Details of the Target
General Information of Target
Probe(s) Labeling This Target
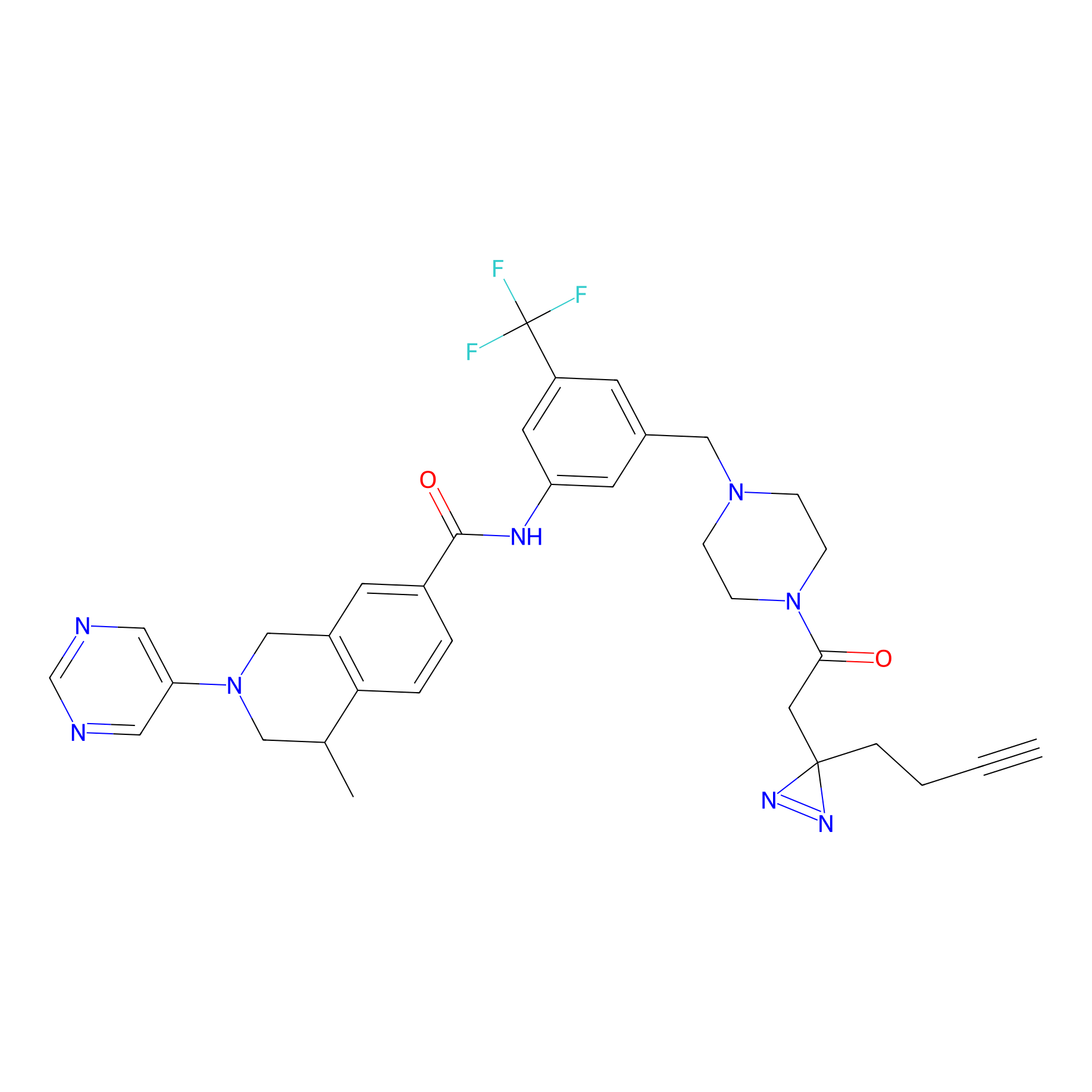
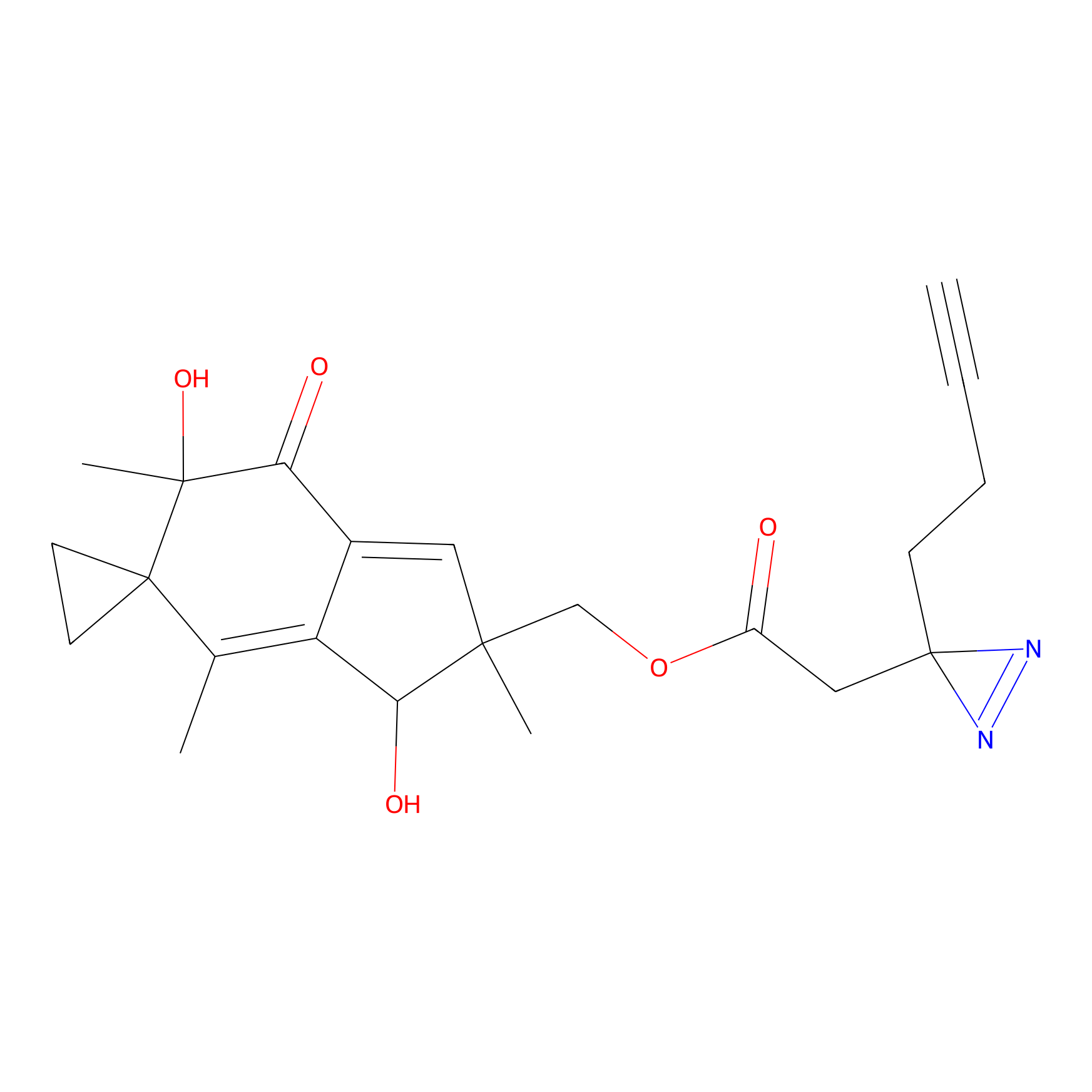
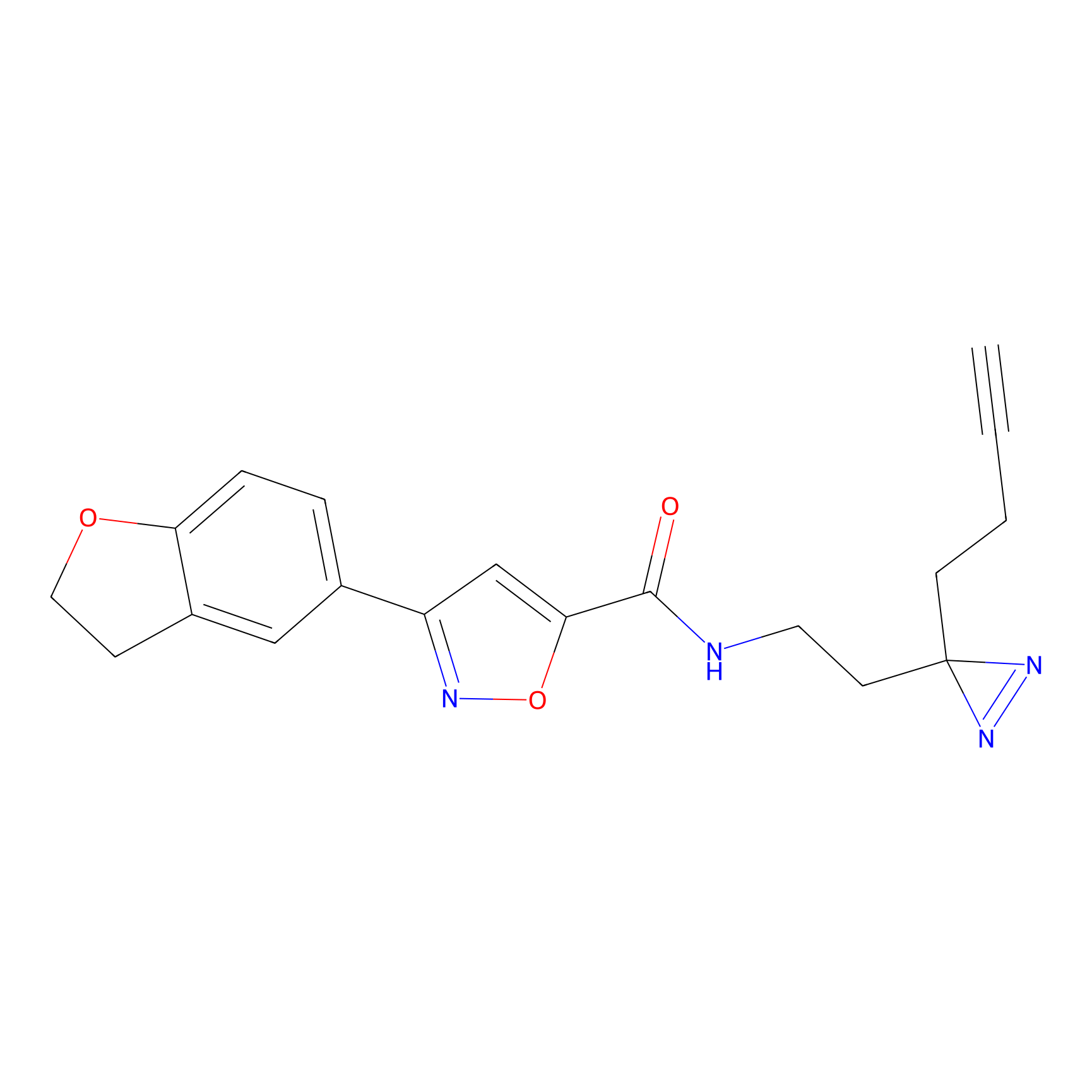
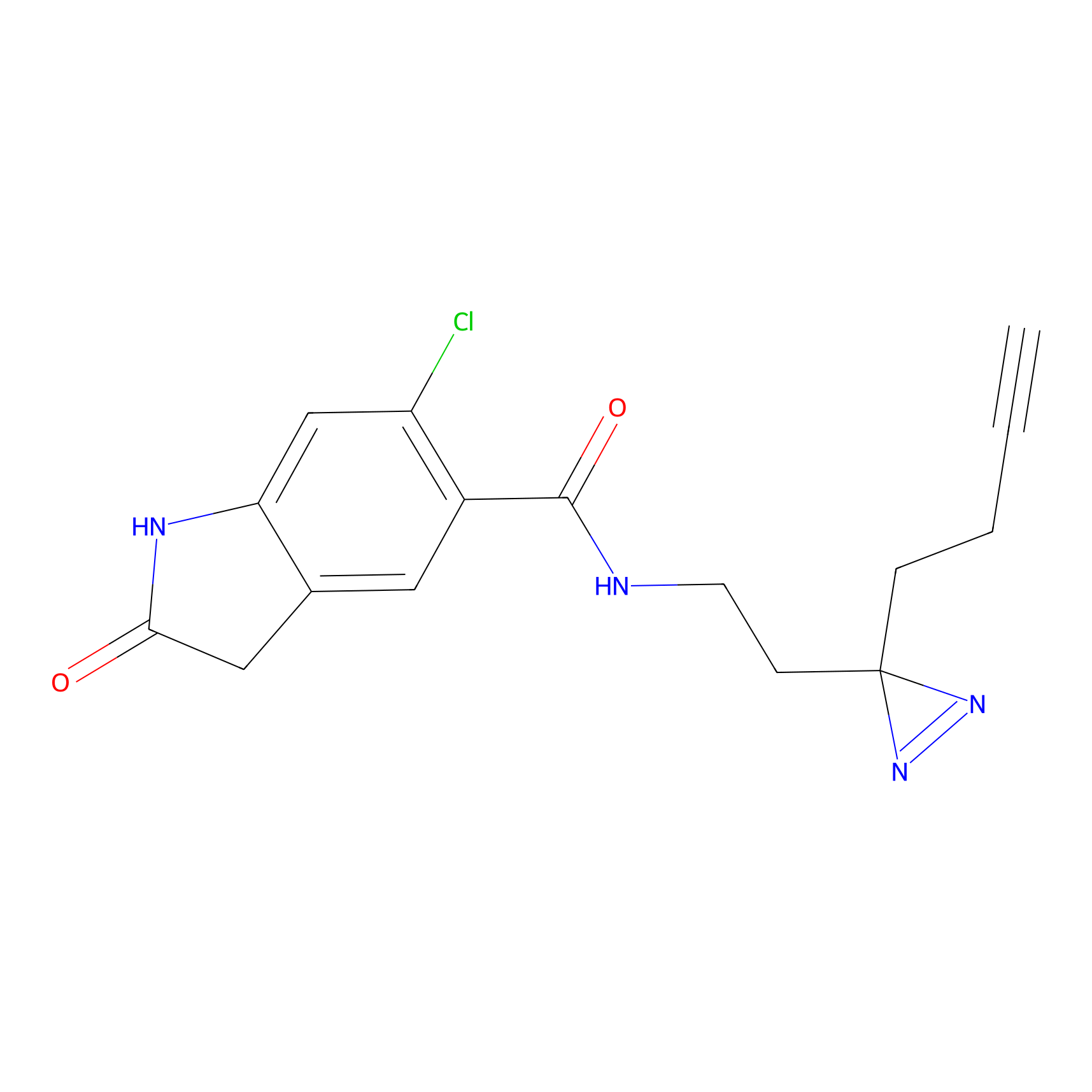
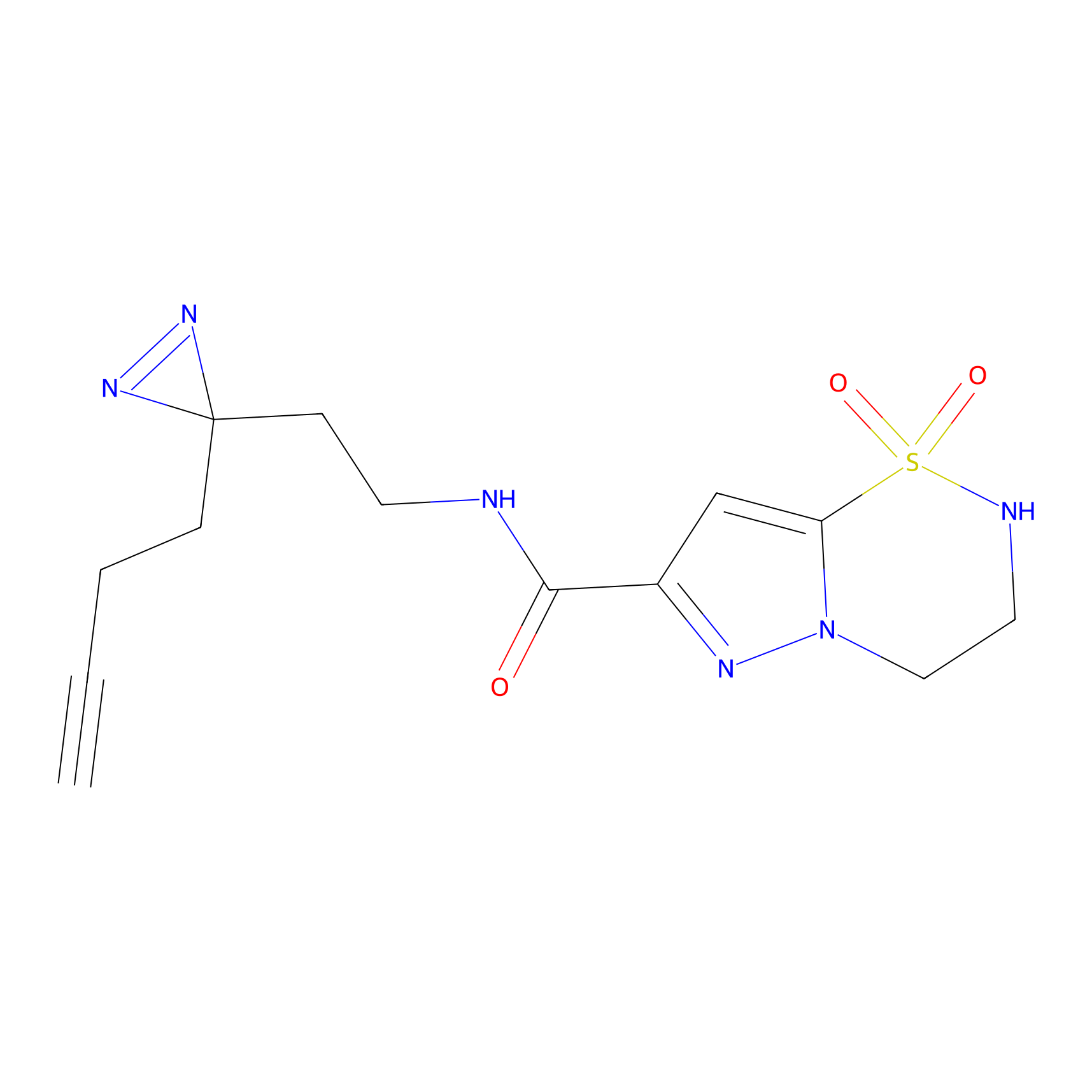
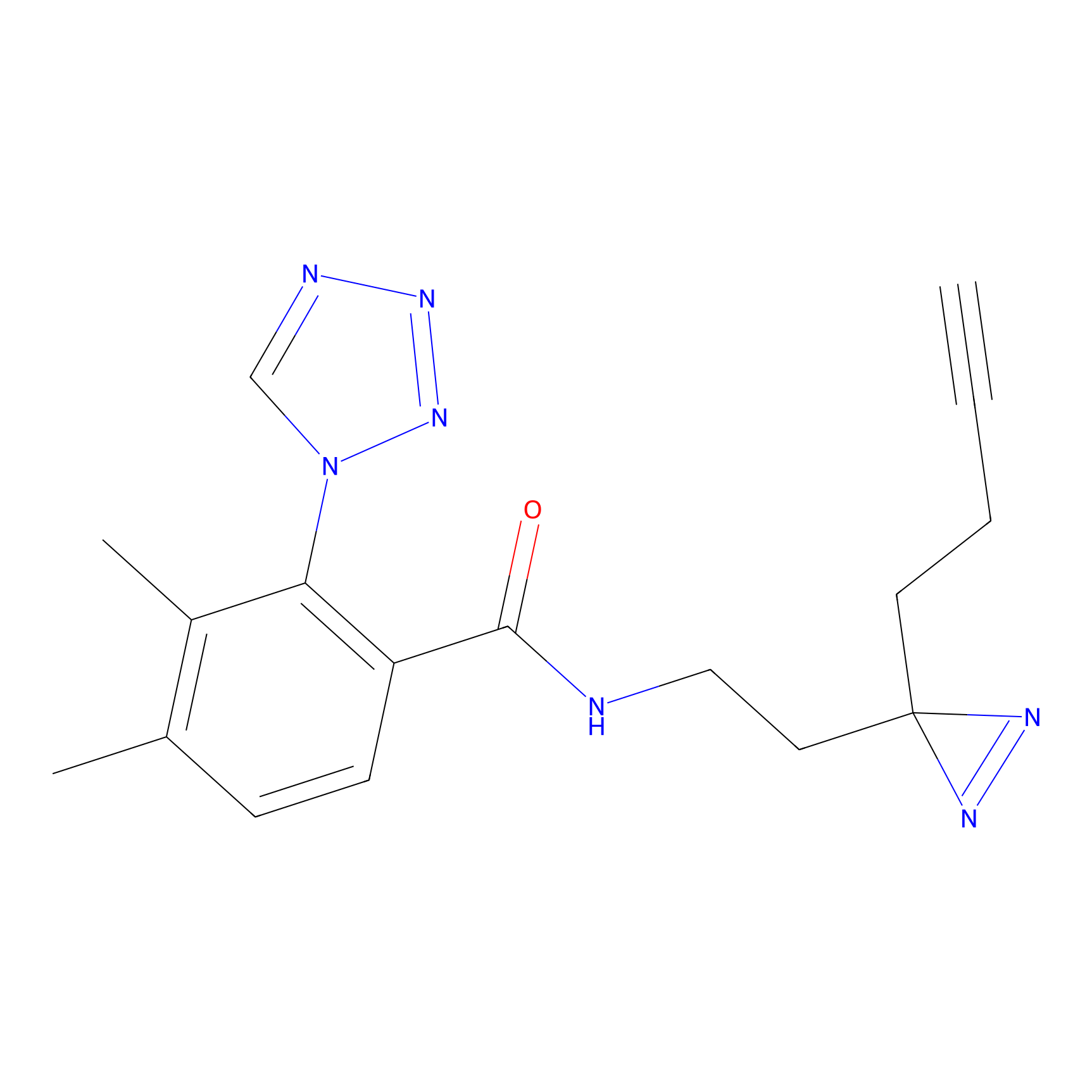
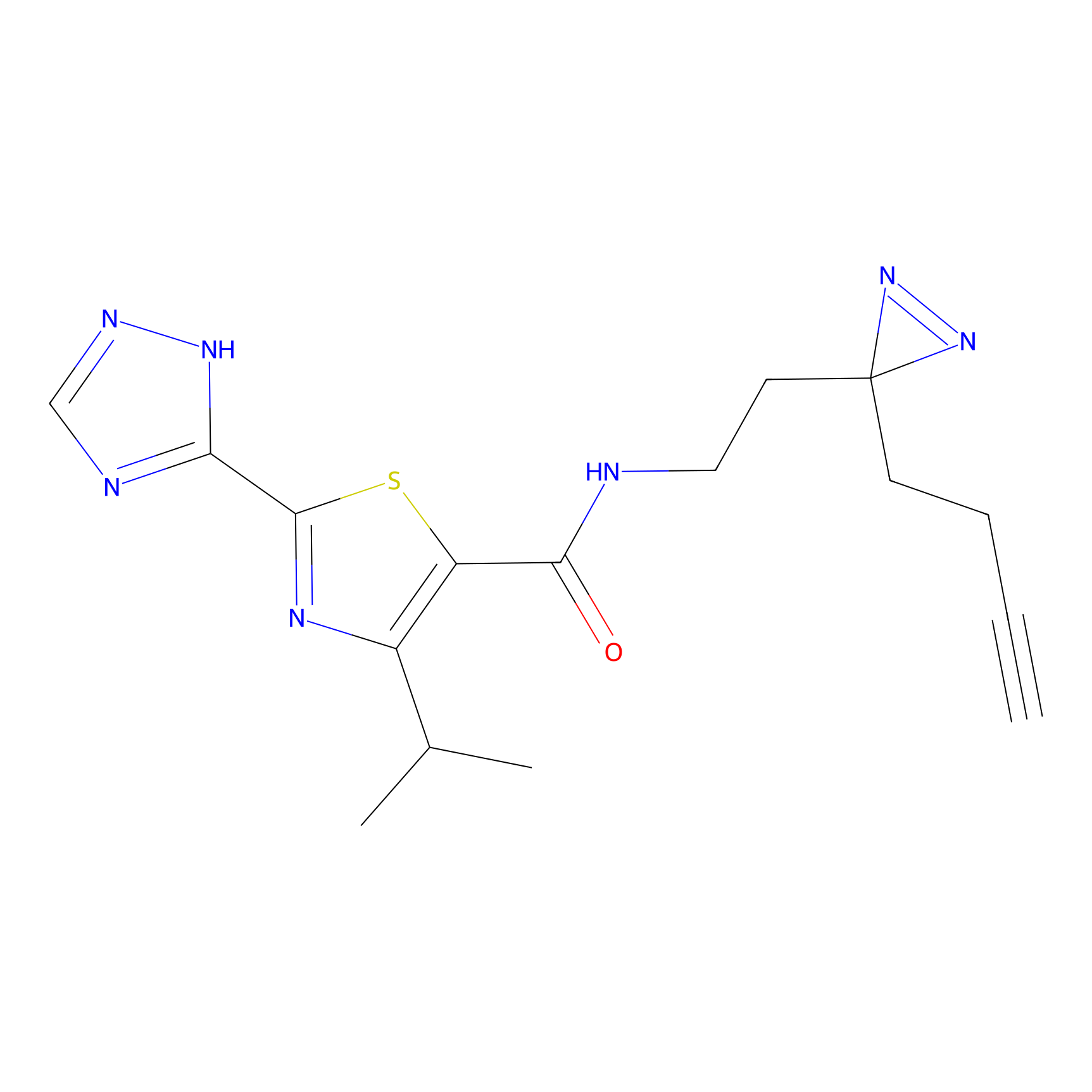
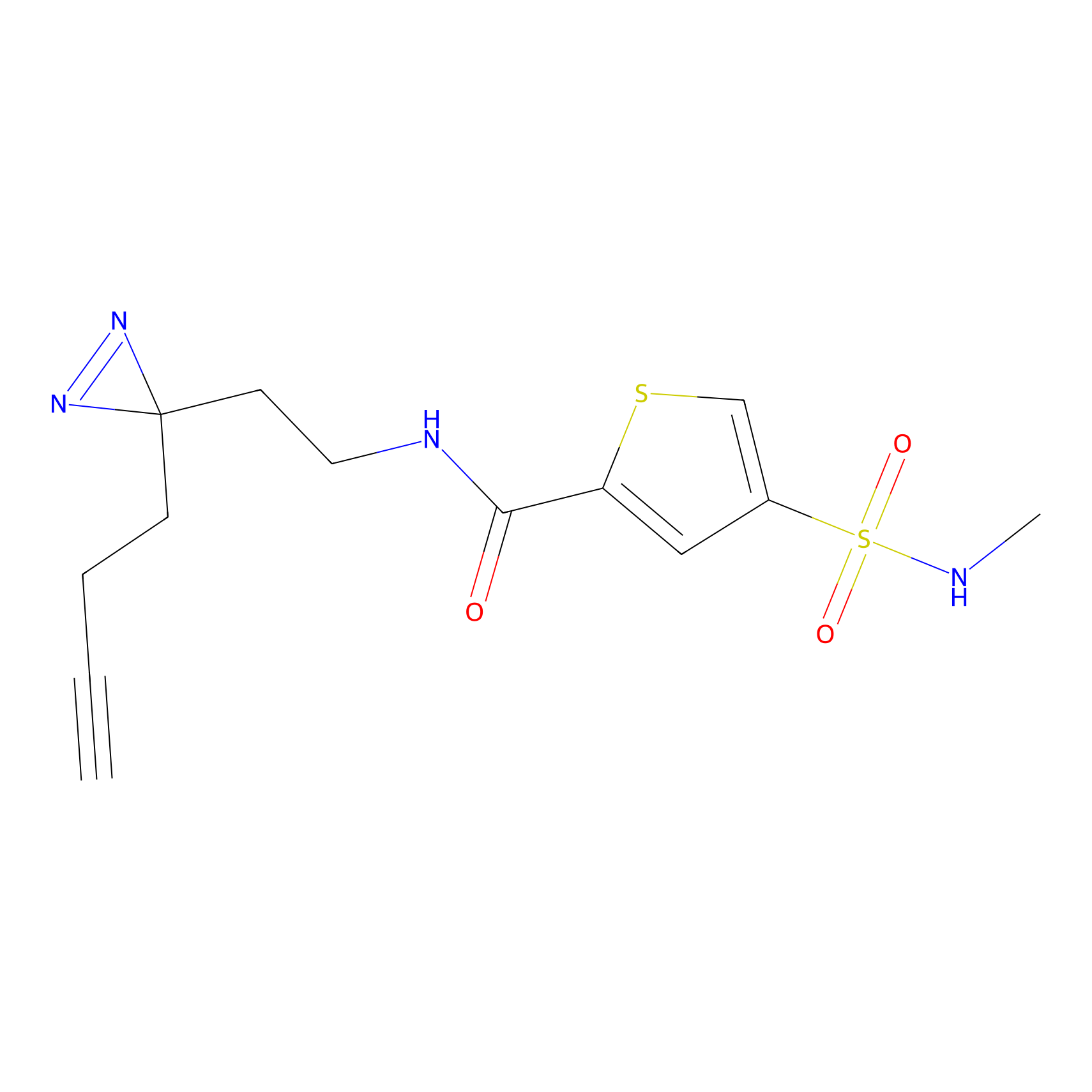
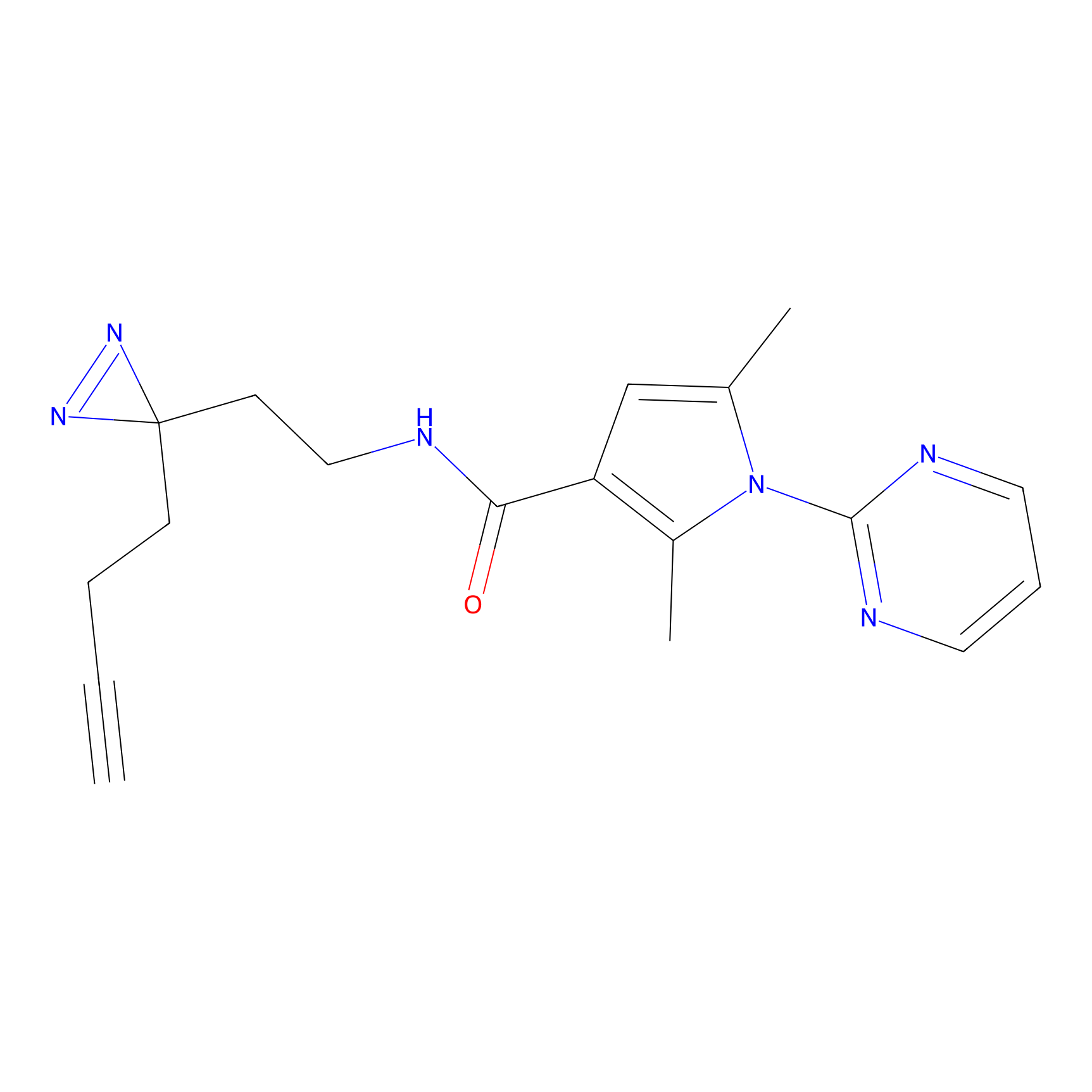
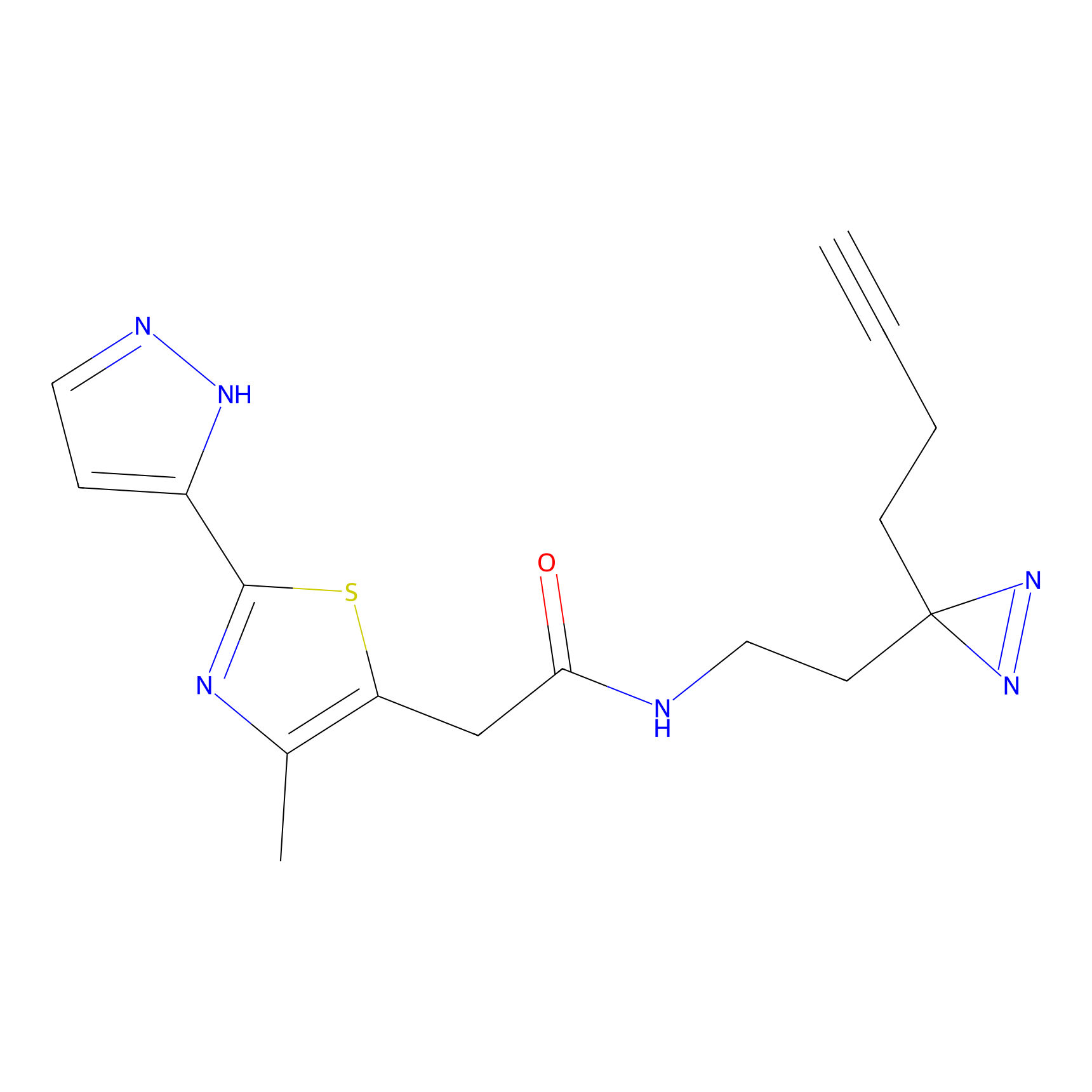
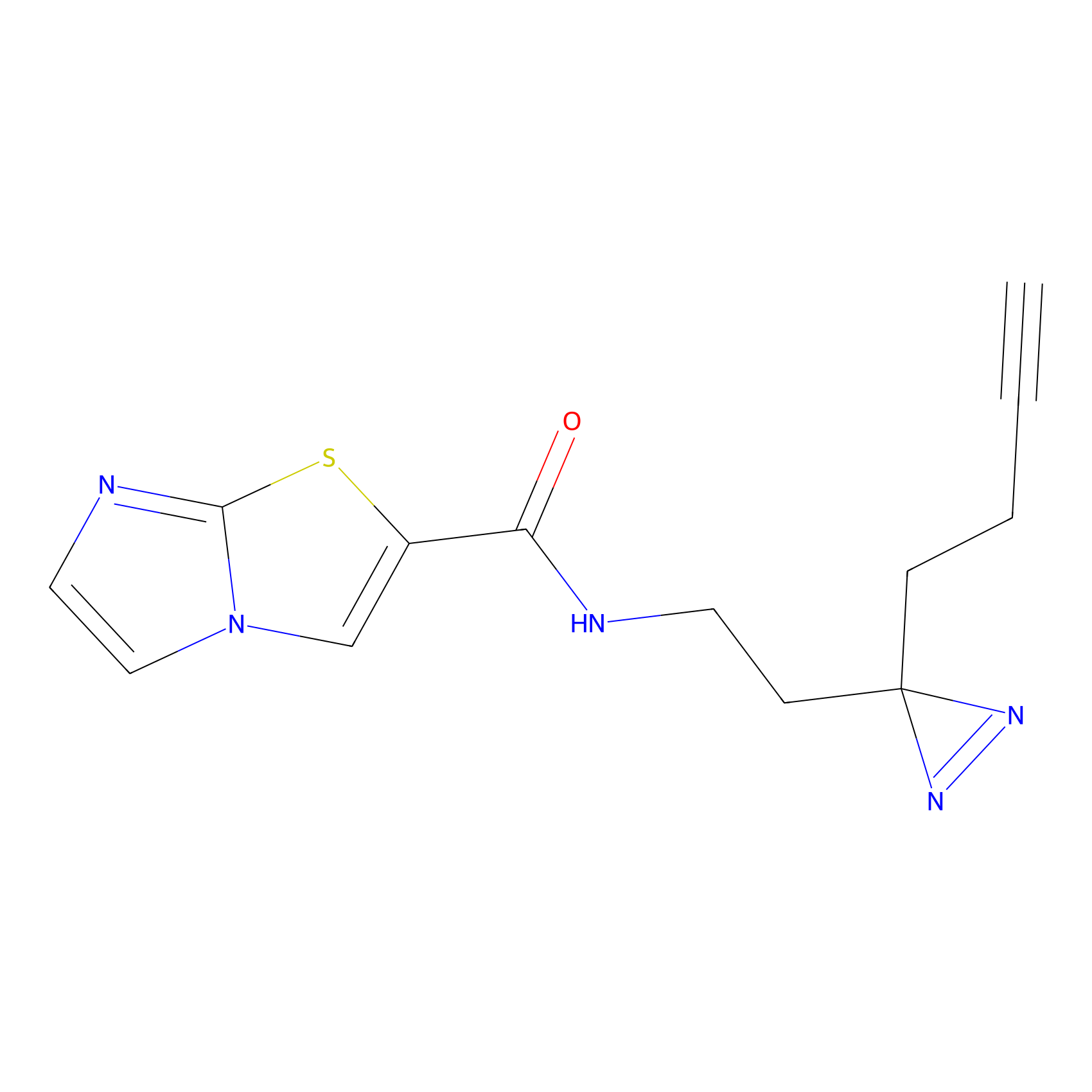
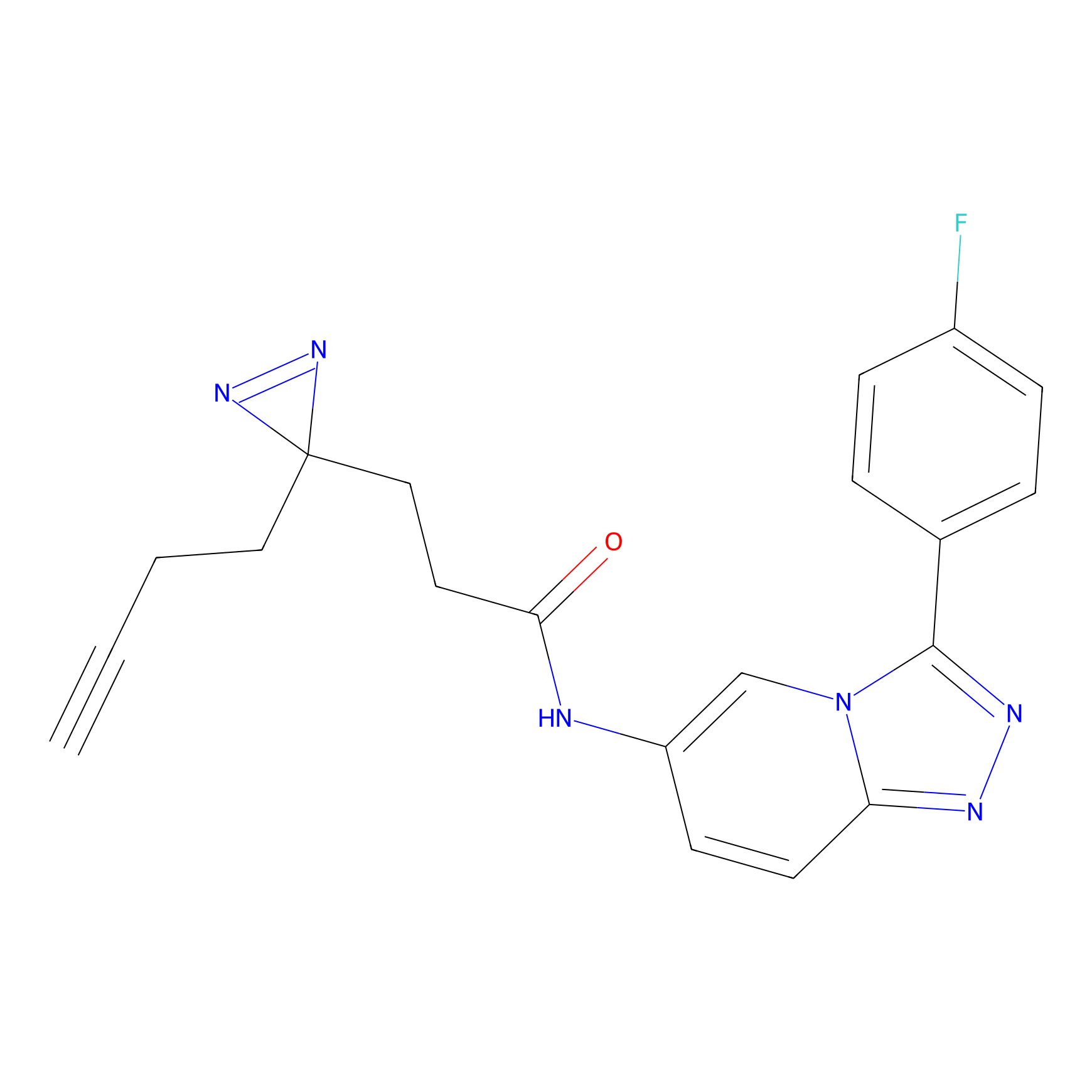
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
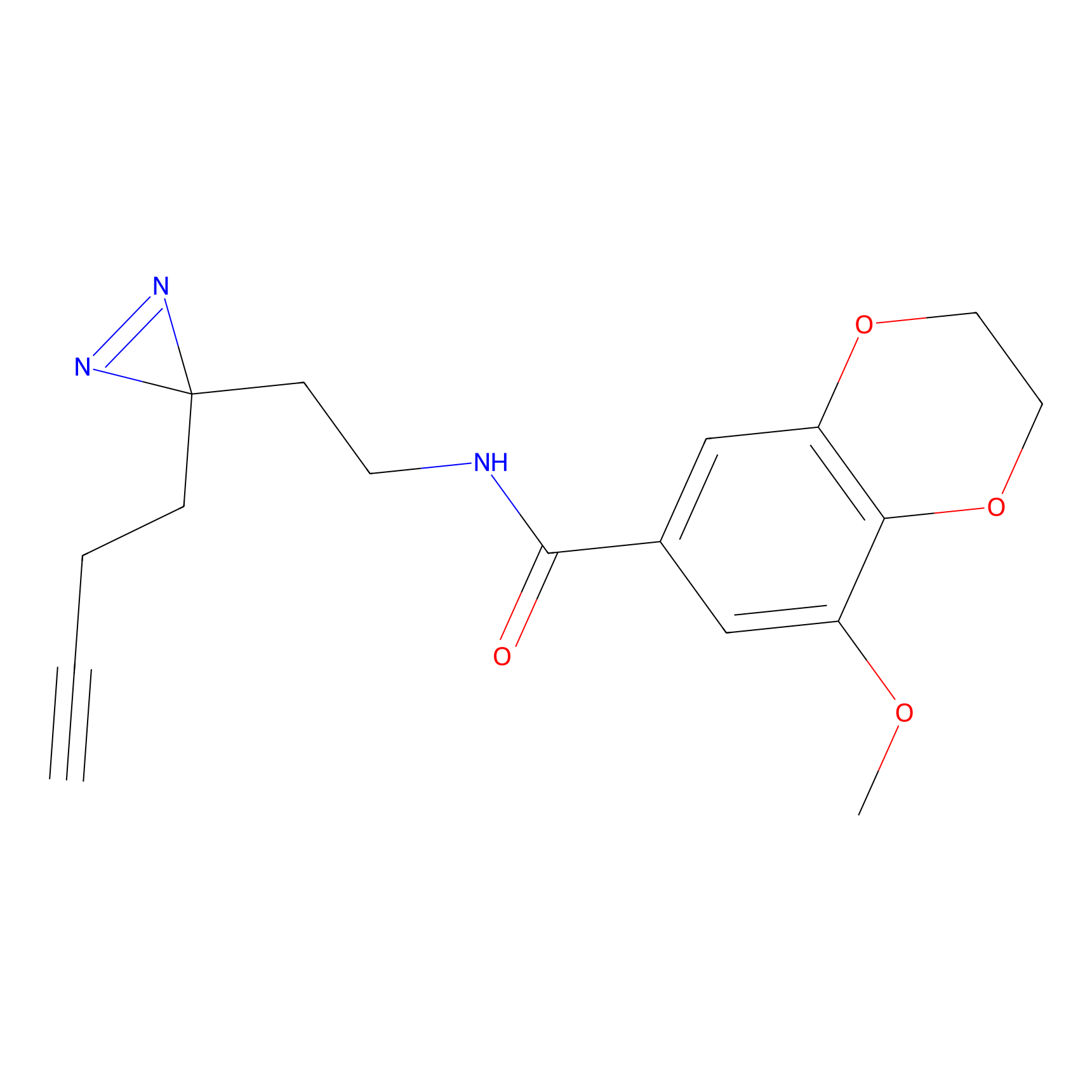
AZ-5 Probe Info |
 |
3.52 | LDD0394 | [1] | |
|
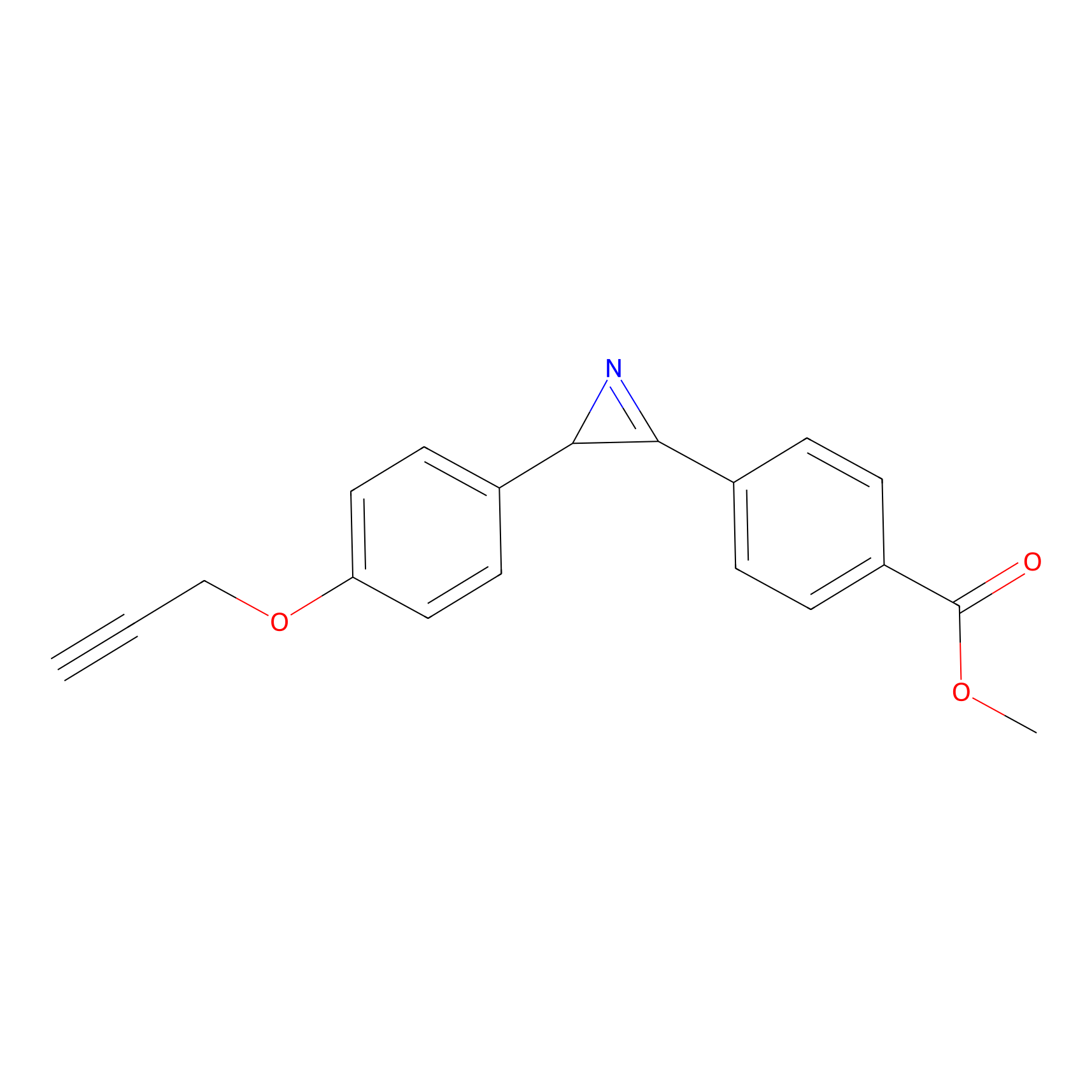
ILS-1 Probe Info |
 |
4.78; 6.61 | LDD0415 | [2] | |
|
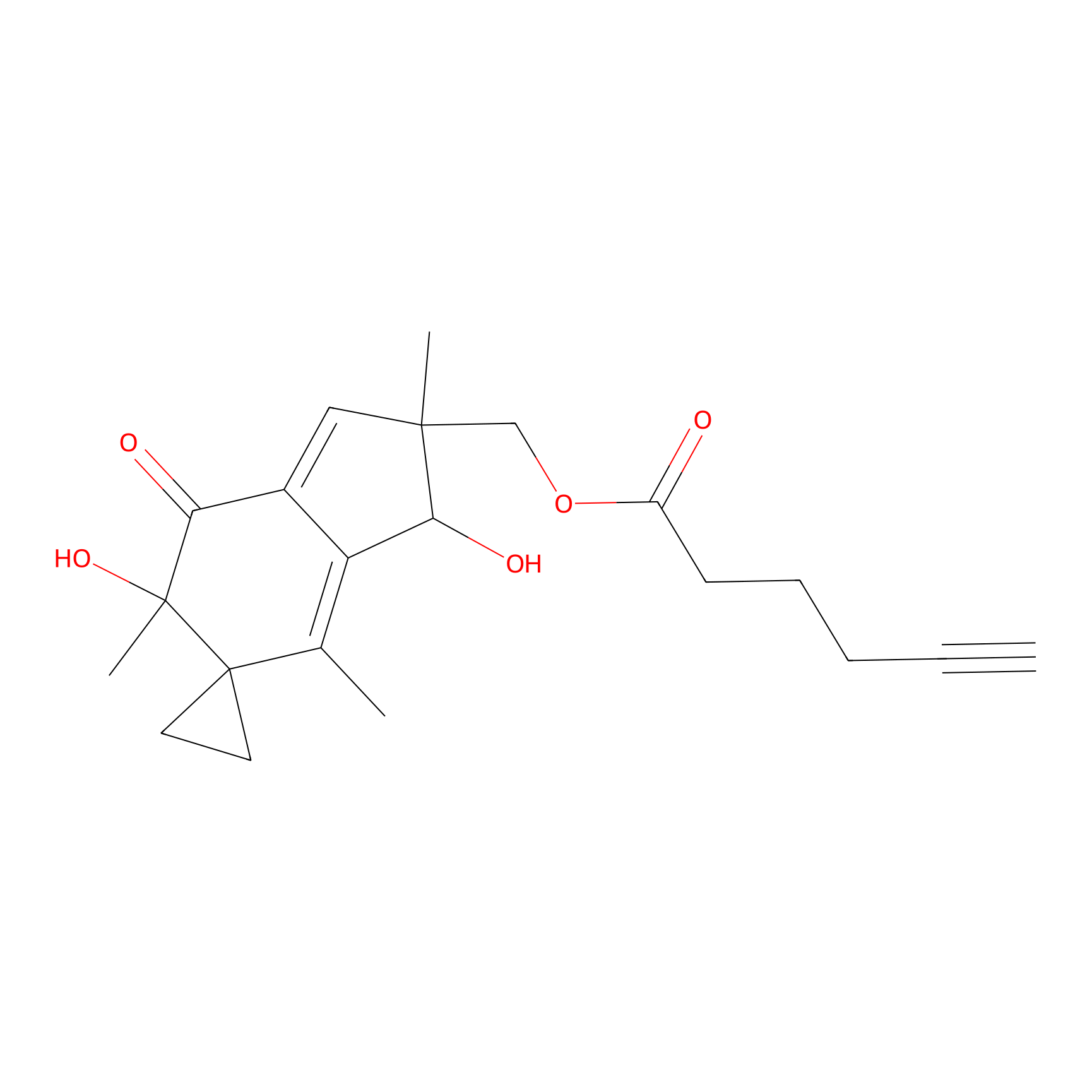
AZ-9 Probe Info |
 |
4.87 | LDD0393 | [1] | |
|
FBPP2 Probe Info |
 |
2.60 | LDD0324 | [3] | |
|
YN-1 Probe Info |
 |
33.21 | LDD0444 | [4] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [4] | |
|
ONAyne Probe Info |
 |
K111(4.12); K167(2.90); K187(1.90); K215(3.78) | LDD0274 | [5] | |
|
OPA-S-S-alkyne Probe Info |
 |
K372(2.29); K187(3.47); K119(6.22); K118(6.22) | LDD3494 | [6] | |
|
5E-2FA Probe Info |
 |
H183(0.00); H344(0.00) | LDD2235 | [7] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [8] | |
|
m-APA Probe Info |
 |
H183(0.00); H344(0.00); H128(0.00) | LDD2231 | [7] | |
|
ATP probe Probe Info |
 |
K131(0.00); K317(0.00); K417(0.00); K207(0.00) | LDD0035 | [9] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [10] | |
|
STPyne Probe Info |
 |
K187(0.00); K426(0.00) | LDD0009 | [10] | |
|
Acrolein Probe Info |
 |
H183(0.00); H128(0.00); H217(0.00) | LDD0217 | [11] | |
|
Crotonaldehyde Probe Info |
 |
H183(0.00); H128(0.00); H217(0.00) | LDD0219 | [11] | |
|
Methacrolein Probe Info |
 |
H217(0.00); H183(0.00) | LDD0218 | [11] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [12] | |
|
HHS-465 Probe Info |
 |
Y13(0.00); Y36(0.00); Y24(0.00) | LDD2240 | [13] | |
|
HHS-475 Probe Info |
 |
Y13(0.85); Y168(0.66); Y24(0.81); Y256(0.78) | LDD2238 | [14] | |
|
HHS-482 Probe Info |
 |
Y13(0.55); Y168(0.46); Y24(0.29); Y256(0.91) | LDD2239 | [14] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H183(0.00); H128(0.00); H217(0.00) | LDD0222 | [11] |
| LDCM0107 | IAA | HeLa | H183(0.00); H128(0.00); H217(0.00) | LDD0221 | [11] |
| LDCM0109 | NEM | HeLa | H183(0.00); H128(0.00); H217(0.00); H344(0.00) | LDD0223 | [11] |
| LDCM0112 | W16 | Hep-G2 | N315(0.61); K317(0.67) | LDD0239 | [12] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Beclin-1 (BECN1) | Beclin family | Q14457 | |||
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Leukocyte immunoglobulin-like receptor subfamily B member 3 (LILRB3) | . | O75022 | |||
Other
References