Details of the Target
General Information of Target
| Target ID | LDTP03614 | |||||
|---|---|---|---|---|---|---|
| Target Name | Heterogeneous nuclear ribonucleoprotein H3 (HNRNPH3) | |||||
| Gene Name | HNRNPH3 | |||||
| Gene ID | 3189 | |||||
| Synonyms |
HNRPH3; Heterogeneous nuclear ribonucleoprotein H3; hnRNP H3; Heterogeneous nuclear ribonucleoprotein 2H9; hnRNP 2H9 |
|||||
| 3D Structure | ||||||
| Sequence |
MDWVMKHNGPNDASDGTVRLRGLPFGCSKEEIVQFFQGLEIVPNGITLTMDYQGRSTGEA
FVQFASKEIAENALGKHKERIGHRYIEIFRSSRSEIKGFYDPPRRLLGQRPGPYDRPIGG RGGYYGAGRGSMYDRMRRGGDGYDGGYGGFDDYGGYNNYGYGNDGFDDRMRDGRGMGGHG YGGAGDASSGFHGGHFVHMRGLPFRATENDIANFFSPLNPIRVHIDIGADGRATGEADVE FVTHEDAVAAMSKDKNNMQHRYIELFLNSTPGGGSGMGGSGMGGYGRDGMDNQGGYGSVG RMGMGNNYSGGYGTPDGLGGYGRGGGGSGGYYGQGGMSGGGWRGMY |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Involved in the splicing process and participates in early heat shock-induced splicing arrest. Due to their great structural variations the different isoforms may possess different functions in the splicing reaction.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
A-EBA Probe Info |
 |
3.86 | LDD0215 | [1] | |
|
ILS-1 Probe Info |
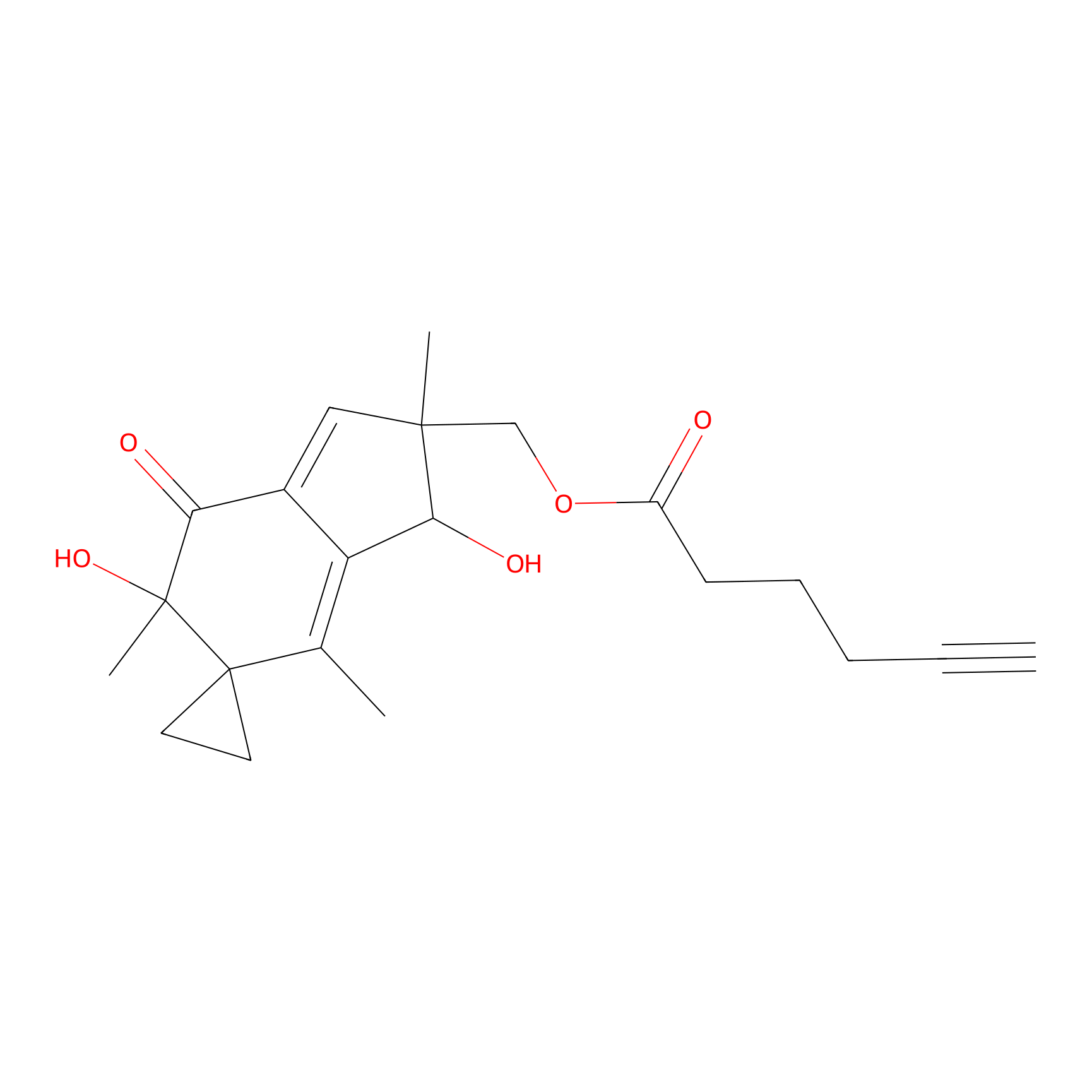 |
2.76 | LDD0415 | [2] | |
|
TH211 Probe Info |
 |
Y296(20.00); Y85(9.01); Y100(7.11); Y262(20.00) | LDD0260 | [3] | |
|
C-Sul Probe Info |
 |
2.41 | LDD0066 | [4] | |
|
TH216 Probe Info |
 |
Y296(20.00); Y85(12.38) | LDD0259 | [3] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
STPyne Probe Info |
 |
K255(10.00); K67(7.18); K97(3.53) | LDD0277 | [6] | |
|
ONAyne Probe Info |
 |
K97(0.00); K255(0.00) | LDD0273 | [6] | |
|
Probe 1 Probe Info |
 |
Y296(63.26); Y312(14.49); Y332(7.93) | LDD3495 | [7] | |
|
m-APA Probe Info |
 |
11.35 | LDD0403 | [8] | |
|
P11 Probe Info |
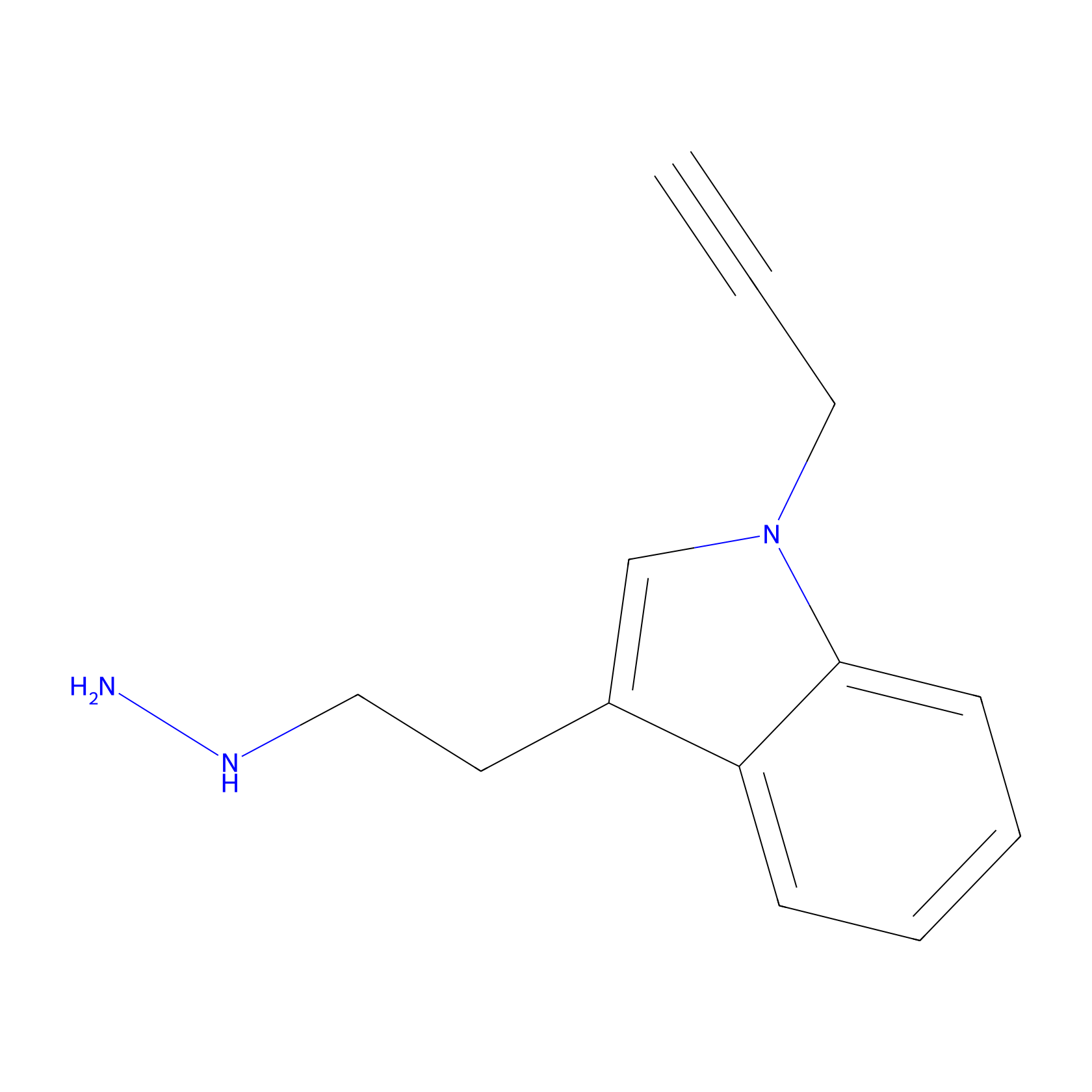 |
1.28 | LDD0211 | [9] | |
|
HPAP Probe Info |
 |
3.77 | LDD0063 | [10] | |
|
5E-2FA Probe Info |
 |
H7(0.00); H224(0.00) | LDD2235 | [11] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [12] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [13] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [14] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [15] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [16] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [16] | |
|
SF Probe Info |
 |
Y100(0.00); Y114(0.00); Y296(0.00) | LDD0028 | [17] | |
|
1c-yne Probe Info |
 |
K97(0.00); K6(0.00); K67(0.00) | LDD0228 | [18] | |
|
Acrolein Probe Info |
 |
H244(0.00); H7(0.00); H224(0.00) | LDD0217 | [19] | |
|
Crotonaldehyde Probe Info |
 |
H224(0.00); H7(0.00) | LDD0219 | [19] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [19] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [20] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [21] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [22] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 11.35 | LDD0403 | [8] |
| LDCM0088 | C45 | HEK-293T | 1.28 | LDD0211 | [9] |
| LDCM0108 | Chloroacetamide | HeLa | H195(0.00); H7(0.00); H244(0.00); H224(0.00) | LDD0222 | [19] |
| LDCM0107 | IAA | HeLa | H244(0.00); H7(0.00); H224(0.00); H195(0.00) | LDD0221 | [19] |
| LDCM0109 | NEM | HeLa | H244(0.00); H7(0.00); H224(0.00); H195(0.00) | LDD0223 | [19] |
| LDCM0014 | Panhematin | K562 | 3.77 | LDD0063 | [10] |
The Interaction Atlas With This Target
References