Details of the Target
General Information of Target
| Target ID | LDTP01542 | |||||
|---|---|---|---|---|---|---|
| Target Name | Golgi SNAP receptor complex member 1 (GOSR1) | |||||
| Gene Name | GOSR1 | |||||
| Gene ID | 9527 | |||||
| Synonyms |
GS28; Golgi SNAP receptor complex member 1; 28 kDa Golgi SNARE protein; 28 kDa cis-Golgi SNARE p28; GOS-28 |
|||||
| 3D Structure | ||||||
| Sequence |
MAAGTSSYWEDLRKQARQLENELDLKLVSFSKLCTSYSHSSTRDGRRDRYSSDTTPLLNG
SSQDRMFETMAIEIEQLLARLTGVNDKMAEYTNSAGVPSLNAALMHTLQRHRDILQDYTH EFHKTKANFMAIRERENLMGSVRKDIESYKSGSGVNNRRTELFLKEHDHLRNSDRLIEET ISIAMATKENMTSQRGMLKSIHSKMNTLANRFPAVNSLIQRINLRKRRDSLILGGVIGIC TILLLLYAFH |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
GOSR1 family
|
|||||
| Subcellular location |
Golgi apparatus membrane
|
|||||
| Function |
Involved in transport from the ER to the Golgi apparatus as well as in intra-Golgi transport. It belongs to a super-family of proteins called t-SNAREs or soluble NSF (N-ethylmaleimide-sensitive factor) attachment protein receptor. May play a protective role against hydrogen peroxide induced cytotoxicity under glutathione depleted conditions in neuronal cells by regulating the intracellular ROS levels via inhibition of p38 MAPK (MAPK11, MAPK12, MAPK13 and MAPK14). Participates in docking and fusion stage of ER to cis-Golgi transport. Plays an important physiological role in VLDL-transport vesicle-Golgi fusion and thus in VLDL delivery to the hepatic cis-Golgi.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DBIA Probe Info |
 |
C34(1.54) | LDD3330 | [1] | |
|
BTD Probe Info |
 |
C34(0.66) | LDD2097 | [2] | |
|
ATP probe Probe Info |
 |
K144(0.00); K124(0.00); K126(0.00) | LDD0199 | [3] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [4] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [4] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [4] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [5] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [5] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [6] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [7] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [5] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [8] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [8] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [8] | |
|
AOyne Probe Info |
 |
14.60 | LDD0443 | [9] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [6] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C34(0.41) | LDD2142 | [2] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C34(0.66) | LDD2112 | [2] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C34(1.17) | LDD2152 | [2] |
| LDCM0214 | AC1 | HEK-293T | C34(0.92) | LDD1507 | [14] |
| LDCM0215 | AC10 | HEK-293T | C34(0.87) | LDD1508 | [14] |
| LDCM0226 | AC11 | HEK-293T | C34(0.93) | LDD1509 | [14] |
| LDCM0237 | AC12 | HEK-293T | C34(1.03) | LDD1510 | [14] |
| LDCM0270 | AC15 | HEK-293T | C34(1.01) | LDD1513 | [14] |
| LDCM0276 | AC17 | HEK-293T | C34(1.12) | LDD1515 | [14] |
| LDCM0277 | AC18 | HEK-293T | C34(0.93) | LDD1516 | [14] |
| LDCM0278 | AC19 | HEK-293T | C34(0.83) | LDD1517 | [14] |
| LDCM0279 | AC2 | HEK-293T | C34(1.09) | LDD1518 | [14] |
| LDCM0280 | AC20 | HEK-293T | C34(1.07) | LDD1519 | [14] |
| LDCM0281 | AC21 | HEK-293T | C34(0.97) | LDD1520 | [14] |
| LDCM0283 | AC23 | HEK-293T | C34(1.14) | LDD1522 | [14] |
| LDCM0284 | AC24 | HEK-293T | C34(1.18) | LDD1523 | [14] |
| LDCM0285 | AC25 | HEK-293T | C34(0.99) | LDD1524 | [14] |
| LDCM0286 | AC26 | HEK-293T | C34(1.08) | LDD1525 | [14] |
| LDCM0287 | AC27 | HEK-293T | C34(0.95) | LDD1526 | [14] |
| LDCM0288 | AC28 | HEK-293T | C34(1.05) | LDD1527 | [14] |
| LDCM0289 | AC29 | HEK-293T | C34(0.98) | LDD1528 | [14] |
| LDCM0290 | AC3 | HEK-293T | C34(1.13) | LDD1529 | [14] |
| LDCM0292 | AC31 | HEK-293T | C34(1.18) | LDD1531 | [14] |
| LDCM0293 | AC32 | HEK-293T | C34(1.03) | LDD1532 | [14] |
| LDCM0294 | AC33 | HEK-293T | C34(1.03) | LDD1533 | [14] |
| LDCM0295 | AC34 | HEK-293T | C34(1.00) | LDD1534 | [14] |
| LDCM0296 | AC35 | HEK-293T | C34(1.06) | LDD1535 | [14] |
| LDCM0297 | AC36 | HEK-293T | C34(0.96) | LDD1536 | [14] |
| LDCM0298 | AC37 | HEK-293T | C34(1.18) | LDD1537 | [14] |
| LDCM0300 | AC39 | HEK-293T | C34(1.01) | LDD1539 | [14] |
| LDCM0301 | AC4 | HEK-293T | C34(1.07) | LDD1540 | [14] |
| LDCM0302 | AC40 | HEK-293T | C34(0.97) | LDD1541 | [14] |
| LDCM0303 | AC41 | HEK-293T | C34(1.04) | LDD1542 | [14] |
| LDCM0304 | AC42 | HEK-293T | C34(0.98) | LDD1543 | [14] |
| LDCM0305 | AC43 | HEK-293T | C34(0.95) | LDD1544 | [14] |
| LDCM0306 | AC44 | HEK-293T | C34(1.12) | LDD1545 | [14] |
| LDCM0307 | AC45 | HEK-293T | C34(1.06) | LDD1546 | [14] |
| LDCM0309 | AC47 | HEK-293T | C34(1.04) | LDD1548 | [14] |
| LDCM0310 | AC48 | HEK-293T | C34(1.00) | LDD1549 | [14] |
| LDCM0311 | AC49 | HEK-293T | C34(0.80) | LDD1550 | [14] |
| LDCM0312 | AC5 | HEK-293T | C34(1.04) | LDD1551 | [14] |
| LDCM0313 | AC50 | HEK-293T | C34(0.89) | LDD1552 | [14] |
| LDCM0314 | AC51 | HEK-293T | C34(0.98) | LDD1553 | [14] |
| LDCM0315 | AC52 | HEK-293T | C34(1.16) | LDD1554 | [14] |
| LDCM0316 | AC53 | HEK-293T | C34(0.96) | LDD1555 | [14] |
| LDCM0318 | AC55 | HEK-293T | C34(0.88) | LDD1557 | [14] |
| LDCM0319 | AC56 | HEK-293T | C34(0.95) | LDD1558 | [14] |
| LDCM0320 | AC57 | HEK-293T | C34(0.91) | LDD1559 | [14] |
| LDCM0321 | AC58 | HEK-293T | C34(0.89) | LDD1560 | [14] |
| LDCM0322 | AC59 | HEK-293T | C34(0.98) | LDD1561 | [14] |
| LDCM0324 | AC60 | HEK-293T | C34(0.90) | LDD1563 | [14] |
| LDCM0325 | AC61 | HEK-293T | C34(0.86) | LDD1564 | [14] |
| LDCM0327 | AC63 | HEK-293T | C34(0.91) | LDD1566 | [14] |
| LDCM0328 | AC64 | HEK-293T | C34(0.96) | LDD1567 | [14] |
| LDCM0334 | AC7 | HEK-293T | C34(1.13) | LDD1568 | [14] |
| LDCM0345 | AC8 | HEK-293T | C34(1.04) | LDD1569 | [14] |
| LDCM0248 | AKOS034007472 | HEK-293T | C34(0.99) | LDD1511 | [14] |
| LDCM0356 | AKOS034007680 | HEK-293T | C34(0.99) | LDD1570 | [14] |
| LDCM0275 | AKOS034007705 | HEK-293T | C34(0.99) | LDD1514 | [14] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [8] |
| LDCM0632 | CL-Sc | Hep-G2 | C34(1.15) | LDD2227 | [6] |
| LDCM0367 | CL1 | HEK-293T | C34(0.94) | LDD1571 | [14] |
| LDCM0369 | CL100 | HEK-293T | C34(1.02) | LDD1573 | [14] |
| LDCM0370 | CL101 | HEK-293T | C34(0.94) | LDD1574 | [14] |
| LDCM0371 | CL102 | HEK-293T | C34(0.92) | LDD1575 | [14] |
| LDCM0373 | CL104 | HEK-293T | C34(0.98) | LDD1577 | [14] |
| LDCM0374 | CL105 | HEK-293T | C34(0.91) | LDD1578 | [14] |
| LDCM0375 | CL106 | HEK-293T | C34(0.95) | LDD1579 | [14] |
| LDCM0377 | CL108 | HEK-293T | C34(0.96) | LDD1581 | [14] |
| LDCM0378 | CL109 | HEK-293T | C34(0.96) | LDD1582 | [14] |
| LDCM0379 | CL11 | HEK-293T | C34(0.95) | LDD1583 | [14] |
| LDCM0380 | CL110 | HEK-293T | C34(0.90) | LDD1584 | [14] |
| LDCM0382 | CL112 | HEK-293T | C34(0.94) | LDD1586 | [14] |
| LDCM0383 | CL113 | HEK-293T | C34(1.06) | LDD1587 | [14] |
| LDCM0384 | CL114 | HEK-293T | C34(0.89) | LDD1588 | [14] |
| LDCM0386 | CL116 | HEK-293T | C34(1.04) | LDD1590 | [14] |
| LDCM0387 | CL117 | HEK-293T | C34(0.99) | LDD1591 | [14] |
| LDCM0388 | CL118 | HEK-293T | C34(0.89) | LDD1592 | [14] |
| LDCM0390 | CL12 | HEK-293T | C34(0.95) | LDD1594 | [14] |
| LDCM0391 | CL120 | HEK-293T | C34(0.98) | LDD1595 | [14] |
| LDCM0392 | CL121 | HEK-293T | C34(0.93) | LDD1596 | [14] |
| LDCM0393 | CL122 | HEK-293T | C34(0.81) | LDD1597 | [14] |
| LDCM0395 | CL124 | HEK-293T | C34(0.95) | LDD1599 | [14] |
| LDCM0396 | CL125 | HEK-293T | C34(0.62) | LDD1600 | [14] |
| LDCM0397 | CL126 | HEK-293T | C34(0.78) | LDD1601 | [14] |
| LDCM0399 | CL128 | HEK-293T | C34(0.99) | LDD1603 | [14] |
| LDCM0400 | CL13 | HEK-293T | C34(0.86) | LDD1604 | [14] |
| LDCM0401 | CL14 | HEK-293T | C34(1.06) | LDD1605 | [14] |
| LDCM0403 | CL16 | HEK-293T | C34(1.05) | LDD1607 | [14] |
| LDCM0404 | CL17 | HEK-293T | C34(0.86) | LDD1608 | [14] |
| LDCM0405 | CL18 | HEK-293T | C34(0.91) | LDD1609 | [14] |
| LDCM0406 | CL19 | HEK-293T | C34(0.85) | LDD1610 | [14] |
| LDCM0407 | CL2 | HEK-293T | C34(0.91) | LDD1611 | [14] |
| LDCM0408 | CL20 | HEK-293T | C34(1.09) | LDD1612 | [14] |
| LDCM0409 | CL21 | HEK-293T | C34(0.93) | LDD1613 | [14] |
| LDCM0411 | CL23 | HEK-293T | C34(1.06) | LDD1615 | [14] |
| LDCM0412 | CL24 | HEK-293T | C34(1.02) | LDD1616 | [14] |
| LDCM0413 | CL25 | HEK-293T | C34(0.74) | LDD1617 | [14] |
| LDCM0414 | CL26 | HEK-293T | C34(0.96) | LDD1618 | [14] |
| LDCM0416 | CL28 | HEK-293T | C34(0.97) | LDD1620 | [14] |
| LDCM0417 | CL29 | HEK-293T | C34(1.14) | LDD1621 | [14] |
| LDCM0419 | CL30 | HEK-293T | C34(1.11) | LDD1623 | [14] |
| LDCM0420 | CL31 | HEK-293T | C34(0.94) | LDD1624 | [14] |
| LDCM0421 | CL32 | HEK-293T | C34(1.07) | LDD1625 | [14] |
| LDCM0422 | CL33 | HEK-293T | C34(0.78) | LDD1626 | [14] |
| LDCM0424 | CL35 | HEK-293T | C34(0.89) | LDD1628 | [14] |
| LDCM0425 | CL36 | HEK-293T | C34(1.18) | LDD1629 | [14] |
| LDCM0426 | CL37 | HEK-293T | C34(0.94) | LDD1630 | [14] |
| LDCM0429 | CL4 | HEK-293T | C34(0.94) | LDD1633 | [14] |
| LDCM0430 | CL40 | HEK-293T | C34(1.01) | LDD1634 | [14] |
| LDCM0431 | CL41 | HEK-293T | C34(1.10) | LDD1635 | [14] |
| LDCM0432 | CL42 | HEK-293T | C34(0.91) | LDD1636 | [14] |
| LDCM0433 | CL43 | HEK-293T | C34(0.96) | LDD1637 | [14] |
| LDCM0434 | CL44 | HEK-293T | C34(1.07) | LDD1638 | [14] |
| LDCM0435 | CL45 | HEK-293T | C34(0.93) | LDD1639 | [14] |
| LDCM0437 | CL47 | HEK-293T | C34(1.05) | LDD1641 | [14] |
| LDCM0438 | CL48 | HEK-293T | C34(1.13) | LDD1642 | [14] |
| LDCM0439 | CL49 | HEK-293T | C34(0.93) | LDD1643 | [14] |
| LDCM0440 | CL5 | HEK-293T | C34(0.85) | LDD1644 | [14] |
| LDCM0441 | CL50 | HEK-293T | C34(1.01) | LDD1645 | [14] |
| LDCM0443 | CL52 | HEK-293T | C34(0.96) | LDD1646 | [14] |
| LDCM0444 | CL53 | HEK-293T | C34(1.01) | LDD1647 | [14] |
| LDCM0445 | CL54 | HEK-293T | C34(0.77) | LDD1648 | [14] |
| LDCM0446 | CL55 | HEK-293T | C34(1.03) | LDD1649 | [14] |
| LDCM0447 | CL56 | HEK-293T | C34(0.95) | LDD1650 | [14] |
| LDCM0448 | CL57 | HEK-293T | C34(1.01) | LDD1651 | [14] |
| LDCM0450 | CL59 | HEK-293T | C34(1.00) | LDD1653 | [14] |
| LDCM0451 | CL6 | HEK-293T | C34(0.74) | LDD1654 | [14] |
| LDCM0452 | CL60 | HEK-293T | C34(1.00) | LDD1655 | [14] |
| LDCM0453 | CL61 | HEK-293T | C34(0.91) | LDD1656 | [14] |
| LDCM0454 | CL62 | HEK-293T | C34(1.14) | LDD1657 | [14] |
| LDCM0456 | CL64 | HEK-293T | C34(1.03) | LDD1659 | [14] |
| LDCM0457 | CL65 | HEK-293T | C34(0.96) | LDD1660 | [14] |
| LDCM0458 | CL66 | HEK-293T | C34(1.00) | LDD1661 | [14] |
| LDCM0459 | CL67 | HEK-293T | C34(1.05) | LDD1662 | [14] |
| LDCM0460 | CL68 | HEK-293T | C34(1.04) | LDD1663 | [14] |
| LDCM0461 | CL69 | HEK-293T | C34(0.85) | LDD1664 | [14] |
| LDCM0462 | CL7 | HEK-293T | C34(0.96) | LDD1665 | [14] |
| LDCM0464 | CL71 | HEK-293T | C34(0.88) | LDD1667 | [14] |
| LDCM0465 | CL72 | HEK-293T | C34(1.04) | LDD1668 | [14] |
| LDCM0466 | CL73 | HEK-293T | C34(0.88) | LDD1669 | [14] |
| LDCM0467 | CL74 | HEK-293T | C34(0.91) | LDD1670 | [14] |
| LDCM0469 | CL76 | HEK-293T | C34(1.03) | LDD1672 | [14] |
| LDCM0470 | CL77 | HEK-293T | C34(0.87) | LDD1673 | [14] |
| LDCM0471 | CL78 | HEK-293T | C34(1.07) | LDD1674 | [14] |
| LDCM0472 | CL79 | HEK-293T | C34(1.08) | LDD1675 | [14] |
| LDCM0473 | CL8 | HEK-293T | C34(0.63) | LDD1676 | [14] |
| LDCM0474 | CL80 | HEK-293T | C34(1.08) | LDD1677 | [14] |
| LDCM0475 | CL81 | HEK-293T | C34(0.91) | LDD1678 | [14] |
| LDCM0477 | CL83 | HEK-293T | C34(1.06) | LDD1680 | [14] |
| LDCM0478 | CL84 | HEK-293T | C34(0.92) | LDD1681 | [14] |
| LDCM0479 | CL85 | HEK-293T | C34(0.64) | LDD1682 | [14] |
| LDCM0480 | CL86 | HEK-293T | C34(0.69) | LDD1683 | [14] |
| LDCM0482 | CL88 | HEK-293T | C34(0.96) | LDD1685 | [14] |
| LDCM0483 | CL89 | HEK-293T | C34(0.78) | LDD1686 | [14] |
| LDCM0484 | CL9 | HEK-293T | C34(1.00) | LDD1687 | [14] |
| LDCM0485 | CL90 | HEK-293T | C34(0.61) | LDD1688 | [14] |
| LDCM0486 | CL91 | HEK-293T | C34(0.86) | LDD1689 | [14] |
| LDCM0487 | CL92 | HEK-293T | C34(0.88) | LDD1690 | [14] |
| LDCM0488 | CL93 | HEK-293T | C34(0.86) | LDD1691 | [14] |
| LDCM0490 | CL95 | HEK-293T | C34(0.83) | LDD1693 | [14] |
| LDCM0491 | CL96 | HEK-293T | C34(0.63) | LDD1694 | [14] |
| LDCM0492 | CL97 | HEK-293T | C34(0.94) | LDD1695 | [14] |
| LDCM0493 | CL98 | HEK-293T | C34(0.92) | LDD1696 | [14] |
| LDCM0625 | F8 | Ramos | C34(1.54) | LDD2187 | [15] |
| LDCM0572 | Fragment10 | Ramos | C34(0.51) | LDD2189 | [15] |
| LDCM0573 | Fragment11 | Ramos | C34(19.10) | LDD2190 | [15] |
| LDCM0574 | Fragment12 | Ramos | C34(0.44) | LDD2191 | [15] |
| LDCM0575 | Fragment13 | Ramos | C34(0.99) | LDD2192 | [15] |
| LDCM0576 | Fragment14 | Ramos | C34(0.76) | LDD2193 | [15] |
| LDCM0579 | Fragment20 | Ramos | C34(0.41) | LDD2194 | [15] |
| LDCM0580 | Fragment21 | Ramos | C34(0.81) | LDD2195 | [15] |
| LDCM0582 | Fragment23 | Ramos | C34(0.90) | LDD2196 | [15] |
| LDCM0578 | Fragment27 | Ramos | C34(0.88) | LDD2197 | [15] |
| LDCM0586 | Fragment28 | Ramos | C34(0.62) | LDD2198 | [15] |
| LDCM0588 | Fragment30 | Ramos | C34(1.06) | LDD2199 | [15] |
| LDCM0589 | Fragment31 | Ramos | C34(0.96) | LDD2200 | [15] |
| LDCM0590 | Fragment32 | Ramos | C34(0.70) | LDD2201 | [15] |
| LDCM0468 | Fragment33 | Ramos | C34(0.90) | LDD2202 | [15] |
| LDCM0596 | Fragment38 | Ramos | C34(1.06) | LDD2203 | [15] |
| LDCM0566 | Fragment4 | Ramos | C34(0.72) | LDD2184 | [15] |
| LDCM0427 | Fragment51 | HEK-293T | C34(1.02) | LDD1631 | [14] |
| LDCM0610 | Fragment52 | Ramos | C34(1.08) | LDD2204 | [15] |
| LDCM0614 | Fragment56 | Ramos | C34(1.47) | LDD2205 | [15] |
| LDCM0569 | Fragment7 | Ramos | C34(0.51) | LDD2186 | [15] |
| LDCM0571 | Fragment9 | Ramos | C34(0.45) | LDD2188 | [15] |
| LDCM0022 | KB02 | HEK-293T | C34(1.07) | LDD1492 | [14] |
| LDCM0023 | KB03 | HEK-293T | C34(0.92) | LDD1497 | [14] |
| LDCM0024 | KB05 | MIA PaCa-2 | C34(1.54) | LDD3330 | [1] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0225 | [8] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C34(0.66) | LDD2097 | [2] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C34(0.88) | LDD2098 | [2] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C34(0.81) | LDD2099 | [2] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C34(0.46) | LDD2100 | [2] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C34(0.57) | LDD2101 | [2] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C34(0.37) | LDD2104 | [2] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C34(1.15) | LDD2105 | [2] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C34(0.35) | LDD2106 | [2] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C34(0.74) | LDD2107 | [2] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C34(0.51) | LDD2108 | [2] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C34(0.47) | LDD2110 | [2] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C34(1.01) | LDD2111 | [2] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C34(0.41) | LDD2115 | [2] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C34(1.34) | LDD2119 | [2] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C34(1.71) | LDD2129 | [2] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C34(0.68) | LDD2135 | [2] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C34(0.83) | LDD2140 | [2] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C34(0.70) | LDD2141 | [2] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C34(0.58) | LDD2146 | [2] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C34(0.44) | LDD2148 | [2] |
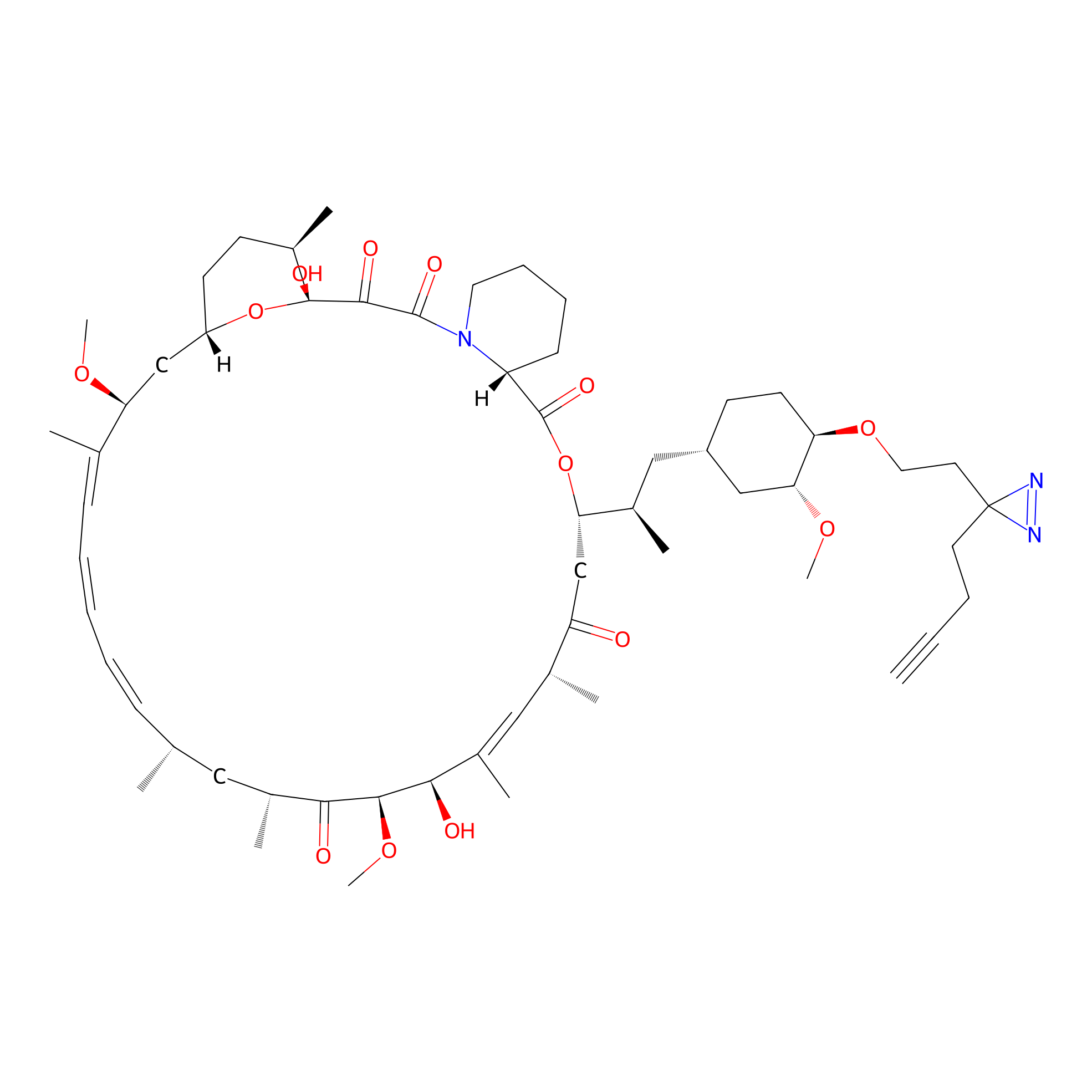
| LDCM0090 | Rapamycin | JHH-7 | 3.80 | LDD0213 | [12] |
References