Details of the Target
General Information of Target
| Target ID | LDTP02609 | |||||
|---|---|---|---|---|---|---|
| Target Name | ADP/ATP translocase 3 (SLC25A6) | |||||
| Gene Name | SLC25A6 | |||||
| Gene ID | 293 | |||||
| Synonyms |
AAC3; ANT3; ADP/ATP translocase 3; ADP,ATP carrier protein 3; ADP,ATP carrier protein, isoform T2; ANT 2; Adenine nucleotide translocator 3; ANT 3; Solute carrier family 25 member 6) [Cleaved into: ADP/ATP translocase 3, N-terminally processed]
|
|||||
| 3D Structure | ||||||
| Sequence |
MTEQAISFAKDFLAGGIAAAISKTAVAPIERVKLLLQVQHASKQIAADKQYKGIVDCIVR
IPKEQGVLSFWRGNLANVIRYFPTQALNFAFKDKYKQIFLGGVDKHTQFWRYFAGNLASG GAAGATSLCFVYPLDFARTRLAADVGKSGTEREFRGLGDCLVKITKSDGIRGLYQGFSVS VQGIIIYRAAYFGVYDTAKGMLPDPKNTHIVVSWMIAQTVTAVAGVVSYPFDTVRRRMMM QSGRKGADIMYTGTVDCWRKIFRDEGGKAFFKGAWSNVLRGMGGAFVLVLYDELKKVI |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Mitochondrial carrier (TC 2.A.29) family
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
ADP:ATP antiporter that mediates import of ADP into the mitochondrial matrix for ATP synthesis, and export of ATP out to fuel the cell. Cycles between the cytoplasmic-open state (c-state) and the matrix-open state (m-state): operates by the alternating access mechanism with a single substrate-binding site intermittently exposed to either the cytosolic (c-state) or matrix (m-state) side of the inner mitochondrial membrane. In addition to its ADP:ATP antiporter activity, also involved in mitochondrial uncoupling and mitochondrial permeability transition pore (mPTP) activity. Plays a role in mitochondrial uncoupling by acting as a proton transporter: proton transport uncouples the proton flows via the electron transport chain and ATP synthase to reduce the efficiency of ATP production and cause mitochondrial thermogenesis. Proton transporter activity is inhibited by ADP:ATP antiporter activity, suggesting that SLC25A6/ANT3 acts as a master regulator of mitochondrial energy output by maintaining a delicate balance between ATP production (ADP:ATP antiporter activity) and thermogenesis (proton transporter activity). Proton transporter activity requires free fatty acids as cofactor, but does not transport it. Also plays a key role in mPTP opening, a non-specific pore that enables free passage of the mitochondrial membranes to solutes of up to 1.5 kDa, and which contributes to cell death. It is however unclear if SLC25A6/ANT3 constitutes a pore-forming component of mPTP or regulates it.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
12.96 | LDD0402 | [2] | |
|
CHEMBL5175495 Probe Info |
 |
9.55 | LDD0196 | [3] | |
|
N1 Probe Info |
 |
13.69 | LDD0242 | [4] | |
|
W1 Probe Info |
 |
10.65 | LDD0235 | [5] | |
|
TH211 Probe Info |
 |
Y251(17.65); Y112(7.74) | LDD0260 | [6] | |
|
C-Sul Probe Info |
 |
4.60 | LDD0066 | [7] | |
|
ONAyne Probe Info |
 |
N.A. | LDD0273 | [8] | |
|
BTD Probe Info |
 |
C160(2.11); C57(1.93) | LDD1700 | [9] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [10] | |
|
Acrolein Probe Info |
 |
H106(0.00); C57(0.00) | LDD0221 | [11] | |
|
DBIA Probe Info |
 |
C160(2.64); C257(2.39) | LDD0080 | [12] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [13] | |
|
ATP probe Probe Info |
 |
K96(0.00); K105(0.00); K43(0.00); K166(0.00) | LDD0199 | [14] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C257(0.00); C57(0.00) | LDD0038 | [15] | |
|
IA-alkyne Probe Info |
 |
C257(0.00); C129(0.00); C160(0.00) | LDD0032 | [16] | |
|
ATP probe Probe Info |
 |
K105(0.00); K272(0.00); K163(0.00); K94(0.00) | LDD0035 | [17] | |
|
IPM Probe Info |
 |
C129(0.00); C160(0.00) | LDD0025 | [18] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [18] | |
|
NAIA_4 Probe Info |
 |
C57(0.00); C129(0.00); C257(0.00) | LDD2226 | [19] | |
|
TFBX Probe Info |
 |
C57(0.00); C257(0.00); C129(0.00) | LDD0027 | [18] | |
|
WYneN Probe Info |
 |
C257(0.00); C160(0.00) | LDD0021 | [20] | |
|
WYneO Probe Info |
 |
C257(0.00); C160(0.00) | LDD0022 | [20] | |
|
Compound 10 Probe Info |
 |
C160(0.00); C57(0.00) | LDD2216 | [21] | |
|
NHS Probe Info |
 |
K63(0.00); K92(0.00); K272(0.00); K147(0.00) | LDD0010 | [20] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0152 | [22] | |
|
SF Probe Info |
 |
K33(0.00); Y81(0.00) | LDD0028 | [23] | |
|
STPyne Probe Info |
 |
K94(0.00); K63(0.00); K163(0.00); K92(0.00) | LDD0009 | [20] | |
|
VSF Probe Info |
 |
C160(0.00); C257(0.00) | LDD0007 | [20] | |
|
Phosphinate-6 Probe Info |
 |
C129(0.00); C160(0.00); C257(0.00) | LDD0018 | [24] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [25] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [26] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [19] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [26] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [26] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
IMP2070 Probe Info |
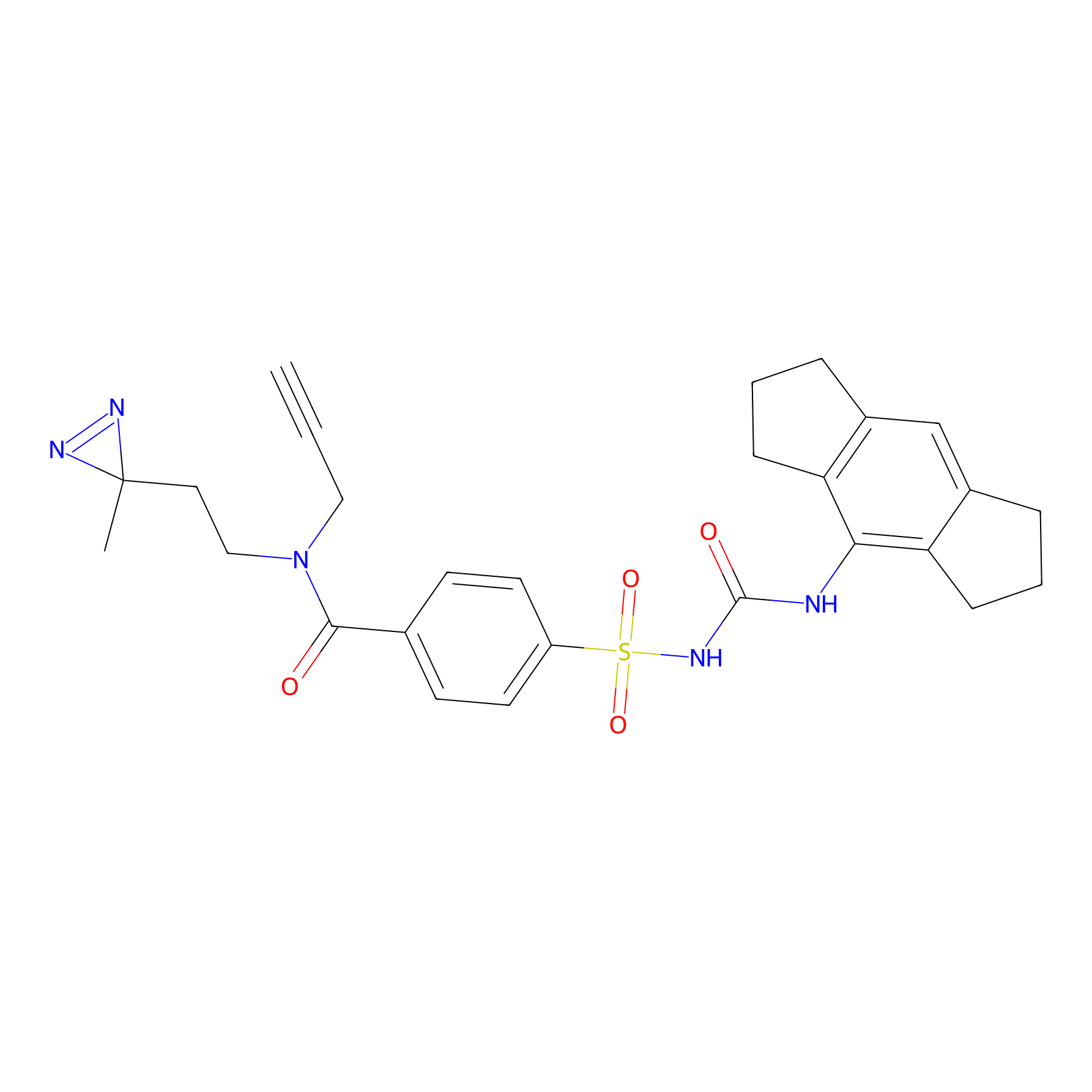 |
1.56 | LDD0367 | [27] | |
|
C022 Probe Info |
 |
10.63 | LDD1728 | [28] | |
|
C040 Probe Info |
 |
10.20 | LDD1740 | [28] | |
|
C056 Probe Info |
 |
18.00 | LDD1753 | [28] | |
|
C091 Probe Info |
 |
11.39 | LDD1782 | [28] | |
|
C092 Probe Info |
 |
16.45 | LDD1783 | [28] | |
|
C094 Probe Info |
 |
25.63 | LDD1785 | [28] | |
|
C106 Probe Info |
 |
20.53 | LDD1793 | [28] | |
|
C112 Probe Info |
 |
17.15 | LDD1799 | [28] | |
|
C134 Probe Info |
 |
26.35 | LDD1816 | [28] | |
|
C143 Probe Info |
 |
16.91 | LDD1825 | [28] | |
|
C153 Probe Info |
 |
15.03 | LDD1834 | [28] | |
|
C160 Probe Info |
 |
7.67 | LDD1840 | [28] | |
|
C161 Probe Info |
 |
12.55 | LDD1841 | [28] | |
|
C163 Probe Info |
 |
6.63 | LDD1843 | [28] | |
|
C178 Probe Info |
 |
14.62 | LDD1857 | [28] | |
|
C196 Probe Info |
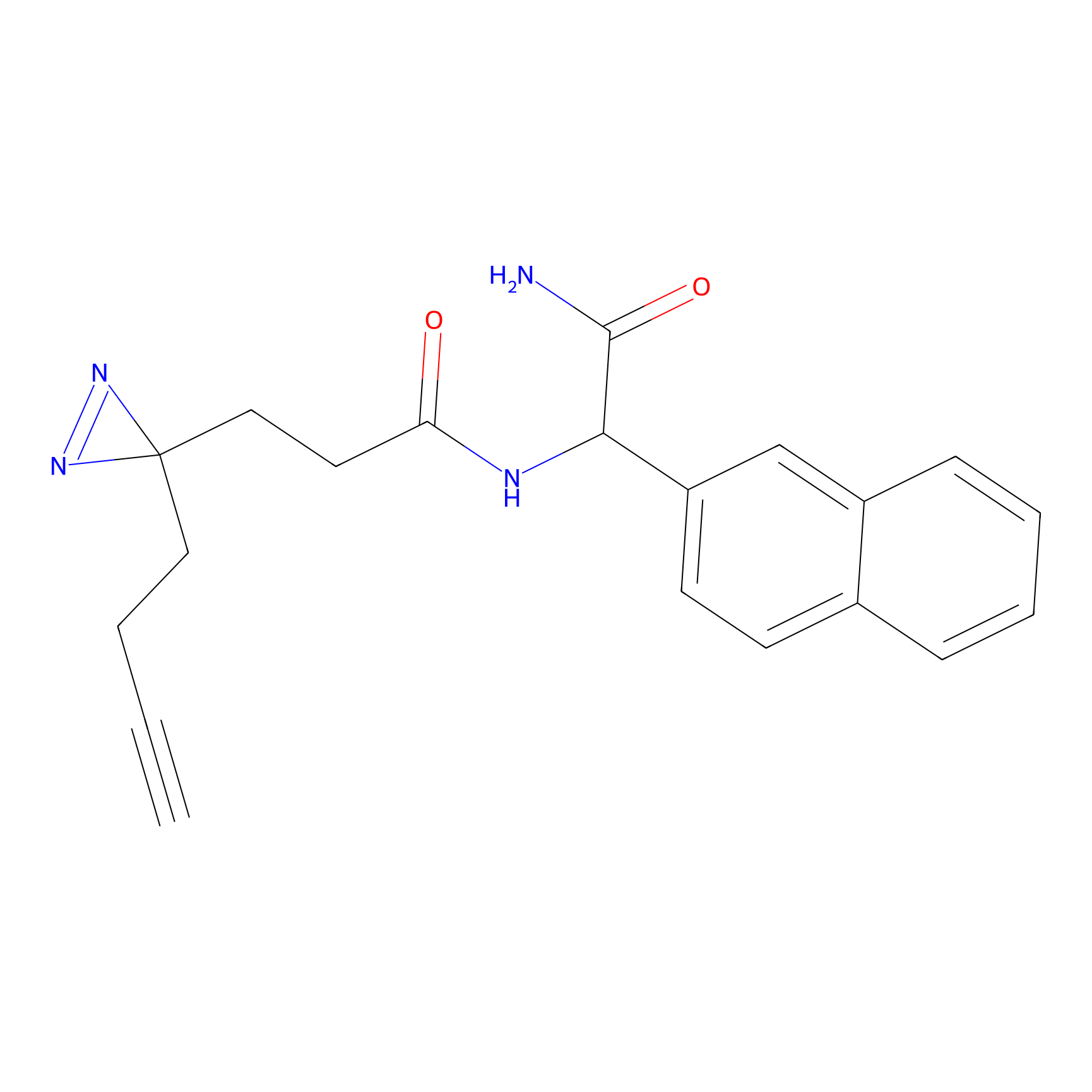 |
12.04 | LDD1872 | [28] | |
|
C201 Probe Info |
 |
26.17 | LDD1877 | [28] | |
|
C218 Probe Info |
 |
11.24 | LDD1892 | [28] | |
|
C219 Probe Info |
 |
5.10 | LDD1893 | [28] | |
|
C220 Probe Info |
 |
20.11 | LDD1894 | [28] | |
|
C225 Probe Info |
 |
5.39 | LDD1898 | [28] | |
|
C228 Probe Info |
 |
26.91 | LDD1901 | [28] | |
|
C231 Probe Info |
 |
16.34 | LDD1904 | [28] | |
|
C235 Probe Info |
 |
17.63 | LDD1908 | [28] | |
|
C246 Probe Info |
 |
12.64 | LDD1919 | [28] | |
|
C277 Probe Info |
 |
11.39 | LDD1947 | [28] | |
|
C285 Probe Info |
 |
22.78 | LDD1955 | [28] | |
|
C287 Probe Info |
 |
8.75 | LDD1957 | [28] | |
|
C289 Probe Info |
 |
35.02 | LDD1959 | [28] | |
|
C293 Probe Info |
 |
27.28 | LDD1963 | [28] | |
|
C296 Probe Info |
 |
12.21 | LDD1966 | [28] | |
|
C310 Probe Info |
 |
19.29 | LDD1977 | [28] | |
|
C339 Probe Info |
 |
5.82 | LDD2002 | [28] | |
|
C349 Probe Info |
 |
17.51 | LDD2010 | [28] | |
|
C350 Probe Info |
 |
66.72 | LDD2011 | [28] | |
|
C362 Probe Info |
 |
29.04 | LDD2023 | [28] | |
|
C366 Probe Info |
 |
7.26 | LDD2027 | [28] | |
|
C388 Probe Info |
 |
45.57 | LDD2047 | [28] | |
|
C403 Probe Info |
 |
13.74 | LDD2061 | [28] | |
|
FFF probe11 Probe Info |
 |
14.81 | LDD0471 | [29] | |
|
FFF probe13 Probe Info |
 |
16.20 | LDD0475 | [29] | |
|
FFF probe14 Probe Info |
 |
14.61 | LDD0477 | [29] | |
|
FFF probe2 Probe Info |
 |
15.88 | LDD0463 | [29] | |
|
FFF probe3 Probe Info |
 |
12.04 | LDD0464 | [29] | |
|
FFF probe4 Probe Info |
 |
8.93 | LDD0466 | [29] | |
|
JN0003 Probe Info |
 |
15.29 | LDD0469 | [29] | |
|
BD-F Probe Info |
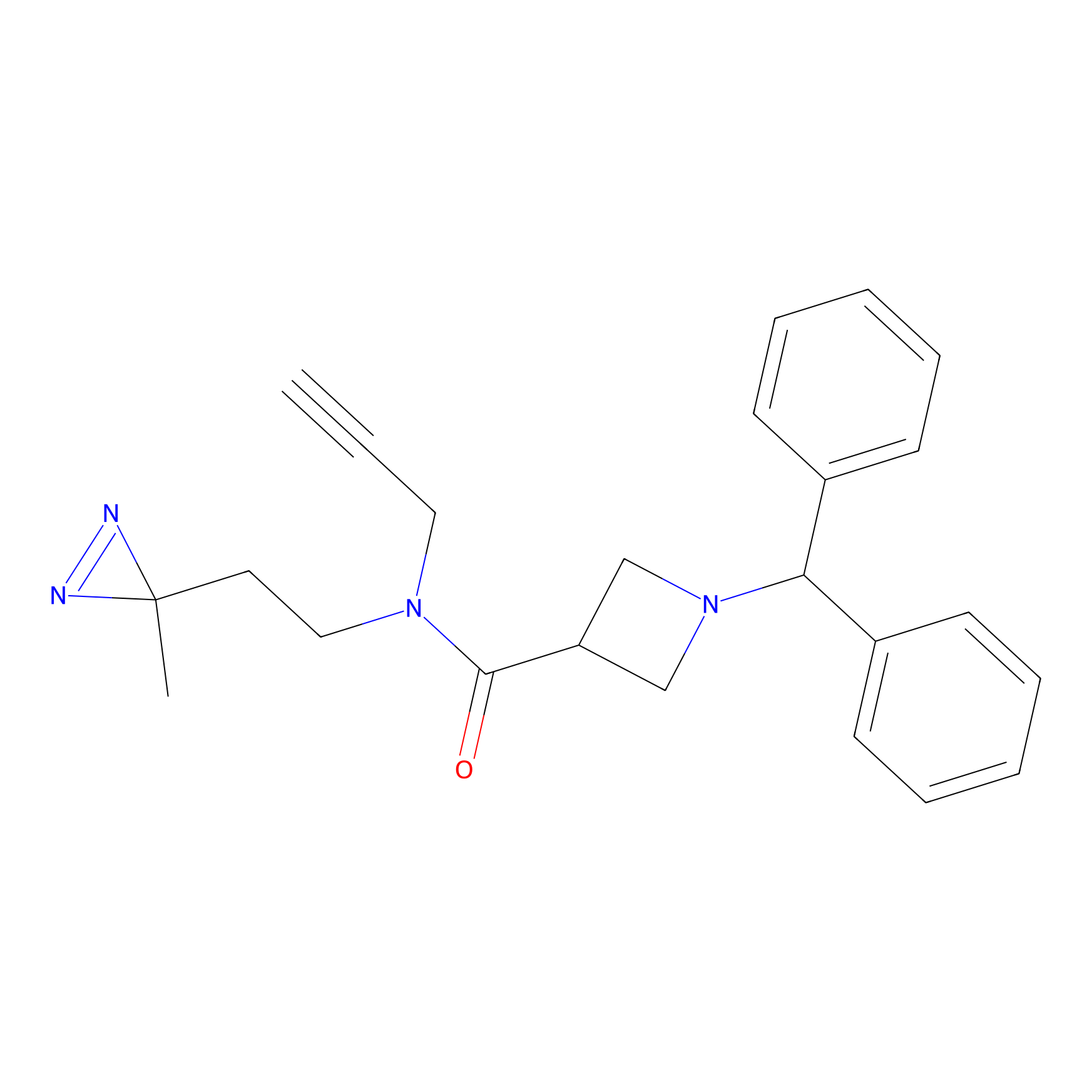 |
D292(0.00); K295(0.00); L294(0.00); E293(0.00) | LDD0024 | [30] | |
|
TM-F Probe Info |
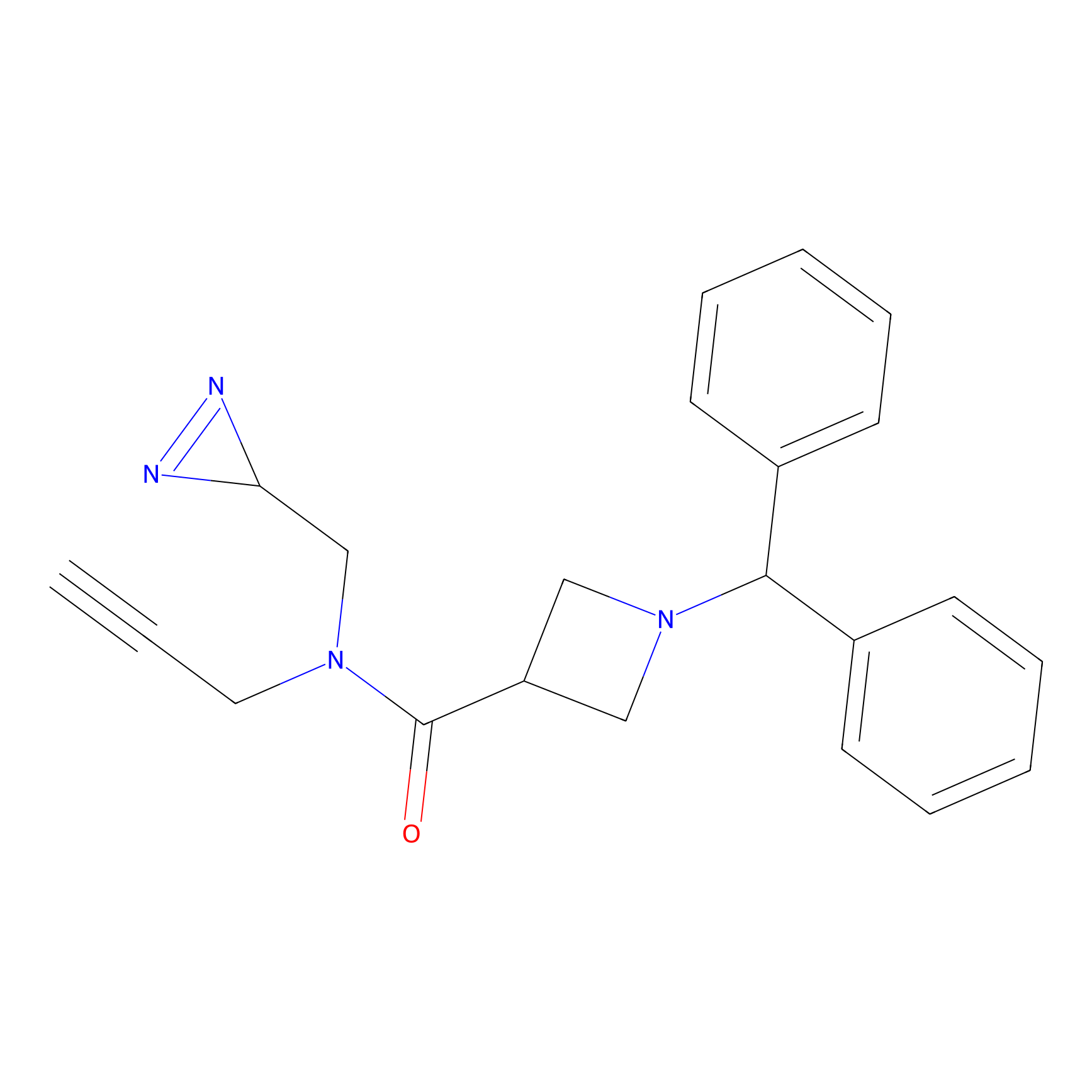 |
G120(0.00); S119(0.00); G124(0.00) | LDD0020 | [30] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [31] | |
|
OEA-DA Probe Info |
 |
3.84 | LDD0046 | [32] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C57(0.67); C160(0.74) | LDD2142 | [9] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C57(1.08); C160(0.95) | LDD2112 | [9] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C57(0.65) | LDD2095 | [9] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C57(1.09); C160(1.24) | LDD2152 | [9] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C160(0.91) | LDD2132 | [9] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C57(0.88) | LDD2131 | [9] |
| LDCM0214 | AC1 | HCT 116 | C129(0.73); C160(1.11); C257(1.05) | LDD0531 | [12] |
| LDCM0215 | AC10 | HCT 116 | C160(1.20); C257(1.13) | LDD0532 | [12] |
| LDCM0216 | AC100 | HCT 116 | C160(1.53); C257(2.23) | LDD0533 | [12] |
| LDCM0217 | AC101 | HCT 116 | C160(0.97); C257(1.43) | LDD0534 | [12] |
| LDCM0218 | AC102 | HCT 116 | C160(1.18); C257(1.41) | LDD0535 | [12] |
| LDCM0219 | AC103 | HCT 116 | C160(1.22); C257(1.64) | LDD0536 | [12] |
| LDCM0220 | AC104 | HCT 116 | C160(1.33); C257(2.14) | LDD0537 | [12] |
| LDCM0221 | AC105 | HCT 116 | C160(1.33); C257(1.67) | LDD0538 | [12] |
| LDCM0222 | AC106 | HCT 116 | C160(1.48); C257(2.40) | LDD0539 | [12] |
| LDCM0223 | AC107 | HCT 116 | C160(1.15); C257(2.43) | LDD0540 | [12] |
| LDCM0224 | AC108 | HCT 116 | C160(1.50); C257(1.47) | LDD0541 | [12] |
| LDCM0225 | AC109 | HCT 116 | C160(1.21); C257(1.83) | LDD0542 | [12] |
| LDCM0226 | AC11 | HCT 116 | C160(0.82); C257(0.87) | LDD0543 | [12] |
| LDCM0227 | AC110 | HCT 116 | C160(1.21); C257(1.57) | LDD0544 | [12] |
| LDCM0228 | AC111 | HCT 116 | C160(1.04); C257(1.73) | LDD0545 | [12] |
| LDCM0229 | AC112 | HCT 116 | C160(1.26); C257(1.86) | LDD0546 | [12] |
| LDCM0230 | AC113 | HCT 116 | C129(0.81); C160(0.82); C257(0.87) | LDD0547 | [12] |
| LDCM0231 | AC114 | HCT 116 | C129(1.05); C160(0.89); C257(0.93) | LDD0548 | [12] |
| LDCM0232 | AC115 | HCT 116 | C129(1.26); C160(0.90); C257(0.90) | LDD0549 | [12] |
| LDCM0233 | AC116 | HCT 116 | C129(1.10); C160(0.73); C257(0.73) | LDD0550 | [12] |
| LDCM0234 | AC117 | HCT 116 | C129(1.09); C160(0.90); C257(0.82) | LDD0551 | [12] |
| LDCM0235 | AC118 | HCT 116 | C129(1.22); C160(0.93); C257(0.95) | LDD0552 | [12] |
| LDCM0236 | AC119 | HCT 116 | C129(0.97); C160(0.77); C257(0.78) | LDD0553 | [12] |
| LDCM0237 | AC12 | HCT 116 | C160(1.22); C257(1.12) | LDD0554 | [12] |
| LDCM0238 | AC120 | HCT 116 | C129(1.67); C160(0.73); C257(0.77) | LDD0555 | [12] |
| LDCM0239 | AC121 | HCT 116 | C129(1.51); C160(1.30); C257(1.29) | LDD0556 | [12] |
| LDCM0240 | AC122 | HCT 116 | C129(0.98); C160(1.03); C257(0.91) | LDD0557 | [12] |
| LDCM0241 | AC123 | HCT 116 | C129(1.90); C160(1.32); C257(1.14) | LDD0558 | [12] |
| LDCM0242 | AC124 | HCT 116 | C129(1.55); C160(1.08); C257(1.00) | LDD0559 | [12] |
| LDCM0243 | AC125 | HCT 116 | C129(1.03); C160(0.91); C257(0.87) | LDD0560 | [12] |
| LDCM0244 | AC126 | HCT 116 | C129(1.42); C160(1.08); C257(1.08) | LDD0561 | [12] |
| LDCM0245 | AC127 | HCT 116 | C129(0.76); C160(1.08); C257(0.95) | LDD0562 | [12] |
| LDCM0246 | AC128 | HEK-293T | C160(0.69); C257(0.86); C57(0.90) | LDD0844 | [12] |
| LDCM0247 | AC129 | HEK-293T | C160(0.63); C257(0.71); C57(1.32) | LDD0845 | [12] |
| LDCM0249 | AC130 | HEK-293T | C160(0.75); C257(0.68); C57(1.22) | LDD0847 | [12] |
| LDCM0250 | AC131 | HEK-293T | C160(0.96); C257(0.67); C57(0.99) | LDD0848 | [12] |
| LDCM0251 | AC132 | HEK-293T | C160(0.60); C257(0.53); C57(1.10) | LDD0849 | [12] |
| LDCM0252 | AC133 | HEK-293T | C160(0.89); C257(0.67); C57(1.35) | LDD0850 | [12] |
| LDCM0253 | AC134 | HEK-293T | C160(0.83); C257(0.93); C57(1.38) | LDD0851 | [12] |
| LDCM0254 | AC135 | HEK-293T | C160(0.97); C257(0.96); C57(1.24) | LDD0852 | [12] |
| LDCM0255 | AC136 | HEK-293T | C160(1.14); C257(0.70); C57(1.57) | LDD0853 | [12] |
| LDCM0256 | AC137 | HEK-293T | C160(0.74); C257(0.75); C57(1.61) | LDD0854 | [12] |
| LDCM0257 | AC138 | HEK-293T | C160(0.86); C257(0.85); C57(1.86) | LDD0855 | [12] |
| LDCM0258 | AC139 | HEK-293T | C160(0.60); C257(0.75); C57(1.62) | LDD0856 | [12] |
| LDCM0259 | AC14 | HCT 116 | C160(0.80); C257(0.83) | LDD0576 | [12] |
| LDCM0260 | AC140 | HEK-293T | C160(0.97); C257(0.79); C57(1.48) | LDD0858 | [12] |
| LDCM0261 | AC141 | HEK-293T | C160(0.88); C257(0.85); C57(1.55) | LDD0859 | [12] |
| LDCM0262 | AC142 | HEK-293T | C160(0.84); C257(0.86); C57(1.34) | LDD0860 | [12] |
| LDCM0263 | AC143 | HCT 116 | C257(1.34) | LDD0580 | [12] |
| LDCM0264 | AC144 | HCT 116 | C257(1.00) | LDD0581 | [12] |
| LDCM0265 | AC145 | HCT 116 | C257(2.03) | LDD0582 | [12] |
| LDCM0266 | AC146 | HCT 116 | C257(1.19) | LDD0583 | [12] |
| LDCM0267 | AC147 | HCT 116 | C257(1.35) | LDD0584 | [12] |
| LDCM0268 | AC148 | HCT 116 | C257(1.40) | LDD0585 | [12] |
| LDCM0269 | AC149 | HCT 116 | C257(1.19) | LDD0586 | [12] |
| LDCM0270 | AC15 | HCT 116 | C160(1.10); C257(1.16) | LDD0587 | [12] |
| LDCM0271 | AC150 | HCT 116 | C257(1.66) | LDD0588 | [12] |
| LDCM0272 | AC151 | HCT 116 | C257(0.86) | LDD0589 | [12] |
| LDCM0273 | AC152 | HCT 116 | C257(1.19) | LDD0590 | [12] |
| LDCM0274 | AC153 | HCT 116 | C257(1.49) | LDD0591 | [12] |
| LDCM0621 | AC154 | HCT 116 | C257(1.72) | LDD2158 | [12] |
| LDCM0622 | AC155 | HCT 116 | C257(1.26) | LDD2159 | [12] |
| LDCM0623 | AC156 | HCT 116 | C257(1.09) | LDD2160 | [12] |
| LDCM0624 | AC157 | HCT 116 | C257(1.43) | LDD2161 | [12] |
| LDCM0276 | AC17 | HCT 116 | C257(1.34); C160(1.39) | LDD0593 | [12] |
| LDCM0277 | AC18 | HCT 116 | C160(1.02); C257(1.18) | LDD0594 | [12] |
| LDCM0278 | AC19 | HCT 116 | C160(1.00); C257(1.04) | LDD0595 | [12] |
| LDCM0279 | AC2 | HCT 116 | C160(0.98); C257(1.01); C129(1.15) | LDD0596 | [12] |
| LDCM0280 | AC20 | HCT 116 | C257(1.28); C160(1.30) | LDD0597 | [12] |
| LDCM0281 | AC21 | HCT 116 | C257(1.24); C160(1.31) | LDD0598 | [12] |
| LDCM0282 | AC22 | HCT 116 | C257(1.26); C160(1.28) | LDD0599 | [12] |
| LDCM0283 | AC23 | HCT 116 | C257(1.31); C160(1.51) | LDD0600 | [12] |
| LDCM0284 | AC24 | HCT 116 | C160(1.40); C257(1.40) | LDD0601 | [12] |
| LDCM0285 | AC25 | HCT 116 | C257(0.75); C160(0.78) | LDD0602 | [12] |
| LDCM0286 | AC26 | HCT 116 | C160(0.79); C257(0.82) | LDD0603 | [12] |
| LDCM0287 | AC27 | HCT 116 | C160(0.70); C257(0.74) | LDD0604 | [12] |
| LDCM0288 | AC28 | HCT 116 | C257(0.78); C160(0.85) | LDD0605 | [12] |
| LDCM0289 | AC29 | HCT 116 | C160(0.69); C257(0.79) | LDD0606 | [12] |
| LDCM0290 | AC3 | HCT 116 | C129(0.88); C257(0.94); C160(0.95) | LDD0607 | [12] |
| LDCM0291 | AC30 | HCT 116 | C160(0.74); C257(0.86) | LDD0608 | [12] |
| LDCM0292 | AC31 | HCT 116 | C257(0.94); C160(0.95) | LDD0609 | [12] |
| LDCM0293 | AC32 | HCT 116 | C257(0.66); C160(0.67) | LDD0610 | [12] |
| LDCM0294 | AC33 | HCT 116 | C257(0.62); C160(0.62) | LDD0611 | [12] |
| LDCM0295 | AC34 | HCT 116 | C160(0.67); C257(0.67) | LDD0612 | [12] |
| LDCM0296 | AC35 | HCT 116 | C257(1.26); C129(1.65); C160(1.67) | LDD0613 | [12] |
| LDCM0297 | AC36 | HCT 116 | C129(0.99); C160(1.04); C257(1.11) | LDD0614 | [12] |
| LDCM0298 | AC37 | HCT 116 | C160(0.75); C257(0.89); C129(0.99) | LDD0615 | [12] |
| LDCM0299 | AC38 | HCT 116 | C160(0.82); C257(0.98); C129(1.13) | LDD0616 | [12] |
| LDCM0300 | AC39 | HCT 116 | C160(1.10); C129(1.12); C257(1.35) | LDD0617 | [12] |
| LDCM0301 | AC4 | HCT 116 | C129(1.16); C257(1.20); C160(1.22) | LDD0618 | [12] |
| LDCM0302 | AC40 | HCT 116 | C160(0.90); C129(0.96); C257(0.99) | LDD0619 | [12] |
| LDCM0303 | AC41 | HCT 116 | C257(0.89); C129(0.91); C160(0.92) | LDD0620 | [12] |
| LDCM0304 | AC42 | HCT 116 | C129(0.82); C160(0.94); C257(1.03) | LDD0621 | [12] |
| LDCM0305 | AC43 | HCT 116 | C160(0.73); C257(0.83); C129(1.22) | LDD0622 | [12] |
| LDCM0306 | AC44 | HCT 116 | C129(0.82); C160(0.83); C257(0.86) | LDD0623 | [12] |
| LDCM0307 | AC45 | HCT 116 | C129(0.72); C160(0.72); C257(0.86) | LDD0624 | [12] |
| LDCM0308 | AC46 | HCT 116 | C257(0.87); C160(0.95); C129(1.04) | LDD0625 | [12] |
| LDCM0309 | AC47 | HCT 116 | C257(1.02); C160(1.13); C129(1.47) | LDD0626 | [12] |
| LDCM0310 | AC48 | HCT 116 | C129(1.00); C257(1.18); C160(1.43) | LDD0627 | [12] |
| LDCM0311 | AC49 | HCT 116 | C160(0.90); C129(0.91); C257(0.93) | LDD0628 | [12] |
| LDCM0312 | AC5 | HCT 116 | C257(0.99); C129(1.04); C160(1.08) | LDD0629 | [12] |
| LDCM0313 | AC50 | HCT 116 | C160(0.91); C257(0.96); C129(1.07) | LDD0630 | [12] |
| LDCM0314 | AC51 | HCT 116 | C160(1.04); C257(1.09); C129(1.21) | LDD0631 | [12] |
| LDCM0315 | AC52 | HCT 116 | C257(0.95); C129(0.97); C160(1.23) | LDD0632 | [12] |
| LDCM0316 | AC53 | HCT 116 | C129(0.82); C257(0.87); C160(0.92) | LDD0633 | [12] |
| LDCM0317 | AC54 | HCT 116 | C129(1.23); C257(1.30); C160(1.39) | LDD0634 | [12] |
| LDCM0318 | AC55 | HCT 116 | C257(0.85); C129(0.94); C160(0.96) | LDD0635 | [12] |
| LDCM0319 | AC56 | HCT 116 | C257(0.96); C129(1.02); C160(1.05) | LDD0636 | [12] |
| LDCM0320 | AC57 | HCT 116 | C160(1.13); C257(1.25) | LDD0637 | [12] |
| LDCM0321 | AC58 | HCT 116 | C160(1.25); C257(1.36) | LDD0638 | [12] |
| LDCM0322 | AC59 | HCT 116 | C160(0.80); C257(0.90) | LDD0639 | [12] |
| LDCM0323 | AC6 | HCT 116 | C160(1.00); C257(1.02) | LDD0640 | [12] |
| LDCM0324 | AC60 | HCT 116 | C160(0.76); C257(1.04) | LDD0641 | [12] |
| LDCM0325 | AC61 | HCT 116 | C257(1.12); C160(1.14) | LDD0642 | [12] |
| LDCM0326 | AC62 | HCT 116 | C160(0.84); C257(0.96) | LDD0643 | [12] |
| LDCM0327 | AC63 | HCT 116 | C257(0.91); C160(0.94) | LDD0644 | [12] |
| LDCM0328 | AC64 | HCT 116 | C160(0.83); C257(0.98) | LDD0645 | [12] |
| LDCM0329 | AC65 | HCT 116 | C160(0.93); C257(1.01) | LDD0646 | [12] |
| LDCM0330 | AC66 | HCT 116 | C160(1.06); C257(1.26) | LDD0647 | [12] |
| LDCM0331 | AC67 | HCT 116 | C257(0.83); C160(0.84) | LDD0648 | [12] |
| LDCM0332 | AC68 | HCT 116 | C257(1.07); C160(1.24) | LDD0649 | [12] |
| LDCM0333 | AC69 | HCT 116 | C257(1.41); C160(1.57) | LDD0650 | [12] |
| LDCM0334 | AC7 | HCT 116 | C160(0.95); C257(1.04) | LDD0651 | [12] |
| LDCM0335 | AC70 | HCT 116 | C257(1.19); C160(1.24) | LDD0652 | [12] |
| LDCM0336 | AC71 | HCT 116 | C257(1.32); C160(1.64) | LDD0653 | [12] |
| LDCM0337 | AC72 | HCT 116 | C257(1.08); C160(1.23) | LDD0654 | [12] |
| LDCM0338 | AC73 | HCT 116 | C257(1.05); C160(1.13) | LDD0655 | [12] |
| LDCM0339 | AC74 | HCT 116 | C257(1.27); C160(1.48) | LDD0656 | [12] |
| LDCM0340 | AC75 | HCT 116 | C257(1.07); C160(1.22) | LDD0657 | [12] |
| LDCM0341 | AC76 | HCT 116 | C257(1.33); C160(1.52) | LDD0658 | [12] |
| LDCM0342 | AC77 | HCT 116 | C257(1.00); C160(1.36) | LDD0659 | [12] |
| LDCM0343 | AC78 | HCT 116 | C257(0.97); C160(1.18) | LDD0660 | [12] |
| LDCM0344 | AC79 | HCT 116 | C257(1.04); C160(1.26) | LDD0661 | [12] |
| LDCM0345 | AC8 | HCT 116 | C160(0.88); C257(0.94) | LDD0662 | [12] |
| LDCM0346 | AC80 | HCT 116 | C257(0.96); C160(1.18) | LDD0663 | [12] |
| LDCM0347 | AC81 | HCT 116 | C257(1.30); C160(1.69) | LDD0664 | [12] |
| LDCM0348 | AC82 | HCT 116 | C257(0.98); C160(1.33) | LDD0665 | [12] |
| LDCM0349 | AC83 | HCT 116 | C160(0.63); C257(0.66); C129(0.96) | LDD0666 | [12] |
| LDCM0350 | AC84 | HCT 116 | C160(0.58); C129(0.64); C257(0.72) | LDD0667 | [12] |
| LDCM0351 | AC85 | HCT 116 | C160(0.65); C257(0.72); C129(0.73) | LDD0668 | [12] |
| LDCM0352 | AC86 | HCT 116 | C160(0.62); C257(0.72); C129(0.80) | LDD0669 | [12] |
| LDCM0353 | AC87 | HCT 116 | C129(0.81); C257(0.98); C160(1.03) | LDD0670 | [12] |
| LDCM0354 | AC88 | HCT 116 | C160(0.93); C129(1.01); C257(1.02) | LDD0671 | [12] |
| LDCM0355 | AC89 | HCT 116 | C160(0.65); C257(0.74); C129(0.76) | LDD0672 | [12] |
| LDCM0357 | AC90 | HCT 116 | C129(0.76); C160(0.83); C257(0.90) | LDD0674 | [12] |
| LDCM0358 | AC91 | HCT 116 | C160(0.69); C129(0.70); C257(0.77) | LDD0675 | [12] |
| LDCM0359 | AC92 | HCT 116 | C129(0.74); C257(0.79); C160(0.81) | LDD0676 | [12] |
| LDCM0360 | AC93 | HCT 116 | C129(0.64); C257(0.76); C160(0.82) | LDD0677 | [12] |
| LDCM0361 | AC94 | HCT 116 | C129(0.59); C160(0.64); C257(0.67) | LDD0678 | [12] |
| LDCM0362 | AC95 | HCT 116 | C129(0.74); C160(0.92); C257(0.93) | LDD0679 | [12] |
| LDCM0363 | AC96 | HCT 116 | C160(0.72); C129(0.74); C257(0.76) | LDD0680 | [12] |
| LDCM0364 | AC97 | HCT 116 | C160(0.61); C257(0.66); C129(0.68) | LDD0681 | [12] |
| LDCM0365 | AC98 | HCT 116 | C160(0.97); C257(0.99) | LDD0682 | [12] |
| LDCM0366 | AC99 | HCT 116 | C257(0.91); C160(1.19) | LDD0683 | [12] |
| LDCM0248 | AKOS034007472 | HCT 116 | C160(1.02); C257(1.00) | LDD0565 | [12] |
| LDCM0356 | AKOS034007680 | HCT 116 | C160(0.79); C257(0.91) | LDD0673 | [12] |
| LDCM0275 | AKOS034007705 | HCT 116 | C160(1.03); C257(1.09) | LDD0592 | [12] |
| LDCM0156 | Aniline | NCI-H1299 | 12.09 | LDD0403 | [2] |
| LDCM0020 | ARS-1620 | HCC44 | C160(1.15) | LDD2171 | [12] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C57(1.00) | LDD2091 | [9] |
| LDCM0108 | Chloroacetamide | HeLa | C57(0.00); H106(0.00) | LDD0222 | [11] |
| LDCM0367 | CL1 | HCT 116 | C129(0.71); C160(0.89); C257(0.92) | LDD0684 | [12] |
| LDCM0368 | CL10 | HCT 116 | C257(0.80); C160(0.86); C129(0.87) | LDD0685 | [12] |
| LDCM0369 | CL100 | HCT 116 | C129(0.89); C257(0.95); C160(0.97) | LDD0686 | [12] |
| LDCM0370 | CL101 | HCT 116 | C160(0.80); C257(0.84) | LDD0687 | [12] |
| LDCM0371 | CL102 | HCT 116 | C160(0.83); C257(0.93) | LDD0688 | [12] |
| LDCM0372 | CL103 | HCT 116 | C160(0.72); C257(0.82) | LDD0689 | [12] |
| LDCM0373 | CL104 | HCT 116 | C160(0.74); C257(0.85) | LDD0690 | [12] |
| LDCM0374 | CL105 | HCT 116 | C257(1.10); C160(1.29) | LDD0691 | [12] |
| LDCM0375 | CL106 | HCT 116 | C257(1.23); C160(1.56) | LDD0692 | [12] |
| LDCM0376 | CL107 | HCT 116 | C257(1.06); C160(1.23) | LDD0693 | [12] |
| LDCM0377 | CL108 | HCT 116 | C257(1.23); C160(1.29) | LDD0694 | [12] |
| LDCM0378 | CL109 | HCT 116 | C257(0.91); C160(0.96) | LDD0695 | [12] |
| LDCM0379 | CL11 | HCT 116 | C129(0.85); C257(0.90); C160(1.23) | LDD0696 | [12] |
| LDCM0380 | CL110 | HCT 116 | C257(1.27); C160(1.41) | LDD0697 | [12] |
| LDCM0381 | CL111 | HCT 116 | C257(1.02); C160(1.08) | LDD0698 | [12] |
| LDCM0382 | CL112 | HCT 116 | C160(0.96); C257(0.96) | LDD0699 | [12] |
| LDCM0383 | CL113 | HCT 116 | C257(0.82); C160(0.87) | LDD0700 | [12] |
| LDCM0384 | CL114 | HCT 116 | C257(0.69); C160(0.81) | LDD0701 | [12] |
| LDCM0385 | CL115 | HCT 116 | C257(0.67); C160(0.69) | LDD0702 | [12] |
| LDCM0386 | CL116 | HCT 116 | C160(0.61); C257(0.61) | LDD0703 | [12] |
| LDCM0387 | CL117 | HCT 116 | C257(1.14); C160(1.52); C129(1.75) | LDD0704 | [12] |
| LDCM0388 | CL118 | HCT 116 | C160(0.89); C257(0.97); C129(1.35) | LDD0705 | [12] |
| LDCM0389 | CL119 | HCT 116 | C129(0.80); C257(0.90); C160(0.93) | LDD0706 | [12] |
| LDCM0390 | CL12 | HCT 116 | C160(0.91); C129(0.97); C257(1.05) | LDD0707 | [12] |
| LDCM0391 | CL120 | HCT 116 | C160(0.85); C257(0.90); C129(0.91) | LDD0708 | [12] |
| LDCM0392 | CL121 | HCT 116 | C160(0.76); C257(0.80); C129(0.86) | LDD0709 | [12] |
| LDCM0393 | CL122 | HCT 116 | C129(0.78); C257(0.91); C160(0.93) | LDD0710 | [12] |
| LDCM0394 | CL123 | HCT 116 | C257(1.01); C129(1.04); C160(1.26) | LDD0711 | [12] |
| LDCM0395 | CL124 | HCT 116 | C129(0.89); C257(0.97); C160(1.04) | LDD0712 | [12] |
| LDCM0396 | CL125 | HCT 116 | C160(0.89); C257(1.03) | LDD0713 | [12] |
| LDCM0397 | CL126 | HCT 116 | C160(0.95); C257(1.06) | LDD0714 | [12] |
| LDCM0398 | CL127 | HCT 116 | C160(0.84); C257(0.98) | LDD0715 | [12] |
| LDCM0399 | CL128 | HCT 116 | C160(1.10); C257(1.10) | LDD0716 | [12] |
| LDCM0400 | CL13 | HCT 116 | C160(0.94); C129(0.98); C257(1.19) | LDD0717 | [12] |
| LDCM0401 | CL14 | HCT 116 | C257(0.52); C160(0.79); C129(0.98) | LDD0718 | [12] |
| LDCM0402 | CL15 | HCT 116 | C160(0.86); C257(0.96); C129(1.34) | LDD0719 | [12] |
| LDCM0403 | CL16 | HCT 116 | C160(0.86); C257(0.91) | LDD0720 | [12] |
| LDCM0404 | CL17 | HCT 116 | C129(0.81); C257(0.94); C160(1.06) | LDD0721 | [12] |
| LDCM0405 | CL18 | HCT 116 | C257(1.00); C160(1.00); C129(1.35) | LDD0722 | [12] |
| LDCM0406 | CL19 | HCT 116 | C160(0.94); C129(1.00); C257(1.12) | LDD0723 | [12] |
| LDCM0407 | CL2 | HCT 116 | C129(0.70); C160(0.91); C257(0.92) | LDD0724 | [12] |
| LDCM0408 | CL20 | HCT 116 | C160(0.83); C129(0.88); C257(0.91) | LDD0725 | [12] |
| LDCM0409 | CL21 | HCT 116 | C129(0.64); C160(1.01); C257(1.03) | LDD0726 | [12] |
| LDCM0410 | CL22 | HCT 116 | C160(0.81); C129(0.91); C257(1.01) | LDD0727 | [12] |
| LDCM0411 | CL23 | HCT 116 | C129(0.75); C257(0.89); C160(0.91) | LDD0728 | [12] |
| LDCM0412 | CL24 | HCT 116 | C129(1.03); C160(1.02); C257(0.95) | LDD0729 | [12] |
| LDCM0413 | CL25 | HCT 116 | C129(1.08); C160(1.06); C257(0.90) | LDD0730 | [12] |
| LDCM0414 | CL26 | HCT 116 | C129(1.17); C160(0.97); C257(0.92) | LDD0731 | [12] |
| LDCM0415 | CL27 | HCT 116 | C129(0.74); C160(0.99); C257(0.90) | LDD0732 | [12] |
| LDCM0416 | CL28 | HCT 116 | C129(0.79); C160(0.93); C257(0.86) | LDD0733 | [12] |
| LDCM0417 | CL29 | HCT 116 | C129(0.81); C160(0.83); C257(0.74) | LDD0734 | [12] |
| LDCM0418 | CL3 | HCT 116 | C129(0.69); C160(0.87); C257(0.95) | LDD0735 | [12] |
| LDCM0419 | CL30 | HCT 116 | C129(1.23); C160(1.20); C257(0.63) | LDD0736 | [12] |
| LDCM0420 | CL31 | HCT 116 | C129(1.08); C160(1.06); C257(1.00) | LDD0737 | [12] |
| LDCM0421 | CL32 | HCT 116 | C160(0.92); C257(0.90) | LDD0738 | [12] |
| LDCM0422 | CL33 | HCT 116 | C160(0.92); C257(0.93) | LDD0739 | [12] |
| LDCM0423 | CL34 | HCT 116 | C160(0.92); C257(0.97) | LDD0740 | [12] |
| LDCM0424 | CL35 | HCT 116 | C160(1.06); C257(0.92) | LDD0741 | [12] |
| LDCM0425 | CL36 | HCT 116 | C160(1.11); C257(1.26) | LDD0742 | [12] |
| LDCM0426 | CL37 | HCT 116 | C160(1.20); C257(1.19) | LDD0743 | [12] |
| LDCM0428 | CL39 | HCT 116 | C160(0.99); C257(0.99) | LDD0745 | [12] |
| LDCM0429 | CL4 | HCT 116 | C129(0.75); C160(0.89); C257(0.95) | LDD0746 | [12] |
| LDCM0430 | CL40 | HCT 116 | C160(1.00); C257(0.90) | LDD0747 | [12] |
| LDCM0431 | CL41 | HCT 116 | C160(0.81); C257(0.89) | LDD0748 | [12] |
| LDCM0432 | CL42 | HCT 116 | C160(1.30); C257(1.19) | LDD0749 | [12] |
| LDCM0433 | CL43 | HCT 116 | C160(0.91); C257(0.88) | LDD0750 | [12] |
| LDCM0434 | CL44 | HCT 116 | C160(1.01); C257(0.86) | LDD0751 | [12] |
| LDCM0435 | CL45 | HCT 116 | C160(1.08); C257(0.93) | LDD0752 | [12] |
| LDCM0436 | CL46 | HCT 116 | C129(1.32); C160(1.01); C257(0.92) | LDD0753 | [12] |
| LDCM0437 | CL47 | HCT 116 | C129(1.61); C160(1.22); C257(1.01) | LDD0754 | [12] |
| LDCM0438 | CL48 | HCT 116 | C129(1.57); C160(1.02); C257(0.96) | LDD0755 | [12] |
| LDCM0439 | CL49 | HCT 116 | C129(1.67); C160(1.11); C257(1.09) | LDD0756 | [12] |
| LDCM0440 | CL5 | HCT 116 | C129(1.16); C160(0.98); C257(1.00) | LDD0757 | [12] |
| LDCM0441 | CL50 | HCT 116 | C129(1.76); C160(1.05); C257(0.96) | LDD0758 | [12] |
| LDCM0442 | CL51 | HCT 116 | C129(1.87); C160(1.20); C257(1.10) | LDD0759 | [12] |
| LDCM0443 | CL52 | HCT 116 | C129(1.44); C160(1.02); C257(0.93) | LDD0760 | [12] |
| LDCM0444 | CL53 | HCT 116 | C129(1.43); C160(1.08); C257(1.02) | LDD0761 | [12] |
| LDCM0445 | CL54 | HCT 116 | C129(0.96); C160(1.09); C257(0.89) | LDD0762 | [12] |
| LDCM0446 | CL55 | HCT 116 | C129(1.11); C160(1.38); C257(1.04) | LDD0763 | [12] |
| LDCM0447 | CL56 | HCT 116 | C129(1.17); C160(1.28); C257(1.08) | LDD0764 | [12] |
| LDCM0448 | CL57 | HCT 116 | C129(1.52); C160(1.37); C257(1.03) | LDD0765 | [12] |
| LDCM0449 | CL58 | HCT 116 | C129(1.22); C160(1.03); C257(0.87) | LDD0766 | [12] |
| LDCM0450 | CL59 | HCT 116 | C129(1.28); C160(1.16); C257(1.04) | LDD0767 | [12] |
| LDCM0451 | CL6 | HCT 116 | C129(0.64); C160(0.75); C257(0.85) | LDD0768 | [12] |
| LDCM0452 | CL60 | HCT 116 | C129(1.69); C160(1.16); C257(0.92) | LDD0769 | [12] |
| LDCM0453 | CL61 | HCT 116 | C257(1.32) | LDD0770 | [12] |
| LDCM0454 | CL62 | HCT 116 | C257(0.87) | LDD0771 | [12] |
| LDCM0455 | CL63 | HCT 116 | C257(0.88) | LDD0772 | [12] |
| LDCM0456 | CL64 | HCT 116 | C257(1.15) | LDD0773 | [12] |
| LDCM0457 | CL65 | HCT 116 | C257(1.01) | LDD0774 | [12] |
| LDCM0458 | CL66 | HCT 116 | C257(0.95) | LDD0775 | [12] |
| LDCM0459 | CL67 | HCT 116 | C257(1.08) | LDD0776 | [12] |
| LDCM0460 | CL68 | HCT 116 | C257(1.30) | LDD0777 | [12] |
| LDCM0461 | CL69 | HCT 116 | C257(0.89) | LDD0778 | [12] |
| LDCM0462 | CL7 | HCT 116 | C129(0.79); C160(1.01); C257(1.02) | LDD0779 | [12] |
| LDCM0463 | CL70 | HCT 116 | C257(0.98) | LDD0780 | [12] |
| LDCM0464 | CL71 | HCT 116 | C257(0.95) | LDD0781 | [12] |
| LDCM0465 | CL72 | HCT 116 | C257(1.07) | LDD0782 | [12] |
| LDCM0466 | CL73 | HCT 116 | C257(1.14) | LDD0783 | [12] |
| LDCM0467 | CL74 | HCT 116 | C257(0.91) | LDD0784 | [12] |
| LDCM0469 | CL76 | HCT 116 | C160(1.22); C257(1.11) | LDD0786 | [12] |
| LDCM0470 | CL77 | HCT 116 | C160(1.18); C257(1.04) | LDD0787 | [12] |
| LDCM0471 | CL78 | HCT 116 | C160(1.12); C257(1.01) | LDD0788 | [12] |
| LDCM0472 | CL79 | HCT 116 | C160(1.05); C257(1.25) | LDD0789 | [12] |
| LDCM0473 | CL8 | HCT 116 | C129(0.62); C160(0.79); C257(0.97) | LDD0790 | [12] |
| LDCM0474 | CL80 | HCT 116 | C160(1.04); C257(1.06) | LDD0791 | [12] |
| LDCM0475 | CL81 | HCT 116 | C160(1.52); C257(1.42) | LDD0792 | [12] |
| LDCM0476 | CL82 | HCT 116 | C160(1.73); C257(1.42) | LDD0793 | [12] |
| LDCM0477 | CL83 | HCT 116 | C160(1.45); C257(1.38) | LDD0794 | [12] |
| LDCM0478 | CL84 | HCT 116 | C160(1.05); C257(1.12) | LDD0795 | [12] |
| LDCM0479 | CL85 | HCT 116 | C160(1.15); C257(1.10) | LDD0796 | [12] |
| LDCM0480 | CL86 | HCT 116 | C160(1.25); C257(1.21) | LDD0797 | [12] |
| LDCM0481 | CL87 | HCT 116 | C160(1.55); C257(1.43) | LDD0798 | [12] |
| LDCM0482 | CL88 | HCT 116 | C160(1.65); C257(1.59) | LDD0799 | [12] |
| LDCM0483 | CL89 | HCT 116 | C160(1.10); C257(1.03) | LDD0800 | [12] |
| LDCM0484 | CL9 | HCT 116 | C129(1.48); C160(0.88); C257(0.96) | LDD0801 | [12] |
| LDCM0485 | CL90 | HCT 116 | C160(1.40); C257(1.39) | LDD0802 | [12] |
| LDCM0486 | CL91 | HCT 116 | C129(1.47); C160(1.09); C257(1.11) | LDD0803 | [12] |
| LDCM0487 | CL92 | HCT 116 | C129(1.10); C160(1.27); C257(1.25) | LDD0804 | [12] |
| LDCM0488 | CL93 | HCT 116 | C129(1.02); C160(1.74); C257(1.50) | LDD0805 | [12] |
| LDCM0489 | CL94 | HCT 116 | C129(1.09); C160(0.81); C257(0.79) | LDD0806 | [12] |
| LDCM0490 | CL95 | HCT 116 | C129(0.88); C160(1.27); C257(1.22) | LDD0807 | [12] |
| LDCM0491 | CL96 | HCT 116 | C129(0.98); C160(0.98); C257(1.03) | LDD0808 | [12] |
| LDCM0492 | CL97 | HCT 116 | C129(1.46); C160(0.91); C257(0.93) | LDD0809 | [12] |
| LDCM0493 | CL98 | HCT 116 | C129(1.05); C160(1.08); C257(1.02) | LDD0810 | [12] |
| LDCM0494 | CL99 | HCT 116 | C129(0.99); C160(1.12); C257(1.15) | LDD0811 | [12] |
| LDCM0495 | E2913 | HEK-293T | C257(1.04); C129(0.89) | LDD1698 | [33] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [10] |
| LDCM0625 | F8 | Ramos | C57(0.91) | LDD2187 | [34] |
| LDCM0573 | Fragment11 | Ramos | C57(0.67) | LDD2190 | [34] |
| LDCM0586 | Fragment28 | Ramos | C57(0.52) | LDD2198 | [34] |
| LDCM0468 | Fragment33 | HCT 116 | C257(1.43) | LDD0785 | [12] |
| LDCM0427 | Fragment51 | HCT 116 | C160(1.25); C257(1.27) | LDD0744 | [12] |
| LDCM0569 | Fragment7 | Ramos | C57(1.94) | LDD2186 | [34] |
| LDCM0015 | HNE | MDA-MB-231 | C57(1.25) | LDD0346 | [34] |
| LDCM0107 | IAA | HeLa | H106(0.00); C57(0.00) | LDD0221 | [11] |
| LDCM0022 | KB02 | HCT 116 | C160(2.64); C257(2.39) | LDD0080 | [12] |
| LDCM0023 | KB03 | HCT 116 | C160(2.52); C257(1.00) | LDD0081 | [12] |
| LDCM0024 | KB05 | HCT 116 | C160(2.20); C257(1.41) | LDD0082 | [12] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C160(0.89) | LDD2121 | [9] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [11] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C57(0.39) | LDD2089 | [9] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C57(1.56) | LDD2090 | [9] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C57(1.22) | LDD2093 | [9] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C57(1.46) | LDD2099 | [9] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C57(0.57) | LDD2100 | [9] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C160(0.85) | LDD2101 | [9] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C57(0.59); C160(0.70) | LDD2104 | [9] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C57(0.94) | LDD2105 | [9] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C57(0.59); C160(0.69) | LDD2106 | [9] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C57(1.18) | LDD2107 | [9] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C57(1.01); C160(0.72) | LDD2108 | [9] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C57(0.86) | LDD2109 | [9] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C57(0.99) | LDD2110 | [9] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C57(1.02) | LDD2114 | [9] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C160(0.63) | LDD2115 | [9] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C57(1.19); C160(1.32) | LDD2118 | [9] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C57(1.12); C160(1.90) | LDD2119 | [9] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C57(0.77); C160(0.65) | LDD2120 | [9] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C57(1.15) | LDD2124 | [9] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C57(0.94) | LDD2125 | [9] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C57(0.94) | LDD2126 | [9] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C57(1.31) | LDD2127 | [9] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C57(1.55); C160(1.16) | LDD2129 | [9] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C57(0.67); C160(0.87) | LDD2133 | [9] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C57(0.59); C160(0.64) | LDD2134 | [9] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C57(1.87); C160(1.34) | LDD2135 | [9] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C57(1.00) | LDD2137 | [9] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C160(2.11); C57(1.93) | LDD1700 | [9] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C57(0.88) | LDD2140 | [9] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C57(1.02) | LDD2141 | [9] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C57(0.89); C160(0.70) | LDD2143 | [9] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C57(1.89); C160(1.99) | LDD2144 | [9] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C160(6.35) | LDD2145 | [9] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C57(1.46) | LDD2146 | [9] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C160(0.55) | LDD2150 | [9] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C57(0.74); C57(0.62) | LDD2206 | [35] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C57(0.88); C57(0.77) | LDD2207 | [35] |
| LDCM0021 | THZ1 | HCT 116 | C160(1.15) | LDD2173 | [12] |
| LDCM0110 | W12 | Hep-G2 | C160(0.75) | LDD0237 | [5] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Leucine-rich repeat serine/threonine-protein kinase 2 (LRRK2) | TKL Ser/Thr protein kinase family | Q5S007 | |||
| E3 ubiquitin-protein ligase TRIM23 (TRIM23) | Arf family | P36406 | |||
Transporter and channel
Other
References
