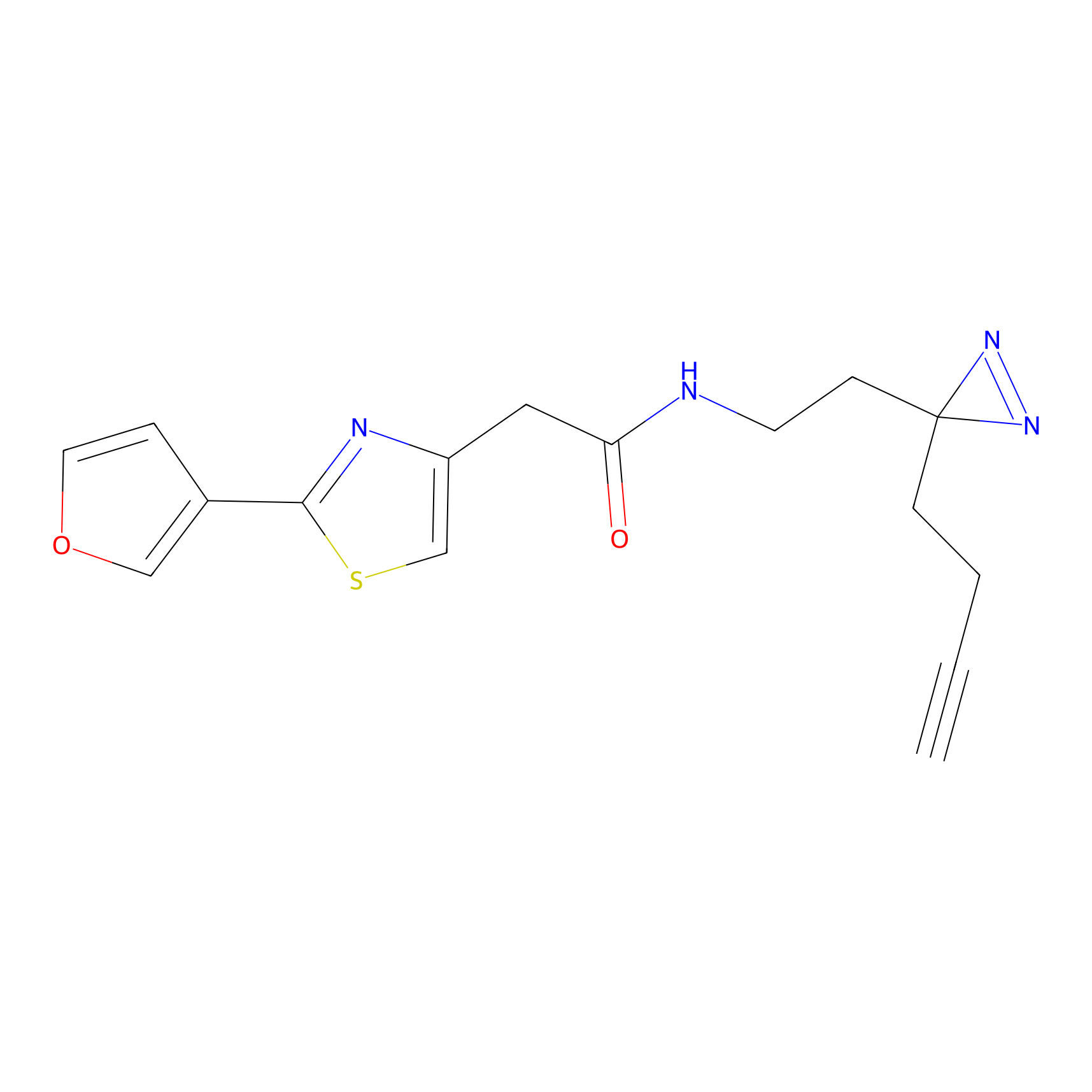
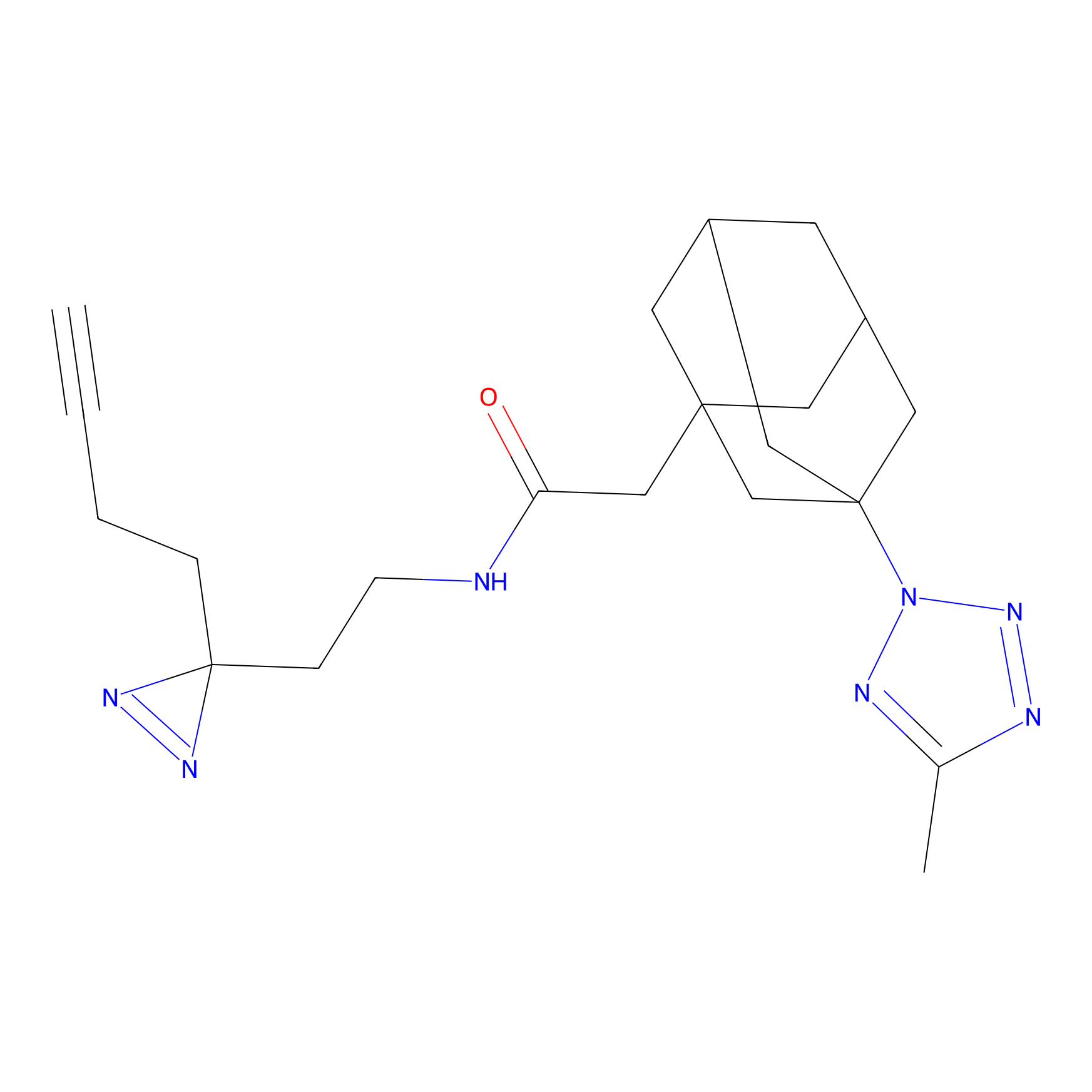
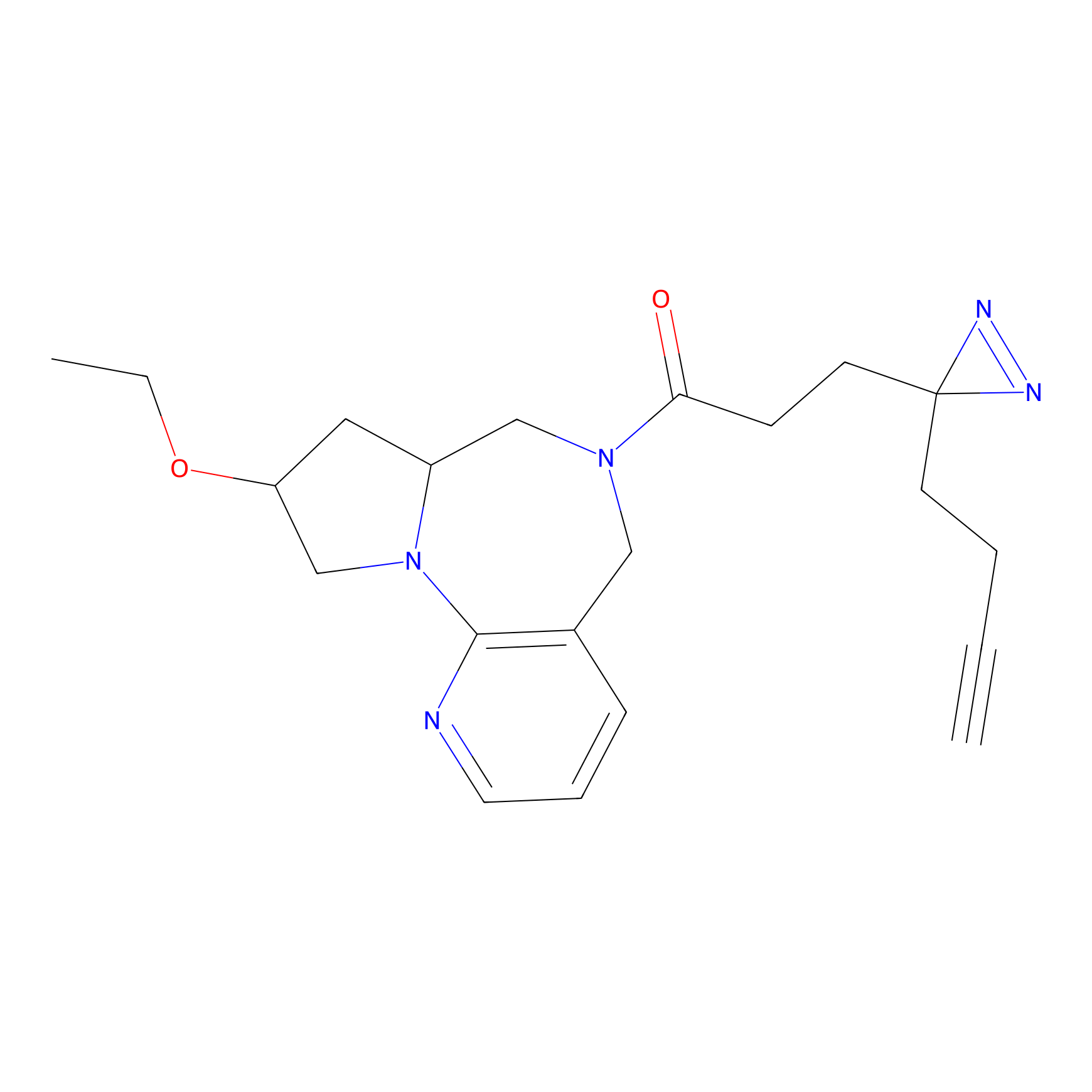
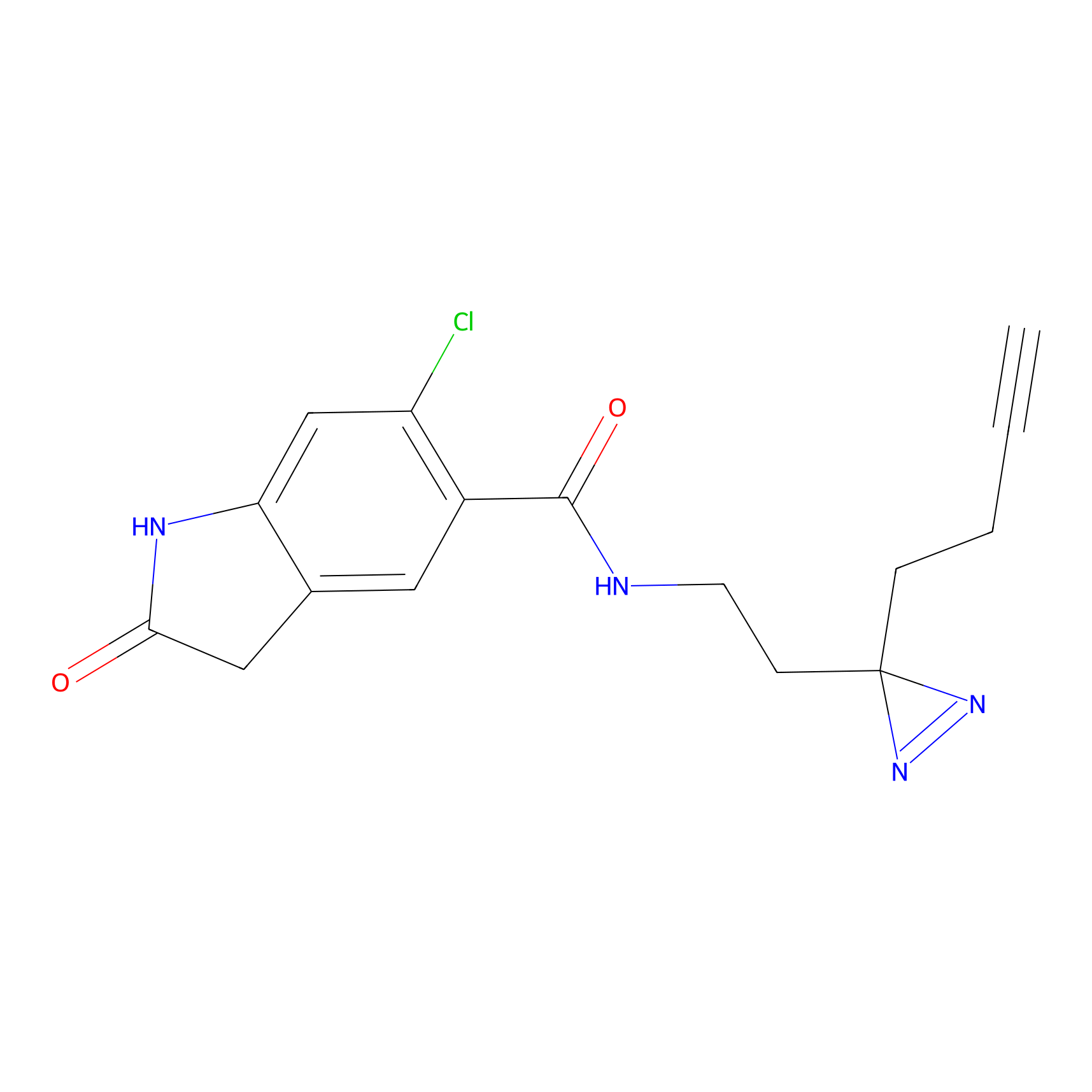
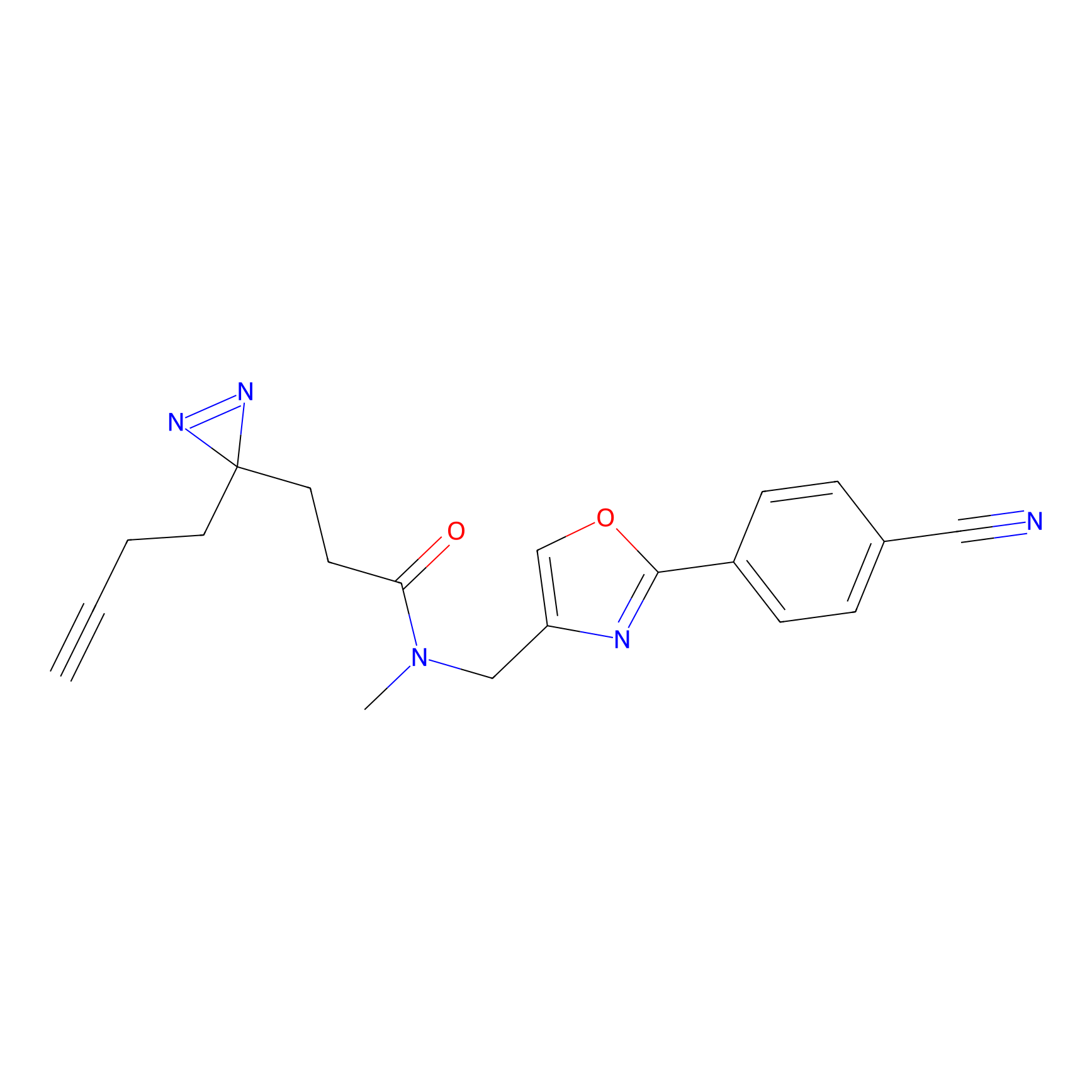
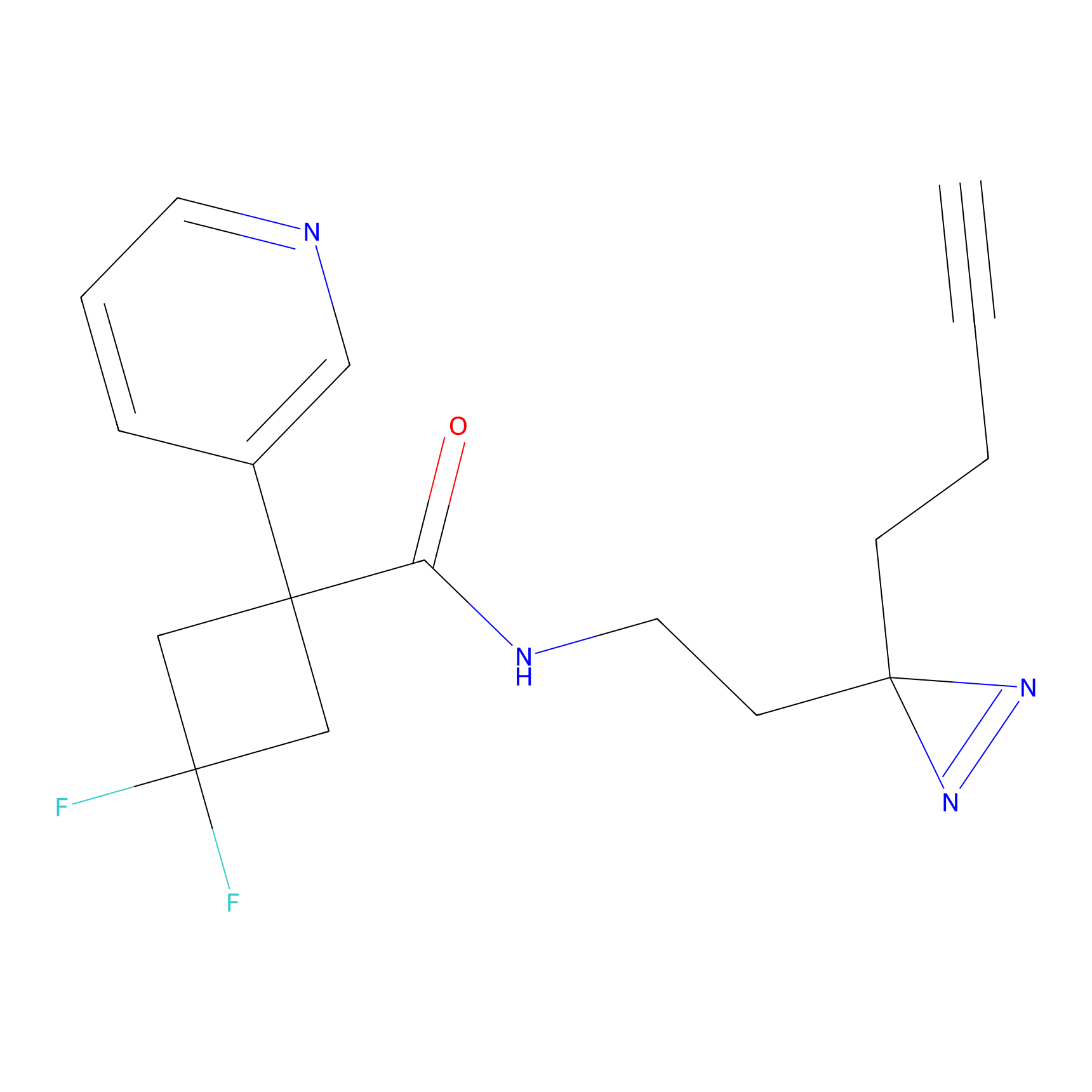
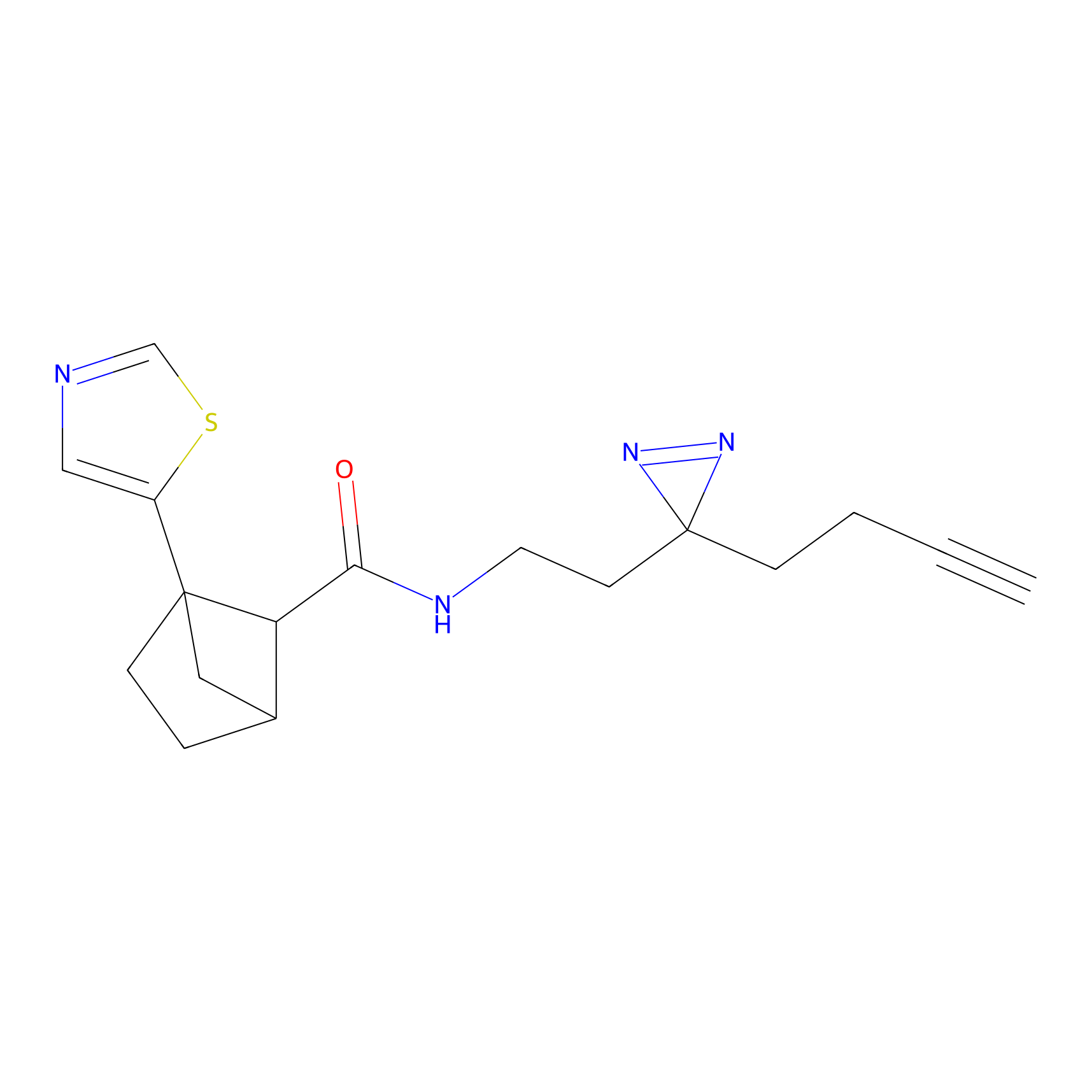
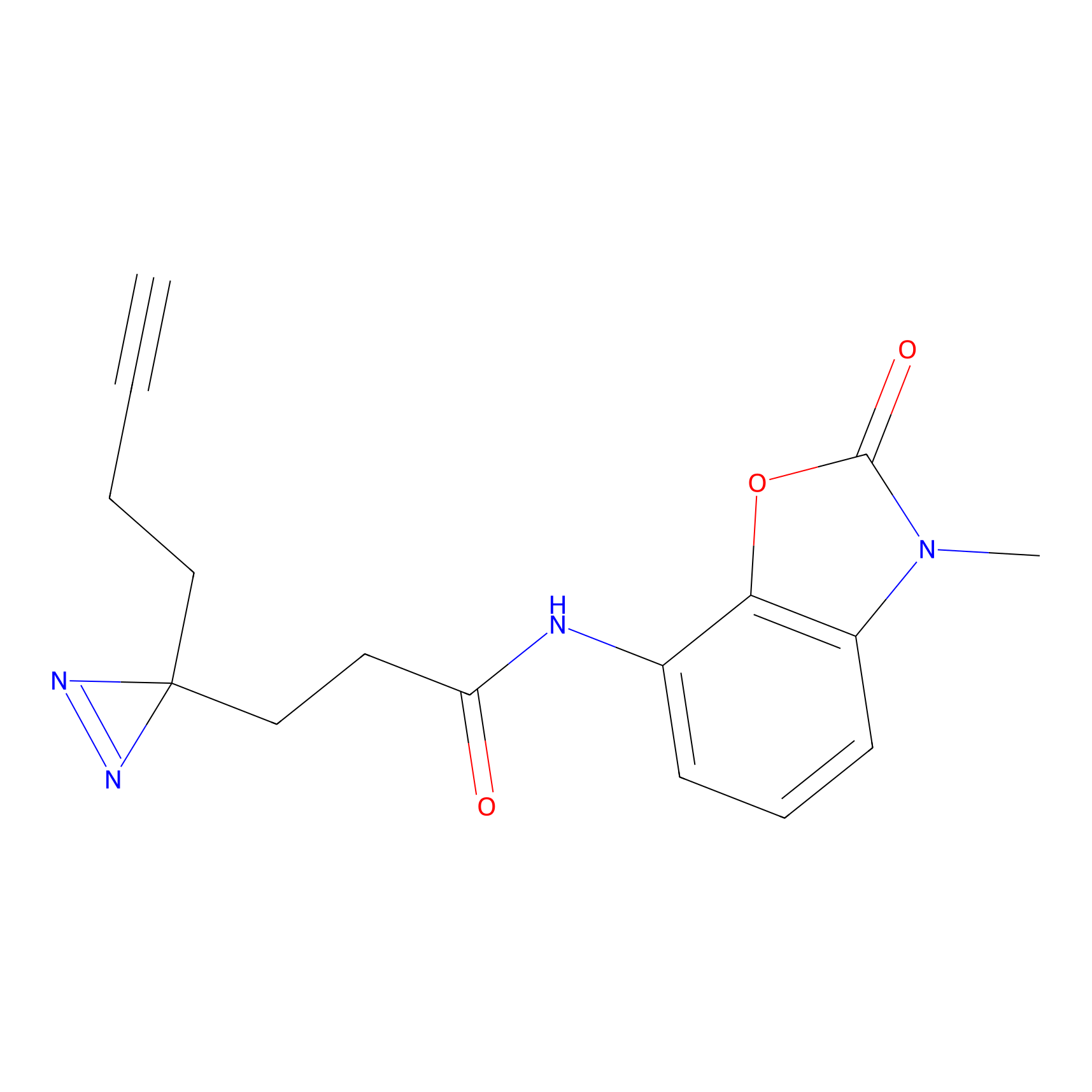
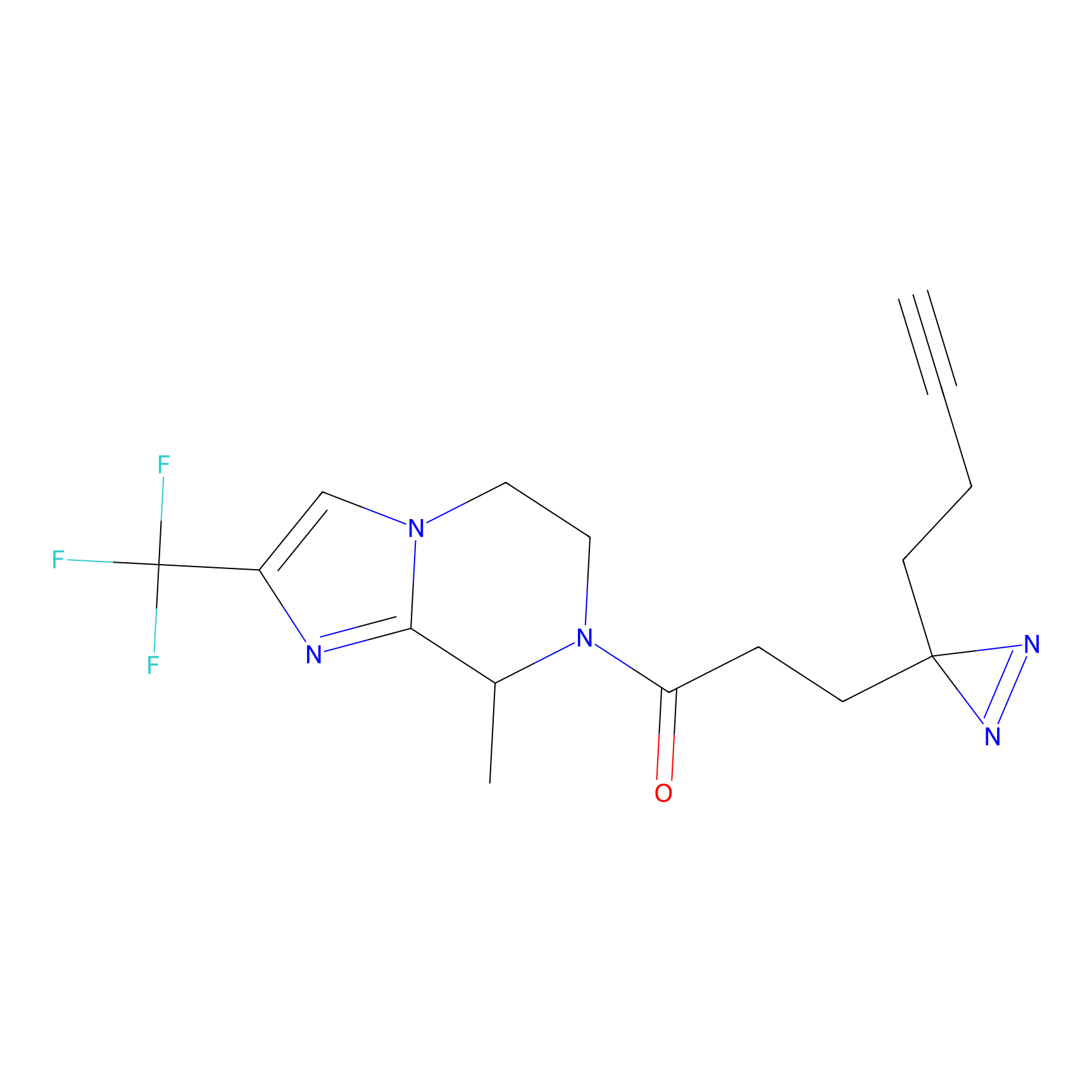
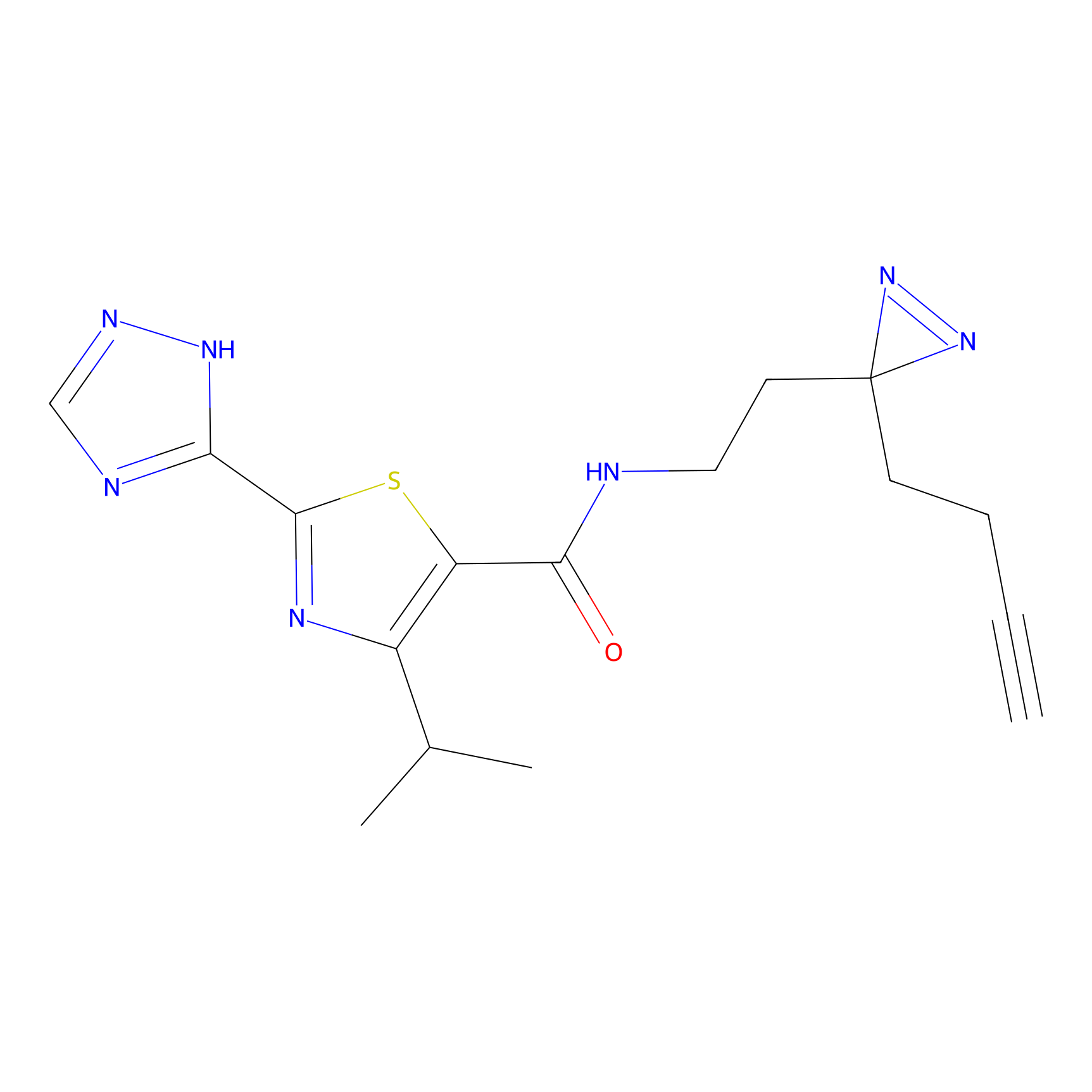
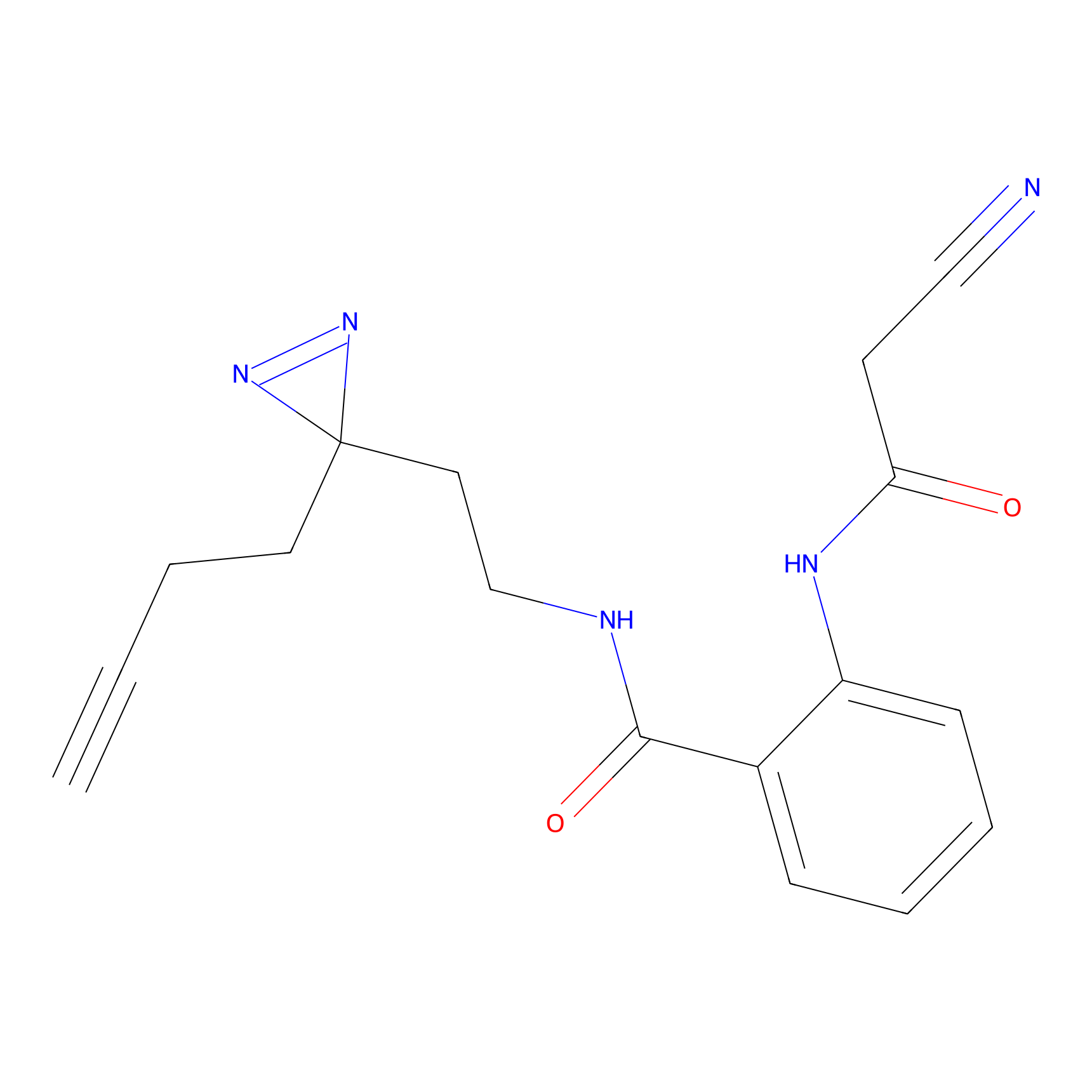
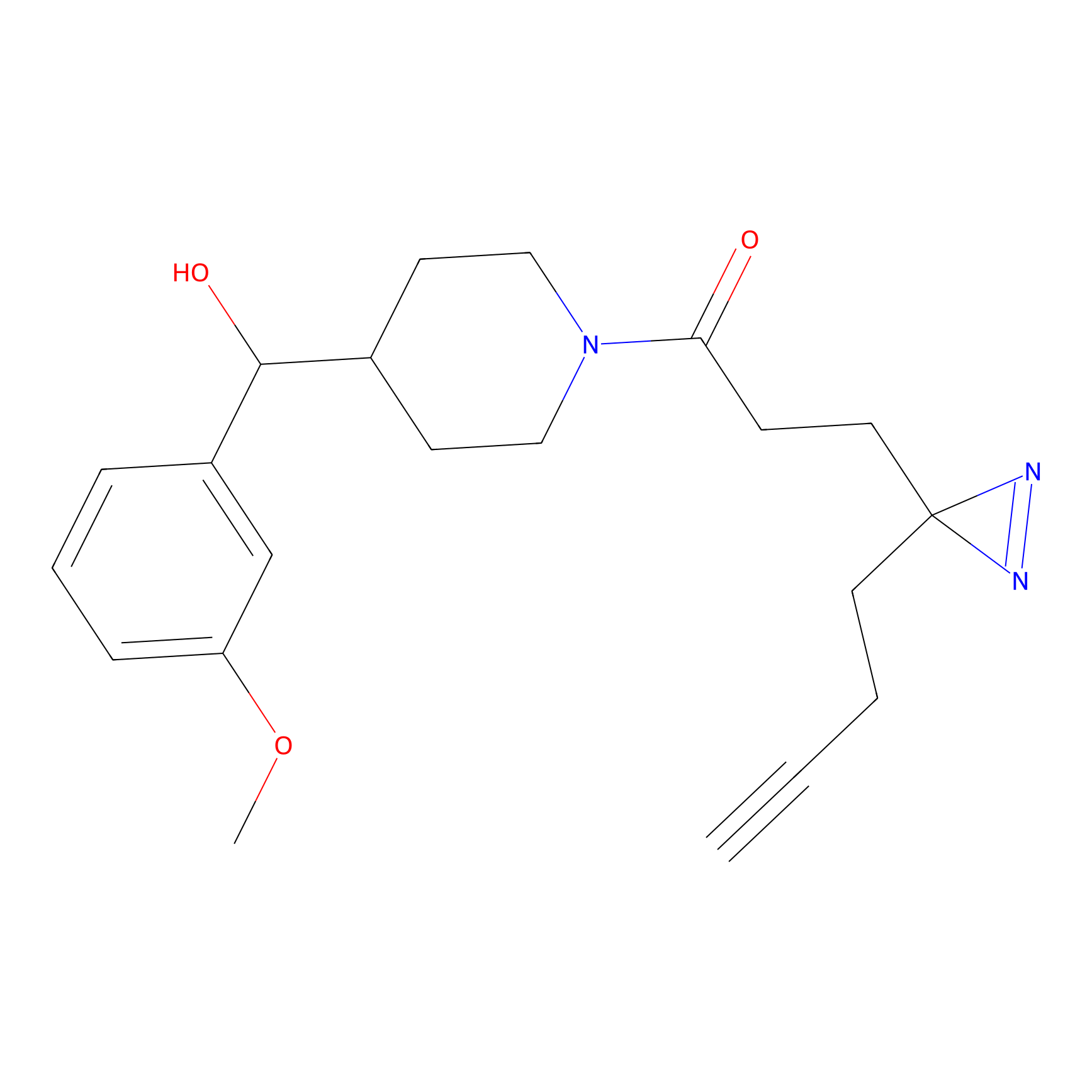
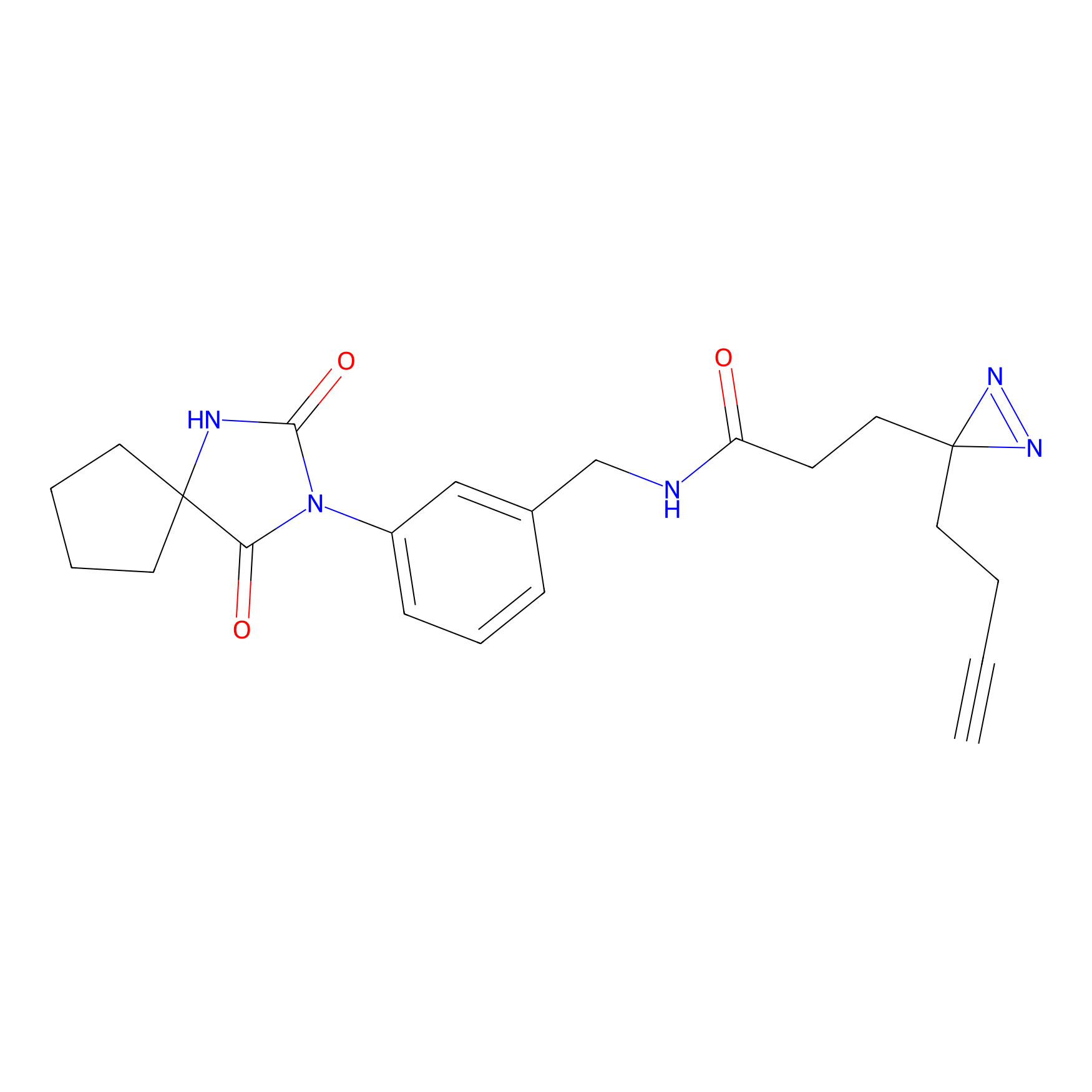
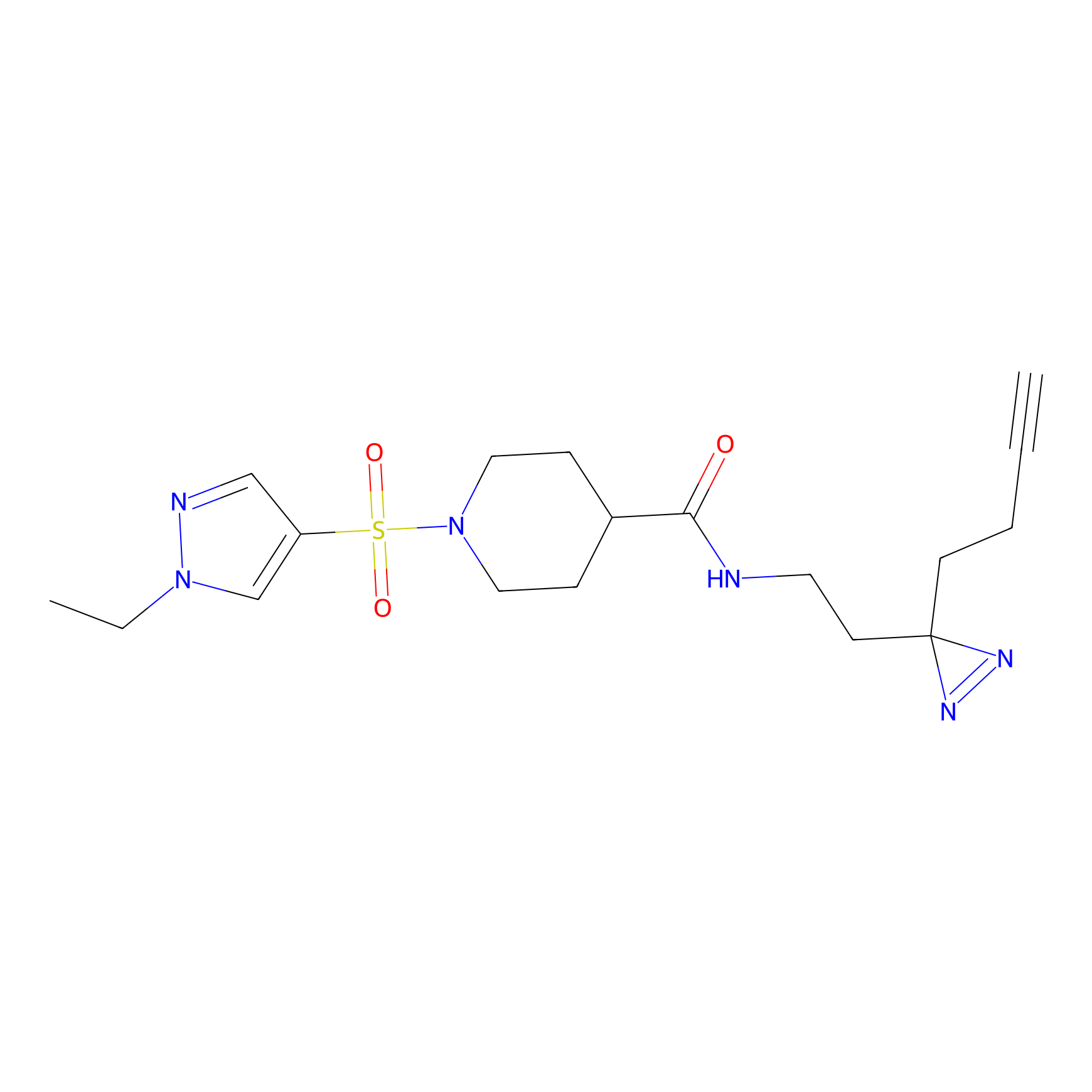
Jackson_1 Probe Info |
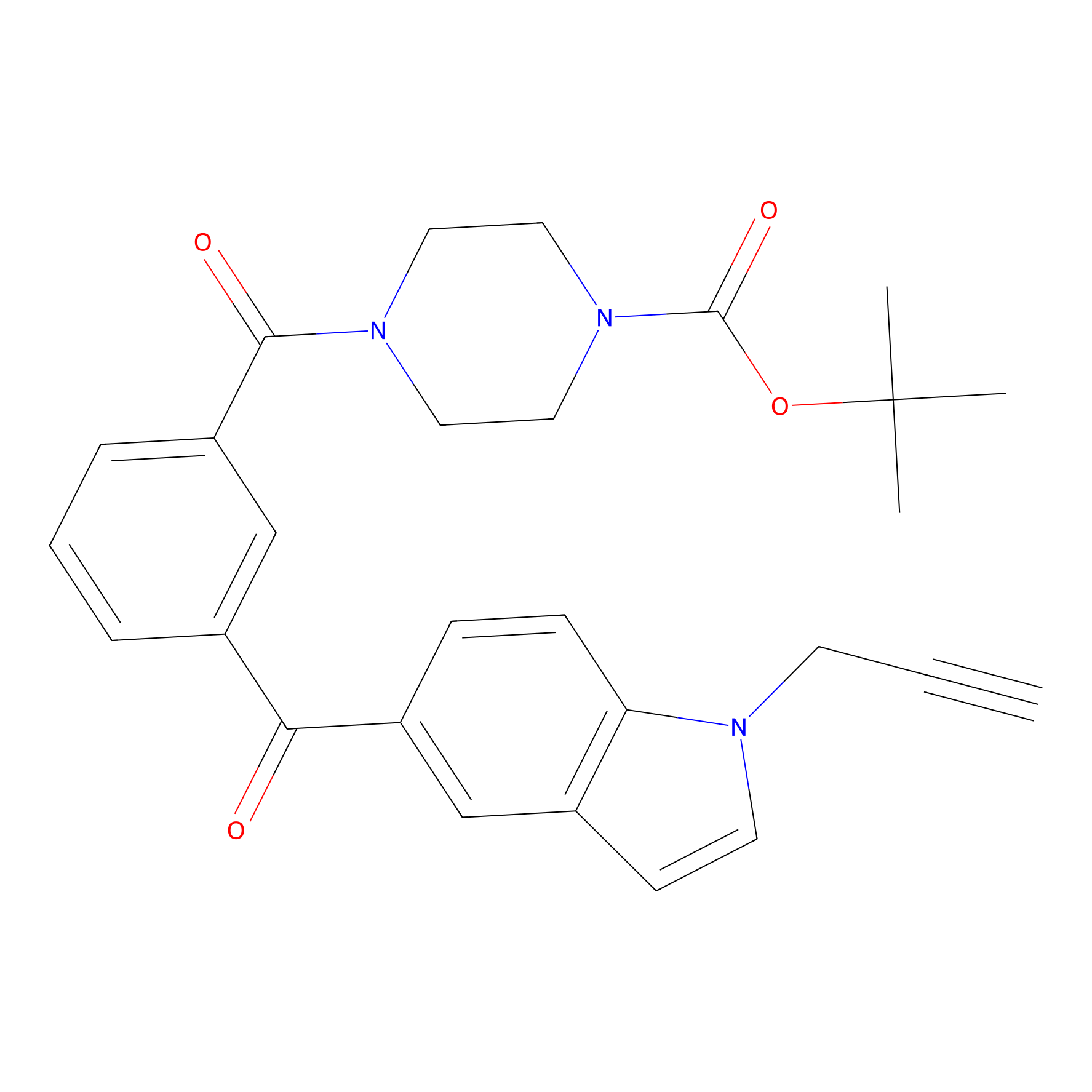 |
2.23 | LDD0120 | [1] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nicotinamide phosphoribosyltransferase (NAMPT) | NAPRTase family | P43490 | |||
The Drug(s) Related To This Target
Approved
Investigative
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| N-formylmethionine | Small molecular drug | DB04464 | |||
| Phenethyl Isothiocyanate | Small molecular drug | DB12695 | |||
References