Details of the Target
General Information of Target
| Target ID | LDTP04789 | |||||
|---|---|---|---|---|---|---|
| Target Name | Synaptojanin-2-binding protein (SYNJ2BP) | |||||
| Gene Name | SYNJ2BP | |||||
| Gene ID | 55333 | |||||
| Synonyms |
OMP25; Synaptojanin-2-binding protein; Mitochondrial outer membrane protein 25 |
|||||
| 3D Structure | ||||||
| Sequence |
MNGRVDYLVTEEEINLTRGPSGLGFNIVGGTDQQYVSNDSGIYVSRIKENGAAALDGRLQ
EGDKILSVNGQDLKNLLHQDAVDLFRNAGYAVSLRVQHRLQVQNGPIGHRGEGDPSGIPI FMVLVPVFALTMVAAWAFMRYRQQL |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Subcellular location |
Mitochondrion outer membrane
|
|||||
| Function | Regulates endocytosis of activin type 2 receptor kinases through the Ral/RALBP1-dependent pathway and may be involved in suppression of activin-induced signal transduction. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
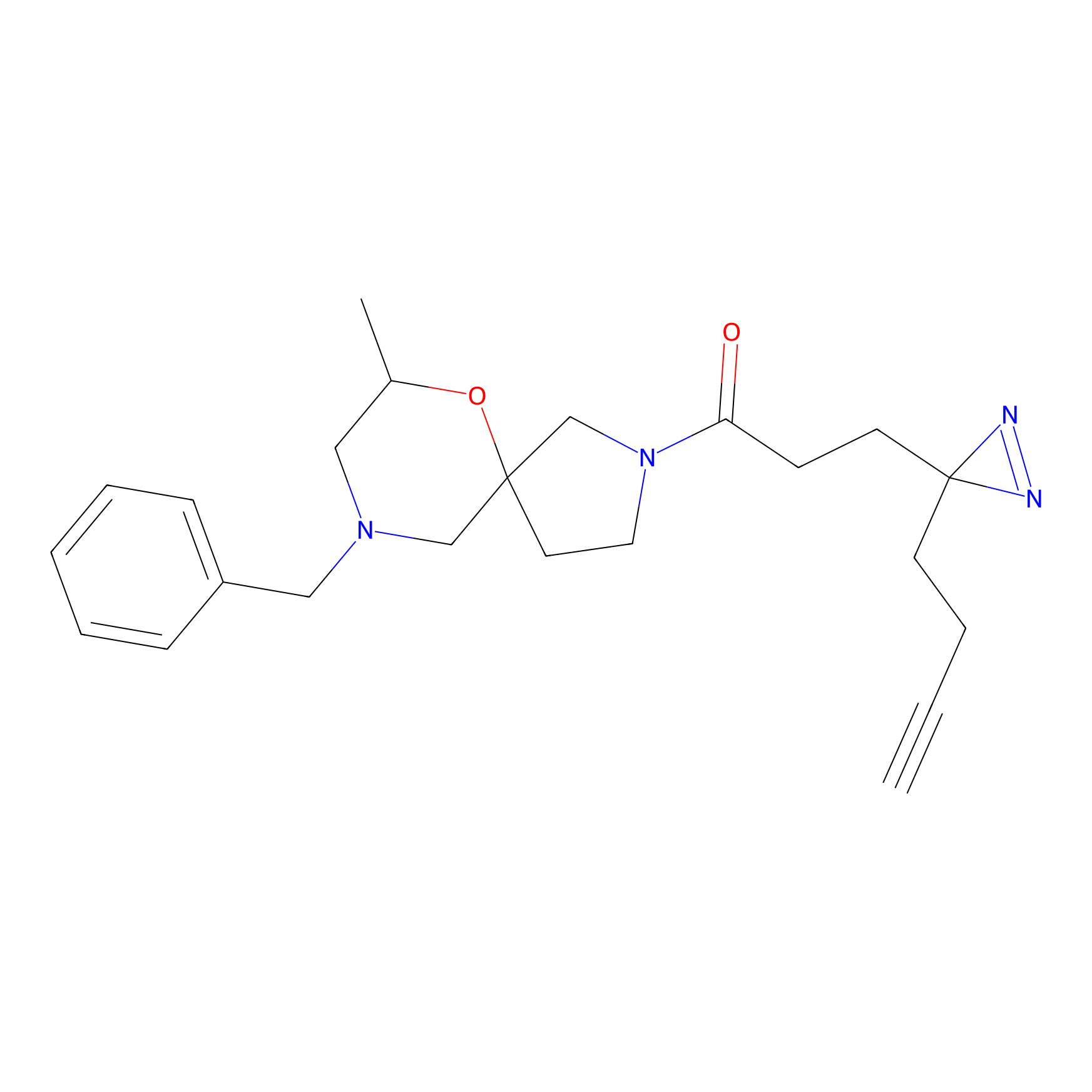
STPyne Probe Info |
 |
K48(4.04) | LDD0277 | [1] | |
|
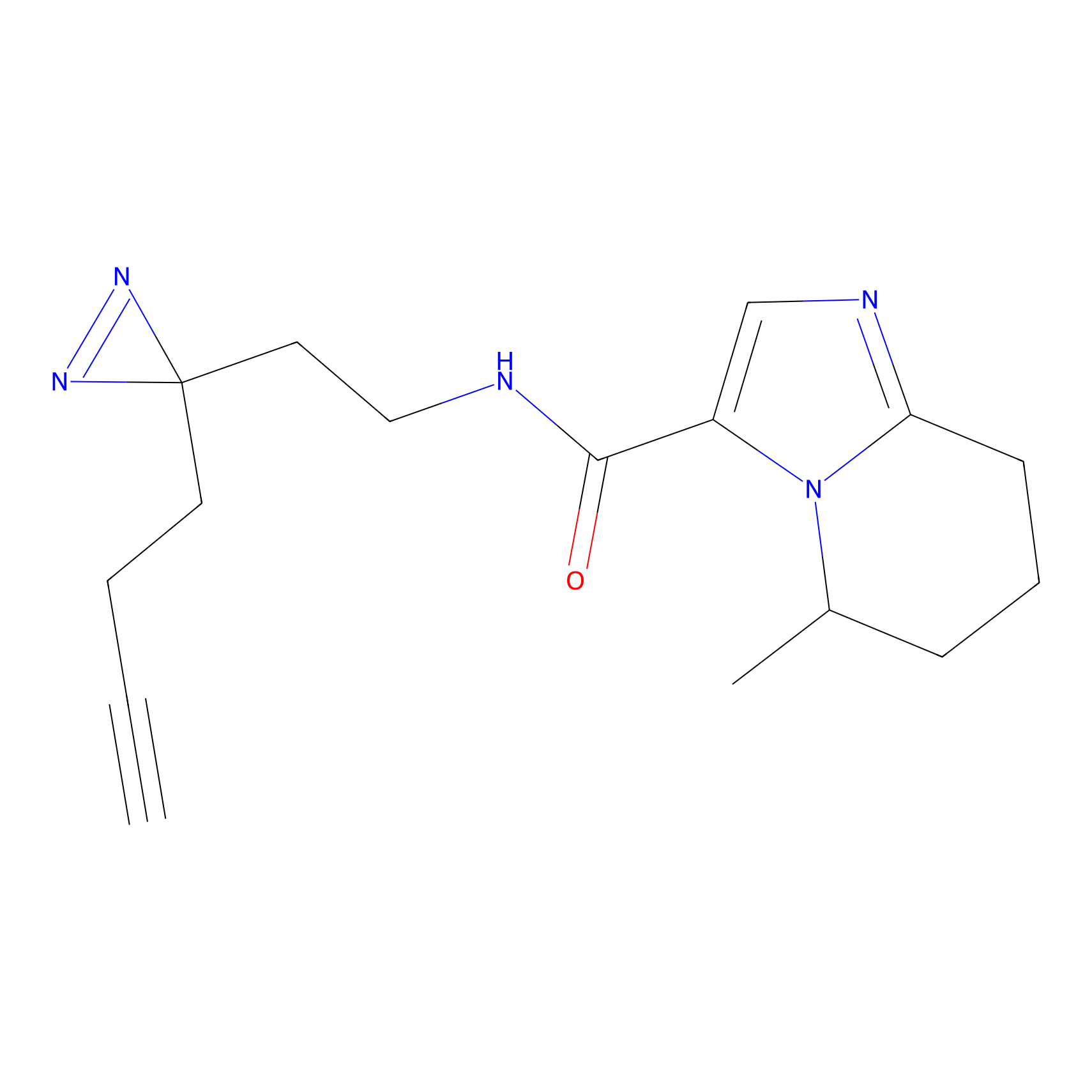
Acrolein Probe Info |
 |
N.A. | LDD0224 | [2] | |
|
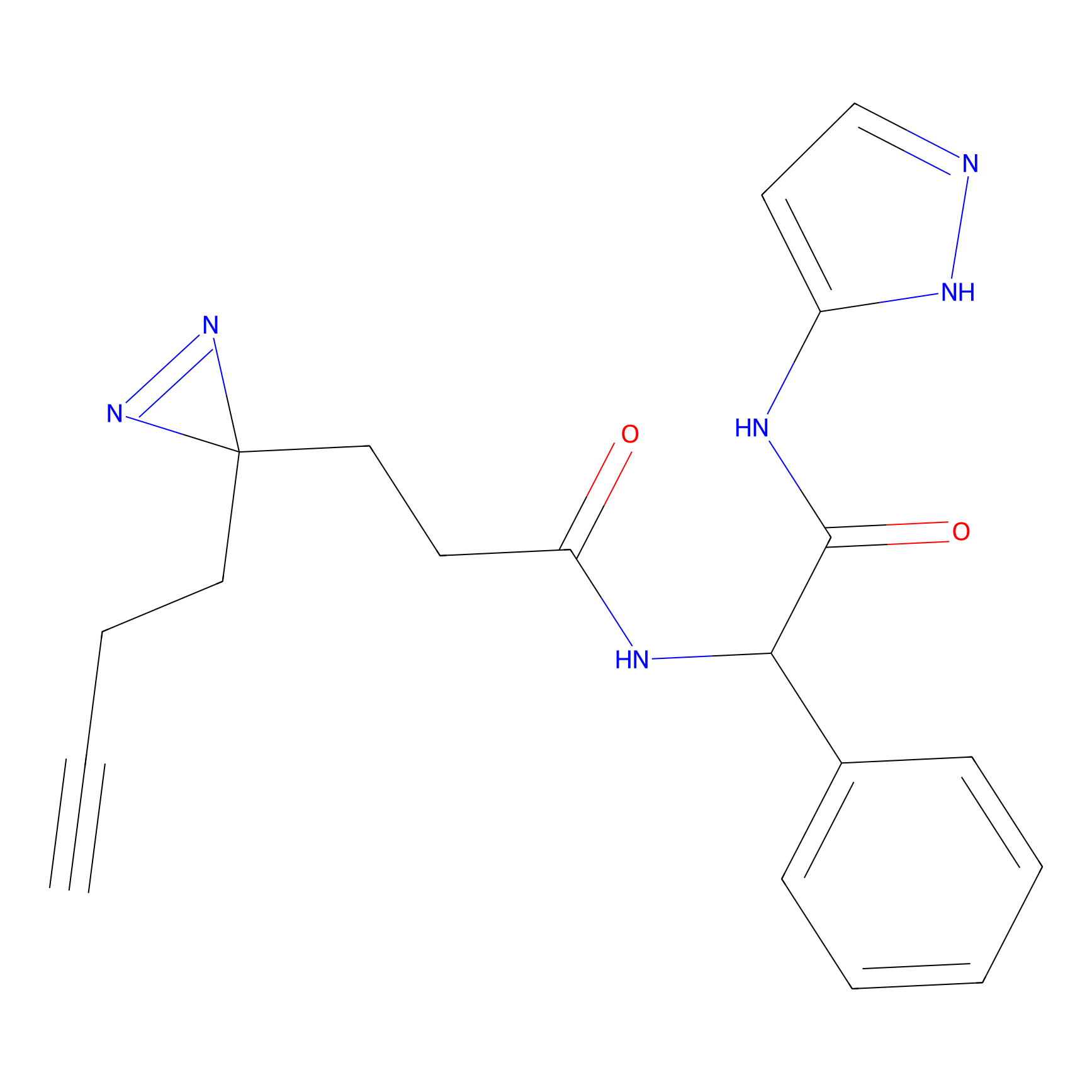
5E-2FA Probe Info |
 |
H109(0.00); H78(0.00) | LDD2235 | [3] | |
|
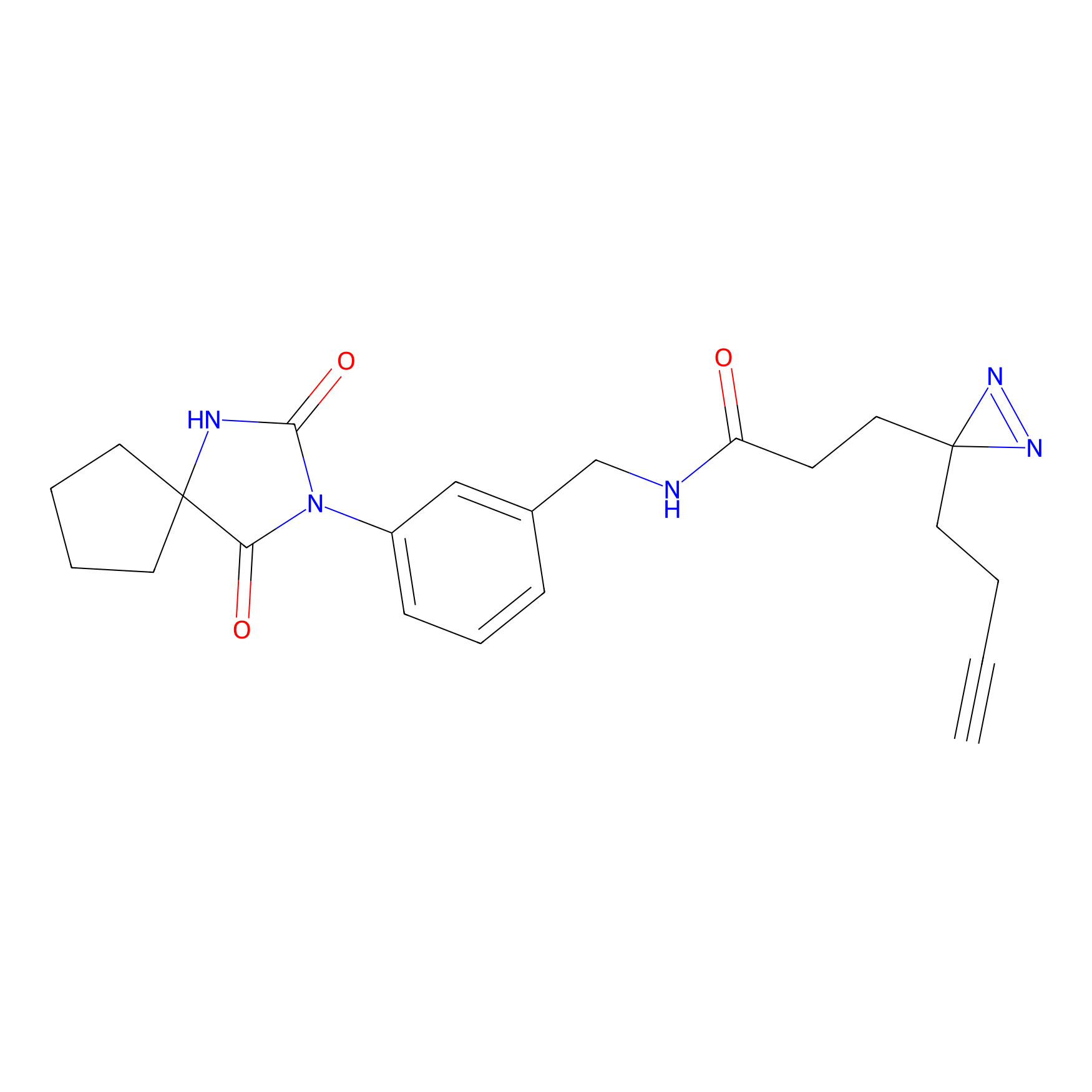
m-APA Probe Info |
 |
N.A. | LDD2231 | [3] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [4] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Cyclic AMP-responsive element-binding protein 3-like protein 1 (CREB3L1) | BZIP family | Q96BA8 | |||
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| B-cell antigen receptor complex-associated protein alpha chain (CD79A) | . | P11912 | |||
Other
References