Details of the Target
General Information of Target
| Target ID | LDTP12867 | |||||
|---|---|---|---|---|---|---|
| Target Name | Very-long-chain enoyl-CoA reductase (TECR) | |||||
| Gene Name | TECR | |||||
| Gene ID | 9524 | |||||
| Synonyms |
GPSN2; SC2; Very-long-chain enoyl-CoA reductase; EC 1.3.1.93; Synaptic glycoprotein SC2; Trans-2,3-enoyl-CoA reductase; TER |
|||||
| 3D Structure | ||||||
| Sequence |
MLLLLLLLPLLWGTKGMEGDRQYGDGYLLQVQELVTVQEGLCVHVPCSFSYPQDGWTDSD
PVHGYWFRAGDRPYQDAPVATNNPDREVQAETQGRFQLLGDIWSNDCSLSIRDARKRDKG SYFFRLERGSMKWSYKSQLNYKTKQLSVFVTALTHRPDILILGTLESGHSRNLTCSVPWA CKQGTPPMISWIGASVSSPGPTTARSSVLTLTPKPQDHGTSLTCQVTLPGTGVTTTSTVR LDVSYPPWNLTMTVFQGDATASTALGNGSSLSVLEGQSLRLVCAVNSNPPARLSWTRGSL TLCPSRSSNPGLLELPRVHVRDEGEFTCRAQNAQGSQHISLSLSLQNEGTGTSRPVSQVT LAAVGGAGATALAFLSFCIIFIIVRSCRKKSARPAAGVGDTGMEDAKAIRGSASQGPLTE SWKDGNPLKKPPPAVAPSSGEEGELHYATLSFHKVKPQDPQGQEATDSEYSEIKIHKRET AETQACLRNHNPSSKEVRG |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Steroid 5-alpha reductase family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Involved in both the production of very long-chain fatty acids for sphingolipid synthesis and the degradation of the sphingosine moiety in sphingolipids through the sphingosine 1-phosphate metabolic pathway. Catalyzes the last of the four reactions of the long-chain fatty acids elongation cycle. This endoplasmic reticulum-bound enzymatic process, allows the addition of 2 carbons to the chain of long- and very long-chain fatty acids/VLCFAs per cycle. This enzyme reduces the trans-2,3-enoyl-CoA fatty acid intermediate to an acyl-CoA that can be further elongated by entering a new cycle of elongation. Thereby, it participates in the production of VLCFAs of different chain lengths that are involved in multiple biological processes as precursors of membrane lipids and lipid mediators. Catalyzes the saturation step of the sphingosine 1-phosphate metabolic pathway, the conversion of trans-2-hexadecenoyl-CoA to palmitoyl-CoA.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| HCT15 | SNV: p.V23M | . | |||
| IM95 | SNV: p.M261T | DBIA Probe Info | |||
| Ishikawa (Heraklio) 02 ER | Deletion: p.A83PfsTer6 | . | |||
| SKMEL2 | SNV: p.T145A | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
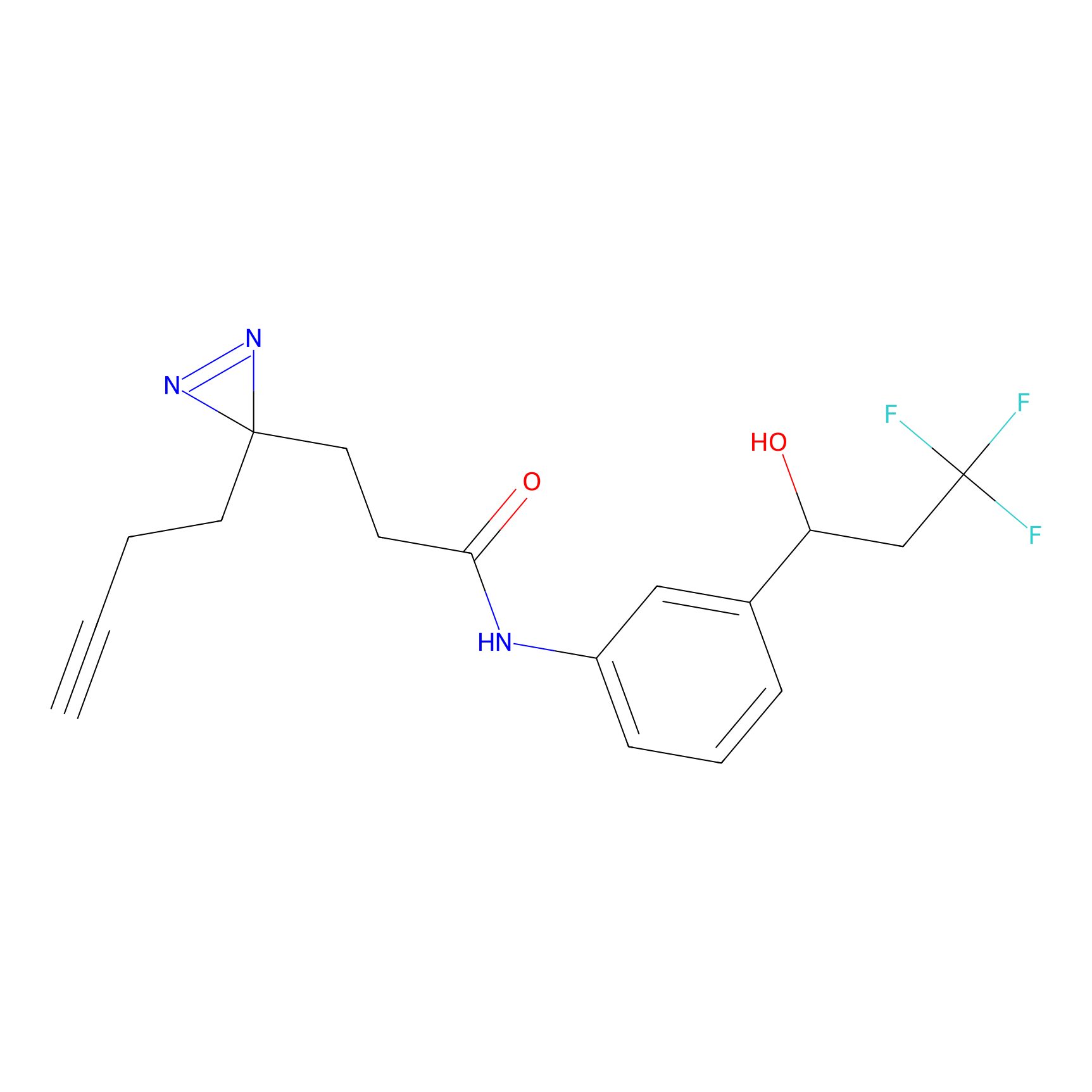
|
m-APA Probe Info |
 |
9.47 | LDD0402 | [1] | |
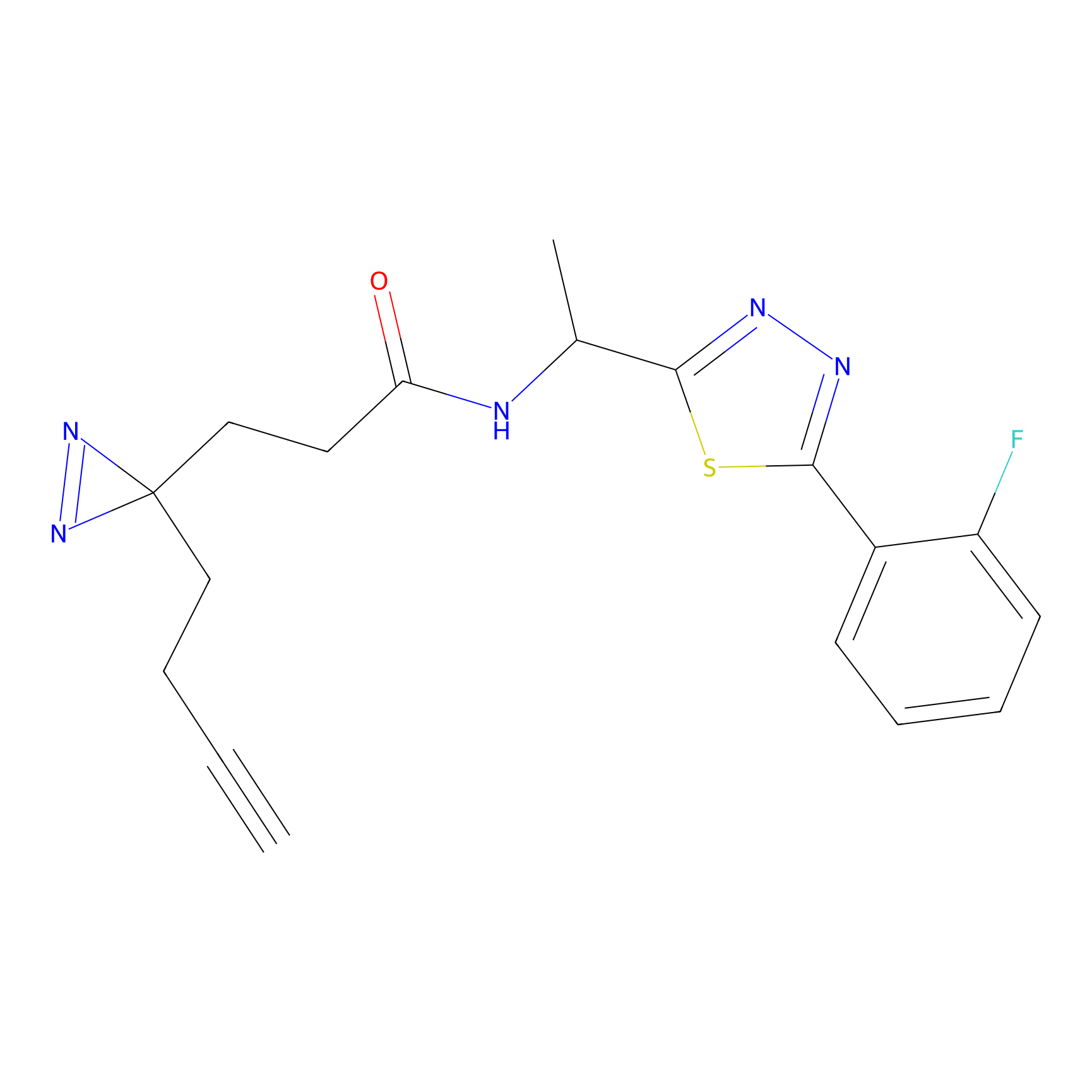
|
HDSF-alk Probe Info |
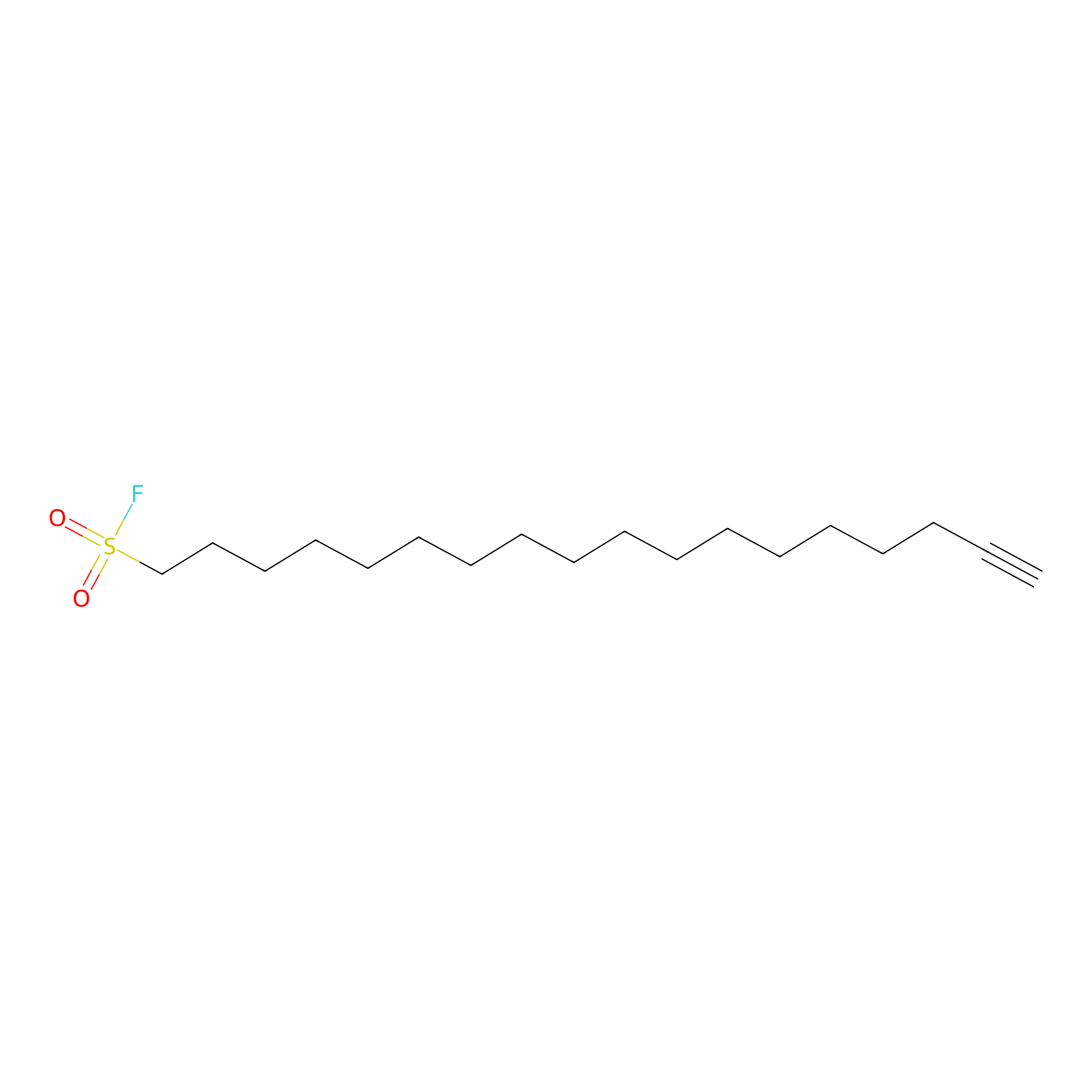 |
2.03 | LDD0197 | [2] | |
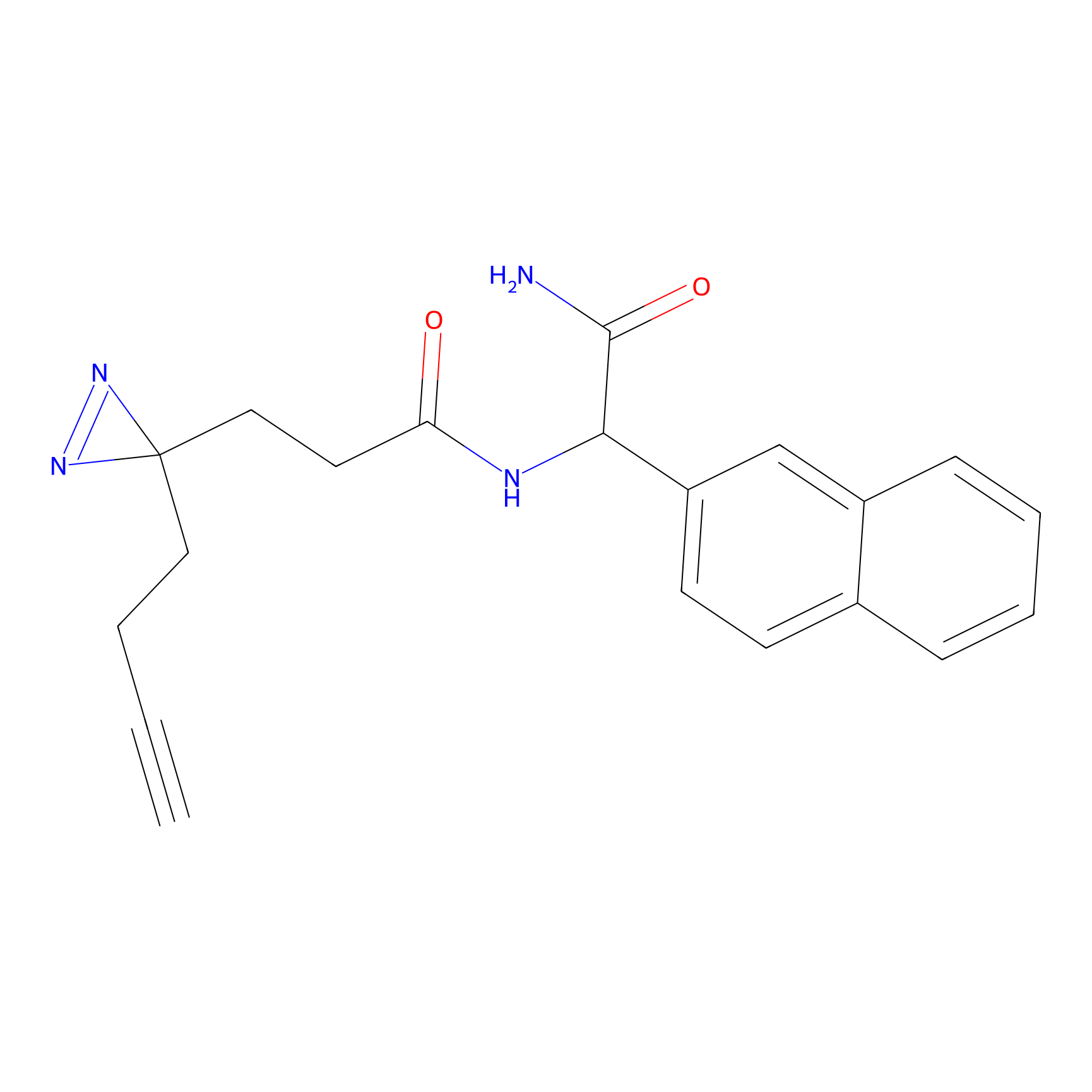
|
P3 Probe Info |
 |
3.42 | LDD0450 | [3] | |
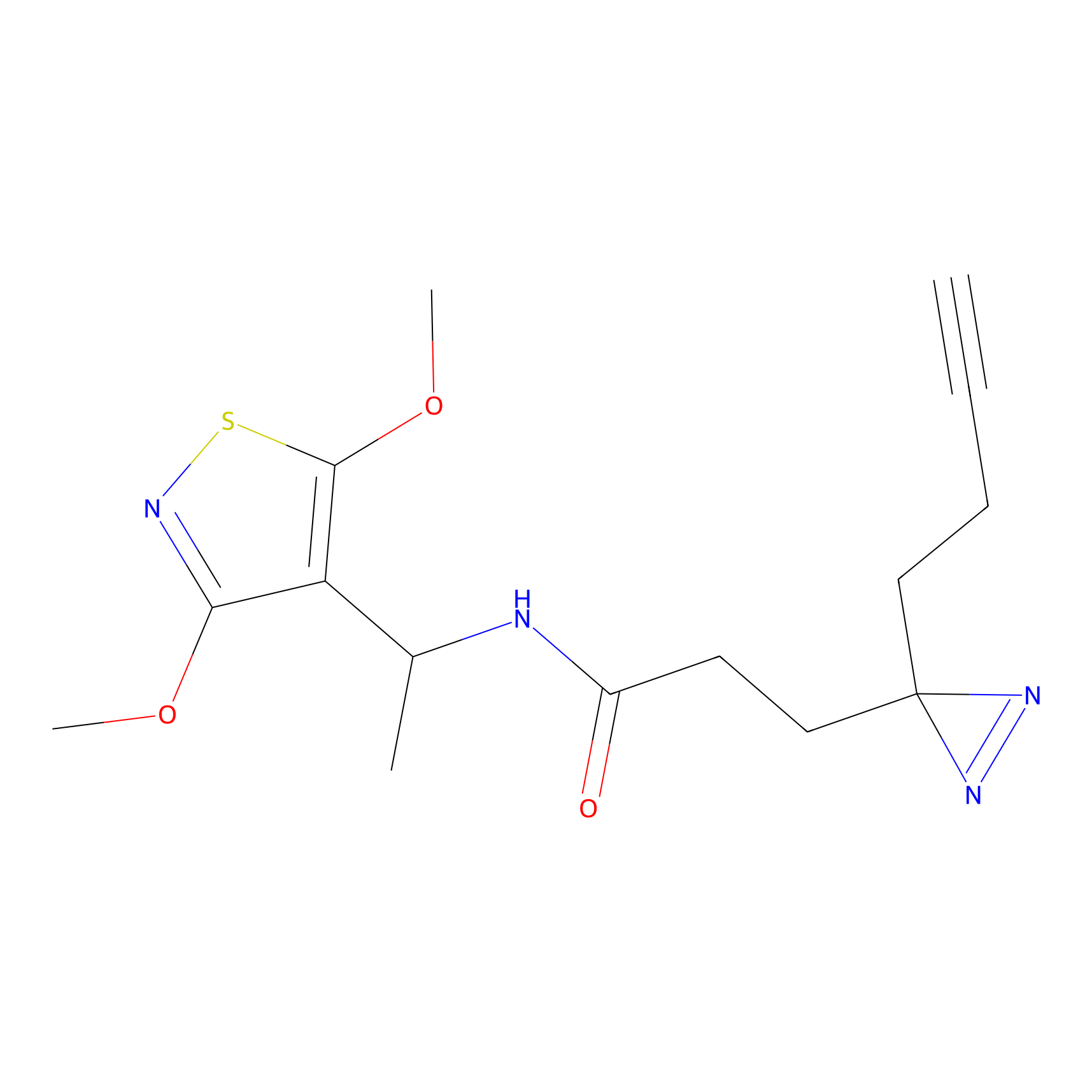
|
CHEMBL5175495 Probe Info |
 |
8.92 | LDD0196 | [4] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [5] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [5] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [5] | |
|
TH211 Probe Info |
 |
Y44(20.00); Y77(20.00); Y4(18.42); Y117(16.81) | LDD0260 | [6] | |
|
TH216 Probe Info |
 |
Y44(16.88) | LDD0259 | [6] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [7] | |
|
ONAyne Probe Info |
 |
N.A. | LDD0273 | [8] | |
|
AF-1 Probe Info |
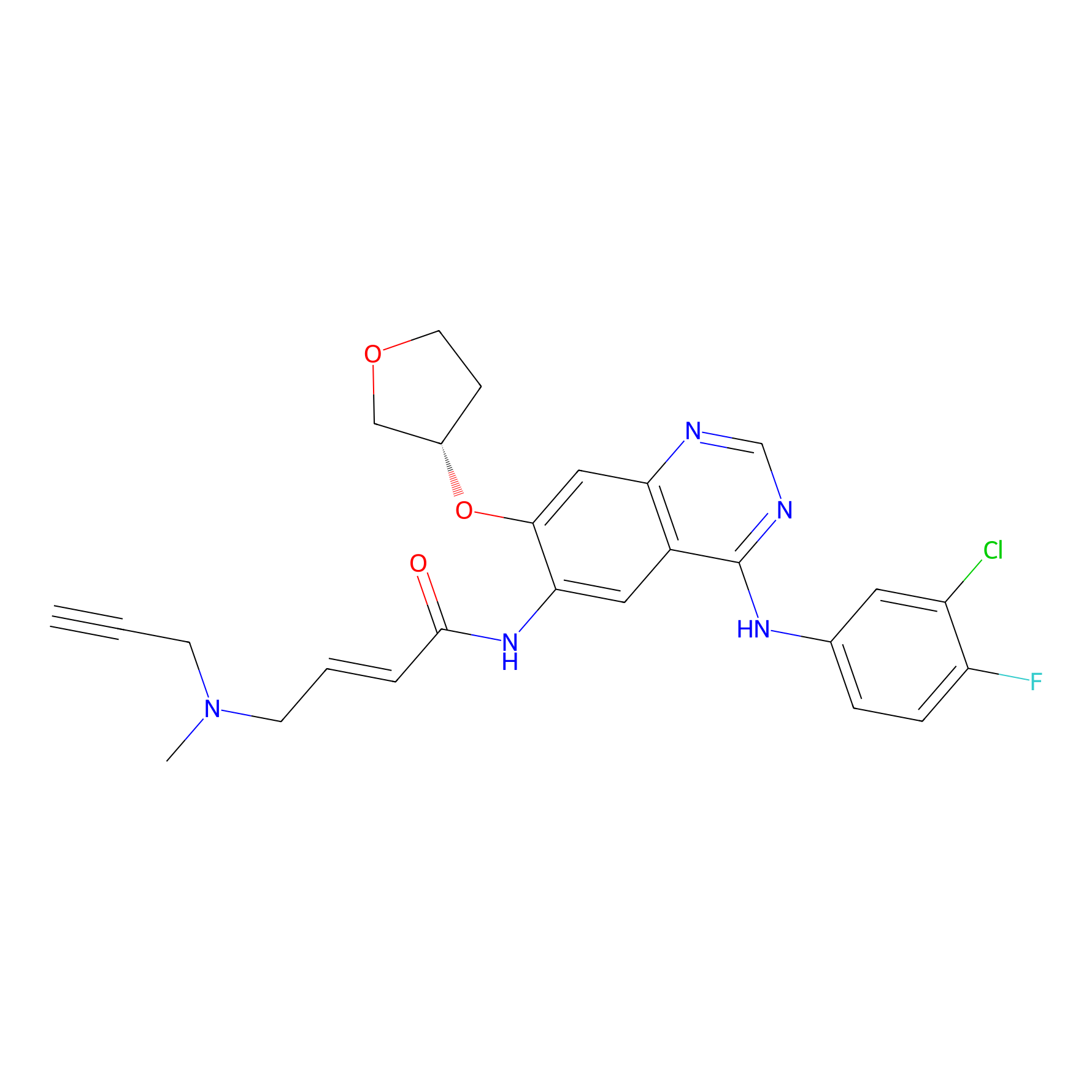 |
10.00 | LDD0421 | [9] | |
|
BTD Probe Info |
 |
C18(2.08) | LDD1700 | [10] | |
|
EA-probe Probe Info |
 |
C18(1.06); C18(1.10) | LDD2210 | [11] | |
|
DBIA Probe Info |
 |
C18(192.49) | LDD0209 | [12] | |
|
ATP probe Probe Info |
 |
K12(0.00); K2(0.00); K60(0.00) | LDD0199 | [13] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [14] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [15] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [16] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [14] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [17] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [18] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [19] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [20] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [21] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [21] | |
|
NHS Probe Info |
 |
K60(0.00); K38(0.00) | LDD0010 | [21] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [22] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [21] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [18] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [21] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [23] | |
|
Acrolein Probe Info |
 |
H3(0.00); C18(0.00); H26(0.00); K2(0.00) | LDD0217 | [24] | |
|
Crotonaldehyde Probe Info |
 |
H153(0.00); H3(0.00); H40(0.00); H149(0.00) | LDD0219 | [24] | |
|
Methacrolein Probe Info |
 |
C18(0.00); K22(0.00); H3(0.00) | LDD0218 | [24] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [25] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [19] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [26] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C18(0.98) | LDD2142 | [10] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C18(1.06) | LDD2112 | [10] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C18(0.68) | LDD2095 | [10] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C18(0.64) | LDD2130 | [10] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C18(1.20) | LDD2117 | [10] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C18(1.41) | LDD2152 | [10] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C18(0.90) | LDD2103 | [10] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C18(0.57) | LDD2132 | [10] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C18(0.64) | LDD2131 | [10] |
| LDCM0214 | AC1 | HEK-293T | C165(0.96); C133(1.15); C131(1.01); C133(1.01) | LDD1507 | [32] |
| LDCM0215 | AC10 | HEK-293T | C131(0.91); C133(0.91) | LDD1508 | [32] |
| LDCM0226 | AC11 | HEK-293T | C165(0.88); C131(1.05); C133(0.84); C131(1.34) | LDD1509 | [32] |
| LDCM0237 | AC12 | HEK-293T | C133(0.86) | LDD1510 | [32] |
| LDCM0259 | AC14 | HEK-293T | C131(1.16) | LDD1512 | [32] |
| LDCM0270 | AC15 | HEK-293T | C165(0.94); C131(0.95); C133(0.95) | LDD1513 | [32] |
| LDCM0276 | AC17 | HEK-293T | C165(1.21); C133(0.98); C131(0.94); C133(0.94) | LDD1515 | [32] |
| LDCM0277 | AC18 | HEK-293T | C131(0.84); C133(0.84) | LDD1516 | [32] |
| LDCM0278 | AC19 | HEK-293T | C165(0.84); C131(0.75); C133(0.71); C131(1.65) | LDD1517 | [32] |
| LDCM0279 | AC2 | HEK-293T | C131(0.91); C133(0.91) | LDD1518 | [32] |
| LDCM0280 | AC20 | HEK-293T | C133(0.89) | LDD1519 | [32] |
| LDCM0282 | AC22 | HEK-293T | C131(1.10) | LDD1521 | [32] |
| LDCM0283 | AC23 | HEK-293T | C165(1.12); C131(0.96); C133(0.96) | LDD1522 | [32] |
| LDCM0285 | AC25 | HEK-293T | C165(1.07); C133(1.02); C131(0.89); C133(0.89) | LDD1524 | [32] |
| LDCM0286 | AC26 | HEK-293T | C131(0.88); C133(0.88) | LDD1525 | [32] |
| LDCM0287 | AC27 | HEK-293T | C165(0.76); C131(0.96); C133(0.90); C131(1.41) | LDD1526 | [32] |
| LDCM0288 | AC28 | HEK-293T | C133(0.87) | LDD1527 | [32] |
| LDCM0290 | AC3 | HEK-293T | C165(0.65); C131(0.93); C133(0.93); C131(1.39) | LDD1529 | [32] |
| LDCM0291 | AC30 | HEK-293T | C131(0.96) | LDD1530 | [32] |
| LDCM0292 | AC31 | HEK-293T | C165(0.87); C131(1.02); C133(1.02) | LDD1531 | [32] |
| LDCM0294 | AC33 | HEK-293T | C165(1.02); C133(1.06); C131(1.06); C133(1.06) | LDD1533 | [32] |
| LDCM0295 | AC34 | HEK-293T | C131(0.80); C133(0.80) | LDD1534 | [32] |
| LDCM0296 | AC35 | HEK-293T | C165(0.71); C131(1.08); C133(0.95); C131(1.04) | LDD1535 | [32] |
| LDCM0297 | AC36 | HEK-293T | C133(0.97) | LDD1536 | [32] |
| LDCM0299 | AC38 | HEK-293T | C131(1.06) | LDD1538 | [32] |
| LDCM0300 | AC39 | HEK-293T | C165(0.85); C131(0.93); C133(0.93) | LDD1539 | [32] |
| LDCM0301 | AC4 | HEK-293T | C133(0.88) | LDD1540 | [32] |
| LDCM0303 | AC41 | HEK-293T | C165(0.85); C133(1.10); C131(1.03); C133(1.03) | LDD1542 | [32] |
| LDCM0304 | AC42 | HEK-293T | C131(0.82); C133(0.82) | LDD1543 | [32] |
| LDCM0305 | AC43 | HEK-293T | C165(0.86); C131(1.09); C133(0.92); C131(1.34) | LDD1544 | [32] |
| LDCM0306 | AC44 | HEK-293T | C133(0.94) | LDD1545 | [32] |
| LDCM0308 | AC46 | HEK-293T | C131(1.18) | LDD1547 | [32] |
| LDCM0309 | AC47 | HEK-293T | C165(1.03); C131(0.95); C133(0.95) | LDD1548 | [32] |
| LDCM0311 | AC49 | HEK-293T | C165(1.22); C133(0.90); C131(0.96); C133(0.96) | LDD1550 | [32] |
| LDCM0313 | AC50 | HEK-293T | C131(0.81); C133(0.81) | LDD1552 | [32] |
| LDCM0314 | AC51 | HEK-293T | C165(0.72); C131(0.98); C133(0.84); C131(1.12) | LDD1553 | [32] |
| LDCM0315 | AC52 | HEK-293T | C133(0.99) | LDD1554 | [32] |
| LDCM0317 | AC54 | HEK-293T | C131(1.01) | LDD1556 | [32] |
| LDCM0318 | AC55 | HEK-293T | C165(1.11); C131(0.93); C133(0.93) | LDD1557 | [32] |
| LDCM0320 | AC57 | HEK-293T | C165(0.94); C133(1.16); C131(1.02); C133(1.02) | LDD1559 | [32] |
| LDCM0321 | AC58 | HEK-293T | C131(0.85); C133(0.85) | LDD1560 | [32] |
| LDCM0322 | AC59 | HEK-293T | C165(0.89); C131(1.00); C133(0.90); C131(1.05) | LDD1561 | [32] |
| LDCM0323 | AC6 | HEK-293T | C131(1.17) | LDD1562 | [32] |
| LDCM0324 | AC60 | HEK-293T | C133(0.97) | LDD1563 | [32] |
| LDCM0326 | AC62 | HEK-293T | C131(1.00) | LDD1565 | [32] |
| LDCM0327 | AC63 | HEK-293T | C165(0.83); C131(0.96); C133(0.96) | LDD1566 | [32] |
| LDCM0334 | AC7 | HEK-293T | C165(0.89); C131(0.95); C133(0.95) | LDD1568 | [32] |
| LDCM0545 | Acetamide | MDA-MB-231 | C18(0.46) | LDD2138 | [10] |
| LDCM0166 | Afatinib | A431 | 10.00 | LDD0421 | [9] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C18(0.85) | LDD2113 | [10] |
| LDCM0356 | AKOS034007680 | HEK-293T | C165(0.93); C133(1.05); C131(0.98); C133(0.98) | LDD1570 | [32] |
| LDCM0156 | Aniline | NCI-H1299 | 11.36 | LDD0403 | [1] |
| LDCM0108 | Chloroacetamide | HeLa | C18(0.00); H3(0.00); H40(0.00); H153(0.00) | LDD0222 | [24] |
| LDCM0632 | CL-Sc | Hep-G2 | C18(0.86) | LDD2227 | [19] |
| LDCM0367 | CL1 | HEK-293T | C165(0.96); C131(1.30); C133(1.30) | LDD1571 | [32] |
| LDCM0368 | CL10 | HEK-293T | C131(3.01) | LDD1572 | [32] |
| LDCM0369 | CL100 | HEK-293T | C131(1.08); C133(1.08) | LDD1573 | [32] |
| LDCM0370 | CL101 | HEK-293T | C165(1.05); C131(1.01); C133(1.01) | LDD1574 | [32] |
| LDCM0371 | CL102 | HEK-293T | C165(1.15) | LDD1575 | [32] |
| LDCM0373 | CL104 | HEK-293T | C131(1.09); C133(1.09) | LDD1577 | [32] |
| LDCM0374 | CL105 | HEK-293T | C165(0.84); C131(1.03); C133(1.03) | LDD1578 | [32] |
| LDCM0375 | CL106 | HEK-293T | C165(1.22) | LDD1579 | [32] |
| LDCM0377 | CL108 | HEK-293T | C131(0.94); C133(0.94) | LDD1581 | [32] |
| LDCM0378 | CL109 | HEK-293T | C165(1.55); C131(1.18); C133(1.18) | LDD1582 | [32] |
| LDCM0379 | CL11 | HEK-293T | C165(1.29); C131(1.67); C133(1.67) | LDD1583 | [32] |
| LDCM0380 | CL110 | HEK-293T | C165(1.41) | LDD1584 | [32] |
| LDCM0382 | CL112 | HEK-293T | C131(1.00); C133(1.00) | LDD1586 | [32] |
| LDCM0383 | CL113 | HEK-293T | C165(0.85); C131(0.87); C133(0.87) | LDD1587 | [32] |
| LDCM0384 | CL114 | HEK-293T | C165(1.65) | LDD1588 | [32] |
| LDCM0386 | CL116 | HEK-293T | C131(0.94); C133(0.94) | LDD1590 | [32] |
| LDCM0387 | CL117 | HEK-293T | C165(0.90); C131(1.02); C133(1.02) | LDD1591 | [32] |
| LDCM0388 | CL118 | HEK-293T | C165(0.90) | LDD1592 | [32] |
| LDCM0391 | CL120 | HEK-293T | C131(1.05); C133(1.05) | LDD1595 | [32] |
| LDCM0392 | CL121 | HEK-293T | C165(0.96); C131(0.88); C133(0.88) | LDD1596 | [32] |
| LDCM0393 | CL122 | HEK-293T | C165(1.08) | LDD1597 | [32] |
| LDCM0395 | CL124 | HEK-293T | C131(1.05); C133(1.05) | LDD1599 | [32] |
| LDCM0396 | CL125 | HEK-293T | C165(1.04); C131(1.01); C133(1.01) | LDD1600 | [32] |
| LDCM0397 | CL126 | HEK-293T | C165(1.06) | LDD1601 | [32] |
| LDCM0399 | CL128 | HEK-293T | C131(0.91); C133(0.91) | LDD1603 | [32] |
| LDCM0400 | CL13 | HEK-293T | C165(1.06); C131(1.00); C133(1.00) | LDD1604 | [32] |
| LDCM0401 | CL14 | HEK-293T | C165(0.99) | LDD1605 | [32] |
| LDCM0403 | CL16 | HEK-293T | C131(1.06); C133(1.06) | LDD1607 | [32] |
| LDCM0404 | CL17 | HEK-293T | C165(1.60); C133(1.63); C131(1.75); C133(1.75) | LDD1608 | [32] |
| LDCM0405 | CL18 | HEK-293T | C131(0.93); C133(0.93) | LDD1609 | [32] |
| LDCM0406 | CL19 | HEK-293T | C165(1.12); C131(1.16); C133(0.91); C131(0.99) | LDD1610 | [32] |
| LDCM0407 | CL2 | HEK-293T | C165(0.94) | LDD1611 | [32] |
| LDCM0408 | CL20 | HEK-293T | C133(1.13) | LDD1612 | [32] |
| LDCM0410 | CL22 | HEK-293T | C131(1.52) | LDD1614 | [32] |
| LDCM0411 | CL23 | HEK-293T | C165(1.04); C131(1.25); C133(1.25) | LDD1615 | [32] |
| LDCM0413 | CL25 | HEK-293T | C165(0.91); C131(1.92); C133(1.92) | LDD1617 | [32] |
| LDCM0414 | CL26 | HEK-293T | C165(1.30) | LDD1618 | [32] |
| LDCM0416 | CL28 | HEK-293T | C131(1.14); C133(1.14) | LDD1620 | [32] |
| LDCM0417 | CL29 | HEK-293T | C165(0.83); C133(0.96); C131(0.94); C133(0.94) | LDD1621 | [32] |
| LDCM0419 | CL30 | HEK-293T | C131(0.87); C133(0.87) | LDD1623 | [32] |
| LDCM0420 | CL31 | HEK-293T | C165(0.95); C131(1.13); C133(0.87); C131(1.23) | LDD1624 | [32] |
| LDCM0421 | CL32 | HEK-293T | C133(1.05) | LDD1625 | [32] |
| LDCM0423 | CL34 | HEK-293T | C131(1.32) | LDD1627 | [32] |
| LDCM0424 | CL35 | HEK-293T | C165(1.21); C131(1.18); C133(1.18) | LDD1628 | [32] |
| LDCM0426 | CL37 | HEK-293T | C165(0.93); C131(1.07); C133(1.07) | LDD1630 | [32] |
| LDCM0429 | CL4 | HEK-293T | C131(1.14); C133(1.14) | LDD1633 | [32] |
| LDCM0430 | CL40 | HEK-293T | C131(0.89); C133(0.89) | LDD1634 | [32] |
| LDCM0431 | CL41 | HEK-293T | C165(1.10); C133(1.37); C131(1.17); C133(1.17) | LDD1635 | [32] |
| LDCM0432 | CL42 | HEK-293T | C131(0.78); C133(0.78) | LDD1636 | [32] |
| LDCM0433 | CL43 | HEK-293T | C165(0.79); C131(0.89); C133(0.93); C131(1.39) | LDD1637 | [32] |
| LDCM0434 | CL44 | HEK-293T | C133(0.99) | LDD1638 | [32] |
| LDCM0436 | CL46 | HEK-293T | C131(1.63) | LDD1640 | [32] |
| LDCM0437 | CL47 | HEK-293T | C165(1.18); C131(1.30); C133(1.30) | LDD1641 | [32] |
| LDCM0439 | CL49 | HEK-293T | C165(1.18); C131(1.01); C133(1.01) | LDD1643 | [32] |
| LDCM0440 | CL5 | HEK-293T | C165(1.05); C133(1.00); C131(1.03); C133(1.03) | LDD1644 | [32] |
| LDCM0441 | CL50 | HEK-293T | C165(1.03) | LDD1645 | [32] |
| LDCM0443 | CL52 | HEK-293T | C131(0.89); C133(0.89) | LDD1646 | [32] |
| LDCM0444 | CL53 | HEK-293T | C165(1.31); C133(1.58); C131(1.57); C133(1.57) | LDD1647 | [32] |
| LDCM0445 | CL54 | HEK-293T | C131(1.71); C133(1.71) | LDD1648 | [32] |
| LDCM0446 | CL55 | HEK-293T | C165(0.95); C131(1.09); C133(0.98); C131(1.02) | LDD1649 | [32] |
| LDCM0447 | CL56 | HEK-293T | C133(0.95) | LDD1650 | [32] |
| LDCM0449 | CL58 | HEK-293T | C131(1.40) | LDD1652 | [32] |
| LDCM0450 | CL59 | HEK-293T | C165(1.25); C131(1.19); C133(1.19) | LDD1653 | [32] |
| LDCM0451 | CL6 | HEK-293T | C131(1.29); C133(1.29) | LDD1654 | [32] |
| LDCM0453 | CL61 | HEK-293T | C165(0.60); C131(0.99); C133(0.99) | LDD1656 | [32] |
| LDCM0454 | CL62 | HEK-293T | C165(0.98) | LDD1657 | [32] |
| LDCM0456 | CL64 | HEK-293T | C131(1.44); C133(1.44) | LDD1659 | [32] |
| LDCM0457 | CL65 | HEK-293T | C165(1.18); C133(0.96); C131(1.02); C133(1.02) | LDD1660 | [32] |
| LDCM0458 | CL66 | HEK-293T | C131(0.90); C133(0.90) | LDD1661 | [32] |
| LDCM0459 | CL67 | HEK-293T | C165(0.81); C131(1.08); C133(1.01); C131(1.11) | LDD1662 | [32] |
| LDCM0460 | CL68 | HEK-293T | C133(0.94) | LDD1663 | [32] |
| LDCM0462 | CL7 | HEK-293T | C165(0.79); C131(1.10); C133(1.01); C131(1.32) | LDD1665 | [32] |
| LDCM0463 | CL70 | HEK-293T | C131(1.34) | LDD1666 | [32] |
| LDCM0464 | CL71 | HEK-293T | C165(1.17); C131(1.30); C133(1.30) | LDD1667 | [32] |
| LDCM0466 | CL73 | HEK-293T | C165(1.00); C131(1.04); C133(1.04) | LDD1669 | [32] |
| LDCM0467 | CL74 | HEK-293T | C165(1.17) | LDD1670 | [32] |
| LDCM0469 | CL76 | HEK-293T | C131(0.92); C133(0.92) | LDD1672 | [32] |
| LDCM0470 | CL77 | HEK-293T | C165(1.31); C133(1.25); C131(1.45); C133(1.45) | LDD1673 | [32] |
| LDCM0471 | CL78 | HEK-293T | C131(0.88); C133(0.88) | LDD1674 | [32] |
| LDCM0472 | CL79 | HEK-293T | C165(0.74); C131(1.03); C133(0.86); C131(0.97) | LDD1675 | [32] |
| LDCM0473 | CL8 | HEK-293T | C133(2.22) | LDD1676 | [32] |
| LDCM0474 | CL80 | HEK-293T | C133(0.83) | LDD1677 | [32] |
| LDCM0476 | CL82 | HEK-293T | C131(1.19) | LDD1679 | [32] |
| LDCM0477 | CL83 | HEK-293T | C165(0.94); C131(1.19); C133(1.19) | LDD1680 | [32] |
| LDCM0479 | CL85 | HEK-293T | C165(0.98); C131(0.98); C133(0.98) | LDD1682 | [32] |
| LDCM0480 | CL86 | HEK-293T | C165(0.88) | LDD1683 | [32] |
| LDCM0482 | CL88 | HEK-293T | C131(0.86); C133(0.86) | LDD1685 | [32] |
| LDCM0483 | CL89 | HEK-293T | C165(0.84); C133(0.97); C131(0.86); C133(0.86) | LDD1686 | [32] |
| LDCM0485 | CL90 | HEK-293T | C131(2.29); C133(2.29) | LDD1688 | [32] |
| LDCM0486 | CL91 | HEK-293T | C165(0.56); C131(0.86); C133(0.81); C131(0.99) | LDD1689 | [32] |
| LDCM0487 | CL92 | HEK-293T | C133(1.10) | LDD1690 | [32] |
| LDCM0489 | CL94 | HEK-293T | C131(1.58) | LDD1692 | [32] |
| LDCM0490 | CL95 | HEK-293T | C165(0.94); C131(1.93); C133(1.93) | LDD1693 | [32] |
| LDCM0492 | CL97 | HEK-293T | C165(1.08); C131(1.23); C133(1.23) | LDD1695 | [32] |
| LDCM0493 | CL98 | HEK-293T | C165(1.05) | LDD1696 | [32] |
| LDCM0185 | Compound 17 | HEK-293T | 10.31 | LDD0512 | [28] |
| LDCM0191 | Compound 21 | HEK-293T | 8.14 | LDD0508 | [28] |
| LDCM0190 | Compound 34 | HEK-293T | 9.47 | LDD0510 | [28] |
| LDCM0192 | Compound 35 | HEK-293T | 5.53 | LDD0509 | [28] |
| LDCM0193 | Compound 36 | HEK-293T | 7.52 | LDD0511 | [28] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C18(4.30) | LDD1702 | [10] |
| LDCM0175 | Ethacrynic acid | HeLa | C18(1.06); C18(1.10) | LDD2210 | [11] |
| LDCM0625 | F8 | Ramos | C18(2.10) | LDD2187 | [33] |
| LDCM0572 | Fragment10 | Ramos | C18(2.46) | LDD2189 | [33] |
| LDCM0574 | Fragment12 | Ramos | C18(3.93) | LDD2191 | [33] |
| LDCM0575 | Fragment13 | Ramos | C18(0.92) | LDD2192 | [33] |
| LDCM0576 | Fragment14 | Ramos | C18(3.39) | LDD2193 | [33] |
| LDCM0580 | Fragment21 | Ramos | C18(0.95) | LDD2195 | [33] |
| LDCM0582 | Fragment23 | Ramos | C18(0.77) | LDD2196 | [33] |
| LDCM0578 | Fragment27 | Ramos | C18(1.00) | LDD2197 | [33] |
| LDCM0586 | Fragment28 | Ramos | C18(0.99) | LDD2198 | [33] |
| LDCM0588 | Fragment30 | Ramos | C18(0.86) | LDD2199 | [33] |
| LDCM0589 | Fragment31 | Ramos | C18(0.86) | LDD2200 | [33] |
| LDCM0590 | Fragment32 | Ramos | C18(3.27) | LDD2201 | [33] |
| LDCM0468 | Fragment33 | Ramos | C18(1.04) | LDD2202 | [33] |
| LDCM0596 | Fragment38 | Ramos | C18(1.27) | LDD2203 | [33] |
| LDCM0566 | Fragment4 | Ramos | C18(2.71) | LDD2184 | [33] |
| LDCM0427 | Fragment51 | HEK-293T | C165(0.92) | LDD1631 | [32] |
| LDCM0610 | Fragment52 | Ramos | C18(1.12) | LDD2204 | [33] |
| LDCM0614 | Fragment56 | Ramos | C18(0.75) | LDD2205 | [33] |
| LDCM0569 | Fragment7 | Ramos | C18(2.56) | LDD2186 | [33] |
| LDCM0571 | Fragment9 | Ramos | C18(7.39) | LDD2188 | [33] |
| LDCM0015 | HNE | MDA-MB-231 | C18(1.10) | LDD0346 | [33] |
| LDCM0107 | IAA | HeLa | H3(0.00); H153(0.00); H40(0.00); H26(0.00) | LDD0221 | [24] |
| LDCM0022 | KB02 | HEK-293T | C131(0.82); C133(0.82) | LDD1492 | [32] |
| LDCM0023 | KB03 | Jurkat | C18(192.49) | LDD0209 | [12] |
| LDCM0024 | KB05 | IGR37 | C18(1.29) | LDD3314 | [34] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C18(1.03) | LDD2102 | [10] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C18(0.62) | LDD2121 | [10] |
| LDCM0109 | NEM | HeLa | H3(0.00); H153(0.00); H40(0.00); H26(0.00) | LDD0223 | [24] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C18(0.97) | LDD2089 | [10] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C18(1.22) | LDD2090 | [10] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C18(0.95) | LDD2092 | [10] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C18(1.25) | LDD2093 | [10] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C18(1.10) | LDD2094 | [10] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C18(0.75) | LDD2097 | [10] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C18(0.94) | LDD2098 | [10] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C18(1.14) | LDD2099 | [10] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C18(0.86) | LDD2100 | [10] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C18(1.05) | LDD2101 | [10] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C18(0.89) | LDD2104 | [10] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C18(1.08) | LDD2105 | [10] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C18(0.66) | LDD2106 | [10] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C18(1.08) | LDD2107 | [10] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C18(0.64) | LDD2108 | [10] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C18(0.97) | LDD2109 | [10] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C18(1.21) | LDD2110 | [10] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C18(1.18) | LDD2111 | [10] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C18(0.58) | LDD2115 | [10] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C18(0.37) | LDD2118 | [10] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C18(1.86) | LDD2119 | [10] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C18(0.78) | LDD2120 | [10] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C18(0.93) | LDD2123 | [10] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C18(1.41) | LDD2124 | [10] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C18(1.05) | LDD2125 | [10] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C18(3.13) | LDD2126 | [10] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C18(1.11) | LDD2127 | [10] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C18(1.07) | LDD2128 | [10] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C18(1.15) | LDD2129 | [10] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C18(0.80) | LDD2133 | [10] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C18(0.53) | LDD2134 | [10] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C18(1.12) | LDD2135 | [10] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C18(1.76) | LDD2136 | [10] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C18(0.87) | LDD2137 | [10] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C18(2.08) | LDD1700 | [10] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C18(1.02) | LDD2140 | [10] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C18(0.42) | LDD2141 | [10] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C18(0.93) | LDD2143 | [10] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C18(2.09) | LDD2144 | [10] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C18(1.01) | LDD2145 | [10] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C18(1.06) | LDD2146 | [10] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C18(2.67) | LDD2147 | [10] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C18(0.49) | LDD2148 | [10] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C18(3.24) | LDD2149 | [10] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C18(0.59) | LDD2150 | [10] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C18(3.21) | LDD2151 | [10] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C18(1.53) | LDD2153 | [10] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C18(1.35); C18(1.01); C18(0.96) | LDD2206 | [35] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C18(0.92); C18(0.74); C18(0.47); C18(0.42) | LDD2207 | [35] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Cyclic AMP-responsive element-binding protein 3-like protein 1 (CREB3L1) | BZIP family | Q96BA8 | |||
| Transcription factor A, mitochondrial (TFAM) | . | Q00059 | |||
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| B-cell antigen receptor complex-associated protein alpha chain (CD79A) | . | P11912 | |||
Other
References