Details of the Target
General Information of Target
| Target ID | LDTP12959 | |||||
|---|---|---|---|---|---|---|
| Target Name | NADH dehydrogenase 1 alpha subcomplex subunit 13 (NDUFA13) | |||||
| Gene Name | NDUFA13 | |||||
| Gene ID | 51079 | |||||
| Synonyms |
GRIM19; NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 13; Cell death regulatory protein GRIM-19; Complex I-B16.6; CI-B16.6; Gene associated with retinoic and interferon-induced mortality 19 protein; GRIM-19; Gene associated with retinoic and IFN-induced mortality 19 protein; NADH-ubiquinone oxidoreductase B16.6 subunit
|
|||||
| 3D Structure | ||||||
| Sequence |
MAGPELLLDSNIRLWVVLPIVIITFFVGMIRHYVSILLQSDKKLTQEQVSDSQVLIRSRV
LRENGKYIPKQSFLTRKYYFNNPEDGFFKKTKRKVVPPSPMTDPTMLTDMMKGNVTNVLP MILIGGWINMTFSGFVTTKVPFPLTLRFKPMLQQGIELLTLDASWVSSASWYFLNVFGLR SIYSLILGQDNAADQSRMMQEQMTGAAMAMPADTNKAFKTEWEALELTDHQWALDDVEEE LMAKDLHFEGMFKKELQTSIF |
|||||
| Target Type |
Clinical trial
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Complex I NDUFA13 subunit family
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Accessory subunit of the mitochondrial membrane respiratory chain NADH dehydrogenase (Complex I), that is believed not to be involved in catalysis. Complex I functions in the transfer of electrons from NADH to the respiratory chain. The immediate electron acceptor for the enzyme is believed to be ubiquinone. Involved in the interferon/all-trans-retinoic acid (IFN/RA) induced cell death. This apoptotic activity is inhibited by interaction with viral IRF1. Prevents the transactivation of STAT3 target genes. May play a role in CARD15-mediated innate mucosal responses and serve to regulate intestinal epithelial cell responses to microbes.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
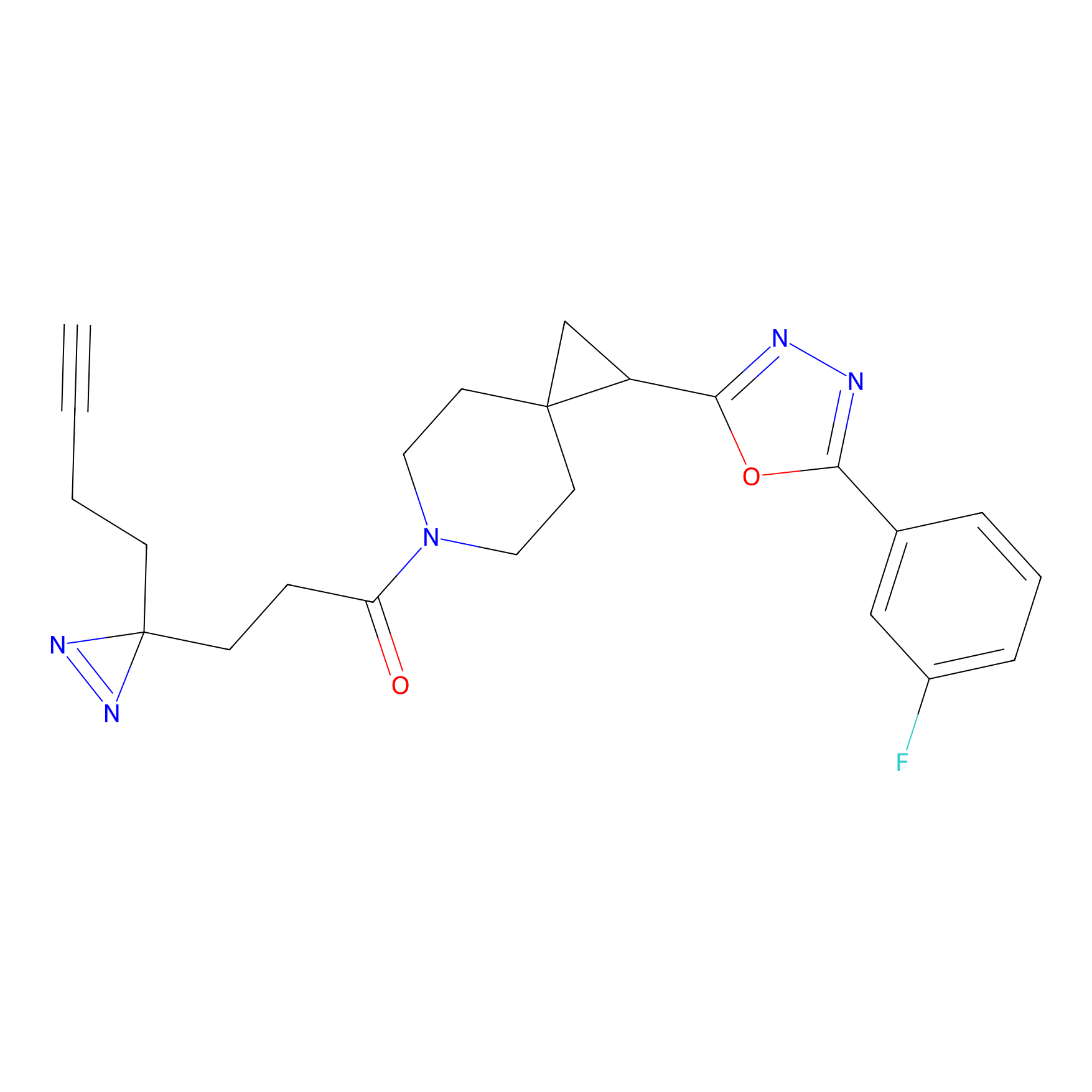
STPyne Probe Info |
 |
K22(1.45) | LDD0277 | [1] | |
|
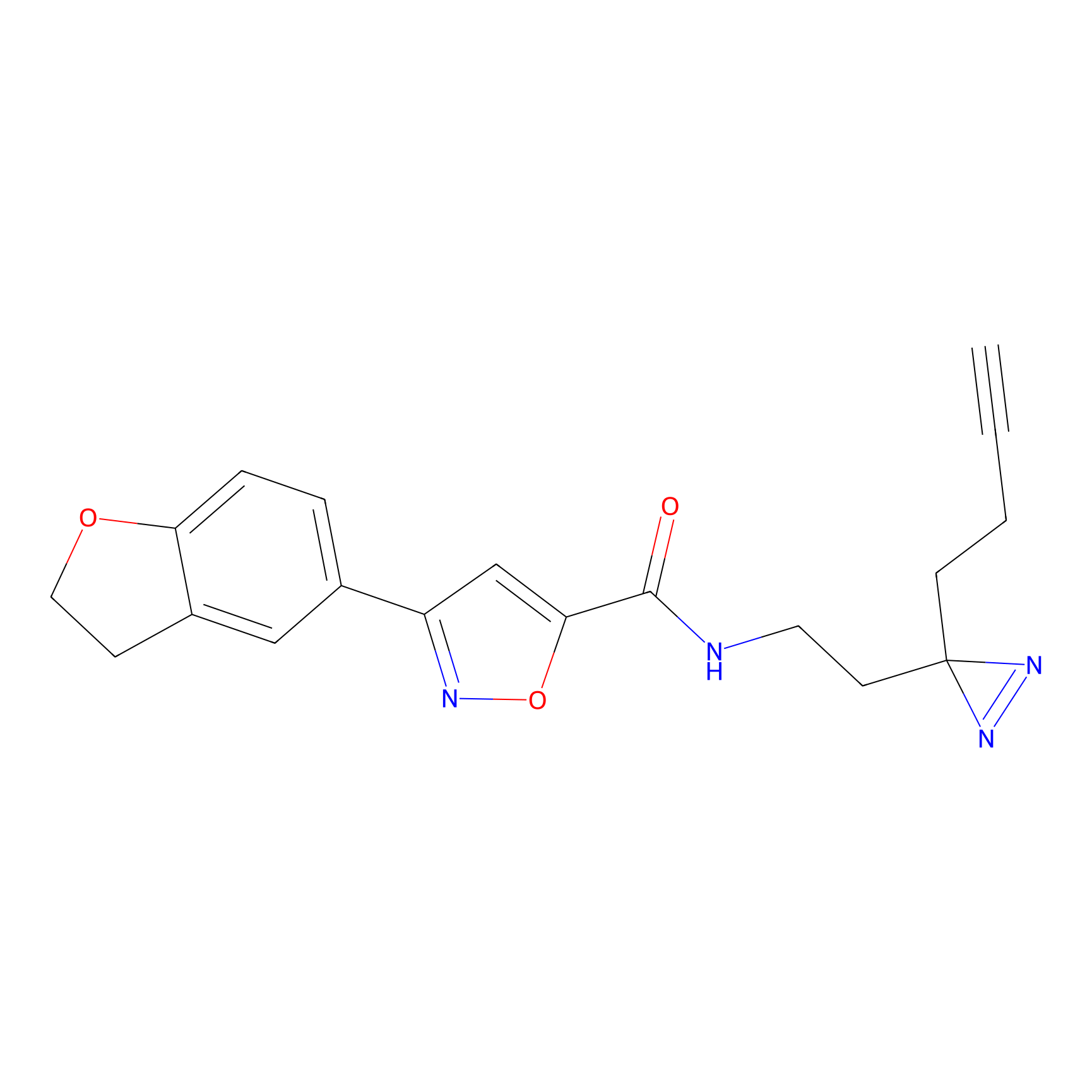
Acrolein Probe Info |
 |
N.A. | LDD0223 | [2] | |
|
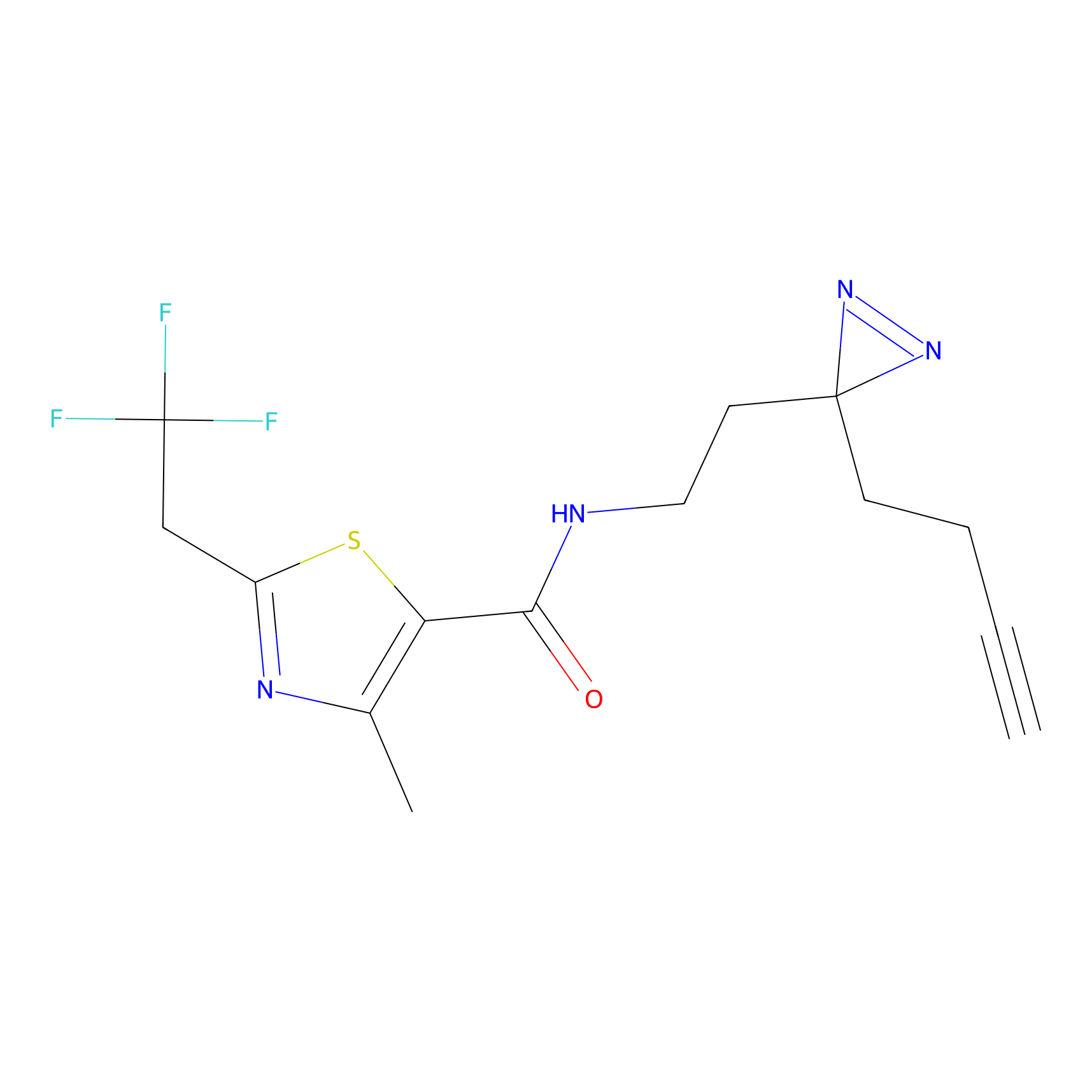
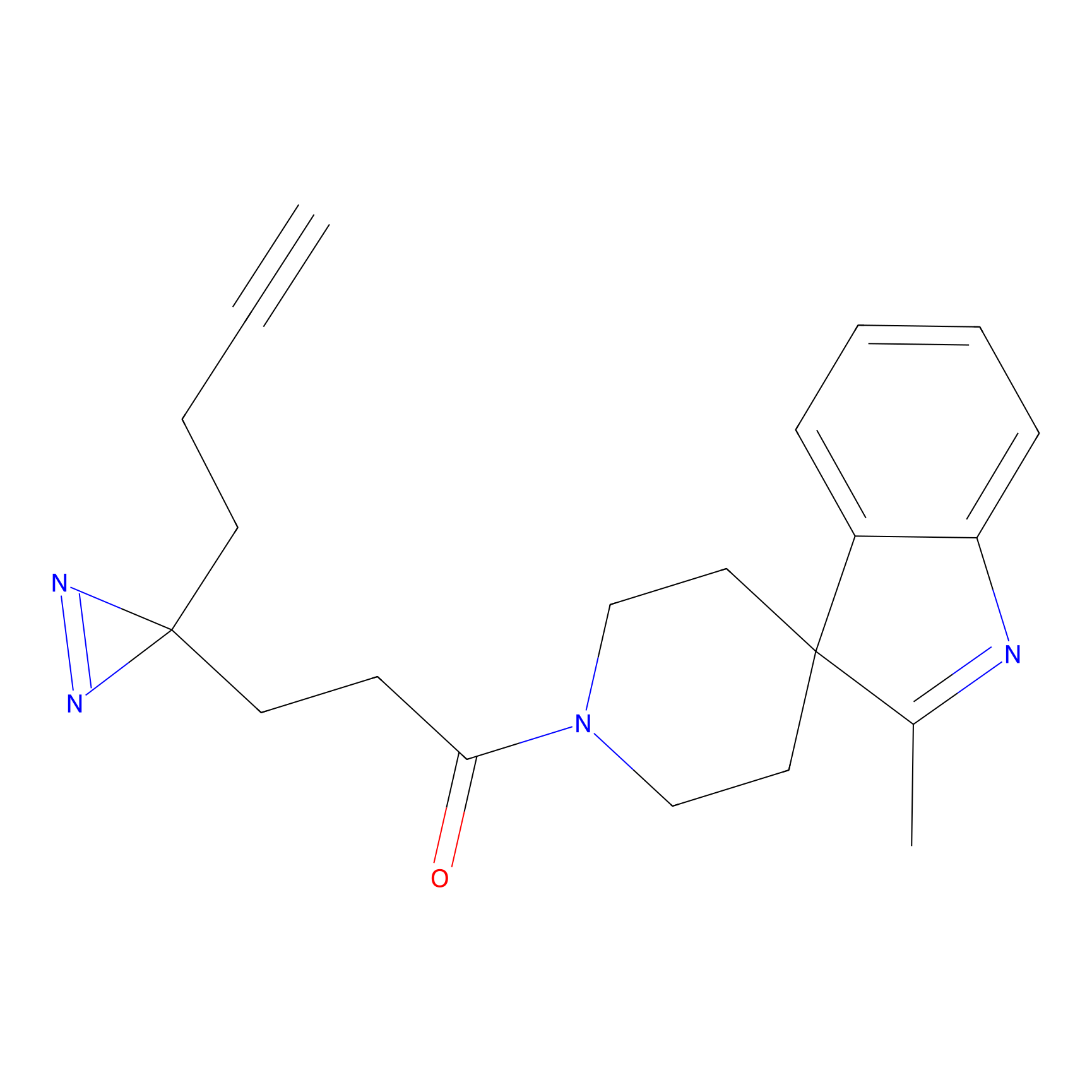
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [3] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Serine protease HTRA2, mitochondrial (HTRA2) | Peptidase S1C family | O43464 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nucleotide-binding oligomerization domain-containing protein 2 (NOD2) | . | Q9HC29 | |||
References