Details of the Target
General Information of Target
| Target ID | LDTP11844 | |||||
|---|---|---|---|---|---|---|
| Target Name | Protein spinster homolog 1 (SPNS1) | |||||
| Gene Name | SPNS1 | |||||
| Gene ID | 83985 | |||||
| Synonyms |
SPIN1; Protein spinster homolog 1; HSpin1; SPNS1; Spinster-like protein 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MLPGCIFLMILLIPQVKEKFILGVEGQQLVRPKKLPLIQKRDTGHTHDDDILKTYEEELL
YEIKLNRKTLVLHLLRSREFLGSNYSETFYSMKGEAFTRHPQIMDHCFYQGSIVHEYDSA ASISTCNGLRGFFRINDQRYLIEPVKYSDEGEHLVFKYNLRVPYGANYSCTELNFTRKTV PGDNESEEDSKIKGIHDEKYVELFIVADDTVYRRNGHPHNKLRNRIWGMVNFVNMIYKTL NIHVTLVGIEIWTHEDKIELYSNIETTLLRFSFWQEKILKTRKDFDHVVLLSGKWLYSHV QGISYPGGMCLPYYSTSIIKDLLPDTNIIANRMAHQLGHNLGMQHDEFPCTCPSGKCVMD SDGSIPALKFSKCSQNQYHQYLKDYKPTCMLNIPFPYNFHDFQFCGNKKLDEGEECDCGP AQECTNPCCDAHTCVLKPGFTCAEGECCESCQIKKAGSICRPAKDECDFPEMCTGHSPAC PKDQFRVNGFPCKNSEGYCFMGKCPTREDQCSELFDDEAIESHDICYKMNTKGNKFGYCK NKENRFLPCEEKDVRCGKIYCTGGELSSLLGEDKTYHLKDPQKNATVKCKTIFLYHDSTD IGLVASGTKCGEGMVCNNGECLNMEKVYISTNCPSQCNENPVDGHGLQCHCEEGQAPVAC EETLHVTNITILVVVLVLVIVGIGVLILLVRYRKCIKLKQVQSPPTETLGVENKGYFGDE QQIRTEPILPEIHFLNKPASKDSRGIADPNQSAK |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Major facilitator superfamily, Spinster (TC 2.A.1.49) family
|
|||||
| Subcellular location |
Lysosome membrane
|
|||||
| Function |
Plays a critical role in the phospholipid salvage pathway from lysosomes to the cytosol. Mediates the rate-limiting, proton-dependent, lysosomal efflux of lysophospholipids, which can then be reacylated by acyltransferases in the endoplasmic reticulum to form phospholipids. Selective for zwitterionic headgroups such as lysophosphatidylcholine (LPC) and lysophosphatidylethanolamine (LPE), can also transport lysophosphatidylglycerol (LPG), but not other anionic lysophospholipids, sphingosine, nor sphingomyelin. Transports lysophospholipids with saturated, monounsaturated, and polyunsaturated fatty acids, such as 1-hexadecanoyl-sn-glycero-3-phosphocholine, 1-(9Z-octadecenoyl)-sn-glycero-3-phosphocholine and 1-(4Z,7Z,10Z,13Z,16Z,19Z-docosahexaenoyl)-sn-glycero-3-phosphocholine, respectively. Can also transport lysoplasmalogen (LPC with a fatty alcohol) such as 1-(1Z-hexadecenyl)-sn-glycero-3-phosphocholine. Lysosomal LPC could function as intracellular signaling messenger. Essential player in lysosomal homeostasis. Crucial for cell survival under conditions of nutrient limitation. May be involved in necrotic or autophagic cell death.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
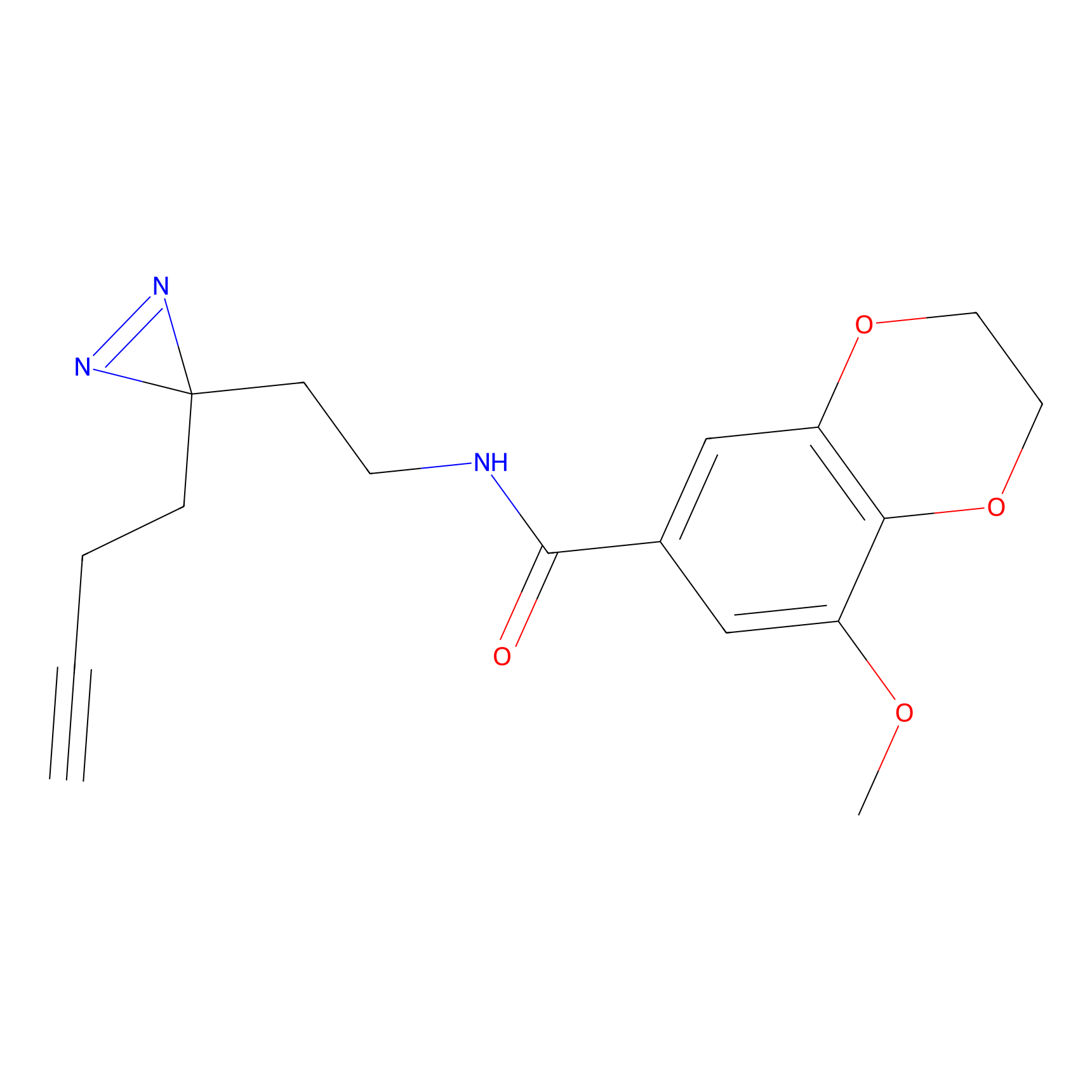
DBIA Probe Info |
 |
C96(1.61) | LDD3442 | [1] | |
|
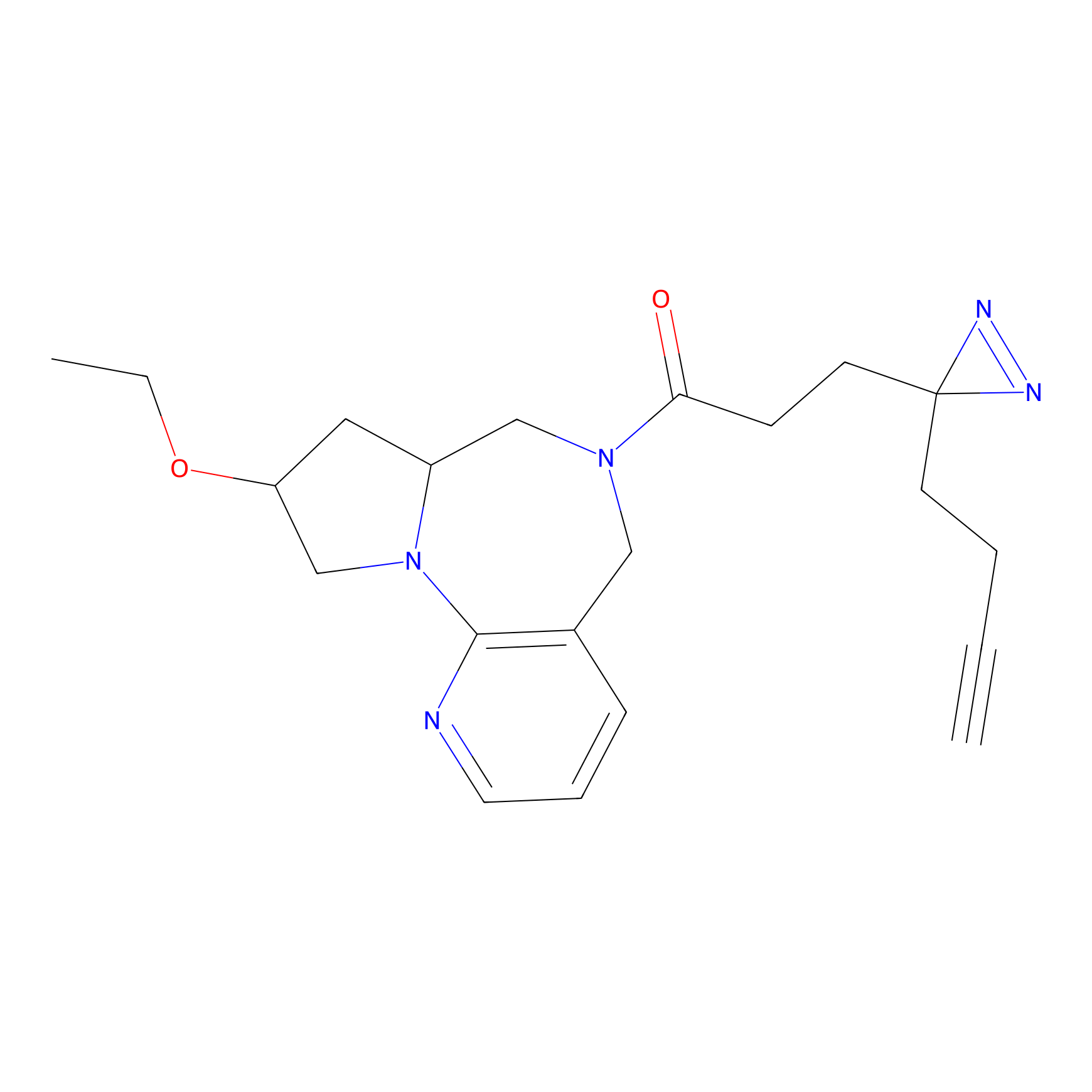
AOyne Probe Info |
 |
15.00 | LDD0443 | [2] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0237 | AC12 | HEK-293T | C376(1.00) | LDD1510 | [8] |
| LDCM0270 | AC15 | HEK-293T | C376(1.41) | LDD1513 | [8] |
| LDCM0280 | AC20 | HEK-293T | C376(0.85) | LDD1519 | [8] |
| LDCM0283 | AC23 | HEK-293T | C376(1.09) | LDD1522 | [8] |
| LDCM0288 | AC28 | HEK-293T | C376(1.23) | LDD1527 | [8] |
| LDCM0292 | AC31 | HEK-293T | C376(1.08) | LDD1531 | [8] |
| LDCM0297 | AC36 | HEK-293T | C376(0.90) | LDD1536 | [8] |
| LDCM0300 | AC39 | HEK-293T | C376(1.14) | LDD1539 | [8] |
| LDCM0301 | AC4 | HEK-293T | C376(0.89) | LDD1540 | [8] |
| LDCM0306 | AC44 | HEK-293T | C376(0.82) | LDD1545 | [8] |
| LDCM0309 | AC47 | HEK-293T | C376(0.99) | LDD1548 | [8] |
| LDCM0315 | AC52 | HEK-293T | C376(0.83) | LDD1554 | [8] |
| LDCM0318 | AC55 | HEK-293T | C376(1.14) | LDD1557 | [8] |
| LDCM0324 | AC60 | HEK-293T | C376(1.13) | LDD1563 | [8] |
| LDCM0327 | AC63 | HEK-293T | C376(1.12) | LDD1566 | [8] |
| LDCM0334 | AC7 | HEK-293T | C376(1.04) | LDD1568 | [8] |
| LDCM0379 | CL11 | HEK-293T | C376(1.47) | LDD1583 | [8] |
| LDCM0408 | CL20 | HEK-293T | C376(1.21) | LDD1612 | [8] |
| LDCM0411 | CL23 | HEK-293T | C376(1.08) | LDD1615 | [8] |
| LDCM0421 | CL32 | HEK-293T | C376(1.10) | LDD1625 | [8] |
| LDCM0424 | CL35 | HEK-293T | C376(0.92) | LDD1628 | [8] |
| LDCM0434 | CL44 | HEK-293T | C376(0.81) | LDD1638 | [8] |
| LDCM0437 | CL47 | HEK-293T | C376(1.15) | LDD1641 | [8] |
| LDCM0447 | CL56 | HEK-293T | C376(1.15) | LDD1650 | [8] |
| LDCM0450 | CL59 | HEK-293T | C376(0.99) | LDD1653 | [8] |
| LDCM0460 | CL68 | HEK-293T | C376(0.97) | LDD1663 | [8] |
| LDCM0464 | CL71 | HEK-293T | C376(1.23) | LDD1667 | [8] |
| LDCM0473 | CL8 | HEK-293T | C376(1.19) | LDD1676 | [8] |
| LDCM0474 | CL80 | HEK-293T | C376(0.82) | LDD1677 | [8] |
| LDCM0477 | CL83 | HEK-293T | C376(1.38) | LDD1680 | [8] |
| LDCM0487 | CL92 | HEK-293T | C376(1.11) | LDD1690 | [8] |
| LDCM0490 | CL95 | HEK-293T | C376(1.26) | LDD1693 | [8] |
| LDCM0022 | KB02 | HuH-1 | C96(1.42) | LDD2374 | [1] |
| LDCM0023 | KB03 | HuH-1 | C96(1.84) | LDD2791 | [1] |
| LDCM0024 | KB05 | SNU-423 | C96(1.61) | LDD3442 | [1] |
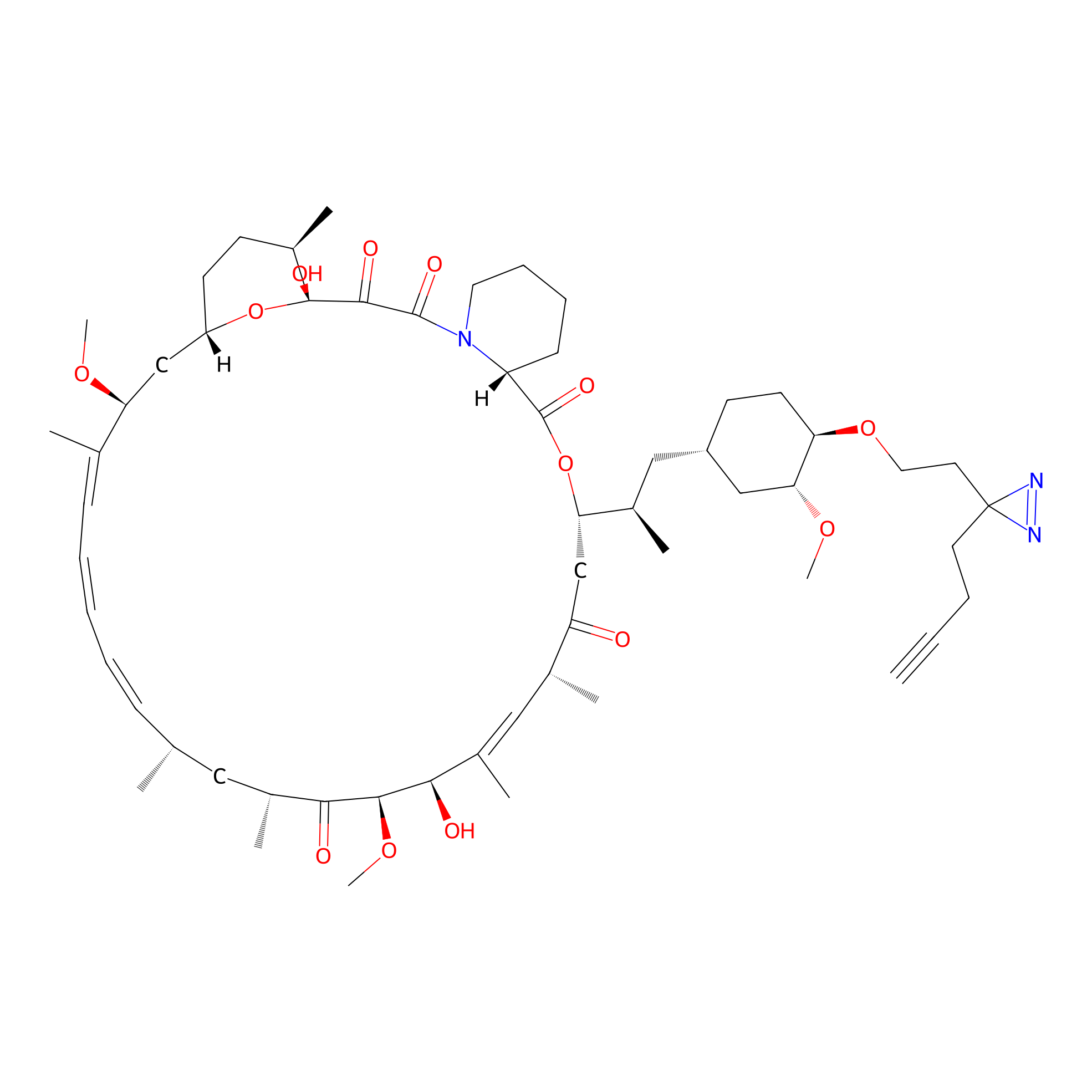
| LDCM0090 | Rapamycin | JHH-7 | 3.35 | LDD0213 | [5] |
The Interaction Atlas With This Target
References