Details of the Target
General Information of Target
| Target ID | LDTP01494 | |||||
|---|---|---|---|---|---|---|
| Target Name | Protein YIF1A (YIF1A) | |||||
| Gene Name | YIF1A | |||||
| Gene ID | 10897 | |||||
| Synonyms |
54TM; HYIF1P; YIF1; Protein YIF1A; 54TMp; YIP1-interacting factor homolog A |
|||||
| 3D Structure | ||||||
| Sequence |
MAYHSGYGAHGSKHRARAAPDPPPLFDDTSGGYSSQPGGYPATGADVAFSVNHLLGDPMA
NVAMAYGSSIASHGKDMVHKELHRFVSVSKLKYFFAVDTAYVAKKLGLLVFPYTHQNWEV QYSRDAPLPPRQDLNAPDLYIPTMAFITYVLLAGMALGIQKRFSPEVLGLCASTALVWVV MEVLALLLGLYLATVRSDLSTFHLLAYSGYKYVGMILSVLTGLLFGSDGYYVALAWTSSA LMYFIVRSLRTAALGPDSMGGPVPRQRLQLYLTLGAAAFQPLIIYWLTFHLVR |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
YIF1 family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function | Possible role in transport between endoplasmic reticulum and Golgi. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
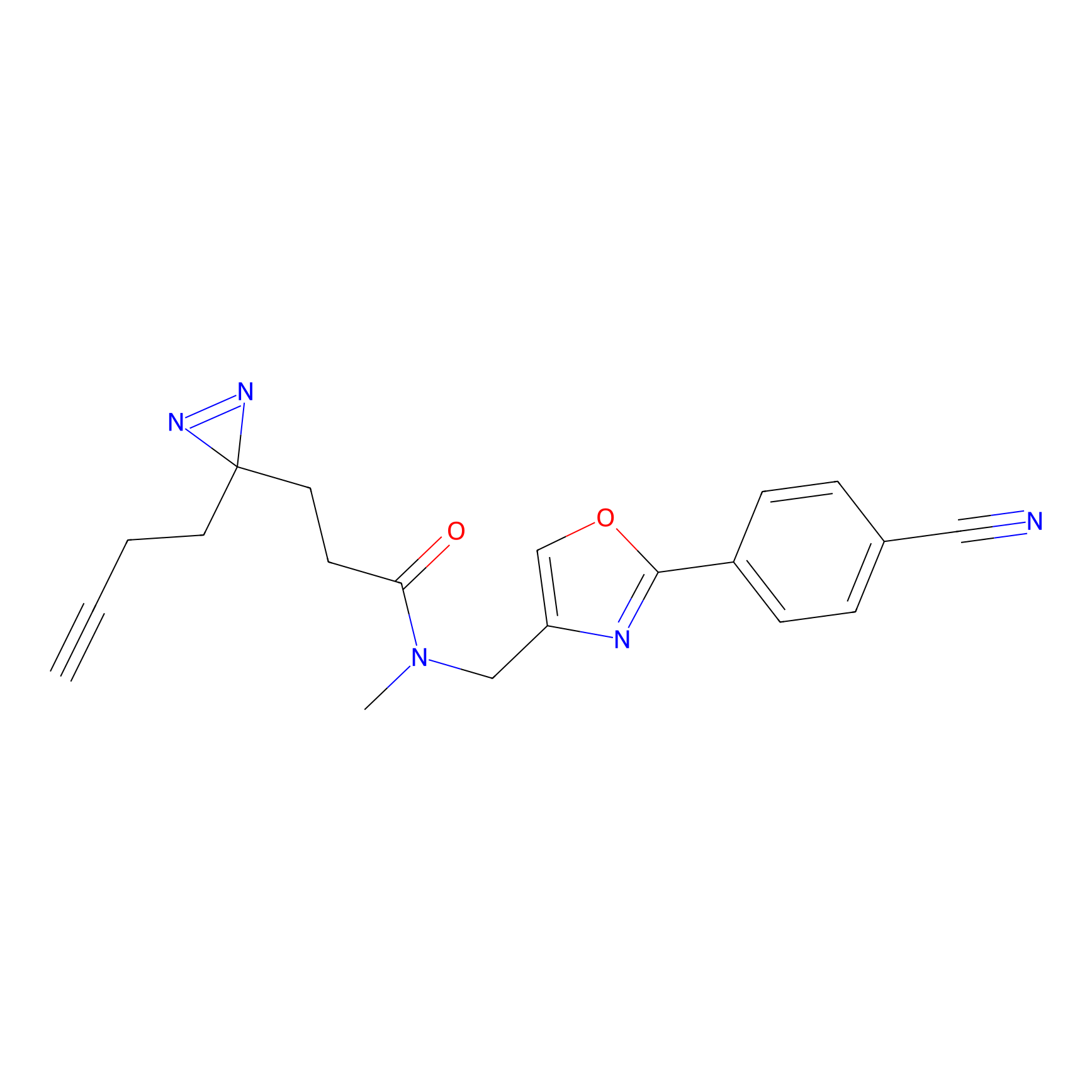
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [1] | |
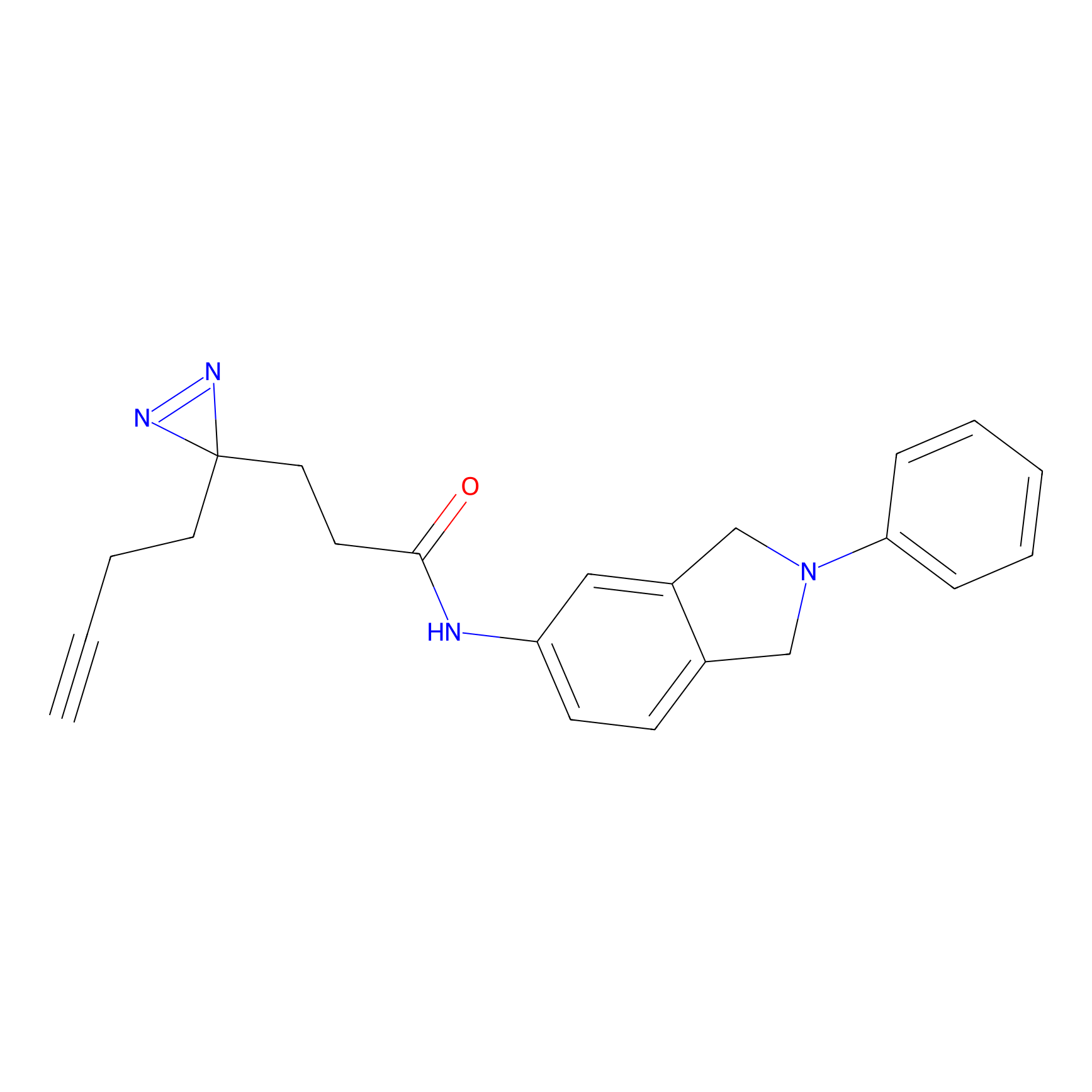
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [2] | |
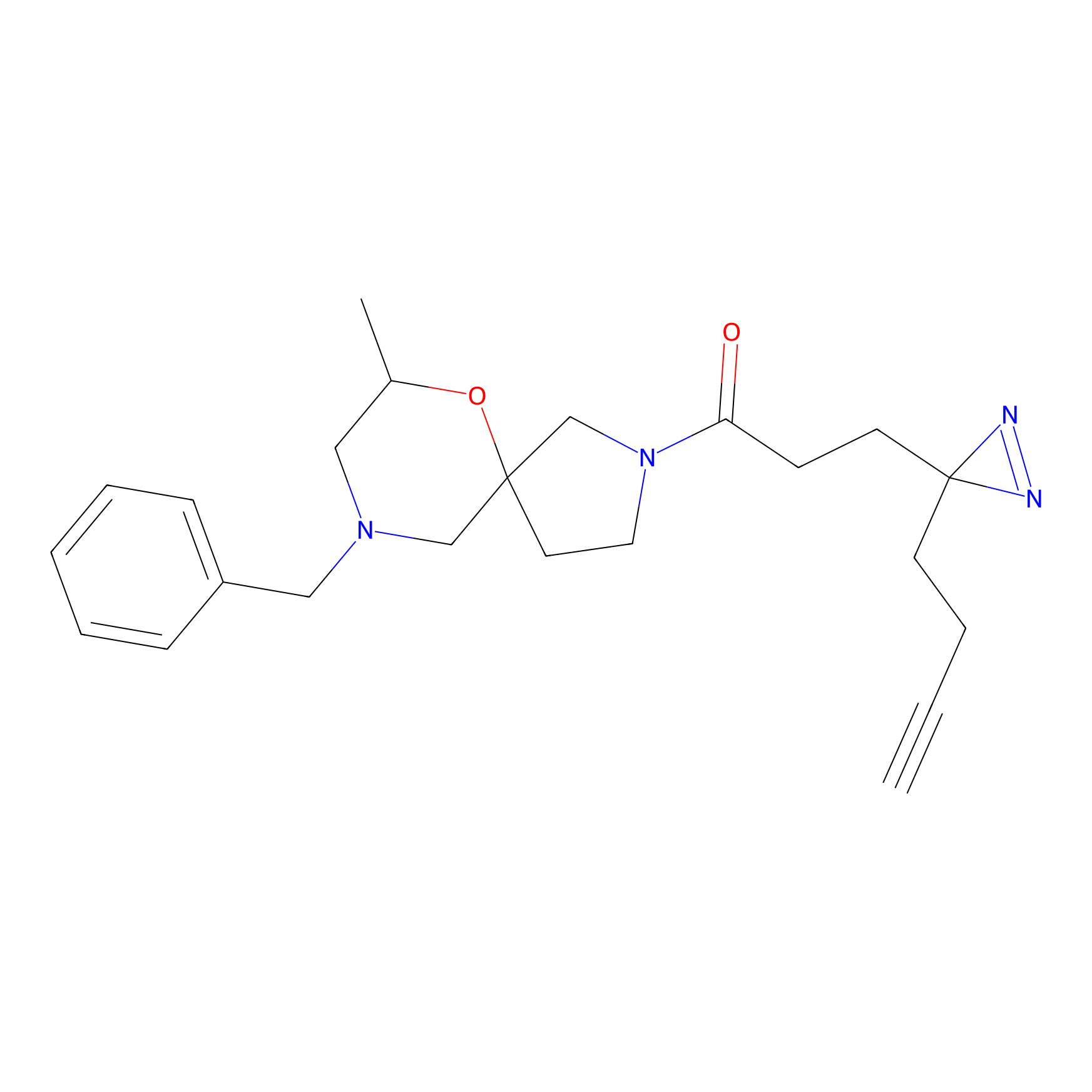
|
SF Probe Info |
 |
Y7(0.00); Y3(0.00) | LDD0028 | [3] | |
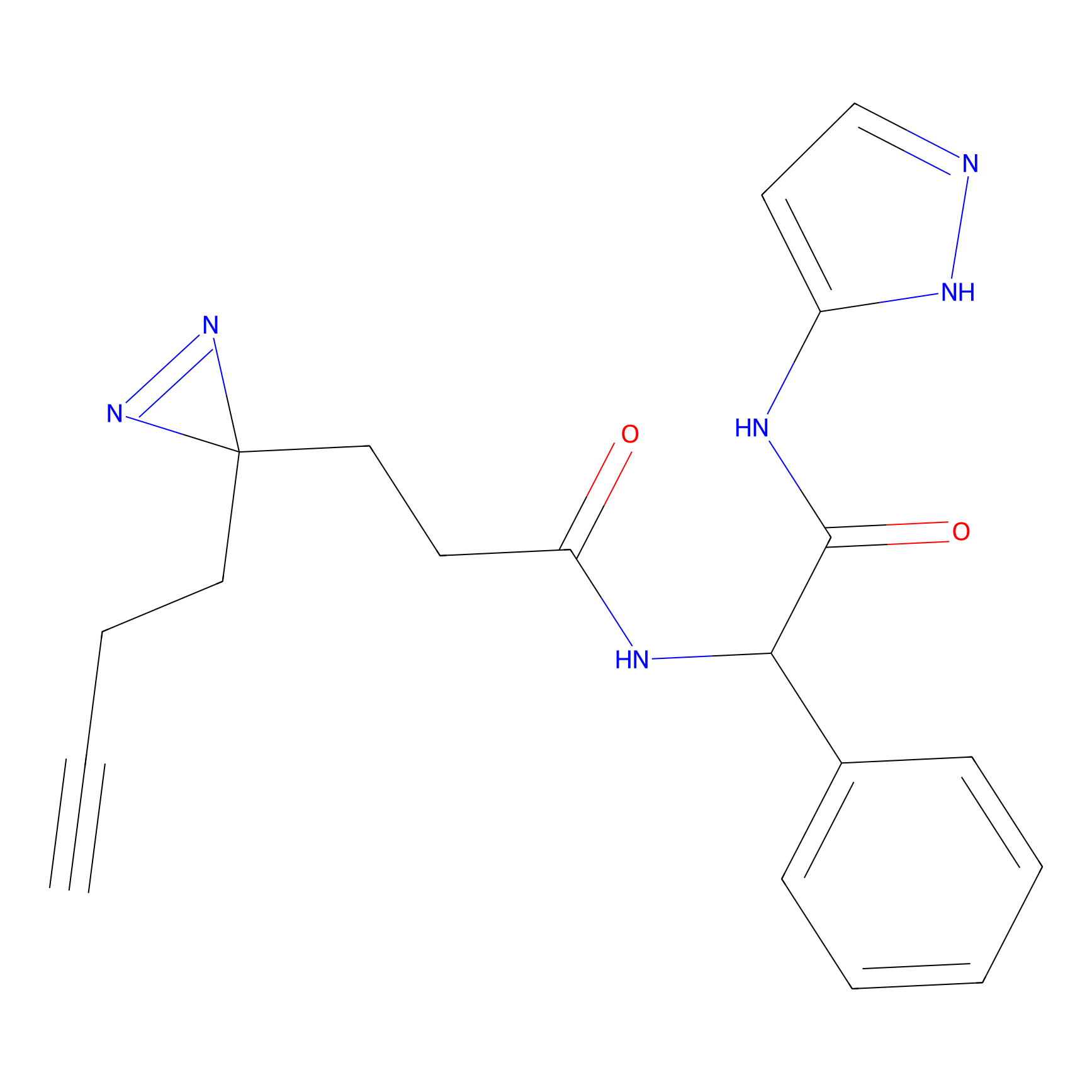
|
AOyne Probe Info |
 |
10.10 | LDD0443 | [4] | |
PAL-AfBPP Probe
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
GPCR
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| B-cell antigen receptor complex-associated protein alpha chain (CD79A) | . | P11912 | |||
Other
References