Details of the Target
General Information of Target
| Target ID | LDTP00235 | |||||
|---|---|---|---|---|---|---|
| Target Name | Membrane-associated progesterone receptor component 1 (PGRMC1) | |||||
| Gene Name | PGRMC1 | |||||
| Gene ID | 10857 | |||||
| Synonyms |
HPR6.6; PGRMC; Membrane-associated progesterone receptor component 1; mPR; Dap1; IZA |
|||||
| 3D Structure | ||||||
| Sequence |
MAAEDVVATGADPSDLESGGLLHEIFTSPLNLLLLGLCIFLLYKIVRGDQPAASGDSDDD
EPPPLPRLKRRDFTPAELRRFDGVQDPRILMAINGKVFDVTKGRKFYGPEGPYGVFAGRD ASRGLATFCLDKEALKDEYDDLSDLTAAQQETLSDWESQFTFKYHHVGKLLKEGEEPTVY SDEEEPKDESARKND |
|||||
| Target Type |
Literature-reported
|
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Cytochrome b5 family, MAPR subfamily
|
|||||
| Subcellular location |
Microsome membrane
|
|||||
| Function |
Component of a progesterone-binding protein complex. Binds progesterone. Has many reported cellular functions (heme homeostasis, interaction with CYPs). Required for the maintenance of uterine histoarchitecture and normal female reproductive lifespan. Intracellular heme chaperone. Regulates heme synthesis via interactions with FECH and acts as a heme donor for at least some hemoproteins. Forms a ternary complex with TMEM97 receptor and low density lipid receptor/LDLR, which increases LDLR-mediated LDL lipoprotein internalization.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
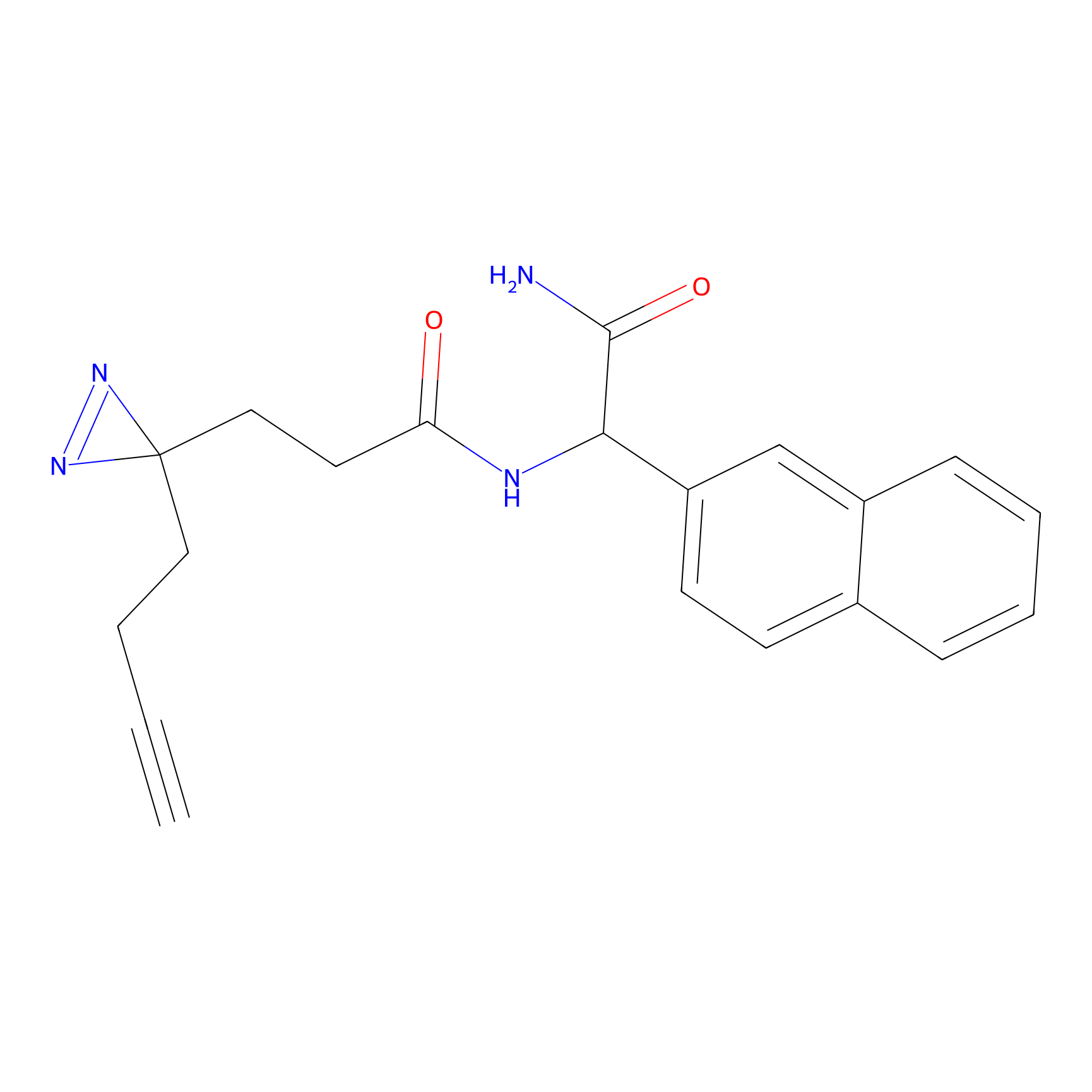
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
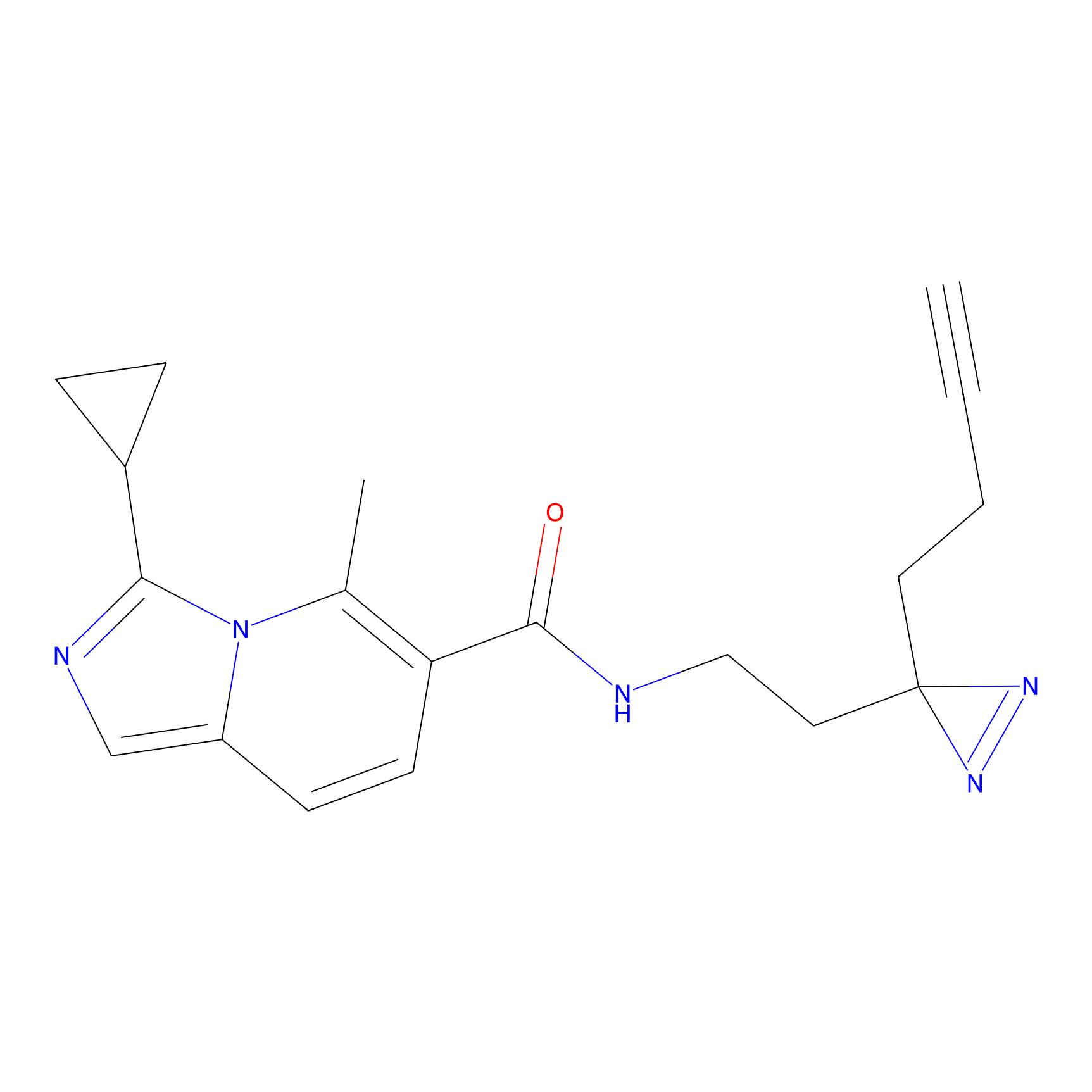
FBPP2 Probe Info |
 |
4.01 | LDD0318 | [2] | |
|
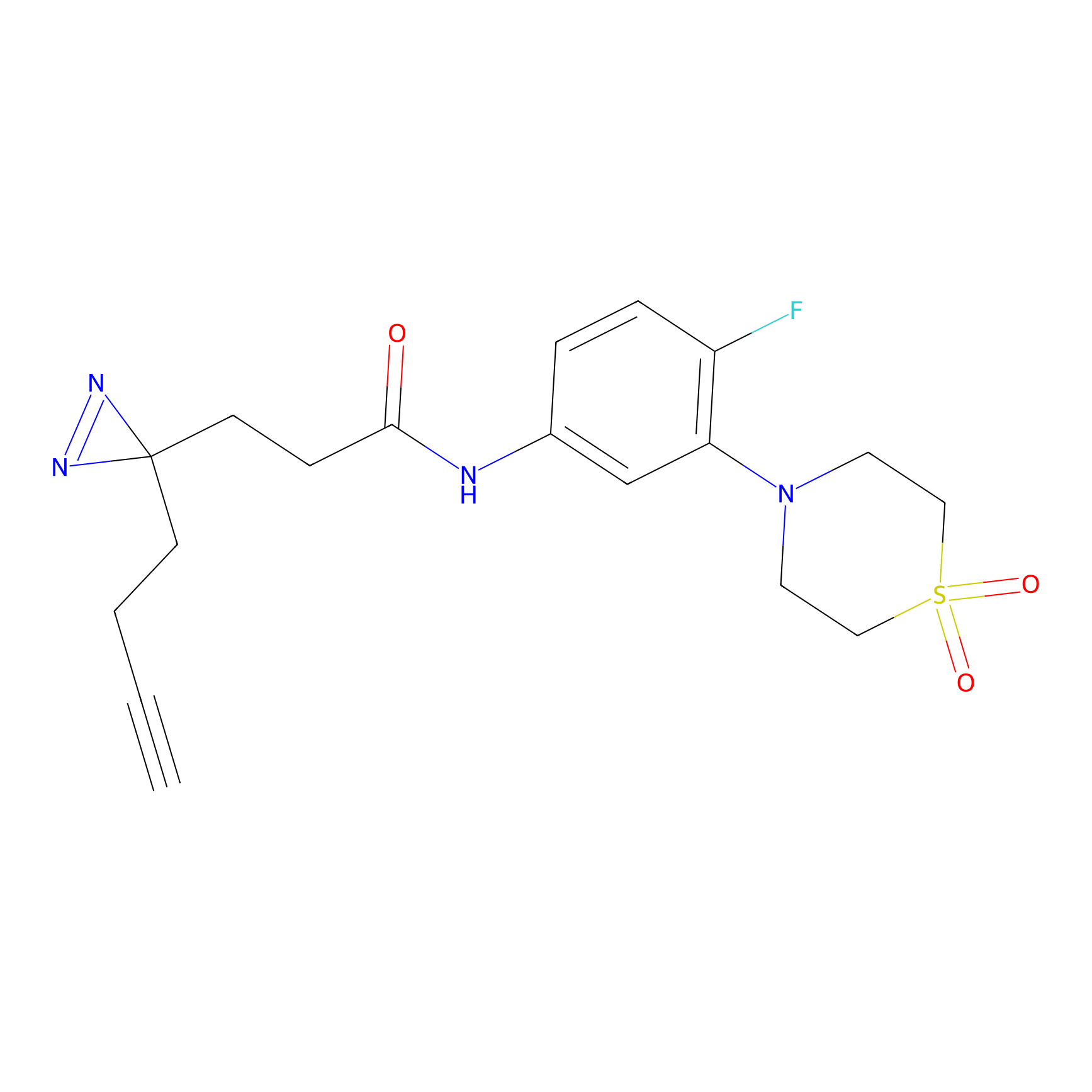
CHEMBL5175495 Probe Info |
 |
10.44 | LDD0196 | [3] | |
|
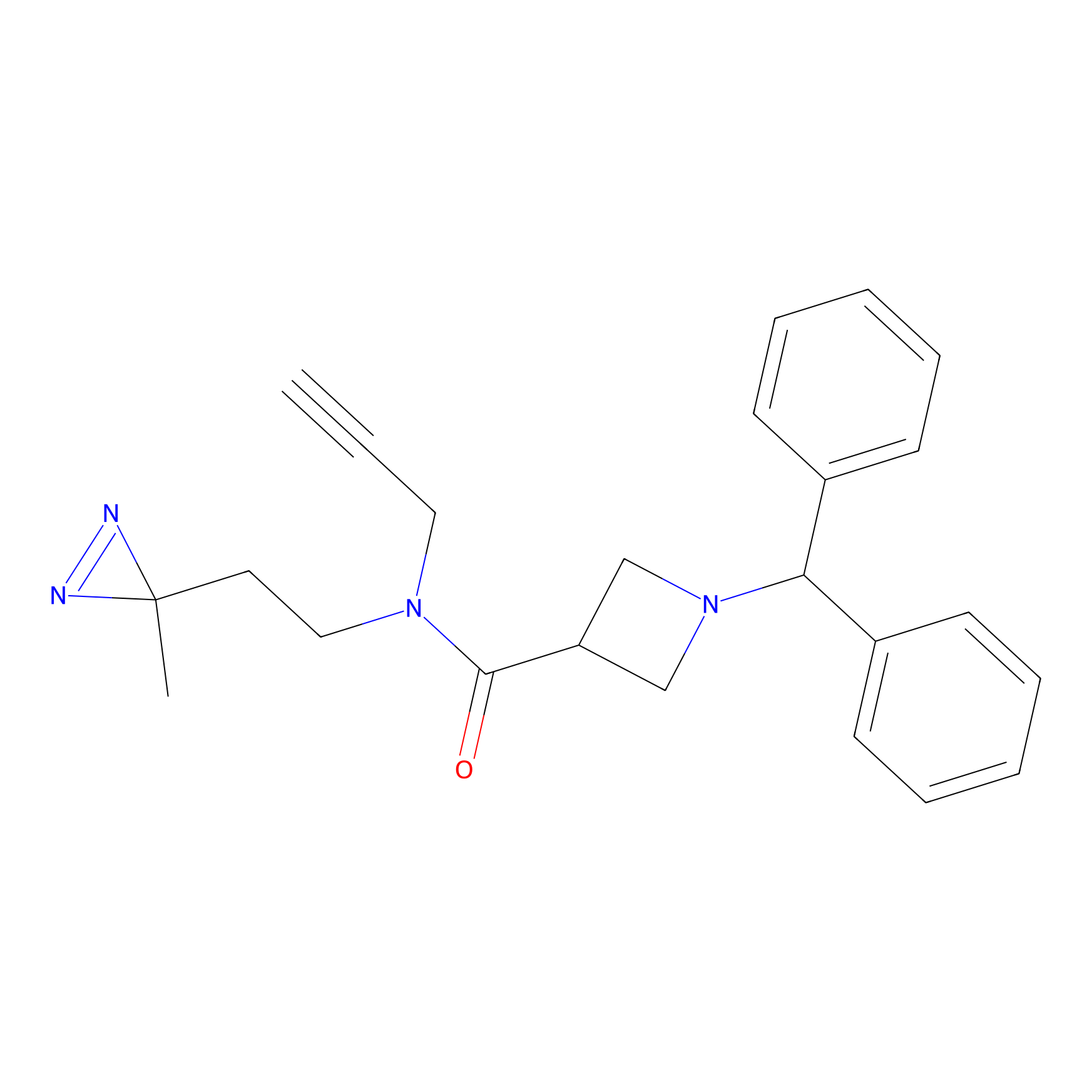
TG42 Probe Info |
 |
4.34 | LDD0326 | [4] | |
|
TH211 Probe Info |
 |
Y113(11.67); Y164(20.00) | LDD0260 | [5] | |
|
FBP2 Probe Info |
 |
4.04 | LDD0317 | [2] | |
|
TH216 Probe Info |
 |
Y113(13.34) | LDD0259 | [5] | |
|
STPyne Probe Info |
 |
K102(7.65); K105(6.03); K96(10.00) | LDD0277 | [6] | |
|
BTD Probe Info |
 |
C129(3.25) | LDD1699 | [7] | |
|
AZ-9 Probe Info |
 |
D131(10.00); E110(10.00) | LDD2209 | [8] | |
|
ONAyne Probe Info |
 |
K102(0.00); K105(0.00) | LDD0273 | [6] | |
|
OPA-S-S-alkyne Probe Info |
 |
K105(2.63); K102(5.04) | LDD3494 | [9] | |
|
Probe 1 Probe Info |
 |
Y113(38.51) | LDD3495 | [10] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [11] | |
|
THZ1-DTB Probe Info |
 |
C129(1.12); C129(1.06) | LDD0460 | [11] | |
|
AHL-Pu-1 Probe Info |
 |
C129(3.56) | LDD0169 | [12] | |
|
HPAP Probe Info |
 |
3.59 | LDD0062 | [13] | |
|
HHS-475 Probe Info |
 |
Y113(0.08) | LDD0264 | [14] | |
|
HHS-465 Probe Info |
 |
Y113(10.00) | LDD2237 | [15] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0221 | [16] | |
|
DBIA Probe Info |
 |
C129(0.98) | LDD1507 | [17] | |
|
ATP probe Probe Info |
 |
K105(0.00); K102(0.00); K169(0.00) | LDD0199 | [18] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [19] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [19] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [19] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [20] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [20] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [21] | |
|
WYneC Probe Info |
 |
N.A. | LDD0014 | [22] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [22] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [23] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [22] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [22] | |
|
OSF Probe Info |
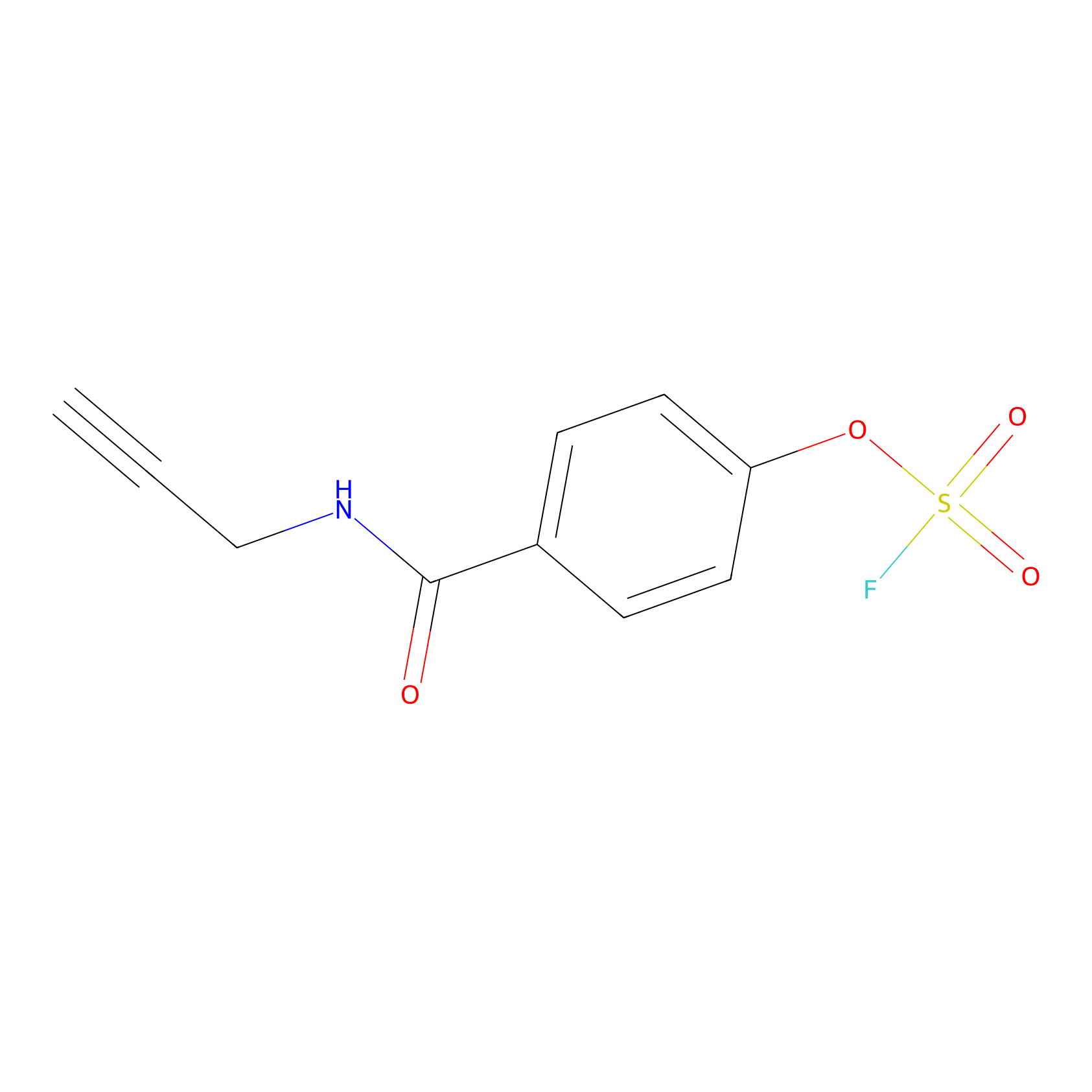 |
N.A. | LDD0029 | [24] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [25] | |
|
SF Probe Info |
 |
Y113(0.00); Y164(0.00); K102(0.00) | LDD0028 | [24] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [26] | |
|
1c-yne Probe Info |
 |
K105(0.00); K163(0.00) | LDD0228 | [23] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [16] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [27] | |
|
AOyne Probe Info |
 |
14.50 | LDD0443 | [28] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [21] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [29] | |
|
HHS-482 Probe Info |
 |
Y113(0.58) | LDD2239 | [15] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C129(0.80) | LDD2142 | [7] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C129(0.82) | LDD2112 | [7] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C129(0.73) | LDD2095 | [7] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C129(1.33) | LDD2152 | [7] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C129(0.84) | LDD2103 | [7] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C129(0.75) | LDD2132 | [7] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C129(3.56) | LDD0169 | [12] |
| LDCM0214 | AC1 | HEK-293T | C129(0.98) | LDD1507 | [17] |
| LDCM0215 | AC10 | HEK-293T | C129(0.91) | LDD1508 | [17] |
| LDCM0226 | AC11 | HEK-293T | C129(0.98) | LDD1509 | [17] |
| LDCM0237 | AC12 | HEK-293T | C129(0.95) | LDD1510 | [17] |
| LDCM0259 | AC14 | HEK-293T | C129(0.93) | LDD1512 | [17] |
| LDCM0270 | AC15 | HEK-293T | C129(0.96) | LDD1513 | [17] |
| LDCM0276 | AC17 | HEK-293T | C129(1.09) | LDD1515 | [17] |
| LDCM0277 | AC18 | HEK-293T | C129(1.01) | LDD1516 | [17] |
| LDCM0278 | AC19 | HEK-293T | C129(0.93) | LDD1517 | [17] |
| LDCM0279 | AC2 | HEK-293T | C129(0.96) | LDD1518 | [17] |
| LDCM0280 | AC20 | HEK-293T | C129(1.05) | LDD1519 | [17] |
| LDCM0281 | AC21 | HEK-293T | C129(1.01) | LDD1520 | [17] |
| LDCM0282 | AC22 | HEK-293T | C129(1.02) | LDD1521 | [17] |
| LDCM0283 | AC23 | HEK-293T | C129(1.07) | LDD1522 | [17] |
| LDCM0284 | AC24 | HEK-293T | C129(1.04) | LDD1523 | [17] |
| LDCM0285 | AC25 | HEK-293T | C129(1.00) | LDD1524 | [17] |
| LDCM0286 | AC26 | HEK-293T | C129(0.89) | LDD1525 | [17] |
| LDCM0287 | AC27 | HEK-293T | C129(0.96) | LDD1526 | [17] |
| LDCM0288 | AC28 | HEK-293T | C129(0.93) | LDD1527 | [17] |
| LDCM0289 | AC29 | HEK-293T | C129(0.91) | LDD1528 | [17] |
| LDCM0290 | AC3 | HEK-293T | C129(0.95) | LDD1529 | [17] |
| LDCM0291 | AC30 | HEK-293T | C129(0.88) | LDD1530 | [17] |
| LDCM0292 | AC31 | HEK-293T | C129(0.95) | LDD1531 | [17] |
| LDCM0293 | AC32 | HEK-293T | C129(1.02) | LDD1532 | [17] |
| LDCM0294 | AC33 | HEK-293T | C129(1.11) | LDD1533 | [17] |
| LDCM0295 | AC34 | HEK-293T | C129(0.99) | LDD1534 | [17] |
| LDCM0296 | AC35 | HEK-293T | C129(1.04) | LDD1535 | [17] |
| LDCM0297 | AC36 | HEK-293T | C129(1.07) | LDD1536 | [17] |
| LDCM0298 | AC37 | HEK-293T | C129(0.88) | LDD1537 | [17] |
| LDCM0299 | AC38 | HEK-293T | C129(0.95) | LDD1538 | [17] |
| LDCM0300 | AC39 | HEK-293T | C129(0.99) | LDD1539 | [17] |
| LDCM0301 | AC4 | HEK-293T | C129(0.91) | LDD1540 | [17] |
| LDCM0302 | AC40 | HEK-293T | C129(0.98) | LDD1541 | [17] |
| LDCM0303 | AC41 | HEK-293T | C129(1.09) | LDD1542 | [17] |
| LDCM0304 | AC42 | HEK-293T | C129(0.88) | LDD1543 | [17] |
| LDCM0305 | AC43 | HEK-293T | C129(1.04) | LDD1544 | [17] |
| LDCM0306 | AC44 | HEK-293T | C129(1.01) | LDD1545 | [17] |
| LDCM0307 | AC45 | HEK-293T | C129(0.89) | LDD1546 | [17] |
| LDCM0308 | AC46 | HEK-293T | C129(0.90) | LDD1547 | [17] |
| LDCM0309 | AC47 | HEK-293T | C129(0.94) | LDD1548 | [17] |
| LDCM0310 | AC48 | HEK-293T | C129(1.04) | LDD1549 | [17] |
| LDCM0311 | AC49 | HEK-293T | C129(1.05) | LDD1550 | [17] |
| LDCM0312 | AC5 | HEK-293T | C129(1.00) | LDD1551 | [17] |
| LDCM0313 | AC50 | HEK-293T | C129(0.93) | LDD1552 | [17] |
| LDCM0314 | AC51 | HEK-293T | C129(1.01) | LDD1553 | [17] |
| LDCM0315 | AC52 | HEK-293T | C129(1.10) | LDD1554 | [17] |
| LDCM0316 | AC53 | HEK-293T | C129(1.06) | LDD1555 | [17] |
| LDCM0317 | AC54 | HEK-293T | C129(1.03) | LDD1556 | [17] |
| LDCM0318 | AC55 | HEK-293T | C129(1.06) | LDD1557 | [17] |
| LDCM0319 | AC56 | HEK-293T | C129(1.04) | LDD1558 | [17] |
| LDCM0320 | AC57 | HEK-293T | C129(1.01) | LDD1559 | [17] |
| LDCM0321 | AC58 | HEK-293T | C129(0.98) | LDD1560 | [17] |
| LDCM0322 | AC59 | HEK-293T | C129(1.02) | LDD1561 | [17] |
| LDCM0323 | AC6 | HEK-293T | C129(0.89) | LDD1562 | [17] |
| LDCM0324 | AC60 | HEK-293T | C129(1.00) | LDD1563 | [17] |
| LDCM0325 | AC61 | HEK-293T | C129(0.92) | LDD1564 | [17] |
| LDCM0326 | AC62 | HEK-293T | C129(0.99) | LDD1565 | [17] |
| LDCM0327 | AC63 | HEK-293T | C129(0.98) | LDD1566 | [17] |
| LDCM0328 | AC64 | HEK-293T | C129(0.97) | LDD1567 | [17] |
| LDCM0334 | AC7 | HEK-293T | C129(0.93) | LDD1568 | [17] |
| LDCM0345 | AC8 | HEK-293T | C129(0.91) | LDD1569 | [17] |
| LDCM0545 | Acetamide | MDA-MB-231 | C129(0.95) | LDD2138 | [7] |
| LDCM0248 | AKOS034007472 | HEK-293T | C129(0.98) | LDD1511 | [17] |
| LDCM0356 | AKOS034007680 | HEK-293T | C129(0.97) | LDD1570 | [17] |
| LDCM0275 | AKOS034007705 | HEK-293T | C129(0.96) | LDD1514 | [17] |
| LDCM0156 | Aniline | NCI-H1299 | 11.98 | LDD0403 | [1] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C129(0.93) | LDD2091 | [7] |
| LDCM0108 | Chloroacetamide | HeLa | C129(0.00); H166(0.00); H165(0.00) | LDD0222 | [16] |
| LDCM0632 | CL-Sc | Hep-G2 | C129(7.29); C129(0.88) | LDD2227 | [21] |
| LDCM0367 | CL1 | HEK-293T | C129(0.89) | LDD1571 | [17] |
| LDCM0368 | CL10 | HEK-293T | C129(0.68) | LDD1572 | [17] |
| LDCM0369 | CL100 | HEK-293T | C129(1.01) | LDD1573 | [17] |
| LDCM0370 | CL101 | HEK-293T | C129(0.97) | LDD1574 | [17] |
| LDCM0371 | CL102 | HEK-293T | C129(0.86) | LDD1575 | [17] |
| LDCM0372 | CL103 | HEK-293T | C129(1.06) | LDD1576 | [17] |
| LDCM0373 | CL104 | HEK-293T | C129(0.92) | LDD1577 | [17] |
| LDCM0374 | CL105 | HEK-293T | C129(1.02) | LDD1578 | [17] |
| LDCM0375 | CL106 | HEK-293T | C129(1.00) | LDD1579 | [17] |
| LDCM0376 | CL107 | HEK-293T | C129(1.10) | LDD1580 | [17] |
| LDCM0377 | CL108 | HEK-293T | C129(1.05) | LDD1581 | [17] |
| LDCM0378 | CL109 | HEK-293T | C129(0.98) | LDD1582 | [17] |
| LDCM0379 | CL11 | HEK-293T | C129(0.76) | LDD1583 | [17] |
| LDCM0380 | CL110 | HEK-293T | C129(0.83) | LDD1584 | [17] |
| LDCM0381 | CL111 | HEK-293T | C129(0.92) | LDD1585 | [17] |
| LDCM0382 | CL112 | HEK-293T | C129(0.86) | LDD1586 | [17] |
| LDCM0383 | CL113 | HEK-293T | C129(1.04) | LDD1587 | [17] |
| LDCM0384 | CL114 | HEK-293T | C129(0.84) | LDD1588 | [17] |
| LDCM0385 | CL115 | HEK-293T | C129(0.92) | LDD1589 | [17] |
| LDCM0386 | CL116 | HEK-293T | C129(1.07) | LDD1590 | [17] |
| LDCM0387 | CL117 | HEK-293T | C129(1.07) | LDD1591 | [17] |
| LDCM0388 | CL118 | HEK-293T | C129(0.93) | LDD1592 | [17] |
| LDCM0389 | CL119 | HEK-293T | C129(0.93) | LDD1593 | [17] |
| LDCM0390 | CL12 | HEK-293T | C129(0.83) | LDD1594 | [17] |
| LDCM0391 | CL120 | HEK-293T | C129(0.98) | LDD1595 | [17] |
| LDCM0392 | CL121 | HEK-293T | C129(1.09) | LDD1596 | [17] |
| LDCM0393 | CL122 | HEK-293T | C129(1.06) | LDD1597 | [17] |
| LDCM0394 | CL123 | HEK-293T | C129(0.96) | LDD1598 | [17] |
| LDCM0395 | CL124 | HEK-293T | C129(0.95) | LDD1599 | [17] |
| LDCM0396 | CL125 | HEK-293T | C129(0.98) | LDD1600 | [17] |
| LDCM0397 | CL126 | HEK-293T | C129(1.02) | LDD1601 | [17] |
| LDCM0398 | CL127 | HEK-293T | C129(1.04) | LDD1602 | [17] |
| LDCM0399 | CL128 | HEK-293T | C129(1.05) | LDD1603 | [17] |
| LDCM0400 | CL13 | HEK-293T | C129(0.94) | LDD1604 | [17] |
| LDCM0401 | CL14 | HEK-293T | C129(0.93) | LDD1605 | [17] |
| LDCM0402 | CL15 | HEK-293T | C129(0.89) | LDD1606 | [17] |
| LDCM0403 | CL16 | HEK-293T | C129(0.97) | LDD1607 | [17] |
| LDCM0404 | CL17 | HEK-293T | C129(0.55) | LDD1608 | [17] |
| LDCM0405 | CL18 | HEK-293T | C129(0.84) | LDD1609 | [17] |
| LDCM0406 | CL19 | HEK-293T | C129(0.96) | LDD1610 | [17] |
| LDCM0407 | CL2 | HEK-293T | C129(0.93) | LDD1611 | [17] |
| LDCM0408 | CL20 | HEK-293T | C129(0.97) | LDD1612 | [17] |
| LDCM0409 | CL21 | HEK-293T | C129(0.82) | LDD1613 | [17] |
| LDCM0410 | CL22 | HEK-293T | C129(0.80) | LDD1614 | [17] |
| LDCM0411 | CL23 | HEK-293T | C129(0.86) | LDD1615 | [17] |
| LDCM0412 | CL24 | HEK-293T | C129(0.86) | LDD1616 | [17] |
| LDCM0413 | CL25 | HEK-293T | C129(1.09) | LDD1617 | [17] |
| LDCM0414 | CL26 | HEK-293T | C129(1.12) | LDD1618 | [17] |
| LDCM0415 | CL27 | HEK-293T | C129(1.06) | LDD1619 | [17] |
| LDCM0416 | CL28 | HEK-293T | C129(0.97) | LDD1620 | [17] |
| LDCM0417 | CL29 | HEK-293T | C129(0.96) | LDD1621 | [17] |
| LDCM0418 | CL3 | HEK-293T | C129(0.84) | LDD1622 | [17] |
| LDCM0419 | CL30 | HEK-293T | C129(0.93) | LDD1623 | [17] |
| LDCM0420 | CL31 | HEK-293T | C129(0.92) | LDD1624 | [17] |
| LDCM0421 | CL32 | HEK-293T | C129(0.91) | LDD1625 | [17] |
| LDCM0422 | CL33 | HEK-293T | C129(0.78) | LDD1626 | [17] |
| LDCM0423 | CL34 | HEK-293T | C129(0.87) | LDD1627 | [17] |
| LDCM0424 | CL35 | HEK-293T | C129(0.91) | LDD1628 | [17] |
| LDCM0425 | CL36 | HEK-293T | C129(0.91) | LDD1629 | [17] |
| LDCM0426 | CL37 | HEK-293T | C129(1.12) | LDD1630 | [17] |
| LDCM0428 | CL39 | HEK-293T | C129(1.17) | LDD1632 | [17] |
| LDCM0429 | CL4 | HEK-293T | C129(0.90) | LDD1633 | [17] |
| LDCM0430 | CL40 | HEK-293T | C129(1.11) | LDD1634 | [17] |
| LDCM0431 | CL41 | HEK-293T | C129(0.93) | LDD1635 | [17] |
| LDCM0432 | CL42 | HEK-293T | C129(0.93) | LDD1636 | [17] |
| LDCM0433 | CL43 | HEK-293T | C129(0.97) | LDD1637 | [17] |
| LDCM0434 | CL44 | HEK-293T | C129(0.82) | LDD1638 | [17] |
| LDCM0435 | CL45 | HEK-293T | C129(0.89) | LDD1639 | [17] |
| LDCM0436 | CL46 | HEK-293T | C129(0.88) | LDD1640 | [17] |
| LDCM0437 | CL47 | HEK-293T | C129(0.89) | LDD1641 | [17] |
| LDCM0438 | CL48 | HEK-293T | C129(1.03) | LDD1642 | [17] |
| LDCM0439 | CL49 | HEK-293T | C129(0.99) | LDD1643 | [17] |
| LDCM0440 | CL5 | HEK-293T | C129(0.85) | LDD1644 | [17] |
| LDCM0441 | CL50 | HEK-293T | C129(1.07) | LDD1645 | [17] |
| LDCM0443 | CL52 | HEK-293T | C129(0.92) | LDD1646 | [17] |
| LDCM0444 | CL53 | HEK-293T | C129(0.82) | LDD1647 | [17] |
| LDCM0445 | CL54 | HEK-293T | C129(0.79) | LDD1648 | [17] |
| LDCM0446 | CL55 | HEK-293T | C129(0.89) | LDD1649 | [17] |
| LDCM0447 | CL56 | HEK-293T | C129(0.85) | LDD1650 | [17] |
| LDCM0448 | CL57 | HEK-293T | C129(0.75) | LDD1651 | [17] |
| LDCM0449 | CL58 | HEK-293T | C129(0.80) | LDD1652 | [17] |
| LDCM0450 | CL59 | HEK-293T | C129(0.84) | LDD1653 | [17] |
| LDCM0451 | CL6 | HEK-293T | C129(0.71) | LDD1654 | [17] |
| LDCM0452 | CL60 | HEK-293T | C129(0.98) | LDD1655 | [17] |
| LDCM0453 | CL61 | HEK-293T | C129(1.11) | LDD1656 | [17] |
| LDCM0454 | CL62 | HEK-293T | C129(1.03) | LDD1657 | [17] |
| LDCM0455 | CL63 | HEK-293T | C129(1.06) | LDD1658 | [17] |
| LDCM0456 | CL64 | HEK-293T | C129(0.87) | LDD1659 | [17] |
| LDCM0457 | CL65 | HEK-293T | C129(1.05) | LDD1660 | [17] |
| LDCM0458 | CL66 | HEK-293T | C129(0.97) | LDD1661 | [17] |
| LDCM0459 | CL67 | HEK-293T | C129(0.90) | LDD1662 | [17] |
| LDCM0460 | CL68 | HEK-293T | C129(0.92) | LDD1663 | [17] |
| LDCM0461 | CL69 | HEK-293T | C129(0.91) | LDD1664 | [17] |
| LDCM0462 | CL7 | HEK-293T | C129(0.90) | LDD1665 | [17] |
| LDCM0463 | CL70 | HEK-293T | C129(0.91) | LDD1666 | [17] |
| LDCM0464 | CL71 | HEK-293T | C129(0.84) | LDD1667 | [17] |
| LDCM0465 | CL72 | HEK-293T | C129(1.11) | LDD1668 | [17] |
| LDCM0466 | CL73 | HEK-293T | C129(0.94) | LDD1669 | [17] |
| LDCM0467 | CL74 | HEK-293T | C129(0.97) | LDD1670 | [17] |
| LDCM0469 | CL76 | HEK-293T | C129(0.96) | LDD1672 | [17] |
| LDCM0470 | CL77 | HEK-293T | C129(0.78) | LDD1673 | [17] |
| LDCM0471 | CL78 | HEK-293T | C129(0.84) | LDD1674 | [17] |
| LDCM0472 | CL79 | HEK-293T | C129(0.89) | LDD1675 | [17] |
| LDCM0473 | CL8 | HEK-293T | C129(0.62) | LDD1676 | [17] |
| LDCM0474 | CL80 | HEK-293T | C129(0.84) | LDD1677 | [17] |
| LDCM0475 | CL81 | HEK-293T | C129(0.93) | LDD1678 | [17] |
| LDCM0476 | CL82 | HEK-293T | C129(0.78) | LDD1679 | [17] |
| LDCM0477 | CL83 | HEK-293T | C129(0.79) | LDD1680 | [17] |
| LDCM0478 | CL84 | HEK-293T | C129(0.81) | LDD1681 | [17] |
| LDCM0479 | CL85 | HEK-293T | C129(1.07) | LDD1682 | [17] |
| LDCM0480 | CL86 | HEK-293T | C129(1.07) | LDD1683 | [17] |
| LDCM0481 | CL87 | HEK-293T | C129(1.03) | LDD1684 | [17] |
| LDCM0482 | CL88 | HEK-293T | C129(1.02) | LDD1685 | [17] |
| LDCM0483 | CL89 | HEK-293T | C129(1.00) | LDD1686 | [17] |
| LDCM0484 | CL9 | HEK-293T | C129(0.72) | LDD1687 | [17] |
| LDCM0485 | CL90 | HEK-293T | C129(0.78) | LDD1688 | [17] |
| LDCM0486 | CL91 | HEK-293T | C129(0.96) | LDD1689 | [17] |
| LDCM0487 | CL92 | HEK-293T | C129(0.96) | LDD1690 | [17] |
| LDCM0488 | CL93 | HEK-293T | C129(0.80) | LDD1691 | [17] |
| LDCM0489 | CL94 | HEK-293T | C129(0.84) | LDD1692 | [17] |
| LDCM0490 | CL95 | HEK-293T | C129(0.75) | LDD1693 | [17] |
| LDCM0491 | CL96 | HEK-293T | C129(0.83) | LDD1694 | [17] |
| LDCM0492 | CL97 | HEK-293T | C129(0.93) | LDD1695 | [17] |
| LDCM0493 | CL98 | HEK-293T | C129(0.91) | LDD1696 | [17] |
| LDCM0494 | CL99 | HEK-293T | C129(0.97) | LDD1697 | [17] |
| LDCM0495 | E2913 | HEK-293T | C129(1.13) | LDD1698 | [17] |
| LDCM0468 | Fragment33 | HEK-293T | C129(0.93) | LDD1671 | [17] |
| LDCM0427 | Fragment51 | HEK-293T | C129(1.08) | LDD1631 | [17] |
| LDCM0116 | HHS-0101 | DM93 | Y113(0.08) | LDD0264 | [14] |
| LDCM0117 | HHS-0201 | DM93 | Y113(1.23) | LDD0265 | [14] |
| LDCM0118 | HHS-0301 | DM93 | Y113(0.24) | LDD0266 | [14] |
| LDCM0119 | HHS-0401 | DM93 | Y113(0.11) | LDD0267 | [14] |
| LDCM0120 | HHS-0701 | DM93 | Y113(0.10) | LDD0268 | [14] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [16] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [11] |
| LDCM0023 | KB03 | MDA-MB-231 | C129(5.43) | LDD1701 | [7] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C129(0.75) | LDD2121 | [7] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0226 | [16] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C129(1.11) | LDD2089 | [7] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C129(1.72) | LDD2093 | [7] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C129(1.42) | LDD2094 | [7] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C129(1.17) | LDD2097 | [7] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C129(1.14) | LDD2101 | [7] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C129(0.92) | LDD2104 | [7] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C129(0.41) | LDD2107 | [7] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C129(0.71) | LDD2108 | [7] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C129(1.48) | LDD2111 | [7] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C129(0.71) | LDD2114 | [7] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C129(0.61) | LDD2115 | [7] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C129(1.26) | LDD2129 | [7] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C129(0.82) | LDD2133 | [7] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C129(1.43) | LDD2135 | [7] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C129(2.55) | LDD1700 | [7] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C129(0.71) | LDD2141 | [7] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C129(2.97) | LDD2144 | [7] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C129(1.87) | LDD2145 | [7] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C129(0.83) | LDD2150 | [7] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C129(1.95) | LDD2153 | [7] |
| LDCM0014 | Panhematin | HEK-293T | 3.59 | LDD0062 | [13] |
| LDCM0021 | THZ1 | HeLa S3 | C129(1.12); C129(1.06) | LDD0460 | [11] |
| LDCM0111 | W14 | Hep-G2 | C129(3.16) | LDD0238 | [27] |
| LDCM0113 | W17 | Hep-G2 | C129(2.46) | LDD0240 | [27] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Endothelial zinc finger protein induced by tumor necrosis factor alpha (ZNF71) | Krueppel C2H2-type zinc-finger protein family | Q9NQZ8 | |||
Other
References