Details of the Target
General Information of Target
| Target ID | LDTP01671 | |||||
|---|---|---|---|---|---|---|
| Target Name | STARD3 N-terminal-like protein (STARD3NL) | |||||
| Gene Name | STARD3NL | |||||
| Gene ID | 83930 | |||||
| Synonyms |
MENTHO; STARD3 N-terminal-like protein; MLN64 N-terminal domain homolog |
|||||
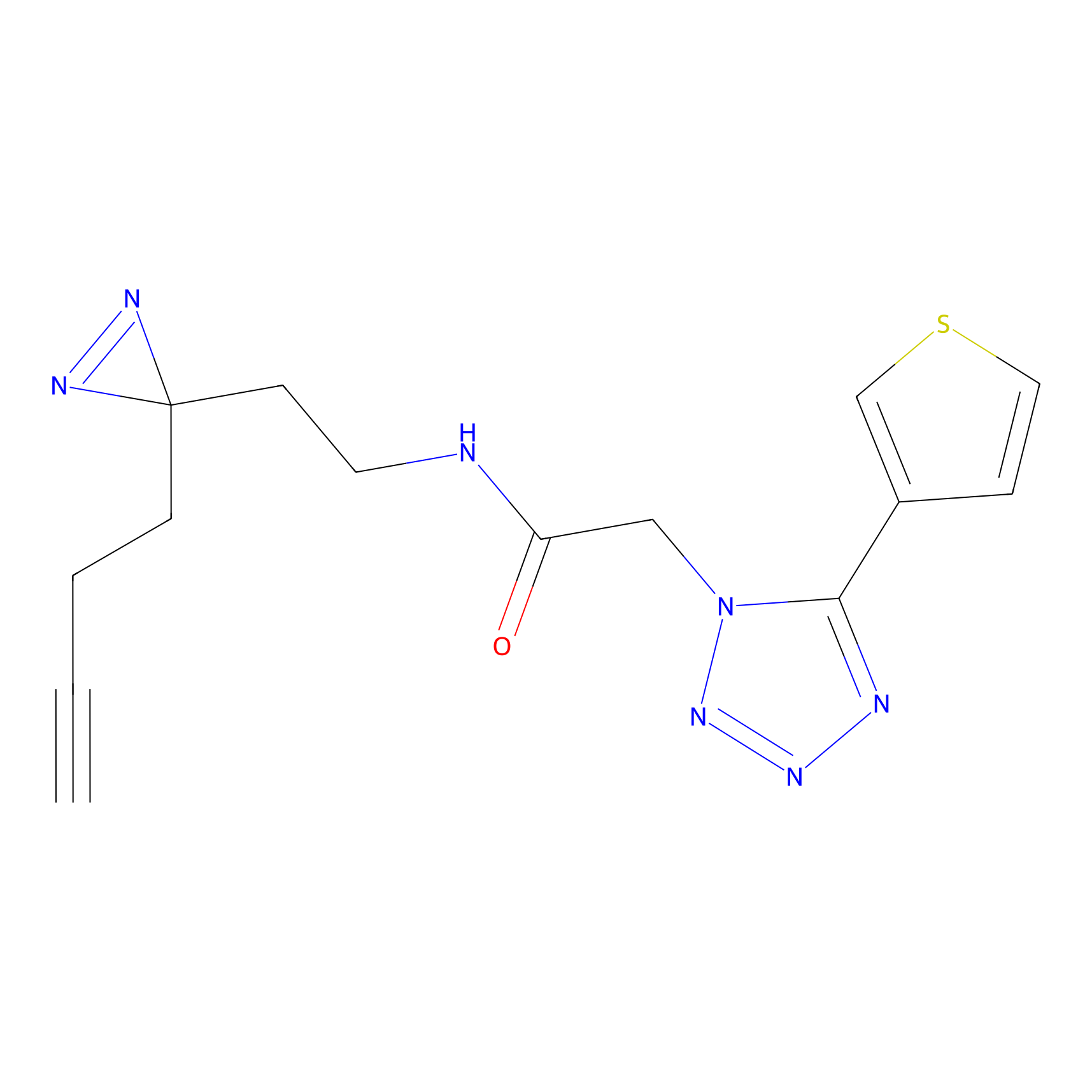
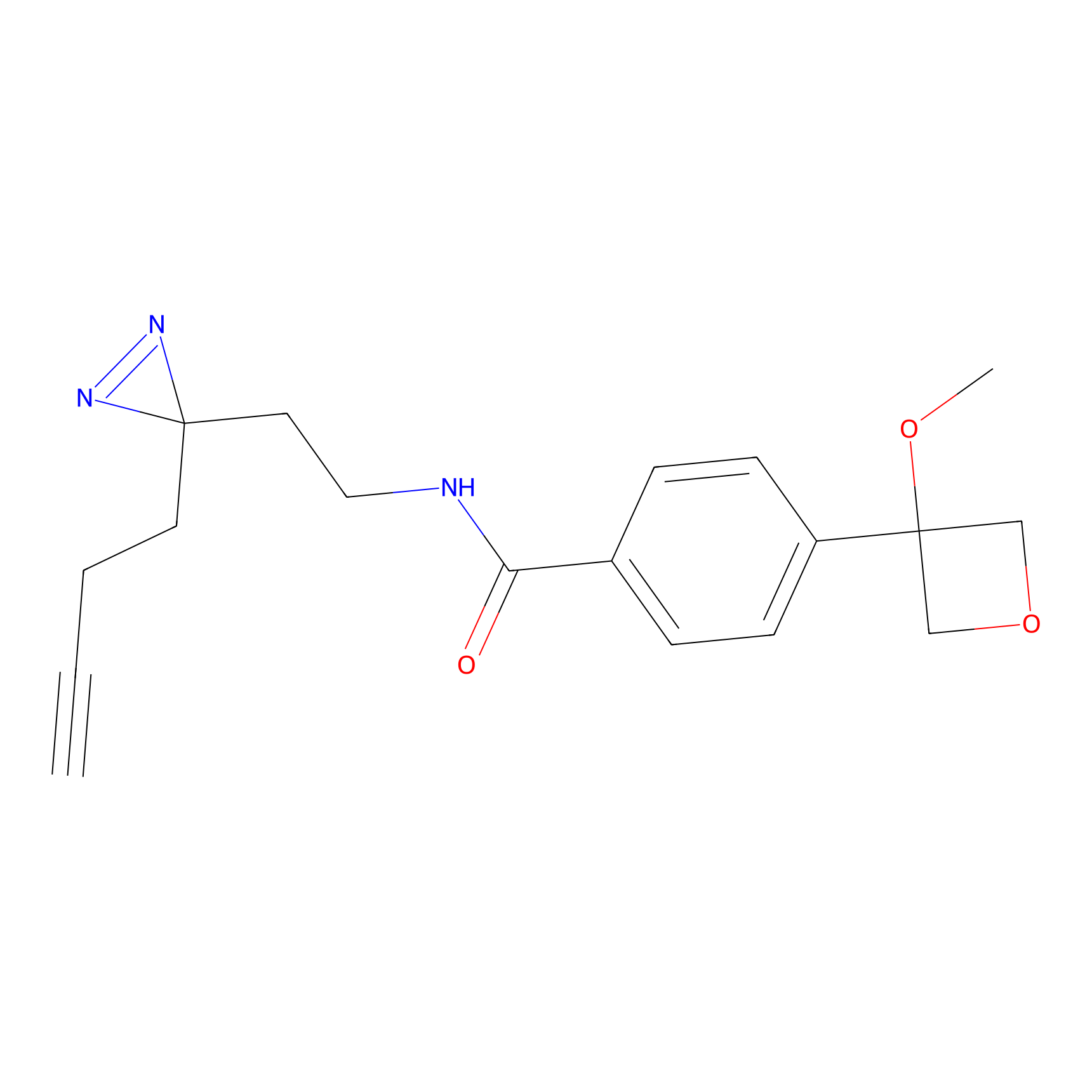
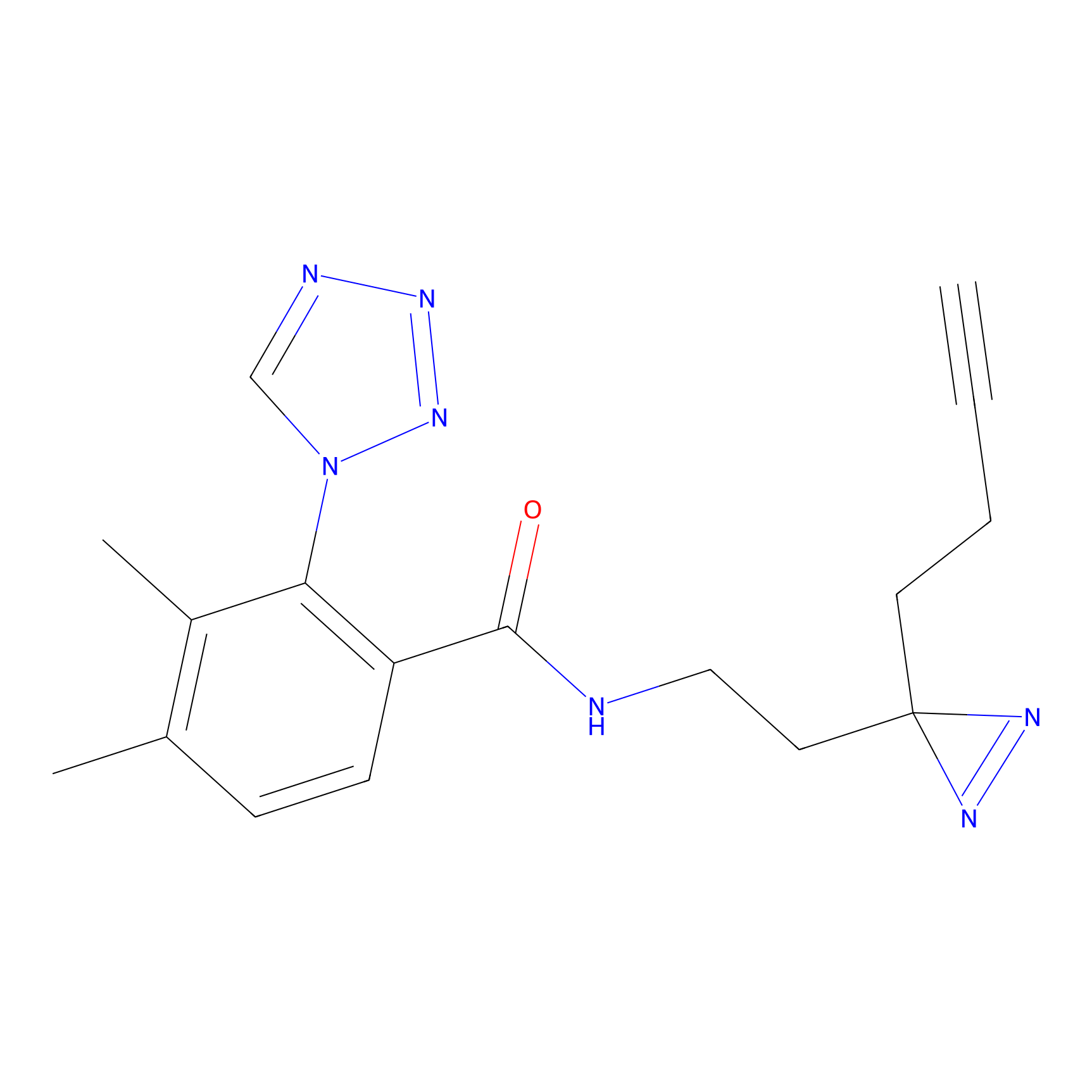
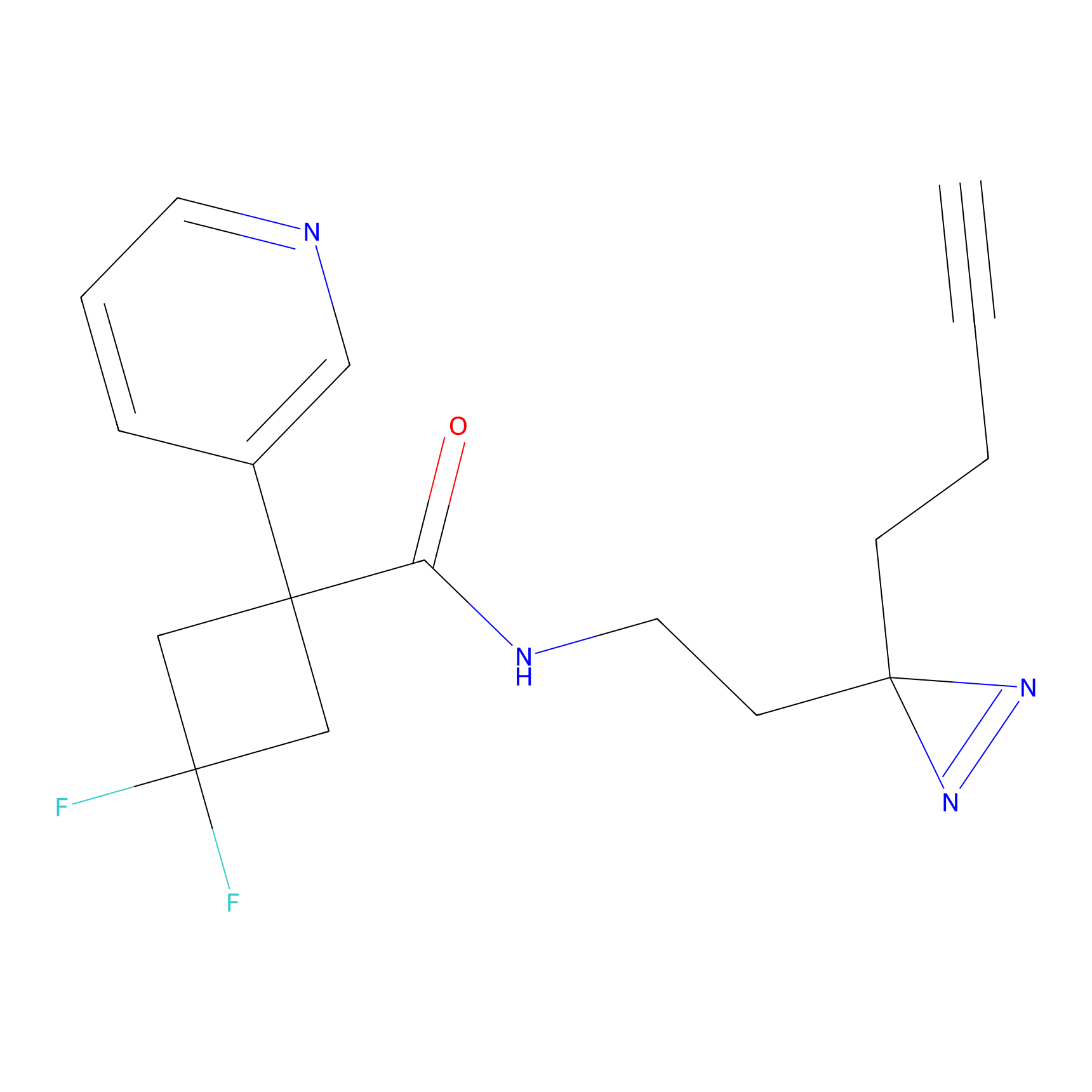
| 3D Structure | ||||||
| Sequence |
MNHLPEDMENALTGSQSSHASLRNIHSINPTQLMARIESYEGREKKGISDVRRTFCLFVT
FDLLFVTLLWIIELNVNGGIENTLEKEVMQYDYYSSYFDIFLLAVFRFKVLILAYAVCRL RHWWAIALTTAVTSAFLLAKVILSKLFSQGAFGYVLPIISFILAWIETWFLDFKVLPQEA EEENRLLIVQDASERAALIPGGLSDGQFYSPPESEAGSEEAEEKQDSEKPLLEL |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
STARD3 family
|
|||||
| Subcellular location |
Late endosome membrane
|
|||||
| Function |
Tethering protein that creates contact site between the endoplasmic reticulum and late endosomes: localizes to late endosome membranes and contacts the endoplasmic reticulum via interaction with VAPA and VAPB.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
6.86 | LDD0318 | [1] | |
PAL-AfBPP Probe
The Interaction Atlas With This Target
References