Details of the Target
General Information of Target
| Target ID | LDTP07667 | |||||
|---|---|---|---|---|---|---|
| Target Name | Transmembrane protein 205 (TMEM205) | |||||
| Gene Name | TMEM205 | |||||
| Gene ID | 374882 | |||||
| Synonyms |
Transmembrane protein 205 |
|||||
| 3D Structure | ||||||
| Sequence |
MEEGGNLGGLIKMVHLLVLSGAWGMQMWVTFVSGFLLFRSLPRHTFGLVQSKLFPFYFHI
SMGCAFINLCILASQHAWAQLTFWEASQLYLLFLSLTLATVNARWLEPRTTAAMWALQTV EKERGLGGEVPGSHQGPDPYRQLREKDPKYSALRQNFFRYHGLSSLCNLGCVLSNGLCLA GLALEIRSL |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
TMEM205 family
|
|||||
| Subcellular location |
Membrane
|
|||||
| Function | In cancer cells, plays a role in resistance to the chemotherapeutic agent cisplatin. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
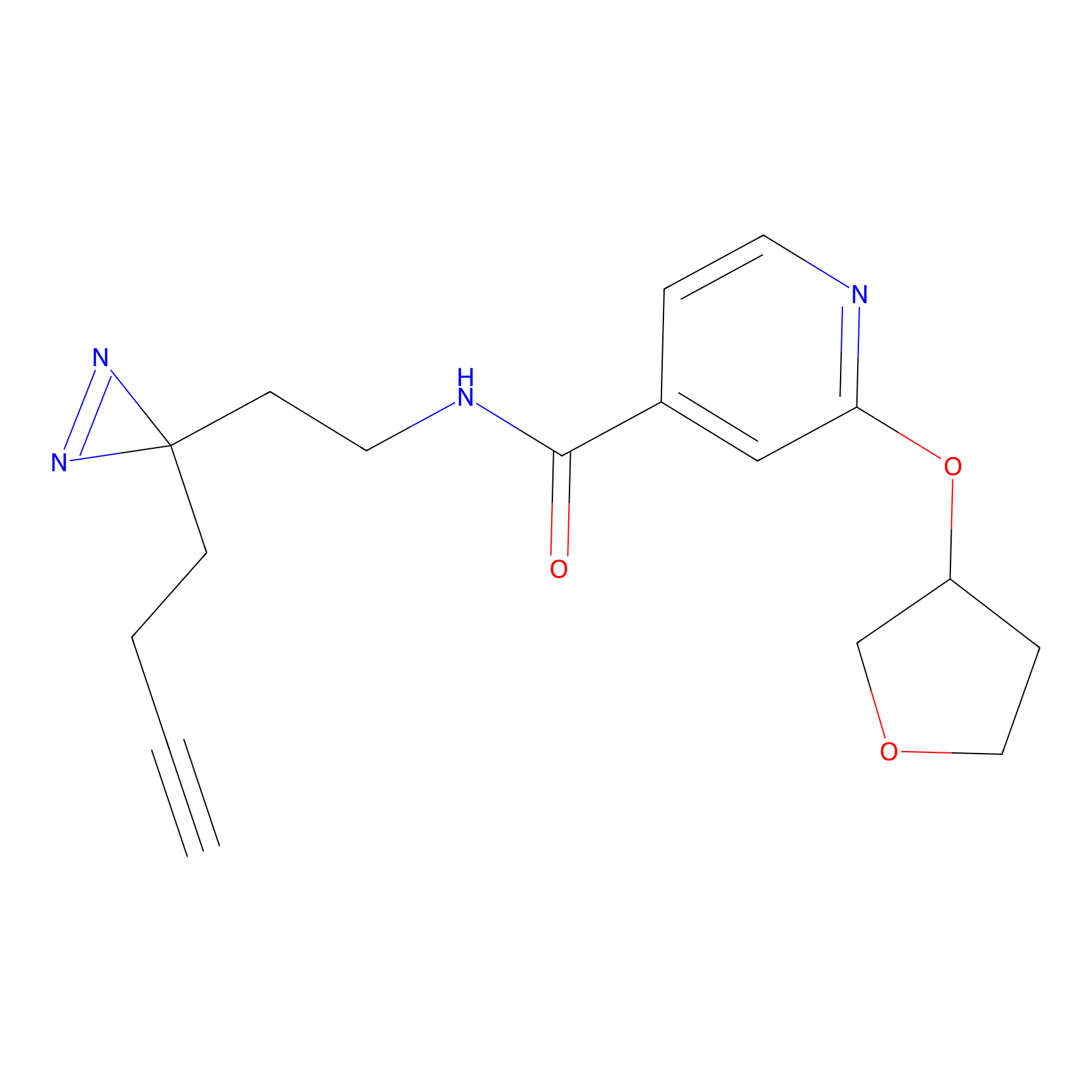
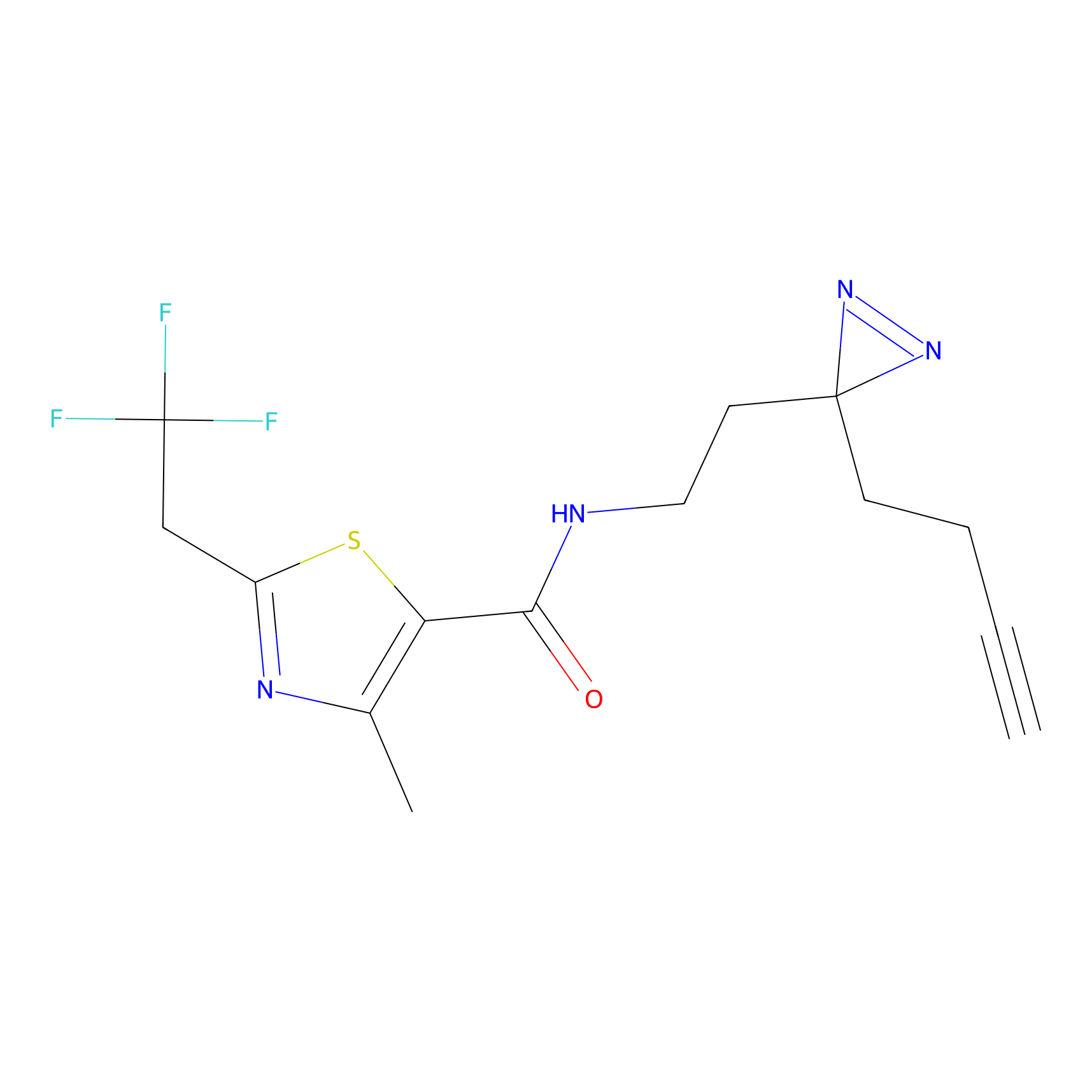
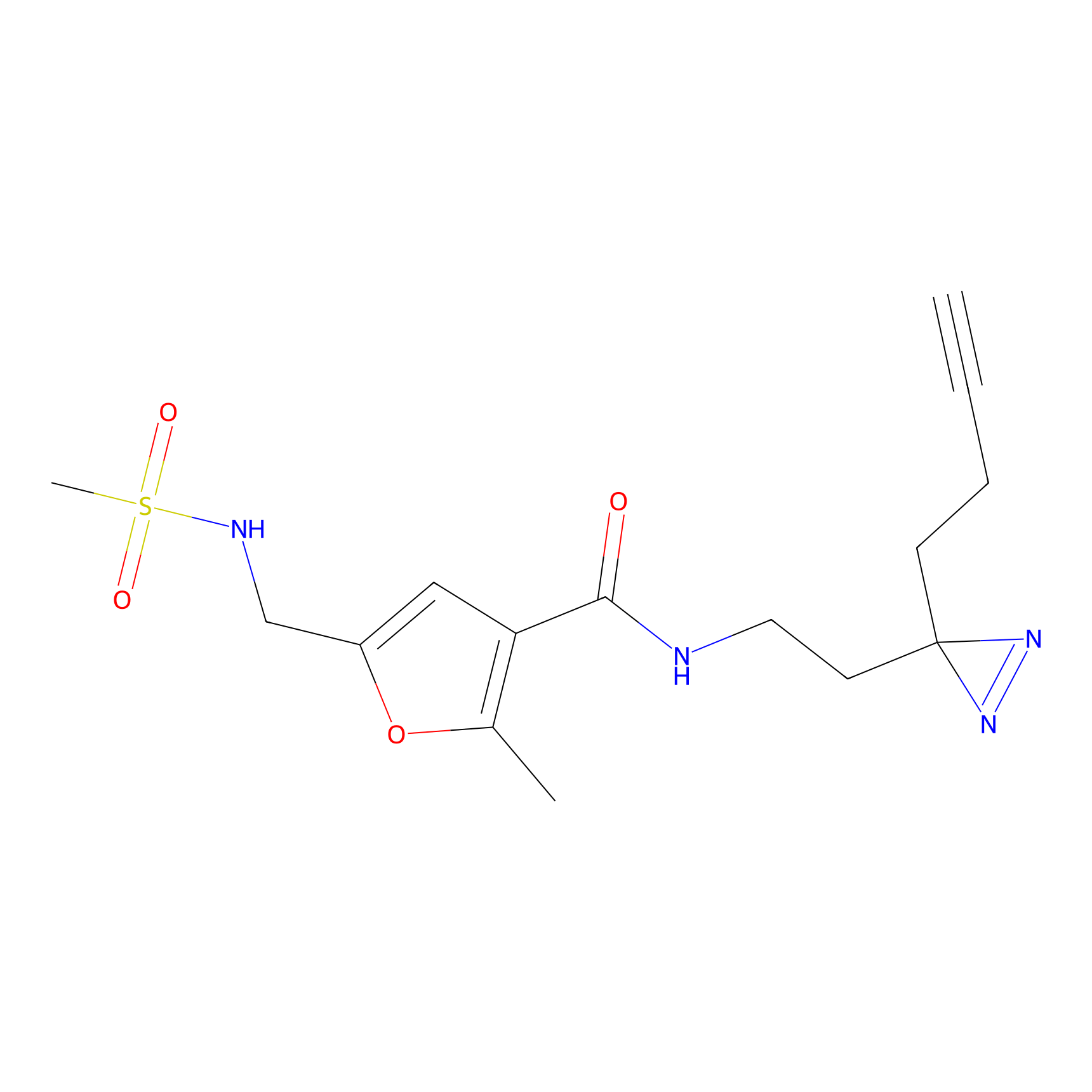
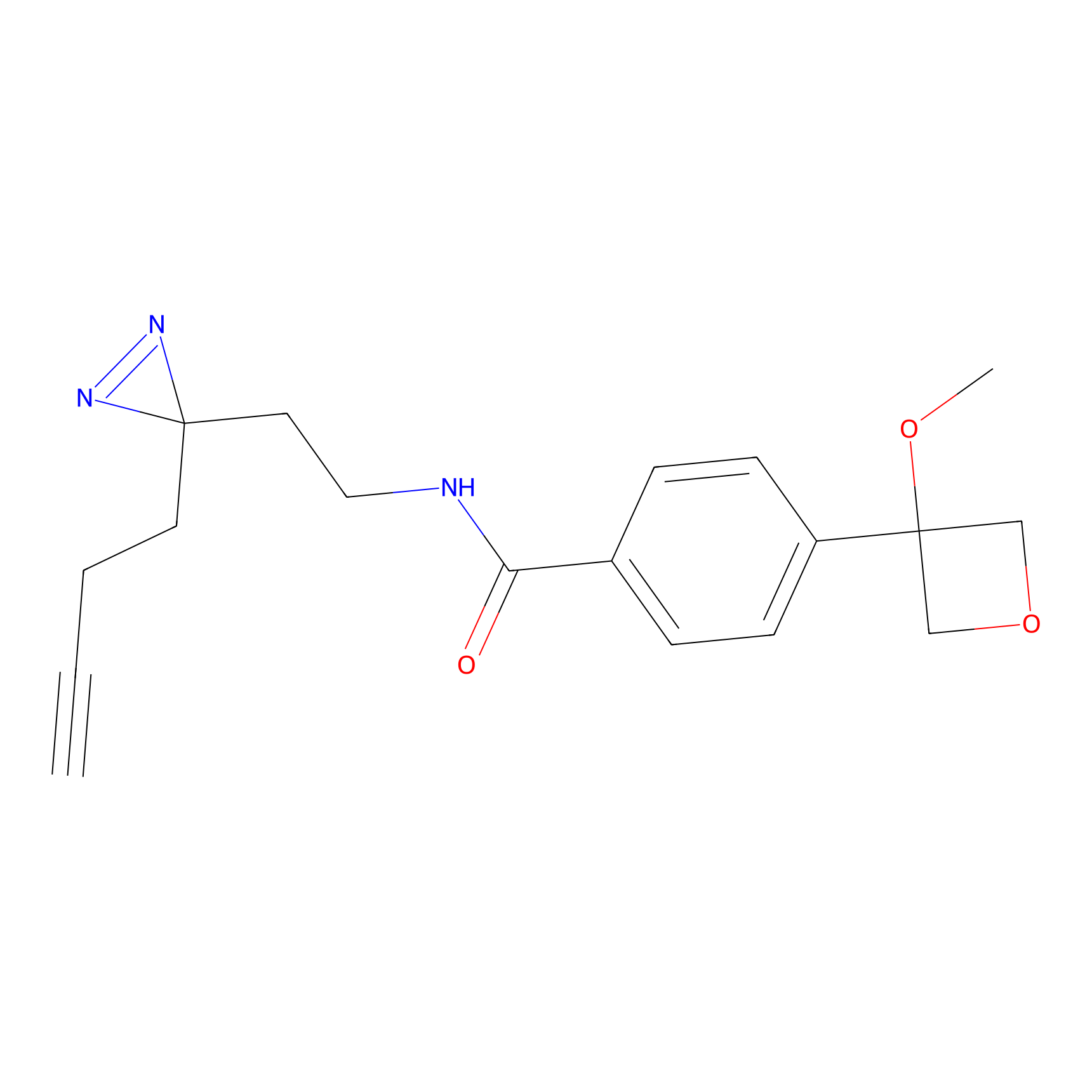
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
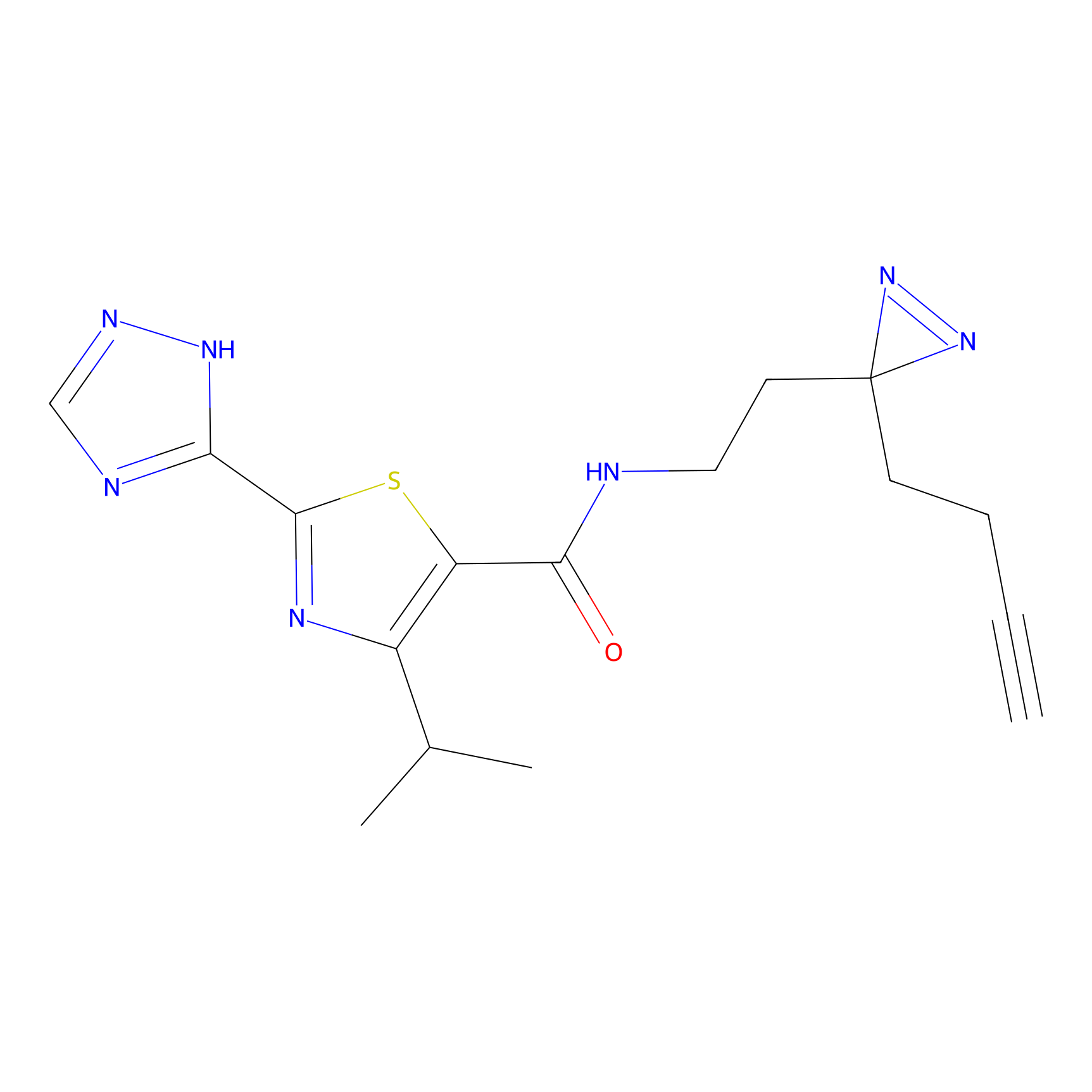
|
TH211 Probe Info |
 |
Y140(11.11) | LDD0260 | [1] | |
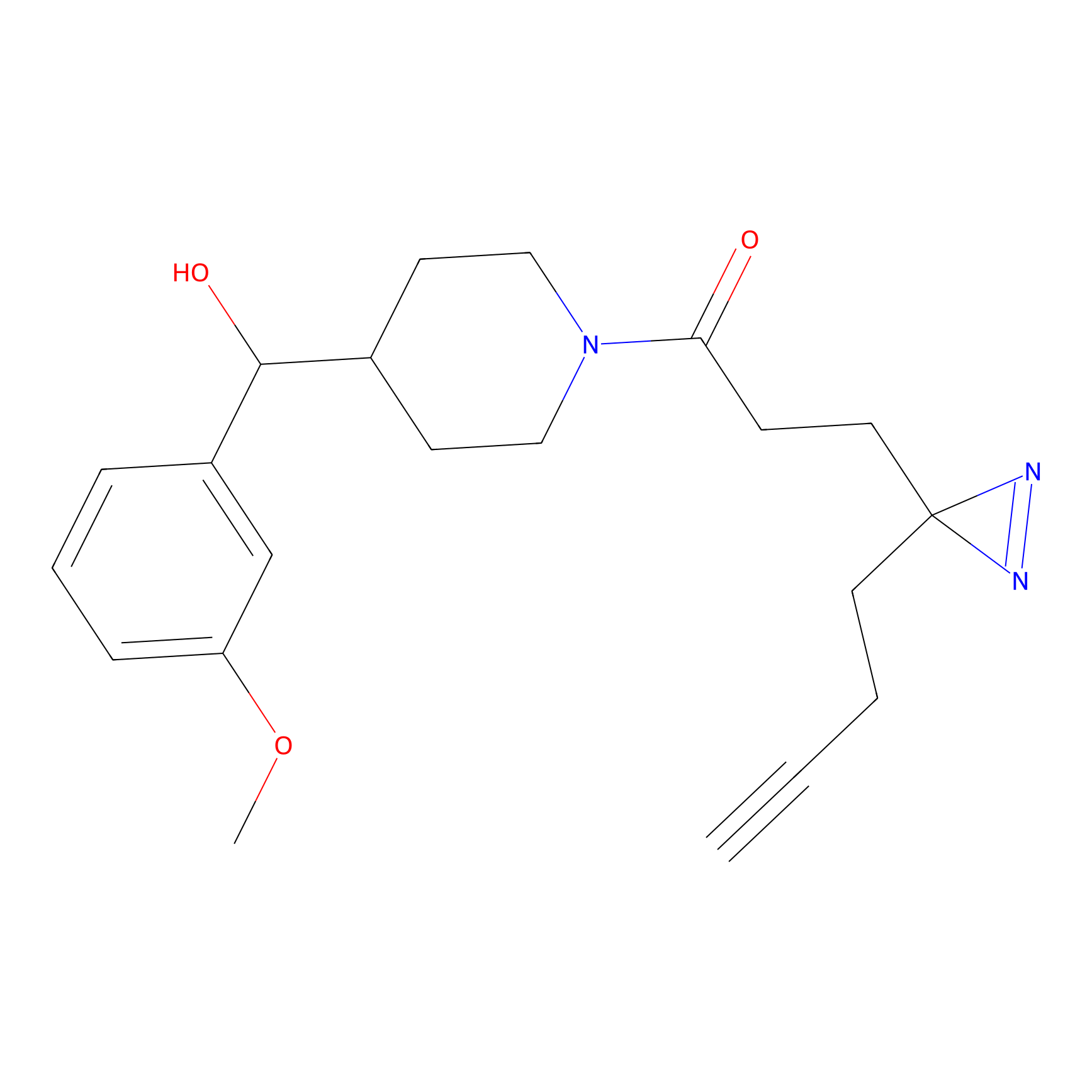
|
C-Sul Probe Info |
 |
4.65 | LDD0066 | [2] | |
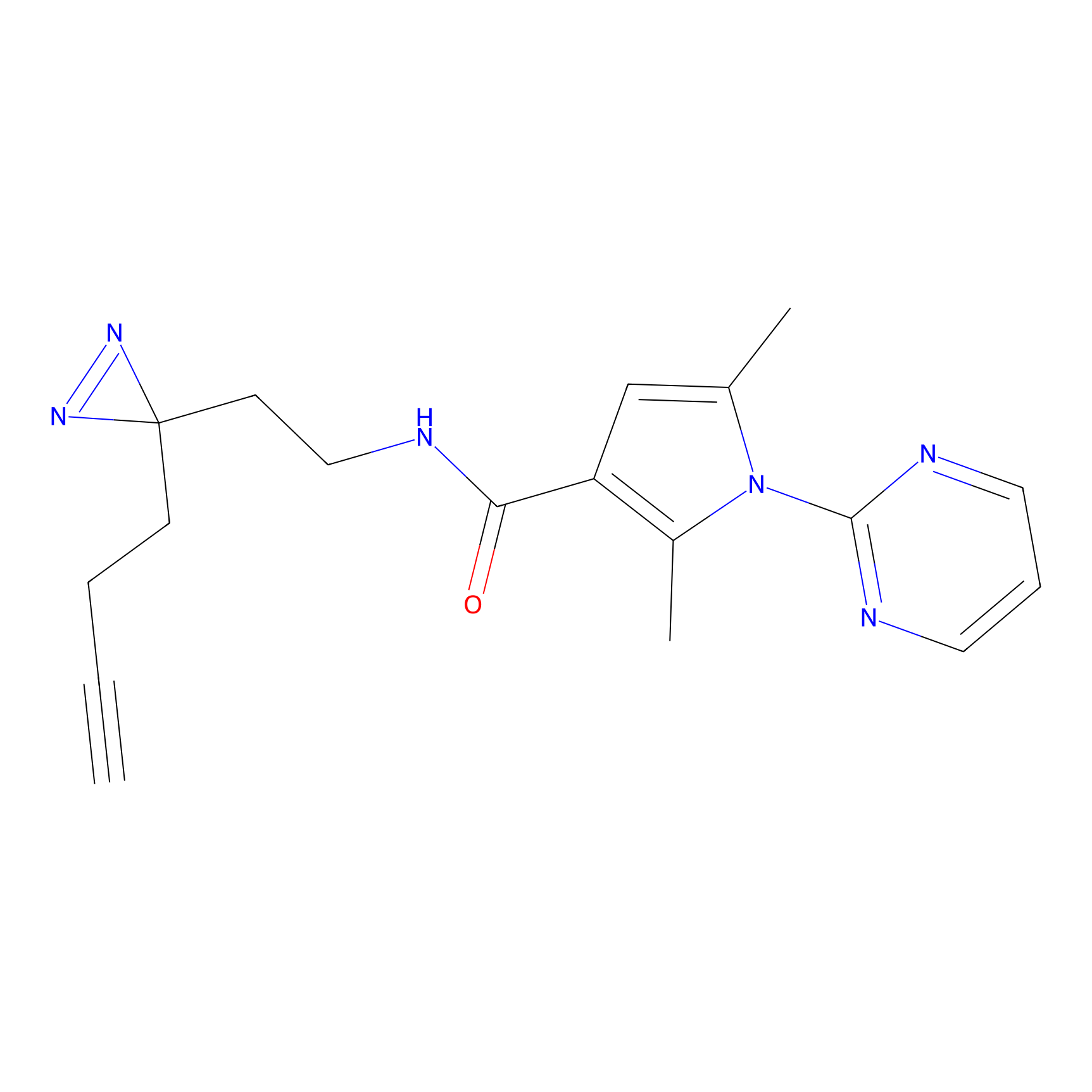
|
STPyne Probe Info |
 |
K149(10.00) | LDD0277 | [3] | |
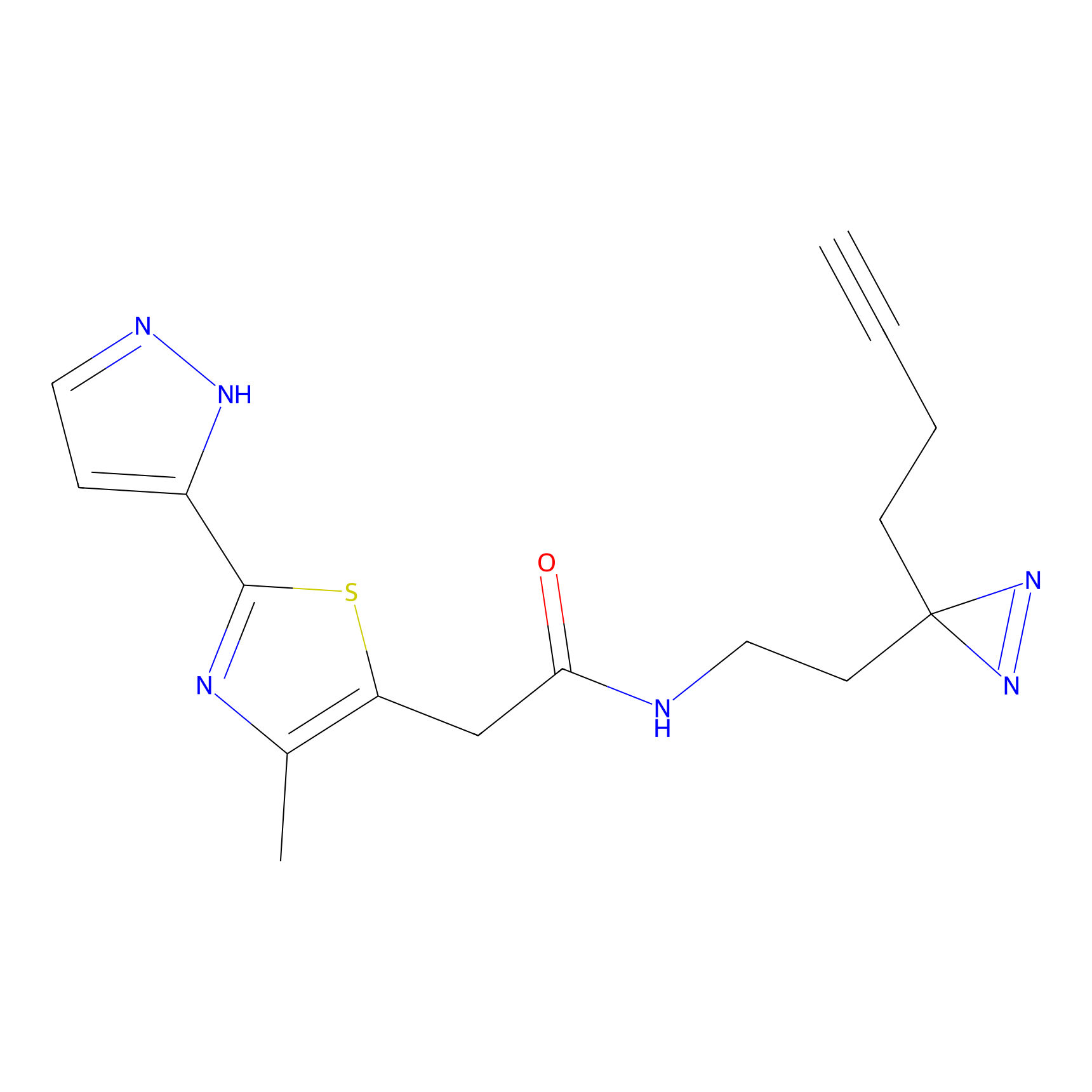
|
Probe 1 Probe Info |
 |
Y140(10.73) | LDD3495 | [4] | |
|
m-APA Probe Info |
 |
13.10 | LDD0403 | [5] | |
|
Jackson_1 Probe Info |
 |
20.00 | LDD0120 | [6] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0221 | [7] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C171(0.00); C178(0.00); C167(0.00) | LDD0038 | [8] | |
|
IA-alkyne Probe Info |
 |
C171(0.00); C178(0.00); C167(0.00) | LDD0036 | [8] | |
|
Lodoacetamide azide Probe Info |
 |
C171(0.00); C178(0.00); C167(0.00) | LDD0037 | [8] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [9] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [7] | |
|
AOyne Probe Info |
 |
10.00 | LDD0443 | [10] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Other
References