Details of the Target
General Information of Target
| Target ID | LDTP06365 | |||||
|---|---|---|---|---|---|---|
| Target Name | Transmembrane emp24 domain-containing protein 2 (TMED2) | |||||
| Gene Name | TMED2 | |||||
| Gene ID | 10959 | |||||
| Synonyms |
RNP24; Transmembrane emp24 domain-containing protein 2; Membrane protein p24A; p24; p24 family protein beta-1; p24beta1 |
|||||
| 3D Structure | ||||||
| Sequence |
MVTLAELLVLLAALLATVSGYFVSIDAHAEECFFERVTSGTKMGLIFEVAEGGFLDIDVE
ITGPDNKGIYKGDRESSGKYTFAAHMDGTYKFCFSNRMSTMTPKIVMFTIDIGEAPKGQD METEAHQNKLEEMINELAVAMTAVKHEQEYMEVRERIHRAINDNTNSRVVLWSFFEALVL VAMTLGQIYYLKRFFEVRRVV |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
EMP24/GP25L family
|
|||||
| Subcellular location |
Cytoplasmic vesicle membrane
|
|||||
| Function |
Involved in vesicular protein trafficking. Mainly functions in the early secretory pathway but also in post-Golgi membranes. Thought to act as cargo receptor at the lumenal side for incorporation of secretory cargo molecules into transport vesicles and to be involved in vesicle coat formation at the cytoplasmic side. In COPII vesicle-mediated anterograde transport involved in the transport of GPI-anchored proteins and proposed to act together with TMED10 as their cargo receptor; the function specifically implies SEC24C and SEC24D of the COPII vesicle coat and lipid raft-like microdomains of the ER. Recognizes GPI anchors structural remodeled in the ER by PGAP1 and MPPE1. In COPI vesicle-mediated retrograde transport inhibits the GTPase-activating activity of ARFGAP1 towards ARF1 thus preventing immature uncoating and allowing cargo selection to take place. Involved in trafficking of G protein-coupled receptors (GPCRs). Regulates F2RL1, OPRM1 and P2RY4 exocytic trafficking from the Golgi to the plasma membrane thus contributing to receptor resensitization. Facilitates CASR maturation and stabilization in the early secretory pathway and increases CASR plasma membrane targeting. Proposed to be involved in organization of intracellular membranes such as the maintenance of the Golgi apparatus. May also play a role in the biosynthesis of secreted cargo such as eventual processing.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
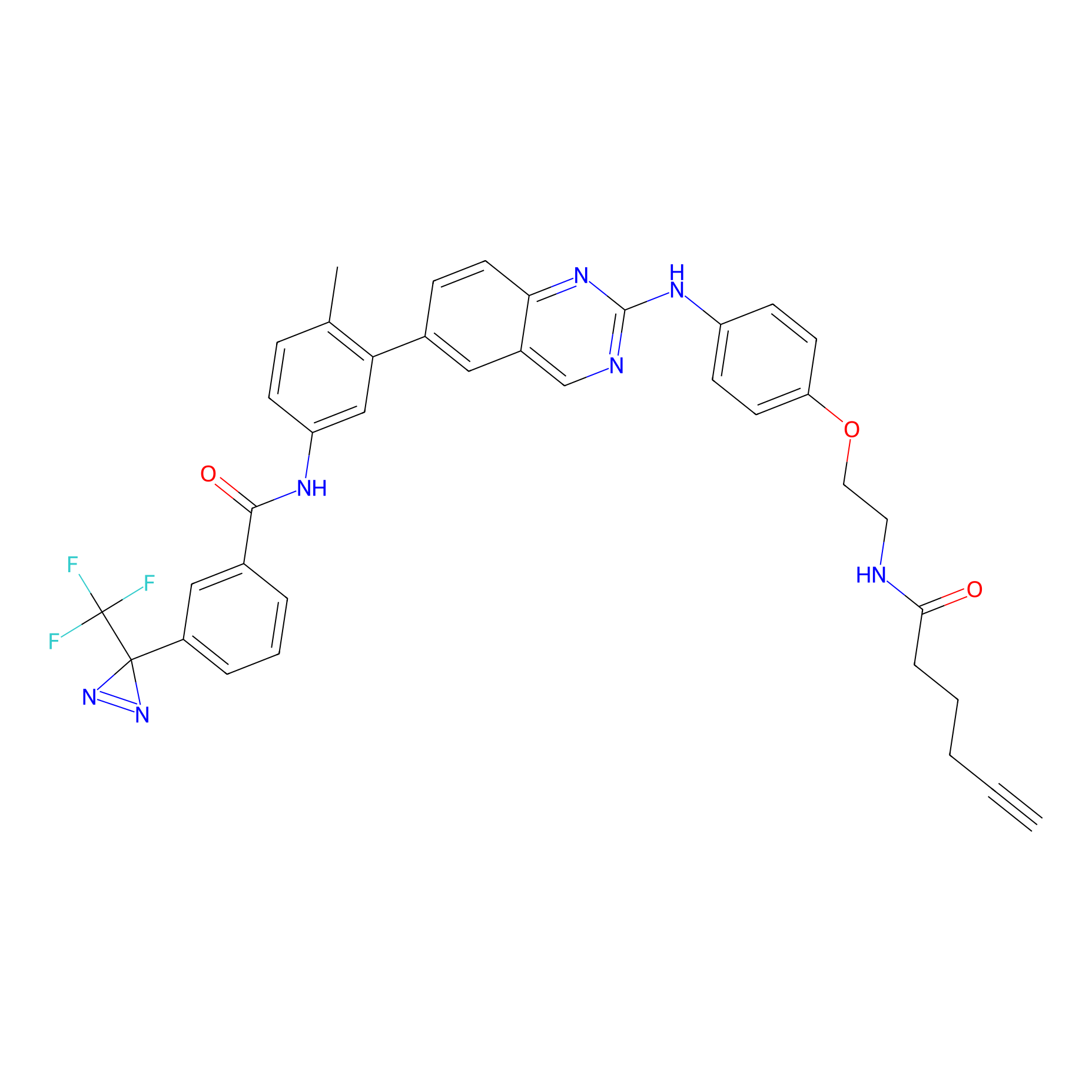
ONAyne Probe Info |
 |
K71(0.84) | LDD0274 | [1] | |
|
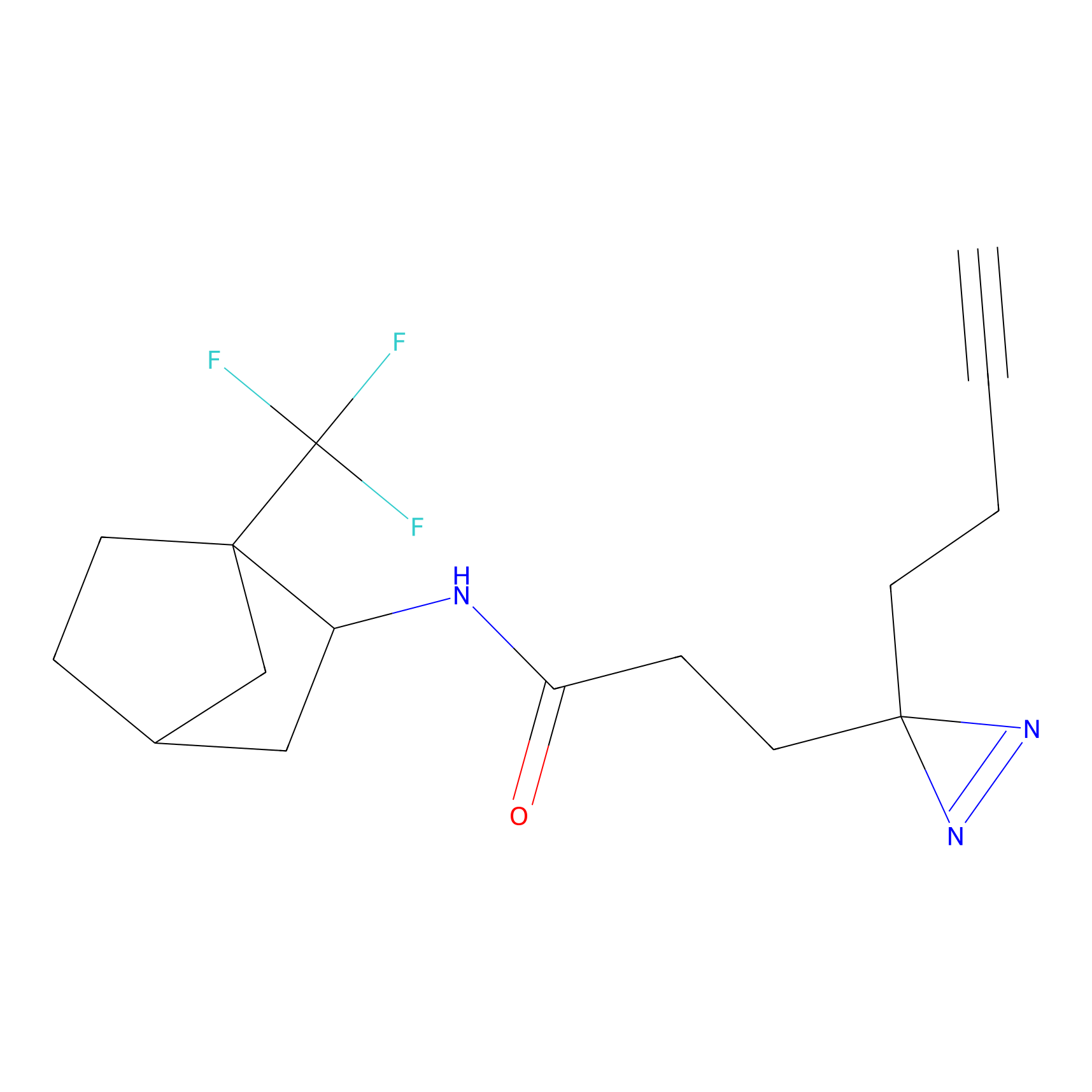
BTD Probe Info |
 |
C93(2.14) | LDD1699 | [2] | |
|
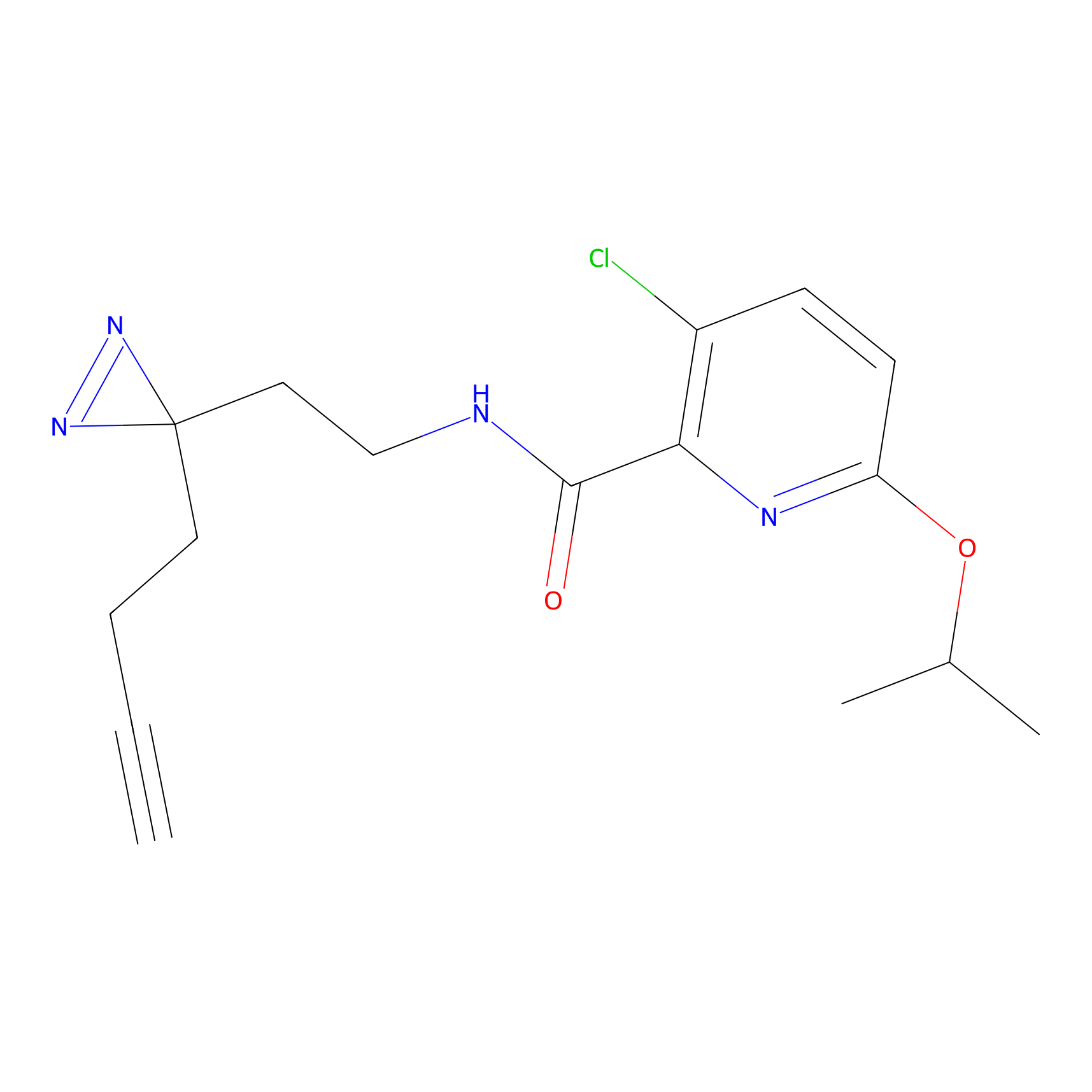
DBIA Probe Info |
 |
C93(1.37) | LDD3338 | [3] | |
|
HPAP Probe Info |
 |
3.16 | LDD0063 | [4] | |
|
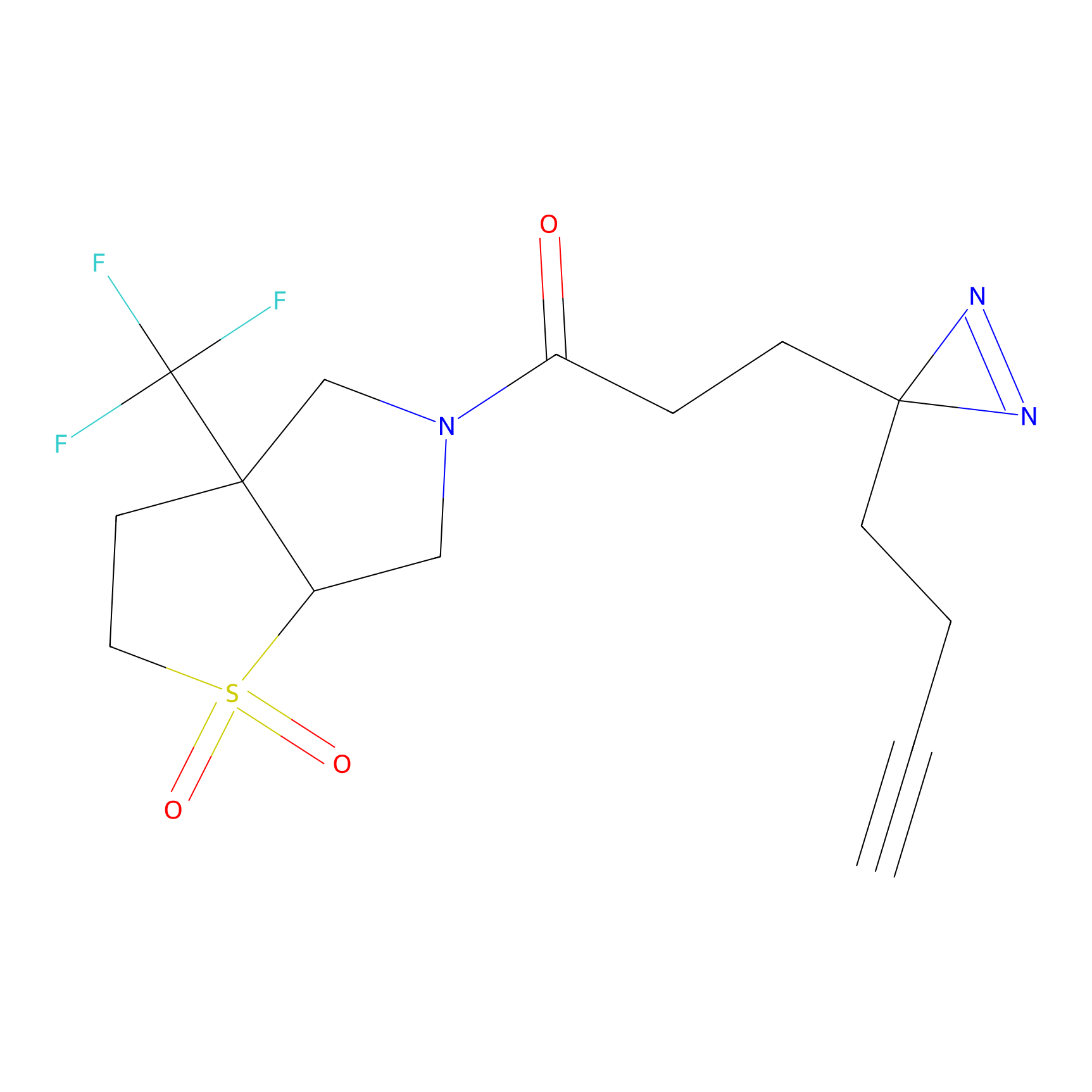
IPM Probe Info |
 |
C93(4.37) | LDD1701 | [2] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [5] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [5] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [6] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [7] | |
|
Acrolein Probe Info |
 |
H146(0.00); H126(0.00) | LDD0217 | [8] | |
|
Crotonaldehyde Probe Info |
 |
H146(0.00); C93(0.00) | LDD0219 | [8] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [9] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C93(0.78) | LDD2142 | [2] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C93(1.24) | LDD2117 | [2] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C93(0.49) | LDD2131 | [2] |
| LDCM0545 | Acetamide | MDA-MB-231 | C93(0.99) | LDD2138 | [2] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C93(1.08) | LDD2113 | [2] |
| LDCM0108 | Chloroacetamide | HeLa | H146(0.00); H126(0.00) | LDD0222 | [8] |
| LDCM0017 | DFG-out-2 | A431 | 4.90 | LDD0075 | [12] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C93(1.51) | LDD1702 | [2] |
| LDCM0107 | IAA | HeLa | H126(0.00); H146(0.00) | LDD0221 | [8] |
| LDCM0022 | KB02 | HCC1806 | C93(1.59) | LDD2341 | [3] |
| LDCM0023 | KB03 | MDA-MB-231 | C93(4.37) | LDD1701 | [2] |
| LDCM0024 | KB05 | MV4-11 | C93(1.37) | LDD3338 | [3] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [8] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C93(1.31) | LDD2089 | [2] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C93(1.53) | LDD2093 | [2] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C93(1.55) | LDD2097 | [2] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C93(1.69) | LDD2099 | [2] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C93(0.72) | LDD2100 | [2] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C93(1.19) | LDD2101 | [2] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C93(1.09) | LDD2105 | [2] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C93(1.05) | LDD2107 | [2] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C93(0.96) | LDD2108 | [2] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C93(1.31) | LDD2109 | [2] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C93(1.14) | LDD2111 | [2] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C93(0.57) | LDD2115 | [2] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C93(0.50) | LDD2120 | [2] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C93(0.90) | LDD2125 | [2] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C93(1.12) | LDD2127 | [2] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C93(0.45) | LDD2128 | [2] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C93(1.18) | LDD2129 | [2] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C93(0.71) | LDD2133 | [2] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C93(0.72) | LDD2134 | [2] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C93(1.05) | LDD2135 | [2] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C93(1.68) | LDD2136 | [2] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C93(3.60) | LDD1700 | [2] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C93(1.29) | LDD2140 | [2] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C93(0.34) | LDD2143 | [2] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C93(1.57) | LDD2144 | [2] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C93(0.49) | LDD2148 | [2] |
| LDCM0014 | Panhematin | K562 | 3.16 | LDD0063 | [4] |
| LDCM0019 | Staurosporine | Hep-G2 | 2.00 | LDD0083 | [13] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Transmembrane emp24 domain-containing protein 10 (TMED10) | EMP24/GP25L family | P49755 | |||
GPCR
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| CD59 glycoprotein (CD59) | . | P13987 | |||
References