Details of the Target
General Information of Target
| Target ID | LDTP03223 | |||||
|---|---|---|---|---|---|---|
| Target Name | Small ribosomal subunit protein uS3 (RPS3) | |||||
| Gene Name | RPS3 | |||||
| Gene ID | 6188 | |||||
| Synonyms |
Small ribosomal subunit protein uS3; 40S ribosomal protein S3; EC 4.2.99.18 |
|||||
| 3D Structure | ||||||
| Sequence |
MAVQISKKRKFVADGIFKAELNEFLTRELAEDGYSGVEVRVTPTRTEIIILATRTQNVLG
EKGRRIRELTAVVQKRFGFPEGSVELYAEKVATRGLCAIAQAESLRYKLLGGLAVRRACY GVLRFIMESGAKGCEVVVSGKLRGQRAKSMKFVDGLMIHSGDPVNYYVDTAVRHVLLRQG VLGIKVKIMLPWDPTGKIGPKKPLPDHVSIVEPKDEILPTTPISEQKGGKPEPPAMPQPV PTA |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Universal ribosomal protein uS3 family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Component of the small ribosomal subunit. The ribosome is a large ribonucleoprotein complex responsible for the synthesis of proteins in the cell. Has endonuclease activity and plays a role in repair of damaged DNA. Cleaves phosphodiester bonds of DNAs containing altered bases with broad specificity and cleaves supercoiled DNA more efficiently than relaxed DNA. Displays high binding affinity for 7,8-dihydro-8-oxoguanine (8-oxoG), a common DNA lesion caused by reactive oxygen species (ROS). Has also been shown to bind with similar affinity to intact and damaged DNA. Stimulates the N-glycosylase activity of the base excision protein OGG1. Enhances the uracil excision activity of UNG1. Also stimulates the cleavage of the phosphodiester backbone by APEX1. When located in the mitochondrion, reduces cellular ROS levels and mitochondrial DNA damage. Has also been shown to negatively regulate DNA repair in cells exposed to hydrogen peroxide. Plays a role in regulating transcription as part of the NF-kappa-B p65-p50 complex where it binds to the RELA/p65 subunit, enhances binding of the complex to DNA and promotes transcription of target genes. Represses its own translation by binding to its cognate mRNA. Binds to and protects TP53/p53 from MDM2-mediated ubiquitination. Involved in spindle formation and chromosome movement during mitosis by regulating microtubule polymerization. Involved in induction of apoptosis through its role in activation of CASP8. Induces neuronal apoptosis by interacting with the E2F1 transcription factor and acting synergistically with it to up-regulate pro-apoptotic proteins BCL2L11/BIM and HRK/Dp5. Interacts with TRADD following exposure to UV radiation and induces apoptosis by caspase-dependent JNK activation.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| JURKAT | SNV: p.I158T | Compound 10 Probe Info | |||
| MOLT4 | SNV: p.R65W | IA-alkyne Probe Info | |||
| OCUM1 | SNV: p.M236T | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [2] | |
|
A-EBA Probe Info |
 |
3.56 | LDD0215 | [3] | |
|
CHEMBL5175495 Probe Info |
 |
6.89 | LDD0196 | [4] | |
|
CY4 Probe Info |
 |
3.74 | LDD0244 | [5] | |
|
N1 Probe Info |
 |
5.62 | LDD0242 | [5] | |
|
TH211 Probe Info |
 |
Y166(12.15) | LDD0257 | [6] | |
|
TH216 Probe Info |
 |
Y167(20.00) | LDD0259 | [6] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [7] | |
|
1oxF11yne Probe Info |
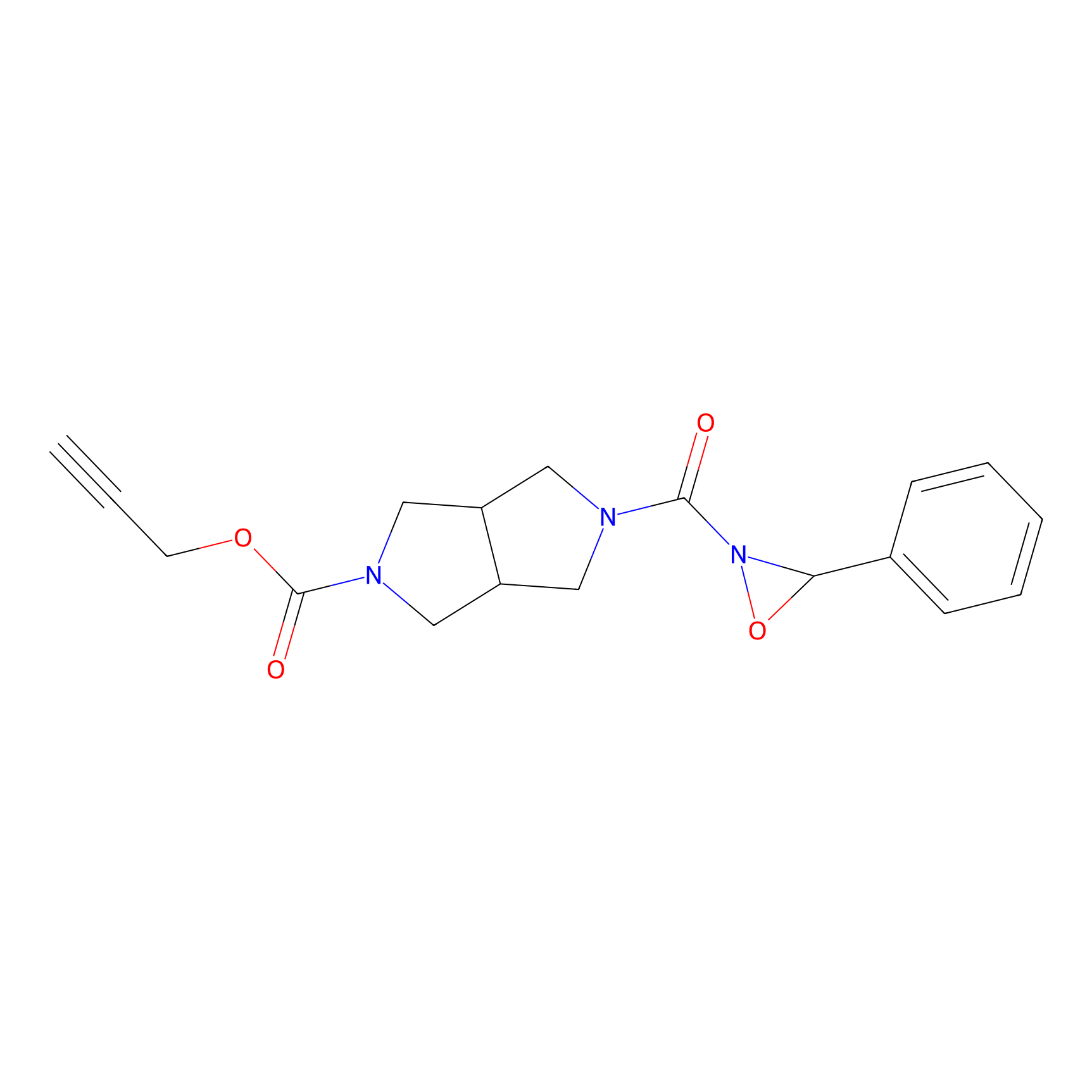 |
N.A. | LDD0193 | [8] | |
|
BTD Probe Info |
 |
C134(2.26); C97(5.17) | LDD1699 | [9] | |
|
AZ-9 Probe Info |
 |
D215(1.48); E89(1.00); E128(1.21); E38(10.00) | LDD2208 | [10] | |
|
ONAyne Probe Info |
 |
K202(0.00); K62(0.00); K75(0.00) | LDD0273 | [11] | |
|
OPA-S-S-alkyne Probe Info |
 |
K75(2.68); K62(3.22); K108(4.95) | LDD3494 | [12] | |
|
Probe 1 Probe Info |
 |
Y34(55.54); Y87(13.02); Y120(19.32) | LDD3495 | [13] | |
|
AF-2 Probe Info |
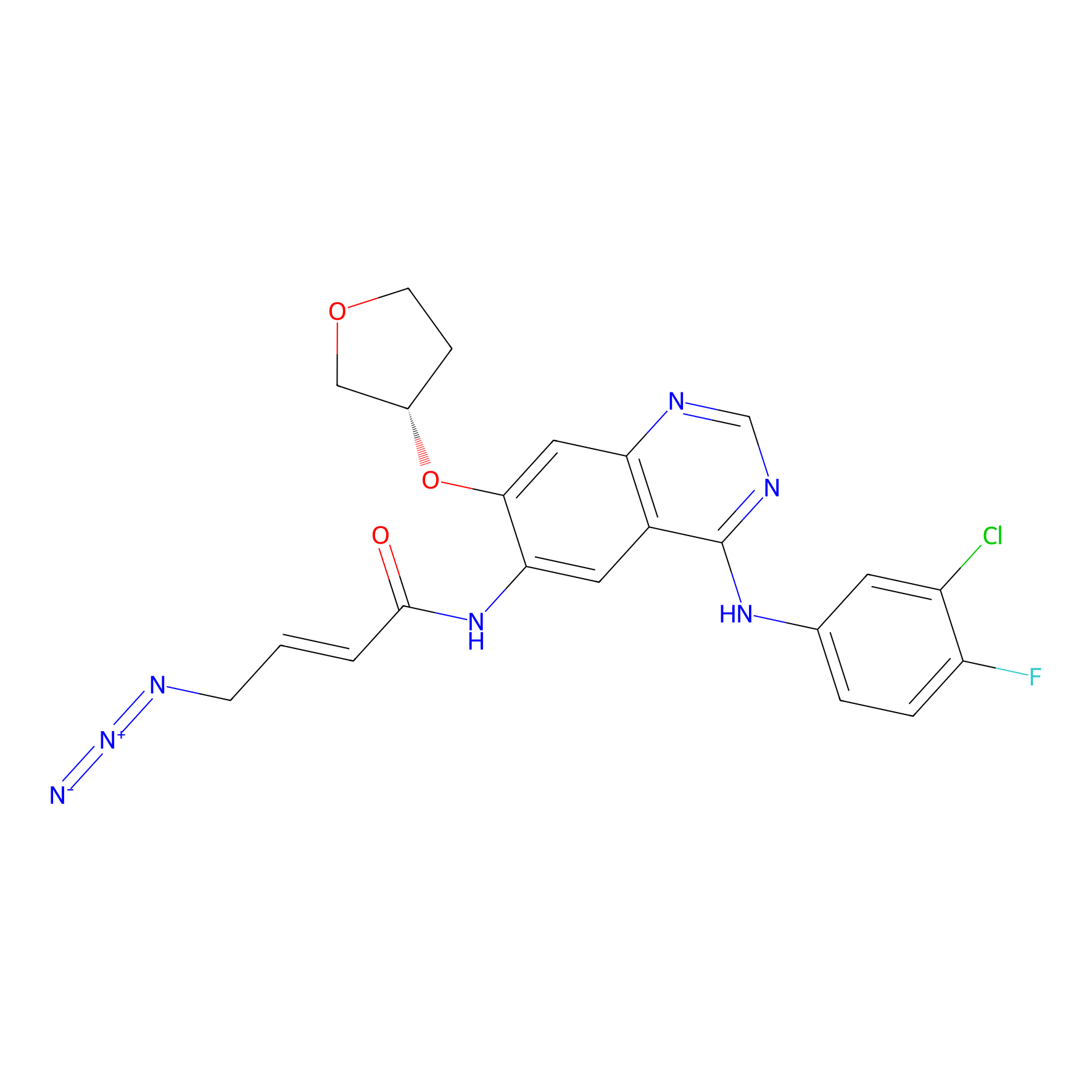 |
1.69 | LDD0422 | [14] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [15] | |
|
THZ1-DTB Probe Info |
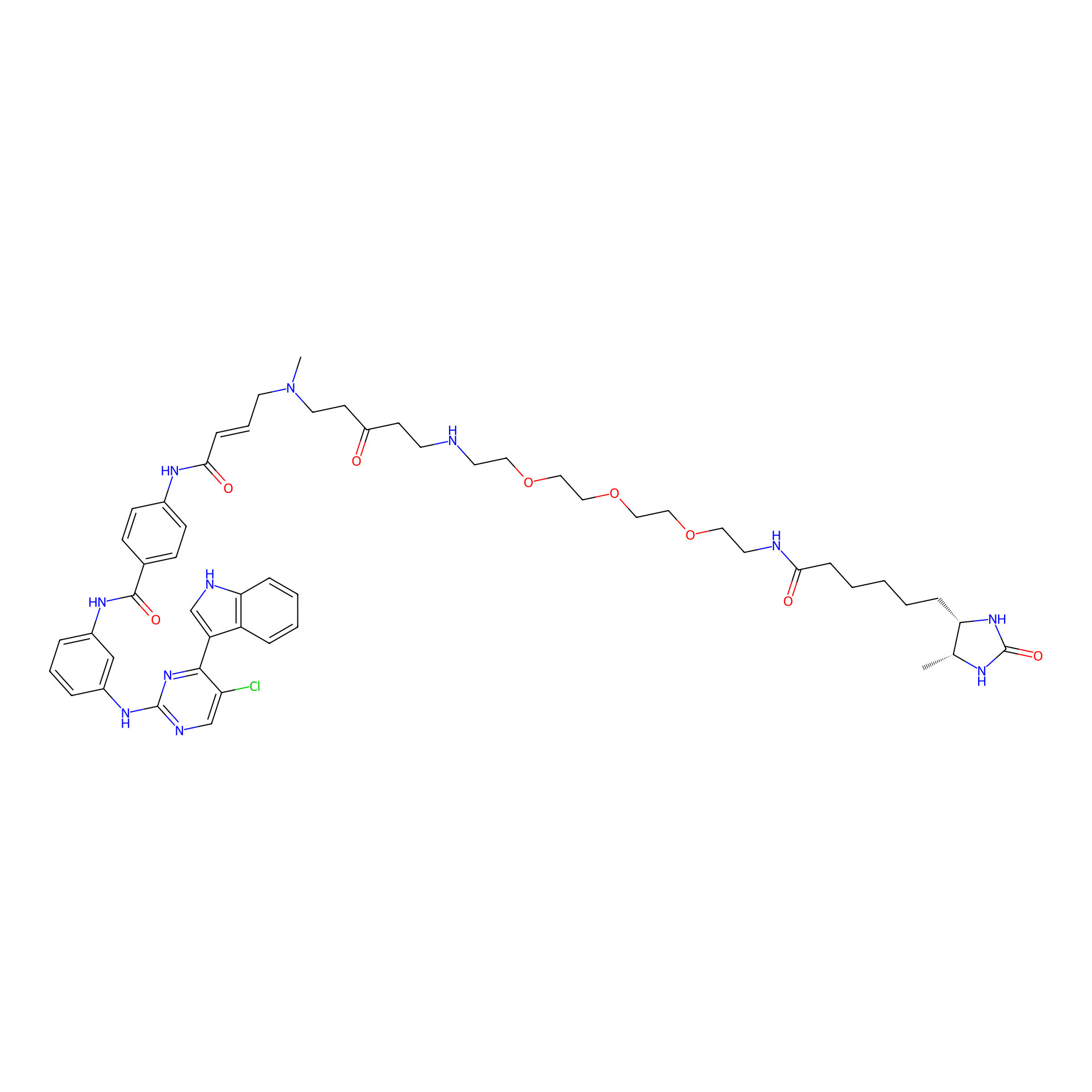 |
C97(1.06) | LDD0460 | [15] | |
|
AHL-Pu-1 Probe Info |
 |
C97(2.59) | LDD0168 | [16] | |
|
EA-probe Probe Info |
 |
C97(1.04) | LDD2210 | [17] | |
|
DBIA Probe Info |
 |
C97(0.94); C134(0.86) | LDD0078 | [18] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [19] | |
|
AMP probe Probe Info |
 |
K108(0.00); K62(0.00); K202(0.00); K201(0.00) | LDD0200 | [20] | |
|
ATP probe Probe Info |
 |
K108(0.00); K62(0.00); K202(0.00); K197(0.00) | LDD0199 | [20] | |
|
CY-1 Probe Info |
 |
K75(0.00); Q74(0.00) | LDD0246 | [5] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C119(0.00); C134(0.00); C97(0.00) | LDD0038 | [21] | |
|
IA-alkyne Probe Info |
 |
C119(0.00); C97(0.00); C134(0.00) | LDD0032 | [22] | |
|
IPIAA_H Probe Info |
 |
C97(0.00); C134(0.00) | LDD0030 | [23] | |
|
IPIAA_L Probe Info |
 |
C134(0.00); C97(0.00) | LDD0031 | [23] | |
|
Lodoacetamide azide Probe Info |
 |
C119(0.00); C134(0.00); C97(0.00) | LDD0037 | [21] | |
|
ATP probe Probe Info |
 |
K151(0.00); K10(0.00); K187(0.00); K75(0.00) | LDD0035 | [24] | |
|
Dyn-2 Probe Info |
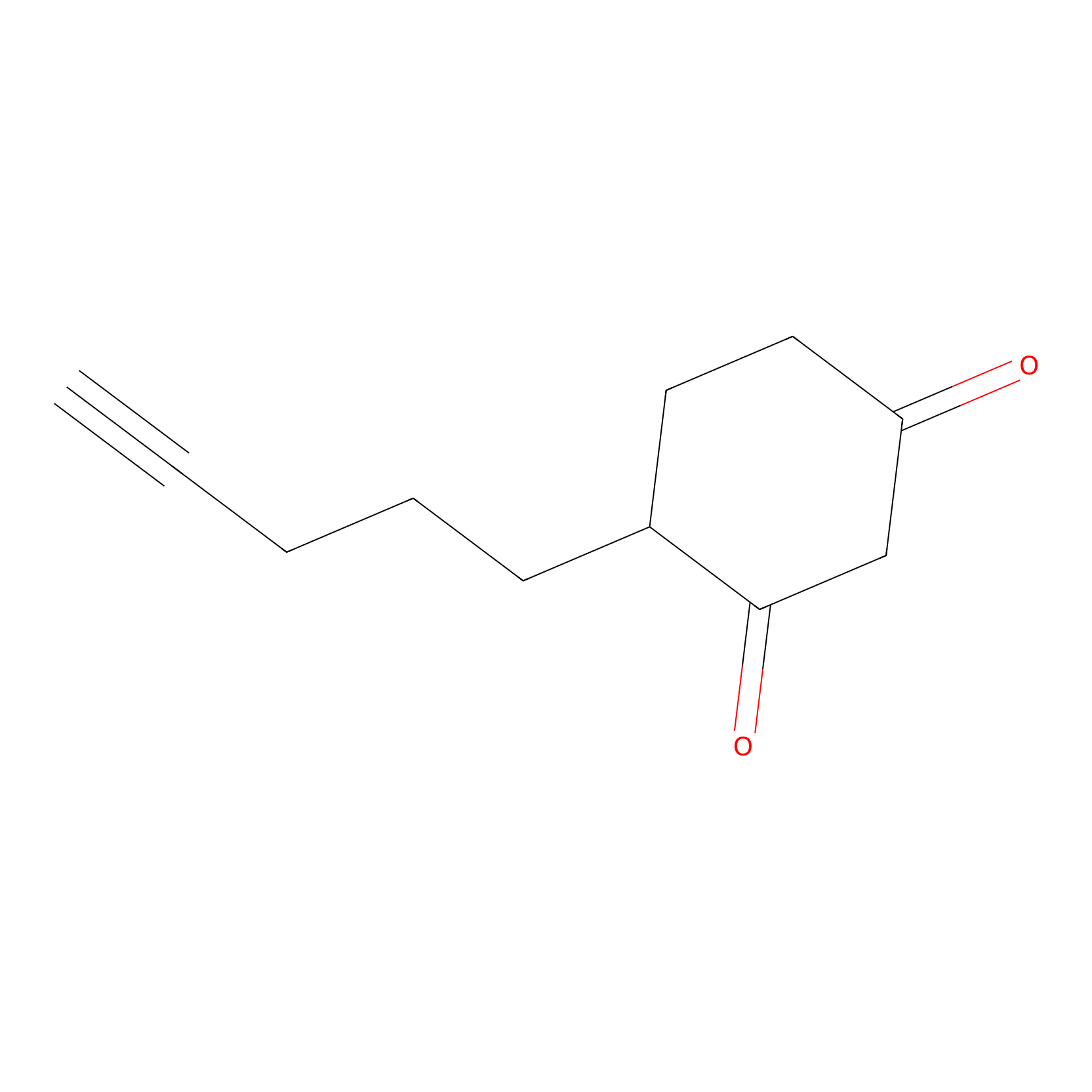 |
N.A. | LDD0013 | [25] | |
|
JW-RF-010 Probe Info |
 |
C134(0.00); C97(0.00) | LDD0026 | [26] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [27] | |
|
TFBX Probe Info |
 |
C134(0.00); C97(0.00) | LDD0027 | [26] | |
|
WYneN Probe Info |
 |
C119(0.00); C97(0.00); C134(0.00) | LDD0021 | [25] | |
|
WYneO Probe Info |
 |
C97(0.00); C134(0.00) | LDD0022 | [25] | |
|
1d-yne Probe Info |
 |
K151(0.00); K230(0.00); K10(0.00) | LDD0356 | [28] | |
|
Compound 10 Probe Info |
 |
C119(0.00); C134(0.00); C97(0.00) | LDD2216 | [29] | |
|
Compound 11 Probe Info |
 |
C134(0.00); C97(0.00) | LDD2213 | [29] | |
|
ENE Probe Info |
 |
C97(0.00); C134(0.00) | LDD0006 | [25] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [25] | |
|
NHS Probe Info |
 |
K230(0.00); K90(0.00); K197(0.00); K108(0.00) | LDD0010 | [25] | |
|
OSF Probe Info |
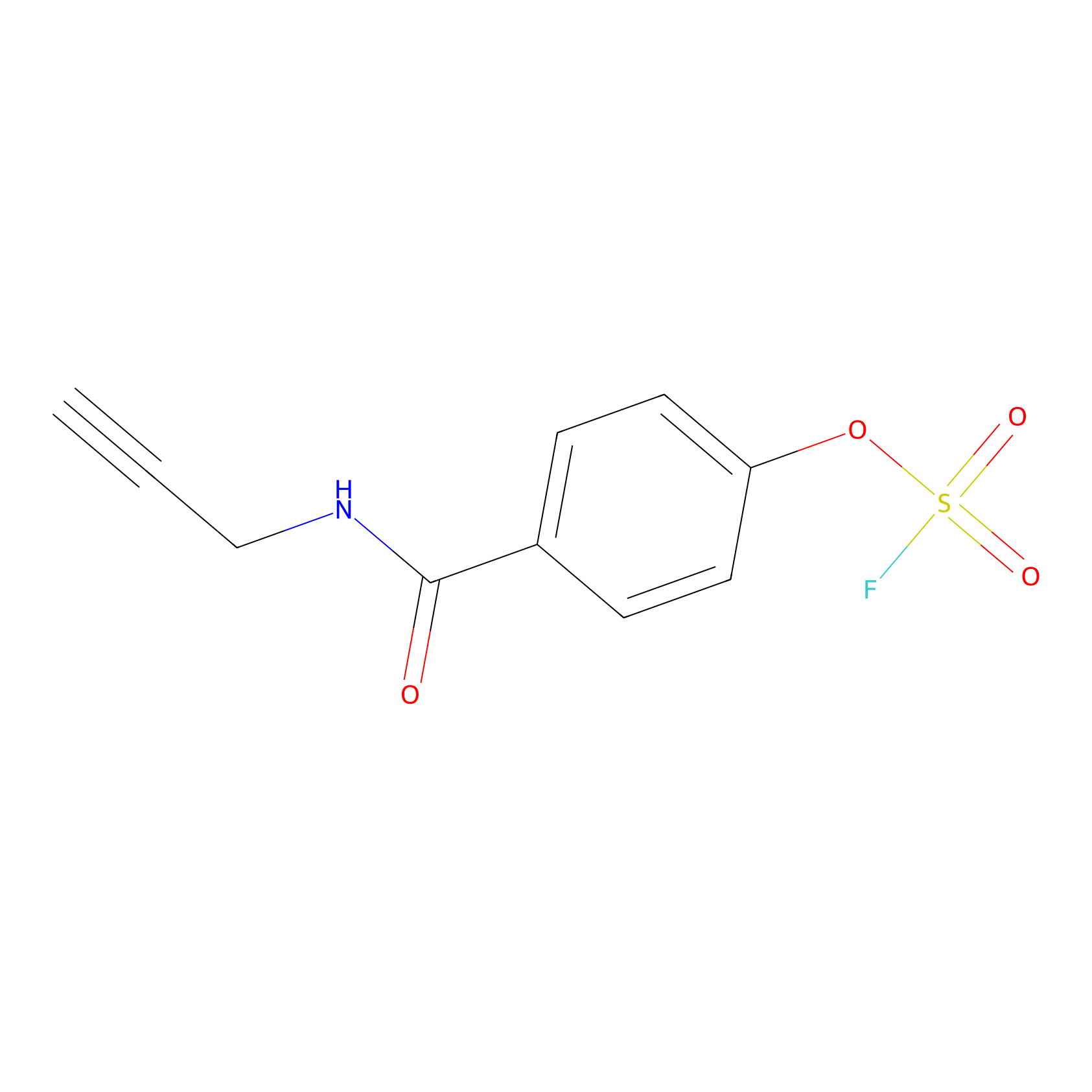 |
N.A. | LDD0029 | [30] | |
|
PF-06672131 Probe Info |
 |
C97(0.00); C134(0.00) | LDD0017 | [31] | |
|
SF Probe Info |
 |
Y120(0.00); Y167(0.00); Y87(0.00); K141(0.00) | LDD0028 | [30] | |
|
STPyne Probe Info |
 |
K230(0.00); K202(0.00) | LDD0009 | [25] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [25] | |
|
Phosphinate-6 Probe Info |
 |
C97(0.00); C119(0.00) | LDD0018 | [32] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [33] | |
|
1c-yne Probe Info |
 |
K201(0.00); K132(0.00); K202(0.00); K227(0.00) | LDD0228 | [28] | |
|
Acrolein Probe Info |
 |
H159(0.00); C97(0.00); H207(0.00); C134(0.00) | LDD0217 | [34] | |
|
Cinnamaldehyde Probe Info |
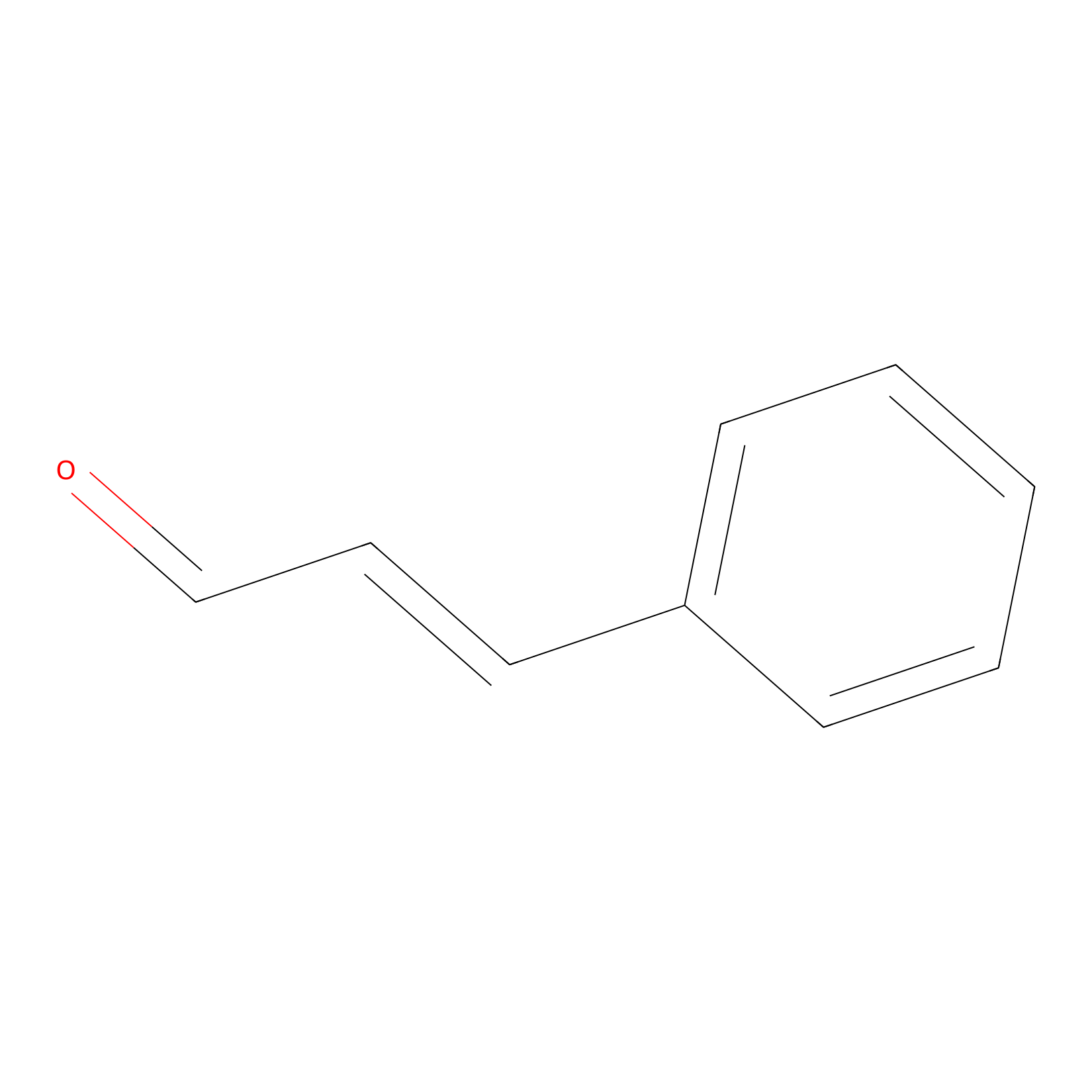 |
N.A. | LDD0220 | [34] | |
|
Crotonaldehyde Probe Info |
 |
C97(0.00); C134(0.00) | LDD0219 | [34] | |
|
Methacrolein Probe Info |
 |
C97(0.00); C134(0.00); C119(0.00) | LDD0218 | [34] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [35] | |
|
AOyne Probe Info |
 |
4.80 | LDD0443 | [36] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [37] | |
|
NAIA_5 Probe Info |
 |
C119(0.00); C134(0.00); C97(0.00) | LDD2223 | [27] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [37] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [37] | |
|
HHS-475 Probe Info |
 |
Y107(1.34); Y166(0.74); Y167(1.07) | LDD2238 | [38] | |
|
HHS-482 Probe Info |
 |
Y107(1.32); Y120(1.03); Y166(1.12); Y167(1.01) | LDD2239 | [38] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C210 Probe Info |
 |
56.10 | LDD1884 | [39] | |
|
C284 Probe Info |
 |
22.78 | LDD1954 | [39] | |
|
C313 Probe Info |
 |
14.03 | LDD1980 | [39] | |
|
C353 Probe Info |
 |
7.26 | LDD2014 | [39] | |
|
C354 Probe Info |
 |
14.83 | LDD2015 | [39] | |
|
FFF probe12 Probe Info |
 |
6.50 | LDD0473 | [40] | |
|
FFF probe13 Probe Info |
 |
8.25 | LDD0475 | [40] | |
|
FFF probe2 Probe Info |
 |
7.60 | LDD0463 | [40] | |
|
FFF probe3 Probe Info |
 |
9.00 | LDD0464 | [40] | |
|
FFF probe6 Probe Info |
 |
5.07 | LDD0468 | [40] | |
|
JN0003 Probe Info |
 |
6.94 | LDD0469 | [40] | |
|
STS-2 Probe Info |
 |
1.07 | LDD0138 | [41] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [42] | |
|
Kambe_26 Probe Info |
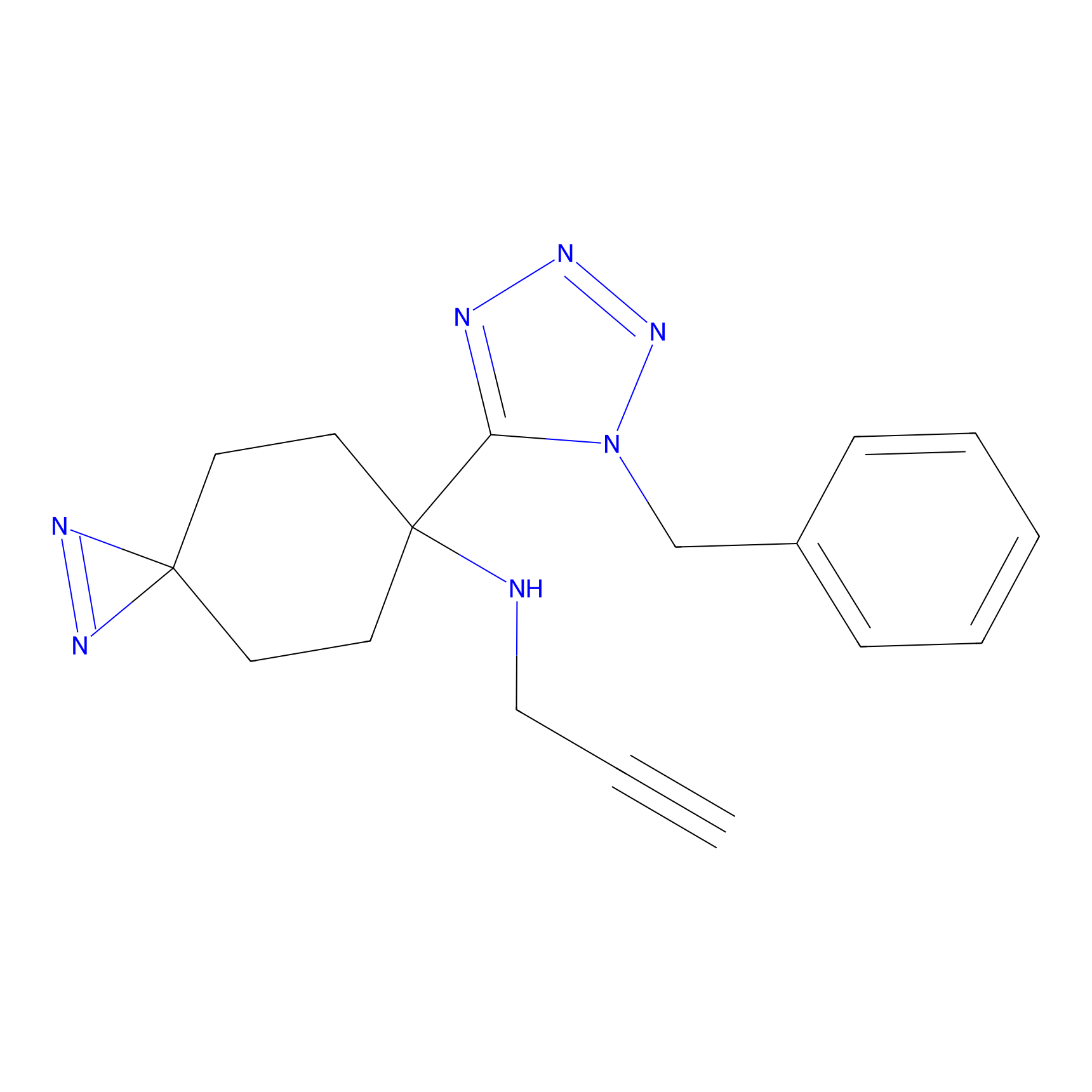 |
20.00 | LDD0127 | [43] | |
|
Diazir Probe Info |
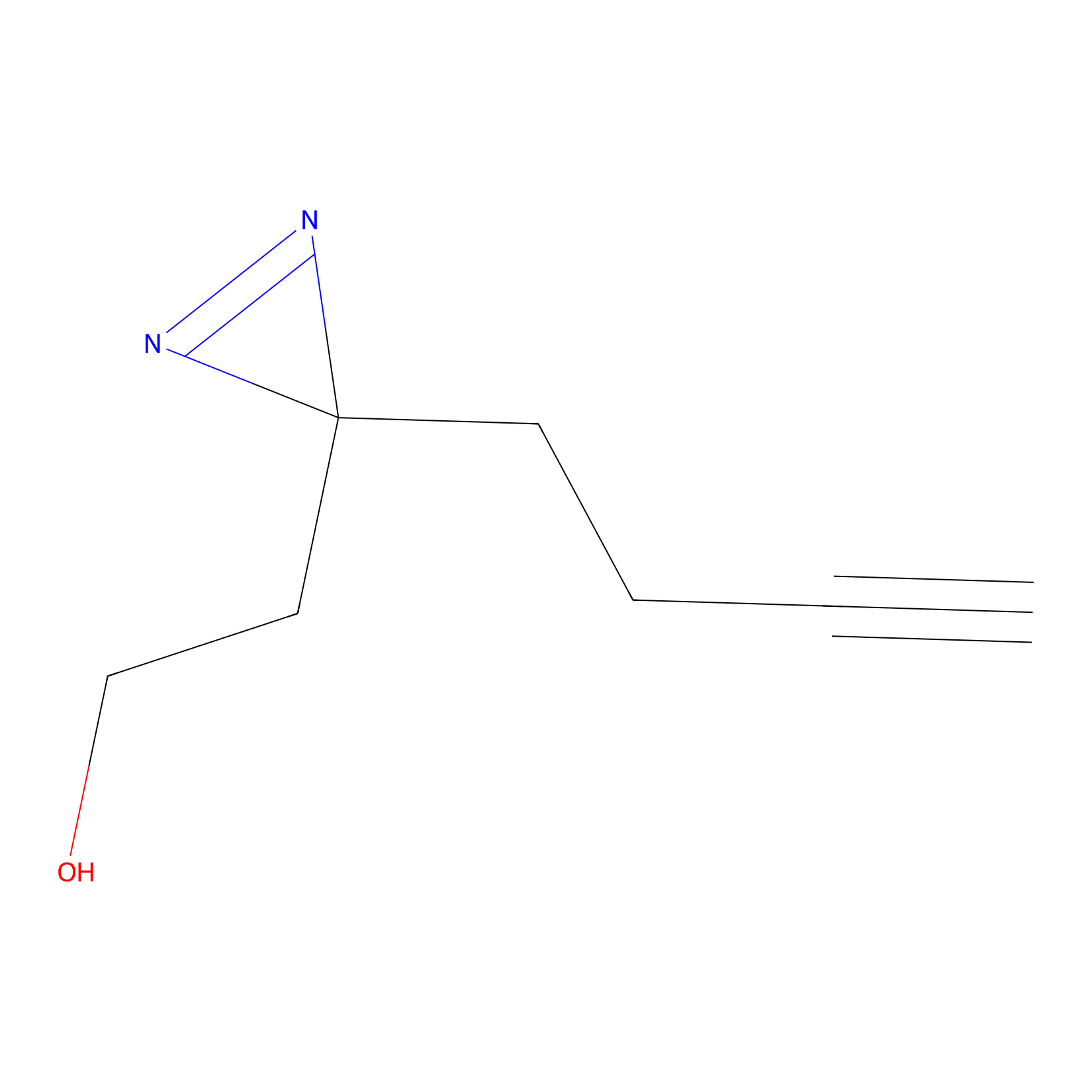 |
N.A. | LDD0011 | [25] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0073 | [44] | |
|
OEA-DA Probe Info |
 |
3.45 | LDD0046 | [45] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0068 | [46] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C134(1.20); C97(0.88); C119(0.73) | LDD2142 | [9] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C134(0.83); C97(1.24); C119(0.85) | LDD2112 | [9] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C134(0.58); C97(0.56); C119(0.56) | LDD2095 | [9] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C134(0.84); C97(0.83); C119(1.06) | LDD2130 | [9] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C134(1.11); C97(1.02); C119(0.98) | LDD2117 | [9] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C134(1.52); C97(0.99); C119(0.94) | LDD2152 | [9] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C134(0.87); C97(1.05); C119(0.98) | LDD2103 | [9] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C134(0.57); C97(0.64); C119(0.58) | LDD2132 | [9] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C134(0.55); C97(0.95); C119(0.52) | LDD2131 | [9] |
| LDCM0025 | 4SU-RNA | HEK-293T | C97(2.59) | LDD0168 | [16] |
| LDCM0563 | Abegg_cp(+)-11 | HeLa | C97(7.13) | LDD0314 | [26] |
| LDCM0561 | Abegg_cp(-)-10 | HeLa | C97(6.33) | LDD0312 | [26] |
| LDCM0562 | Abegg_cp(-)-11 | HeLa | C97(7.59) | LDD0313 | [26] |
| LDCM0214 | AC1 | HEK-293T | C119(0.94); C134(0.92); C97(1.00) | LDD1507 | [47] |
| LDCM0215 | AC10 | HEK-293T | C119(0.91); C134(0.98); C97(1.18) | LDD1508 | [47] |
| LDCM0226 | AC11 | HEK-293T | C119(0.98); C134(1.03) | LDD1509 | [47] |
| LDCM0237 | AC12 | HEK-293T | C119(1.01); C134(0.98); C97(0.99) | LDD1510 | [47] |
| LDCM0259 | AC14 | HEK-293T | C119(0.96); C134(1.11); C97(0.95) | LDD1512 | [47] |
| LDCM0270 | AC15 | HEK-293T | C119(0.94); C134(0.97) | LDD1513 | [47] |
| LDCM0276 | AC17 | HEK-293T | C119(1.00); C134(0.98); C97(0.91) | LDD1515 | [47] |
| LDCM0277 | AC18 | HEK-293T | C119(0.98); C134(0.99); C97(0.91) | LDD1516 | [47] |
| LDCM0278 | AC19 | HEK-293T | C119(0.99); C134(0.99) | LDD1517 | [47] |
| LDCM0279 | AC2 | HEK-293T | C119(0.92); C134(0.93); C97(1.09) | LDD1518 | [47] |
| LDCM0280 | AC20 | HEK-293T | C119(1.01); C134(0.99); C97(0.95) | LDD1519 | [47] |
| LDCM0281 | AC21 | HEK-293T | C119(1.04); C134(1.08) | LDD1520 | [47] |
| LDCM0282 | AC22 | HEK-293T | C119(1.04); C134(1.08); C97(0.99) | LDD1521 | [47] |
| LDCM0283 | AC23 | HEK-293T | C119(0.96); C134(0.93) | LDD1522 | [47] |
| LDCM0284 | AC24 | HEK-293T | C119(0.97); C134(0.95); C97(0.87) | LDD1523 | [47] |
| LDCM0285 | AC25 | HEK-293T | C119(1.01); C134(0.96); C97(1.09) | LDD1524 | [47] |
| LDCM0286 | AC26 | HEK-293T | C119(1.02); C134(1.02); C97(1.06) | LDD1525 | [47] |
| LDCM0287 | AC27 | HEK-293T | C119(1.03); C134(1.05) | LDD1526 | [47] |
| LDCM0288 | AC28 | HEK-293T | C119(1.07); C134(1.02); C97(1.28) | LDD1527 | [47] |
| LDCM0289 | AC29 | HEK-293T | C119(1.06); C134(1.04) | LDD1528 | [47] |
| LDCM0290 | AC3 | HEK-293T | C119(1.00); C134(1.00) | LDD1529 | [47] |
| LDCM0291 | AC30 | HEK-293T | C119(1.05); C134(1.09); C97(0.94) | LDD1530 | [47] |
| LDCM0292 | AC31 | HEK-293T | C119(1.02); C134(0.99) | LDD1531 | [47] |
| LDCM0293 | AC32 | HEK-293T | C119(1.00); C134(1.01); C97(0.96) | LDD1532 | [47] |
| LDCM0294 | AC33 | HEK-293T | C119(0.91); C134(0.84); C97(1.01) | LDD1533 | [47] |
| LDCM0295 | AC34 | HEK-293T | C119(0.95); C134(0.97); C97(0.97) | LDD1534 | [47] |
| LDCM0296 | AC35 | HEK-293T | C119(0.95); C134(1.02) | LDD1535 | [47] |
| LDCM0297 | AC36 | HEK-293T | C119(0.93); C134(0.90); C97(1.06) | LDD1536 | [47] |
| LDCM0298 | AC37 | HEK-293T | C119(0.91); C134(0.99) | LDD1537 | [47] |
| LDCM0299 | AC38 | HEK-293T | C119(0.92); C134(1.11); C97(0.87) | LDD1538 | [47] |
| LDCM0300 | AC39 | HEK-293T | C119(0.96); C134(0.90) | LDD1539 | [47] |
| LDCM0301 | AC4 | HEK-293T | C119(0.98); C134(0.97); C97(1.03) | LDD1540 | [47] |
| LDCM0302 | AC40 | HEK-293T | C119(0.94); C134(0.98); C97(1.09) | LDD1541 | [47] |
| LDCM0303 | AC41 | HEK-293T | C119(0.91); C134(0.89); C97(0.97) | LDD1542 | [47] |
| LDCM0304 | AC42 | HEK-293T | C119(0.90); C134(0.94); C97(1.15) | LDD1543 | [47] |
| LDCM0305 | AC43 | HEK-293T | C119(0.95); C134(0.98) | LDD1544 | [47] |
| LDCM0306 | AC44 | HEK-293T | C119(0.94); C134(0.91); C97(1.18) | LDD1545 | [47] |
| LDCM0307 | AC45 | HEK-293T | C119(0.92); C134(1.02) | LDD1546 | [47] |
| LDCM0308 | AC46 | HEK-293T | C119(0.92); C134(1.06); C97(0.91) | LDD1547 | [47] |
| LDCM0309 | AC47 | HEK-293T | C119(0.97); C134(0.93) | LDD1548 | [47] |
| LDCM0310 | AC48 | HEK-293T | C119(0.92); C134(0.90); C97(1.13) | LDD1549 | [47] |
| LDCM0311 | AC49 | HEK-293T | C119(0.98); C134(0.89); C97(1.13) | LDD1550 | [47] |
| LDCM0312 | AC5 | HEK-293T | C119(1.01); C134(1.04) | LDD1551 | [47] |
| LDCM0313 | AC50 | HEK-293T | C119(0.94); C134(0.96); C97(0.99) | LDD1552 | [47] |
| LDCM0314 | AC51 | HEK-293T | C119(0.90); C134(0.98) | LDD1553 | [47] |
| LDCM0315 | AC52 | HEK-293T | C119(0.97); C134(0.92); C97(0.99) | LDD1554 | [47] |
| LDCM0316 | AC53 | HEK-293T | C119(0.89); C134(0.99) | LDD1555 | [47] |
| LDCM0317 | AC54 | HEK-293T | C119(0.98); C134(1.08); C97(0.94) | LDD1556 | [47] |
| LDCM0318 | AC55 | HEK-293T | C119(0.96); C134(0.94) | LDD1557 | [47] |
| LDCM0319 | AC56 | HEK-293T | C119(0.95); C134(0.99); C97(1.00) | LDD1558 | [47] |
| LDCM0320 | AC57 | HEK-293T | C119(0.90); C134(0.89); C97(1.31) | LDD1559 | [47] |
| LDCM0321 | AC58 | HEK-293T | C119(0.95); C134(0.97); C97(1.13) | LDD1560 | [47] |
| LDCM0322 | AC59 | HEK-293T | C119(0.91); C134(0.98) | LDD1561 | [47] |
| LDCM0323 | AC6 | HEK-293T | C119(0.95); C134(1.04); C97(0.98) | LDD1562 | [47] |
| LDCM0324 | AC60 | HEK-293T | C119(0.95); C134(0.96); C97(1.07) | LDD1563 | [47] |
| LDCM0325 | AC61 | HEK-293T | C119(0.91); C134(1.00) | LDD1564 | [47] |
| LDCM0326 | AC62 | HEK-293T | C119(0.94); C134(1.02); C97(1.03) | LDD1565 | [47] |
| LDCM0327 | AC63 | HEK-293T | C119(1.02); C134(0.97) | LDD1566 | [47] |
| LDCM0328 | AC64 | HEK-293T | C119(0.90); C134(0.94); C97(1.17) | LDD1567 | [47] |
| LDCM0334 | AC7 | HEK-293T | C119(0.90); C134(0.92) | LDD1568 | [47] |
| LDCM0345 | AC8 | HEK-293T | C119(0.95); C134(0.99); C97(1.31) | LDD1569 | [47] |
| LDCM0545 | Acetamide | MDA-MB-231 | C134(0.59); C97(0.46); C119(0.39) | LDD2138 | [9] |
| LDCM0166 | Afatinib | A431 | 1.69 | LDD0422 | [14] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C134(1.13); C97(0.77); C119(1.15) | LDD2113 | [9] |
| LDCM0248 | AKOS034007472 | HEK-293T | C119(0.91); C134(0.98) | LDD1511 | [47] |
| LDCM0356 | AKOS034007680 | HEK-293T | C119(0.94); C134(0.95); C97(1.23) | LDD1570 | [47] |
| LDCM0275 | AKOS034007705 | HEK-293T | C119(0.97); C134(0.99); C97(1.13) | LDD1514 | [47] |
| LDCM0156 | Aniline | NCI-H1299 | 12.36 | LDD0403 | [2] |
| LDCM0020 | ARS-1620 | HCC44 | C97(0.94); C134(0.86) | LDD0078 | [18] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C134(0.65); C119(0.69) | LDD2091 | [9] |
| LDCM0630 | CCW28-3 | 231MFP | C97(1.57) | LDD2214 | [48] |
| LDCM0108 | Chloroacetamide | HeLa | H159(0.00); H207(0.00); C97(0.00); C134(0.00) | LDD0222 | [34] |
| LDCM0632 | CL-Sc | Hep-G2 | C134(1.06); C97(1.00); C119(0.90); C97(0.69) | LDD2227 | [27] |
| LDCM0367 | CL1 | HEK-293T | C119(0.90); C134(0.88) | LDD1571 | [47] |
| LDCM0368 | CL10 | HEK-293T | C119(0.86); C134(0.93); C97(4.85) | LDD1572 | [47] |
| LDCM0369 | CL100 | HEK-293T | C119(0.98); C134(0.93) | LDD1573 | [47] |
| LDCM0370 | CL101 | HEK-293T | C119(0.93); C134(0.95) | LDD1574 | [47] |
| LDCM0371 | CL102 | HEK-293T | C119(0.99); C134(0.90) | LDD1575 | [47] |
| LDCM0372 | CL103 | HEK-293T | C119(0.98); C134(0.97) | LDD1576 | [47] |
| LDCM0373 | CL104 | HEK-293T | C119(1.00); C134(0.93) | LDD1577 | [47] |
| LDCM0374 | CL105 | HEK-293T | C119(0.95); C134(0.96) | LDD1578 | [47] |
| LDCM0375 | CL106 | HEK-293T | C119(1.01); C134(0.99) | LDD1579 | [47] |
| LDCM0376 | CL107 | HEK-293T | C119(1.05); C134(1.00) | LDD1580 | [47] |
| LDCM0377 | CL108 | HEK-293T | C119(0.99); C134(0.94) | LDD1581 | [47] |
| LDCM0378 | CL109 | HEK-293T | C119(1.00); C134(1.01) | LDD1582 | [47] |
| LDCM0379 | CL11 | HEK-293T | C119(1.11); C134(1.16) | LDD1583 | [47] |
| LDCM0380 | CL110 | HEK-293T | C119(0.97); C134(0.92) | LDD1584 | [47] |
| LDCM0381 | CL111 | HEK-293T | C119(1.02); C134(0.97) | LDD1585 | [47] |
| LDCM0382 | CL112 | HEK-293T | C119(0.90); C134(0.91) | LDD1586 | [47] |
| LDCM0383 | CL113 | HEK-293T | C119(0.95); C134(0.99) | LDD1587 | [47] |
| LDCM0384 | CL114 | HEK-293T | C119(0.98); C134(0.86) | LDD1588 | [47] |
| LDCM0385 | CL115 | HEK-293T | C119(0.92); C134(0.92) | LDD1589 | [47] |
| LDCM0386 | CL116 | HEK-293T | C119(0.94); C134(0.93) | LDD1590 | [47] |
| LDCM0387 | CL117 | HEK-293T | C119(0.89); C134(0.89) | LDD1591 | [47] |
| LDCM0388 | CL118 | HEK-293T | C119(0.94); C134(0.98) | LDD1592 | [47] |
| LDCM0389 | CL119 | HEK-293T | C119(1.00); C134(0.91) | LDD1593 | [47] |
| LDCM0390 | CL12 | HEK-293T | C119(1.05); C134(1.06); C97(1.34) | LDD1594 | [47] |
| LDCM0391 | CL120 | HEK-293T | C119(0.93); C134(0.92) | LDD1595 | [47] |
| LDCM0392 | CL121 | HEK-293T | C119(0.96); C134(0.90) | LDD1596 | [47] |
| LDCM0393 | CL122 | HEK-293T | C119(0.98); C134(0.97) | LDD1597 | [47] |
| LDCM0394 | CL123 | HEK-293T | C119(0.88); C134(0.82) | LDD1598 | [47] |
| LDCM0395 | CL124 | HEK-293T | C119(0.91); C134(0.87) | LDD1599 | [47] |
| LDCM0396 | CL125 | HEK-293T | C119(0.96); C134(0.99) | LDD1600 | [47] |
| LDCM0397 | CL126 | HEK-293T | C119(0.93); C134(0.90) | LDD1601 | [47] |
| LDCM0398 | CL127 | HEK-293T | C119(0.94); C134(0.89) | LDD1602 | [47] |
| LDCM0399 | CL128 | HEK-293T | C119(0.91); C134(0.91) | LDD1603 | [47] |
| LDCM0400 | CL13 | HEK-293T | C119(0.87); C134(0.93) | LDD1604 | [47] |
| LDCM0401 | CL14 | HEK-293T | C119(0.93); C134(1.06) | LDD1605 | [47] |
| LDCM0402 | CL15 | HEK-293T | C119(0.91); C134(0.89) | LDD1606 | [47] |
| LDCM0403 | CL16 | HEK-293T | C119(0.94); C134(1.08) | LDD1607 | [47] |
| LDCM0404 | CL17 | HEK-293T | C119(0.82); C134(0.86); C97(2.01) | LDD1608 | [47] |
| LDCM0405 | CL18 | HEK-293T | C119(0.96); C134(1.12); C97(1.37) | LDD1609 | [47] |
| LDCM0406 | CL19 | HEK-293T | C119(0.96); C134(1.17) | LDD1610 | [47] |
| LDCM0407 | CL2 | HEK-293T | C119(0.97); C134(1.13) | LDD1611 | [47] |
| LDCM0408 | CL20 | HEK-293T | C119(0.99); C134(1.02); C97(1.13) | LDD1612 | [47] |
| LDCM0409 | CL21 | HEK-293T | C119(0.96); C134(1.19) | LDD1613 | [47] |
| LDCM0410 | CL22 | HEK-293T | C119(0.97); C134(1.15); C97(1.78) | LDD1614 | [47] |
| LDCM0411 | CL23 | HEK-293T | C119(1.05); C134(1.10) | LDD1615 | [47] |
| LDCM0412 | CL24 | HEK-293T | C119(1.02); C134(1.09); C97(1.31) | LDD1616 | [47] |
| LDCM0413 | CL25 | HEK-293T | C119(1.10); C134(1.00) | LDD1617 | [47] |
| LDCM0414 | CL26 | HEK-293T | C119(1.08); C134(1.08) | LDD1618 | [47] |
| LDCM0415 | CL27 | HEK-293T | C119(1.04); C134(1.01) | LDD1619 | [47] |
| LDCM0416 | CL28 | HEK-293T | C119(1.01); C134(1.03) | LDD1620 | [47] |
| LDCM0417 | CL29 | HEK-293T | C119(1.04); C134(1.05); C97(1.15) | LDD1621 | [47] |
| LDCM0418 | CL3 | HEK-293T | C119(0.95); C134(0.99) | LDD1622 | [47] |
| LDCM0419 | CL30 | HEK-293T | C119(1.09); C134(1.04); C97(1.20) | LDD1623 | [47] |
| LDCM0420 | CL31 | HEK-293T | C119(1.03); C134(1.05) | LDD1624 | [47] |
| LDCM0421 | CL32 | HEK-293T | C119(1.09); C134(1.06); C97(1.16) | LDD1625 | [47] |
| LDCM0422 | CL33 | HEK-293T | C119(1.70); C134(1.28) | LDD1626 | [47] |
| LDCM0423 | CL34 | HEK-293T | C119(1.06); C134(1.01); C97(1.62) | LDD1627 | [47] |
| LDCM0424 | CL35 | HEK-293T | C119(1.11); C134(1.10) | LDD1628 | [47] |
| LDCM0425 | CL36 | HEK-293T | C119(1.11); C134(0.94); C97(1.34) | LDD1629 | [47] |
| LDCM0426 | CL37 | HEK-293T | C119(1.05); C134(0.96) | LDD1630 | [47] |
| LDCM0428 | CL39 | HEK-293T | C119(0.91); C134(0.89) | LDD1632 | [47] |
| LDCM0429 | CL4 | HEK-293T | C119(0.93); C134(0.94) | LDD1633 | [47] |
| LDCM0430 | CL40 | HEK-293T | C119(1.02); C134(1.01) | LDD1634 | [47] |
| LDCM0431 | CL41 | HEK-293T | C119(0.93); C134(0.94); C97(1.23) | LDD1635 | [47] |
| LDCM0432 | CL42 | HEK-293T | C119(1.00); C134(1.05); C97(0.99) | LDD1636 | [47] |
| LDCM0433 | CL43 | HEK-293T | C119(1.01); C134(1.05) | LDD1637 | [47] |
| LDCM0434 | CL44 | HEK-293T | C119(1.11); C134(1.13); C97(1.08) | LDD1638 | [47] |
| LDCM0435 | CL45 | HEK-293T | C119(0.95); C134(0.93) | LDD1639 | [47] |
| LDCM0436 | CL46 | HEK-293T | C119(1.10); C134(0.95); C97(1.46) | LDD1640 | [47] |
| LDCM0437 | CL47 | HEK-293T | C119(1.02); C134(1.07) | LDD1641 | [47] |
| LDCM0438 | CL48 | HEK-293T | C119(1.10); C134(0.94); C97(1.16) | LDD1642 | [47] |
| LDCM0439 | CL49 | HEK-293T | C119(1.04); C134(0.96) | LDD1643 | [47] |
| LDCM0440 | CL5 | HEK-293T | C119(0.91); C134(0.92); C97(1.27) | LDD1644 | [47] |
| LDCM0441 | CL50 | HEK-293T | C119(1.04); C134(0.98) | LDD1645 | [47] |
| LDCM0443 | CL52 | HEK-293T | C119(1.04); C134(0.98) | LDD1646 | [47] |
| LDCM0444 | CL53 | HEK-293T | C119(0.93); C134(0.83); C97(1.22) | LDD1647 | [47] |
| LDCM0445 | CL54 | HEK-293T | C119(0.96); C134(0.90); C97(1.66) | LDD1648 | [47] |
| LDCM0446 | CL55 | HEK-293T | C119(1.05); C134(1.16) | LDD1649 | [47] |
| LDCM0447 | CL56 | HEK-293T | C119(1.07); C134(0.95); C97(1.14) | LDD1650 | [47] |
| LDCM0448 | CL57 | HEK-293T | C119(0.83); C134(0.98) | LDD1651 | [47] |
| LDCM0449 | CL58 | HEK-293T | C119(0.97); C134(0.98); C97(1.84) | LDD1652 | [47] |
| LDCM0450 | CL59 | HEK-293T | C119(1.03); C134(1.04) | LDD1653 | [47] |
| LDCM0451 | CL6 | HEK-293T | C119(0.92); C134(0.97); C97(1.92) | LDD1654 | [47] |
| LDCM0452 | CL60 | HEK-293T | C119(1.02); C134(0.99); C97(1.69) | LDD1655 | [47] |
| LDCM0453 | CL61 | HEK-293T | C119(0.93); C134(0.99) | LDD1656 | [47] |
| LDCM0454 | CL62 | HEK-293T | C119(1.01); C134(1.04) | LDD1657 | [47] |
| LDCM0455 | CL63 | HEK-293T | C119(0.96); C134(0.98) | LDD1658 | [47] |
| LDCM0456 | CL64 | HEK-293T | C119(1.01); C134(0.88) | LDD1659 | [47] |
| LDCM0457 | CL65 | HEK-293T | C119(0.96); C134(0.89); C97(1.22) | LDD1660 | [47] |
| LDCM0458 | CL66 | HEK-293T | C119(0.95); C134(0.94); C97(1.52) | LDD1661 | [47] |
| LDCM0459 | CL67 | HEK-293T | C119(1.04); C134(1.10) | LDD1662 | [47] |
| LDCM0460 | CL68 | HEK-293T | C119(0.98); C134(0.98); C97(1.12) | LDD1663 | [47] |
| LDCM0461 | CL69 | HEK-293T | C119(0.95); C134(1.03) | LDD1664 | [47] |
| LDCM0462 | CL7 | HEK-293T | C119(0.94); C134(1.15) | LDD1665 | [47] |
| LDCM0463 | CL70 | HEK-293T | C119(1.03); C134(1.05); C97(1.20) | LDD1666 | [47] |
| LDCM0464 | CL71 | HEK-293T | C119(1.01); C134(1.03) | LDD1667 | [47] |
| LDCM0465 | CL72 | HEK-293T | C119(1.05); C134(1.05); C97(1.28) | LDD1668 | [47] |
| LDCM0466 | CL73 | HEK-293T | C119(0.96); C134(0.89) | LDD1669 | [47] |
| LDCM0467 | CL74 | HEK-293T | C119(0.98); C134(1.05) | LDD1670 | [47] |
| LDCM0469 | CL76 | HEK-293T | C119(0.98); C134(0.94) | LDD1672 | [47] |
| LDCM0470 | CL77 | HEK-293T | C119(0.94); C134(0.82); C97(1.32) | LDD1673 | [47] |
| LDCM0471 | CL78 | HEK-293T | C119(1.02); C134(1.02); C97(1.16) | LDD1674 | [47] |
| LDCM0472 | CL79 | HEK-293T | C119(1.03); C134(1.07) | LDD1675 | [47] |
| LDCM0473 | CL8 | HEK-293T | C119(1.08); C134(0.84); C97(0.90) | LDD1676 | [47] |
| LDCM0474 | CL80 | HEK-293T | C119(1.08); C134(1.03); C97(1.00) | LDD1677 | [47] |
| LDCM0475 | CL81 | HEK-293T | C119(1.05); C134(1.13) | LDD1678 | [47] |
| LDCM0476 | CL82 | HEK-293T | C119(1.02); C134(1.06); C97(1.22) | LDD1679 | [47] |
| LDCM0477 | CL83 | HEK-293T | C119(1.07); C134(1.01) | LDD1680 | [47] |
| LDCM0478 | CL84 | HEK-293T | C119(1.04); C134(0.94); C97(1.60) | LDD1681 | [47] |
| LDCM0479 | CL85 | HEK-293T | C119(0.93); C134(1.06) | LDD1682 | [47] |
| LDCM0480 | CL86 | HEK-293T | C119(0.99); C134(1.00) | LDD1683 | [47] |
| LDCM0481 | CL87 | HEK-293T | C119(0.90); C134(0.86) | LDD1684 | [47] |
| LDCM0482 | CL88 | HEK-293T | C119(0.92); C134(0.91) | LDD1685 | [47] |
| LDCM0483 | CL89 | HEK-293T | C119(0.94); C134(0.91); C97(1.24) | LDD1686 | [47] |
| LDCM0484 | CL9 | HEK-293T | C119(0.98); C134(1.24) | LDD1687 | [47] |
| LDCM0485 | CL90 | HEK-293T | C119(1.35); C134(1.09); C97(2.49) | LDD1688 | [47] |
| LDCM0486 | CL91 | HEK-293T | C119(0.98); C134(1.06) | LDD1689 | [47] |
| LDCM0487 | CL92 | HEK-293T | C119(0.95); C134(0.90); C97(1.10) | LDD1690 | [47] |
| LDCM0488 | CL93 | HEK-293T | C119(0.94); C134(1.03) | LDD1691 | [47] |
| LDCM0489 | CL94 | HEK-293T | C119(0.92); C134(0.96); C97(1.37) | LDD1692 | [47] |
| LDCM0490 | CL95 | HEK-293T | C119(0.98); C134(0.79) | LDD1693 | [47] |
| LDCM0491 | CL96 | HEK-293T | C119(0.97); C134(0.90); C97(1.30) | LDD1694 | [47] |
| LDCM0492 | CL97 | HEK-293T | C119(0.95); C134(0.93) | LDD1695 | [47] |
| LDCM0493 | CL98 | HEK-293T | C119(1.06); C134(0.97) | LDD1696 | [47] |
| LDCM0494 | CL99 | HEK-293T | C119(0.92); C134(0.87) | LDD1697 | [47] |
| LDCM0634 | CY-0357 | Hep-G2 | C119(1.00); C97(0.71) | LDD2228 | [27] |
| LDCM0495 | E2913 | HEK-293T | C119(0.88); C134(0.83) | LDD1698 | [47] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C119(2.70); C97(1.33) | LDD1702 | [9] |
| LDCM0175 | Ethacrynic acid | HeLa | C97(1.04) | LDD2210 | [17] |
| LDCM0625 | F8 | Ramos | C119(0.71); C134(0.45); C97(0.81) | LDD2187 | [49] |
| LDCM0572 | Fragment10 | MDA-MB-231 | C97(3.44) | LDD1389 | [50] |
| LDCM0573 | Fragment11 | MDA-MB-231 | C97(4.78) | LDD1391 | [50] |
| LDCM0574 | Fragment12 | MDA-MB-231 | C97(7.17) | LDD1393 | [50] |
| LDCM0575 | Fragment13 | MDA-MB-231 | C97(1.23) | LDD1395 | [50] |
| LDCM0576 | Fragment14 | MDA-MB-231 | C97(1.46) | LDD1397 | [50] |
| LDCM0577 | Fragment15 | MDA-MB-231 | C97(1.01) | LDD1399 | [50] |
| LDCM0579 | Fragment20 | MDA-MB-231 | C97(2.54) | LDD1402 | [50] |
| LDCM0580 | Fragment21 | MDA-MB-231 | C97(2.16) | LDD1404 | [50] |
| LDCM0581 | Fragment22 | MDA-MB-231 | C97(1.25) | LDD1406 | [50] |
| LDCM0582 | Fragment23 | MDA-MB-231 | C97(1.61) | LDD1408 | [50] |
| LDCM0583 | Fragment24 | Ramos | C97(1.54) | LDD1410 | [50] |
| LDCM0584 | Fragment25 | MDA-MB-231 | C97(1.10) | LDD1411 | [50] |
| LDCM0585 | Fragment26 | Ramos | C97(0.92) | LDD1412 | [50] |
| LDCM0578 | Fragment27 | MDA-MB-231 | C97(0.90) | LDD1401 | [50] |
| LDCM0586 | Fragment28 | MDA-MB-231 | C97(0.98) | LDD1415 | [50] |
| LDCM0587 | Fragment29 | MDA-MB-231 | C97(1.18) | LDD1417 | [50] |
| LDCM0588 | Fragment30 | MDA-MB-231 | C97(1.14) | LDD1419 | [50] |
| LDCM0589 | Fragment31 | MDA-MB-231 | C97(1.35) | LDD1421 | [50] |
| LDCM0590 | Fragment32 | MDA-MB-231 | C97(3.19) | LDD1423 | [50] |
| LDCM0468 | Fragment33 | MDA-MB-231 | C97(1.28) | LDD1425 | [50] |
| LDCM0592 | Fragment34 | MDA-MB-231 | C97(0.89) | LDD1427 | [50] |
| LDCM0593 | Fragment35 | MDA-MB-231 | C97(0.88) | LDD1429 | [50] |
| LDCM0594 | Fragment36 | MDA-MB-231 | C97(4.00) | LDD1431 | [50] |
| LDCM0595 | Fragment37 | Ramos | C97(1.22) | LDD1432 | [50] |
| LDCM0596 | Fragment38 | MDA-MB-231 | C97(1.17) | LDD1433 | [50] |
| LDCM0597 | Fragment39 | MDA-MB-231 | C97(3.08) | LDD1435 | [50] |
| LDCM0566 | Fragment4 | MDA-MB-231 | C97(2.79) | LDD1378 | [50] |
| LDCM0598 | Fragment40 | MDA-MB-231 | C97(0.89) | LDD1436 | [50] |
| LDCM0599 | Fragment41 | MDA-MB-231 | C97(1.20) | LDD1438 | [50] |
| LDCM0600 | Fragment42 | Ramos | C97(0.96) | LDD1440 | [50] |
| LDCM0601 | Fragment43 | MDA-MB-231 | C97(4.14) | LDD1441 | [50] |
| LDCM0602 | Fragment44 | MDA-MB-231 | C97(0.80) | LDD1443 | [50] |
| LDCM0603 | Fragment45 | MDA-MB-231 | C97(3.06) | LDD1444 | [50] |
| LDCM0604 | Fragment46 | MDA-MB-231 | C97(0.96) | LDD1445 | [50] |
| LDCM0605 | Fragment47 | MDA-MB-231 | C97(1.16) | LDD1446 | [50] |
| LDCM0606 | Fragment48 | MDA-MB-231 | C97(1.18) | LDD1447 | [50] |
| LDCM0607 | Fragment49 | MDA-MB-231 | C97(18.46) | LDD1448 | [50] |
| LDCM0608 | Fragment50 | MDA-MB-231 | C97(1.83) | LDD1449 | [50] |
| LDCM0427 | Fragment51 | MDA-MB-231 | C97(1.89) | LDD1450 | [50] |
| LDCM0610 | Fragment52 | MDA-MB-231 | C97(1.27) | LDD1452 | [50] |
| LDCM0611 | Fragment53 | MDA-MB-231 | C97(1.02) | LDD1454 | [50] |
| LDCM0612 | Fragment54 | MDA-MB-231 | C97(1.17) | LDD1456 | [50] |
| LDCM0613 | Fragment55 | MDA-MB-231 | C97(2.09) | LDD1457 | [50] |
| LDCM0614 | Fragment56 | MDA-MB-231 | C97(1.01) | LDD1458 | [50] |
| LDCM0568 | Fragment6 | MDA-MB-231 | C97(1.00) | LDD1382 | [50] |
| LDCM0569 | Fragment7 | MDA-MB-231 | C97(2.41) | LDD1383 | [50] |
| LDCM0570 | Fragment8 | MDA-MB-231 | C97(2.40) | LDD1385 | [50] |
| LDCM0571 | Fragment9 | MDA-MB-231 | C97(1.12) | LDD1387 | [50] |
| LDCM0015 | HNE | MDA-MB-231 | C97(1.01); C134(0.98) | LDD0346 | [49] |
| LDCM0107 | IAA | HeLa | H207(0.00); C97(0.00); H159(0.00); C134(0.00) | LDD0221 | [34] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [15] |
| LDCM0073 | Kambe_cp3 | PC-3 | 20.00 | LDD0127 | [43] |
| LDCM0078 | kambe_cp70 | PC-3 | 2.08 | LDD0134 | [43] |
| LDCM0022 | KB02 | MDA-MB-231 | C97(7.31) | LDD1374 | [50] |
| LDCM0023 | KB03 | Jurkat | C97(8.65); C134(6.77); C119(79.86) | LDD0209 | [51] |
| LDCM0024 | KB05 | COLO792 | C150(2.96) | LDD3310 | [52] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C97(1.07) | LDD2102 | [9] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C134(0.89); C97(0.54); C119(0.87) | LDD2121 | [9] |
| LDCM0109 | NEM | HeLa | H159(0.00); H207(0.00); C97(0.00) | LDD0223 | [34] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C134(0.77); C97(0.57); C119(0.93) | LDD2089 | [9] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C134(0.79); C97(1.30) | LDD2090 | [9] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C134(1.04); C97(0.92); C119(1.06) | LDD2092 | [9] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C134(1.21); C97(1.23); C119(1.24) | LDD2093 | [9] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C134(1.31); C97(1.06); C119(1.54) | LDD2094 | [9] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C97(0.25); C119(0.26) | LDD2096 | [9] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C134(1.72); C97(0.96); C119(1.31) | LDD2097 | [9] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C134(1.07); C97(0.97) | LDD2098 | [9] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C134(1.30); C97(0.90); C119(1.72) | LDD2099 | [9] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C119(0.66) | LDD2100 | [9] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C134(0.97); C97(0.61) | LDD2101 | [9] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C134(0.65); C97(0.95); C119(0.61) | LDD2104 | [9] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C134(0.75); C97(1.55); C119(1.21) | LDD2105 | [9] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C134(0.68); C97(0.93); C119(1.10) | LDD2106 | [9] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C134(1.08); C97(0.95); C119(1.02) | LDD2107 | [9] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C134(0.89); C97(0.87); C119(0.79) | LDD2108 | [9] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C134(0.59); C97(0.86); C119(0.73) | LDD2109 | [9] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C134(0.90); C97(1.99); C119(1.09) | LDD2110 | [9] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C134(1.00); C97(1.01); C119(0.73) | LDD2111 | [9] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C134(0.54); C97(0.83); C119(0.68) | LDD2114 | [9] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C134(0.58); C97(0.59); C119(0.63) | LDD2115 | [9] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C134(1.25); C97(0.33); C119(0.69) | LDD2116 | [9] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C134(0.72); C97(0.35); C119(0.62) | LDD2118 | [9] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C134(1.99); C97(1.92); C119(1.54) | LDD2119 | [9] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C134(0.64); C97(0.66); C119(0.63) | LDD2120 | [9] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C134(0.67); C97(0.31); C119(0.24) | LDD2122 | [9] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C134(0.84); C97(0.92); C119(0.85) | LDD2123 | [9] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C134(0.94); C97(0.26); C119(0.26) | LDD2124 | [9] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C134(0.86); C97(0.87); C119(0.84) | LDD2125 | [9] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C134(0.22); C97(0.28); C119(0.34) | LDD2126 | [9] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C134(1.03); C97(0.89); C119(1.02) | LDD2127 | [9] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C134(0.84); C97(0.79); C119(0.43) | LDD2128 | [9] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C134(1.25); C97(1.00); C119(1.33) | LDD2129 | [9] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C134(0.65); C97(0.57); C119(0.77) | LDD2133 | [9] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C134(0.74); C97(0.52); C119(0.60) | LDD2134 | [9] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C134(1.43); C97(1.06) | LDD2135 | [9] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C134(1.19); C97(0.98); C119(1.32) | LDD2136 | [9] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C134(0.89); C97(0.85); C119(0.95) | LDD2137 | [9] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C119(2.70); C97(1.43) | LDD1700 | [9] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C134(0.94); C97(0.80); C119(0.85) | LDD2140 | [9] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C134(0.75); C97(0.84); C119(0.70) | LDD2141 | [9] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C134(0.89); C97(0.71); C119(0.51) | LDD2143 | [9] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C134(2.60); C97(2.31); C119(1.98) | LDD2144 | [9] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C97(6.24); C119(4.09) | LDD2145 | [9] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C134(1.12); C97(1.11); C119(1.08) | LDD2146 | [9] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C134(1.10); C97(1.17) | LDD2147 | [9] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C134(0.52); C97(0.58); C119(0.53) | LDD2148 | [9] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C134(1.01); C97(0.31) | LDD2149 | [9] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C134(0.58); C97(0.43); C119(0.58) | LDD2150 | [9] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C134(0.99); C97(0.37); C119(0.27) | LDD2151 | [9] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C97(2.26) | LDD2153 | [9] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C134(0.95); C97(0.72) | LDD2207 | [53] |
| LDCM0131 | RA190 | MM1.R | C134(1.41); C119(1.31); C97(1.12) | LDD0304 | [54] |
| LDCM0021 | THZ1 | HeLa S3 | C97(1.06) | LDD0460 | [15] |
| LDCM0111 | W14 | Hep-G2 | S160(0.59) | LDD0238 | [35] |
| LDCM0112 | W16 | Hep-G2 | Q74(0.67); K75(0.88); R76(0.89) | LDD0239 | [35] |
| LDCM0113 | W17 | Hep-G2 | C97(2.64) | LDD0240 | [35] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| 14-3-3 protein zeta/delta (YWHAZ) | 14-3-3 family | P63104 | |||
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
| Cellular tumor antigen p53 (TP53) | P53 family | P04637 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| NF-kappa-B inhibitor alpha (NFKBIA) | NF-kappa-B inhibitor family | P25963 | |||
| Nuclear factor NF-kappa-B p105 subunit (NFKB1) | . | P19838 | |||
| Transcription factor p65 (RELA) | . | Q04206 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Protein LTV1 homolog (LTV1) | LTV1 family | Q96GA3 | |||
| NEDD8 (NEDD8) | Ubiquitin family | Q15843 | |||
References
