Details of the Target
General Information of Target
| Target ID | LDTP00907 | |||||
|---|---|---|---|---|---|---|
| Target Name | Retinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta (PDE6D) | |||||
| Gene Name | PDE6D | |||||
| Gene ID | 5147 | |||||
| Synonyms |
PDED; Retinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta; GMP-PDE delta; Protein p17 |
|||||
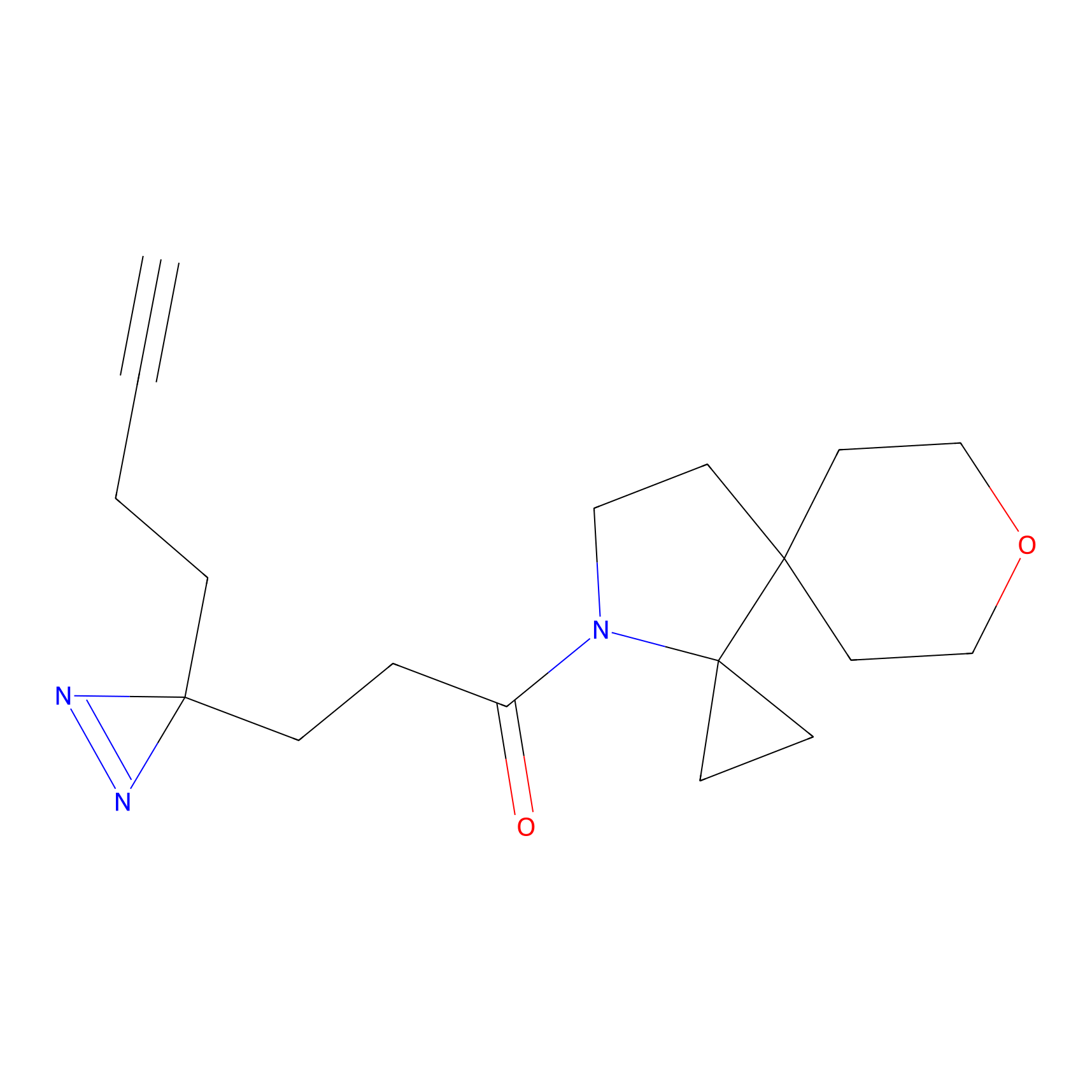
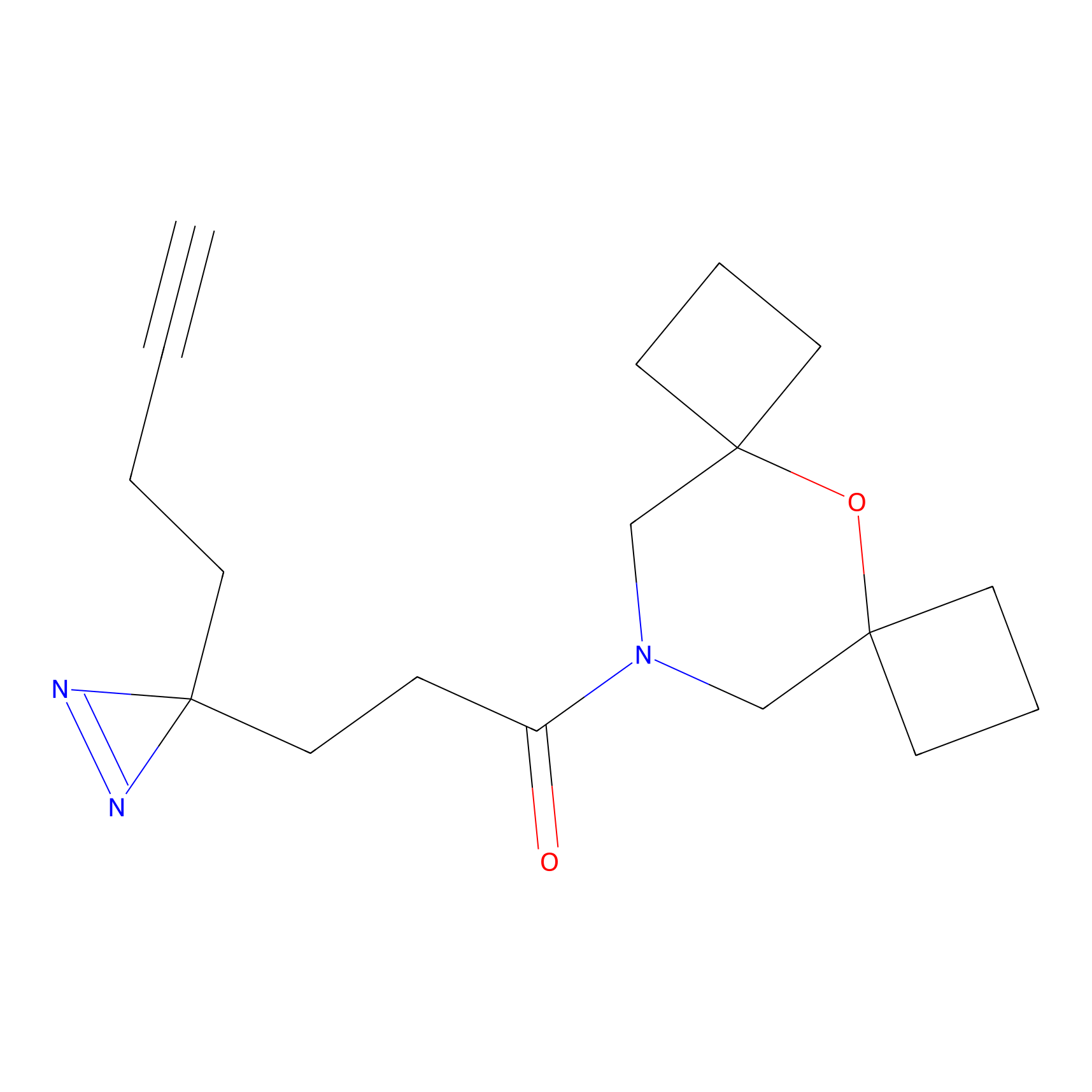
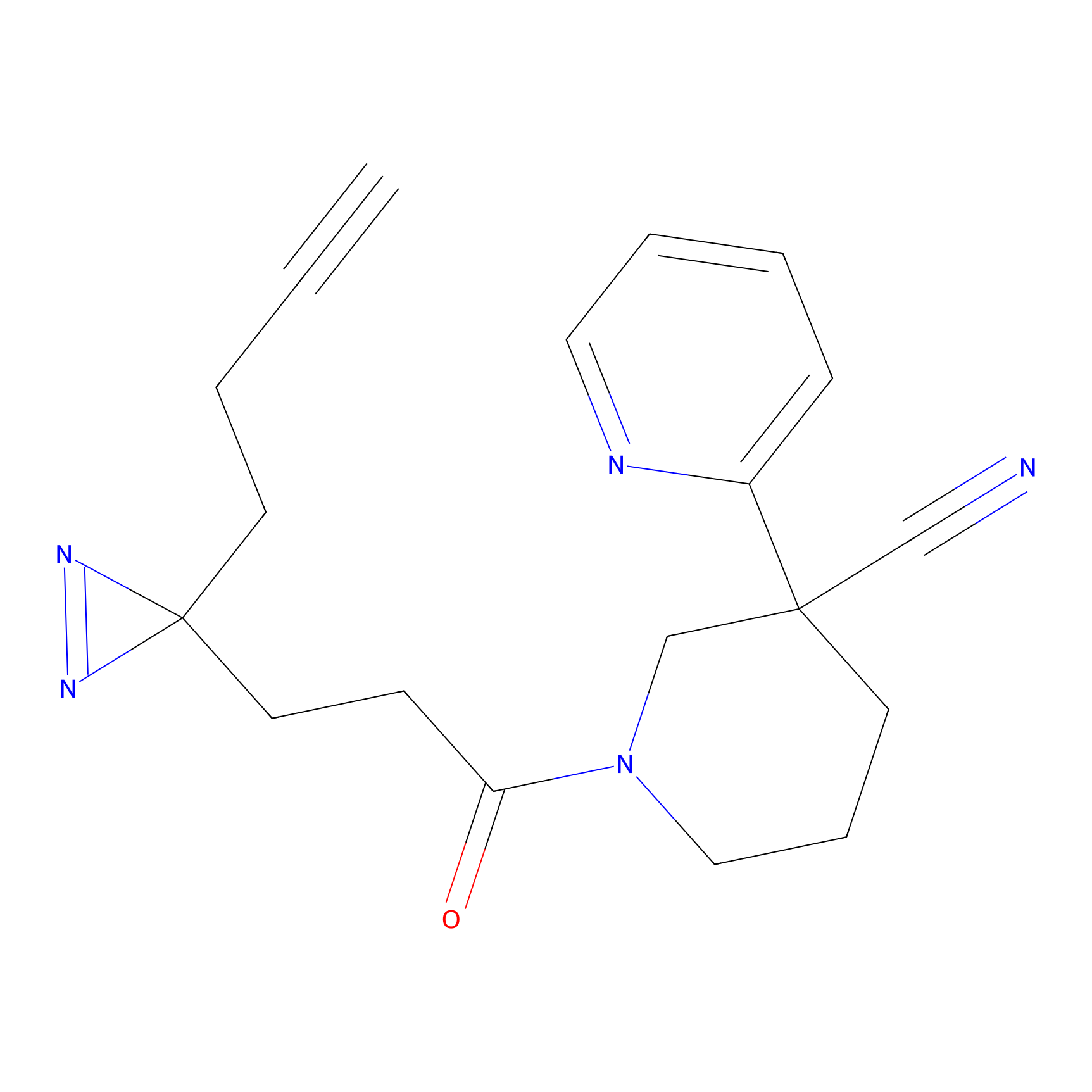
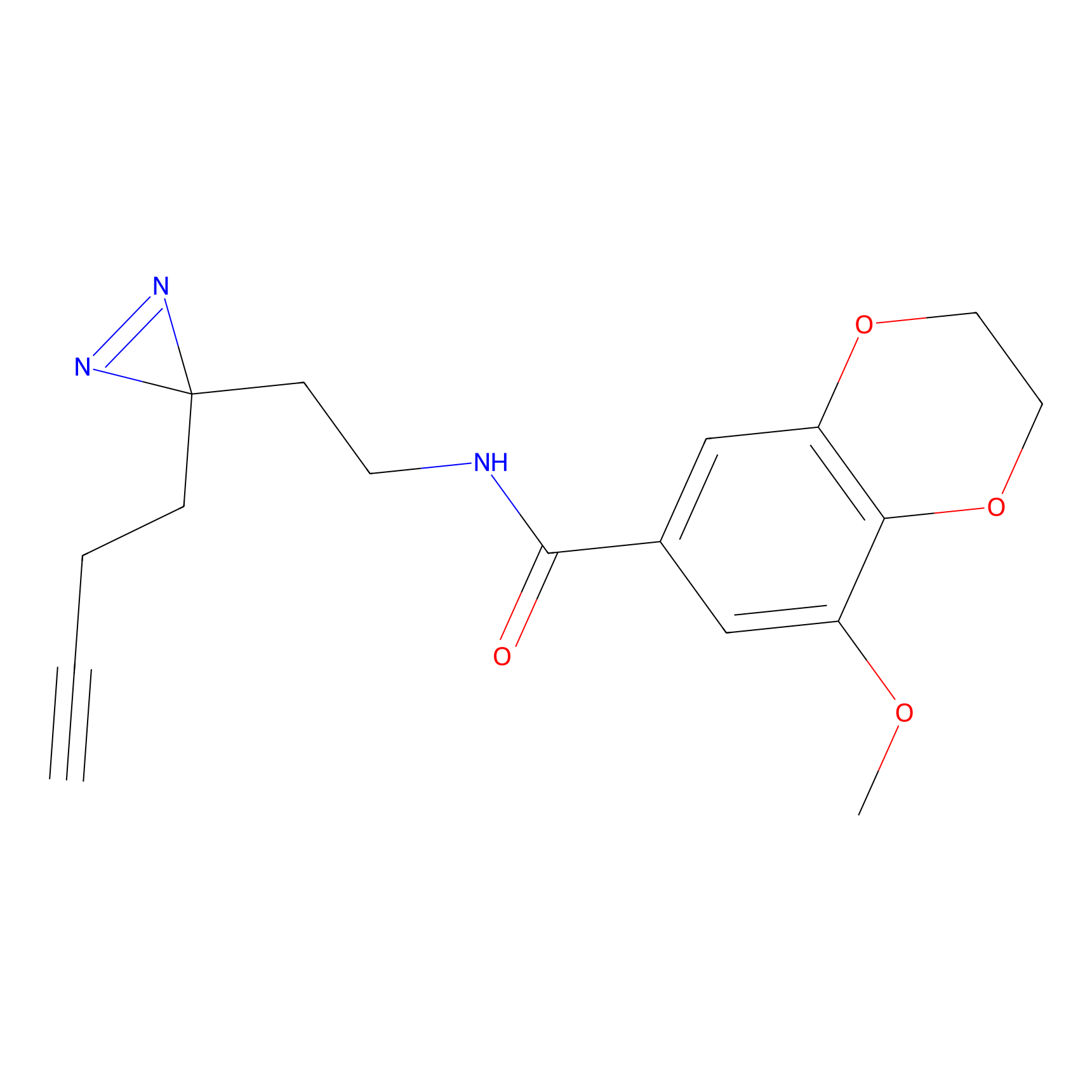
| 3D Structure | ||||||
| Sequence |
MSAKDERAREILRGFKLNWMNLRDAETGKILWQGTEDLSVPGVEHEARVPKKILKCKAVS
RELNFSSTEQMEKFRLEQKVYFKGQCLEEWFFEFGFVIPNSTNTWQSLIEAAPESQMMPA SVLTGNVIIETKFFDDDLLVSTSRVRLFYV |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
PDE6D/unc-119 family
|
|||||
| Subcellular location |
Cytoplasm, cytosol
|
|||||
| Function |
Promotes the release of prenylated target proteins from cellular membranes. Modulates the activity of prenylated or palmitoylated Ras family members by regulating their subcellular location. Required for normal ciliary targeting of farnesylated target proteins, such as INPP5E. Modulates the subcellular location of target proteins by acting as a GTP specific dissociation inhibitor (GDI). Increases the affinity of ARL3 for GTP by several orders of magnitude. Stabilizes ARL3-GTP by decreasing the nucleotide dissociation rate.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
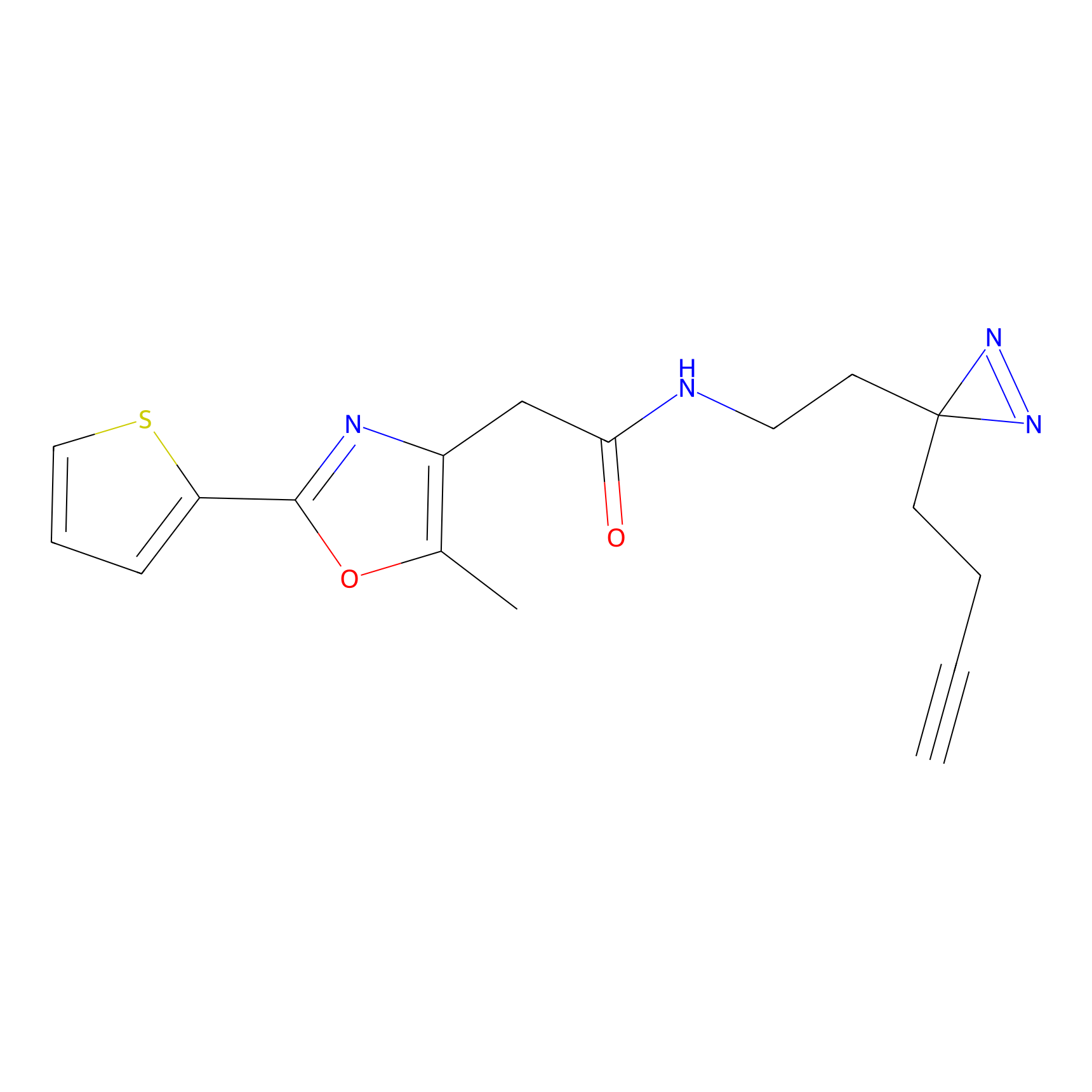
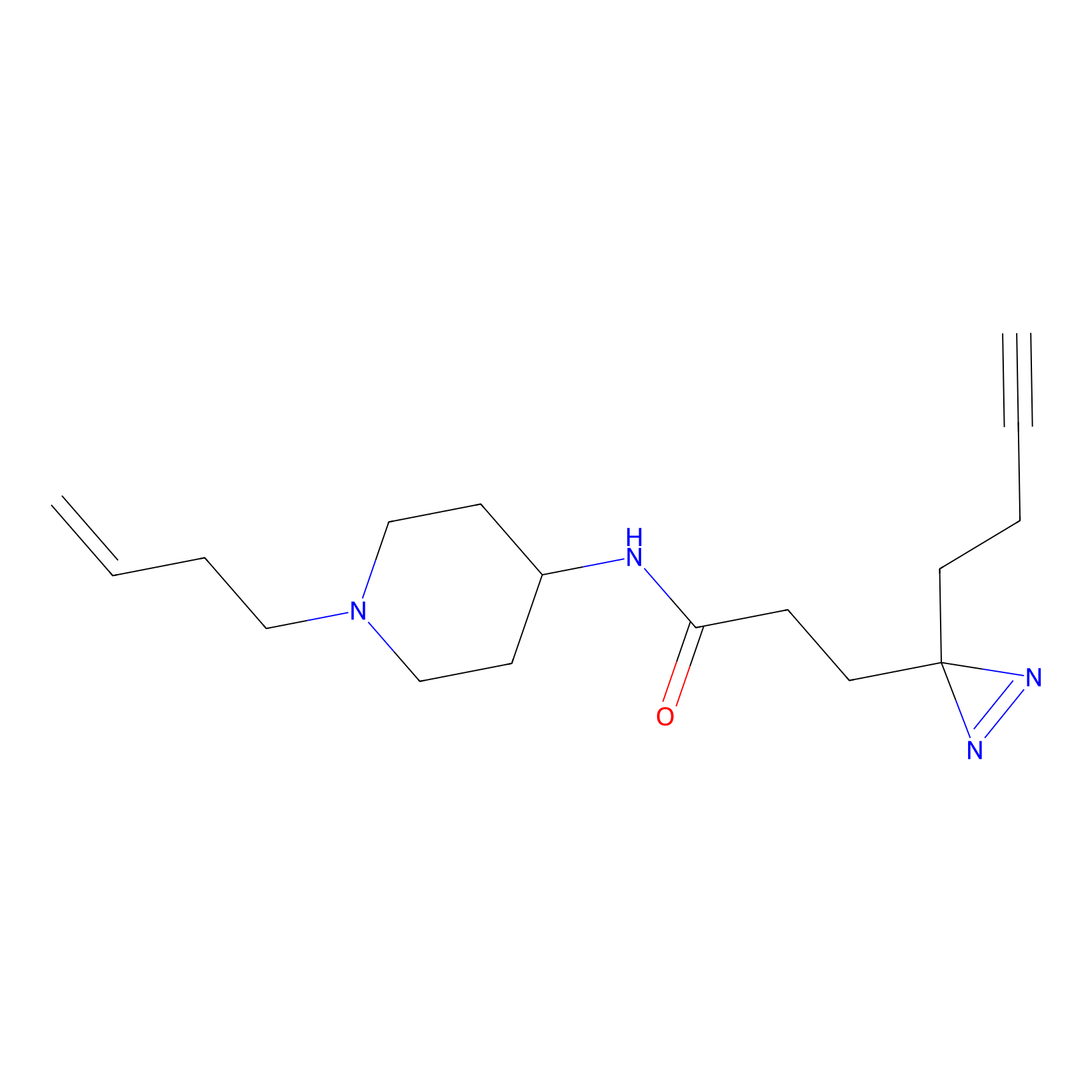
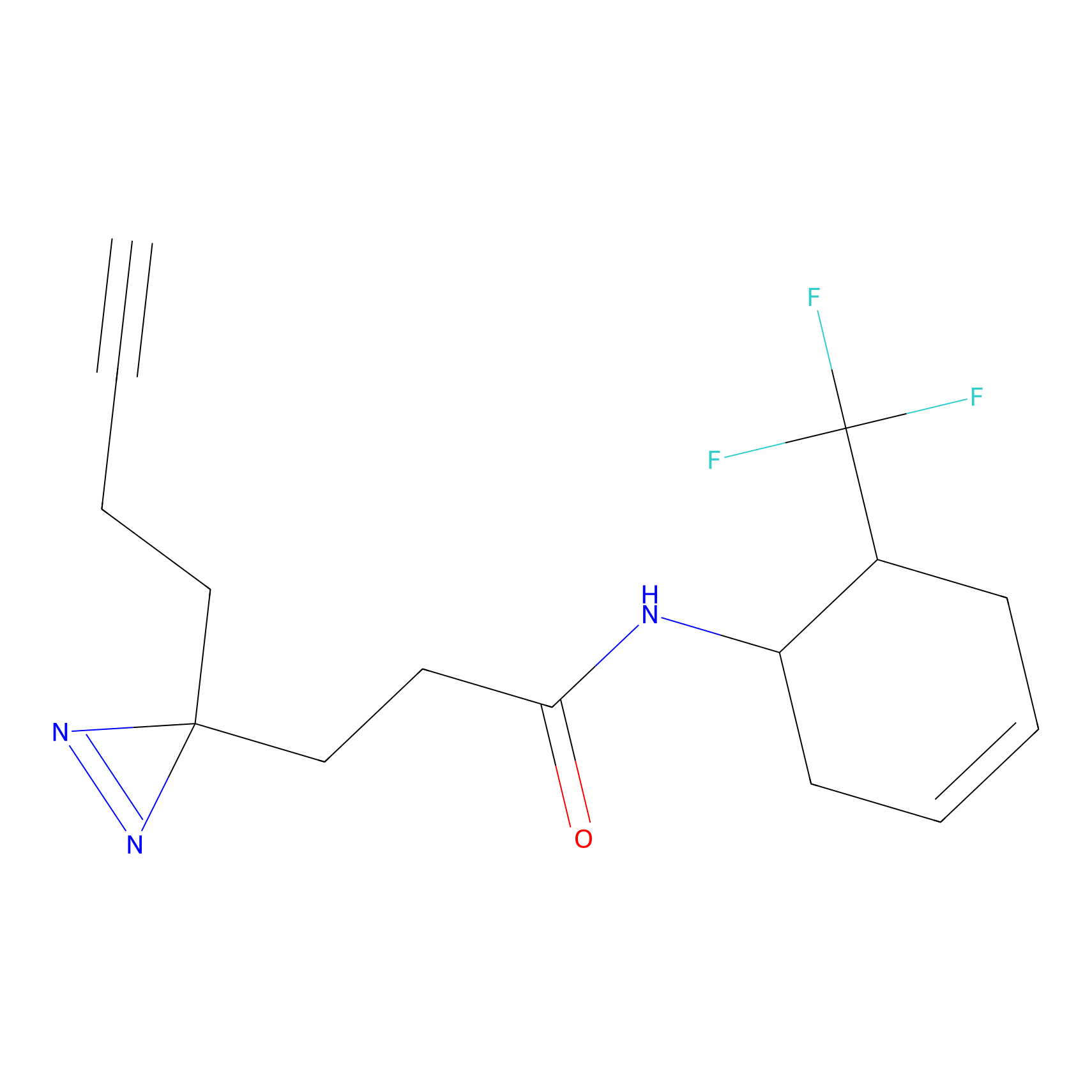
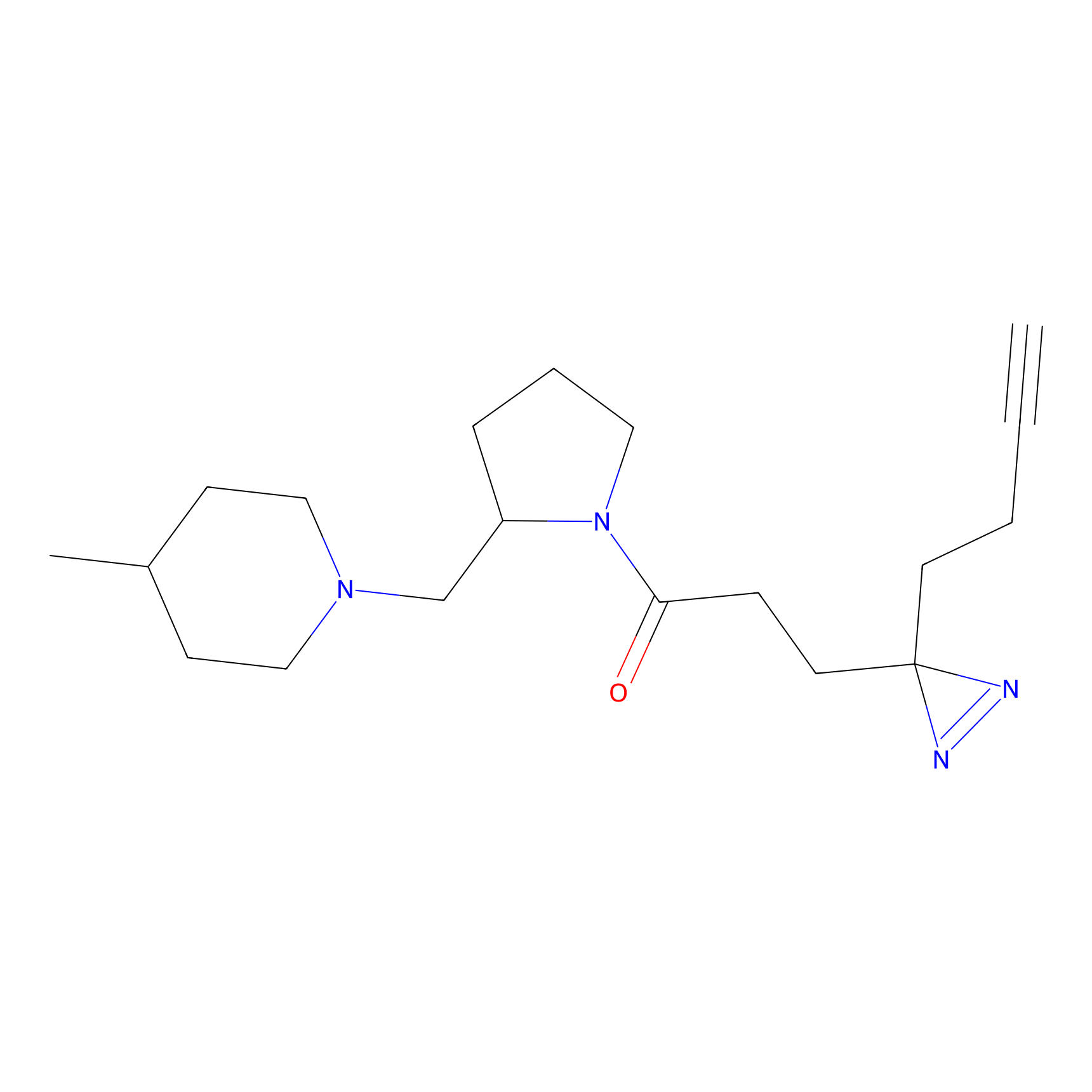
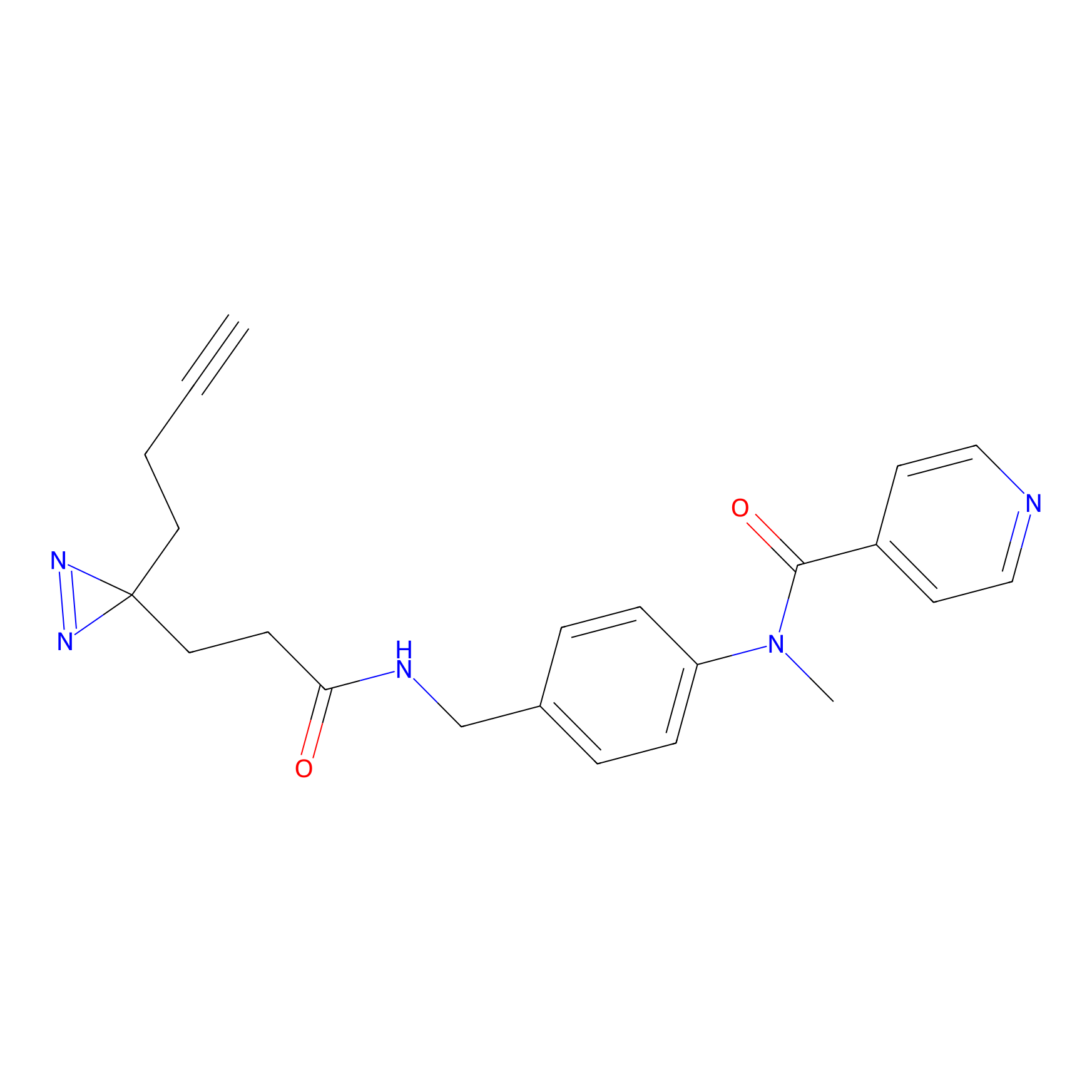
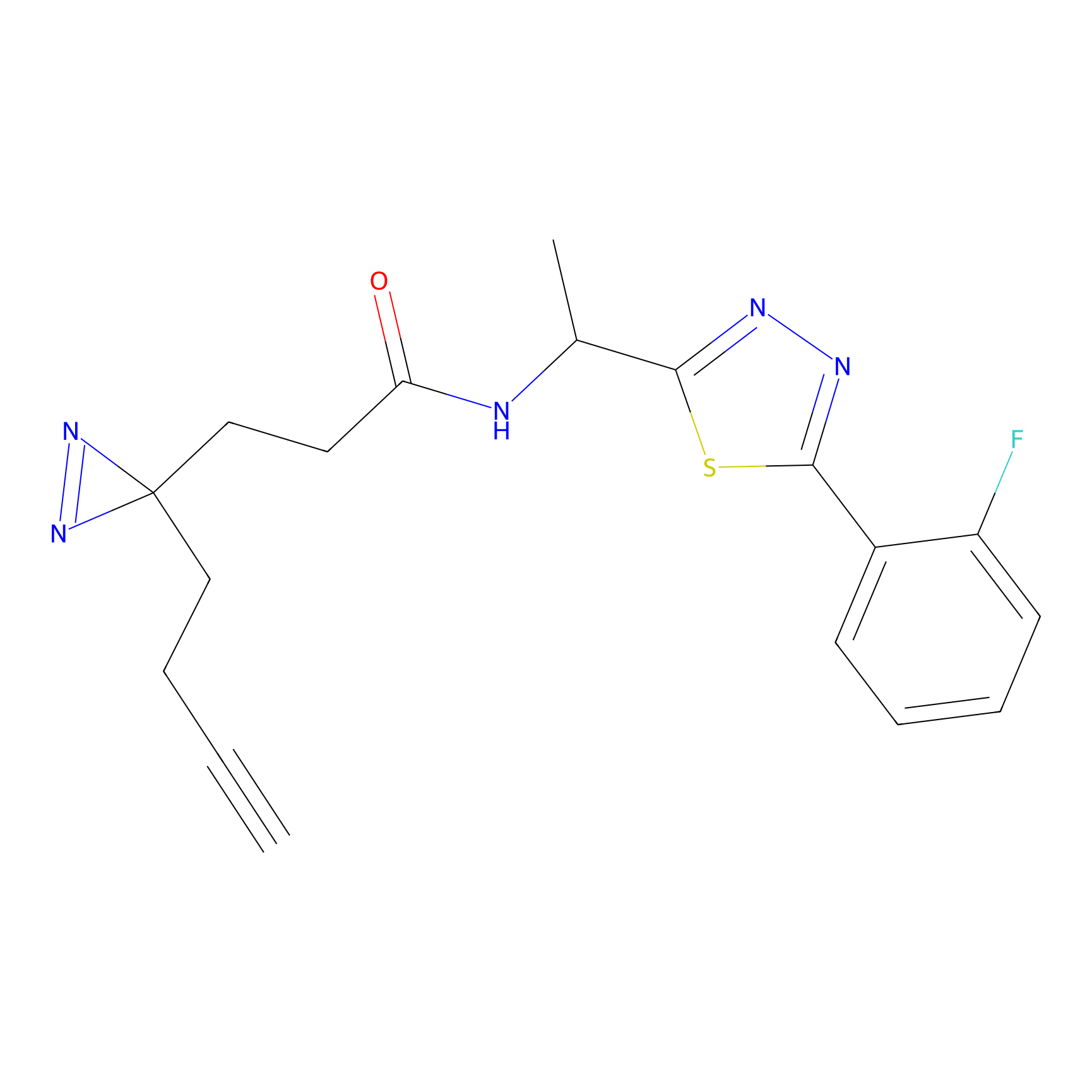
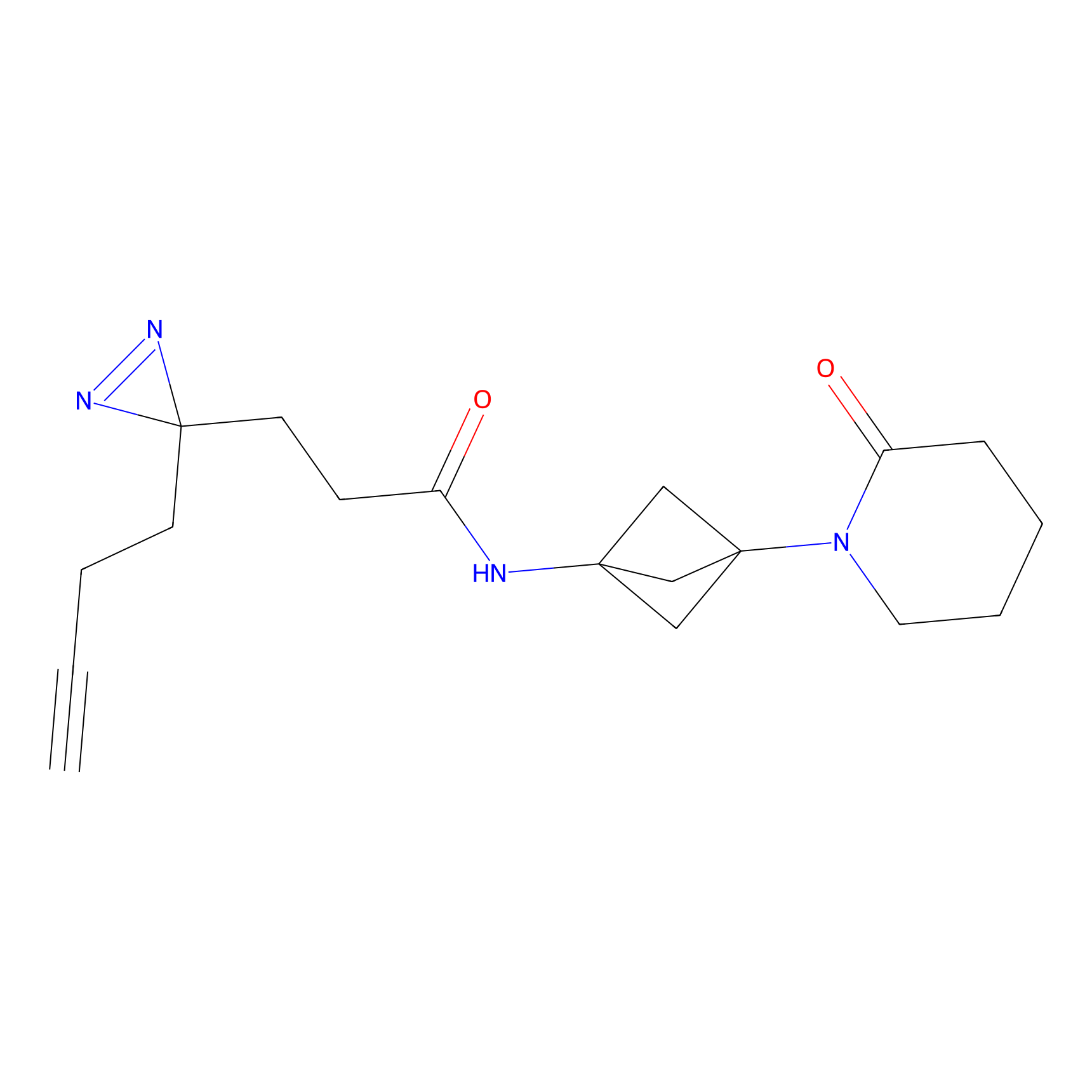
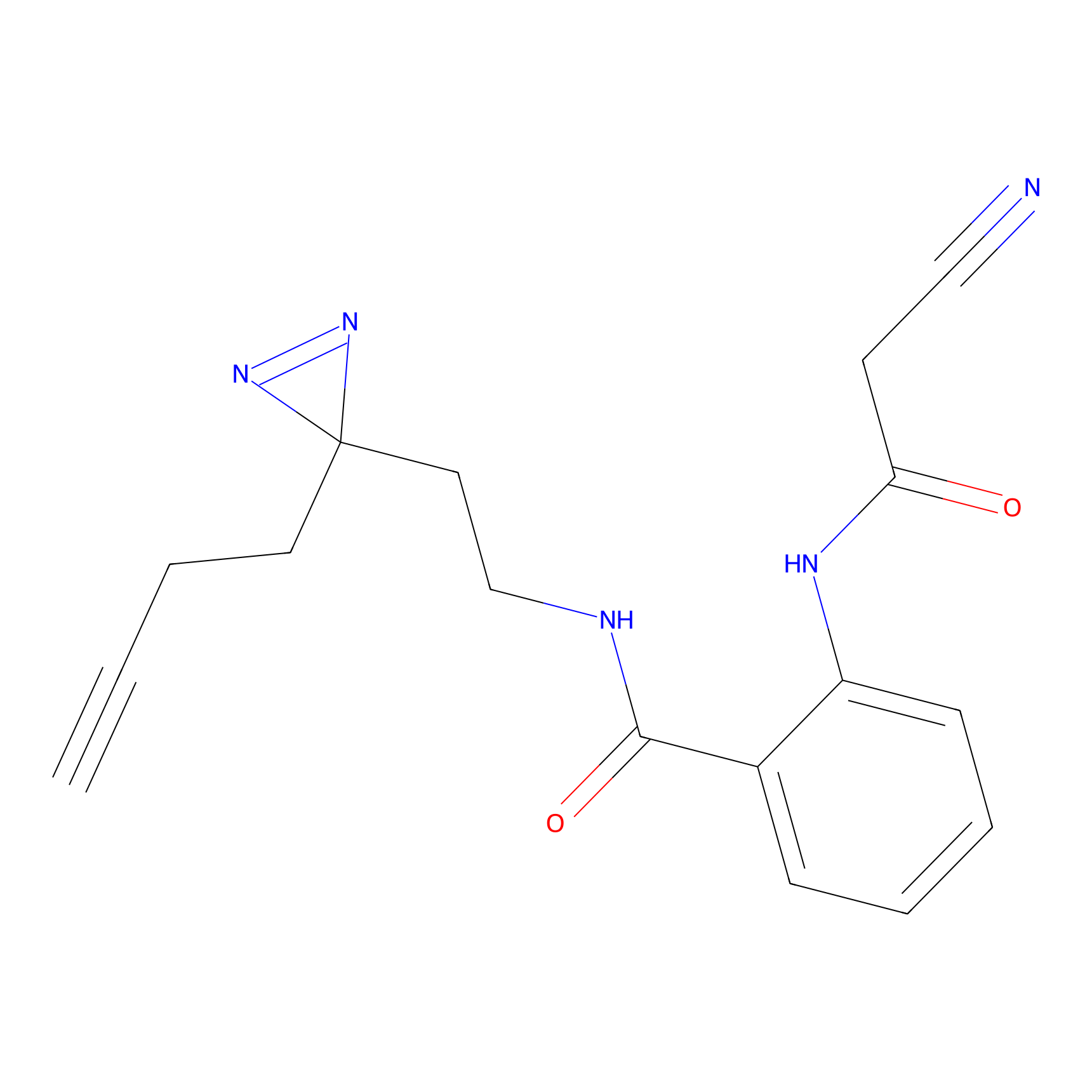
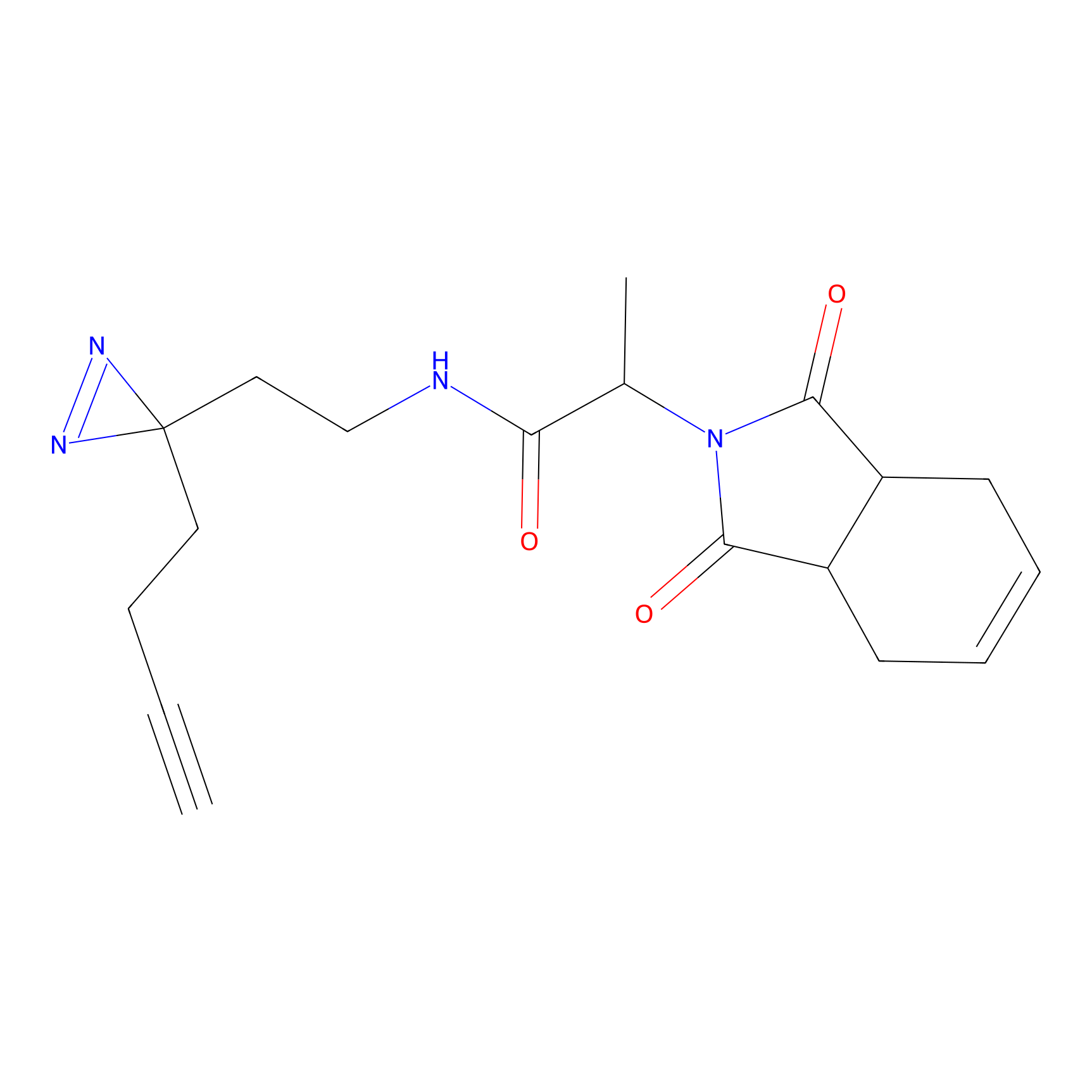
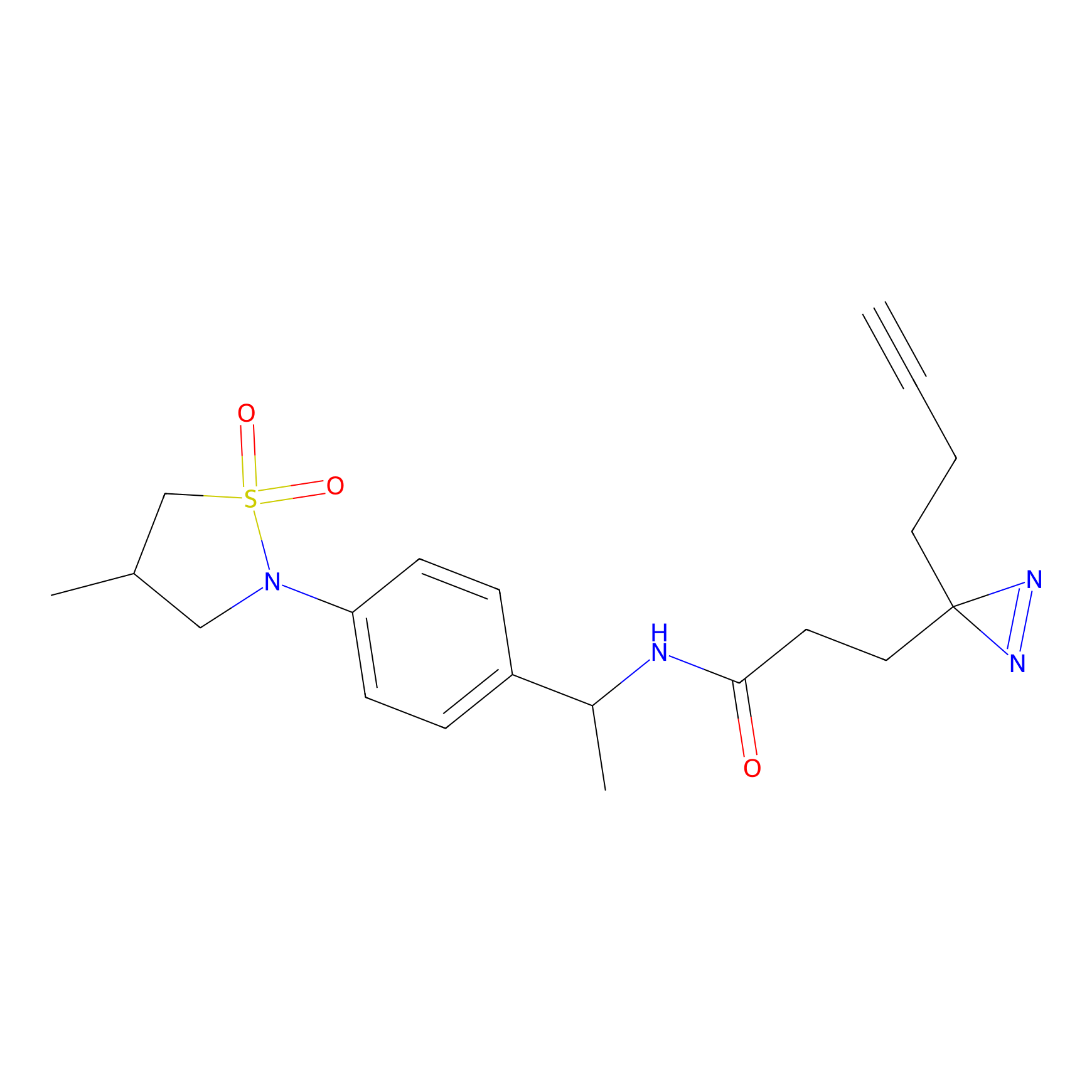
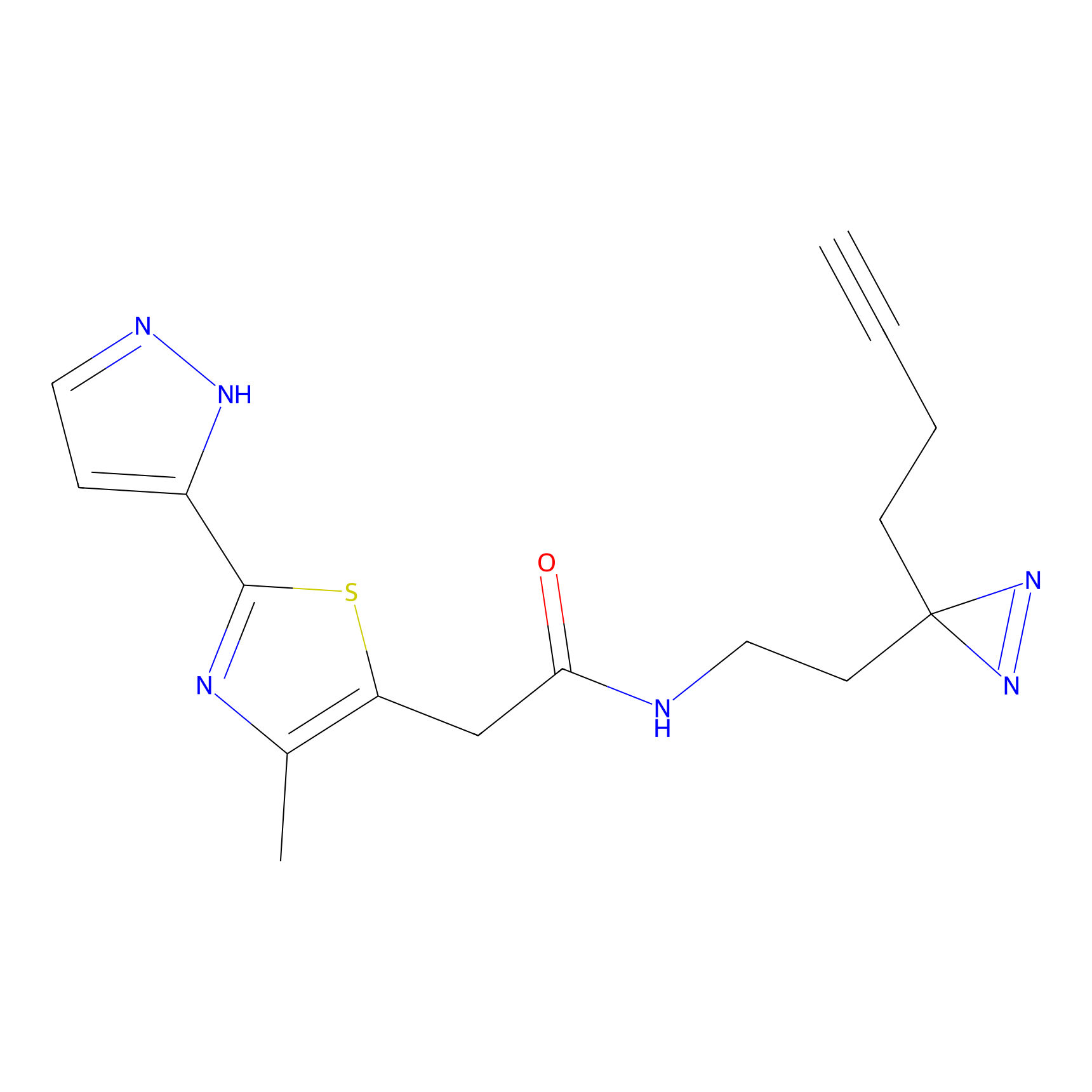
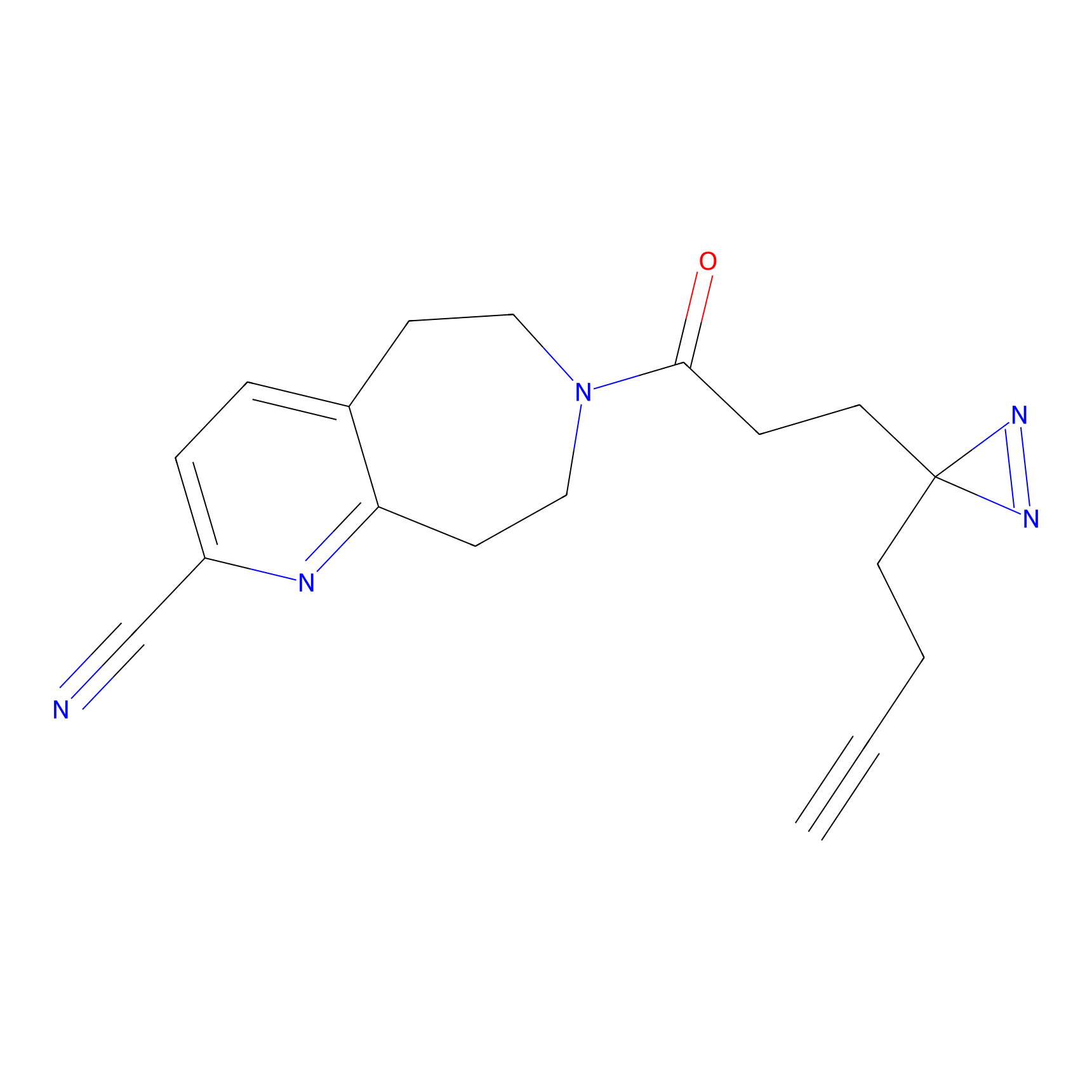
Probe(s) Labeling This Target
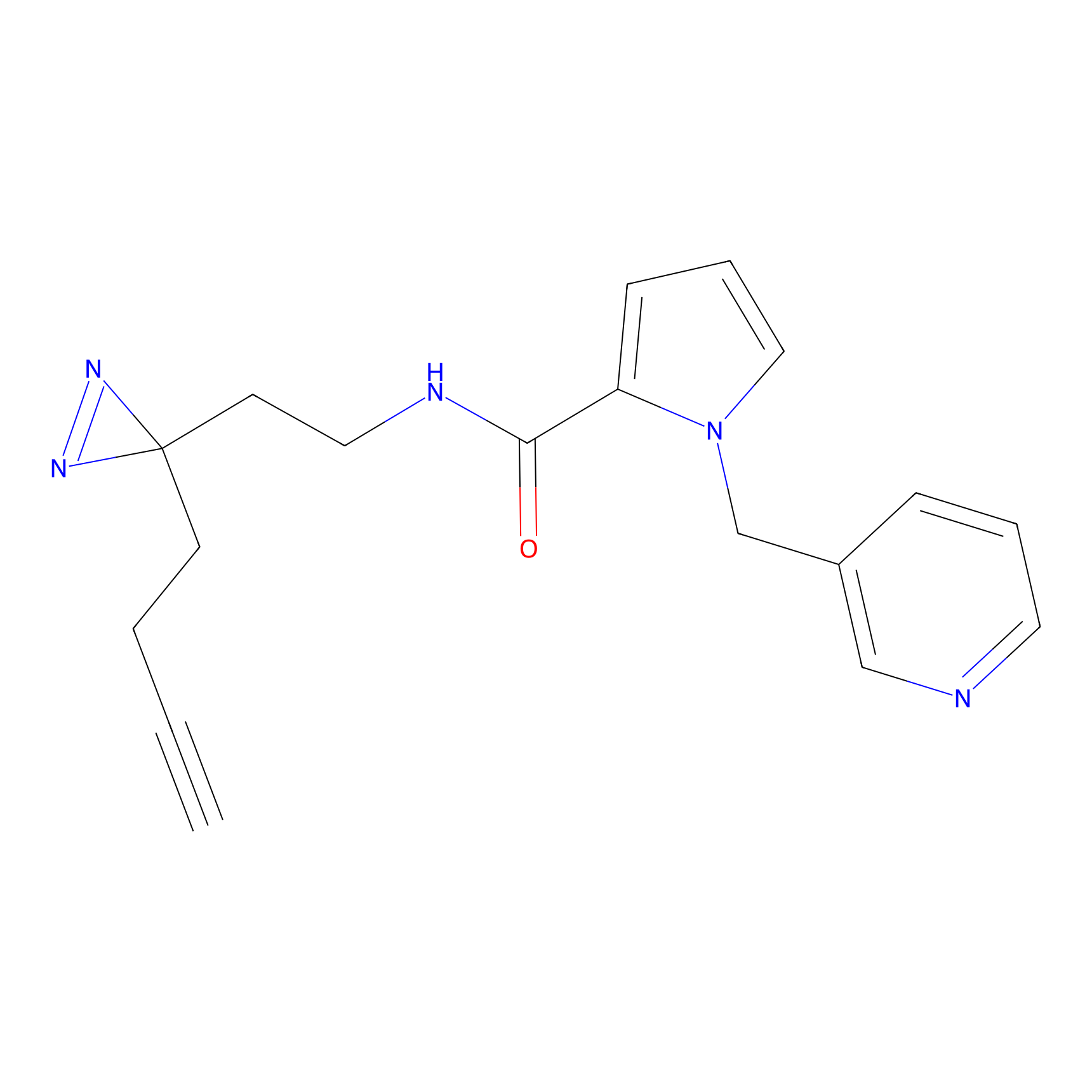
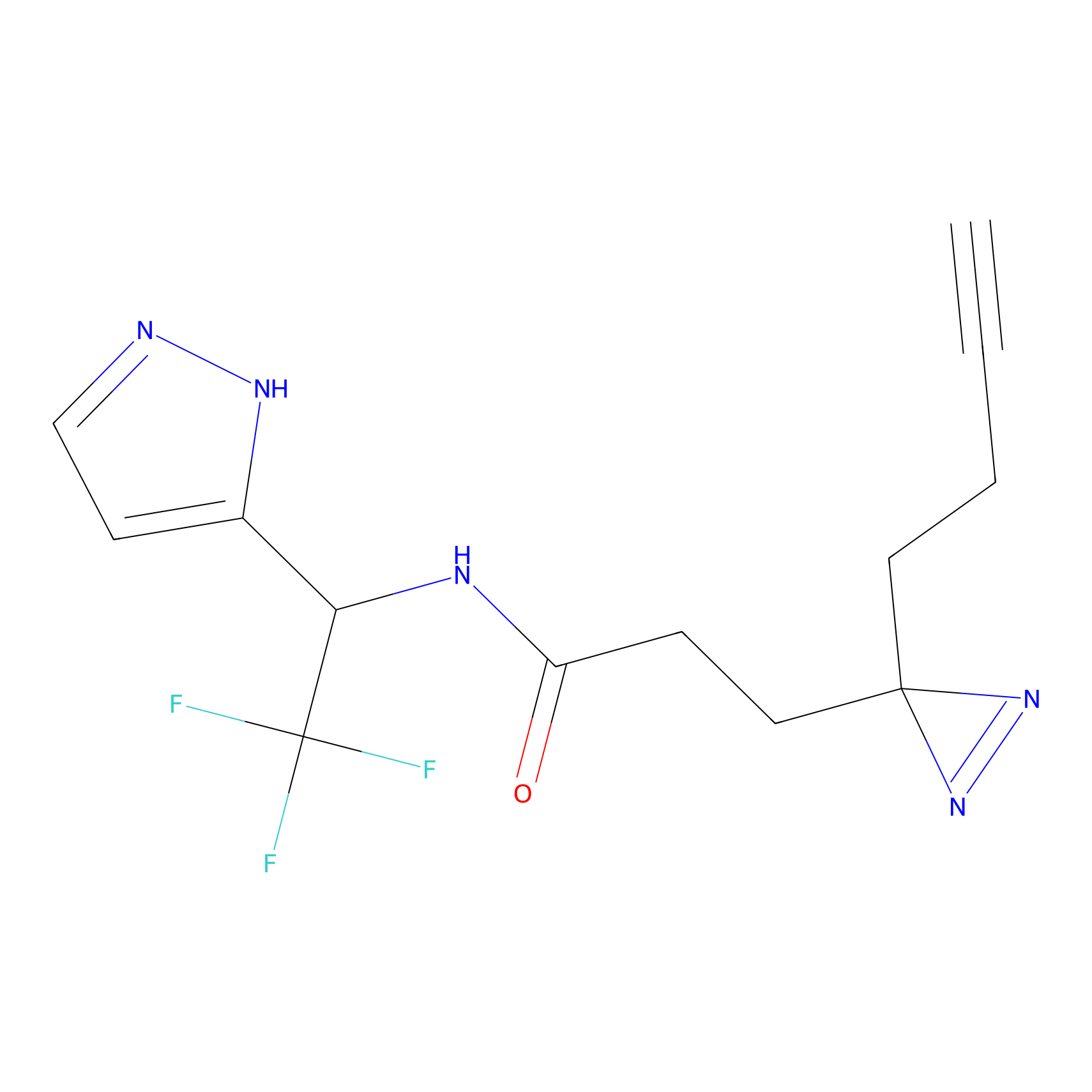
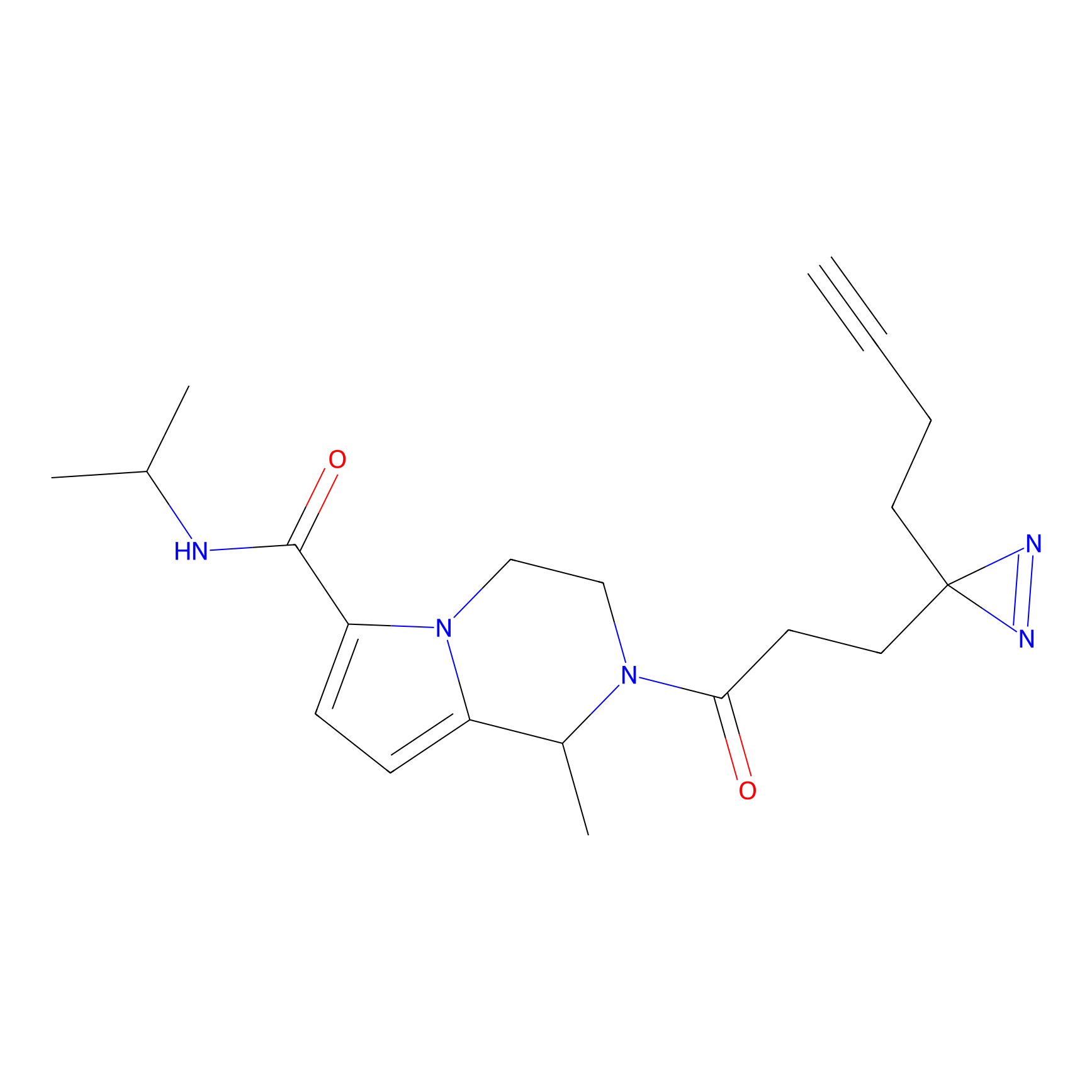
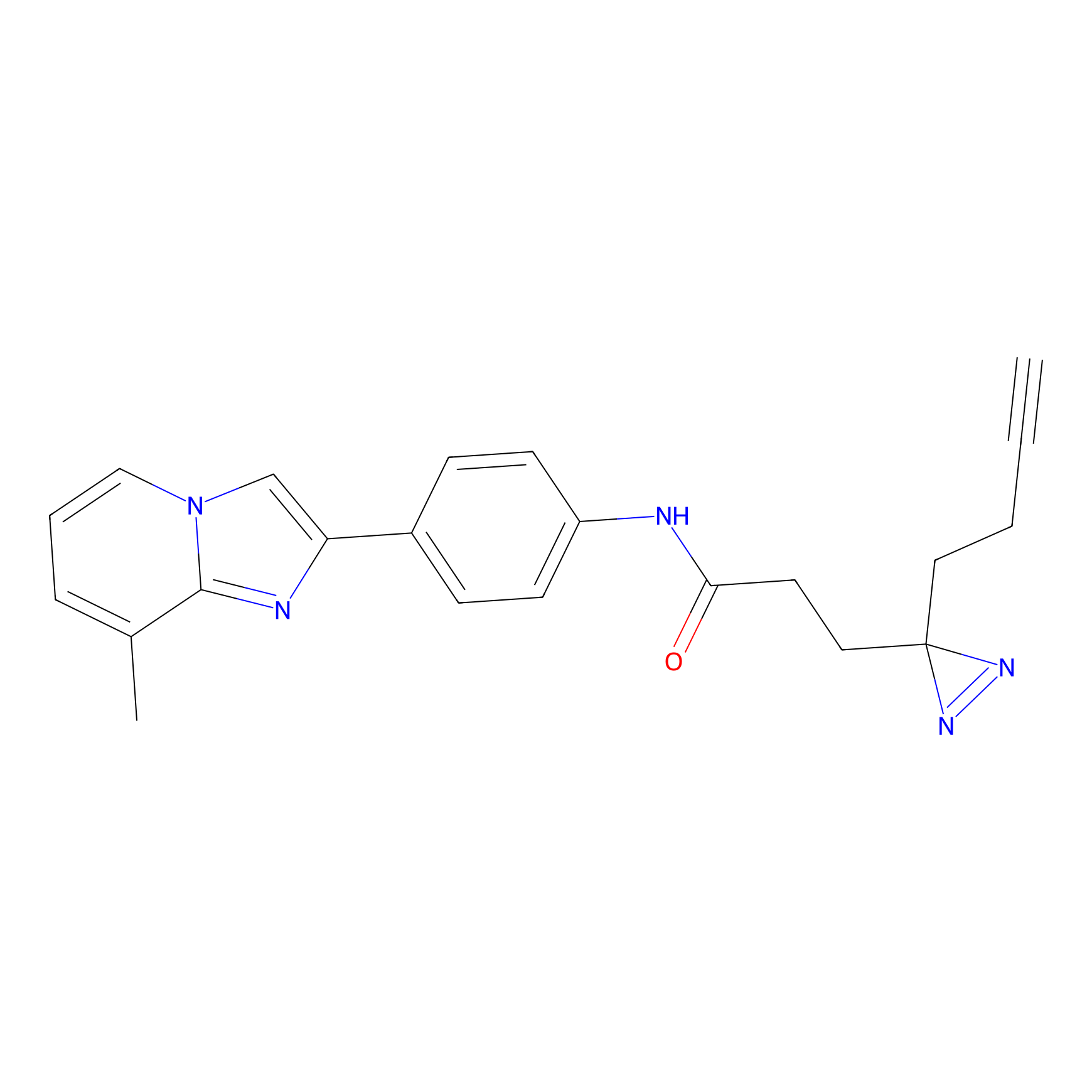
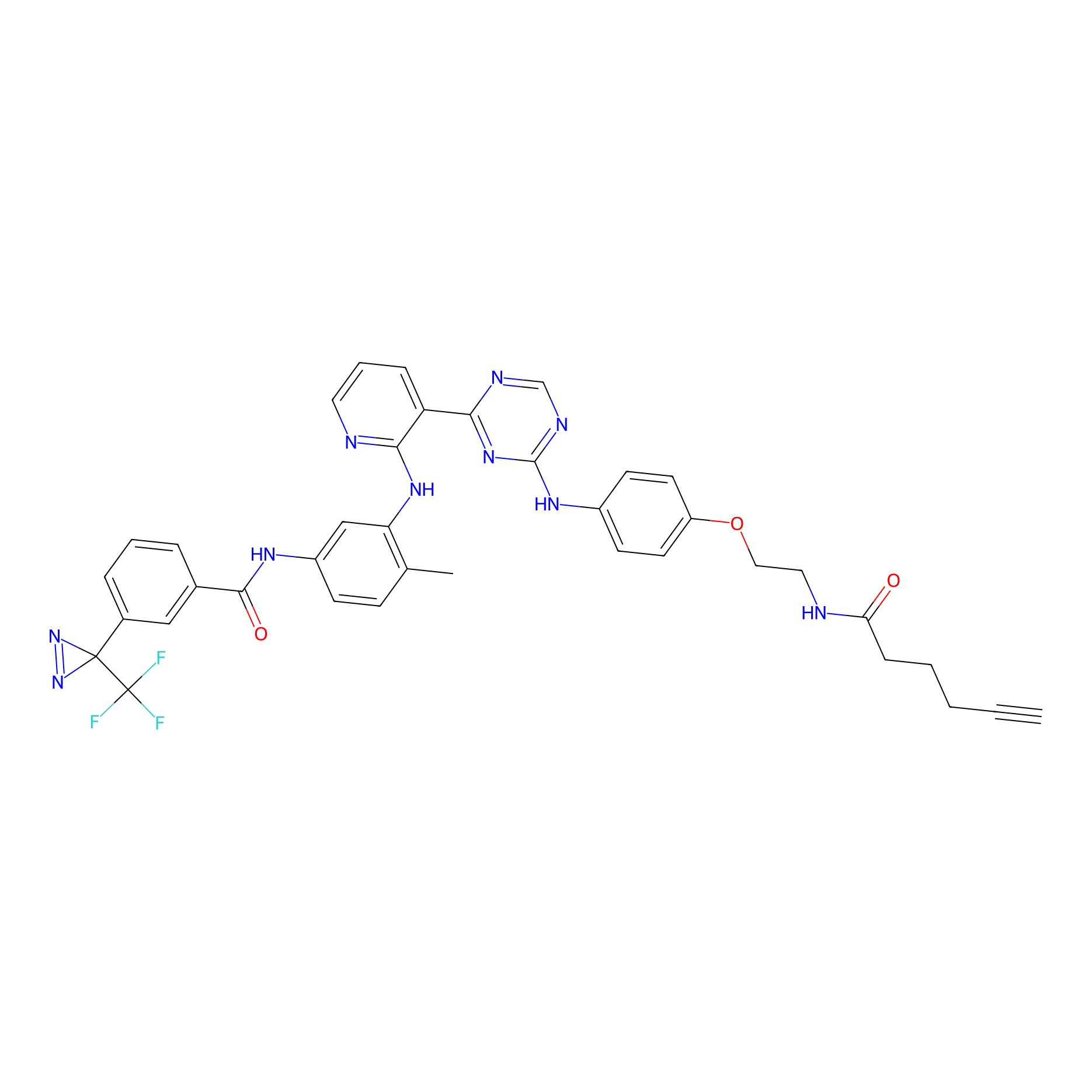
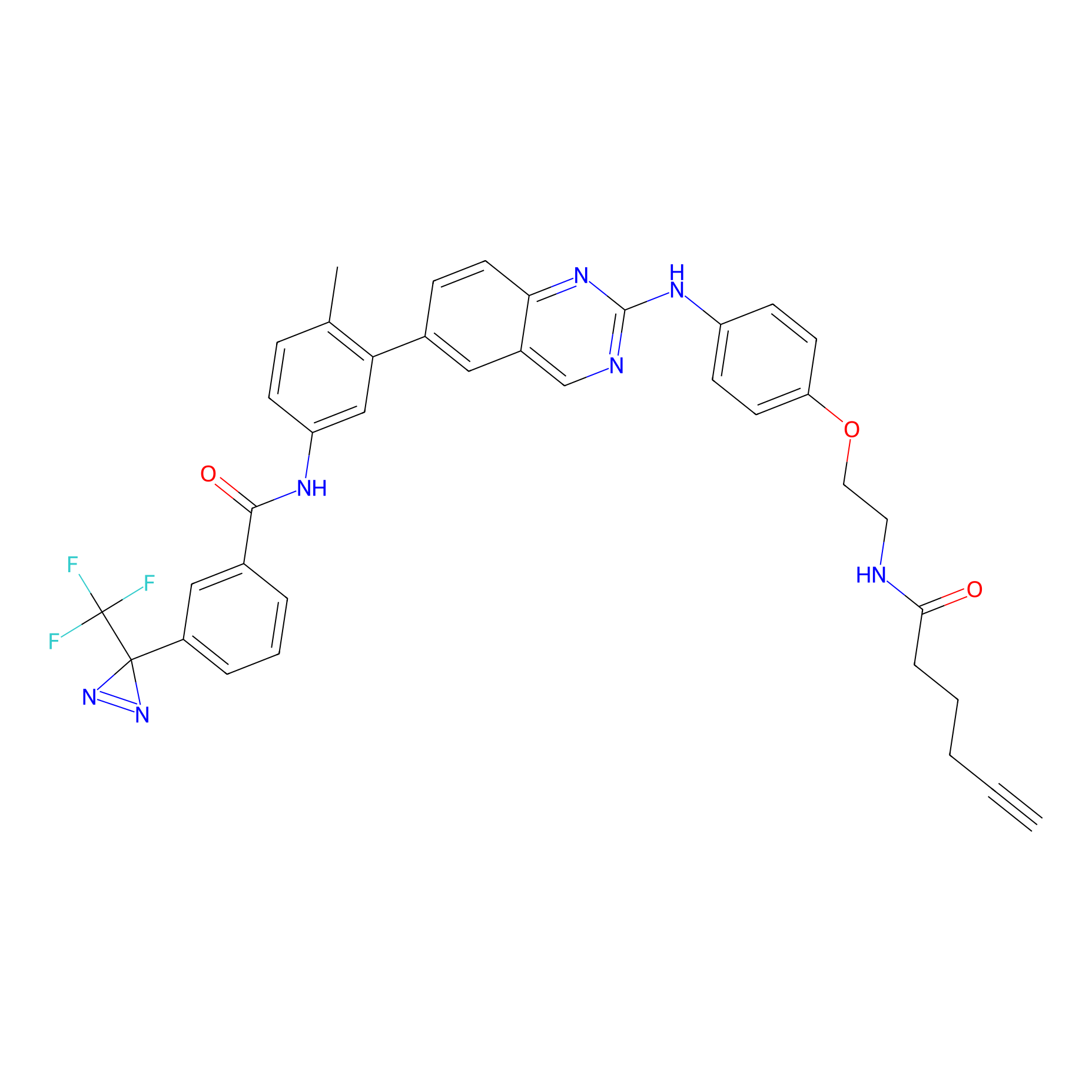
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Homeobox protein orthopedia (OTP) | Paired homeobox family | Q5XKR4 | |||
References