Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
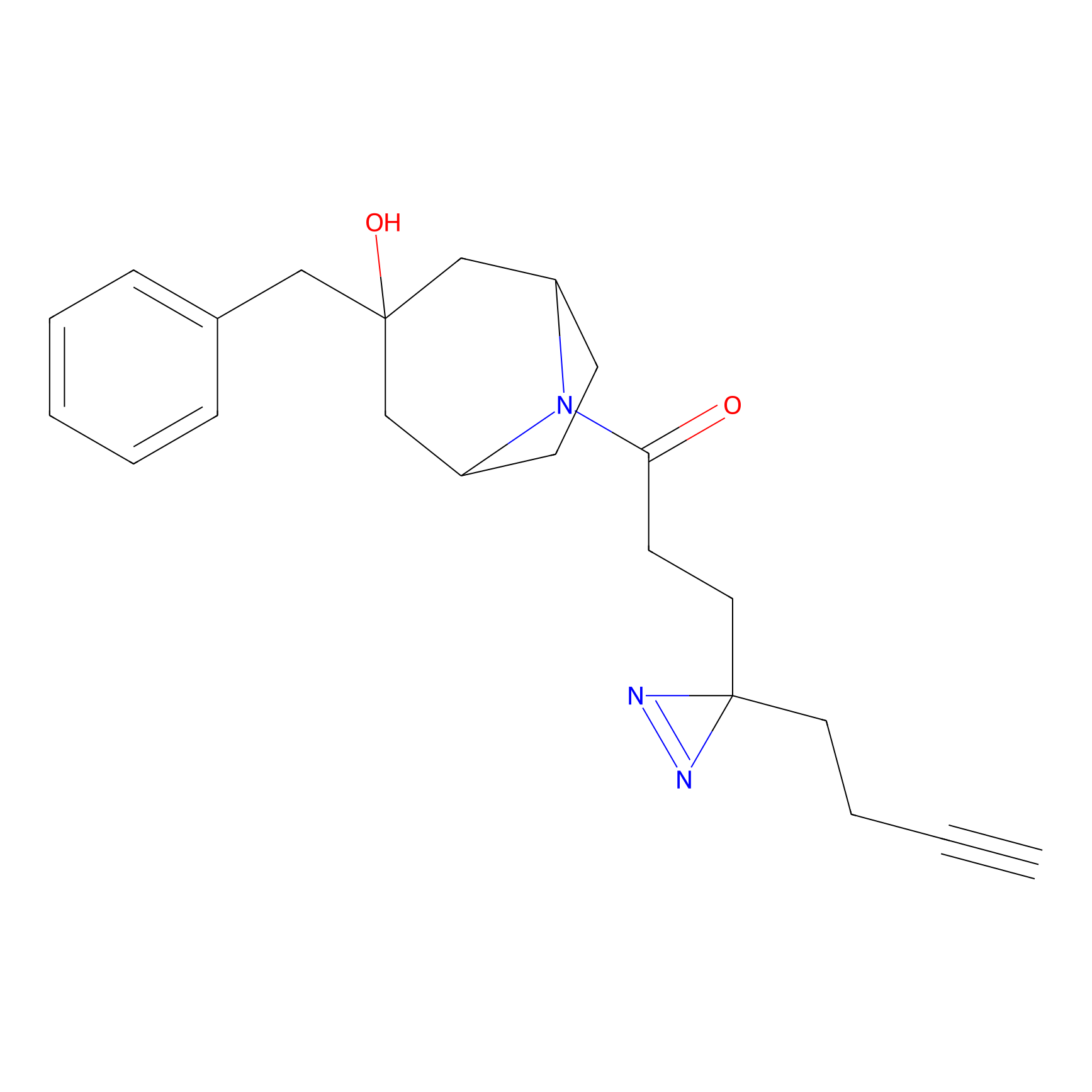
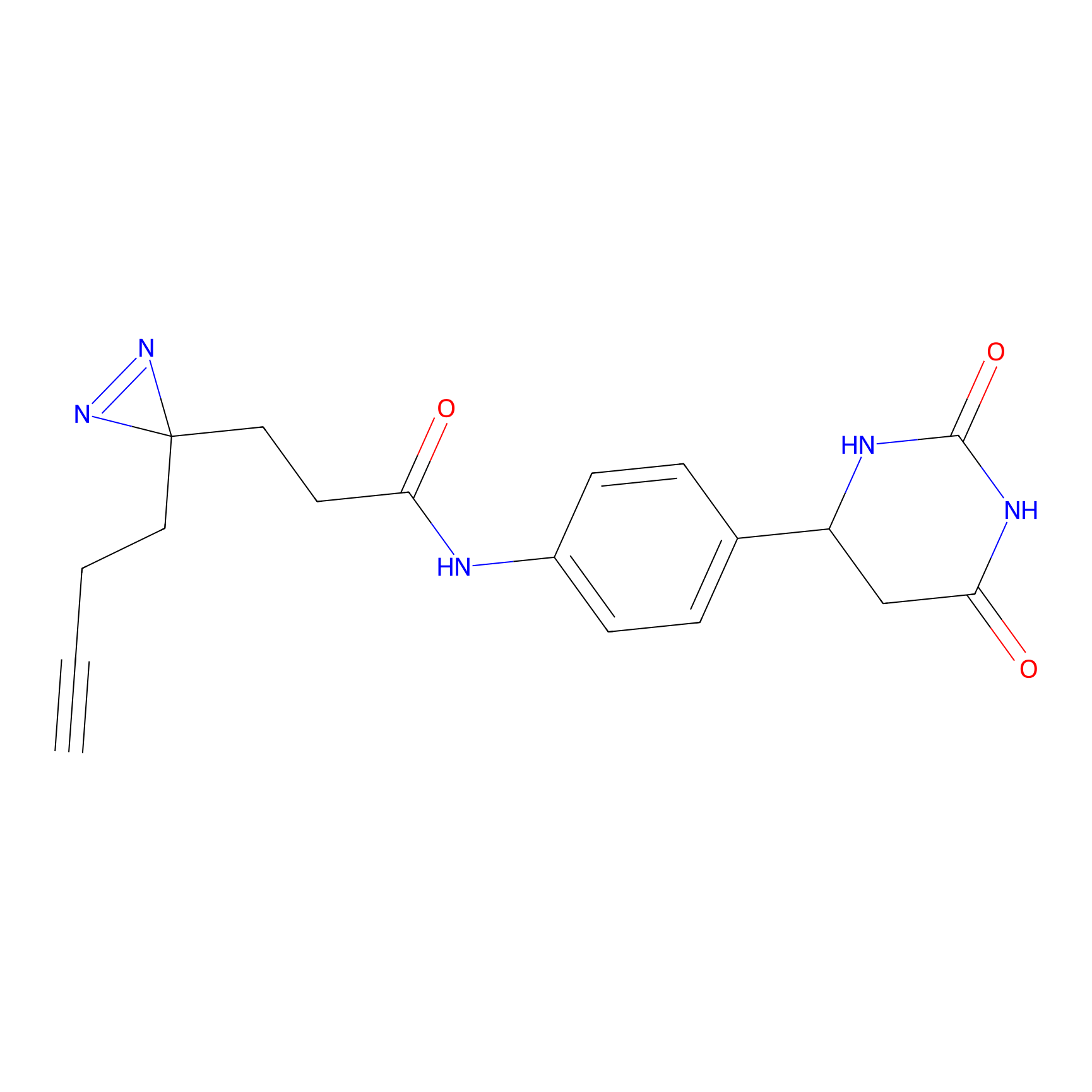
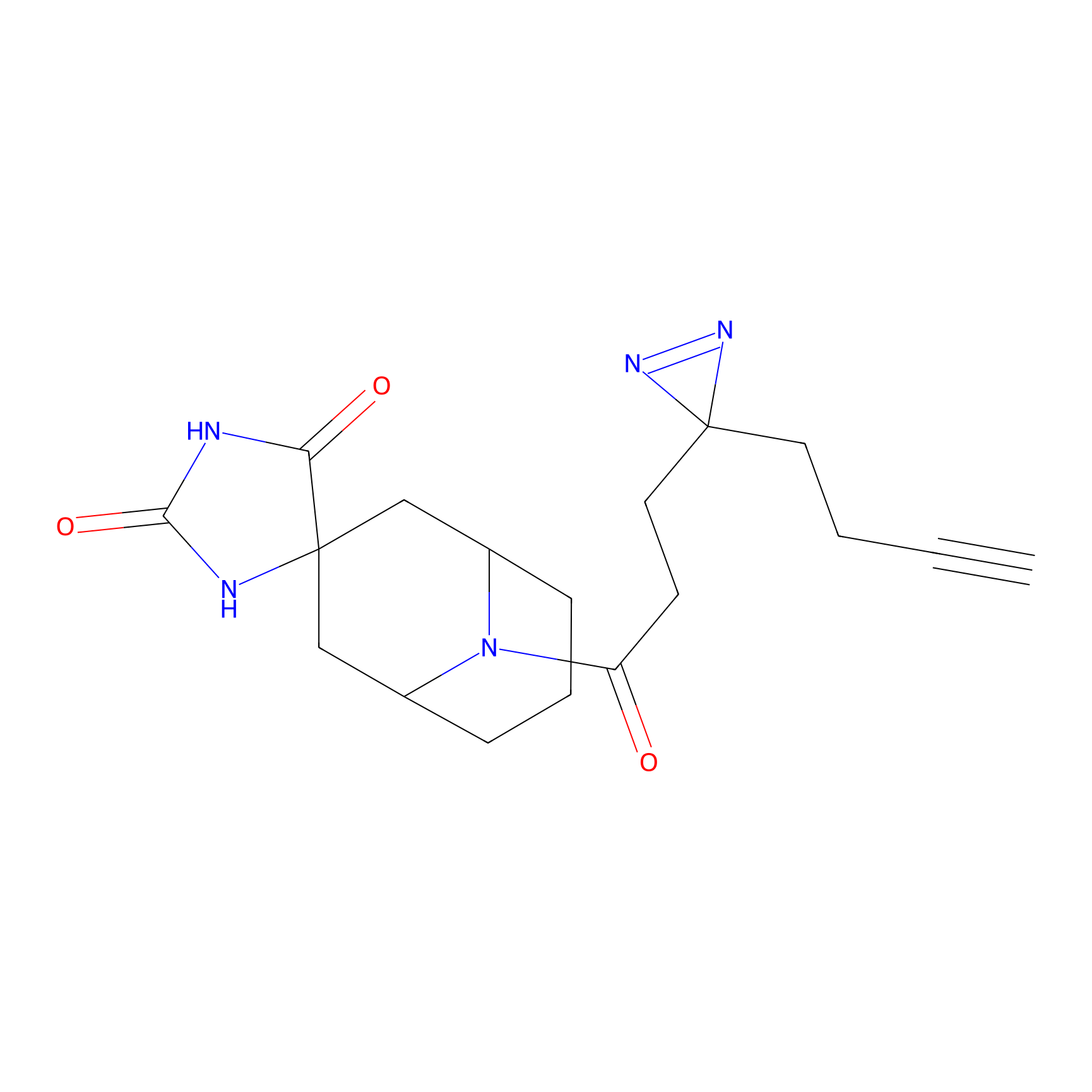
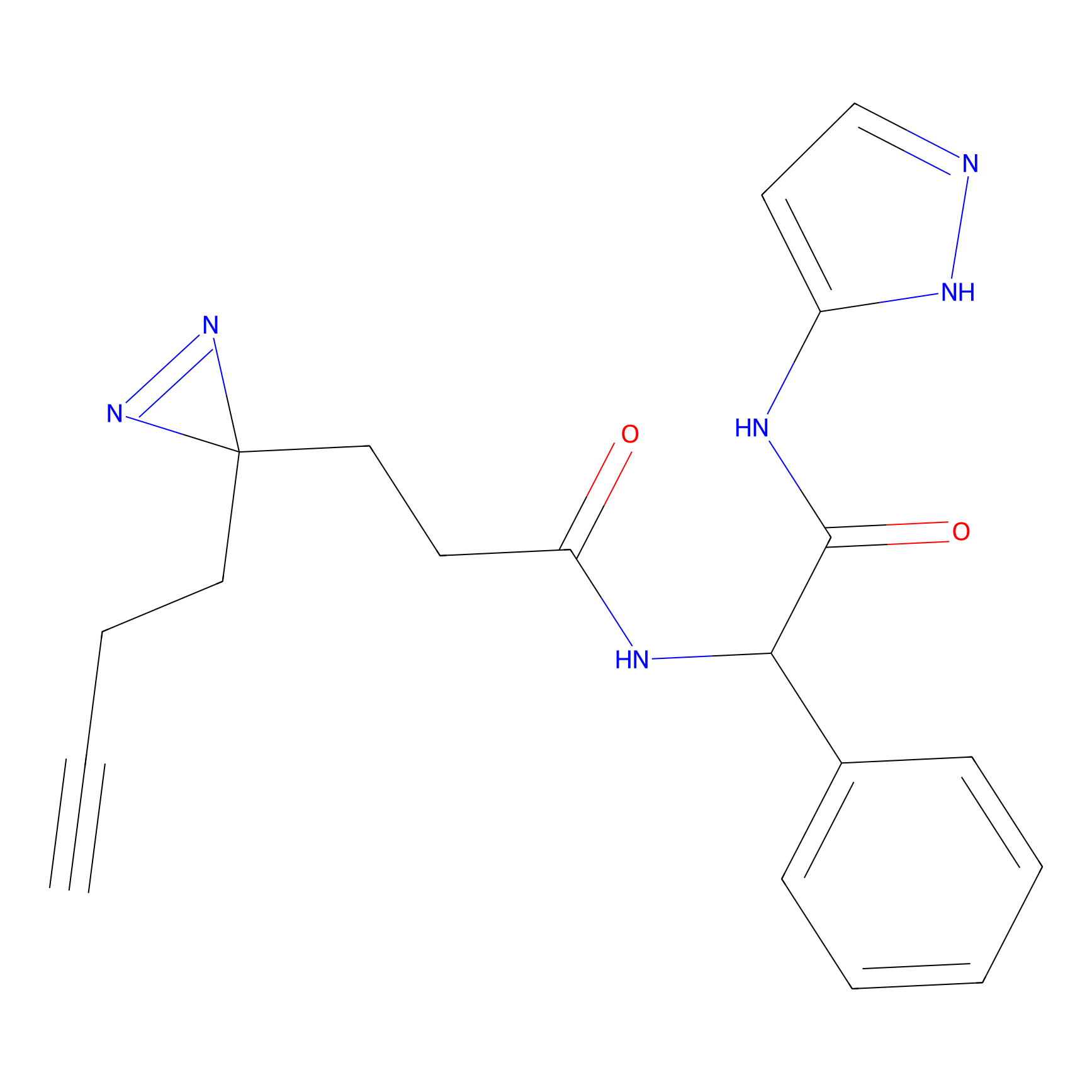
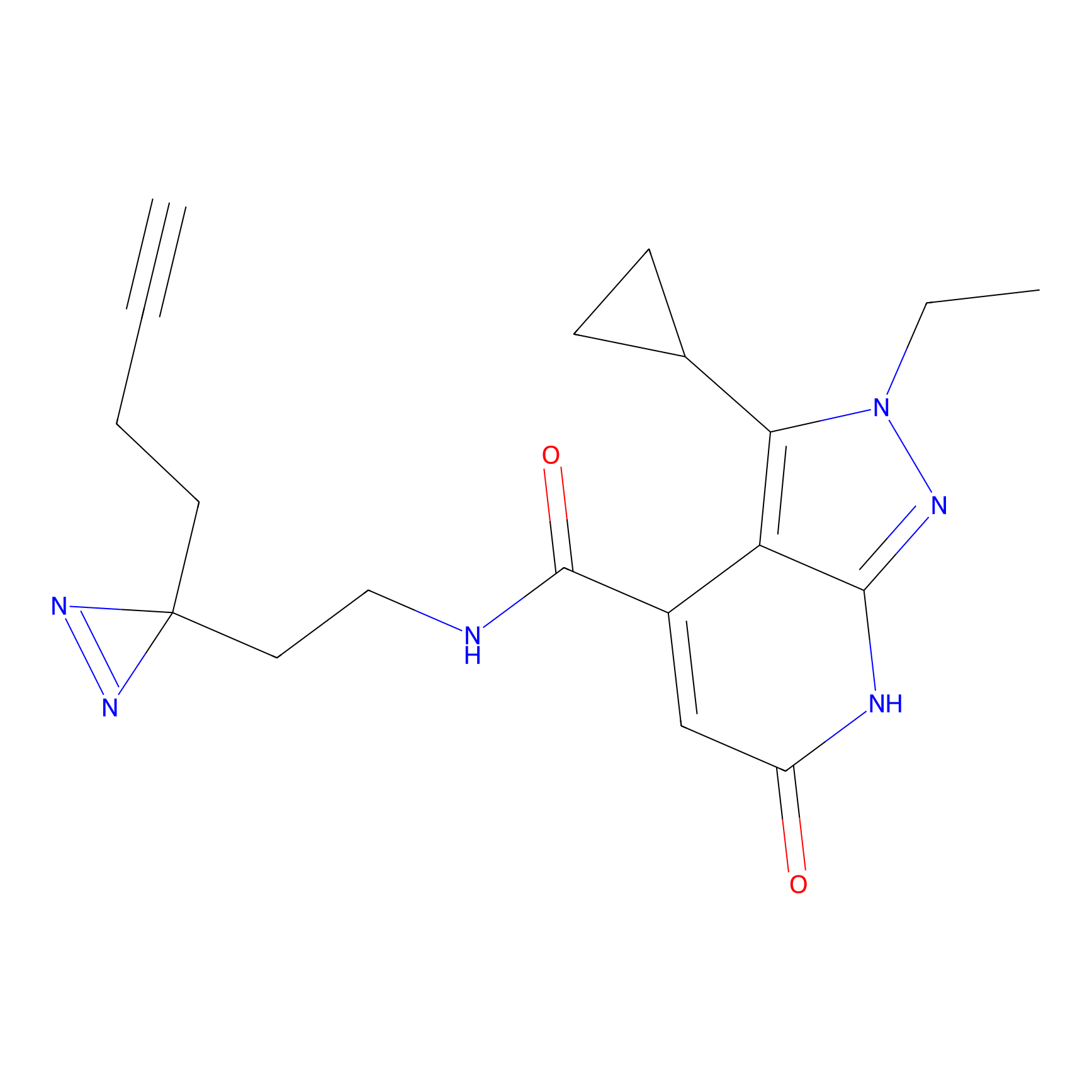
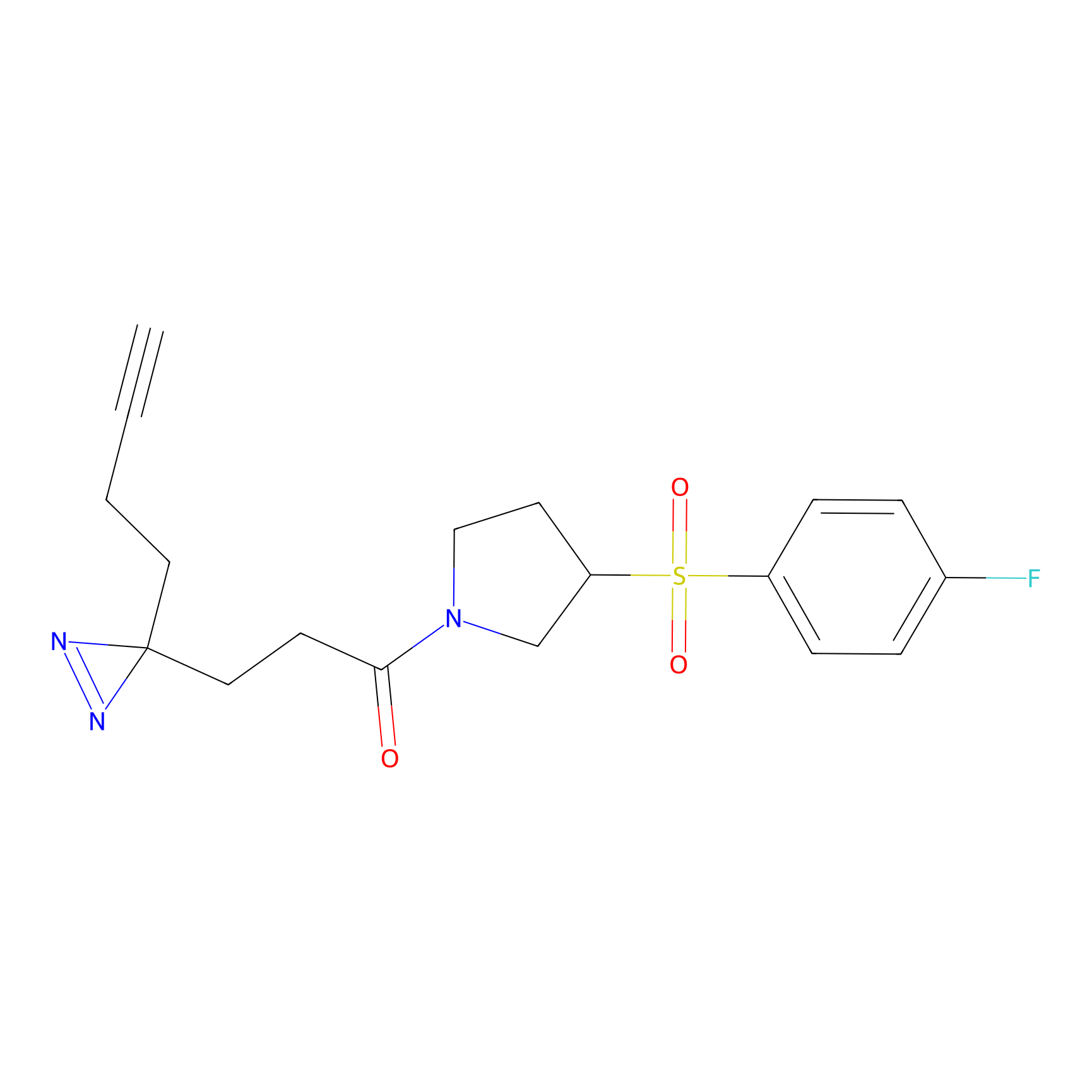
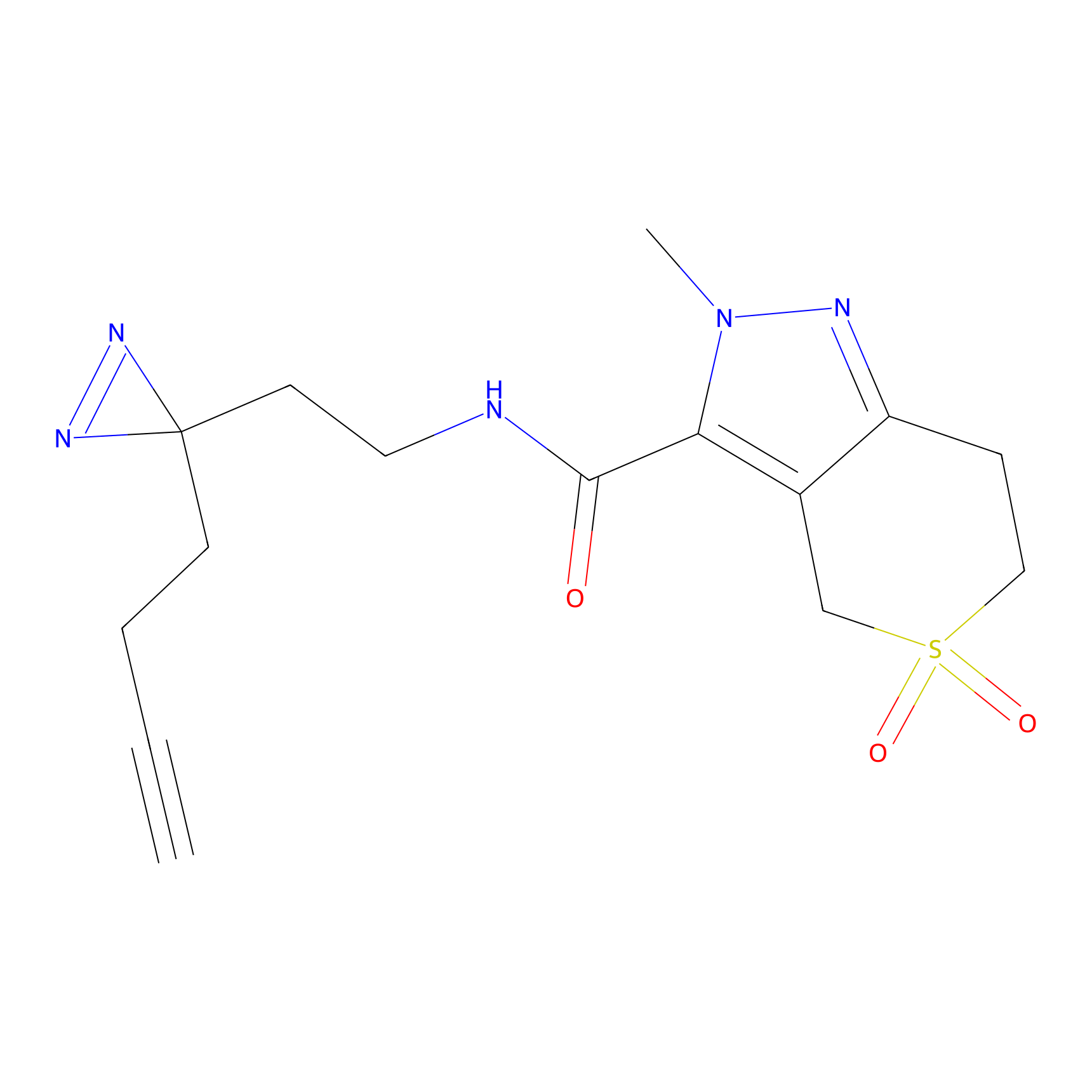
Alkylaryl probe 3 Probe Info |
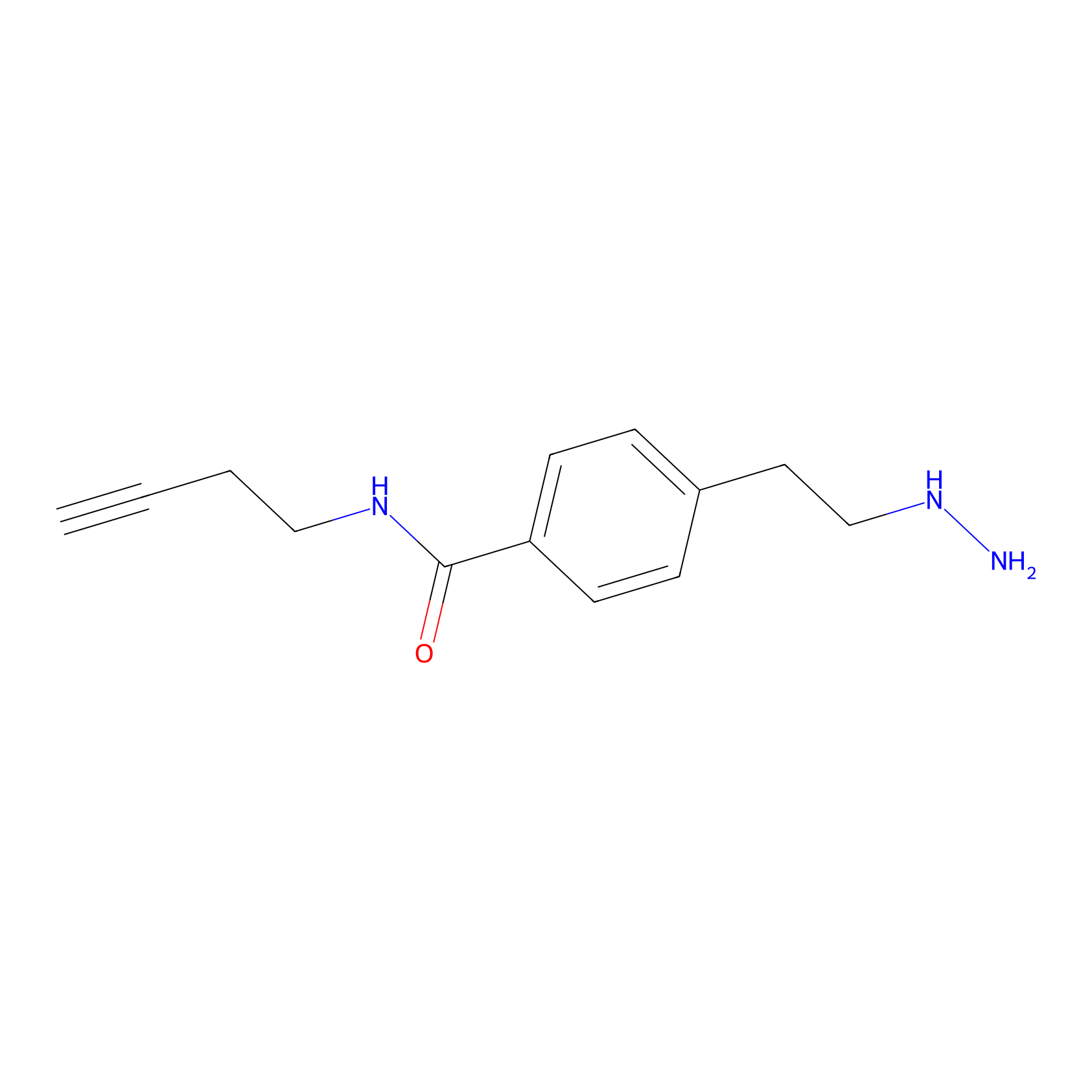 |
5.96 | LDD0381 | [1] | |
|
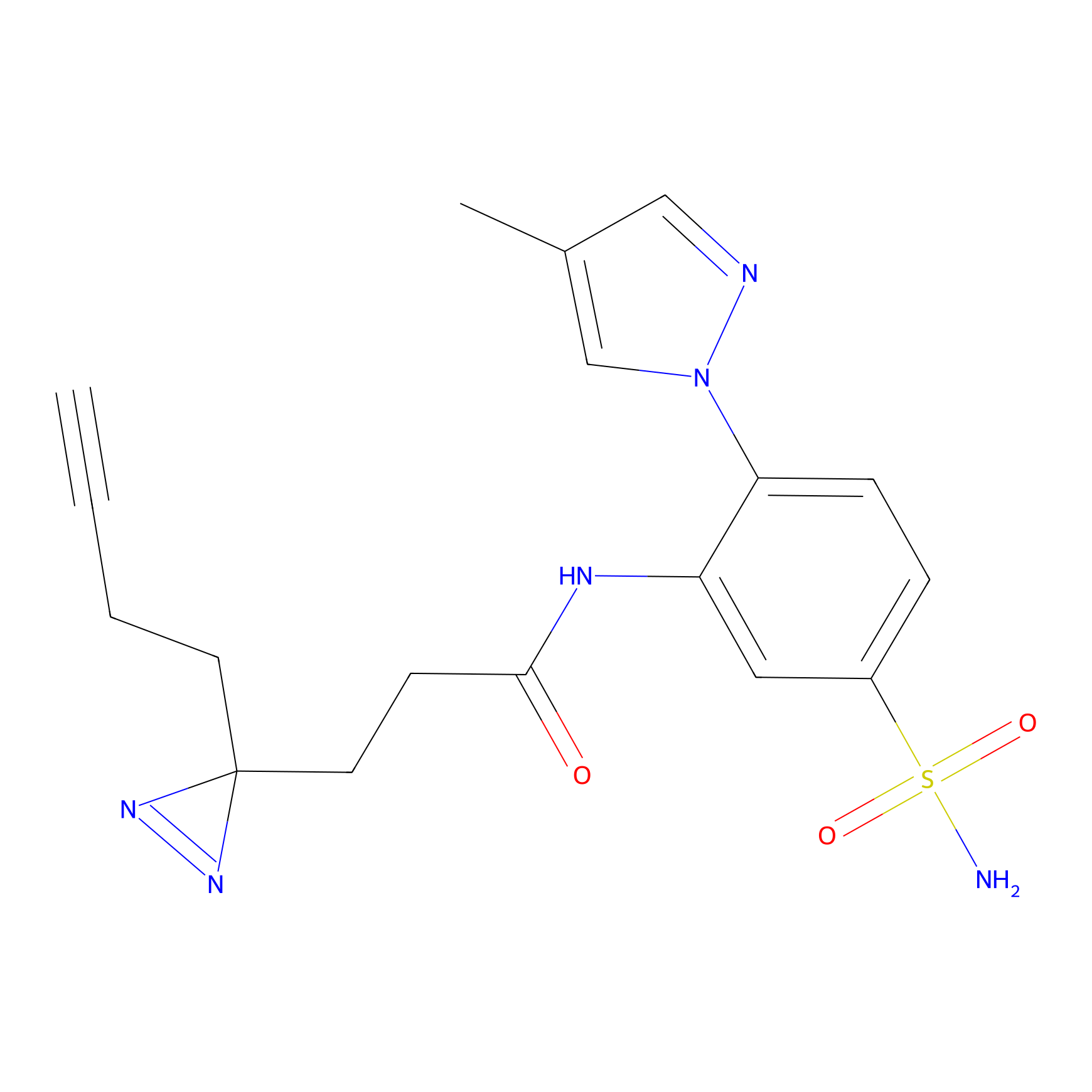
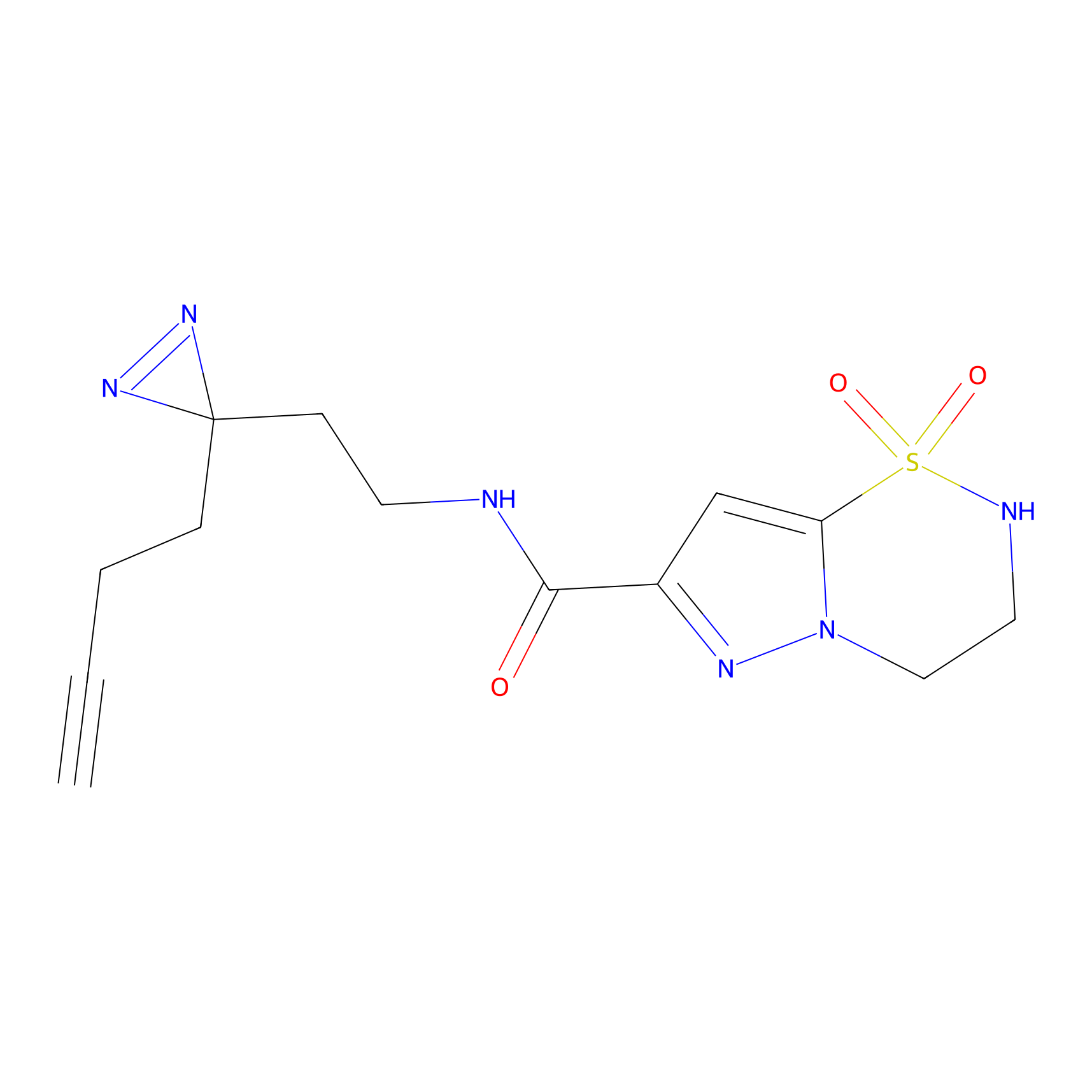
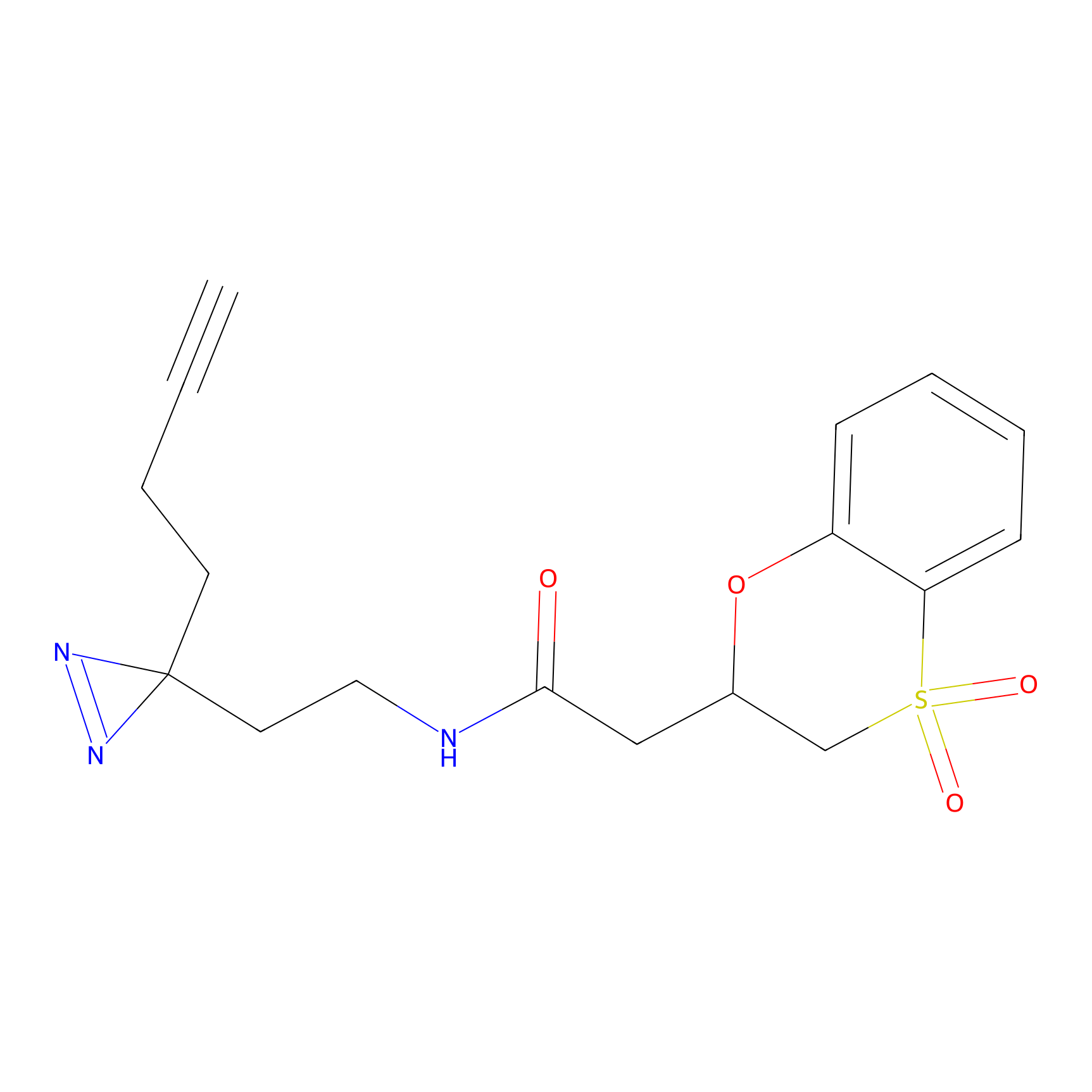
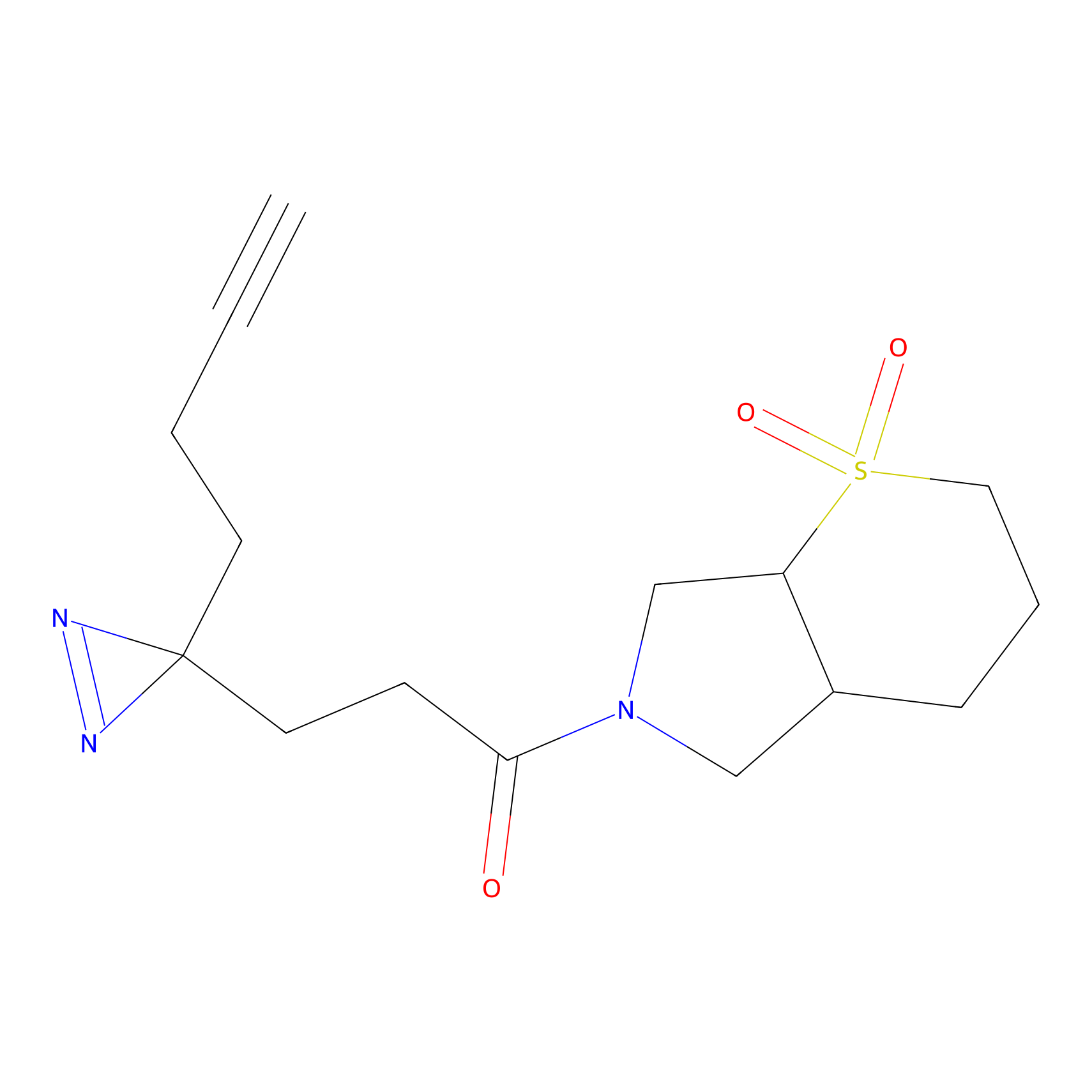
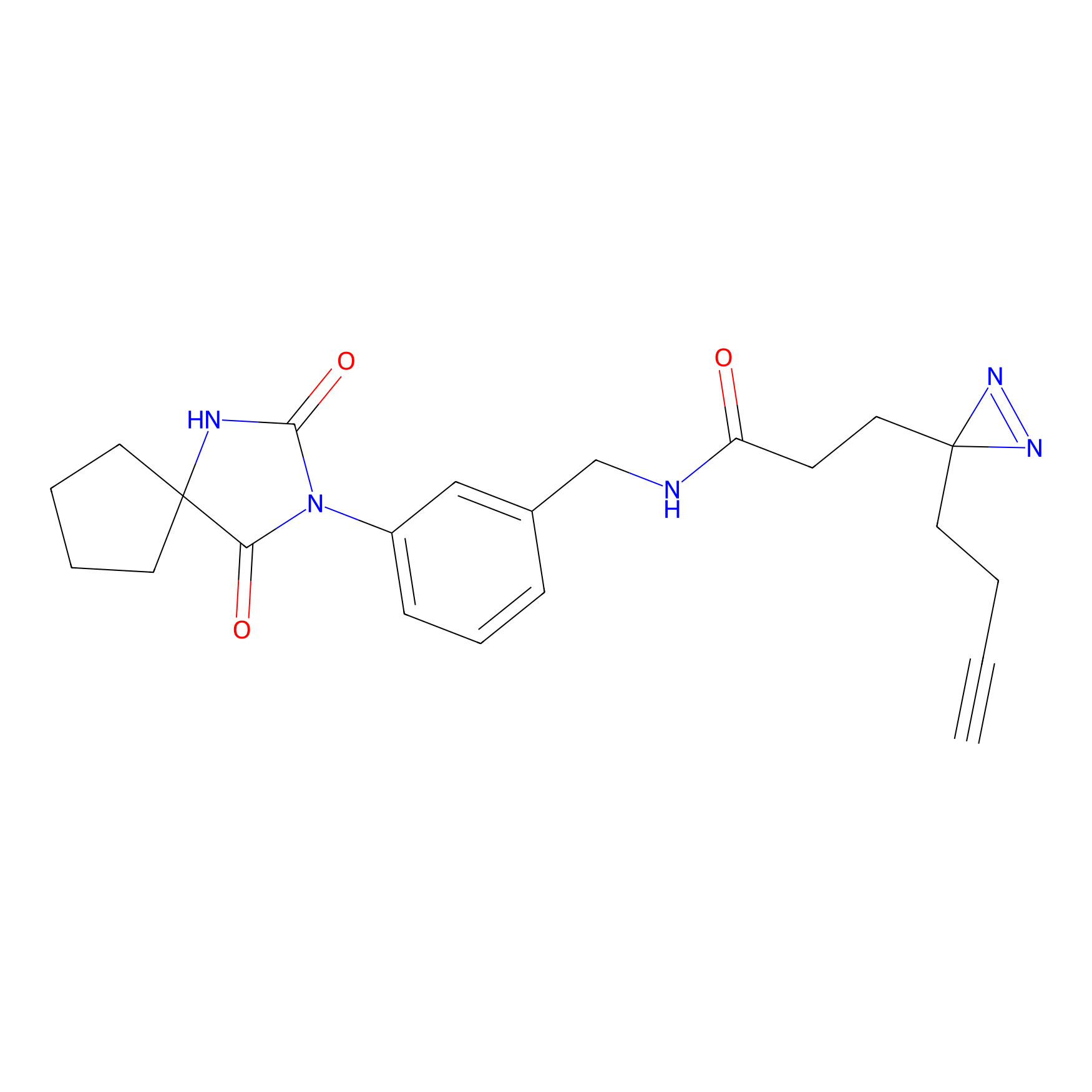
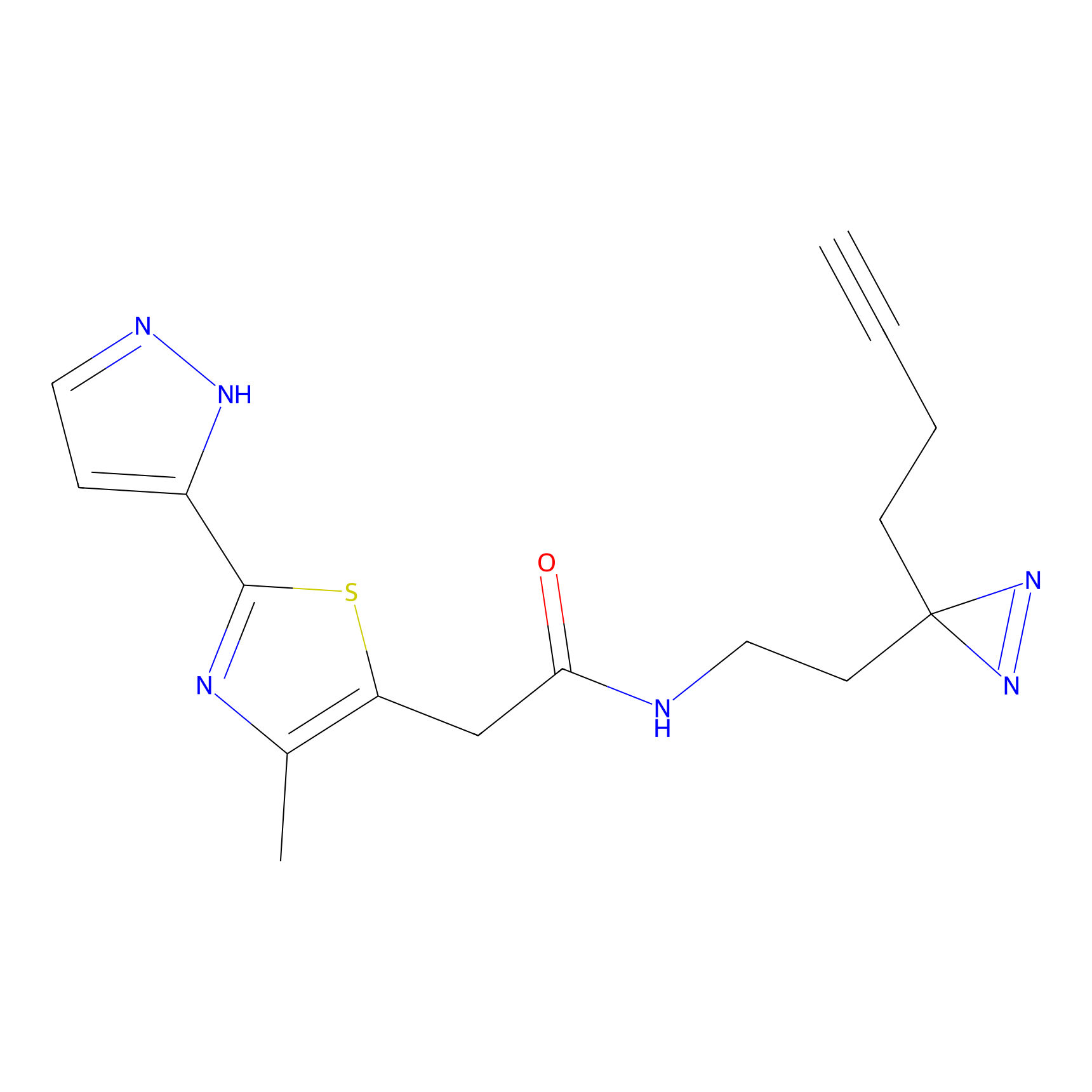
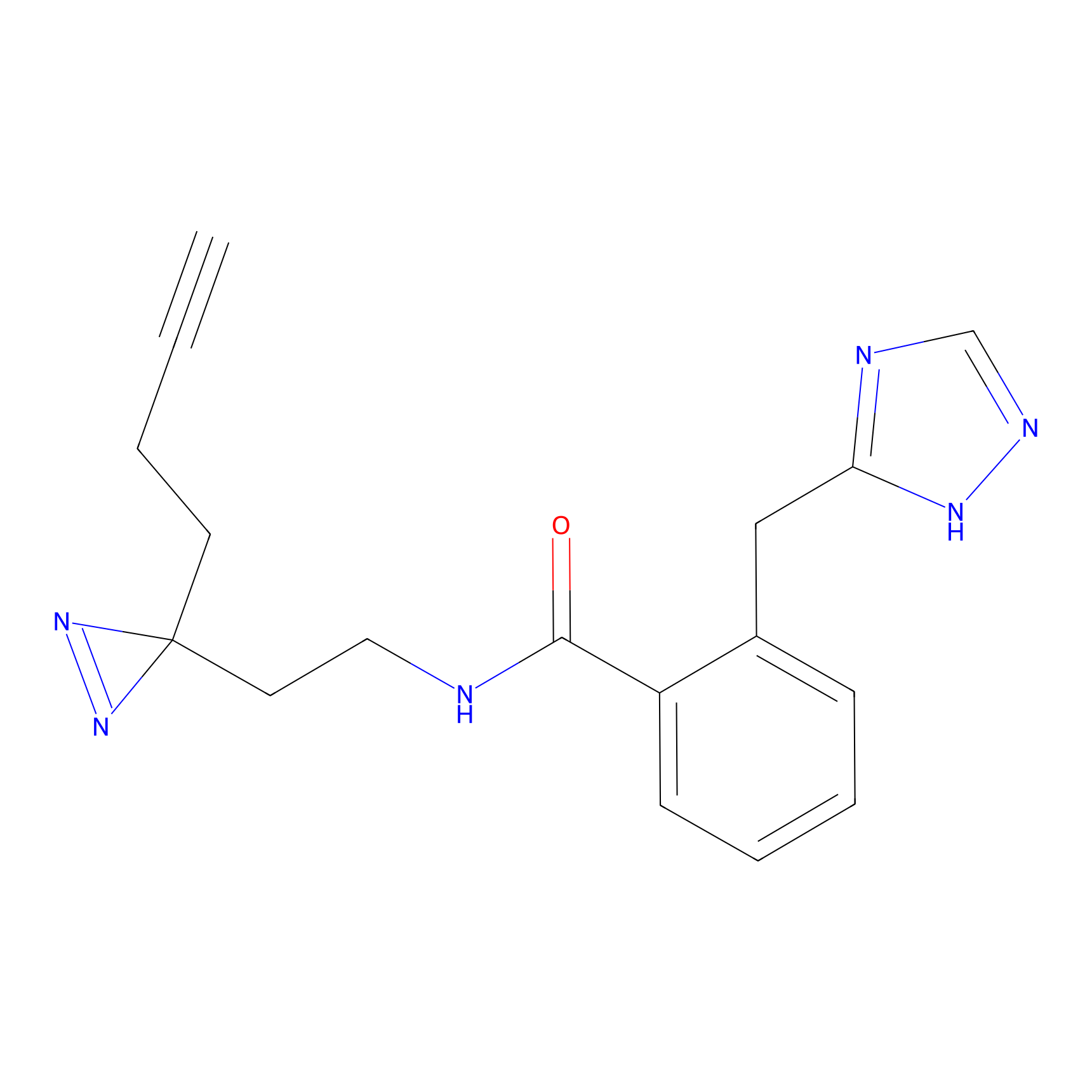
FBPP2 Probe Info |
 |
362.11 | LDD0318 | [2] | |
|
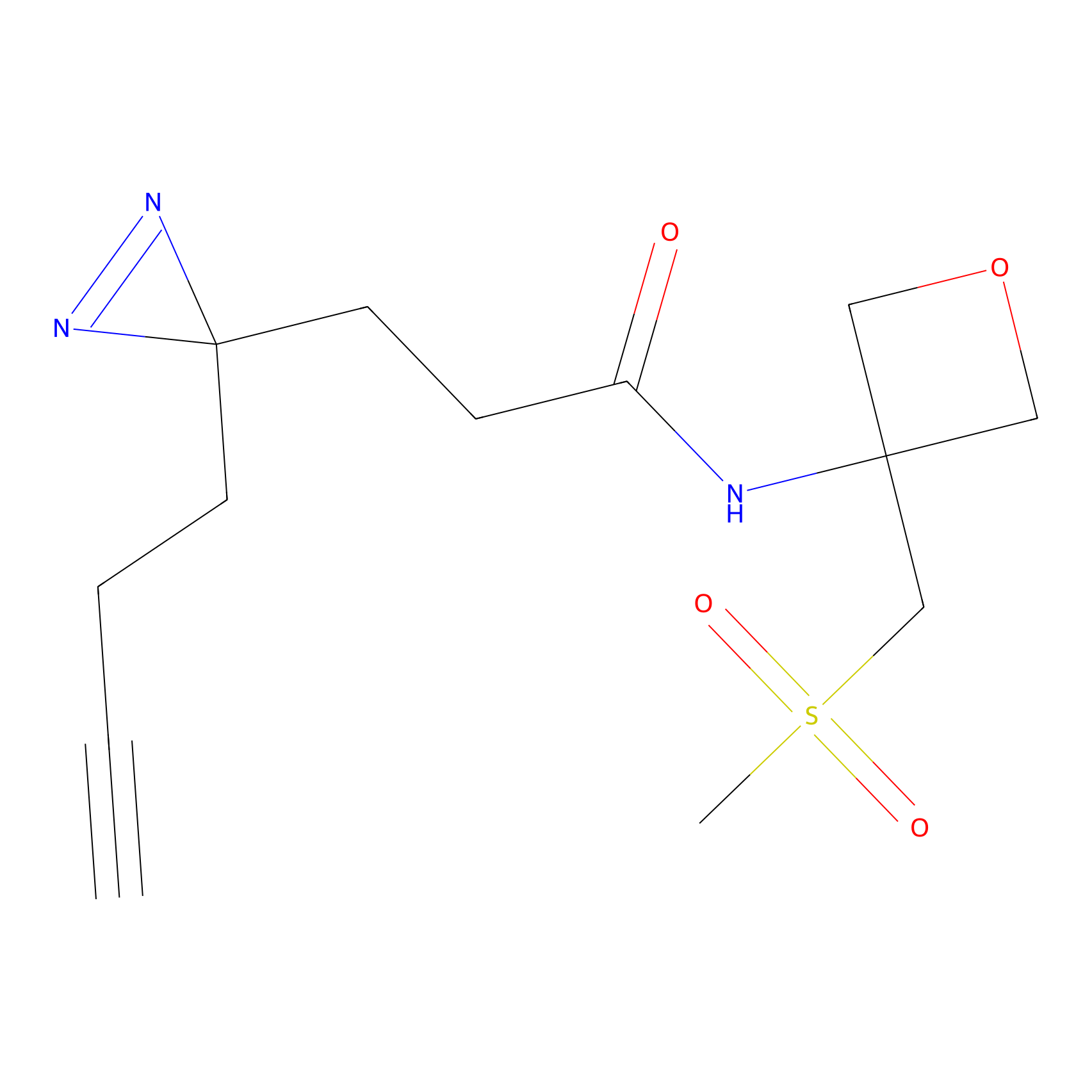
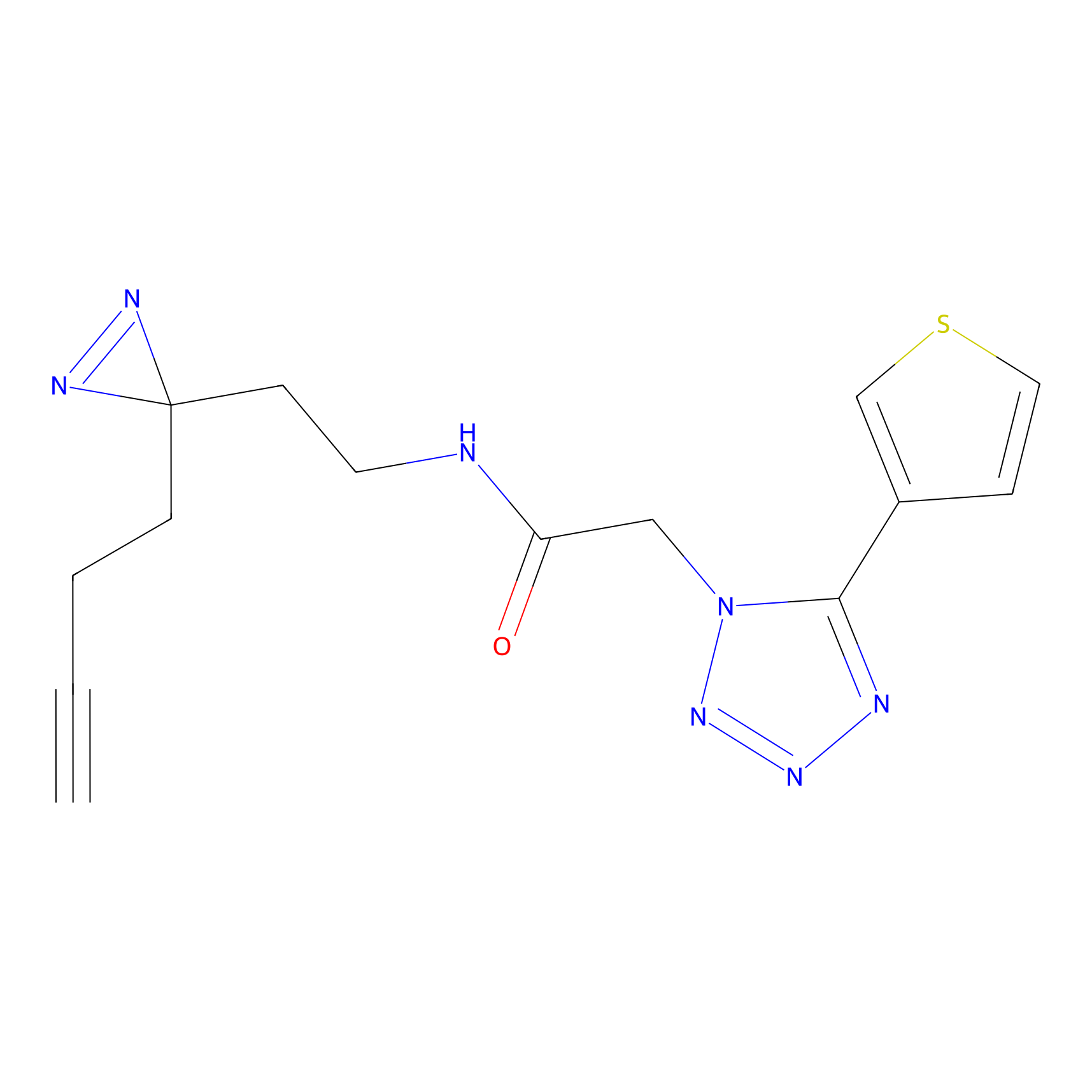
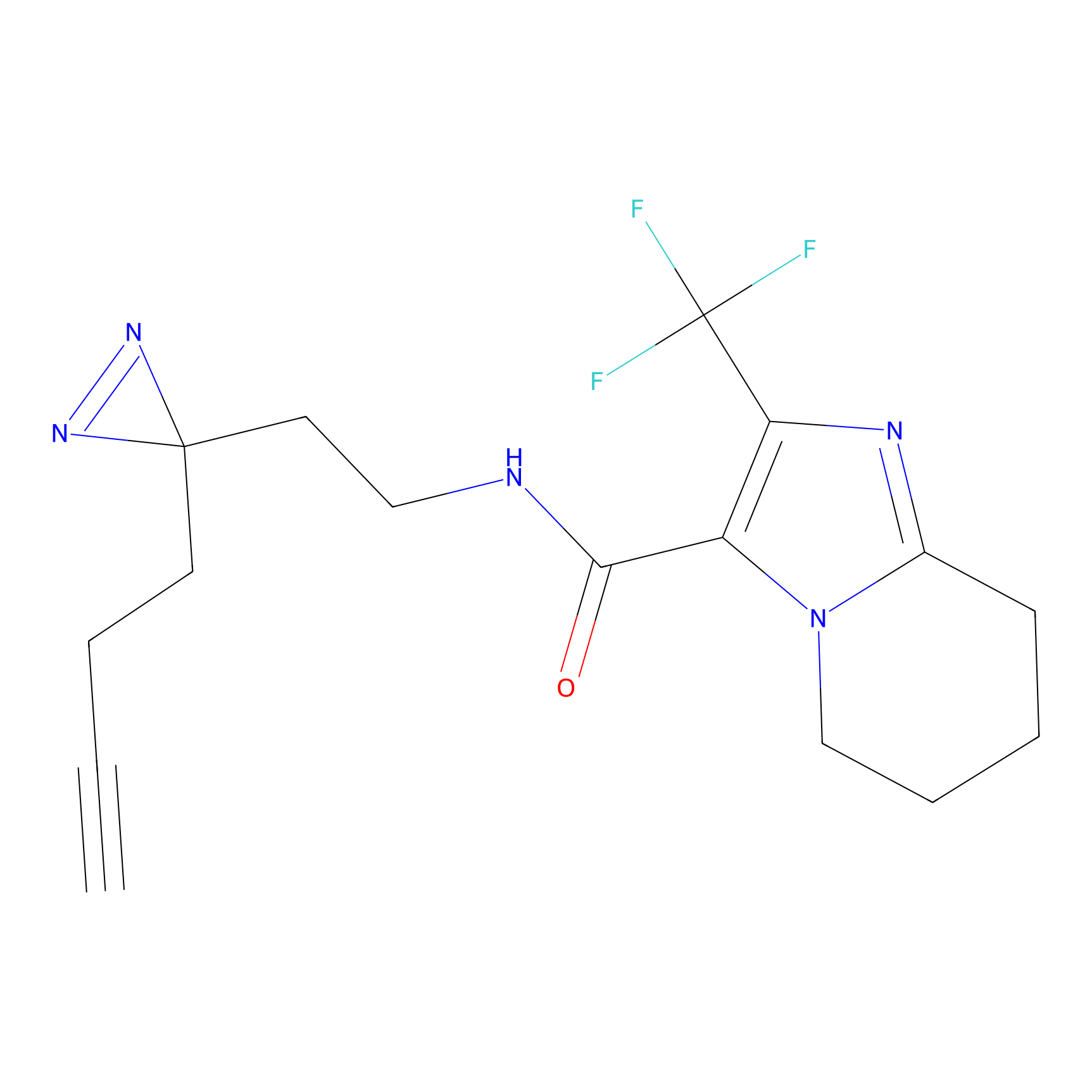
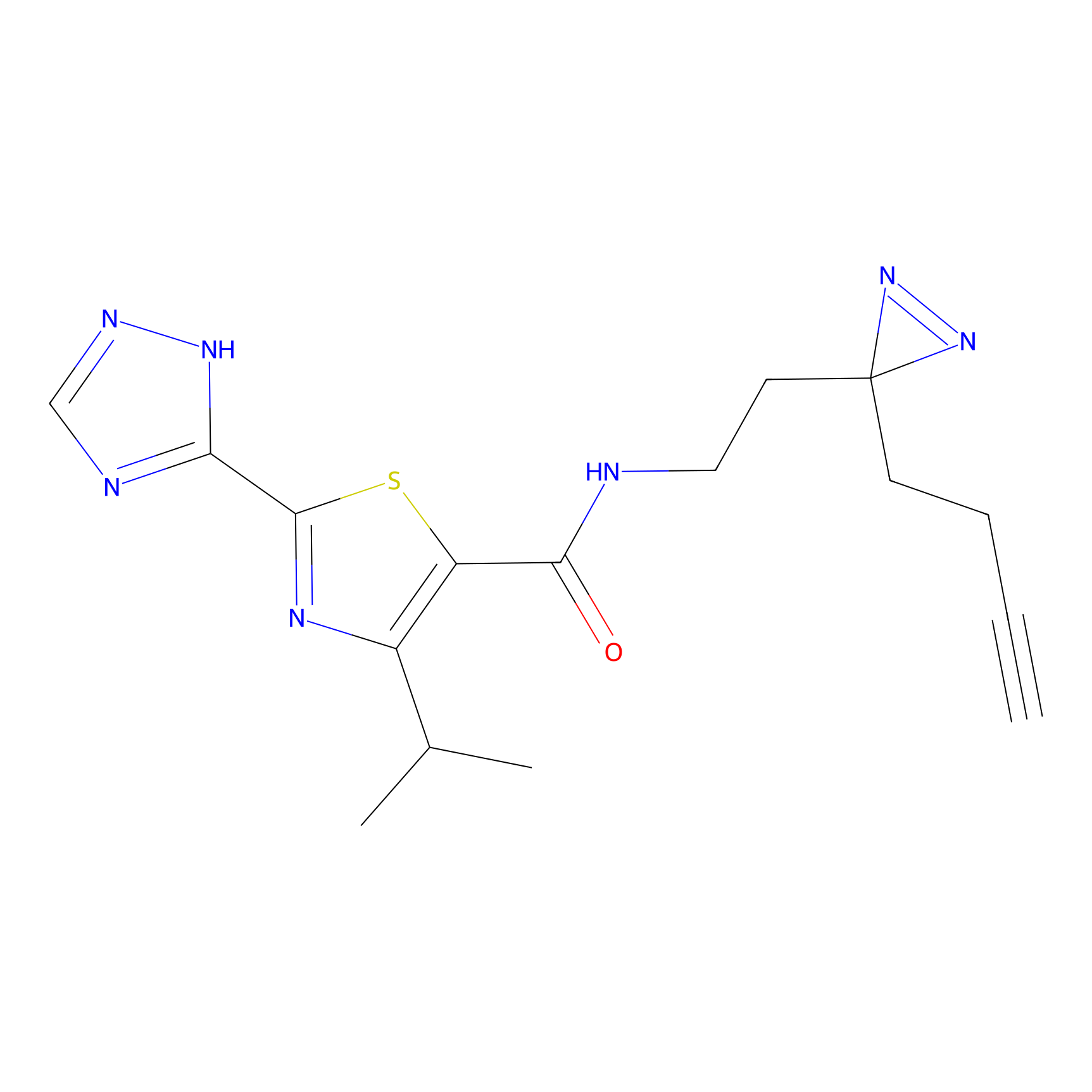
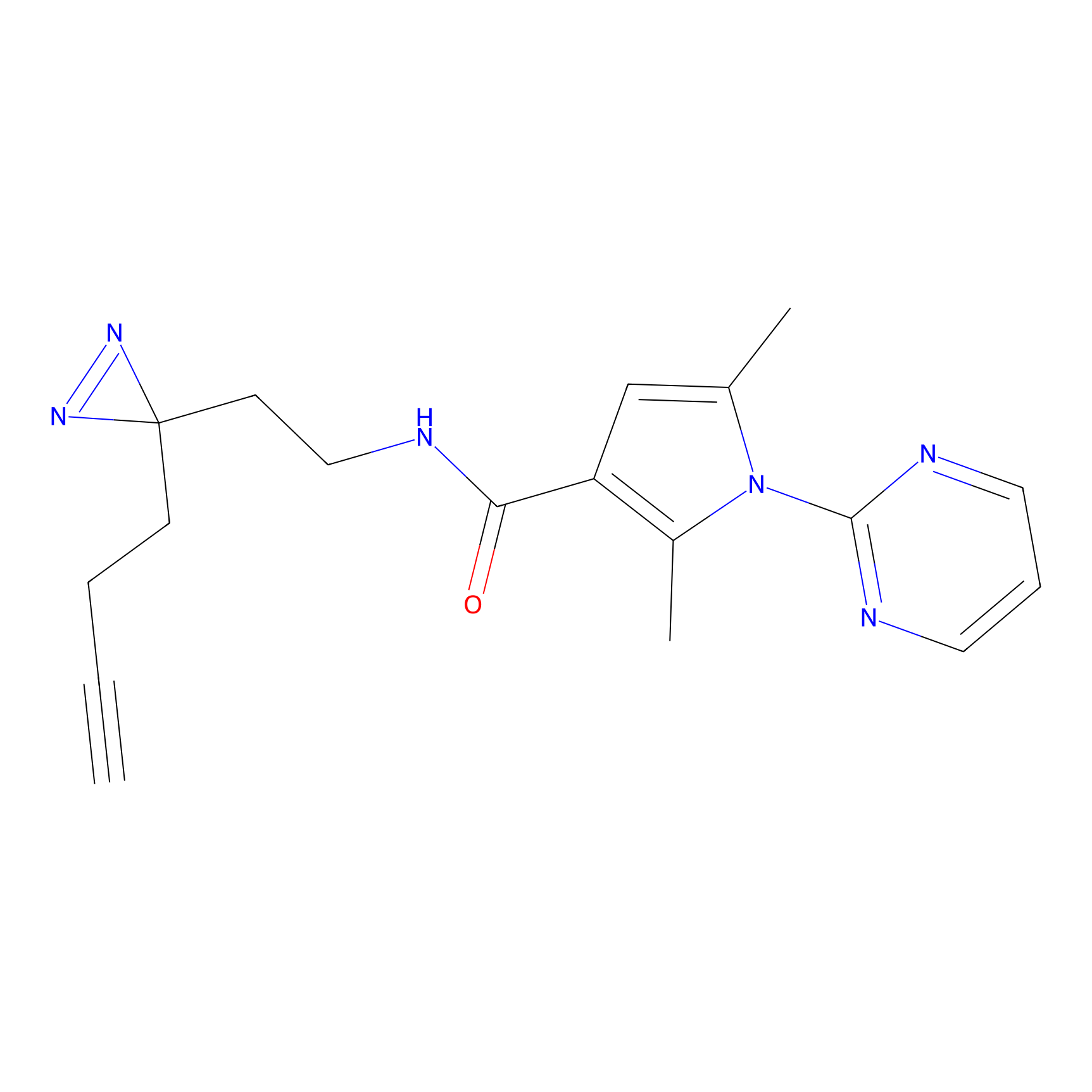
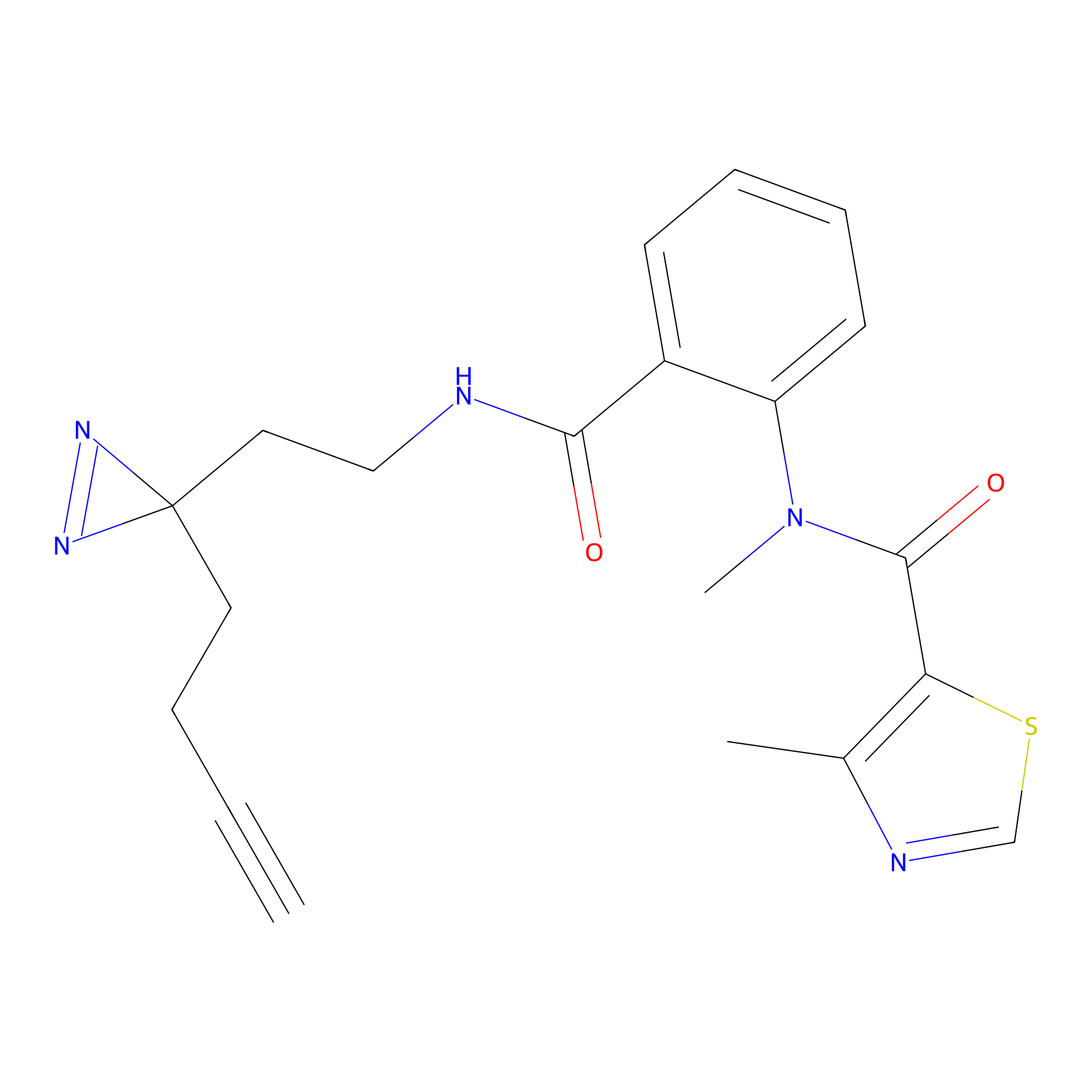
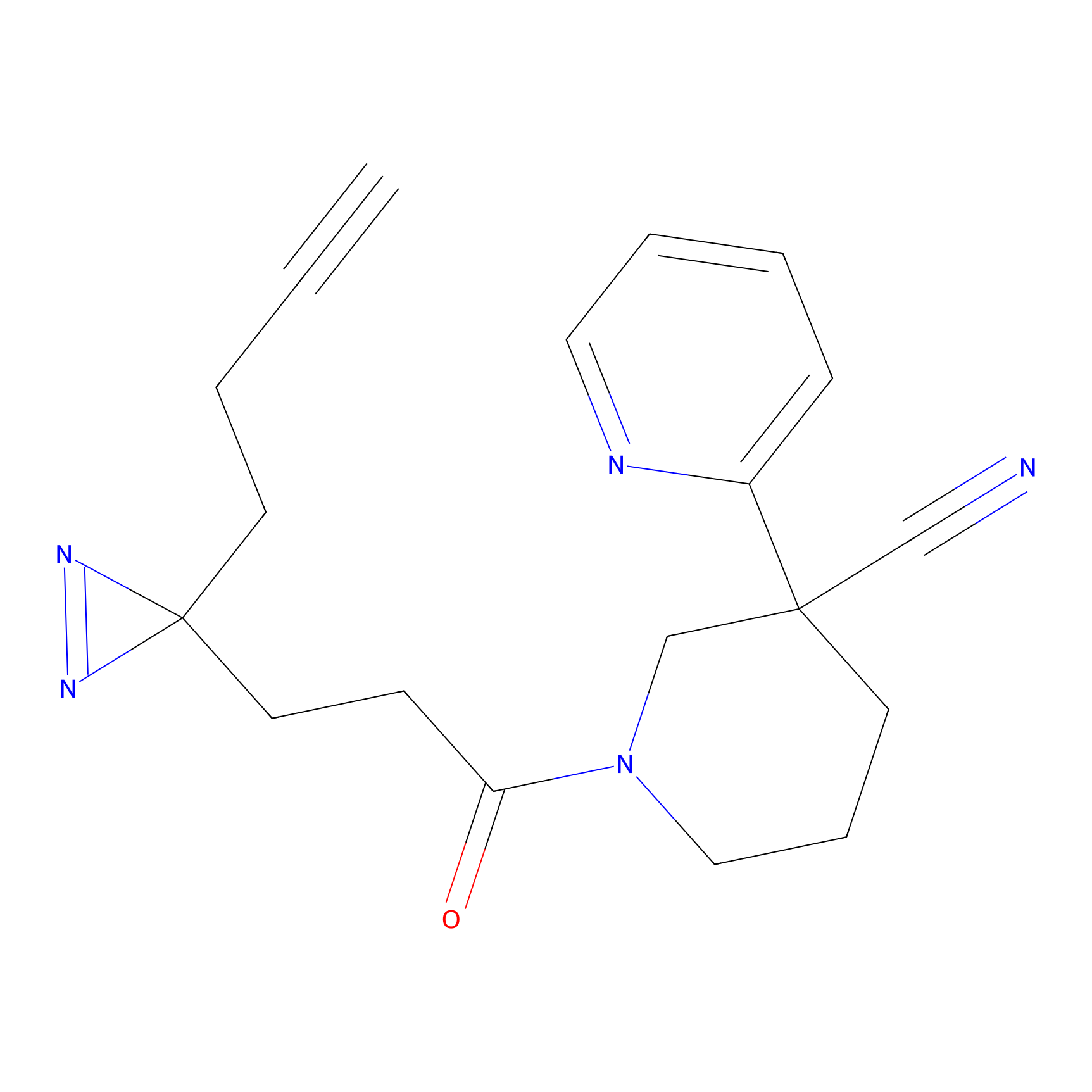
C-Sul Probe Info |
 |
8.50 | LDD0066 | [3] | |
|
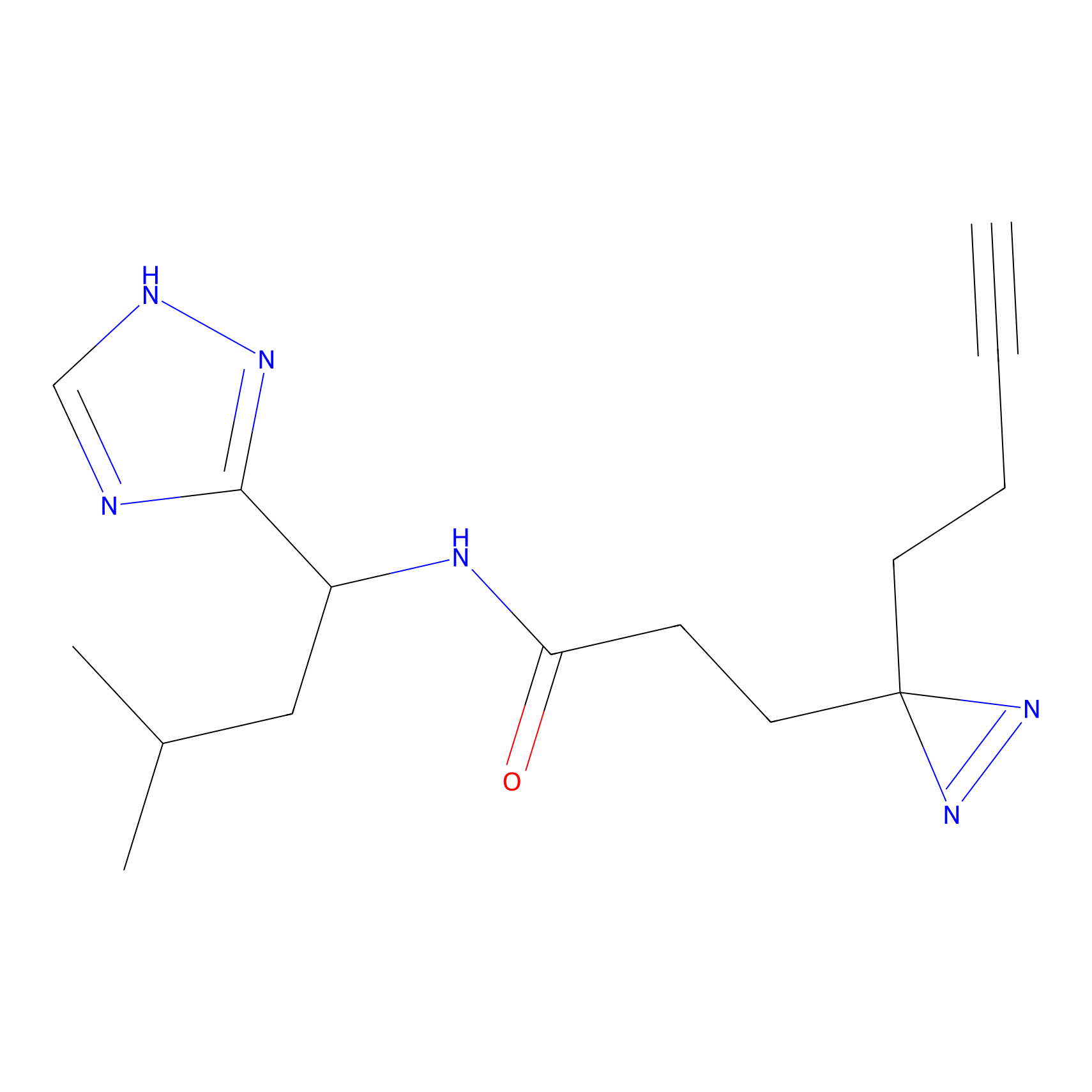
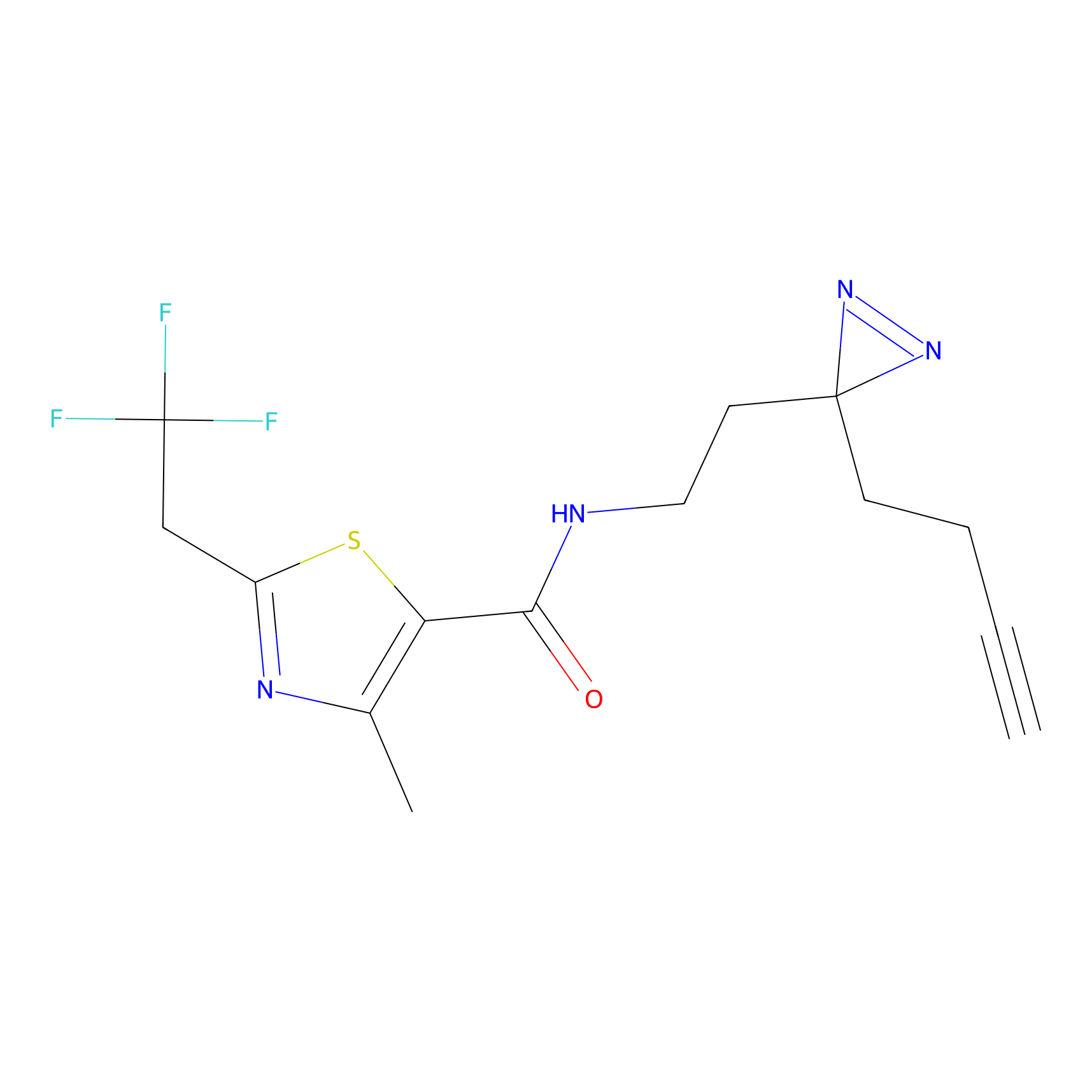
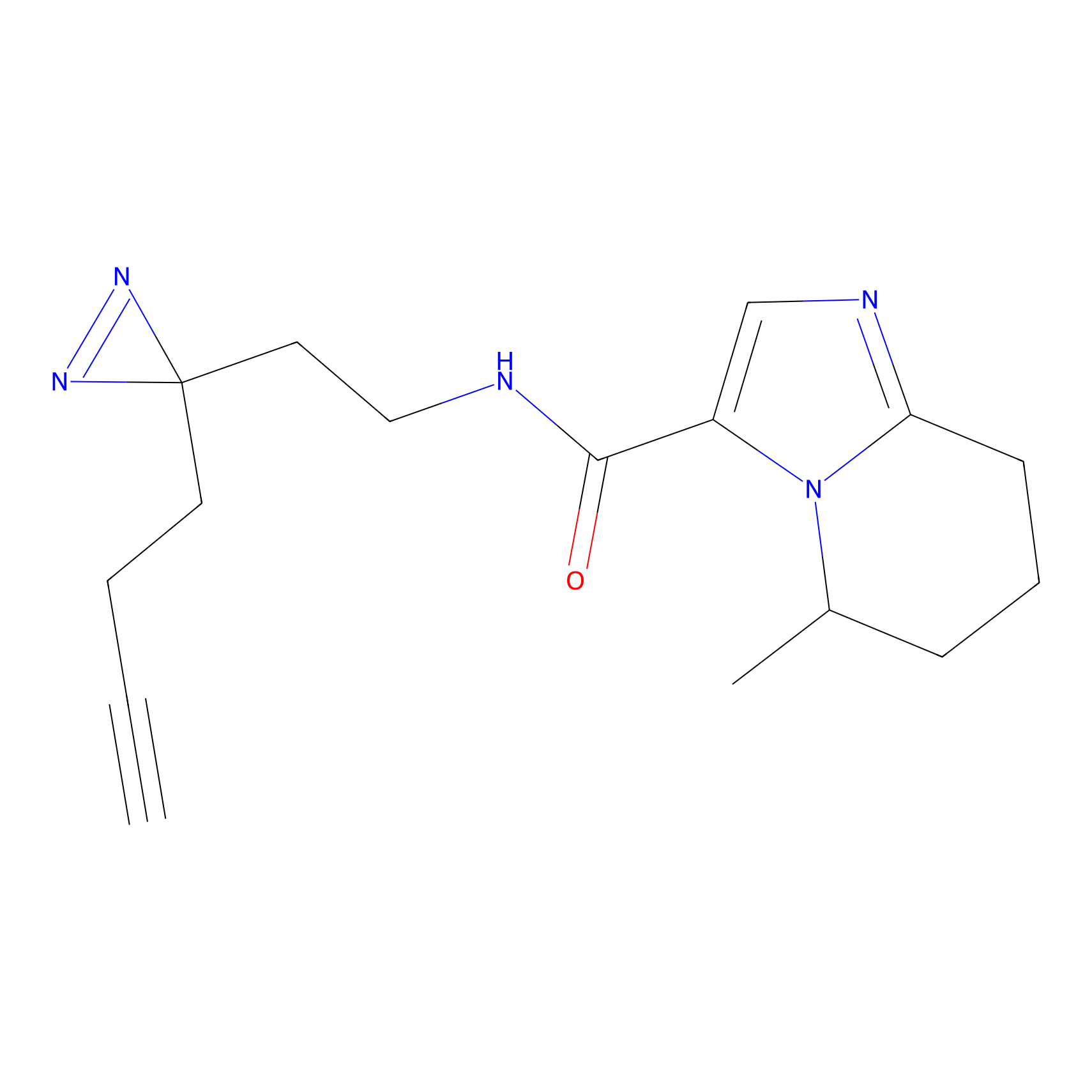
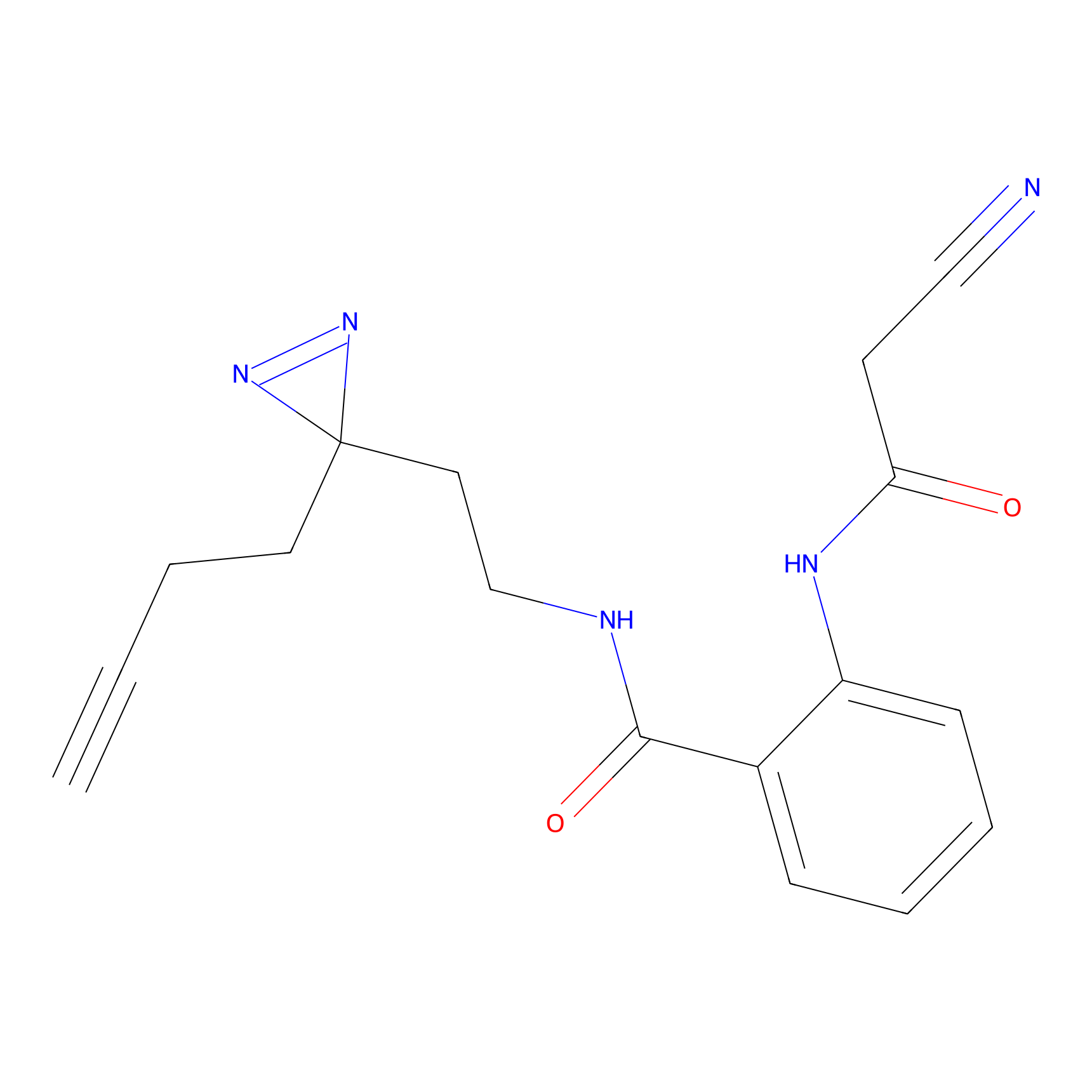
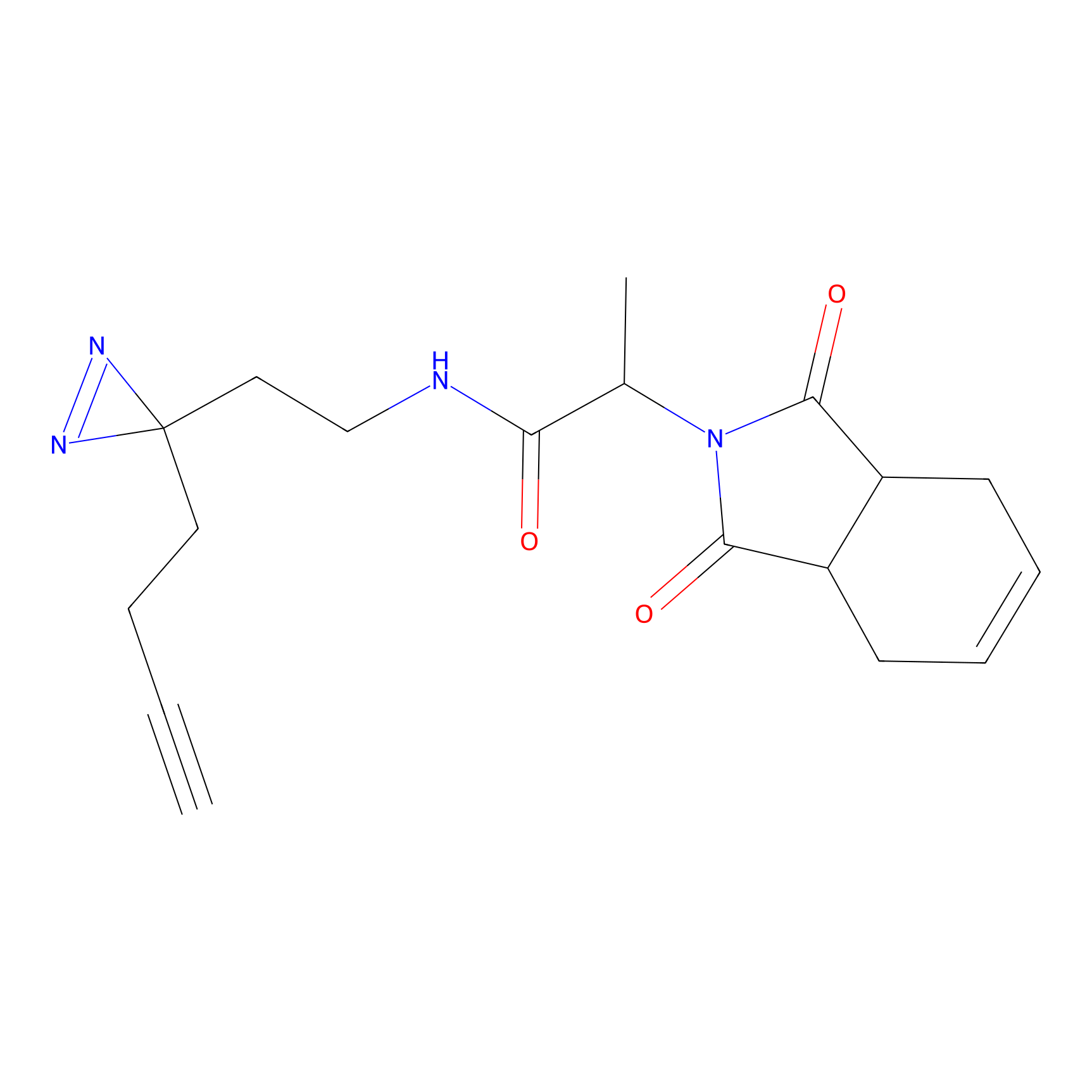
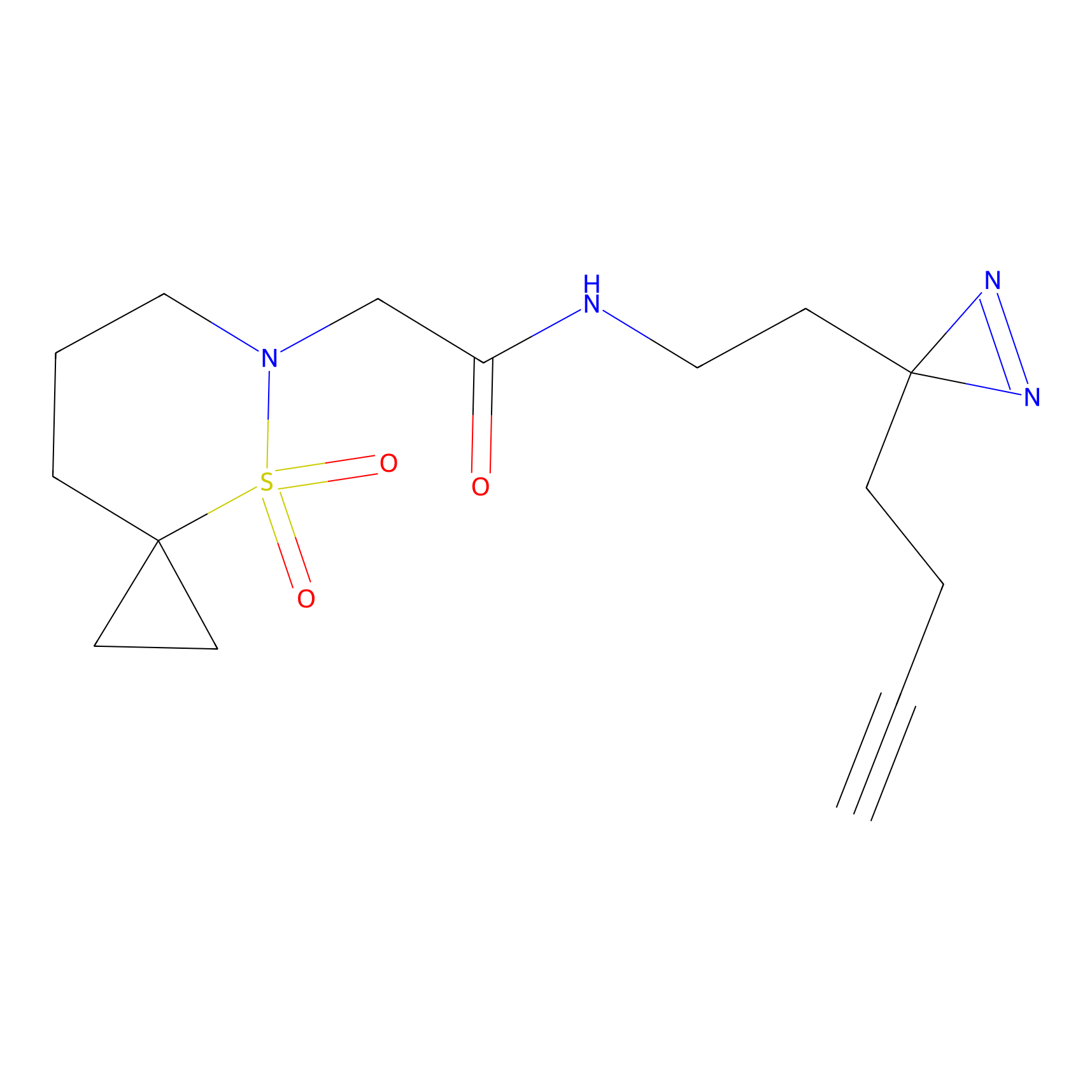
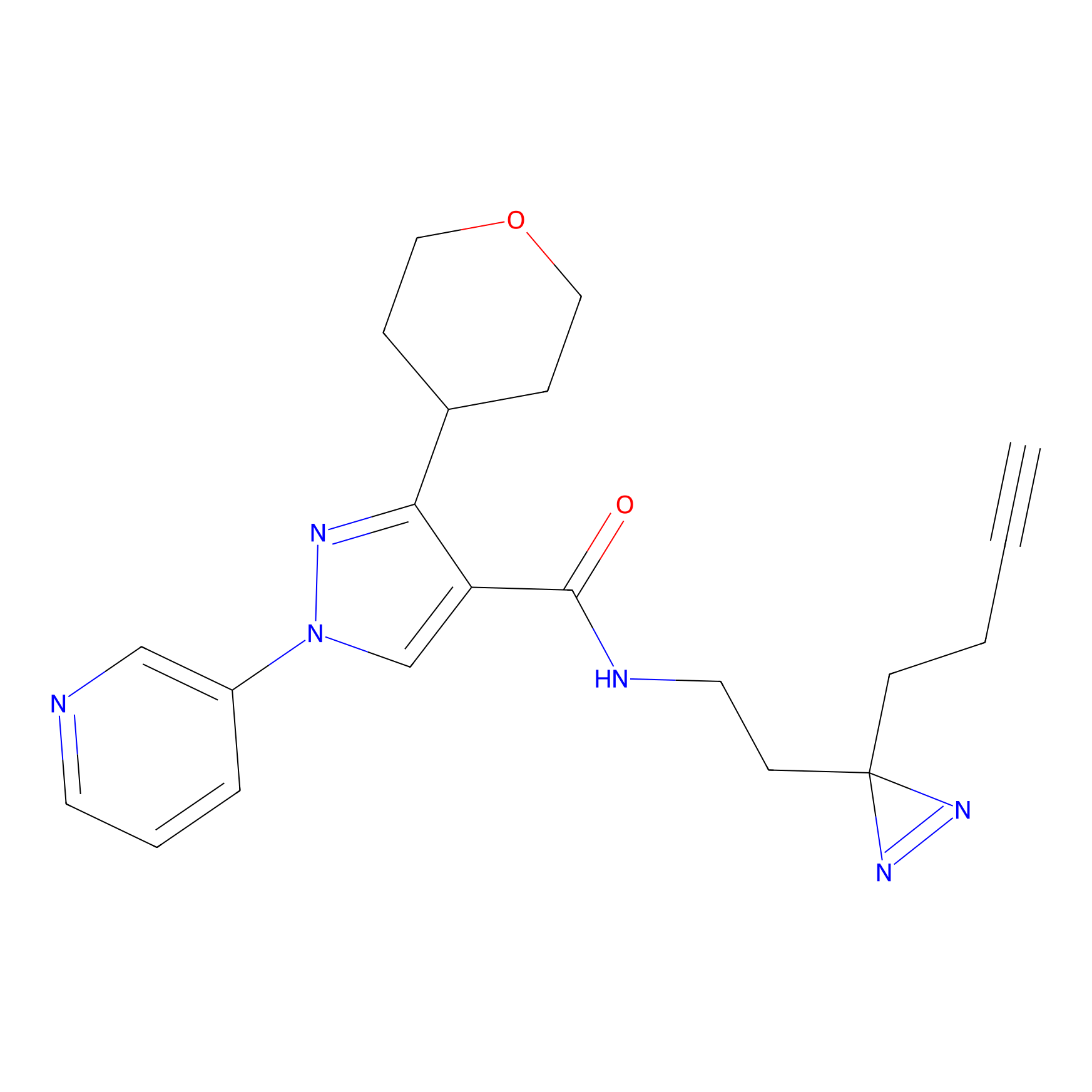
TH211 Probe Info |
 |
Y137(18.79); Y88(12.03) | LDD0257 | [4] | |
|
TH214 Probe Info |
 |
Y137(20.00) | LDD0258 | [4] | |
|
TH216 Probe Info |
 |
Y137(20.00); Y88(6.83) | LDD0259 | [4] | |
|
OPA-S-S-alkyne Probe Info |
 |
K136(0.96) | LDD3494 | [5] | |
|
Probe 1 Probe Info |
 |
Y55(28.65); Y88(180.36); Y142(16.69) | LDD3495 | [6] | |
|
P11 Probe Info |
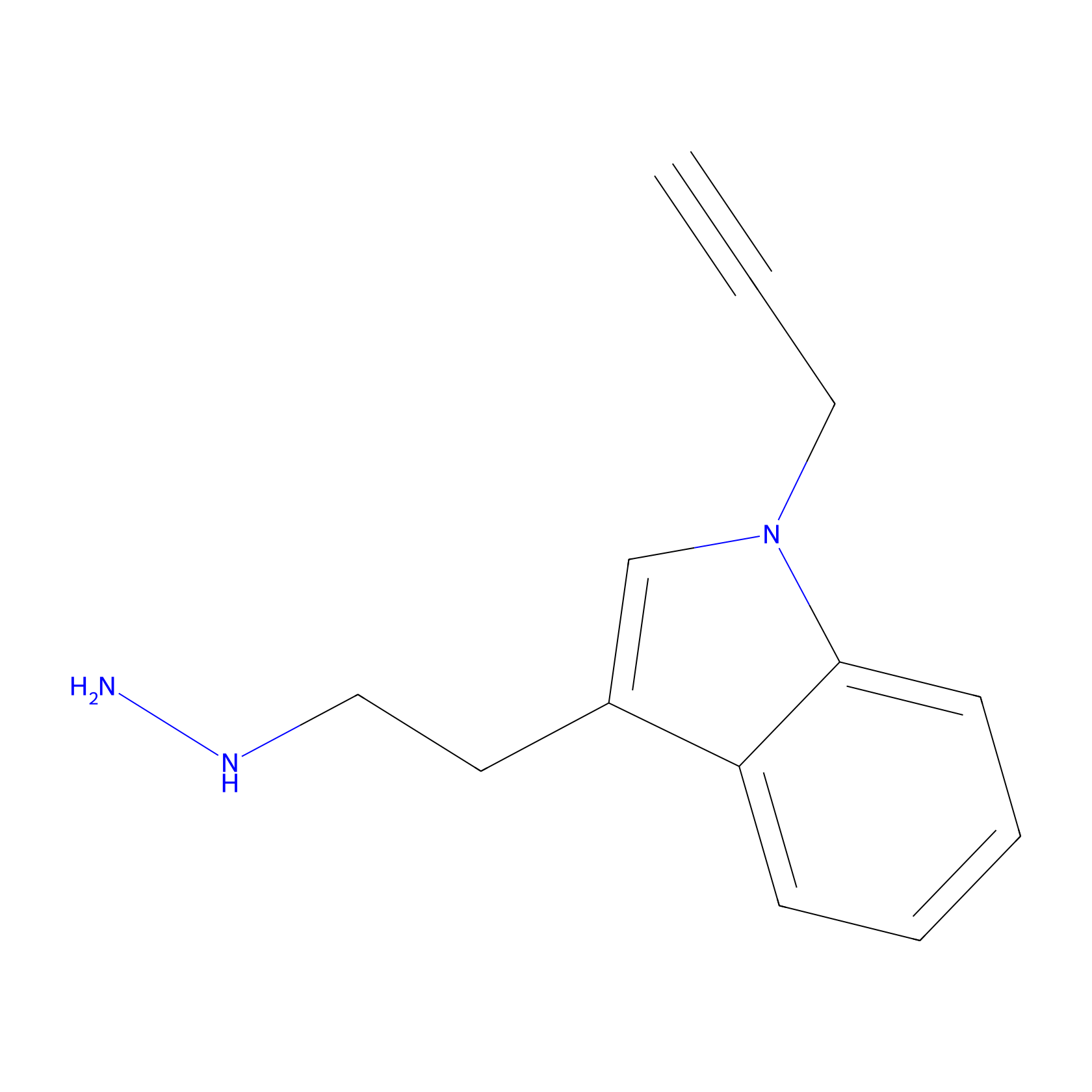 |
15.60 | LDD0201 | [7] | |
|
HPAP Probe Info |
 |
3.56 | LDD0062 | [8] | |
|
HHS-475 Probe Info |
 |
Y88(1.12); Y82(2.36) | LDD0264 | [9] | |
|
AMP probe Probe Info |
 |
N.A. | LDD0200 | [10] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [10] | |
|
SF Probe Info |
 |
Y88(0.00); Y82(0.00) | LDD0028 | [11] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0357 | [12] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [13] | |
|
HHS-482 Probe Info |
 |
Y88(0.65) | LDD2239 | [14] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0088 | C45 | HEK-293T | 15.60 | LDD0201 | [7] |
| LDCM0189 | Compound 16 | HEK-293T | 40.00 | LDD0492 | [16] |
| LDCM0185 | Compound 17 | HEK-293T | 11.81 | LDD0504 | [16] |
| LDCM0182 | Compound 18 | HEK-293T | 6.99 | LDD0501 | [16] |
| LDCM0184 | Compound 20 | HEK-293T | 6.83 | LDD0503 | [16] |
| LDCM0191 | Compound 21 | HEK-293T | 9.13 | LDD0493 | [16] |
| LDCM0187 | Compound 32 | HEK-293T | 27.69 | LDD0507 | [16] |
| LDCM0190 | Compound 34 | HEK-293T | 16.67 | LDD0497 | [16] |
| LDCM0192 | Compound 35 | HEK-293T | 11.05 | LDD0491 | [16] |
| LDCM0193 | Compound 36 | HEK-293T | 7.75 | LDD0494 | [16] |
| LDCM0194 | Compound 37 | HEK-293T | 12.20 | LDD0498 | [16] |
| LDCM0195 | Compound 38 | HEK-293T | 11.43 | LDD0499 | [16] |
| LDCM0196 | Compound 39 | HEK-293T | 11.11 | LDD0496 | [16] |
| LDCM0197 | Compound 40 | HEK-293T | 13.33 | LDD0495 | [16] |
| LDCM0181 | Compound 41 | HEK-293T | 5.24 | LDD0502 | [16] |
| LDCM0183 | Compound 42 | HEK-293T | 3.86 | LDD0500 | [16] |
| LDCM0116 | HHS-0101 | DM93 | Y88(1.12); Y82(2.36) | LDD0264 | [9] |
| LDCM0117 | HHS-0201 | DM93 | Y88(1.43); Y82(3.42) | LDD0265 | [9] |
| LDCM0118 | HHS-0301 | DM93 | Y88(0.61); Y82(0.97) | LDD0266 | [9] |
| LDCM0119 | HHS-0401 | DM93 | Y88(0.73); Y82(1.66) | LDD0267 | [9] |
| LDCM0120 | HHS-0701 | DM93 | Y88(1.02); Y82(8.30) | LDD0268 | [9] |
| LDCM0073 | Kambe_cp3 | PC-3 | 3.23 | LDD0129 | [17] |
| LDCM0079 | kambe_cp64 | PC-3 | 2.29 | LDD0135 | [17] |
| LDCM0078 | kambe_cp70 | PC-3 | 2.05 | LDD0134 | [17] |
| LDCM0014 | Panhematin | HEK-293T | 3.56 | LDD0062 | [8] |
References