Details of the Target
General Information of Target
| Target ID | LDTP11911 | |||||
|---|---|---|---|---|---|---|
| Target Name | Golgi phosphoprotein 3 (GOLPH3) | |||||
| Gene Name | GOLPH3 | |||||
| Gene ID | 64083 | |||||
| Synonyms |
GPP34; Golgi phosphoprotein 3; Coat protein GPP34; Mitochondrial DNA absence factor; MIDAS |
|||||
| 3D Structure | ||||||
| Sequence |
MTTLTHRARRTEISKNSEKKMESEEDSNWEKSPDNEDSGDSKDIRLTLMEEVLLLGLKDK
EGYTSFWNDCISSGLRGGILIELAMRGRIYLEPPTMRKKRLLDRKVLLKSDSPTGDVLLD ETLKHIKATEPTETVQTWIELLTGETWNPFKLQYQLRNVRERIAKNLVEKGILTTEKQNF LLFDMTTHPVTNTTEKQRLVKKLQDSVLERWVNDPQRMDKRTLALLVLAHSSDVLENVFS SLTDDKYDVAMNRAKDLVELDPEVEGTKPSATEMIWAVLAAFNKS |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
GOLPH3/VPS74 family
|
|||||
| Subcellular location |
Golgi apparatus, Golgi stack membrane
|
|||||
| Function |
Phosphatidylinositol-4-phosphate-binding protein that links Golgi membranes to the cytoskeleton and may participate in the tensile force required for vesicle budding from the Golgi. Thereby, may play a role in Golgi membrane trafficking and could indirectly give its flattened shape to the Golgi apparatus. May also bind to the coatomer to regulate Golgi membrane trafficking. May play a role in anterograde transport from the Golgi to the plasma membrane and regulate secretion. Has also been involved in the control of the localization of Golgi enzymes through interaction with their cytoplasmic part. May play an indirect role in cell migration. Has also been involved in the modulation of mTOR signaling. May also be involved in the regulation of mitochondrial lipids biosynthesis.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STPyne Probe Info |
 |
K138(9.20) | LDD0277 | [1] | |
|
m-APA Probe Info |
 |
11.74 | LDD0403 | [2] | |
|
SAHA-CA-4PAP Probe Info |
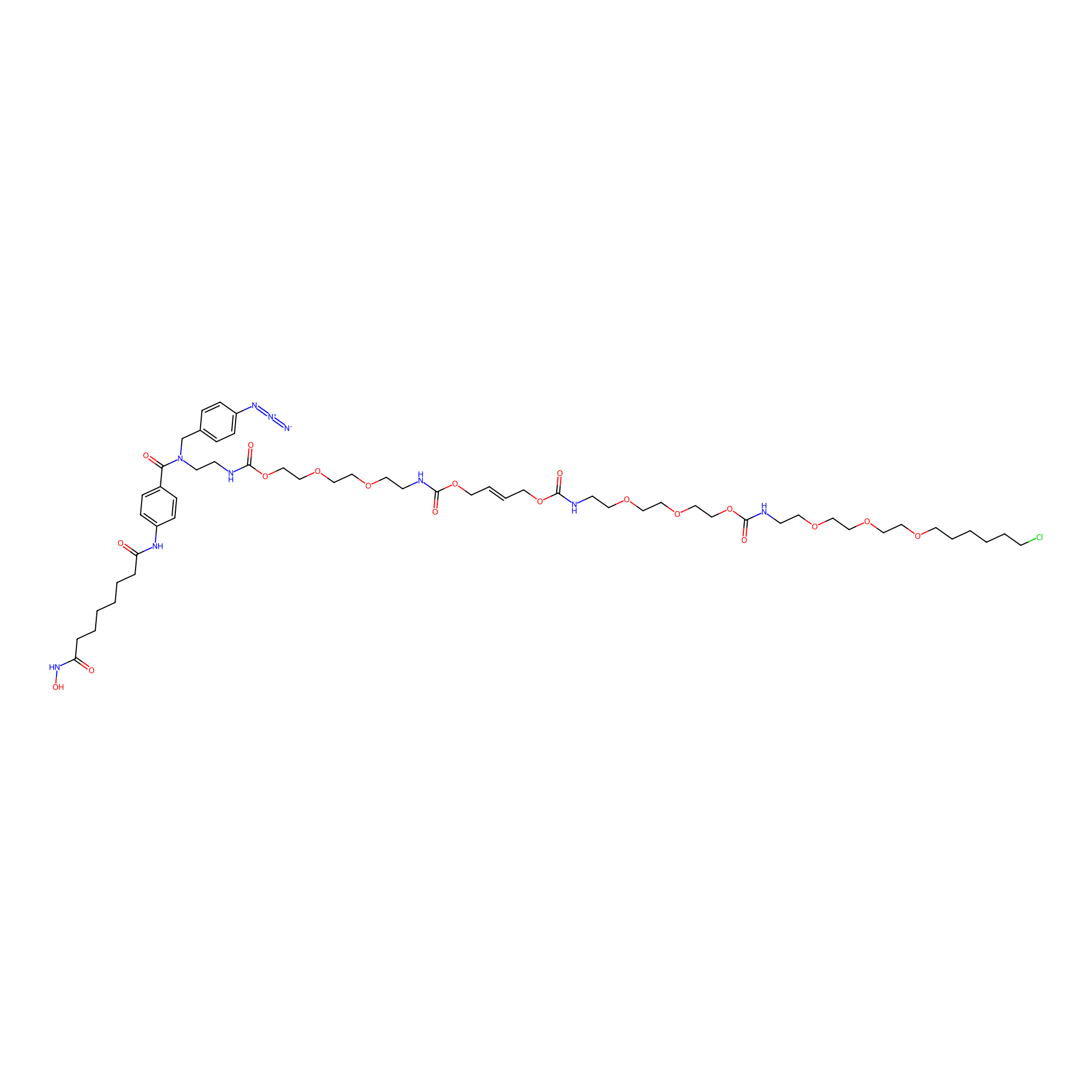 |
4.50 | LDD0360 | [3] | |
|
BTD Probe Info |
 |
C108(1.04) | LDD2093 | [4] | |
|
DBIA Probe Info |
 |
C84(1.14); C108(0.99) | LDD0078 | [5] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [6] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C84(0.00); C92(0.00); C108(0.00); C280(0.00) | LDD0038 | [7] | |
|
IA-alkyne Probe Info |
 |
C84(0.00); C92(0.00); C108(0.00); C280(0.00) | LDD0036 | [7] | |
|
Lodoacetamide azide Probe Info |
 |
C84(0.00); C108(0.00); C280(0.00) | LDD0037 | [7] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [8] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [8] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [8] | |
|
Phosphinate-6 Probe Info |
 |
C108(0.00); C122(0.00) | LDD0018 | [9] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [10] | |
|
W1 Probe Info |
 |
C108(0.00); C280(0.00) | LDD0236 | [11] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [12] | |
|
NAIA_5 Probe Info |
 |
C92(0.00); C108(0.00); C280(0.00); C122(0.00) | LDD2223 | [13] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C108(0.82) | LDD2117 | [4] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C108(1.38) | LDD2152 | [4] |
| LDCM0214 | AC1 | HCT 116 | C108(1.04); C84(1.07); C92(1.38) | LDD0531 | [5] |
| LDCM0215 | AC10 | HCT 116 | C108(1.20); C84(1.18); C92(1.00) | LDD0532 | [5] |
| LDCM0216 | AC100 | HCT 116 | C108(1.11); C84(1.04) | LDD0533 | [5] |
| LDCM0217 | AC101 | HCT 116 | C108(0.96); C84(1.09) | LDD0534 | [5] |
| LDCM0218 | AC102 | HCT 116 | C108(0.99); C84(0.94) | LDD0535 | [5] |
| LDCM0219 | AC103 | HCT 116 | C108(1.15); C84(1.12) | LDD0536 | [5] |
| LDCM0220 | AC104 | HCT 116 | C108(1.14); C84(0.95) | LDD0537 | [5] |
| LDCM0221 | AC105 | HCT 116 | C108(1.12); C84(1.37) | LDD0538 | [5] |
| LDCM0222 | AC106 | HCT 116 | C108(1.31); C84(1.12) | LDD0539 | [5] |
| LDCM0223 | AC107 | HCT 116 | C108(1.16); C84(1.05) | LDD0540 | [5] |
| LDCM0224 | AC108 | HCT 116 | C108(1.21); C84(1.25) | LDD0541 | [5] |
| LDCM0225 | AC109 | HCT 116 | C108(0.94); C84(1.11) | LDD0542 | [5] |
| LDCM0226 | AC11 | HCT 116 | C108(1.28); C84(1.18); C92(1.57) | LDD0543 | [5] |
| LDCM0227 | AC110 | HCT 116 | C108(1.13); C84(1.07) | LDD0544 | [5] |
| LDCM0228 | AC111 | HCT 116 | C108(1.09); C84(1.11) | LDD0545 | [5] |
| LDCM0229 | AC112 | HCT 116 | C108(1.10); C84(0.90) | LDD0546 | [5] |
| LDCM0230 | AC113 | HCT 116 | C108(1.07); C84(0.86) | LDD0547 | [5] |
| LDCM0231 | AC114 | HCT 116 | C108(1.03); C84(0.99) | LDD0548 | [5] |
| LDCM0232 | AC115 | HCT 116 | C108(1.12); C84(1.14) | LDD0549 | [5] |
| LDCM0233 | AC116 | HCT 116 | C108(1.27); C84(1.16) | LDD0550 | [5] |
| LDCM0234 | AC117 | HCT 116 | C108(1.09); C84(1.09) | LDD0551 | [5] |
| LDCM0235 | AC118 | HCT 116 | C108(1.07); C84(0.99) | LDD0552 | [5] |
| LDCM0236 | AC119 | HCT 116 | C108(1.14); C84(1.22) | LDD0553 | [5] |
| LDCM0237 | AC12 | HCT 116 | C108(1.04); C84(1.07); C92(1.19) | LDD0554 | [5] |
| LDCM0238 | AC120 | HCT 116 | C108(1.10); C84(1.05) | LDD0555 | [5] |
| LDCM0239 | AC121 | HCT 116 | C108(1.00); C84(1.00) | LDD0556 | [5] |
| LDCM0240 | AC122 | HCT 116 | C108(1.22); C84(1.20) | LDD0557 | [5] |
| LDCM0241 | AC123 | HCT 116 | C108(0.97); C84(0.97) | LDD0558 | [5] |
| LDCM0242 | AC124 | HCT 116 | C108(1.08); C84(1.05) | LDD0559 | [5] |
| LDCM0243 | AC125 | HCT 116 | C108(1.18); C84(1.18) | LDD0560 | [5] |
| LDCM0244 | AC126 | HCT 116 | C108(1.18); C84(1.19) | LDD0561 | [5] |
| LDCM0245 | AC127 | HCT 116 | C108(1.22); C84(1.05) | LDD0562 | [5] |
| LDCM0246 | AC128 | HCT 116 | C108(1.44); C84(1.16) | LDD0563 | [5] |
| LDCM0247 | AC129 | HCT 116 | C108(1.05); C84(0.88) | LDD0564 | [5] |
| LDCM0249 | AC130 | HCT 116 | C108(1.40); C84(1.11) | LDD0566 | [5] |
| LDCM0250 | AC131 | HCT 116 | C108(0.93); C84(0.91) | LDD0567 | [5] |
| LDCM0251 | AC132 | HCT 116 | C108(1.38); C84(1.13) | LDD0568 | [5] |
| LDCM0252 | AC133 | HCT 116 | C108(1.37); C84(1.08) | LDD0569 | [5] |
| LDCM0253 | AC134 | HCT 116 | C108(1.61); C84(1.12) | LDD0570 | [5] |
| LDCM0254 | AC135 | HCT 116 | C108(1.38); C84(1.06) | LDD0571 | [5] |
| LDCM0255 | AC136 | HCT 116 | C108(1.38); C84(1.05) | LDD0572 | [5] |
| LDCM0256 | AC137 | HCT 116 | C108(1.52); C84(1.08) | LDD0573 | [5] |
| LDCM0257 | AC138 | HCT 116 | C108(1.64); C84(1.08) | LDD0574 | [5] |
| LDCM0258 | AC139 | HCT 116 | C108(1.44); C84(1.16) | LDD0575 | [5] |
| LDCM0259 | AC14 | HCT 116 | C108(0.95); C84(1.33); C92(1.14) | LDD0576 | [5] |
| LDCM0260 | AC140 | HCT 116 | C108(1.53); C84(1.35) | LDD0577 | [5] |
| LDCM0261 | AC141 | HCT 116 | C108(1.63); C84(1.17) | LDD0578 | [5] |
| LDCM0262 | AC142 | HCT 116 | C108(1.76); C84(1.12) | LDD0579 | [5] |
| LDCM0263 | AC143 | HCT 116 | C108(0.92); C84(1.10); C92(1.03) | LDD0580 | [5] |
| LDCM0264 | AC144 | HCT 116 | C92(1.06); C108(1.26); C84(1.27) | LDD0581 | [5] |
| LDCM0265 | AC145 | HCT 116 | C92(0.95); C108(1.12); C84(1.16) | LDD0582 | [5] |
| LDCM0266 | AC146 | HCT 116 | C92(0.76); C108(1.00); C84(1.21) | LDD0583 | [5] |
| LDCM0267 | AC147 | HCT 116 | C84(1.09); C108(1.13); C92(1.23) | LDD0584 | [5] |
| LDCM0268 | AC148 | HCT 116 | C92(0.90); C108(1.38); C84(1.57) | LDD0585 | [5] |
| LDCM0269 | AC149 | HCT 116 | C92(0.86); C108(1.16); C84(1.24) | LDD0586 | [5] |
| LDCM0270 | AC15 | HCT 116 | C108(1.07); C92(1.14); C84(1.20) | LDD0587 | [5] |
| LDCM0271 | AC150 | HCT 116 | C92(0.94); C84(1.06); C108(1.32) | LDD0588 | [5] |
| LDCM0272 | AC151 | HCT 116 | C92(0.93); C84(1.15); C108(1.78) | LDD0589 | [5] |
| LDCM0273 | AC152 | HCT 116 | C92(0.94); C108(1.03); C84(1.30) | LDD0590 | [5] |
| LDCM0274 | AC153 | HCT 116 | C92(0.89); C108(1.47); C84(1.52) | LDD0591 | [5] |
| LDCM0621 | AC154 | HCT 116 | C108(1.11); C84(1.23); C92(1.01) | LDD2158 | [5] |
| LDCM0622 | AC155 | HCT 116 | C108(1.45); C84(1.13); C92(0.88) | LDD2159 | [5] |
| LDCM0623 | AC156 | HCT 116 | C108(0.90); C84(0.90); C92(0.91) | LDD2160 | [5] |
| LDCM0624 | AC157 | HCT 116 | C108(0.83); C84(0.85); C92(0.98) | LDD2161 | [5] |
| LDCM0276 | AC17 | HCT 116 | C84(1.06); C108(1.19) | LDD0593 | [5] |
| LDCM0277 | AC18 | HCT 116 | C84(0.87); C108(1.28) | LDD0594 | [5] |
| LDCM0278 | AC19 | HCT 116 | C84(1.08); C108(1.11) | LDD0595 | [5] |
| LDCM0279 | AC2 | HCT 116 | C92(1.06); C108(1.12); C84(1.16) | LDD0596 | [5] |
| LDCM0280 | AC20 | HCT 116 | C84(0.98); C108(1.16) | LDD0597 | [5] |
| LDCM0281 | AC21 | HCT 116 | C84(0.94); C108(1.06) | LDD0598 | [5] |
| LDCM0282 | AC22 | HCT 116 | C84(0.92); C108(1.08) | LDD0599 | [5] |
| LDCM0283 | AC23 | HCT 116 | C84(0.98); C108(1.08) | LDD0600 | [5] |
| LDCM0284 | AC24 | HCT 116 | C84(0.93); C108(1.08) | LDD0601 | [5] |
| LDCM0285 | AC25 | HCT 116 | C84(1.14); C108(1.15); C92(1.15) | LDD0602 | [5] |
| LDCM0286 | AC26 | HCT 116 | C108(1.21); C84(1.21); C92(1.23) | LDD0603 | [5] |
| LDCM0287 | AC27 | HCT 116 | C92(1.15); C108(1.17); C84(1.30) | LDD0604 | [5] |
| LDCM0288 | AC28 | HCT 116 | C84(1.15); C108(1.16); C92(1.24) | LDD0605 | [5] |
| LDCM0289 | AC29 | HCT 116 | C92(0.83); C84(1.28); C108(1.29) | LDD0606 | [5] |
| LDCM0290 | AC3 | HCT 116 | C108(1.12); C84(1.19); C92(1.24) | LDD0607 | [5] |
| LDCM0291 | AC30 | HCT 116 | C92(0.86); C108(1.25); C84(1.27) | LDD0608 | [5] |
| LDCM0292 | AC31 | HCT 116 | C92(1.17); C108(1.23); C84(1.27) | LDD0609 | [5] |
| LDCM0293 | AC32 | HCT 116 | C92(0.94); C108(1.34); C84(1.48) | LDD0610 | [5] |
| LDCM0294 | AC33 | HCT 116 | C92(1.05); C108(1.24); C84(1.40) | LDD0611 | [5] |
| LDCM0295 | AC34 | HCT 116 | C92(1.13); C84(1.37); C108(1.45) | LDD0612 | [5] |
| LDCM0296 | AC35 | HCT 116 | C108(0.86); C84(0.86) | LDD0613 | [5] |
| LDCM0297 | AC36 | HCT 116 | C108(0.92); C84(1.29) | LDD0614 | [5] |
| LDCM0298 | AC37 | HCT 116 | C108(0.97); C84(1.01) | LDD0615 | [5] |
| LDCM0299 | AC38 | HCT 116 | C108(0.96); C84(1.07) | LDD0616 | [5] |
| LDCM0300 | AC39 | HCT 116 | C108(1.07); C84(1.15) | LDD0617 | [5] |
| LDCM0301 | AC4 | HCT 116 | C108(1.04); C84(1.15); C92(1.56) | LDD0618 | [5] |
| LDCM0302 | AC40 | HCT 116 | C108(1.05); C84(1.11) | LDD0619 | [5] |
| LDCM0303 | AC41 | HCT 116 | C108(1.04); C84(1.24) | LDD0620 | [5] |
| LDCM0304 | AC42 | HCT 116 | C108(1.05); C84(1.09) | LDD0621 | [5] |
| LDCM0305 | AC43 | HCT 116 | C108(1.03); C84(1.25) | LDD0622 | [5] |
| LDCM0306 | AC44 | HCT 116 | C108(1.00); C84(1.06) | LDD0623 | [5] |
| LDCM0307 | AC45 | HCT 116 | C84(1.12); C108(1.16) | LDD0624 | [5] |
| LDCM0308 | AC46 | HCT 116 | C92(0.63); C108(1.10); C84(1.51) | LDD0625 | [5] |
| LDCM0309 | AC47 | HCT 116 | C92(0.86); C108(1.06); C84(1.41) | LDD0626 | [5] |
| LDCM0310 | AC48 | HCT 116 | C92(0.64); C108(1.11); C84(1.48) | LDD0627 | [5] |
| LDCM0311 | AC49 | HCT 116 | C92(0.58); C108(1.18); C84(1.43) | LDD0628 | [5] |
| LDCM0312 | AC5 | HCT 116 | C108(1.06); C84(1.32); C92(1.60) | LDD0629 | [5] |
| LDCM0313 | AC50 | HCT 116 | C92(0.57); C108(1.11); C84(1.45) | LDD0630 | [5] |
| LDCM0314 | AC51 | HCT 116 | C92(0.63); C108(0.94); C84(0.96) | LDD0631 | [5] |
| LDCM0315 | AC52 | HCT 116 | C92(0.62); C108(1.04); C84(1.34) | LDD0632 | [5] |
| LDCM0316 | AC53 | HCT 116 | C92(0.91); C108(0.96); C84(1.18) | LDD0633 | [5] |
| LDCM0317 | AC54 | HCT 116 | C92(0.71); C108(1.04); C84(1.44) | LDD0634 | [5] |
| LDCM0318 | AC55 | HCT 116 | C108(1.00); C84(1.22); C92(1.42) | LDD0635 | [5] |
| LDCM0319 | AC56 | HCT 116 | C92(0.81); C108(0.95); C84(1.65) | LDD0636 | [5] |
| LDCM0320 | AC57 | HCT 116 | C92(1.02); C84(1.28); C108(1.47) | LDD0637 | [5] |
| LDCM0321 | AC58 | HCT 116 | C92(1.01); C108(1.41); C84(1.41) | LDD0638 | [5] |
| LDCM0322 | AC59 | HCT 116 | C92(1.03); C108(1.57); C84(1.69) | LDD0639 | [5] |
| LDCM0323 | AC6 | HCT 116 | C92(0.79); C108(1.06); C84(1.24) | LDD0640 | [5] |
| LDCM0324 | AC60 | HCT 116 | C92(0.81); C108(1.33); C84(1.51) | LDD0641 | [5] |
| LDCM0325 | AC61 | HCT 116 | C92(0.86); C84(1.27); C108(1.35) | LDD0642 | [5] |
| LDCM0326 | AC62 | HCT 116 | C92(0.94); C84(1.25); C108(1.53) | LDD0643 | [5] |
| LDCM0327 | AC63 | HCT 116 | C92(0.99); C108(1.51); C84(1.53) | LDD0644 | [5] |
| LDCM0328 | AC64 | HCT 116 | C92(0.92); C84(1.61); C108(1.67) | LDD0645 | [5] |
| LDCM0329 | AC65 | HCT 116 | C92(1.05); C84(1.54); C108(1.83) | LDD0646 | [5] |
| LDCM0330 | AC66 | HCT 116 | C92(1.10); C108(1.58); C84(1.59) | LDD0647 | [5] |
| LDCM0331 | AC67 | HCT 116 | C92(0.90); C84(1.67); C108(1.90) | LDD0648 | [5] |
| LDCM0332 | AC68 | HCT 116 | C92(0.81); C84(1.06); C108(1.16) | LDD0649 | [5] |
| LDCM0333 | AC69 | HCT 116 | C92(0.85); C108(0.95); C84(1.03) | LDD0650 | [5] |
| LDCM0334 | AC7 | HCT 116 | C92(0.87); C108(1.11); C84(1.17) | LDD0651 | [5] |
| LDCM0335 | AC70 | HCT 116 | C92(0.68); C84(1.05); C108(1.19) | LDD0652 | [5] |
| LDCM0336 | AC71 | HCT 116 | C92(0.89); C108(1.00); C84(1.21) | LDD0653 | [5] |
| LDCM0337 | AC72 | HCT 116 | C92(0.77); C108(1.10); C84(1.21) | LDD0654 | [5] |
| LDCM0338 | AC73 | HCT 116 | C92(0.67); C108(1.32); C84(1.35) | LDD0655 | [5] |
| LDCM0339 | AC74 | HCT 116 | C92(1.15); C108(1.23); C84(1.82) | LDD0656 | [5] |
| LDCM0340 | AC75 | HCT 116 | C92(0.81); C84(1.33); C108(1.42) | LDD0657 | [5] |
| LDCM0341 | AC76 | HCT 116 | C92(1.05); C108(1.10); C84(1.14) | LDD0658 | [5] |
| LDCM0342 | AC77 | HCT 116 | C92(0.74); C108(1.06); C84(1.37) | LDD0659 | [5] |
| LDCM0343 | AC78 | HCT 116 | C92(0.69); C108(0.98); C84(1.19) | LDD0660 | [5] |
| LDCM0344 | AC79 | HCT 116 | C92(0.70); C108(1.14); C84(1.22) | LDD0661 | [5] |
| LDCM0345 | AC8 | HCT 116 | C92(0.90); C84(0.99); C108(1.19) | LDD0662 | [5] |
| LDCM0346 | AC80 | HCT 116 | C108(1.11); C92(1.13); C84(1.31) | LDD0663 | [5] |
| LDCM0347 | AC81 | HCT 116 | C92(0.82); C108(1.08); C84(1.18) | LDD0664 | [5] |
| LDCM0348 | AC82 | HCT 116 | C92(1.05); C84(1.32); C108(1.47) | LDD0665 | [5] |
| LDCM0349 | AC83 | HCT 116 | C84(1.27); C108(1.29) | LDD0666 | [5] |
| LDCM0350 | AC84 | HCT 116 | C84(1.36); C108(1.41) | LDD0667 | [5] |
| LDCM0351 | AC85 | HCT 116 | C84(1.09); C108(1.11) | LDD0668 | [5] |
| LDCM0352 | AC86 | HCT 116 | C108(1.17); C84(1.28) | LDD0669 | [5] |
| LDCM0353 | AC87 | HCT 116 | C108(1.05); C84(1.08) | LDD0670 | [5] |
| LDCM0354 | AC88 | HCT 116 | C84(1.08); C108(1.12) | LDD0671 | [5] |
| LDCM0355 | AC89 | HCT 116 | C108(1.20); C84(1.23) | LDD0672 | [5] |
| LDCM0357 | AC90 | HCT 116 | C108(1.01); C84(1.04) | LDD0674 | [5] |
| LDCM0358 | AC91 | HCT 116 | C108(1.23); C84(1.41) | LDD0675 | [5] |
| LDCM0359 | AC92 | HCT 116 | C108(1.23); C84(1.43) | LDD0676 | [5] |
| LDCM0360 | AC93 | HCT 116 | C84(0.96); C108(1.16) | LDD0677 | [5] |
| LDCM0361 | AC94 | HCT 116 | C84(0.94); C108(1.07) | LDD0678 | [5] |
| LDCM0362 | AC95 | HCT 116 | C108(1.09); C84(1.14) | LDD0679 | [5] |
| LDCM0363 | AC96 | HCT 116 | C108(1.12); C84(1.32) | LDD0680 | [5] |
| LDCM0364 | AC97 | HCT 116 | C84(1.09); C108(1.20) | LDD0681 | [5] |
| LDCM0365 | AC98 | HCT 116 | C84(1.32); C108(1.37) | LDD0682 | [5] |
| LDCM0366 | AC99 | HCT 116 | C84(0.94); C108(1.14) | LDD0683 | [5] |
| LDCM0248 | AKOS034007472 | HCT 116 | C108(1.23); C84(1.19); C92(1.29) | LDD0565 | [5] |
| LDCM0356 | AKOS034007680 | HCT 116 | C108(1.07); C84(1.22); C92(1.25) | LDD0673 | [5] |
| LDCM0275 | AKOS034007705 | HCT 116 | C92(0.86); C108(1.18); C84(1.63) | LDD0592 | [5] |
| LDCM0156 | Aniline | NCI-H1299 | 11.74 | LDD0403 | [2] |
| LDCM0020 | ARS-1620 | HCC44 | C84(1.14); C108(0.99) | LDD0078 | [5] |
| LDCM0630 | CCW28-3 | 231MFP | C84(0.25) | LDD2214 | [18] |
| LDCM0108 | Chloroacetamide | HeLa | C280(0.00); C108(0.00) | LDD0222 | [10] |
| LDCM0632 | CL-Sc | Hep-G2 | C108(20.00); C280(1.76); C108(0.42) | LDD2227 | [13] |
| LDCM0367 | CL1 | HCT 116 | C84(0.93); C92(0.95); C108(1.03) | LDD0684 | [5] |
| LDCM0368 | CL10 | HCT 116 | C92(0.75); C108(1.05); C84(1.28) | LDD0685 | [5] |
| LDCM0369 | CL100 | HCT 116 | C84(1.03); C108(1.08); C92(1.27) | LDD0686 | [5] |
| LDCM0370 | CL101 | HCT 116 | C92(0.91); C108(1.02); C84(1.28) | LDD0687 | [5] |
| LDCM0371 | CL102 | HCT 116 | C92(0.86); C108(0.89); C84(1.21) | LDD0688 | [5] |
| LDCM0372 | CL103 | HCT 116 | C108(0.94); C84(1.00); C92(1.23) | LDD0689 | [5] |
| LDCM0373 | CL104 | HCT 116 | C92(0.91); C108(1.03); C84(1.04) | LDD0690 | [5] |
| LDCM0374 | CL105 | HCT 116 | C84(1.15); C108(1.24) | LDD0691 | [5] |
| LDCM0375 | CL106 | HCT 116 | C84(1.29); C108(1.29) | LDD0692 | [5] |
| LDCM0376 | CL107 | HCT 116 | C84(1.26); C108(1.27) | LDD0693 | [5] |
| LDCM0377 | CL108 | HCT 116 | C84(1.19); C108(1.41) | LDD0694 | [5] |
| LDCM0378 | CL109 | HCT 116 | C84(1.11); C108(1.15) | LDD0695 | [5] |
| LDCM0379 | CL11 | HCT 116 | C108(0.81); C92(0.97); C84(1.24) | LDD0696 | [5] |
| LDCM0380 | CL110 | HCT 116 | C84(1.16); C108(1.20) | LDD0697 | [5] |
| LDCM0381 | CL111 | HCT 116 | C84(1.03); C108(1.12) | LDD0698 | [5] |
| LDCM0382 | CL112 | HCT 116 | C84(0.91); C108(0.99); C92(1.09) | LDD0699 | [5] |
| LDCM0383 | CL113 | HCT 116 | C92(0.88); C84(1.19); C108(1.38) | LDD0700 | [5] |
| LDCM0384 | CL114 | HCT 116 | C92(0.87); C84(1.14); C108(1.25) | LDD0701 | [5] |
| LDCM0385 | CL115 | HCT 116 | C84(1.09); C92(1.13); C108(1.23) | LDD0702 | [5] |
| LDCM0386 | CL116 | HCT 116 | C108(1.20); C84(1.31); C92(1.36) | LDD0703 | [5] |
| LDCM0387 | CL117 | HCT 116 | C108(1.22); C84(1.27) | LDD0704 | [5] |
| LDCM0388 | CL118 | HCT 116 | C108(1.09); C84(1.15) | LDD0705 | [5] |
| LDCM0389 | CL119 | HCT 116 | C84(1.07); C108(1.12) | LDD0706 | [5] |
| LDCM0390 | CL12 | HCT 116 | C92(0.80); C108(1.00); C84(1.37) | LDD0707 | [5] |
| LDCM0391 | CL120 | HCT 116 | C108(1.10); C84(1.13) | LDD0708 | [5] |
| LDCM0392 | CL121 | HCT 116 | C92(0.73); C108(0.86); C84(1.10) | LDD0709 | [5] |
| LDCM0393 | CL122 | HCT 116 | C92(0.71); C108(0.89); C84(1.53) | LDD0710 | [5] |
| LDCM0394 | CL123 | HCT 116 | C108(0.48); C92(0.67); C84(1.53) | LDD0711 | [5] |
| LDCM0395 | CL124 | HCT 116 | C92(0.66); C108(1.10); C84(1.73) | LDD0712 | [5] |
| LDCM0396 | CL125 | HCT 116 | C84(1.09); C92(1.16); C108(1.23) | LDD0713 | [5] |
| LDCM0397 | CL126 | HCT 116 | C92(0.77); C108(1.17); C84(1.54) | LDD0714 | [5] |
| LDCM0398 | CL127 | HCT 116 | C108(1.18); C84(1.22); C92(1.29) | LDD0715 | [5] |
| LDCM0399 | CL128 | HCT 116 | C92(1.04); C84(1.31); C108(1.43) | LDD0716 | [5] |
| LDCM0400 | CL13 | HCT 116 | C92(0.80); C108(0.97); C84(1.28) | LDD0717 | [5] |
| LDCM0401 | CL14 | HCT 116 | C92(0.82); C108(0.97); C84(1.12) | LDD0718 | [5] |
| LDCM0402 | CL15 | HCT 116 | C92(0.92); C108(0.96); C84(1.27) | LDD0719 | [5] |
| LDCM0403 | CL16 | HCT 116 | C92(1.05); C84(1.07); C108(1.40) | LDD0720 | [5] |
| LDCM0404 | CL17 | HCT 116 | C92(0.97); C108(0.99); C84(1.01) | LDD0721 | [5] |
| LDCM0405 | CL18 | HCT 116 | C92(0.94); C84(1.11); C108(1.24) | LDD0722 | [5] |
| LDCM0406 | CL19 | HCT 116 | C92(0.81); C108(1.05); C84(1.21) | LDD0723 | [5] |
| LDCM0407 | CL2 | HCT 116 | C92(0.86); C84(0.97); C108(1.08) | LDD0724 | [5] |
| LDCM0408 | CL20 | HCT 116 | C92(0.99); C108(1.24); C84(1.42) | LDD0725 | [5] |
| LDCM0409 | CL21 | HCT 116 | C92(0.87); C108(1.22); C84(1.26) | LDD0726 | [5] |
| LDCM0410 | CL22 | HCT 116 | C92(1.14); C108(1.26); C84(1.52) | LDD0727 | [5] |
| LDCM0411 | CL23 | HCT 116 | C92(0.90); C108(1.12); C84(1.15) | LDD0728 | [5] |
| LDCM0412 | CL24 | HCT 116 | C92(0.98); C84(1.13); C108(1.62) | LDD0729 | [5] |
| LDCM0413 | CL25 | HCT 116 | C108(1.29); C84(1.39); C92(0.70) | LDD0730 | [5] |
| LDCM0414 | CL26 | HCT 116 | C108(1.47); C84(1.17); C92(0.94) | LDD0731 | [5] |
| LDCM0415 | CL27 | HCT 116 | C108(1.41); C84(1.20); C92(0.91) | LDD0732 | [5] |
| LDCM0416 | CL28 | HCT 116 | C108(1.78); C84(1.13); C92(0.94) | LDD0733 | [5] |
| LDCM0417 | CL29 | HCT 116 | C108(1.22); C84(1.21); C92(0.89) | LDD0734 | [5] |
| LDCM0418 | CL3 | HCT 116 | C108(0.87); C84(0.90); C92(0.82) | LDD0735 | [5] |
| LDCM0419 | CL30 | HCT 116 | C108(1.61); C84(1.30); C92(0.92) | LDD0736 | [5] |
| LDCM0420 | CL31 | HCT 116 | C108(0.97); C84(1.11) | LDD0737 | [5] |
| LDCM0421 | CL32 | HCT 116 | C108(1.18); C84(1.16) | LDD0738 | [5] |
| LDCM0422 | CL33 | HCT 116 | C108(1.13); C84(1.51) | LDD0739 | [5] |
| LDCM0423 | CL34 | HCT 116 | C108(0.94); C84(1.35) | LDD0740 | [5] |
| LDCM0424 | CL35 | HCT 116 | C108(1.49); C84(1.27) | LDD0741 | [5] |
| LDCM0425 | CL36 | HCT 116 | C108(0.92); C84(1.29) | LDD0742 | [5] |
| LDCM0426 | CL37 | HCT 116 | C108(1.15); C84(1.26) | LDD0743 | [5] |
| LDCM0428 | CL39 | HCT 116 | C108(1.14); C84(1.29) | LDD0745 | [5] |
| LDCM0429 | CL4 | HCT 116 | C108(0.98); C84(1.09); C92(1.04) | LDD0746 | [5] |
| LDCM0430 | CL40 | HCT 116 | C108(1.02); C84(1.22) | LDD0747 | [5] |
| LDCM0431 | CL41 | HCT 116 | C108(0.81); C84(1.20) | LDD0748 | [5] |
| LDCM0432 | CL42 | HCT 116 | C108(0.75); C84(1.52) | LDD0749 | [5] |
| LDCM0433 | CL43 | HCT 116 | C108(1.08); C84(1.34) | LDD0750 | [5] |
| LDCM0434 | CL44 | HCT 116 | C108(1.33); C84(1.23) | LDD0751 | [5] |
| LDCM0435 | CL45 | HCT 116 | C108(1.30); C84(1.40) | LDD0752 | [5] |
| LDCM0436 | CL46 | HCT 116 | C108(0.94); C84(0.81) | LDD0753 | [5] |
| LDCM0437 | CL47 | HCT 116 | C108(0.98); C84(0.65) | LDD0754 | [5] |
| LDCM0438 | CL48 | HCT 116 | C108(0.94); C84(0.65) | LDD0755 | [5] |
| LDCM0439 | CL49 | HCT 116 | C108(0.96); C84(0.76) | LDD0756 | [5] |
| LDCM0440 | CL5 | HCT 116 | C108(1.01); C84(1.12); C92(0.88) | LDD0757 | [5] |
| LDCM0441 | CL50 | HCT 116 | C108(0.93); C84(0.71) | LDD0758 | [5] |
| LDCM0442 | CL51 | HCT 116 | C108(0.99); C84(0.81) | LDD0759 | [5] |
| LDCM0443 | CL52 | HCT 116 | C108(1.05); C84(0.73) | LDD0760 | [5] |
| LDCM0444 | CL53 | HCT 116 | C108(0.87); C84(0.62) | LDD0761 | [5] |
| LDCM0445 | CL54 | HCT 116 | C108(0.91); C84(0.72) | LDD0762 | [5] |
| LDCM0446 | CL55 | HCT 116 | C108(0.94); C84(0.71) | LDD0763 | [5] |
| LDCM0447 | CL56 | HCT 116 | C108(1.01); C84(0.85) | LDD0764 | [5] |
| LDCM0448 | CL57 | HCT 116 | C108(0.94); C84(0.66) | LDD0765 | [5] |
| LDCM0449 | CL58 | HCT 116 | C108(0.96); C84(0.68) | LDD0766 | [5] |
| LDCM0450 | CL59 | HCT 116 | C108(1.00); C84(0.78) | LDD0767 | [5] |
| LDCM0451 | CL6 | HCT 116 | C108(1.12); C84(1.06); C92(0.74) | LDD0768 | [5] |
| LDCM0452 | CL60 | HCT 116 | C108(1.06); C84(0.79) | LDD0769 | [5] |
| LDCM0453 | CL61 | HCT 116 | C108(1.04); C84(1.11); C92(0.70) | LDD0770 | [5] |
| LDCM0454 | CL62 | HCT 116 | C108(0.88); C84(1.08); C92(0.79) | LDD0771 | [5] |
| LDCM0455 | CL63 | HCT 116 | C108(0.83); C84(1.11); C92(0.76) | LDD0772 | [5] |
| LDCM0456 | CL64 | HCT 116 | C108(1.01); C84(1.31); C92(0.91) | LDD0773 | [5] |
| LDCM0457 | CL65 | HCT 116 | C108(1.03); C84(1.05); C92(0.86) | LDD0774 | [5] |
| LDCM0458 | CL66 | HCT 116 | C108(1.03); C84(1.42); C92(0.94) | LDD0775 | [5] |
| LDCM0459 | CL67 | HCT 116 | C108(0.84); C84(1.21); C92(0.93) | LDD0776 | [5] |
| LDCM0460 | CL68 | HCT 116 | C108(0.80); C84(1.21); C92(0.70) | LDD0777 | [5] |
| LDCM0461 | CL69 | HCT 116 | C108(0.97); C84(1.13); C92(0.55) | LDD0778 | [5] |
| LDCM0462 | CL7 | HCT 116 | C108(1.07); C84(1.22); C92(0.91) | LDD0779 | [5] |
| LDCM0463 | CL70 | HCT 116 | C108(0.89); C84(1.15); C92(0.65) | LDD0780 | [5] |
| LDCM0464 | CL71 | HCT 116 | C108(1.00); C84(1.20); C92(0.58) | LDD0781 | [5] |
| LDCM0465 | CL72 | HCT 116 | C108(0.98); C84(1.12); C92(0.56) | LDD0782 | [5] |
| LDCM0466 | CL73 | HCT 116 | C108(0.95); C84(1.21); C92(0.78) | LDD0783 | [5] |
| LDCM0467 | CL74 | HCT 116 | C108(0.82); C84(1.20); C92(0.73) | LDD0784 | [5] |
| LDCM0469 | CL76 | HCT 116 | C108(0.94); C84(1.03) | LDD0786 | [5] |
| LDCM0470 | CL77 | HCT 116 | C108(0.81); C84(0.95) | LDD0787 | [5] |
| LDCM0471 | CL78 | HCT 116 | C108(0.98); C84(1.02) | LDD0788 | [5] |
| LDCM0472 | CL79 | HCT 116 | C108(1.01); C84(1.20) | LDD0789 | [5] |
| LDCM0473 | CL8 | HCT 116 | C108(0.93); C84(1.37); C92(0.59) | LDD0790 | [5] |
| LDCM0474 | CL80 | HCT 116 | C108(1.02); C84(1.12) | LDD0791 | [5] |
| LDCM0475 | CL81 | HCT 116 | C108(0.96); C84(1.12) | LDD0792 | [5] |
| LDCM0476 | CL82 | HCT 116 | C108(1.03); C84(1.42) | LDD0793 | [5] |
| LDCM0477 | CL83 | HCT 116 | C108(0.89); C84(1.39) | LDD0794 | [5] |
| LDCM0478 | CL84 | HCT 116 | C108(1.03); C84(1.45) | LDD0795 | [5] |
| LDCM0479 | CL85 | HCT 116 | C108(1.03); C84(1.23) | LDD0796 | [5] |
| LDCM0480 | CL86 | HCT 116 | C108(0.98); C84(1.02) | LDD0797 | [5] |
| LDCM0481 | CL87 | HCT 116 | C108(0.95); C84(1.28) | LDD0798 | [5] |
| LDCM0482 | CL88 | HCT 116 | C108(0.89); C84(1.26) | LDD0799 | [5] |
| LDCM0483 | CL89 | HCT 116 | C108(0.86); C84(1.73) | LDD0800 | [5] |
| LDCM0484 | CL9 | HCT 116 | C108(1.07); C84(1.18); C92(0.77) | LDD0801 | [5] |
| LDCM0485 | CL90 | HCT 116 | C108(0.82); C84(1.17) | LDD0802 | [5] |
| LDCM0486 | CL91 | HCT 116 | C108(1.11); C84(1.11); C92(0.86) | LDD0803 | [5] |
| LDCM0487 | CL92 | HCT 116 | C108(0.97); C84(1.11); C92(1.05) | LDD0804 | [5] |
| LDCM0488 | CL93 | HCT 116 | C108(1.05); C84(1.01); C92(1.38) | LDD0805 | [5] |
| LDCM0489 | CL94 | HCT 116 | C108(0.89); C84(1.15); C92(2.09) | LDD0806 | [5] |
| LDCM0490 | CL95 | HCT 116 | C108(0.97); C84(1.14); C92(1.11) | LDD0807 | [5] |
| LDCM0491 | CL96 | HCT 116 | C108(1.05); C84(1.18); C92(1.03) | LDD0808 | [5] |
| LDCM0492 | CL97 | HCT 116 | C108(1.16); C84(1.23); C92(0.99) | LDD0809 | [5] |
| LDCM0493 | CL98 | HCT 116 | C108(1.09); C84(1.10); C92(1.02) | LDD0810 | [5] |
| LDCM0494 | CL99 | HCT 116 | C108(1.03); C84(1.10); C92(1.18) | LDD0811 | [5] |
| LDCM0634 | CY-0357 | Hep-G2 | C108(0.78) | LDD2228 | [13] |
| LDCM0495 | E2913 | HEK-293T | C108(1.12); C84(1.00); C92(0.98); C280(0.97) | LDD1698 | [19] |
| LDCM0625 | F8 | Ramos | C108(1.36); C280(1.05); C84(1.35) | LDD2187 | [20] |
| LDCM0572 | Fragment10 | Ramos | C108(1.13) | LDD2189 | [20] |
| LDCM0573 | Fragment11 | Ramos | C108(0.90); C280(0.73); C84(0.55) | LDD2190 | [20] |
| LDCM0574 | Fragment12 | Ramos | C108(1.07) | LDD2191 | [20] |
| LDCM0575 | Fragment13 | Ramos | C108(1.06) | LDD2192 | [20] |
| LDCM0576 | Fragment14 | Ramos | C108(0.69); C280(0.76); C84(1.27) | LDD2193 | [20] |
| LDCM0579 | Fragment20 | Ramos | C108(0.98) | LDD2194 | [20] |
| LDCM0580 | Fragment21 | Ramos | C108(0.78) | LDD2195 | [20] |
| LDCM0582 | Fragment23 | Ramos | C108(0.79) | LDD2196 | [20] |
| LDCM0578 | Fragment27 | Ramos | C108(0.99) | LDD2197 | [20] |
| LDCM0586 | Fragment28 | Ramos | C108(0.87); C280(0.68) | LDD2198 | [20] |
| LDCM0588 | Fragment30 | Ramos | C108(1.15) | LDD2199 | [20] |
| LDCM0589 | Fragment31 | Ramos | C108(1.10) | LDD2200 | [20] |
| LDCM0590 | Fragment32 | Ramos | C108(1.56) | LDD2201 | [20] |
| LDCM0468 | Fragment33 | HCT 116 | C108(0.92); C84(1.31); C92(0.61) | LDD0785 | [5] |
| LDCM0596 | Fragment38 | Ramos | C108(0.97) | LDD2203 | [20] |
| LDCM0566 | Fragment4 | Ramos | C108(1.03); C280(0.94); C84(0.60) | LDD2184 | [20] |
| LDCM0427 | Fragment51 | HCT 116 | C108(1.11); C84(1.51) | LDD0744 | [5] |
| LDCM0610 | Fragment52 | Ramos | C108(0.95) | LDD2204 | [20] |
| LDCM0614 | Fragment56 | Ramos | C108(1.11) | LDD2205 | [20] |
| LDCM0569 | Fragment7 | Ramos | C108(0.84); C280(0.70); C84(0.66) | LDD2186 | [20] |
| LDCM0571 | Fragment9 | Ramos | C108(1.11) | LDD2188 | [20] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [10] |
| LDCM0022 | KB02 | HCT 116 | C84(1.06) | LDD0080 | [5] |
| LDCM0023 | KB03 | HCT 116 | C84(2.23) | LDD0081 | [5] |
| LDCM0024 | KB05 | HCT 116 | C84(1.79) | LDD0082 | [5] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [10] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C108(1.04) | LDD2093 | [4] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C108(0.87) | LDD2096 | [4] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C108(1.03) | LDD2099 | [4] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C108(1.04) | LDD2109 | [4] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C108(1.40) | LDD2111 | [4] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C108(0.77) | LDD2123 | [4] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C108(0.91) | LDD2125 | [4] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C108(1.38) | LDD2127 | [4] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C108(1.20) | LDD2129 | [4] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C108(1.13) | LDD2135 | [4] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C108(1.93) | LDD2136 | [4] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C108(1.01) | LDD2137 | [4] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C108(0.94) | LDD2140 | [4] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C108(0.94) | LDD2146 | [4] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C108(2.99) | LDD2206 | [21] |
| LDCM0131 | RA190 | MM1.R | C84(1.25) | LDD0304 | [22] |
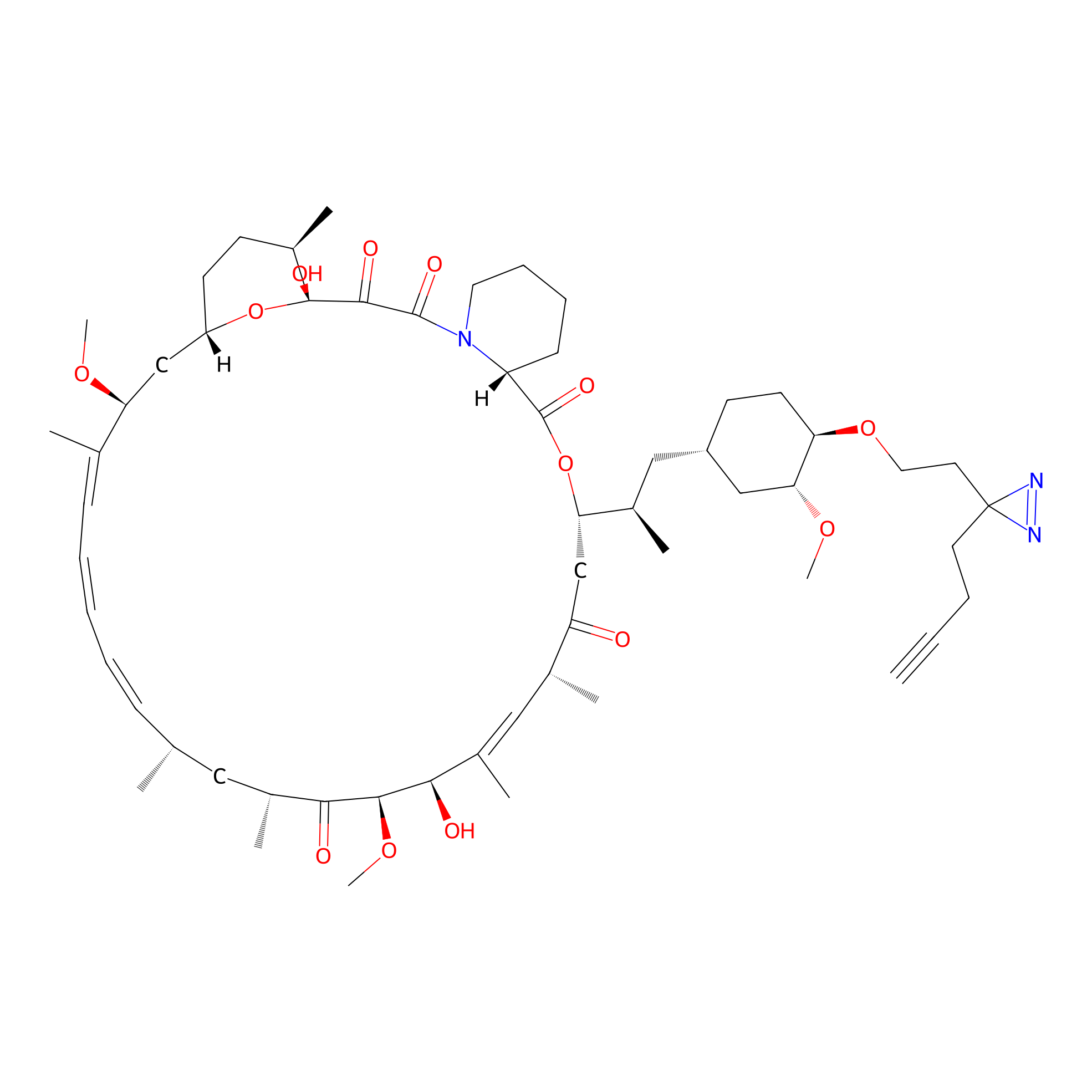
| LDCM0090 | Rapamycin | JHH-7 | 4.67 | LDD0213 | [16] |
| LDCM0096 | SAHA | K562 | 4.50 | LDD0360 | [3] |
| LDCM0021 | THZ1 | HCT 116 | C108(1.21); C84(1.07) | LDD2173 | [5] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| ADP-ribosylation factor-like protein 6-interacting protein 1 (ARL6IP1) | ARL6ip family | Q15041 | |||
References