Details of the Target
General Information of Target
Probe(s) Labeling This Target
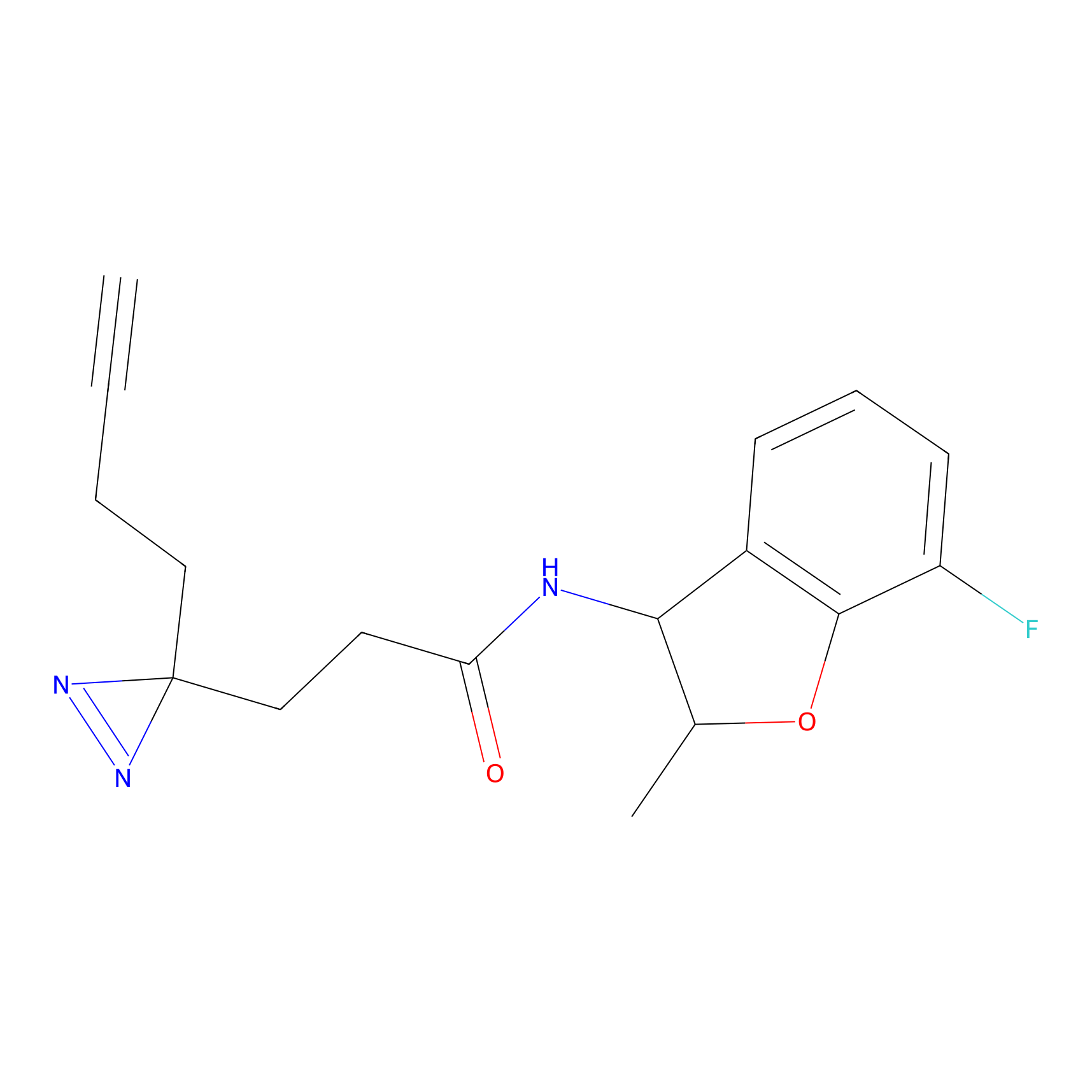
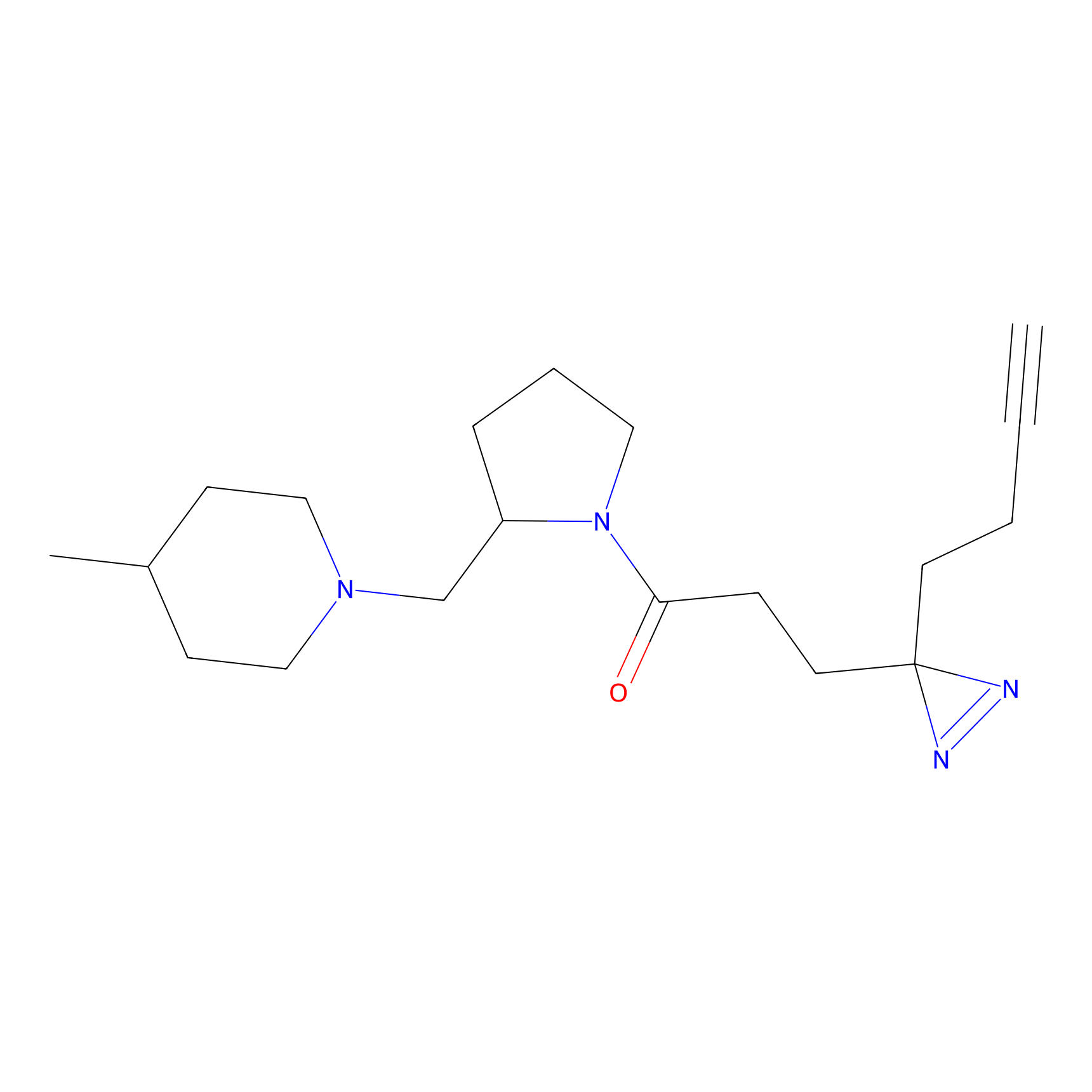
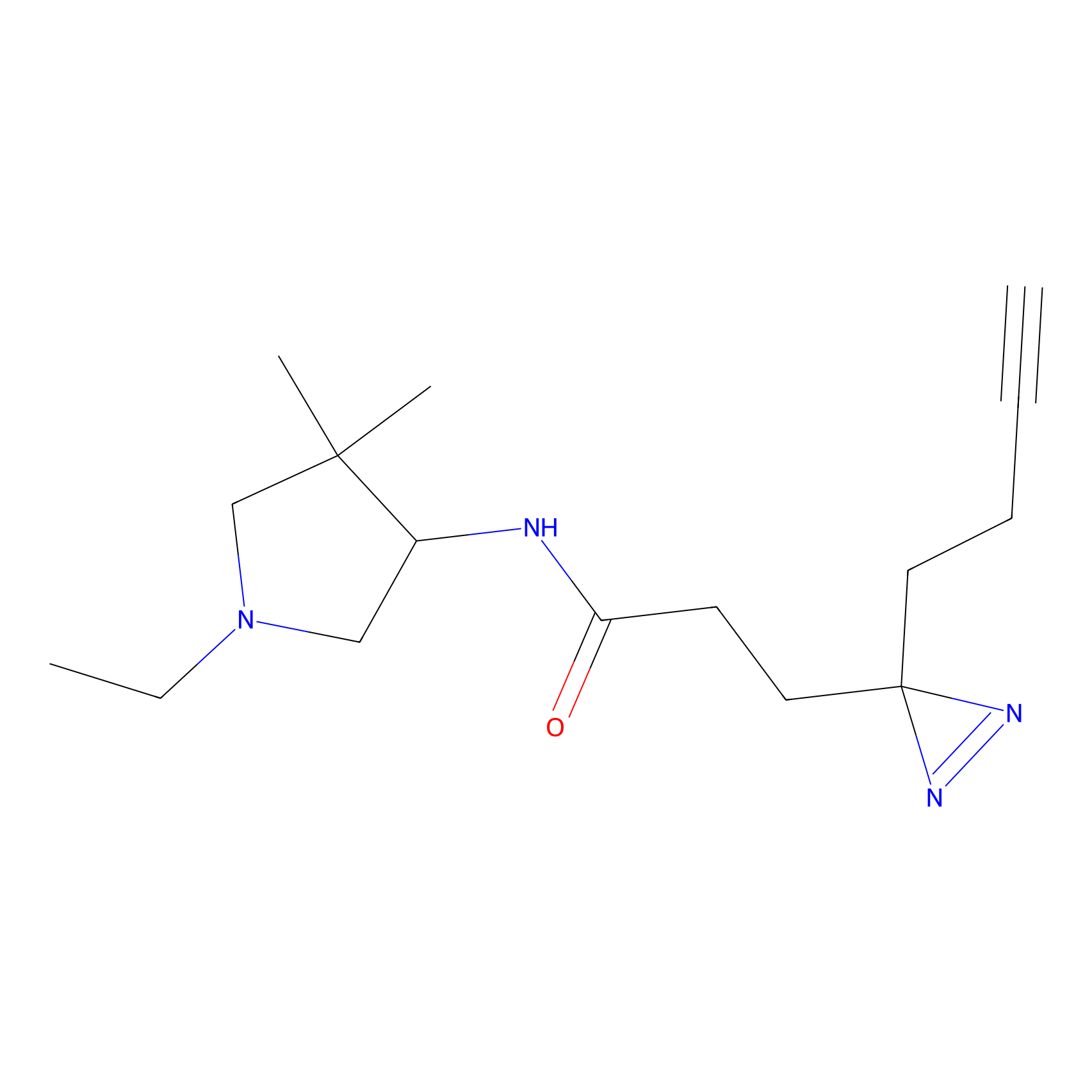
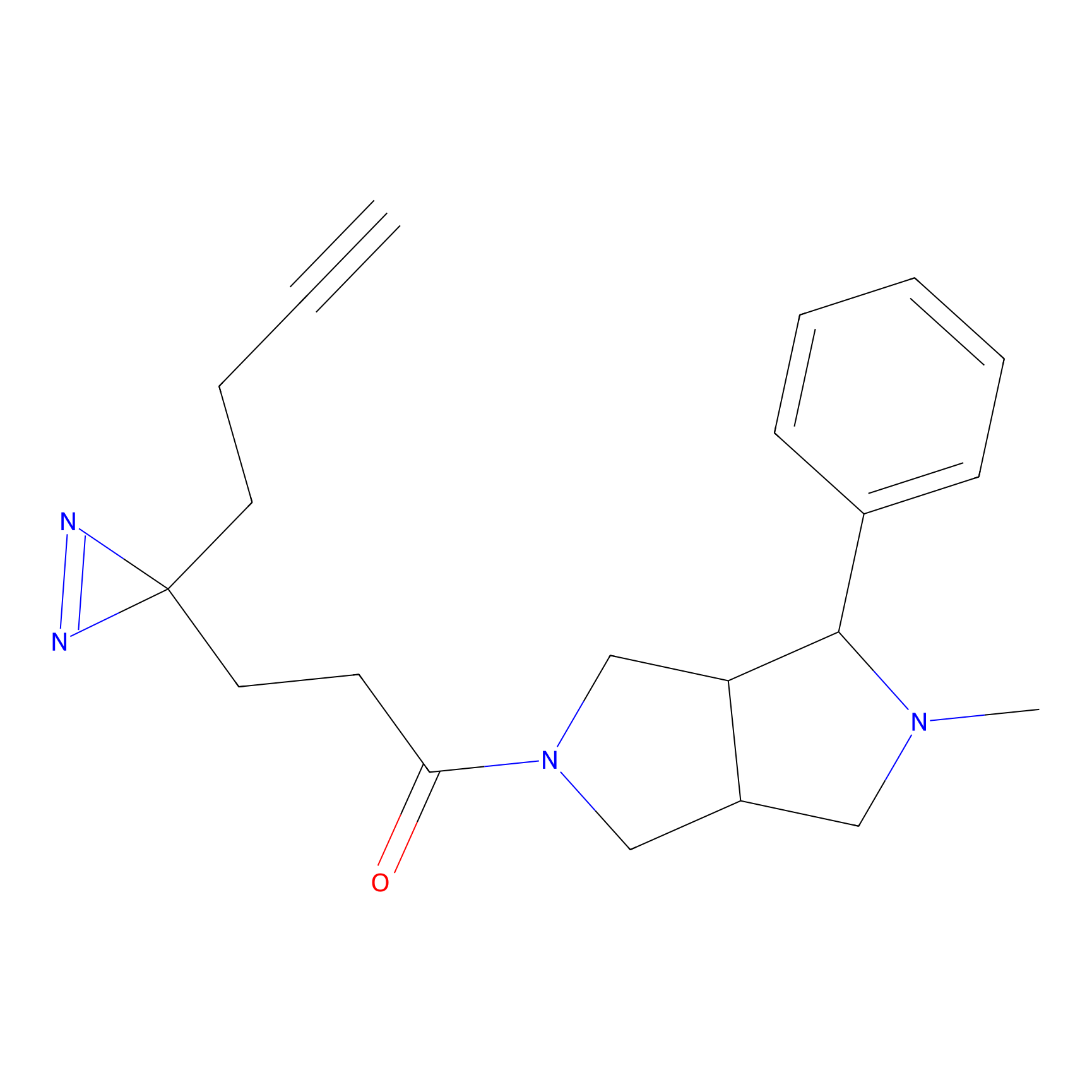
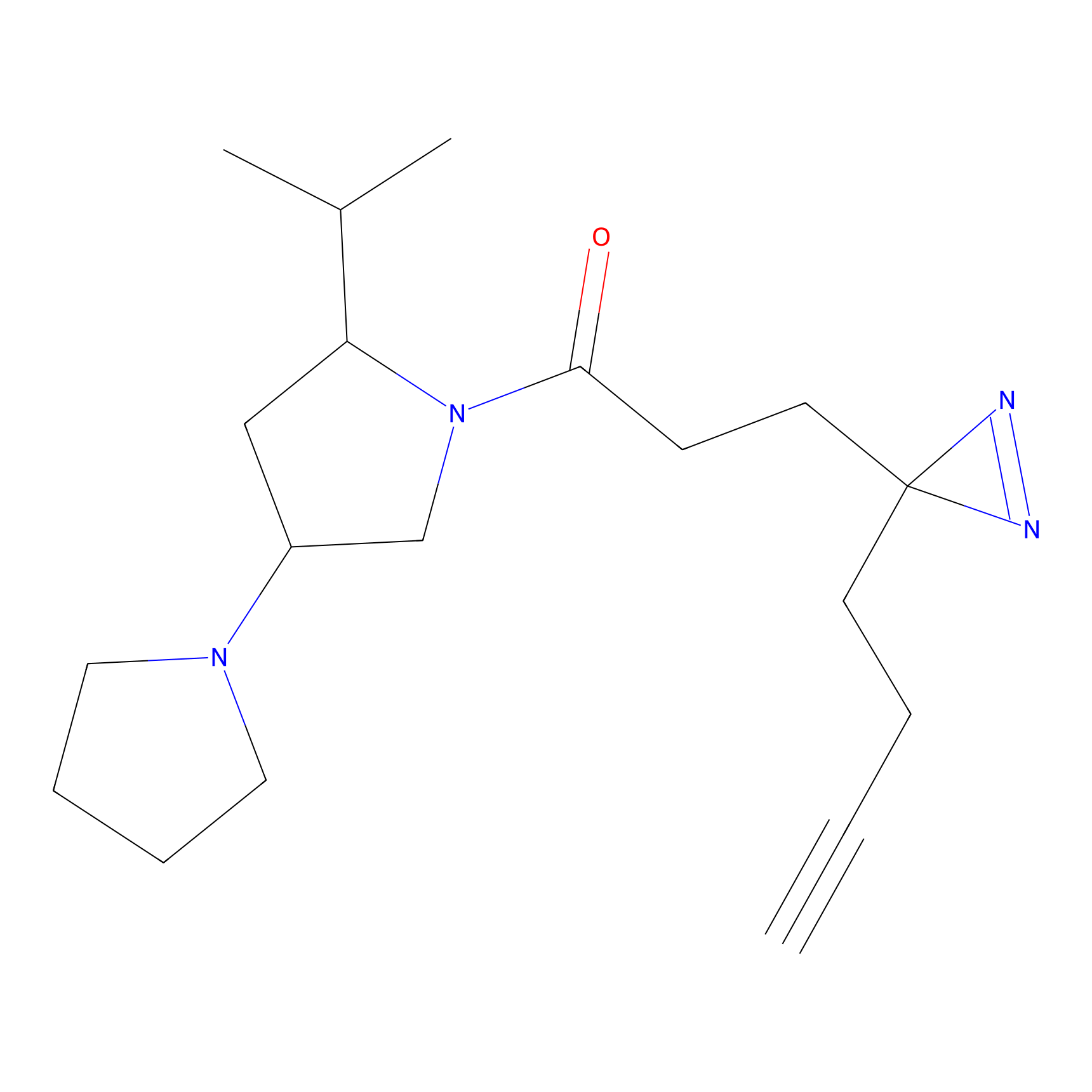
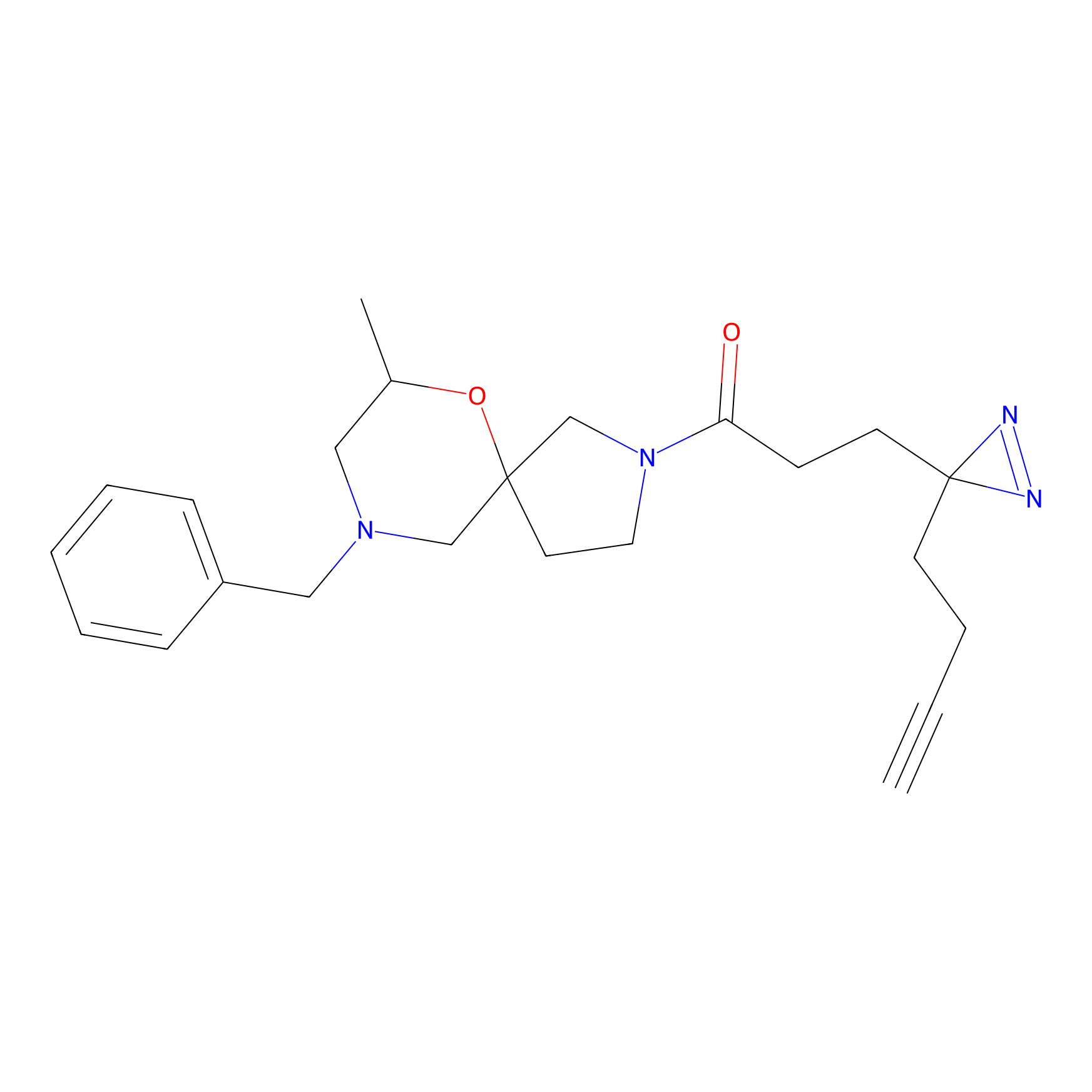
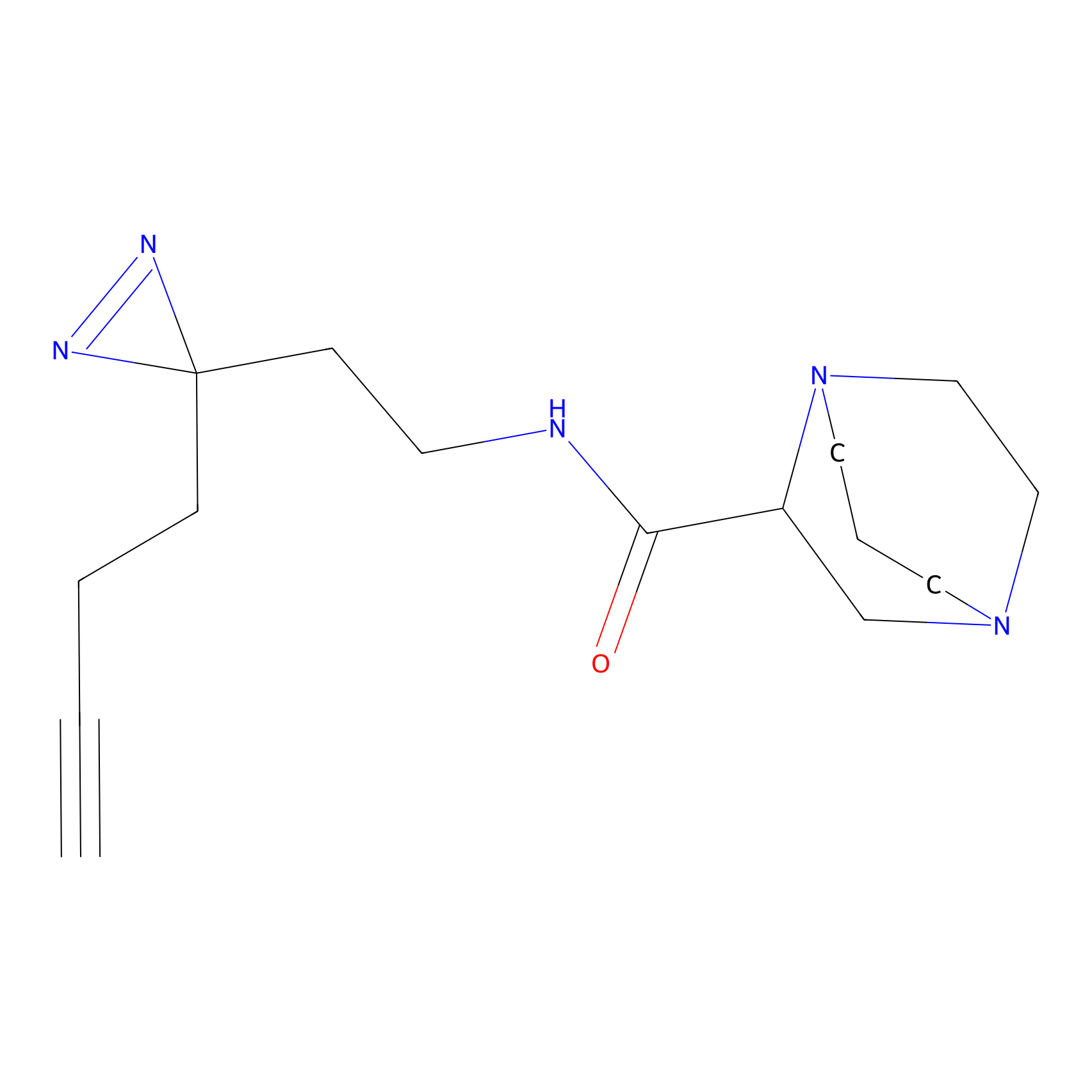
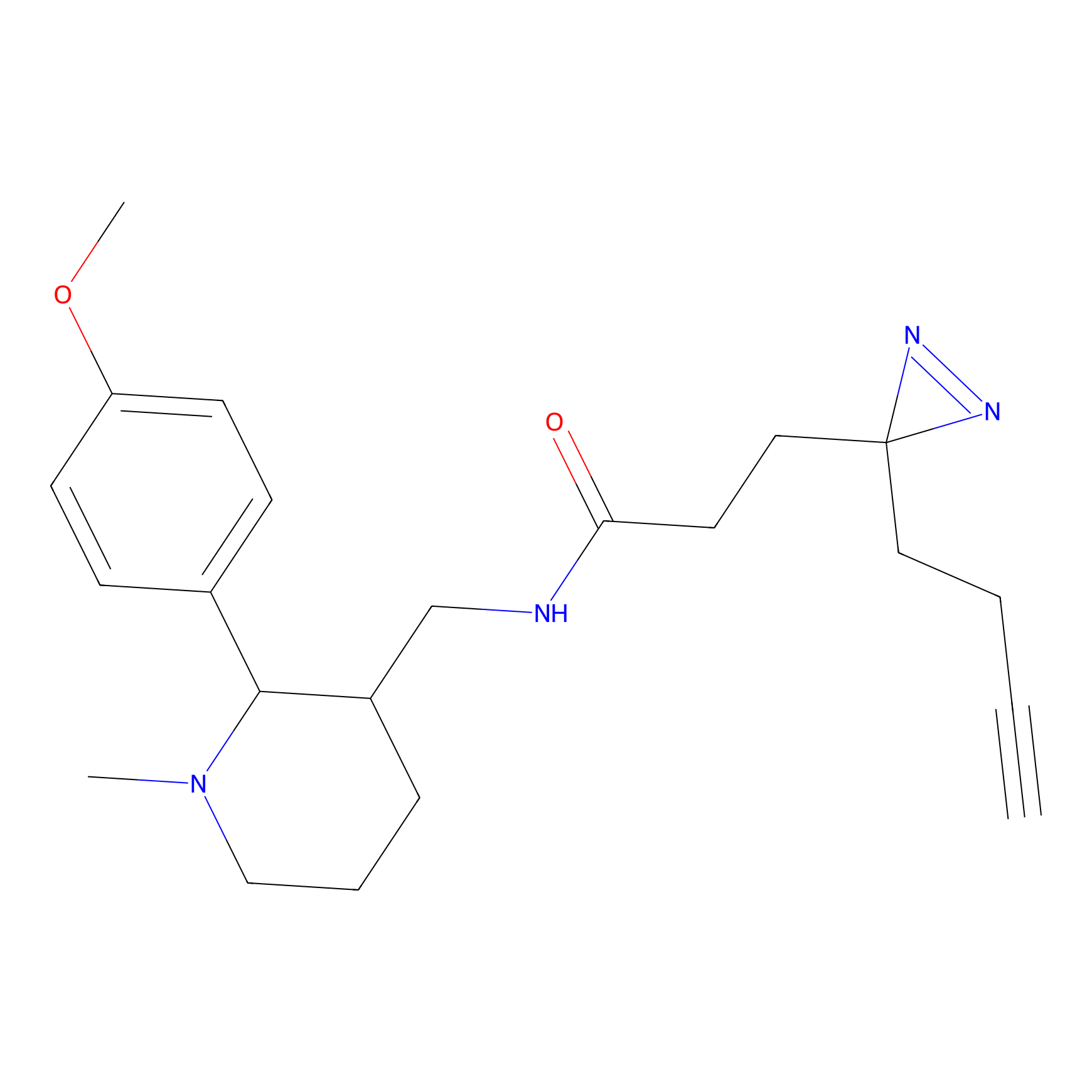
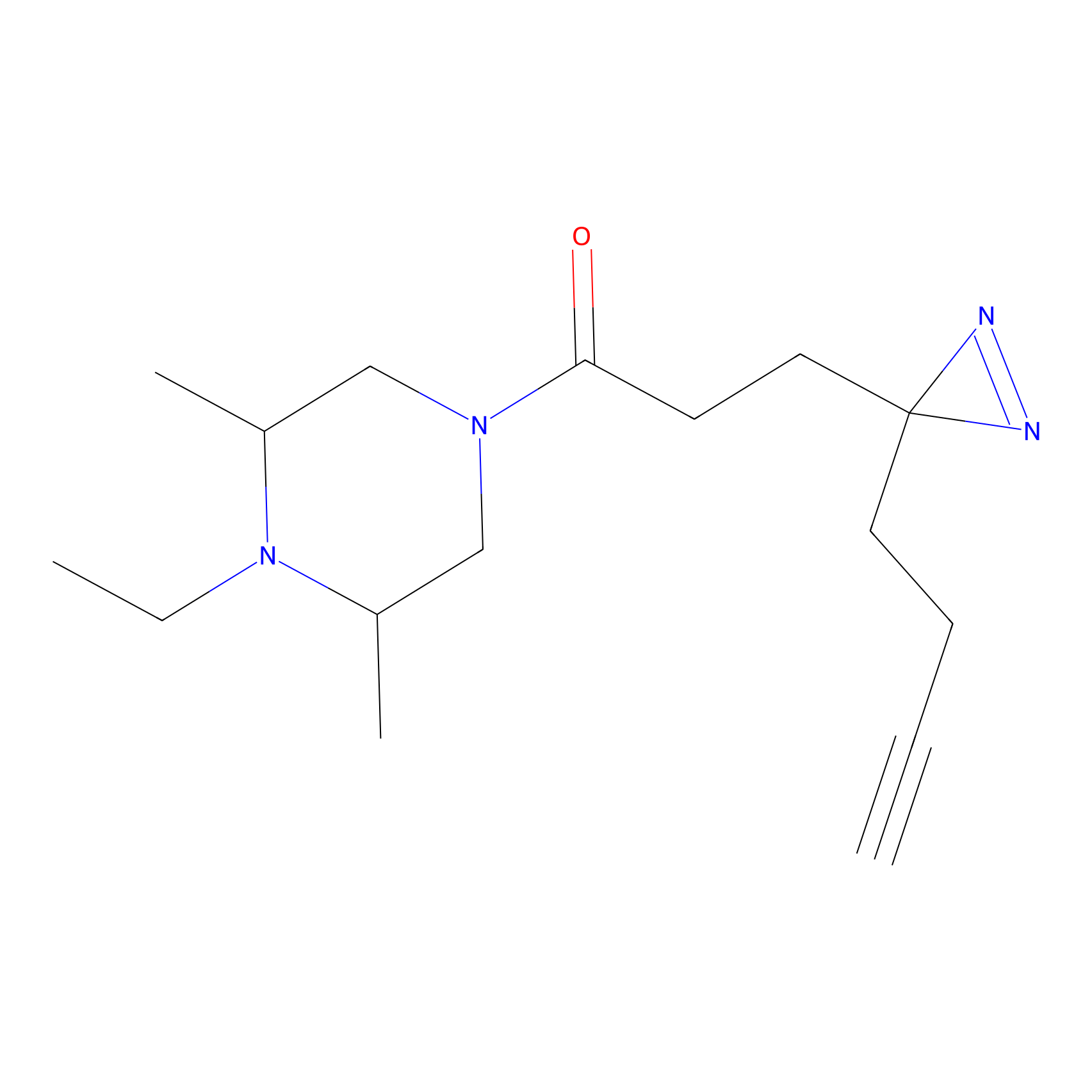
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
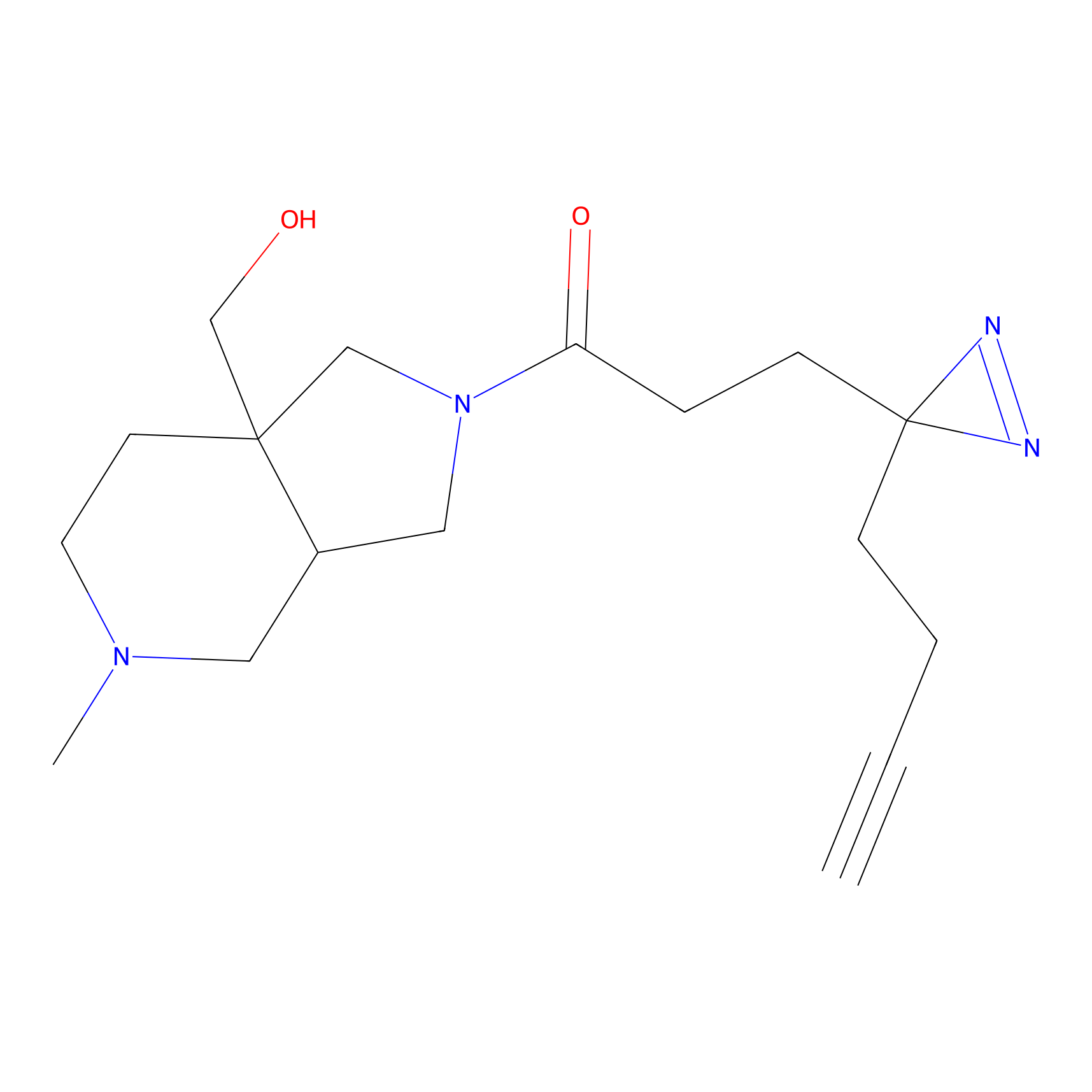
Alkylaryl probe 1 Probe Info |
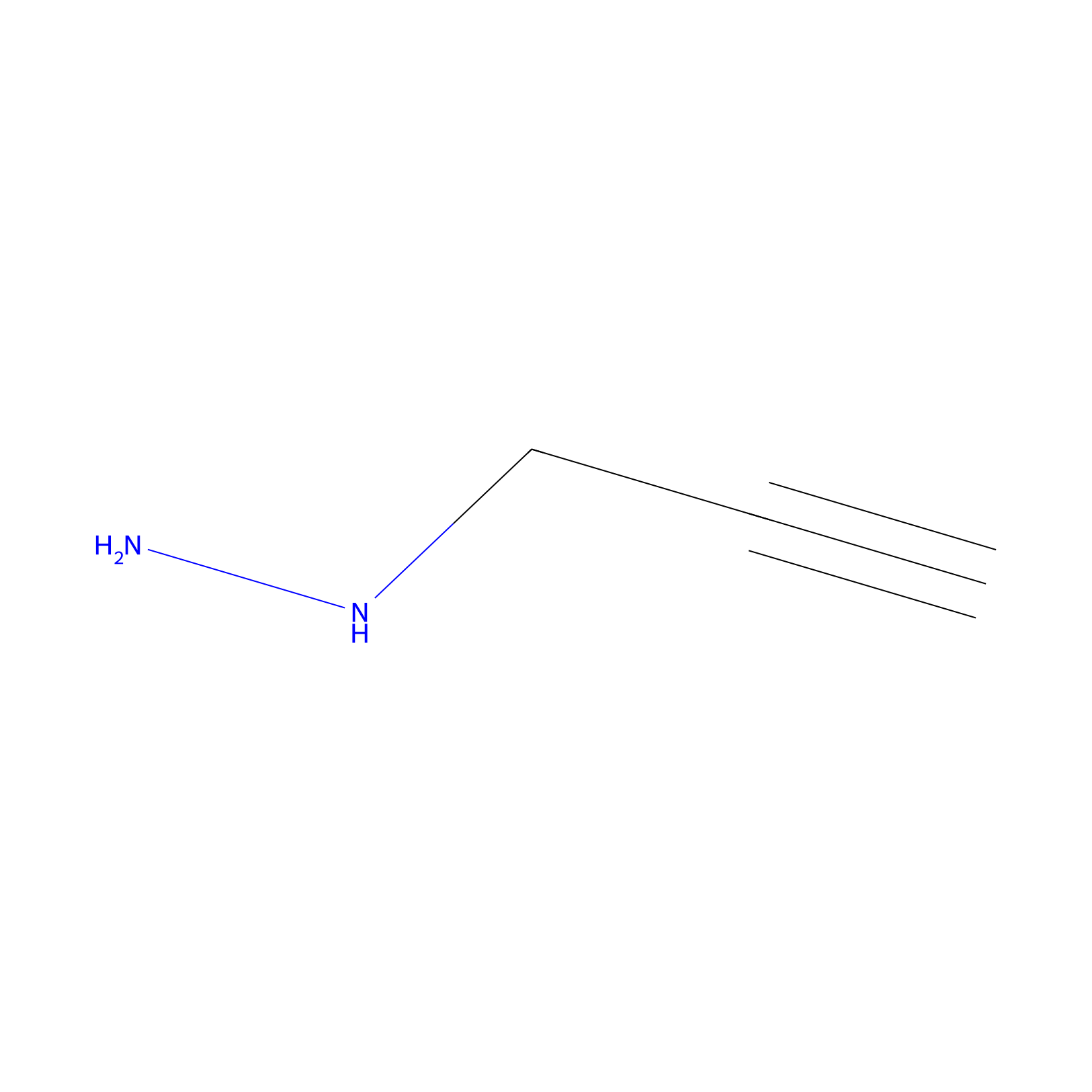 |
9.00 | LDD0387 | [2] | |
|
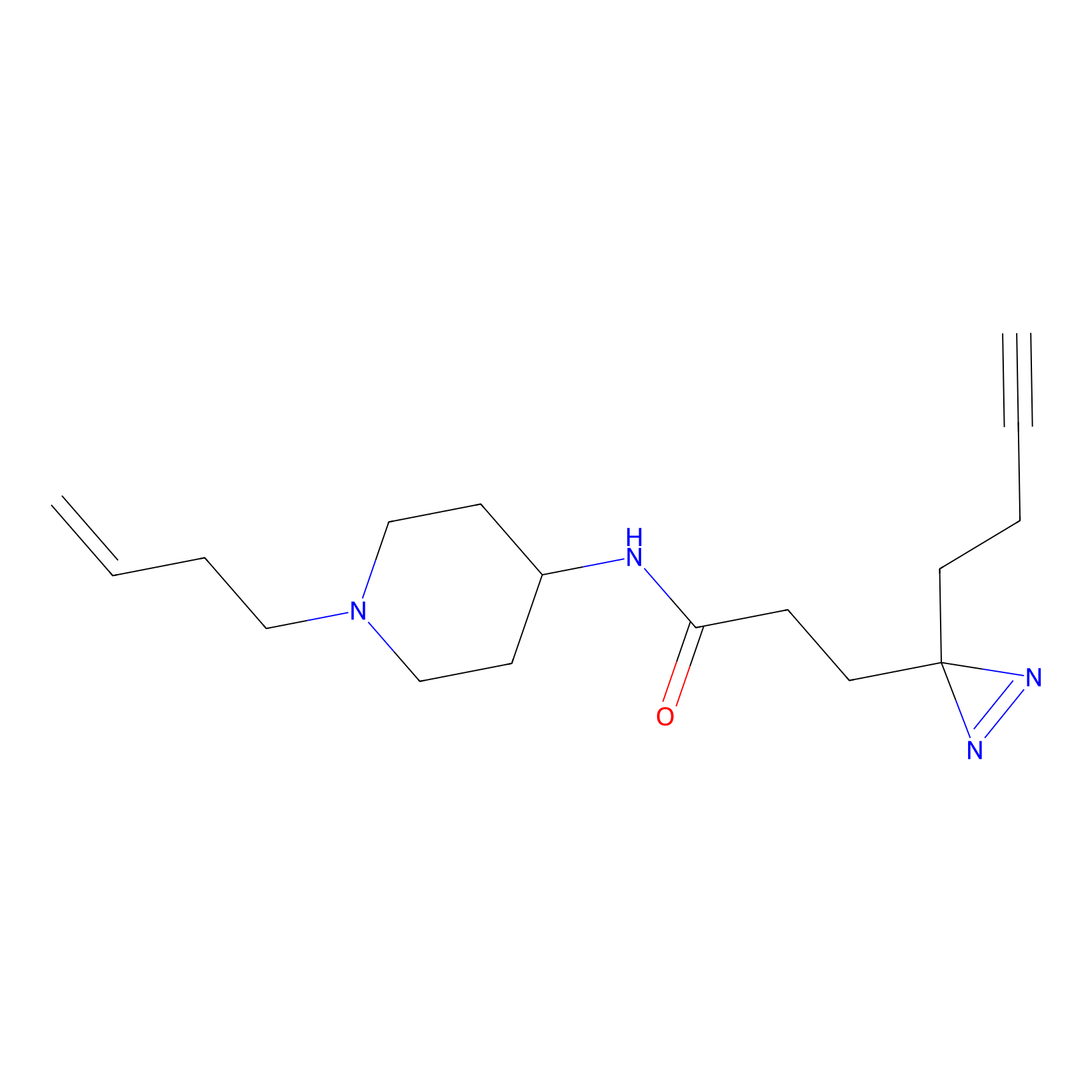
Alkylaryl probe 2 Probe Info |
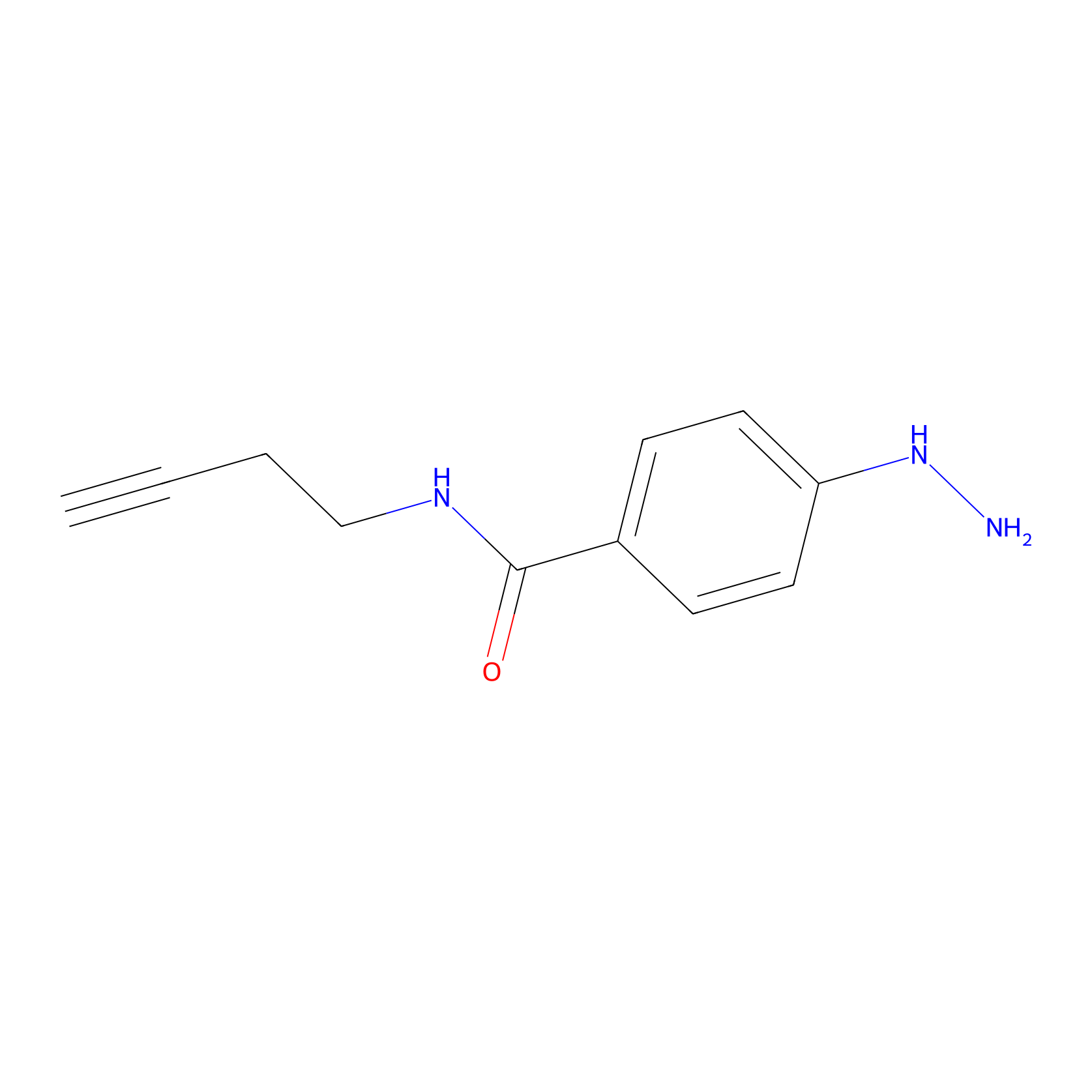 |
20.00 | LDD0391 | [2] | |
|
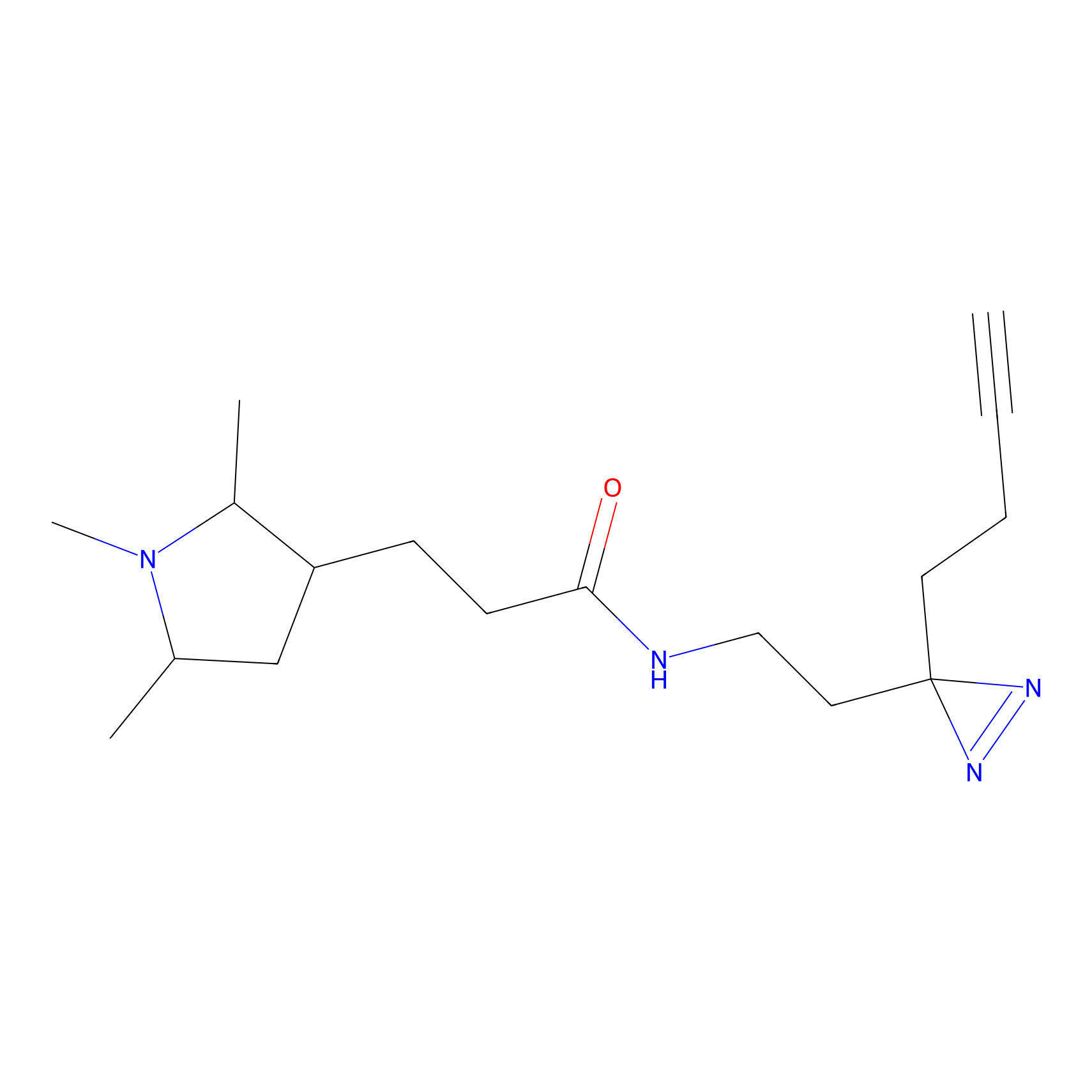
FBPP2 Probe Info |
 |
5.97 | LDD0318 | [3] | |
|
TH216 Probe Info |
 |
Y407(17.27) | LDD0259 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0222 | [6] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [7] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [7] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [8] | |
|
HHS-482 Probe Info |
 |
Y458(0.38) | LDD2239 | [9] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Drug(s) Related To This Target
Approved
Phase 3
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Avalglucosidase Alfa | Recombinant protein | D34SXJ | |||
| Deoxynojirimycin | Small molecular drug | D0S5LT | |||
| Bmn-701 | . | D05JEG | |||
Phase 2
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Salacinol | Small molecular drug | D07TSW | |||
Phase 1
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Gz402666 | . | D0Q1UB | |||
Preclinical
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| 5-(4-(4-acetylphenyl)Piperazin-1-ylsulfonyl)-6-chloroindolin-2-one | Small molecular drug | DW68KF | |||
Investigative
References