Details of the Target
General Information of Target
| Target ID | LDTP04634 | |||||
|---|---|---|---|---|---|---|
| Target Name | Alpha-N-acetylglucosaminidase (NAGLU) | |||||
| Gene Name | NAGLU | |||||
| Gene ID | 4669 | |||||
| Synonyms |
UFHSD1; Alpha-N-acetylglucosaminidase; EC 3.2.1.50; N-acetyl-alpha-glucosaminidase; NAG) [Cleaved into: Alpha-N-acetylglucosaminidase 82 kDa form; Alpha-N-acetylglucosaminidase 77 kDa form] |
|||||
| 3D Structure | ||||||
| Sequence |
MEAVAVAAAVGVLLLAGAGGAAGDEAREAAAVRALVARLLGPGPAADFSVSVERALAAKP
GLDTYSLGGGGAARVRVRGSTGVAAAAGLHRYLRDFCGCHVAWSGSQLRLPRPLPAVPGE LTEATPNRYRYYQNVCTQSYSFVWWDWARWEREIDWMALNGINLALAWSGQEAIWQRVYL ALGLTQAEINEFFTGPAFLAWGRMGNLHTWDGPLPPSWHIKQLYLQHRVLDQMRSFGMTP VLPAFAGHVPEAVTRVFPQVNVTKMGSWGHFNCSYSCSFLLAPEDPIFPIIGSLFLRELI KEFGTDHIYGADTFNEMQPPSSEPSYLAAATTAVYEAMTAVDTEAVWLLQGWLFQHQPQF WGPAQIRAVLGAVPRGRLLVLDLFAESQPVYTRTASFQGQPFIWCMLHNFGGNHGLFGAL EAVNGGPEAARLFPNSTMVGTGMAPEGISQNEVVYSLMAELGWRKDPVPDLAAWVTSFAA RRYGVSHPDAGAAWRLLLRSVYNCSGEACRGHNRSPLVRRPSLQMNTSIWYNRSDVFEAW RLLLTSAPSLATSPAFRYDLLDLTRQAVQELVSLYYEEARSAYLSKELASLLRAGGVLAY ELLPALDEVLASDSRFLLGSWLEQARAAAVSEAEADFYEQNSRYQLTLWGPEGNILDYAN KQLAGLVANYYTPRWRLFLEALVDSVAQGIPFQQHQFDKNVFQLEQAFVLSKQRYPSQPR GDTVDLAKKIFLKYYPRWVAGSW |
|||||
| Target Type |
Successful
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Glycosyl hydrolase 89 family
|
|||||
| Subcellular location |
Lysosome
|
|||||
| Function | Involved in the degradation of heparan sulfate. | |||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
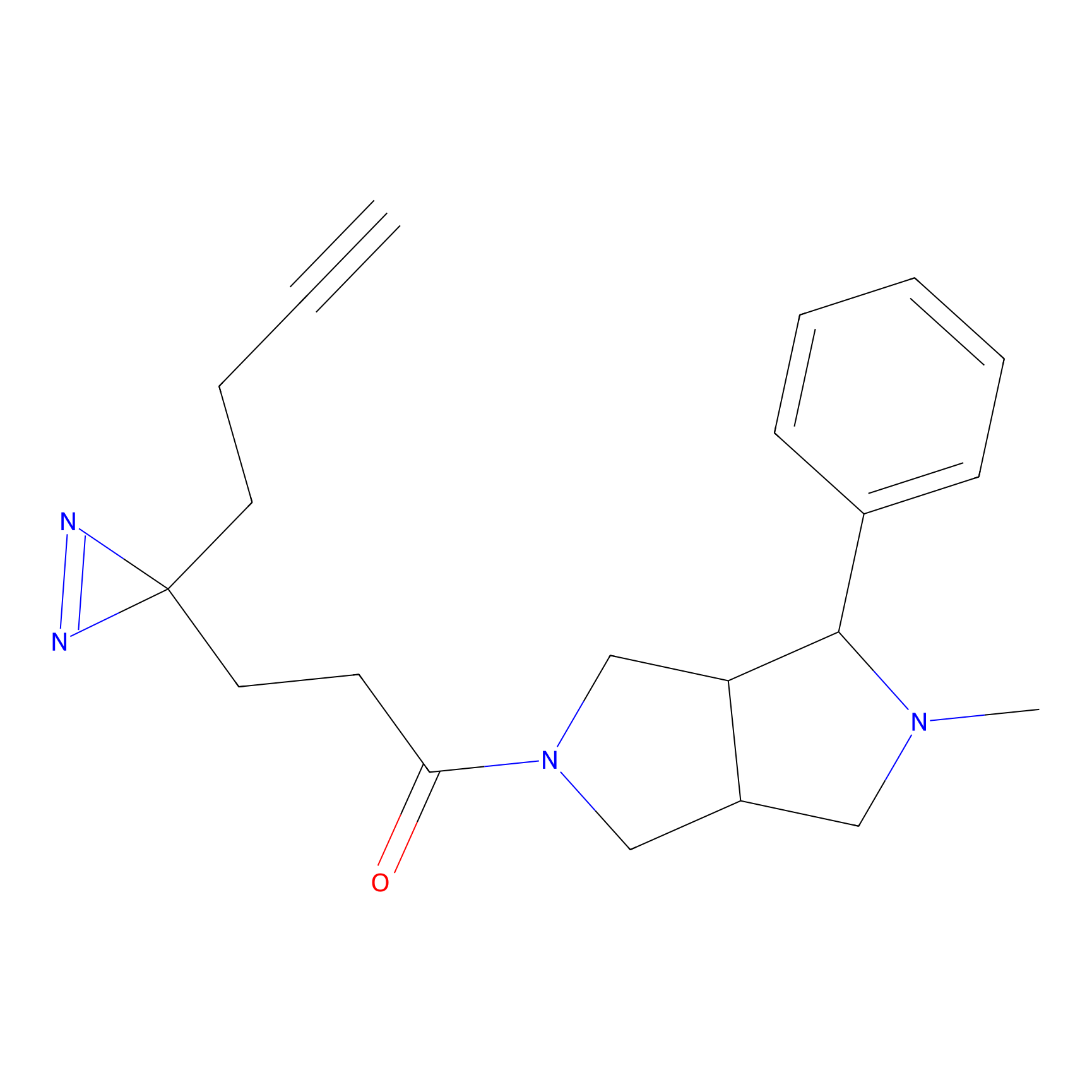
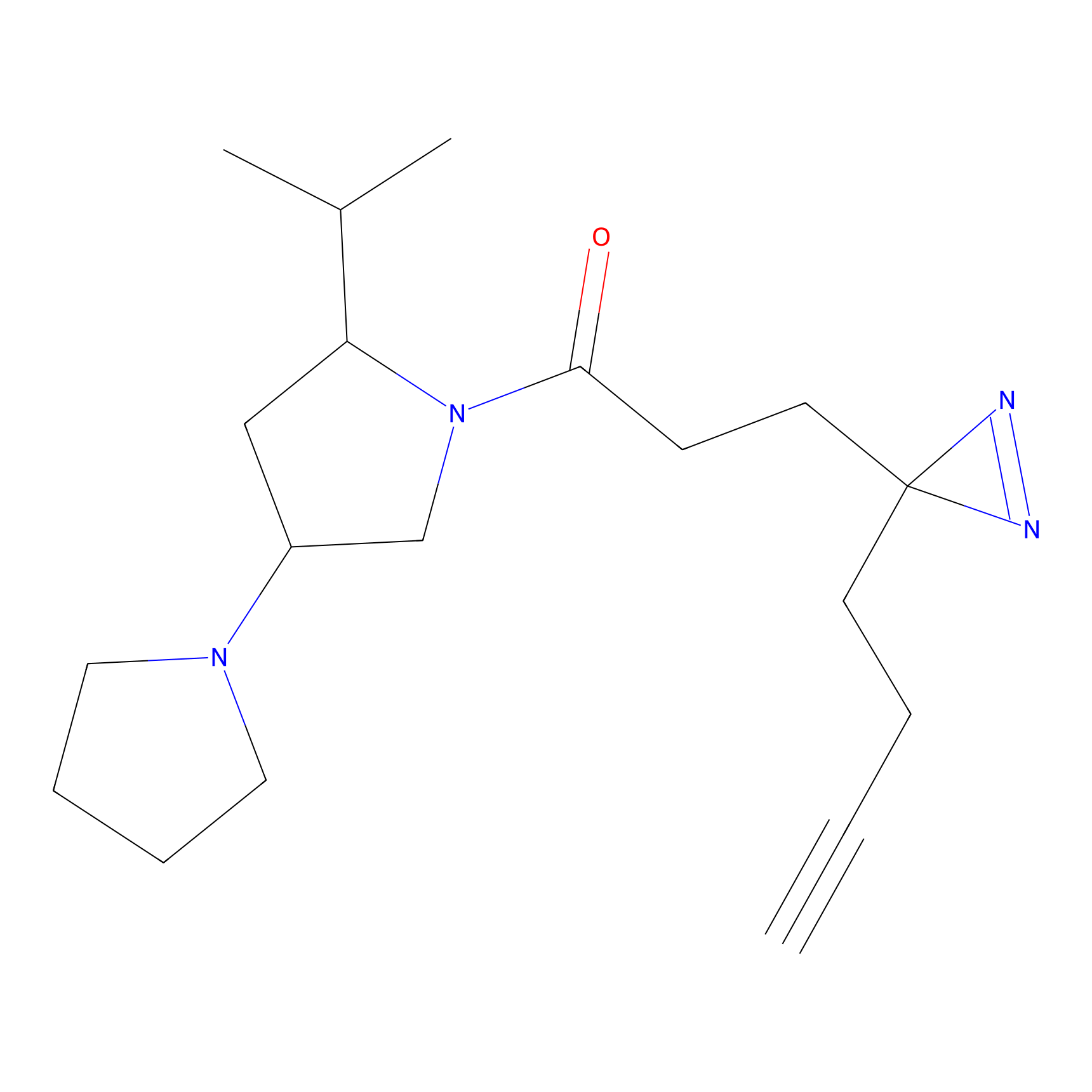
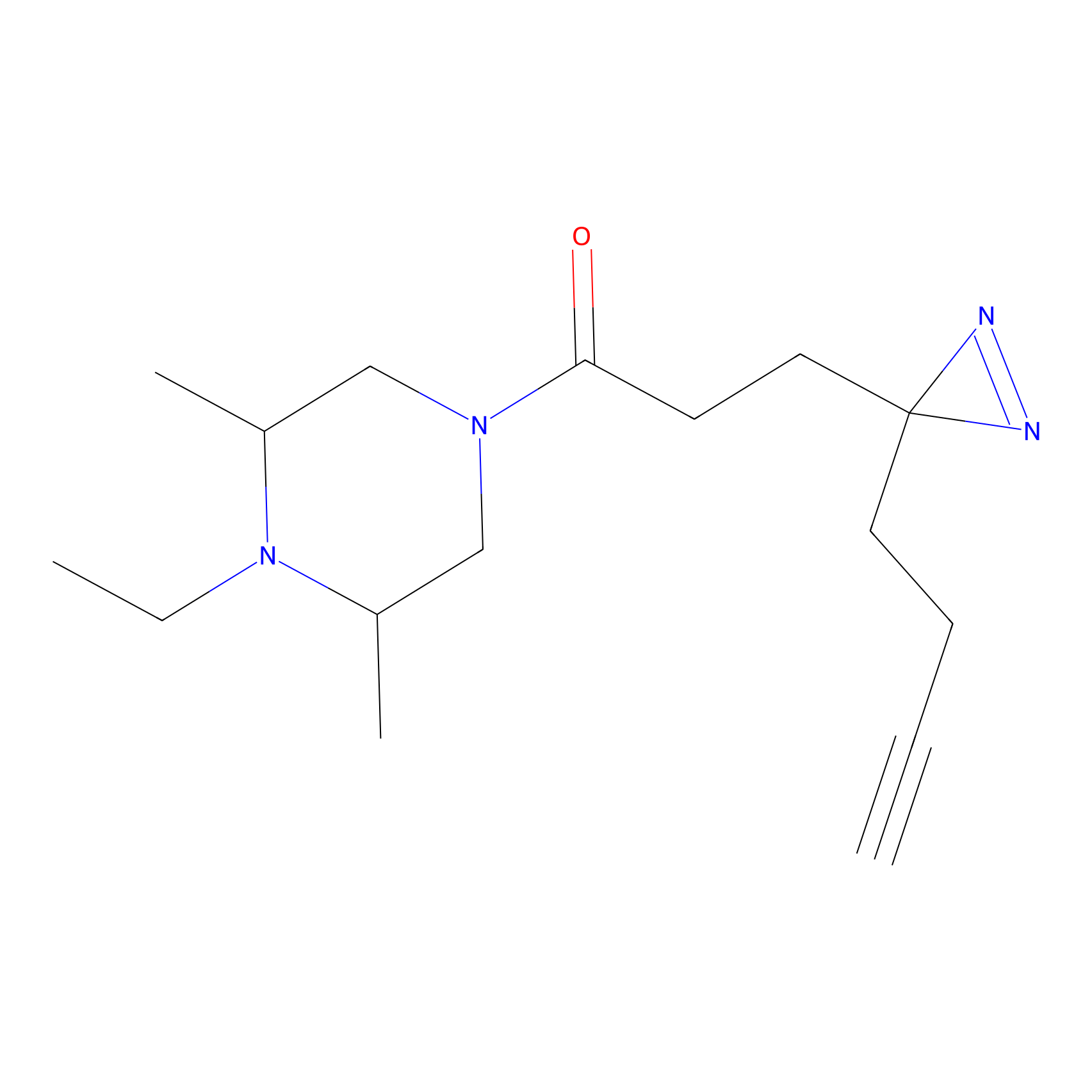
STPyne Probe Info |
 |
K728(2.13) | LDD0277 | [1] | |
|
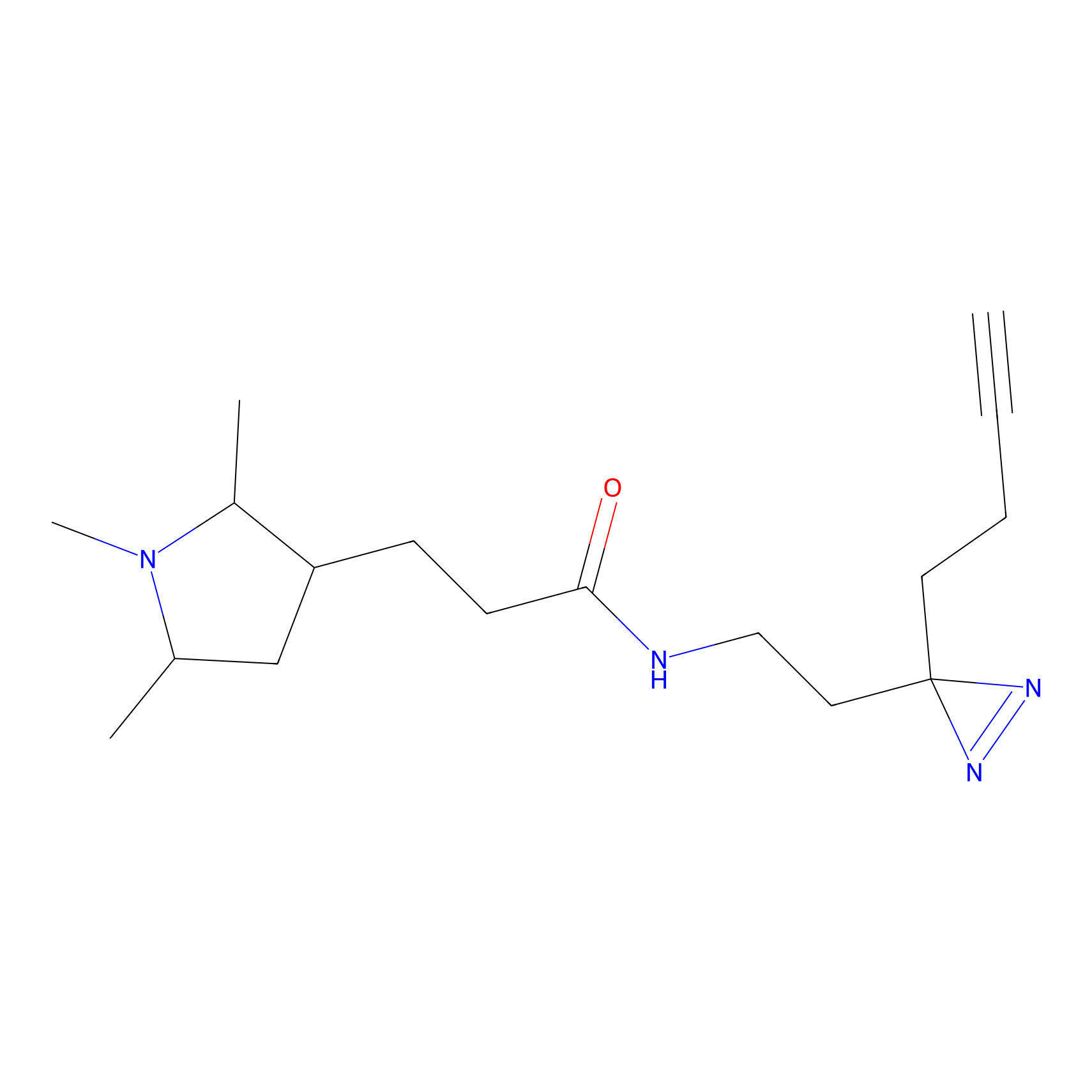
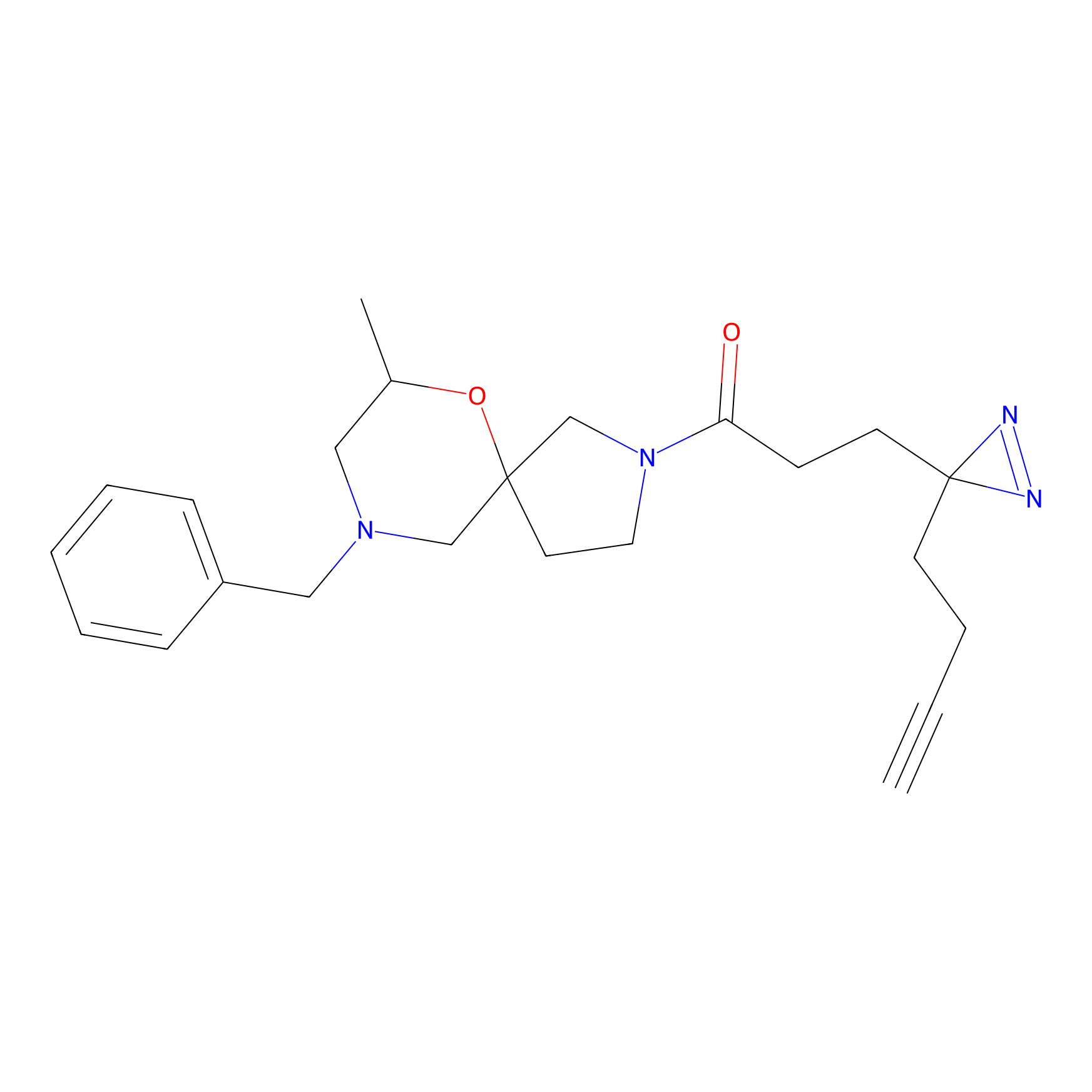
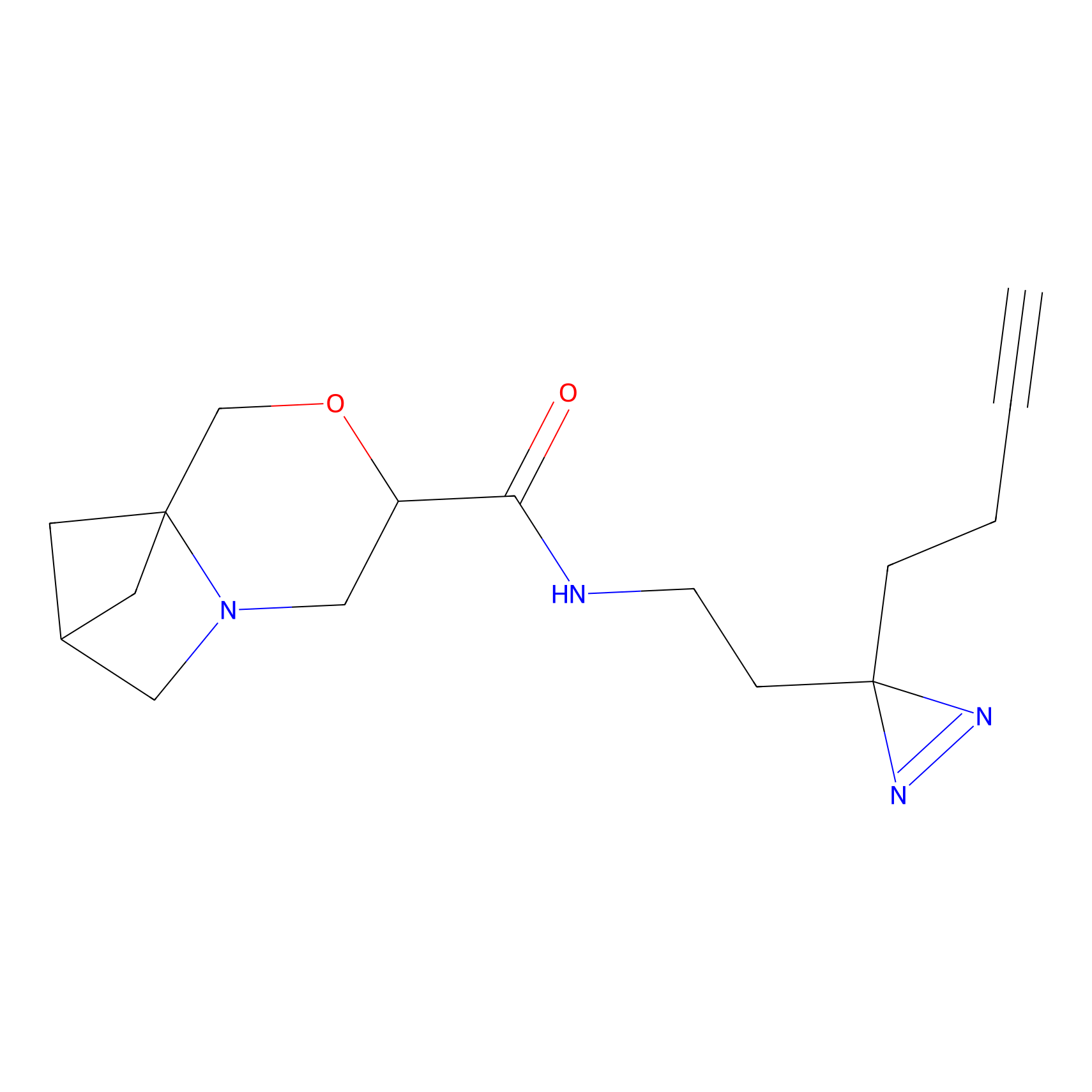
Probe 1 Probe Info |
 |
Y638(23.26) | LDD3495 | [2] | |
|
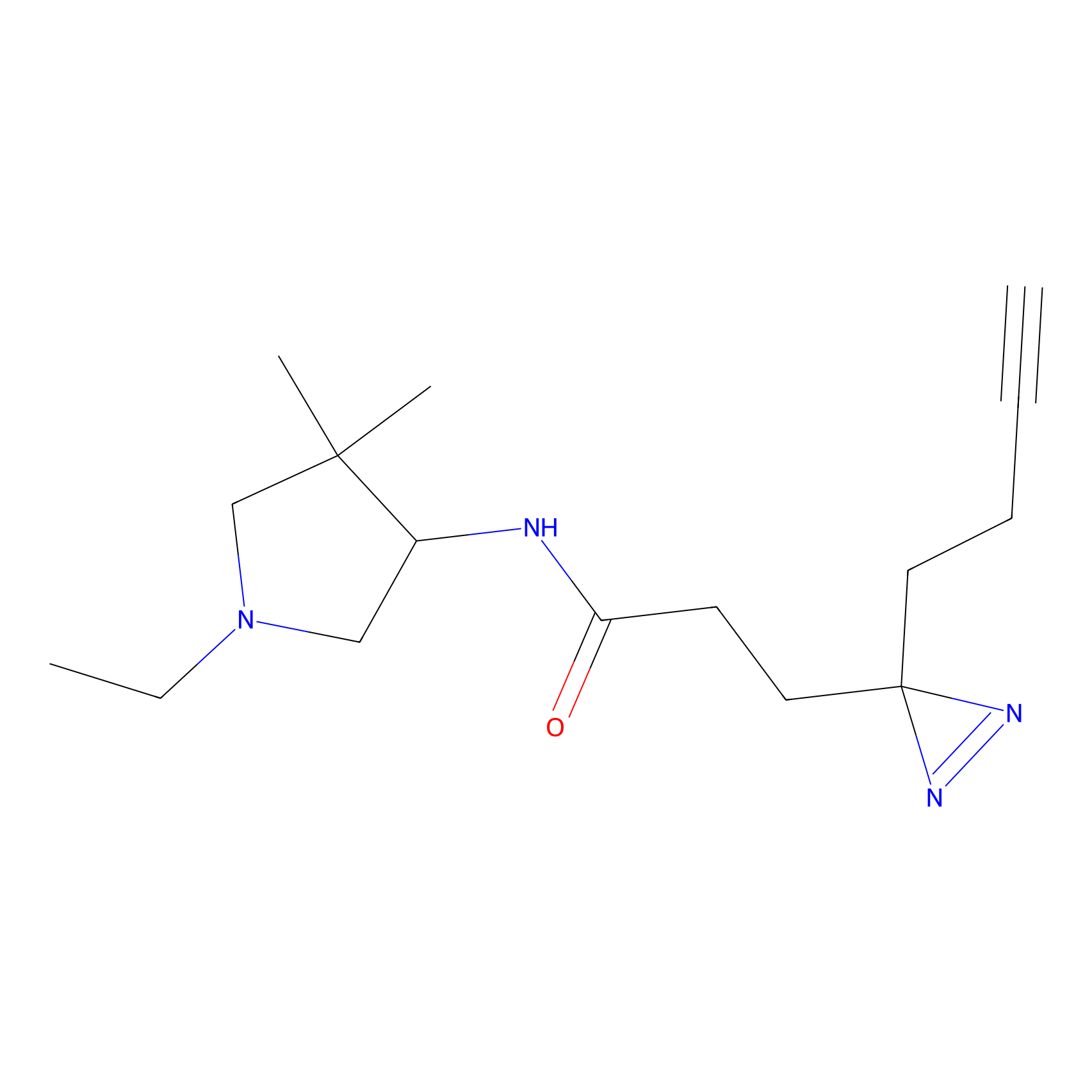
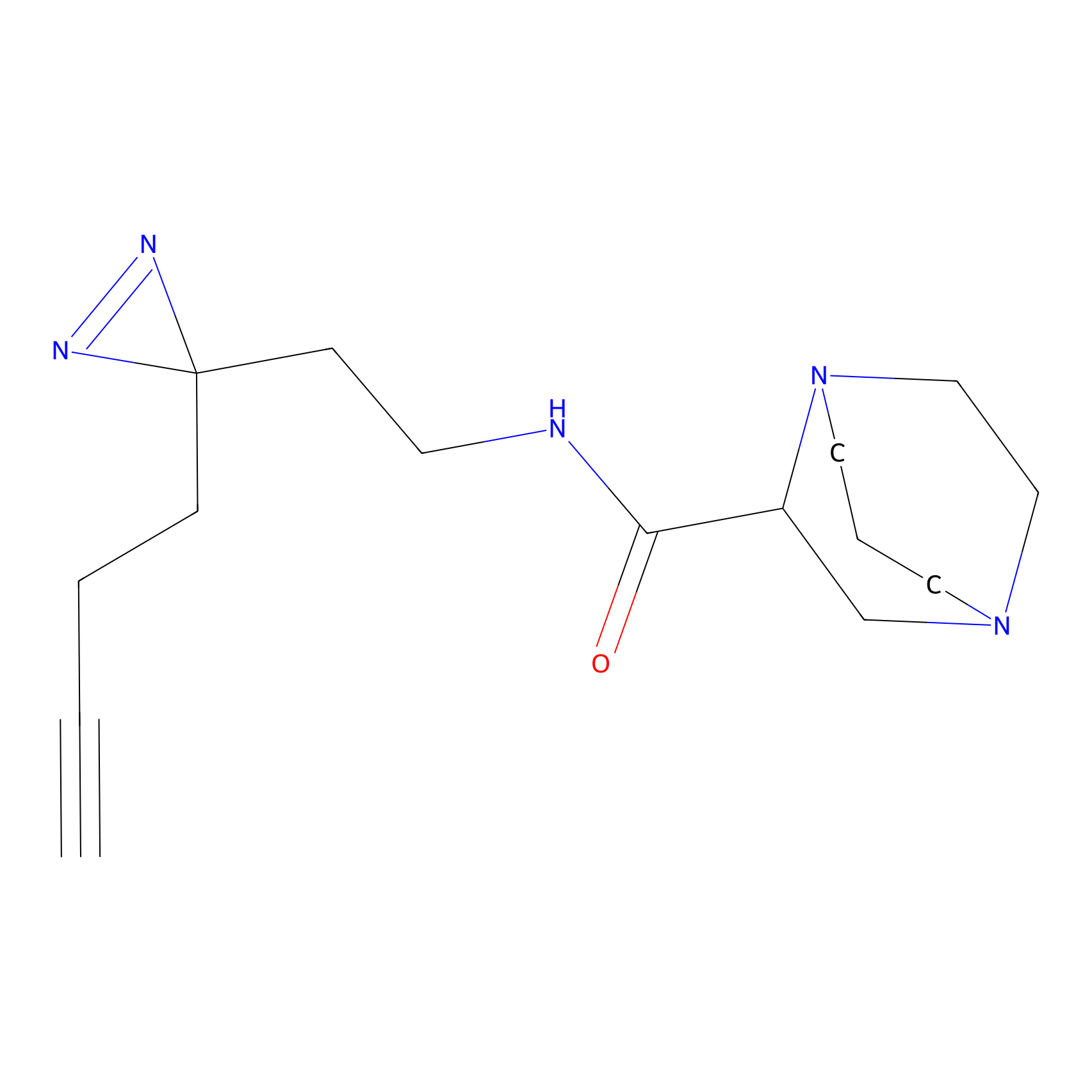
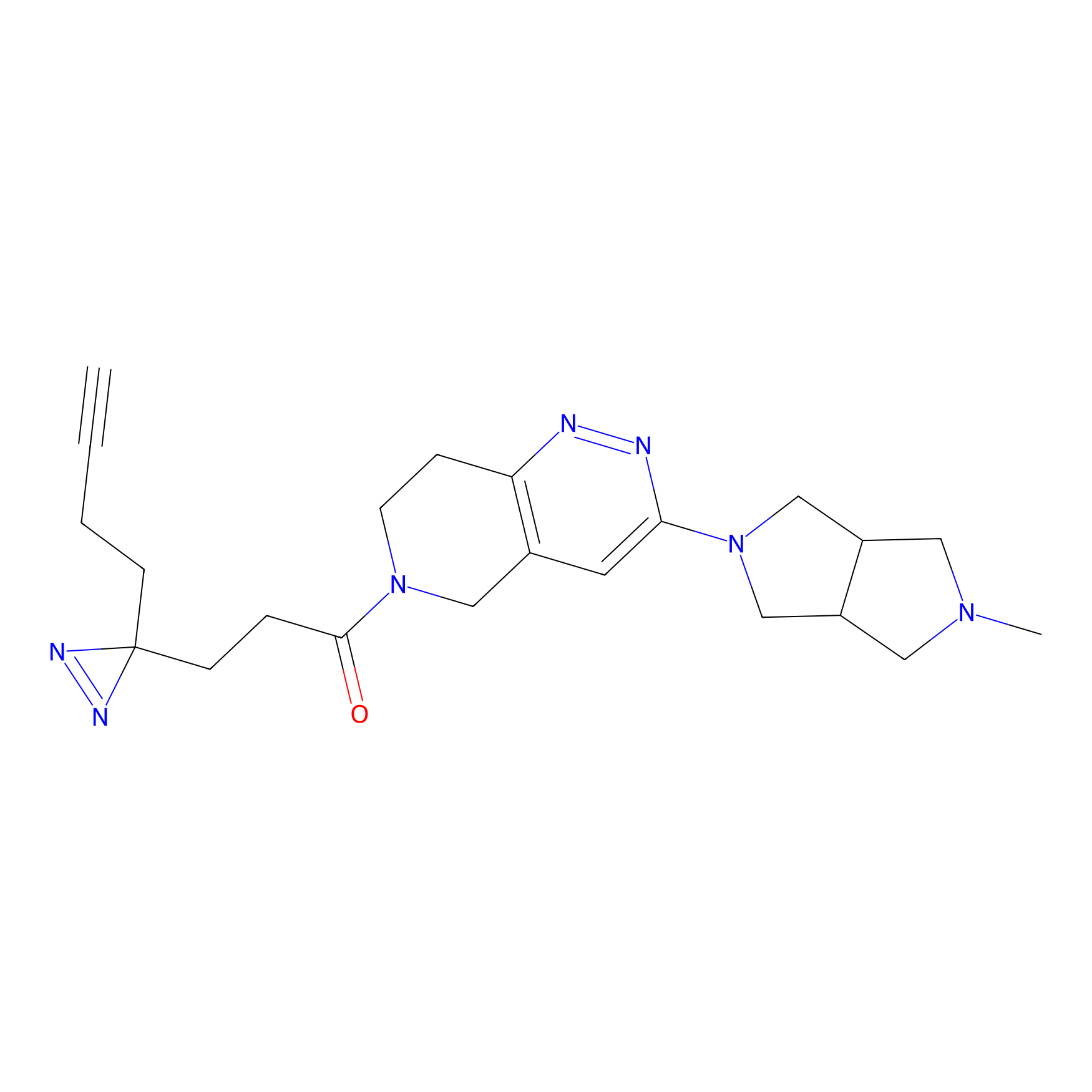
FBPP2 Probe Info |
 |
2.07 | LDD0054 | [3] | |
|
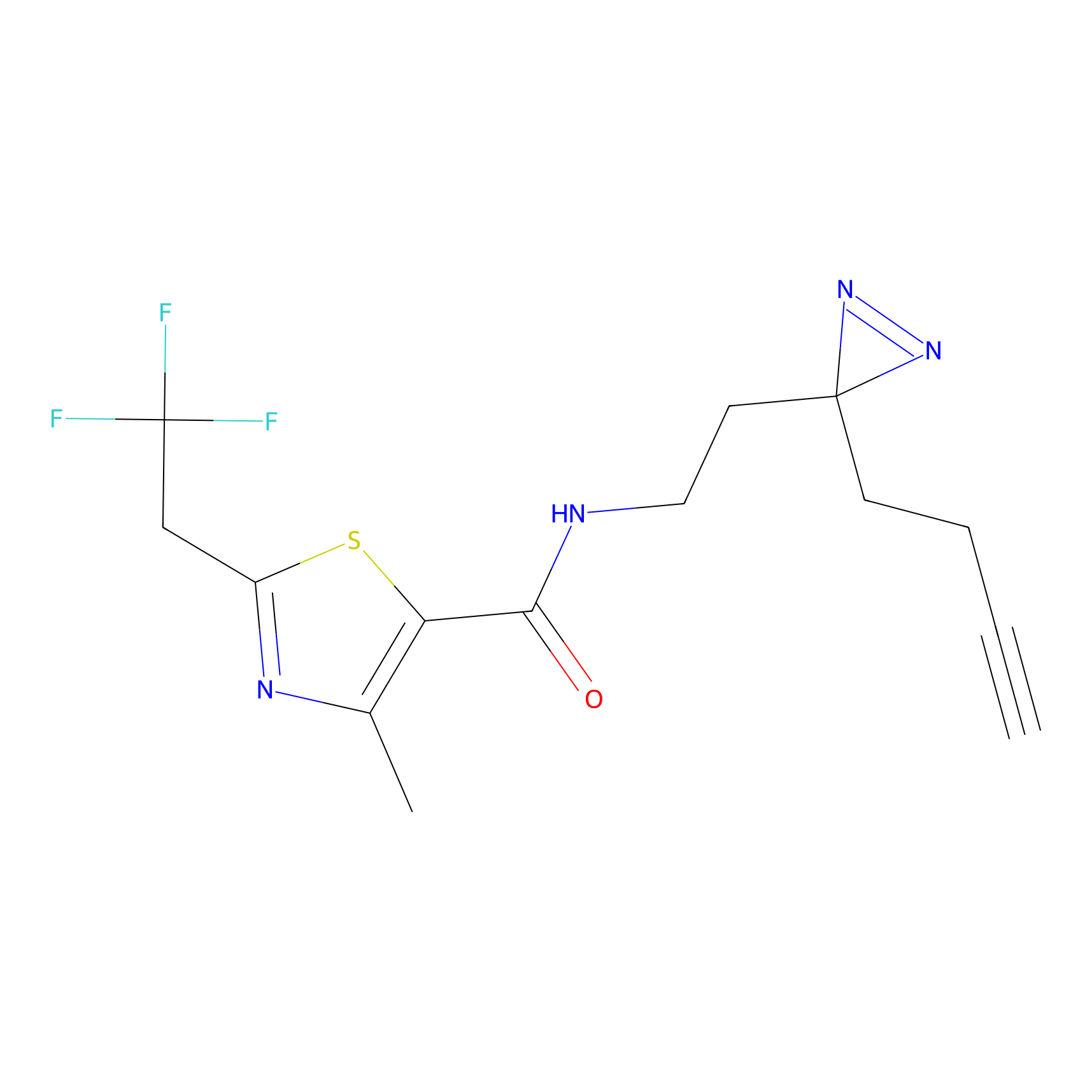
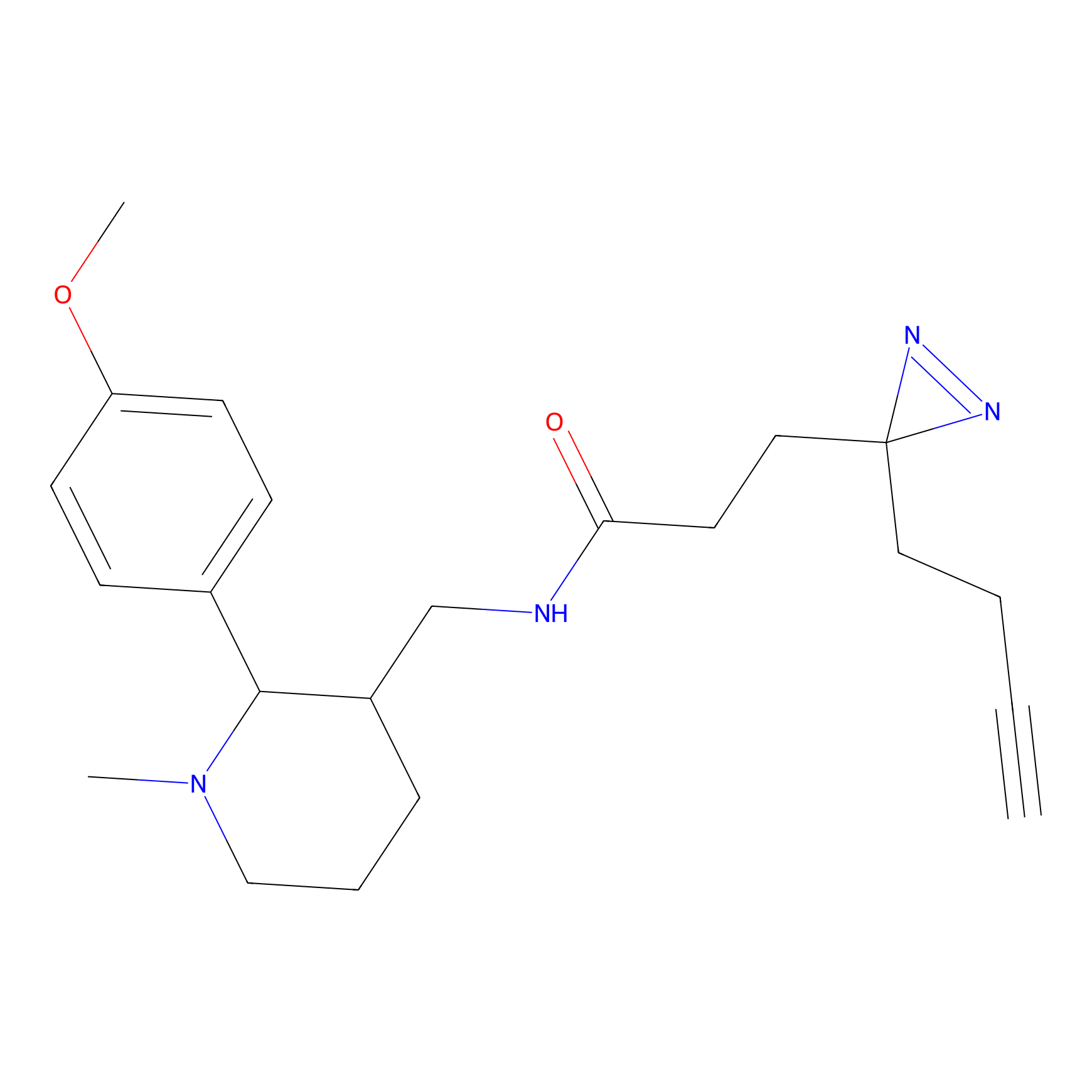
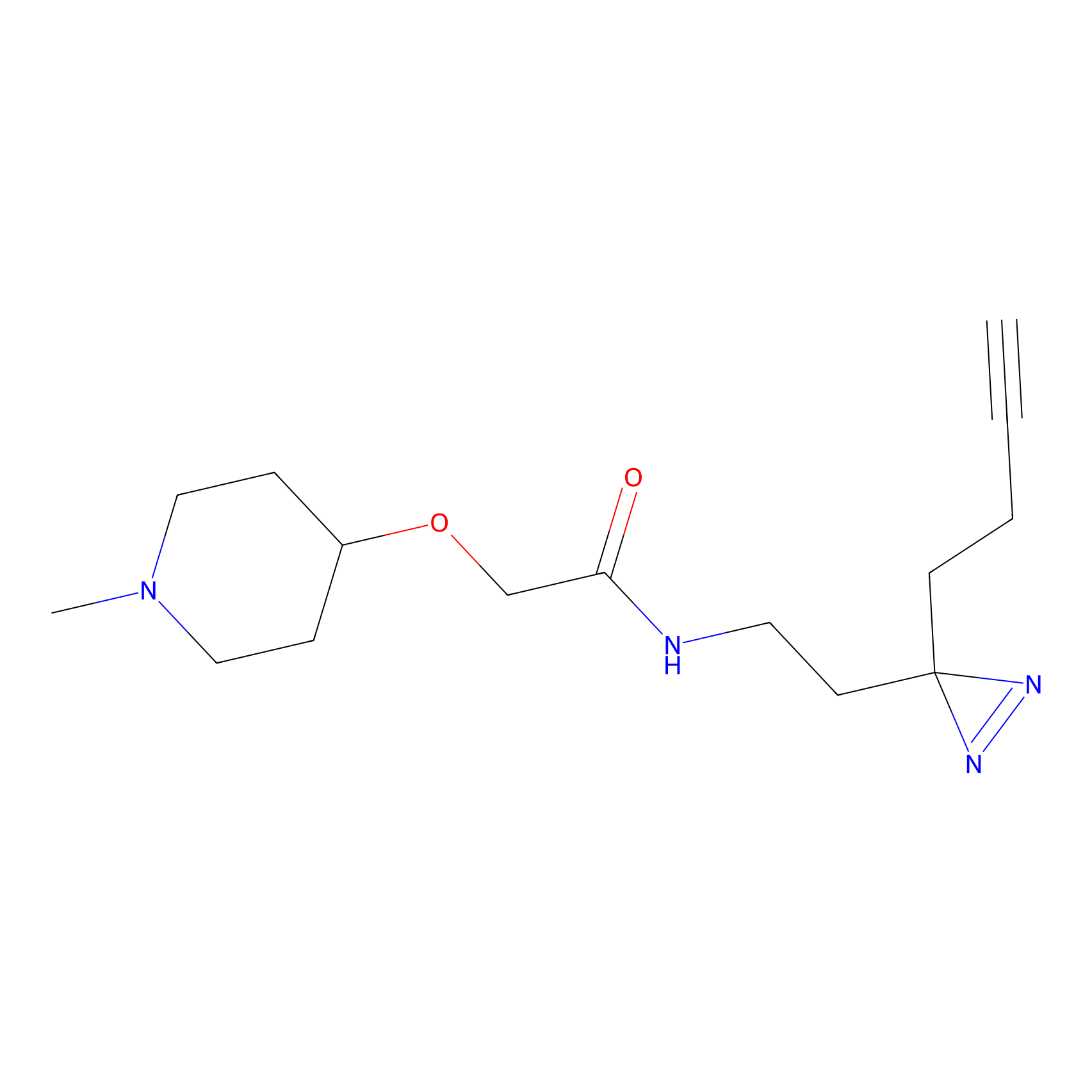
DBIA Probe Info |
 |
C97(0.93); C99(0.93) | LDD0078 | [4] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0020 | ARS-1620 | HCC44 | C97(0.93); C99(0.93) | LDD0078 | [4] |
| LDCM0185 | Compound 17 | HEK-293T | 6.27 | LDD0512 | [6] |
| LDCM0191 | Compound 21 | HEK-293T | 11.56 | LDD0508 | [6] |
| LDCM0190 | Compound 34 | HEK-293T | 9.30 | LDD0510 | [6] |
| LDCM0192 | Compound 35 | HEK-293T | 9.09 | LDD0509 | [6] |
| LDCM0193 | Compound 36 | HEK-293T | 9.52 | LDD0511 | [6] |
| LDCM0008 | Tranylcypromine | SH-SY5Y | 2.07 | LDD0054 | [3] |
The Interaction Atlas With This Target
References