Details of the Target
General Information of Target
| Target ID | LDTP05029 | |||||
|---|---|---|---|---|---|---|
| Target Name | Peptidyl-prolyl cis-trans isomerase FKBP1A (FKBP1A) | |||||
| Gene Name | FKBP1A | |||||
| Gene ID | 2280 | |||||
| Synonyms |
FKBP1; FKBP12; Peptidyl-prolyl cis-trans isomerase FKBP1A; PPIase FKBP1A; EC 5.2.1.8; 12 kDa FK506-binding protein; 12 kDa FKBP; FKBP-12; Calstabin-1; FK506-binding protein 1A; FKBP-1A; Immunophilin FKBP12; Rotamase
|
|||||
| 3D Structure | ||||||
| Sequence |
MGVQVETISPGDGRTFPKRGQTCVVHYTGMLEDGKKFDSSRDRNKPFKFMLGKQEVIRGW
EEGVAQMSVGQRAKLTISPDYAYGATGHPGIIPPHATLVFDVELLKLE |
|||||
| Target Type |
Successful
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
FKBP-type PPIase family, FKBP1 subfamily
|
|||||
| Subcellular location |
Cytoplasm, cytosol
|
|||||
| Function |
Keeps in an inactive conformation TGFBR1, the TGF-beta type I serine/threonine kinase receptor, preventing TGF-beta receptor activation in absence of ligand. Recruits SMAD7 to ACVR1B which prevents the association of SMAD2 and SMAD3 with the activin receptor complex, thereby blocking the activin signal. May modulate the RYR1 calcium channel activity. PPIases accelerate the folding of proteins. It catalyzes the cis-trans isomerization of proline imidic peptide bonds in oligopeptides.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
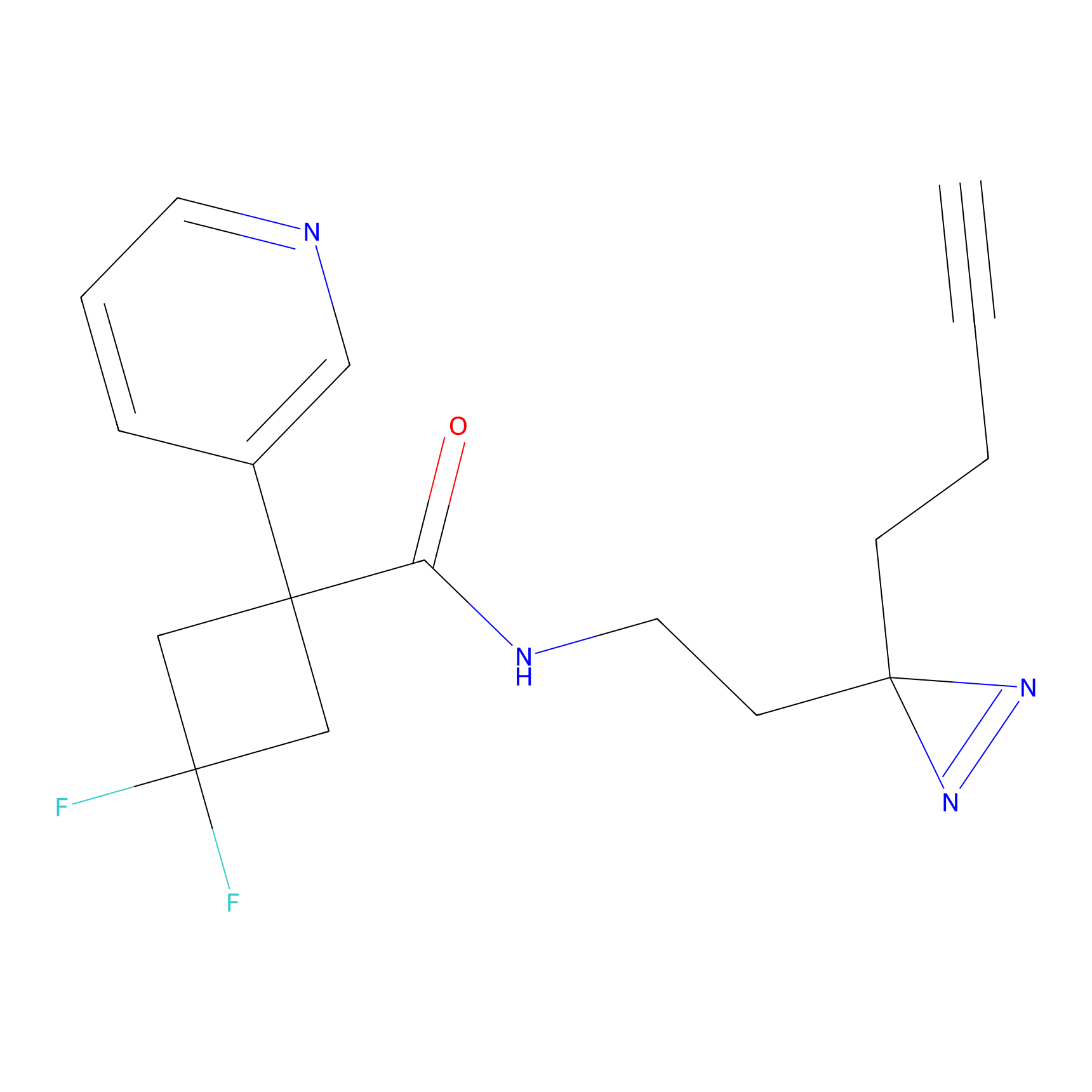
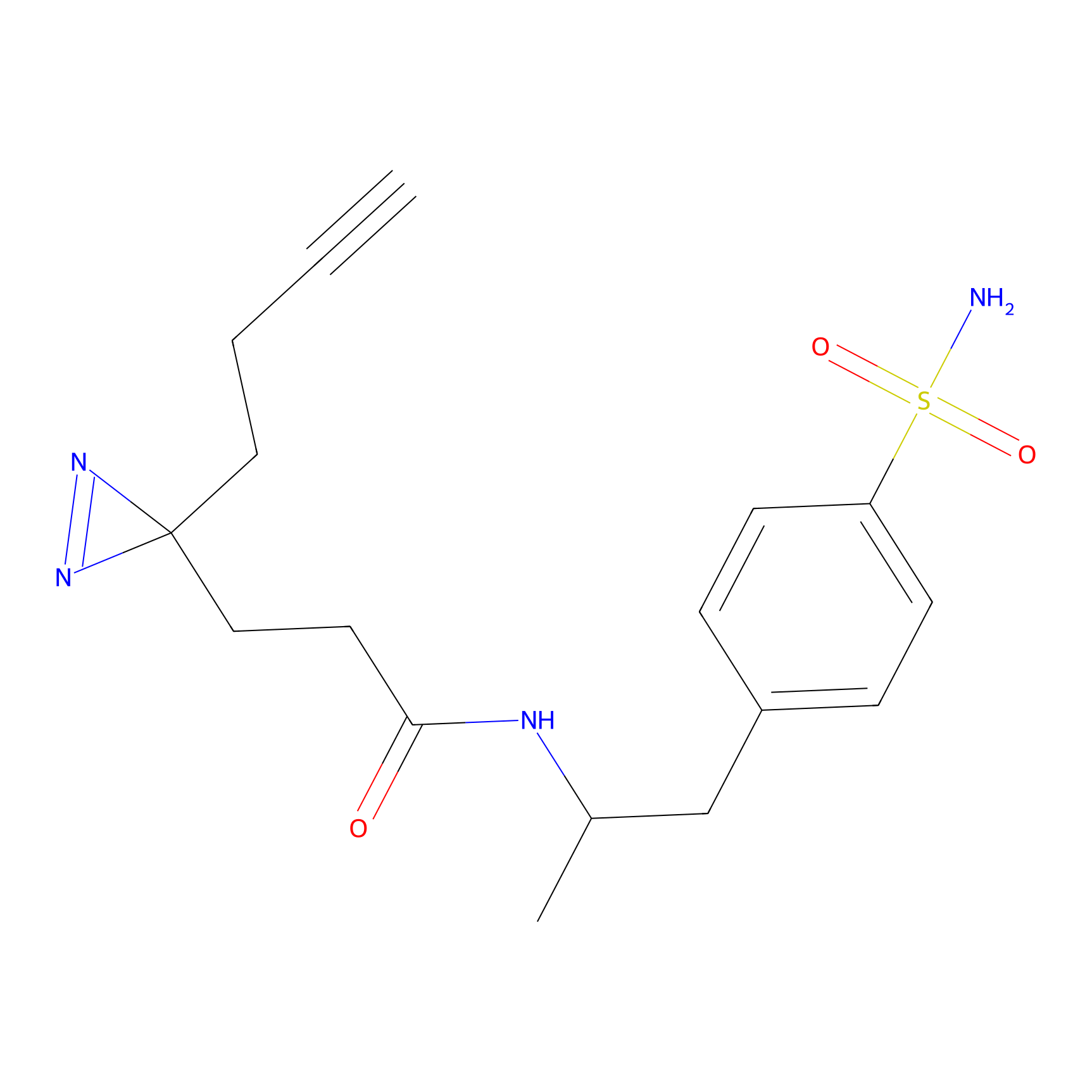
TH211 Probe Info |
 |
Y27(5.62) | LDD0260 | [1] | |
|
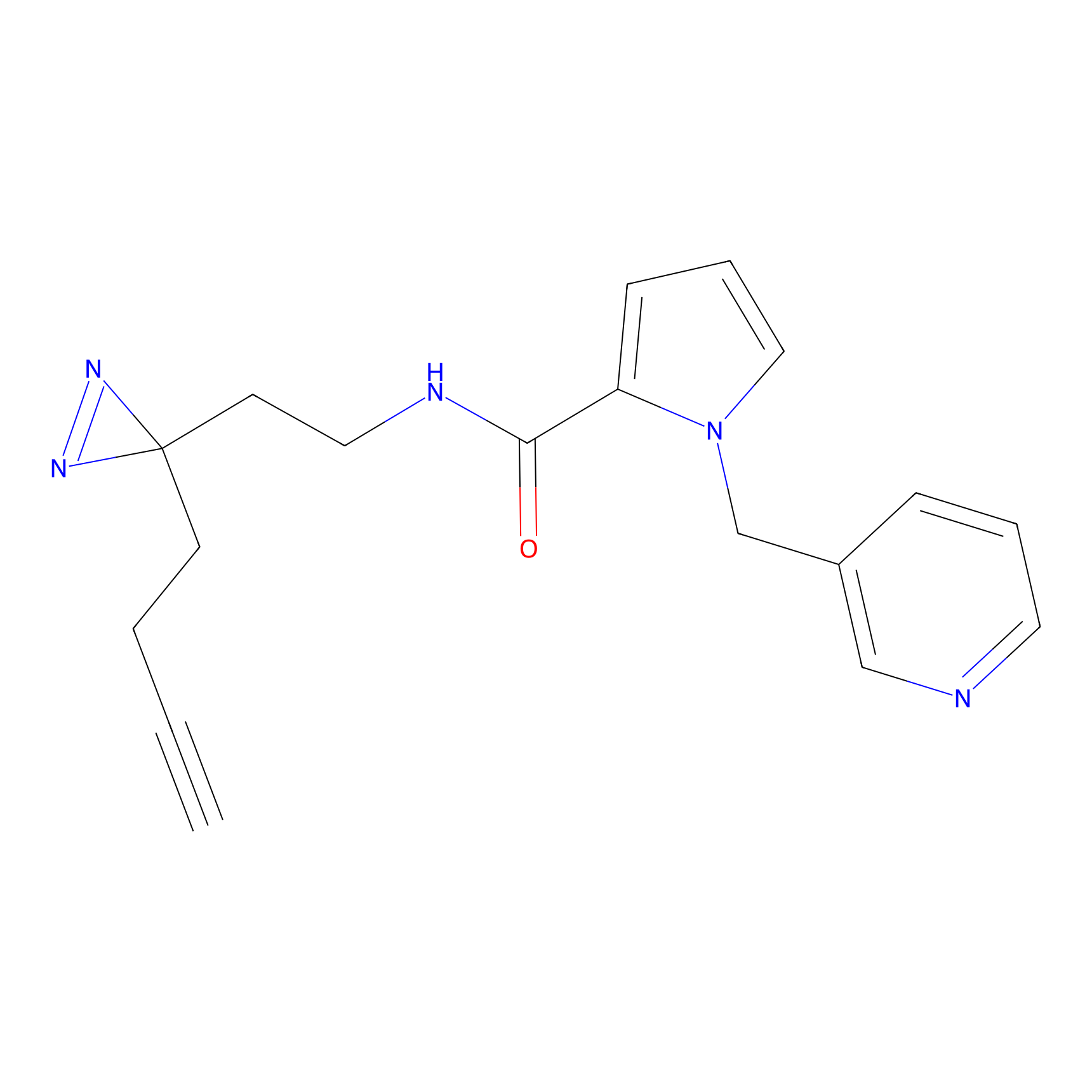
TH214 Probe Info |
 |
Y27(13.90) | LDD0258 | [1] | |
|
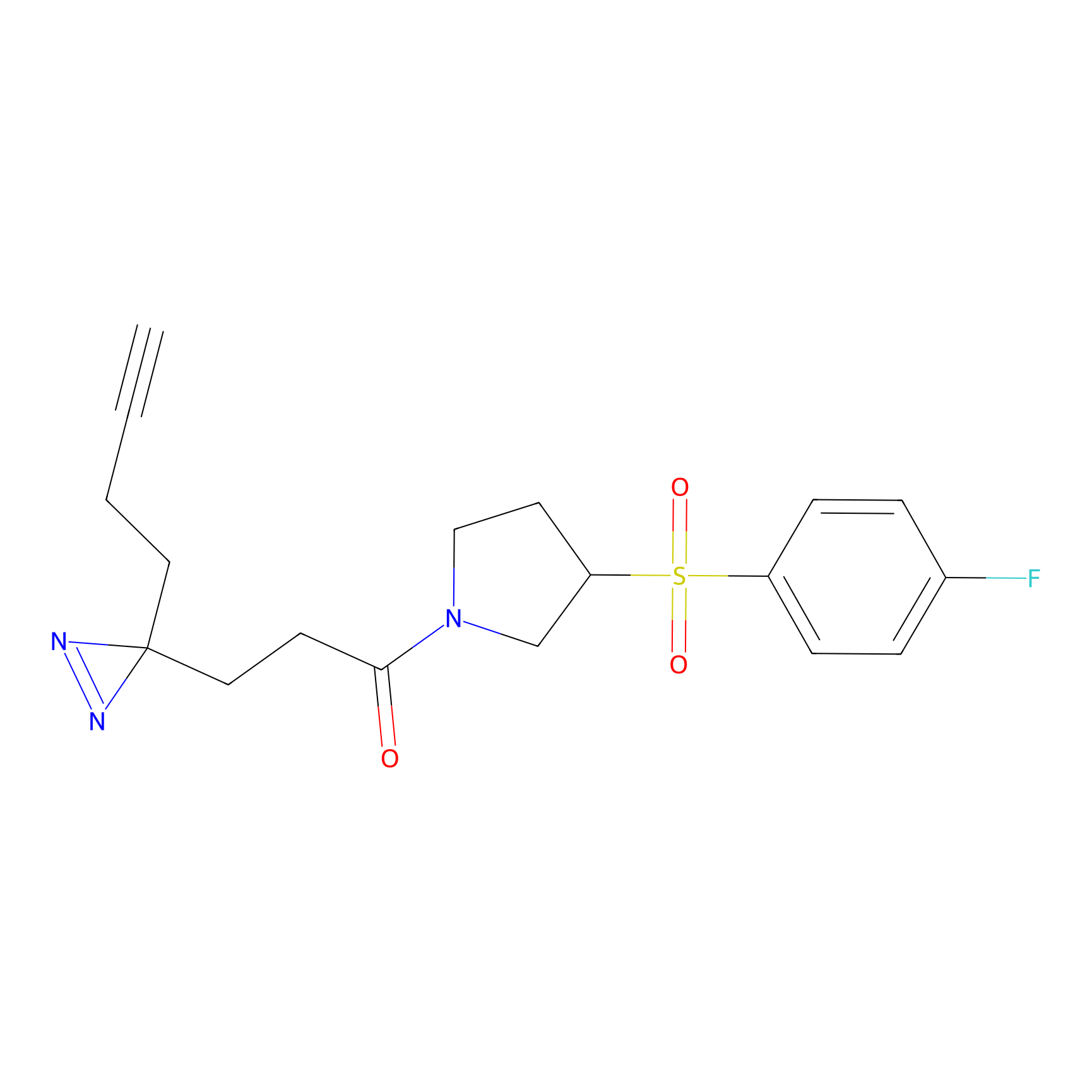
TH216 Probe Info |
 |
Y27(14.04) | LDD0259 | [1] | |
|
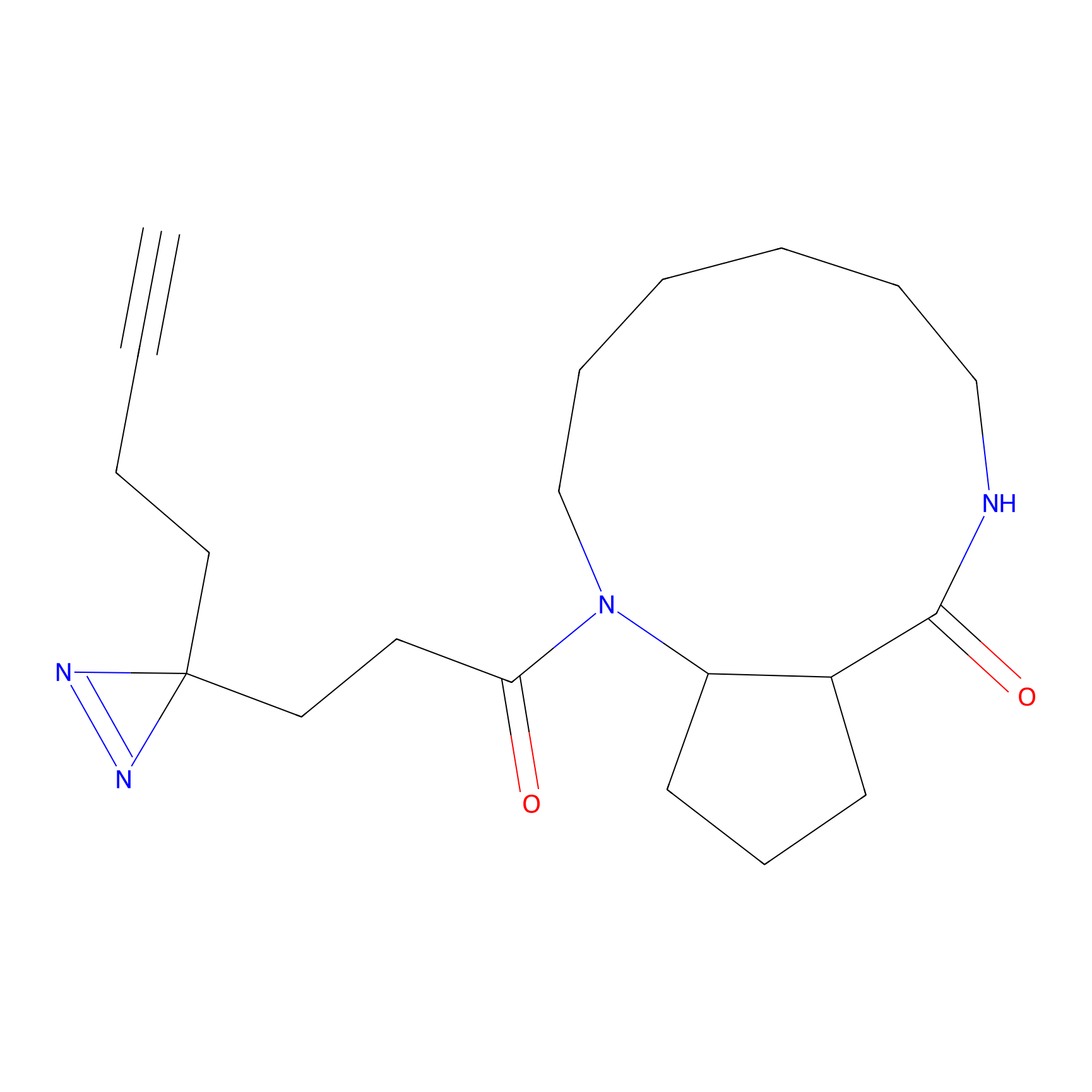
ONAyne Probe Info |
 |
K36(4.85); K53(4.41) | LDD0274 | [2] | |
|
STPyne Probe Info |
 |
K48(7.82); K53(9.21) | LDD0277 | [2] | |
|
DBIA Probe Info |
 |
C23(1.13) | LDD3397 | [3] | |
|
BTD Probe Info |
 |
C23(2.77) | LDD1700 | [4] | |
|
HHS-482 Probe Info |
 |
Y27(1.44); Y83(0.96) | LDD0285 | [5] | |
|
HHS-475 Probe Info |
 |
Y27(0.77); Y83(0.94); Y81(1.00) | LDD0264 | [6] | |
|
HHS-465 Probe Info |
 |
Y27(9.10); Y81(5.81); Y83(4.78) | LDD2237 | [7] | |
|
5E-2FA Probe Info |
 |
H26(0.00); H88(0.00); H95(0.00) | LDD2235 | [8] | |
|
ATP probe Probe Info |
 |
K35(0.00); K36(0.00); K53(0.00) | LDD0199 | [9] | |
|
m-APA Probe Info |
 |
H26(0.00); H88(0.00) | LDD2231 | [8] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [10] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [10] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [10] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [11] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [12] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [13] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [14] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C23(0.35) | LDD2142 | [4] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C23(0.51) | LDD2112 | [4] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C23(1.40) | LDD2117 | [4] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C23(1.56) | LDD2152 | [4] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C23(1.09) | LDD2103 | [4] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C23(0.45) | LDD2132 | [4] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C23(0.68) | LDD2131 | [4] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C23(0.57) | LDD2113 | [4] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C23(0.46) | LDD2091 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [13] |
| LDCM0168 | Crenolanib | NCI-H1703 | 3.54 | LDD0429 | [17] |
| LDCM0625 | F8 | Ramos | C23(3.32) | LDD2187 | [20] |
| LDCM0572 | Fragment10 | Ramos | C23(0.28) | LDD2189 | [20] |
| LDCM0573 | Fragment11 | Ramos | C23(0.02) | LDD2190 | [20] |
| LDCM0574 | Fragment12 | Ramos | C23(0.45) | LDD2191 | [20] |
| LDCM0575 | Fragment13 | Ramos | C23(0.53) | LDD2192 | [20] |
| LDCM0576 | Fragment14 | Ramos | C23(0.77) | LDD2193 | [20] |
| LDCM0579 | Fragment20 | Ramos | C23(0.33) | LDD2194 | [20] |
| LDCM0586 | Fragment28 | Ramos | C23(0.72) | LDD2198 | [20] |
| LDCM0588 | Fragment30 | Ramos | C23(0.92) | LDD2199 | [20] |
| LDCM0590 | Fragment32 | Ramos | C23(0.42) | LDD2201 | [20] |
| LDCM0566 | Fragment4 | Ramos | C23(0.53) | LDD2184 | [20] |
| LDCM0614 | Fragment56 | Ramos | C23(0.41) | LDD2205 | [20] |
| LDCM0569 | Fragment7 | Ramos | C23(0.49) | LDD2186 | [20] |
| LDCM0571 | Fragment9 | Ramos | C23(0.19) | LDD2188 | [20] |
| LDCM0116 | HHS-0101 | DM93 | Y27(0.77); Y83(0.94); Y81(1.00) | LDD0264 | [6] |
| LDCM0117 | HHS-0201 | DM93 | Y27(0.73); Y83(0.91); Y81(1.58) | LDD0265 | [6] |
| LDCM0118 | HHS-0301 | DM93 | Y27(0.65); Y81(0.81); Y83(1.15) | LDD0266 | [6] |
| LDCM0119 | HHS-0401 | DM93 | Y27(0.92); Y81(1.05); Y83(1.11) | LDD0267 | [6] |
| LDCM0120 | HHS-0701 | DM93 | Y27(0.89); Y83(1.00); Y81(1.31) | LDD0268 | [6] |
| LDCM0123 | JWB131 | DM93 | Y27(1.44); Y83(0.96) | LDD0285 | [5] |
| LDCM0124 | JWB142 | DM93 | Y27(0.56); Y83(0.65) | LDD0286 | [5] |
| LDCM0125 | JWB146 | DM93 | Y27(1.16); Y83(1.44) | LDD0287 | [5] |
| LDCM0126 | JWB150 | DM93 | Y27(3.56); Y83(4.15) | LDD0288 | [5] |
| LDCM0127 | JWB152 | DM93 | Y27(2.21); Y83(1.34) | LDD0289 | [5] |
| LDCM0128 | JWB198 | DM93 | Y27(1.57); Y83(1.74) | LDD0290 | [5] |
| LDCM0129 | JWB202 | DM93 | Y27(0.40) | LDD0291 | [5] |
| LDCM0130 | JWB211 | DM93 | Y27(0.81); Y83(0.81) | LDD0292 | [5] |
| LDCM0022 | KB02 | HEK-293T | C23(1.03) | LDD1492 | [21] |
| LDCM0023 | KB03 | HEK-293T | C23(0.98) | LDD1497 | [21] |
| LDCM0024 | KB05 | PF-382 | C23(1.13) | LDD3397 | [3] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C23(1.32) | LDD2102 | [4] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C23(1.38) | LDD2090 | [4] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C23(1.42) | LDD2092 | [4] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C23(1.52) | LDD2093 | [4] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C23(2.33) | LDD2094 | [4] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C23(0.63) | LDD2096 | [4] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C23(0.75) | LDD2097 | [4] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C23(0.74) | LDD2098 | [4] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C23(1.18) | LDD2099 | [4] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C23(0.40) | LDD2100 | [4] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C23(0.89) | LDD2101 | [4] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C23(0.42) | LDD2104 | [4] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C23(0.26) | LDD2106 | [4] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C23(1.23) | LDD2107 | [4] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C23(0.56) | LDD2108 | [4] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C23(0.32) | LDD2110 | [4] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C23(1.26) | LDD2111 | [4] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C23(0.50) | LDD2114 | [4] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C23(0.44) | LDD2115 | [4] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C23(0.89) | LDD2116 | [4] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C23(1.03) | LDD2118 | [4] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C23(1.48) | LDD2119 | [4] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C23(0.91) | LDD2120 | [4] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C23(0.73) | LDD2122 | [4] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C23(0.86) | LDD2123 | [4] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C23(0.87) | LDD2124 | [4] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C23(1.00) | LDD2125 | [4] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C23(0.92) | LDD2126 | [4] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C23(1.29) | LDD2127 | [4] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C23(1.04) | LDD2128 | [4] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C23(1.20) | LDD2129 | [4] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C23(0.57) | LDD2133 | [4] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C23(0.60) | LDD2134 | [4] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C23(1.45) | LDD2135 | [4] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C23(1.27) | LDD2136 | [4] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C23(1.10) | LDD2137 | [4] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C23(2.77) | LDD1700 | [4] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C23(0.99) | LDD2140 | [4] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C23(0.42) | LDD2141 | [4] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C23(1.02) | LDD2143 | [4] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C23(3.07) | LDD2144 | [4] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C23(0.20) | LDD2145 | [4] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C23(1.38) | LDD2146 | [4] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C23(1.53) | LDD2147 | [4] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C23(0.44) | LDD2148 | [4] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C23(1.12) | LDD2149 | [4] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C23(0.76) | LDD2150 | [4] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C23(0.71) | LDD2151 | [4] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C23(1.70) | LDD2153 | [4] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Amyloid-beta precursor protein (APP) | APP family | P05067 | |||
| Ryanodine receptor 2 (RYR2) | Ryanodine receptor family | Q92736 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Mothers against decapentaplegic homolog 7 (SMAD7) | Dwarfin/SMAD family | O15105 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Alpha-hemoglobin-stabilizing protein (AHSP) | AHSP family | Q9NZD4 | |||
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Pimecrolimus | Small molecular drug | DB00337 | |||
| Tacrolimus | Small molecular drug | DB00864 | |||
Phase 2
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Gpi-1485 | Small molecular drug | D0N0AR | |||
| Ap1903 | . | D0K8TO | |||
| Sdz-281-240 | . | D0O5OF | |||
Investigative
Discontinued
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Gpi-1046 | Small molecular drug | D04PGB | |||
References