Details of the Target
General Information of Target
| Target ID | LDTP03182 | |||||
|---|---|---|---|---|---|---|
| Target Name | Heterogeneous nuclear ribonucleoproteins A2/B1 (HNRNPA2B1) | |||||
| Gene Name | HNRNPA2B1 | |||||
| Gene ID | 3181 | |||||
| Synonyms |
HNRPA2B1; Heterogeneous nuclear ribonucleoproteins A2/B1; hnRNP A2/B1 |
|||||
| 3D Structure | ||||||
| Sequence |
MEKTLETVPLERKKREKEQFRKLFIGGLSFETTEESLRNYYEQWGKLTDCVVMRDPASKR
SRGFGFVTFSSMAEVDAAMAARPHSIDGRVVEPKRAVAREESGKPGAHVTVKKLFVGGIK EDTEEHHLRDYFEEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDKIVLQKYHTINGH NAEVRKALSRQEMQEVQSSRSGRGGNFGFGDSRGGGGNFGPGPGSNFRGGSDGYGSGRGF GDGYNGYGGGPGGGNFGGSPGYGGGRGGYGGGGPGYGNQGGGYGGGYDNYGGGNYGSGNY NDFGNYNQQPSNYGPMKSGNFGGSRNMGGPYGGGNYGPGGSGGSGGYGGRSRY |
|||||
| Target Type |
Literature-reported
|
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Heterogeneous nuclear ribonucleoprotein (hnRNP) that associates with nascent pre-mRNAs, packaging them into hnRNP particles. The hnRNP particle arrangement on nascent hnRNA is non-random and sequence-dependent and serves to condense and stabilize the transcripts and minimize tangling and knotting. Packaging plays a role in various processes such as transcription, pre-mRNA processing, RNA nuclear export, subcellular location, mRNA translation and stability of mature mRNAs. Forms hnRNP particles with at least 20 other different hnRNP and heterogeneous nuclear RNA in the nucleus. Involved in transport of specific mRNAs to the cytoplasm in oligodendrocytes and neurons: acts by specifically recognizing and binding the A2RE (21 nucleotide hnRNP A2 response element) or the A2RE11 (derivative 11 nucleotide oligonucleotide) sequence motifs present on some mRNAs, and promotes their transport to the cytoplasm. Specifically binds single-stranded telomeric DNA sequences, protecting telomeric DNA repeat against endonuclease digestion. Also binds other RNA molecules, such as primary miRNA (pri-miRNAs): acts as a nuclear 'reader' of the N6-methyladenosine (m6A) mark by specifically recognizing and binding a subset of nuclear m6A-containing pri-miRNAs. Binding to m6A-containing pri-miRNAs promotes pri-miRNA processing by enhancing binding of DGCR8 to pri-miRNA transcripts. Involved in miRNA sorting into exosomes following sumoylation, possibly by binding (m6A)-containing pre-miRNAs. Acts as a regulator of efficiency of mRNA splicing, possibly by binding to m6A-containing pre-mRNAs. Plays a role in the splicing of pyruvate kinase PKM by binding repressively to sequences flanking PKM exon 9, inhibiting exon 9 inclusion and resulting in exon 10 inclusion and production of the PKM M2 isoform. Also plays a role in the activation of the innate immune response. Mechanistically, senses the presence of viral DNA in the nucleus, homodimerizes and is demethylated by JMJD6. In turn, translocates to the cytoplasm where it activates the TBK1-IRF3 pathway, leading to interferon alpha/beta production.; (Microbial infection) Involved in the transport of HIV-1 genomic RNA out of the nucleus, to the microtubule organizing center (MTOC), and then from the MTOC to the cytoplasm: acts by specifically recognizing and binding the A2RE (21 nucleotide hnRNP A2 response element) sequence motifs present on HIV-1 genomic RNA, and promotes its transport.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P3 Probe Info |
 |
1.90 | LDD0450 | [1] | |
|
P8 Probe Info |
 |
1.94 | LDD0455 | [1] | |
|
A-EBA Probe Info |
 |
3.07 | LDD0215 | [2] | |
|
CY4 Probe Info |
 |
3.86 | LDD0244 | [3] | |
|
N1 Probe Info |
 |
2.24 | LDD0242 | [3] | |
|
C-Sul Probe Info |
 |
6.70 | LDD0066 | [4] | |
|
TH211 Probe Info |
 |
Y244(20.00); Y247(20.00); Y262(20.00); Y331(20.00) | LDD0257 | [5] | |
|
TH214 Probe Info |
 |
Y174(20.00) | LDD0258 | [5] | |
|
TH216 Probe Info |
 |
Y174(20.00); Y234(20.00); Y336(20.00) | LDD0259 | [5] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [6] | |
|
1oxF11yne Probe Info |
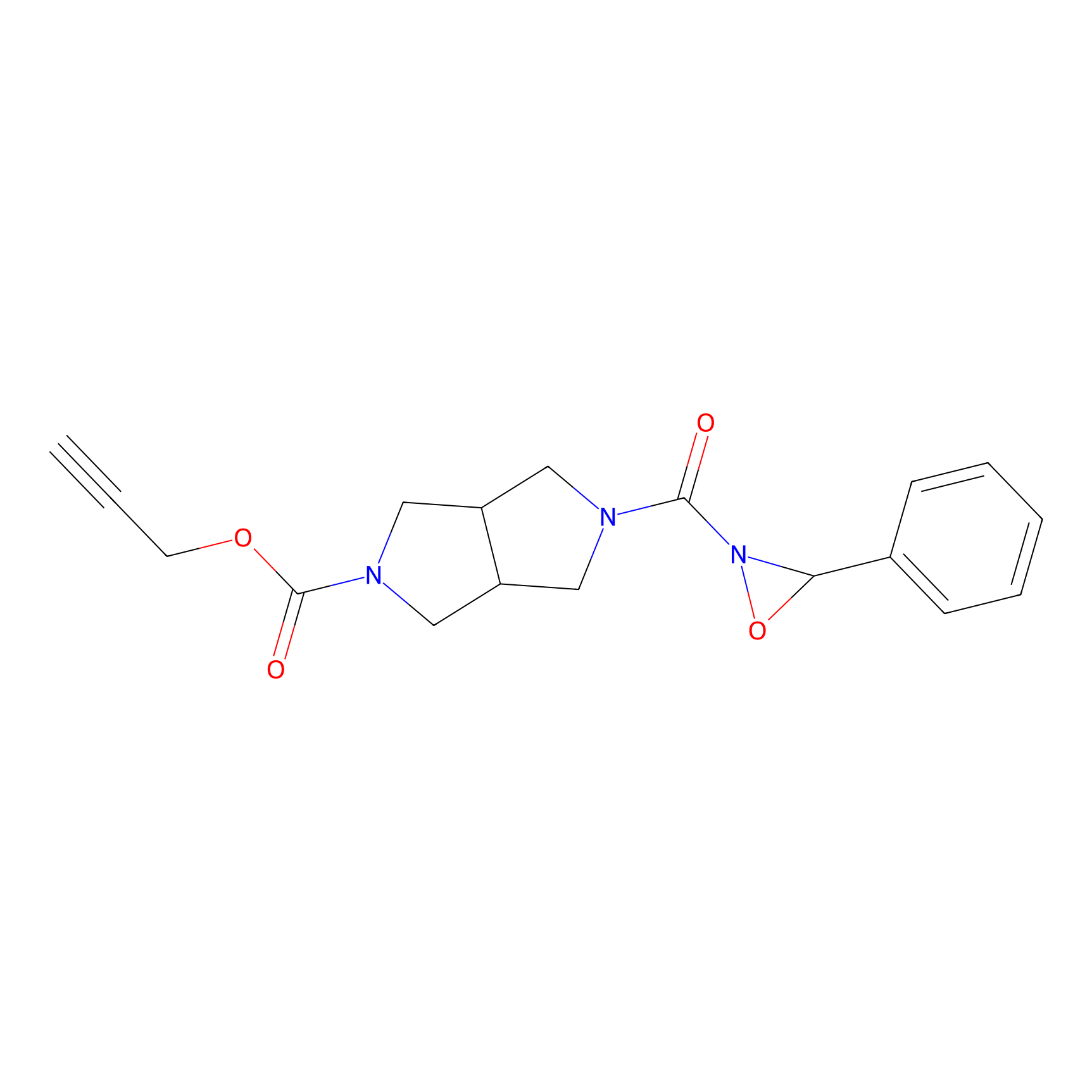 |
N.A. | LDD0193 | [7] | |
|
AZ-9 Probe Info |
 |
E134(1.02); E42(0.91); D211(1.22); E142(0.42) | LDD2208 | [8] | |
|
ONAyne Probe Info |
 |
K17(0.00); K104(0.00); K59(0.00); K112(0.00) | LDD0273 | [9] | |
|
OPA-S-S-alkyne Probe Info |
 |
K120(5.33); K173(7.67) | LDD3494 | [10] | |
|
Probe 1 Probe Info |
 |
Y40(55.53); Y41(28.20); Y131(18.54); Y135(47.82) | LDD3495 | [11] | |
|
BTD Probe Info |
 |
C50(2.07) | LDD1700 | [12] | |
|
HHS-482 Probe Info |
 |
Y174(1.01); Y234(0.87); Y331(0.80); Y336(0.81) | LDD0285 | [13] | |
|
HHS-475 Probe Info |
 |
Y331(0.71); Y174(0.74); Y347(0.74); Y336(0.75) | LDD0264 | [14] | |
|
HHS-465 Probe Info |
 |
Y174(8.02); Y234(10.00); Y336(10.00); Y347(10.00) | LDD2237 | [15] | |
|
DBIA Probe Info |
 |
C50(2.14) | LDD0531 | [16] | |
|
5E-2FA Probe Info |
 |
H108(0.00); H163(0.00); H84(0.00); H180(0.00) | LDD2235 | [17] | |
|
ATP probe Probe Info |
 |
K22(0.00); K173(0.00); K104(0.00); K112(0.00) | LDD0199 | [18] | |
|
m-APA Probe Info |
 |
H163(0.00); H126(0.00); H180(0.00); H108(0.00) | LDD2231 | [17] | |
|
CY-1 Probe Info |
 |
M327(0.00); Y336(0.00); N335(0.00) | LDD0246 | [3] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [19] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [20] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [21] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [21] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [19] | |
|
2PCA Probe Info |
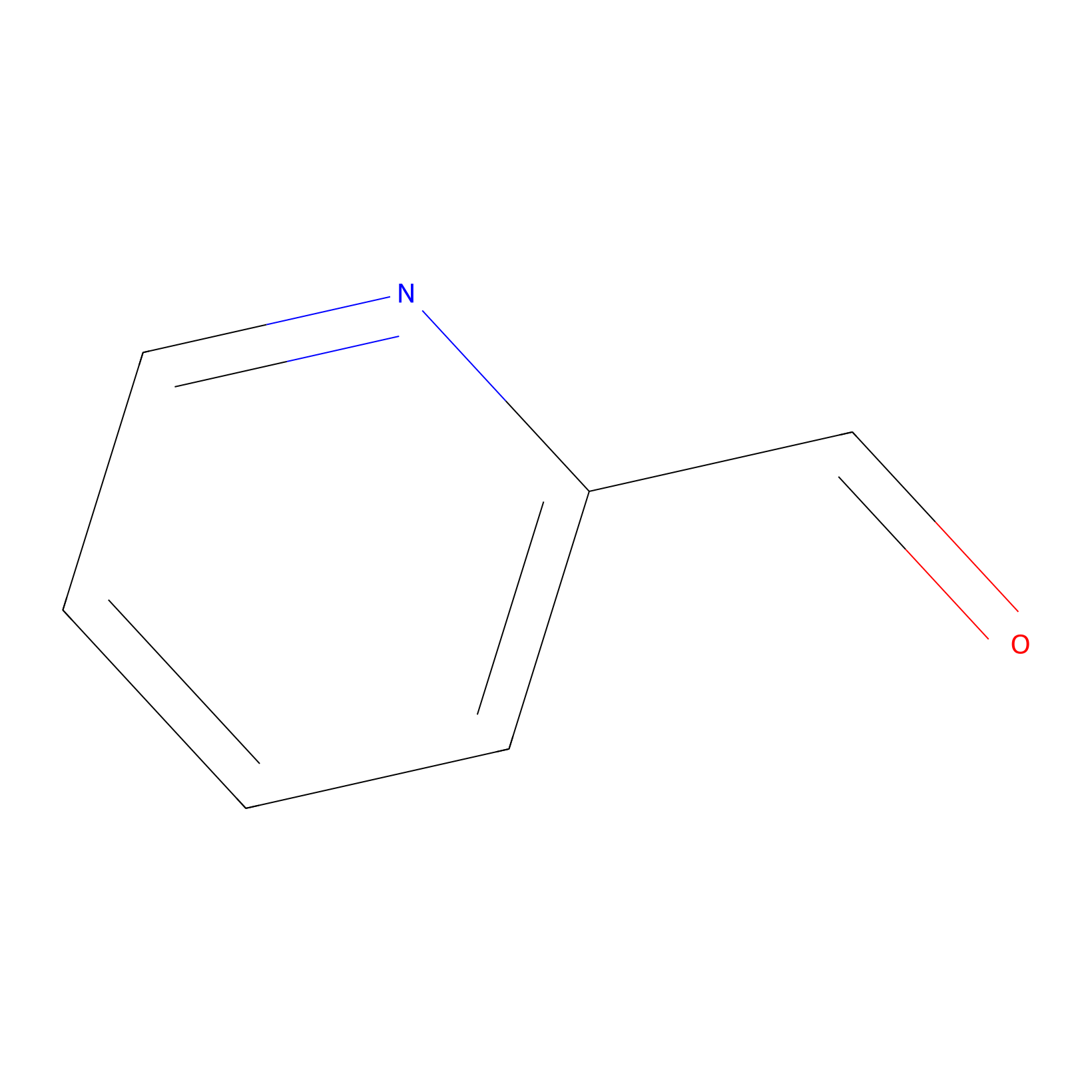 |
N.A. | LDD0034 | [22] | |
|
ATP probe Probe Info |
 |
K173(0.00); K168(0.00); K59(0.00) | LDD0035 | [23] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [24] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [24] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [25] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [25] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [26] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [27] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [25] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [25] | |
|
NHS Probe Info |
 |
K113(0.00); K112(0.00); K3(0.00); K104(0.00) | LDD0010 | [25] | |
|
OSF Probe Info |
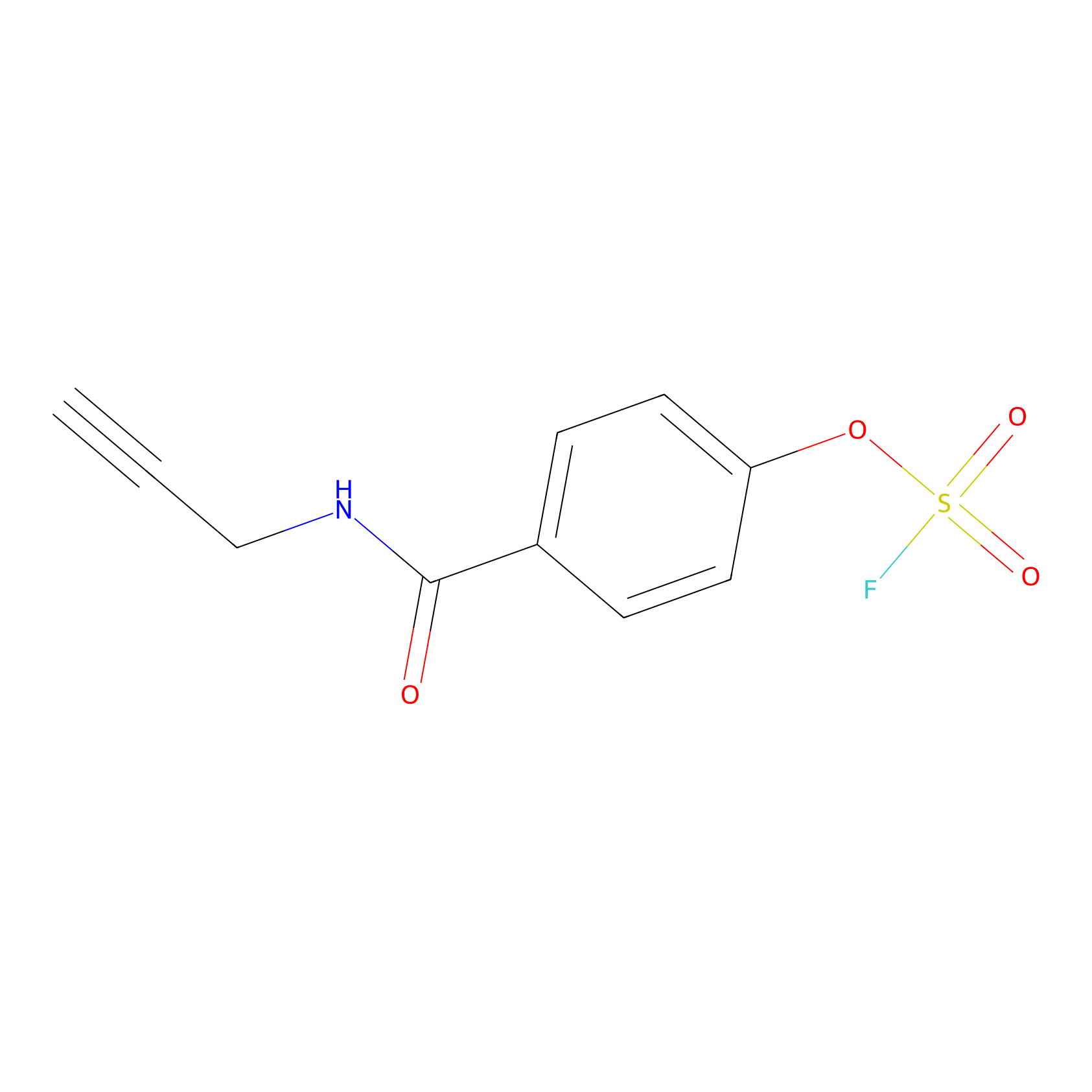 |
Y234(0.00); Y336(0.00); Y331(0.00) | LDD0029 | [28] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0152 | [29] | |
|
PPMS Probe Info |
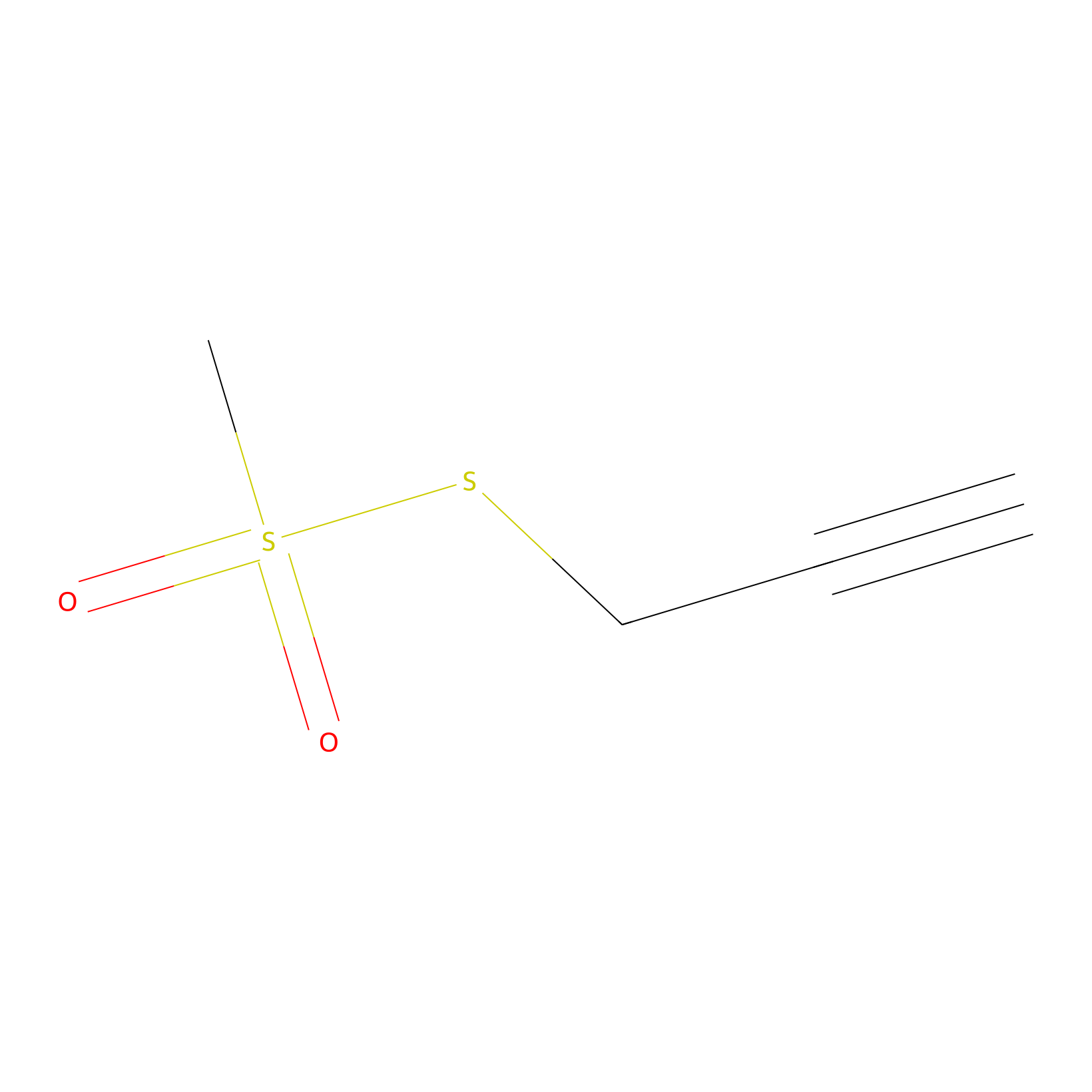 |
N.A. | LDD0008 | [25] | |
|
SF Probe Info |
 |
Y331(0.00); Y295(0.00); Y244(0.00); K59(0.00) | LDD0028 | [28] | |
|
STPyne Probe Info |
 |
K112(0.00); K113(0.00) | LDD0009 | [25] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [25] | |
|
YN-1 Probe Info |
 |
N.A. | LDD0447 | [6] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [30] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [31] | |
|
1c-yne Probe Info |
 |
K46(0.00); K173(0.00); K22(0.00); K168(0.00) | LDD0228 | [26] | |
|
Acrolein Probe Info |
 |
C50(0.00); H163(0.00); H180(0.00); H108(0.00) | LDD0217 | [32] | |
|
Cinnamaldehyde Probe Info |
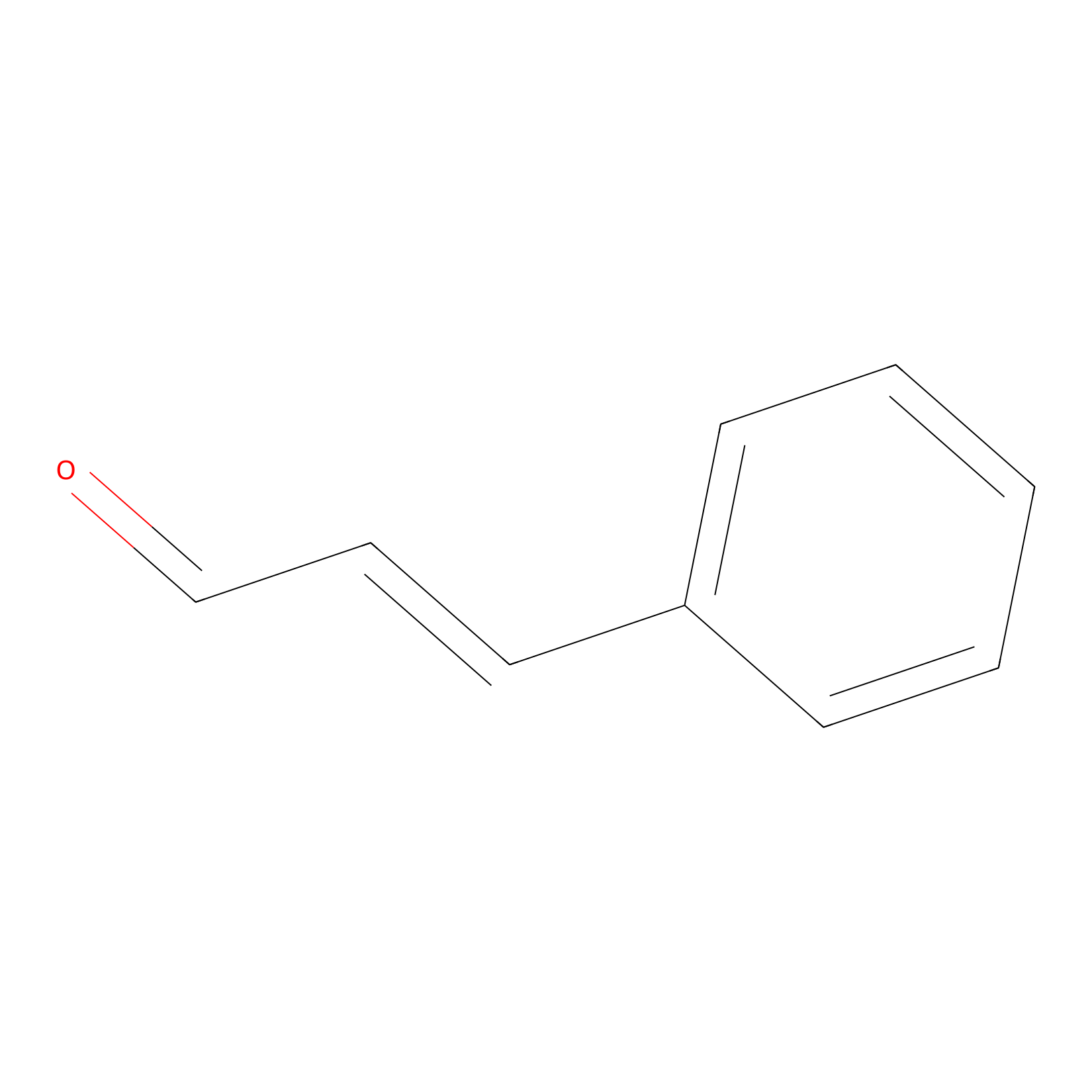 |
N.A. | LDD0220 | [32] | |
|
Crotonaldehyde Probe Info |
 |
C50(0.00); H175(0.00); H180(0.00); H108(0.00) | LDD0219 | [32] | |
|
Methacrolein Probe Info |
 |
C50(0.00); H180(0.00); H163(0.00); H108(0.00) | LDD0218 | [32] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [33] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [34] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [35] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [34] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C165 Probe Info |
 |
17.27 | LDD1845 | [36] | |
|
C191 Probe Info |
 |
11.63 | LDD1868 | [36] | |
|
C218 Probe Info |
 |
11.79 | LDD1892 | [36] | |
|
C220 Probe Info |
 |
12.47 | LDD1894 | [36] | |
|
C232 Probe Info |
 |
37.53 | LDD1905 | [36] | |
|
C338 Probe Info |
 |
18.77 | LDD2001 | [36] | |
|
C346 Probe Info |
 |
10.63 | LDD2007 | [36] | |
|
C348 Probe Info |
 |
10.41 | LDD2009 | [36] | |
|
C353 Probe Info |
 |
5.78 | LDD2014 | [36] | |
|
C355 Probe Info |
 |
22.47 | LDD2016 | [36] | |
|
C361 Probe Info |
 |
18.25 | LDD2022 | [36] | |
|
C382 Probe Info |
 |
8.57 | LDD2041 | [36] | |
|
C391 Probe Info |
 |
13.64 | LDD2050 | [36] | |
|
FFF probe11 Probe Info |
 |
13.86 | LDD0471 | [37] | |
|
FFF probe12 Probe Info |
 |
6.08 | LDD0473 | [37] | |
|
FFF probe13 Probe Info |
 |
16.00 | LDD0475 | [37] | |
|
FFF probe14 Probe Info |
 |
10.96 | LDD0477 | [37] | |
|
FFF probe2 Probe Info |
 |
18.09 | LDD0463 | [37] | |
|
FFF probe3 Probe Info |
 |
11.65 | LDD0464 | [37] | |
|
FFF probe6 Probe Info |
 |
7.82 | LDD0468 | [37] | |
|
FFF probe9 Probe Info |
 |
5.83 | LDD0470 | [37] | |
|
JN0003 Probe Info |
 |
13.71 | LDD0469 | [37] | |
|
STS-2 Probe Info |
 |
1.44 | LDD0139 | [38] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [39] | |
|
Diazir Probe Info |
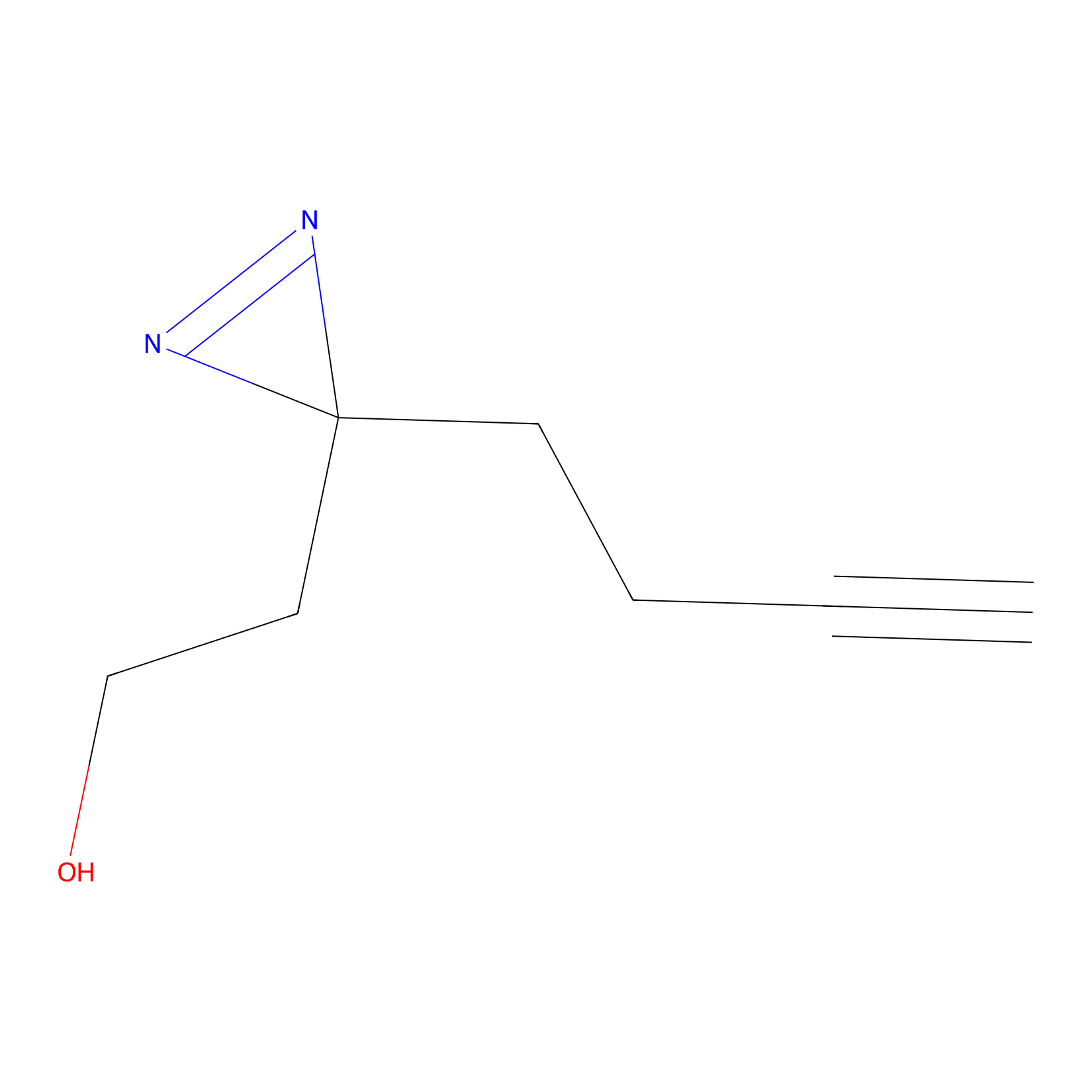 |
Y347(0.00); E195(0.00) | LDD0011 | [25] | |
|
BD-F Probe Info |
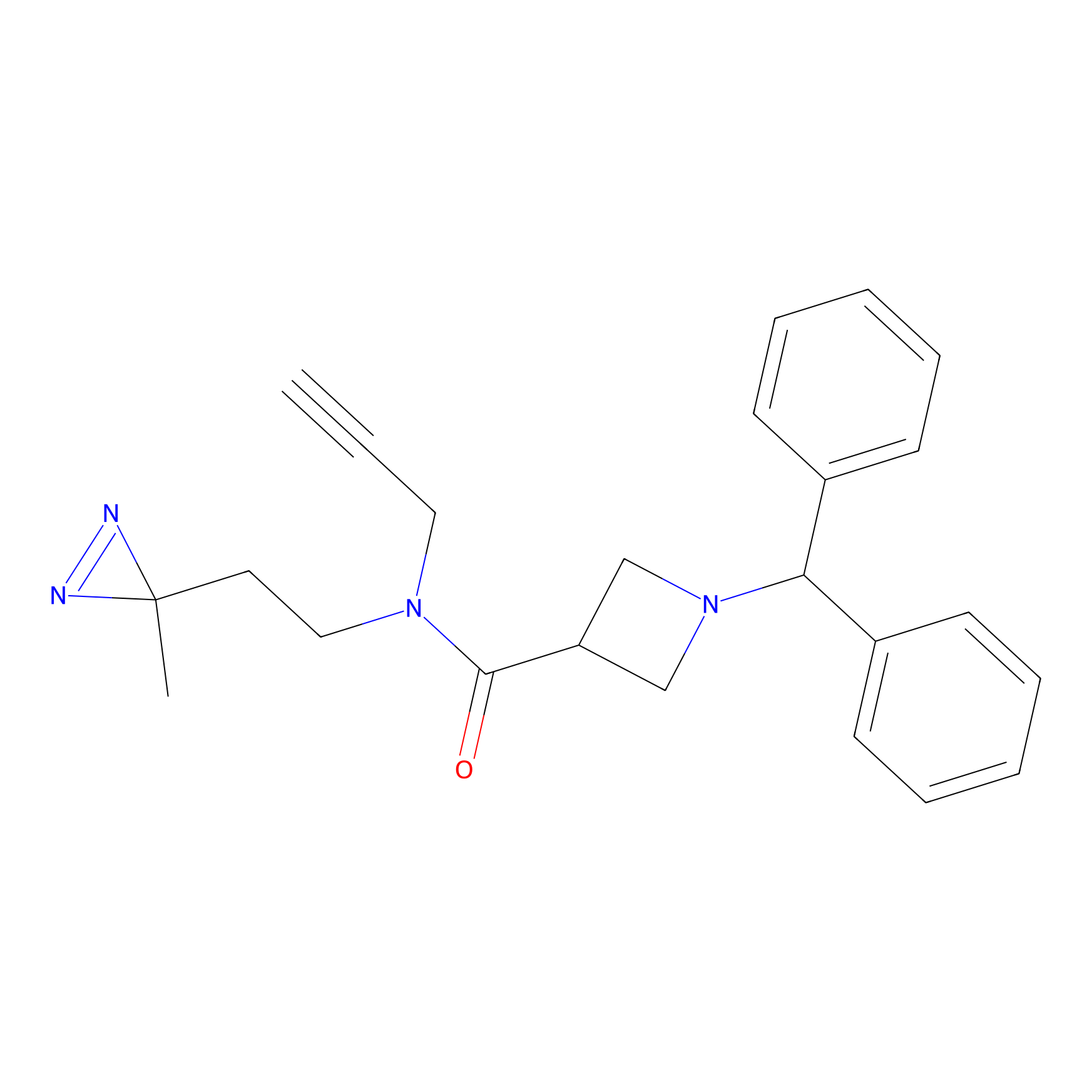 |
F256(0.00); G337(0.00); P251(0.00); G243(0.00) | LDD0024 | [40] | |
|
LD-F Probe Info |
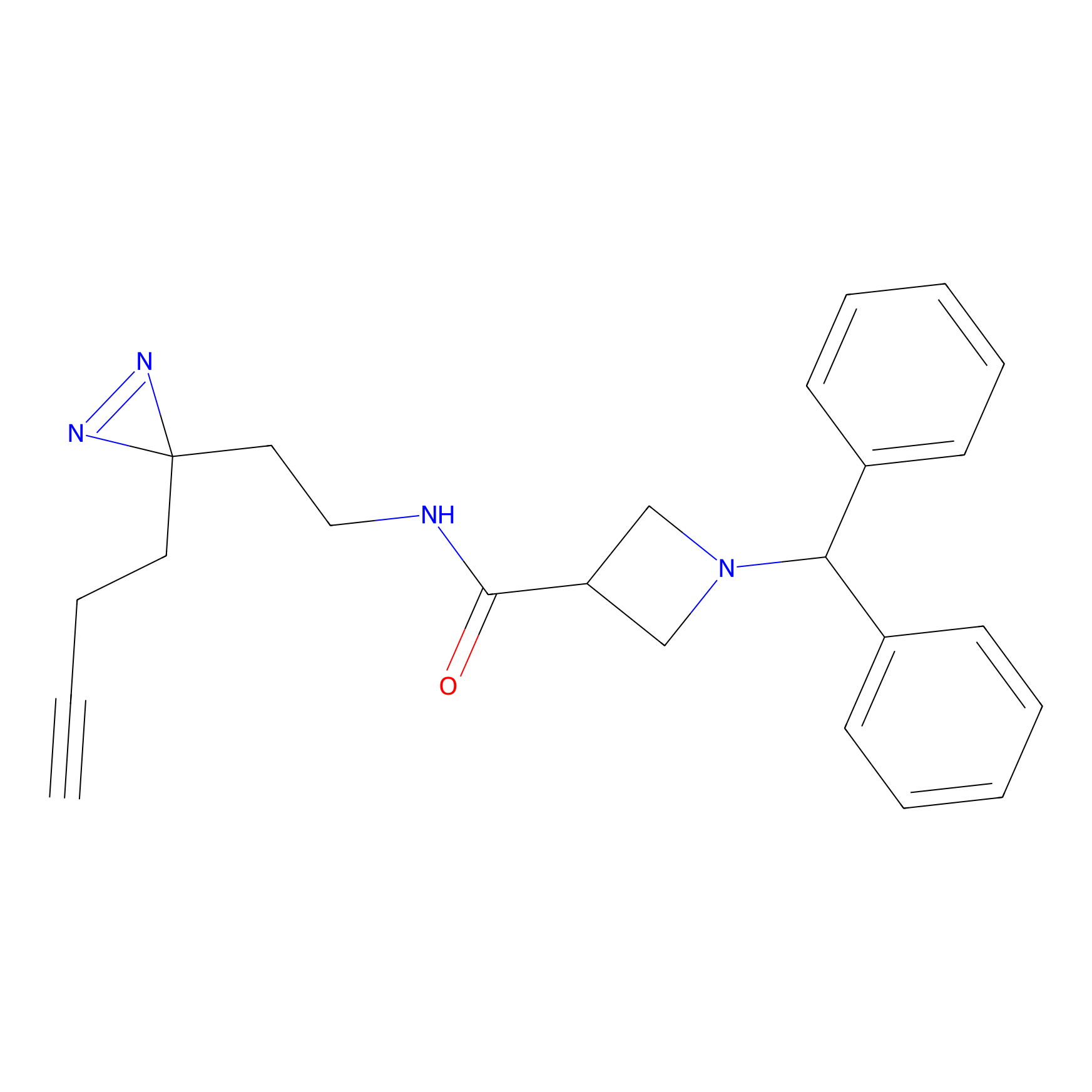 |
G337(0.00); G334(0.00); N335(0.00); N245(0.00) | LDD0015 | [40] | |
|
Photocelecoxib Probe Info |
 |
N.A. | LDD0153 | [41] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [42] | |
|
OEA-DA Probe Info |
 |
4.24 | LDD0046 | [43] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0069 | [44] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C50(0.81) | LDD2112 | [12] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C50(0.49) | LDD2095 | [12] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C50(1.10) | LDD2130 | [12] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C50(0.87) | LDD2117 | [12] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C50(0.39) | LDD2132 | [12] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C50(0.88) | LDD2131 | [12] |
| LDCM0214 | AC1 | HCT 116 | C50(2.14) | LDD0531 | [16] |
| LDCM0215 | AC10 | HCT 116 | C50(1.26) | LDD0532 | [16] |
| LDCM0216 | AC100 | HCT 116 | C50(0.90) | LDD0533 | [16] |
| LDCM0217 | AC101 | HCT 116 | C50(0.70) | LDD0534 | [16] |
| LDCM0218 | AC102 | HCT 116 | C50(0.73) | LDD0535 | [16] |
| LDCM0219 | AC103 | HCT 116 | C50(0.49) | LDD0536 | [16] |
| LDCM0220 | AC104 | HCT 116 | C50(0.63) | LDD0537 | [16] |
| LDCM0221 | AC105 | HCT 116 | C50(0.66) | LDD0538 | [16] |
| LDCM0222 | AC106 | HCT 116 | C50(0.54) | LDD0539 | [16] |
| LDCM0223 | AC107 | HCT 116 | C50(0.52) | LDD0540 | [16] |
| LDCM0224 | AC108 | HCT 116 | C50(0.59) | LDD0541 | [16] |
| LDCM0225 | AC109 | HCT 116 | C50(0.63) | LDD0542 | [16] |
| LDCM0226 | AC11 | HCT 116 | C50(2.03) | LDD0543 | [16] |
| LDCM0227 | AC110 | HCT 116 | C50(0.52) | LDD0544 | [16] |
| LDCM0228 | AC111 | HCT 116 | C50(0.38) | LDD0545 | [16] |
| LDCM0229 | AC112 | HCT 116 | C50(0.40) | LDD0546 | [16] |
| LDCM0230 | AC113 | HCT 116 | C50(1.33) | LDD0547 | [16] |
| LDCM0231 | AC114 | HCT 116 | C50(1.29) | LDD0548 | [16] |
| LDCM0232 | AC115 | HCT 116 | C50(1.06) | LDD0549 | [16] |
| LDCM0233 | AC116 | HCT 116 | C50(1.16) | LDD0550 | [16] |
| LDCM0234 | AC117 | HCT 116 | C50(1.39) | LDD0551 | [16] |
| LDCM0235 | AC118 | HCT 116 | C50(1.56) | LDD0552 | [16] |
| LDCM0236 | AC119 | HCT 116 | C50(1.54) | LDD0553 | [16] |
| LDCM0237 | AC12 | HCT 116 | C50(1.69) | LDD0554 | [16] |
| LDCM0238 | AC120 | HCT 116 | C50(1.13) | LDD0555 | [16] |
| LDCM0239 | AC121 | HCT 116 | C50(1.16) | LDD0556 | [16] |
| LDCM0240 | AC122 | HCT 116 | C50(1.31) | LDD0557 | [16] |
| LDCM0241 | AC123 | HCT 116 | C50(1.26) | LDD0558 | [16] |
| LDCM0242 | AC124 | HCT 116 | C50(1.44) | LDD0559 | [16] |
| LDCM0243 | AC125 | HCT 116 | C50(1.28) | LDD0560 | [16] |
| LDCM0244 | AC126 | HCT 116 | C50(1.08) | LDD0561 | [16] |
| LDCM0245 | AC127 | HCT 116 | C50(1.37) | LDD0562 | [16] |
| LDCM0246 | AC128 | HCT 116 | C50(1.29) | LDD0563 | [16] |
| LDCM0247 | AC129 | HCT 116 | C50(1.34) | LDD0564 | [16] |
| LDCM0249 | AC130 | HCT 116 | C50(1.43) | LDD0566 | [16] |
| LDCM0250 | AC131 | HCT 116 | C50(1.27) | LDD0567 | [16] |
| LDCM0251 | AC132 | HCT 116 | C50(0.98) | LDD0568 | [16] |
| LDCM0252 | AC133 | HCT 116 | C50(1.51) | LDD0569 | [16] |
| LDCM0253 | AC134 | HCT 116 | C50(1.09) | LDD0570 | [16] |
| LDCM0254 | AC135 | HCT 116 | C50(1.15) | LDD0571 | [16] |
| LDCM0255 | AC136 | HCT 116 | C50(1.19) | LDD0572 | [16] |
| LDCM0256 | AC137 | HCT 116 | C50(1.04) | LDD0573 | [16] |
| LDCM0257 | AC138 | HCT 116 | C50(0.63) | LDD0574 | [16] |
| LDCM0258 | AC139 | HCT 116 | C50(0.79) | LDD0575 | [16] |
| LDCM0259 | AC14 | HCT 116 | C50(1.26) | LDD0576 | [16] |
| LDCM0260 | AC140 | HCT 116 | C50(0.58) | LDD0577 | [16] |
| LDCM0261 | AC141 | HCT 116 | C50(0.60) | LDD0578 | [16] |
| LDCM0262 | AC142 | HCT 116 | C50(1.17) | LDD0579 | [16] |
| LDCM0263 | AC143 | HCT 116 | C50(1.06) | LDD0580 | [16] |
| LDCM0264 | AC144 | HCT 116 | C50(0.76) | LDD0581 | [16] |
| LDCM0265 | AC145 | HCT 116 | C50(0.91) | LDD0582 | [16] |
| LDCM0266 | AC146 | HCT 116 | C50(1.08) | LDD0583 | [16] |
| LDCM0267 | AC147 | HCT 116 | C50(0.77) | LDD0584 | [16] |
| LDCM0268 | AC148 | HCT 116 | C50(0.52) | LDD0585 | [16] |
| LDCM0269 | AC149 | HCT 116 | C50(0.77) | LDD0586 | [16] |
| LDCM0270 | AC15 | HCT 116 | C50(1.29) | LDD0587 | [16] |
| LDCM0271 | AC150 | HCT 116 | C50(2.51) | LDD0588 | [16] |
| LDCM0272 | AC151 | HCT 116 | C50(1.09) | LDD0589 | [16] |
| LDCM0273 | AC152 | HCT 116 | C50(0.82) | LDD0590 | [16] |
| LDCM0274 | AC153 | HCT 116 | C50(0.77) | LDD0591 | [16] |
| LDCM0621 | AC154 | HCT 116 | C50(0.92) | LDD2158 | [16] |
| LDCM0622 | AC155 | HCT 116 | C50(1.34) | LDD2159 | [16] |
| LDCM0623 | AC156 | HCT 116 | C50(1.41) | LDD2160 | [16] |
| LDCM0624 | AC157 | HCT 116 | C50(1.13) | LDD2161 | [16] |
| LDCM0276 | AC17 | HCT 116 | C50(1.18) | LDD0593 | [16] |
| LDCM0277 | AC18 | HCT 116 | C50(1.02) | LDD0594 | [16] |
| LDCM0278 | AC19 | HCT 116 | C50(1.07) | LDD0595 | [16] |
| LDCM0279 | AC2 | HCT 116 | C50(1.31) | LDD0596 | [16] |
| LDCM0280 | AC20 | HCT 116 | C50(1.08) | LDD0597 | [16] |
| LDCM0281 | AC21 | HCT 116 | C50(0.99) | LDD0598 | [16] |
| LDCM0282 | AC22 | HCT 116 | C50(1.15) | LDD0599 | [16] |
| LDCM0283 | AC23 | HCT 116 | C50(1.04) | LDD0600 | [16] |
| LDCM0284 | AC24 | HCT 116 | C50(0.97) | LDD0601 | [16] |
| LDCM0285 | AC25 | HCT 116 | C50(1.06) | LDD0602 | [16] |
| LDCM0286 | AC26 | HCT 116 | C50(0.92) | LDD0603 | [16] |
| LDCM0287 | AC27 | HCT 116 | C50(0.89) | LDD0604 | [16] |
| LDCM0288 | AC28 | HCT 116 | C50(0.86) | LDD0605 | [16] |
| LDCM0289 | AC29 | HCT 116 | C50(0.94) | LDD0606 | [16] |
| LDCM0290 | AC3 | HCT 116 | C50(1.99) | LDD0607 | [16] |
| LDCM0291 | AC30 | HCT 116 | C50(0.69) | LDD0608 | [16] |
| LDCM0292 | AC31 | HCT 116 | C50(0.95) | LDD0609 | [16] |
| LDCM0293 | AC32 | HCT 116 | C50(0.69) | LDD0610 | [16] |
| LDCM0294 | AC33 | HCT 116 | C50(0.59) | LDD0611 | [16] |
| LDCM0295 | AC34 | HCT 116 | C50(0.52) | LDD0612 | [16] |
| LDCM0296 | AC35 | HCT 116 | C50(1.65) | LDD0613 | [16] |
| LDCM0297 | AC36 | HCT 116 | C50(1.23) | LDD0614 | [16] |
| LDCM0298 | AC37 | HCT 116 | C50(1.43) | LDD0615 | [16] |
| LDCM0299 | AC38 | HCT 116 | C50(1.10) | LDD0616 | [16] |
| LDCM0300 | AC39 | HCT 116 | C50(1.02) | LDD0617 | [16] |
| LDCM0301 | AC4 | HCT 116 | C50(2.03) | LDD0618 | [16] |
| LDCM0302 | AC40 | HCT 116 | C50(0.98) | LDD0619 | [16] |
| LDCM0303 | AC41 | HCT 116 | C50(1.09) | LDD0620 | [16] |
| LDCM0304 | AC42 | HCT 116 | C50(0.96) | LDD0621 | [16] |
| LDCM0305 | AC43 | HCT 116 | C50(0.88) | LDD0622 | [16] |
| LDCM0306 | AC44 | HCT 116 | C50(1.29) | LDD0623 | [16] |
| LDCM0307 | AC45 | HCT 116 | C50(1.48) | LDD0624 | [16] |
| LDCM0308 | AC46 | HCT 116 | C50(0.80) | LDD0625 | [16] |
| LDCM0309 | AC47 | HCT 116 | C50(1.26) | LDD0626 | [16] |
| LDCM0310 | AC48 | HCT 116 | C50(0.88) | LDD0627 | [16] |
| LDCM0311 | AC49 | HCT 116 | C50(0.75) | LDD0628 | [16] |
| LDCM0312 | AC5 | HCT 116 | C50(1.98) | LDD0629 | [16] |
| LDCM0313 | AC50 | HCT 116 | C50(0.73) | LDD0630 | [16] |
| LDCM0314 | AC51 | HCT 116 | C50(0.81) | LDD0631 | [16] |
| LDCM0315 | AC52 | HCT 116 | C50(0.86) | LDD0632 | [16] |
| LDCM0316 | AC53 | HCT 116 | C50(1.36) | LDD0633 | [16] |
| LDCM0317 | AC54 | HCT 116 | C50(1.01) | LDD0634 | [16] |
| LDCM0318 | AC55 | HCT 116 | C50(1.23) | LDD0635 | [16] |
| LDCM0319 | AC56 | HCT 116 | C50(0.95) | LDD0636 | [16] |
| LDCM0320 | AC57 | HCT 116 | C50(1.11) | LDD0637 | [16] |
| LDCM0321 | AC58 | HCT 116 | C50(0.77) | LDD0638 | [16] |
| LDCM0322 | AC59 | HCT 116 | C50(0.82) | LDD0639 | [16] |
| LDCM0323 | AC6 | HCT 116 | C50(0.96) | LDD0640 | [16] |
| LDCM0324 | AC60 | HCT 116 | C50(1.06) | LDD0641 | [16] |
| LDCM0325 | AC61 | HCT 116 | C50(1.15) | LDD0642 | [16] |
| LDCM0326 | AC62 | HCT 116 | C50(0.80) | LDD0643 | [16] |
| LDCM0327 | AC63 | HCT 116 | C50(1.02) | LDD0644 | [16] |
| LDCM0328 | AC64 | HCT 116 | C50(0.87) | LDD0645 | [16] |
| LDCM0329 | AC65 | HCT 116 | C50(0.77) | LDD0646 | [16] |
| LDCM0330 | AC66 | HCT 116 | C50(0.95) | LDD0647 | [16] |
| LDCM0331 | AC67 | HCT 116 | C50(0.53) | LDD0648 | [16] |
| LDCM0332 | AC68 | HCT 116 | C50(0.90) | LDD0649 | [16] |
| LDCM0333 | AC69 | HCT 116 | C50(0.86) | LDD0650 | [16] |
| LDCM0334 | AC7 | HCT 116 | C50(1.23) | LDD0651 | [16] |
| LDCM0335 | AC70 | HCT 116 | C50(0.61) | LDD0652 | [16] |
| LDCM0336 | AC71 | HCT 116 | C50(1.59) | LDD0653 | [16] |
| LDCM0337 | AC72 | HCT 116 | C50(1.03) | LDD0654 | [16] |
| LDCM0338 | AC73 | HCT 116 | C50(0.37) | LDD0655 | [16] |
| LDCM0339 | AC74 | HCT 116 | C50(0.51) | LDD0656 | [16] |
| LDCM0340 | AC75 | HCT 116 | C50(0.48) | LDD0657 | [16] |
| LDCM0341 | AC76 | HCT 116 | C50(0.90) | LDD0658 | [16] |
| LDCM0342 | AC77 | HCT 116 | C50(0.67) | LDD0659 | [16] |
| LDCM0343 | AC78 | HCT 116 | C50(1.35) | LDD0660 | [16] |
| LDCM0344 | AC79 | HCT 116 | C50(1.13) | LDD0661 | [16] |
| LDCM0345 | AC8 | HCT 116 | C50(0.81) | LDD0662 | [16] |
| LDCM0346 | AC80 | HCT 116 | C50(0.91) | LDD0663 | [16] |
| LDCM0347 | AC81 | HCT 116 | C50(1.02) | LDD0664 | [16] |
| LDCM0348 | AC82 | HCT 116 | C50(0.49) | LDD0665 | [16] |
| LDCM0349 | AC83 | HCT 116 | C50(0.64) | LDD0666 | [16] |
| LDCM0350 | AC84 | HCT 116 | C50(0.66) | LDD0667 | [16] |
| LDCM0351 | AC85 | HCT 116 | C50(0.88) | LDD0668 | [16] |
| LDCM0352 | AC86 | HCT 116 | C50(0.88) | LDD0669 | [16] |
| LDCM0353 | AC87 | HCT 116 | C50(1.05) | LDD0670 | [16] |
| LDCM0354 | AC88 | HCT 116 | C50(0.81) | LDD0671 | [16] |
| LDCM0355 | AC89 | HCT 116 | C50(0.73) | LDD0672 | [16] |
| LDCM0357 | AC90 | HCT 116 | C50(1.68) | LDD0674 | [16] |
| LDCM0358 | AC91 | HCT 116 | C50(0.65) | LDD0675 | [16] |
| LDCM0359 | AC92 | HCT 116 | C50(0.57) | LDD0676 | [16] |
| LDCM0360 | AC93 | HCT 116 | C50(0.84) | LDD0677 | [16] |
| LDCM0361 | AC94 | HCT 116 | C50(1.05) | LDD0678 | [16] |
| LDCM0362 | AC95 | HCT 116 | C50(1.27) | LDD0679 | [16] |
| LDCM0363 | AC96 | HCT 116 | C50(0.88) | LDD0680 | [16] |
| LDCM0364 | AC97 | HCT 116 | C50(0.71) | LDD0681 | [16] |
| LDCM0365 | AC98 | HCT 116 | C50(0.29) | LDD0682 | [16] |
| LDCM0366 | AC99 | HCT 116 | C50(0.65) | LDD0683 | [16] |
| LDCM0545 | Acetamide | MDA-MB-231 | C50(1.05) | LDD2138 | [12] |
| LDCM0248 | AKOS034007472 | HCT 116 | C50(1.21) | LDD0565 | [16] |
| LDCM0356 | AKOS034007680 | HCT 116 | C50(1.36) | LDD0673 | [16] |
| LDCM0275 | AKOS034007705 | HCT 116 | C50(0.87) | LDD0592 | [16] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C50(0.86) | LDD2091 | [12] |
| LDCM0108 | Chloroacetamide | HeLa | C50(0.00); H180(0.00); H163(0.00); H108(0.00) | LDD0222 | [32] |
| LDCM0367 | CL1 | HCT 116 | C50(0.84) | LDD0684 | [16] |
| LDCM0368 | CL10 | HCT 116 | C50(1.39) | LDD0685 | [16] |
| LDCM0369 | CL100 | HCT 116 | C50(1.34) | LDD0686 | [16] |
| LDCM0370 | CL101 | HCT 116 | C50(1.08) | LDD0687 | [16] |
| LDCM0371 | CL102 | HCT 116 | C50(1.52) | LDD0688 | [16] |
| LDCM0372 | CL103 | HCT 116 | C50(1.88) | LDD0689 | [16] |
| LDCM0373 | CL104 | HCT 116 | C50(1.10) | LDD0690 | [16] |
| LDCM0374 | CL105 | HCT 116 | C50(0.87) | LDD0691 | [16] |
| LDCM0375 | CL106 | HCT 116 | C50(0.72) | LDD0692 | [16] |
| LDCM0376 | CL107 | HCT 116 | C50(0.80) | LDD0693 | [16] |
| LDCM0377 | CL108 | HCT 116 | C50(0.61) | LDD0694 | [16] |
| LDCM0378 | CL109 | HCT 116 | C50(0.85) | LDD0695 | [16] |
| LDCM0379 | CL11 | HCT 116 | C50(1.51) | LDD0696 | [16] |
| LDCM0380 | CL110 | HCT 116 | C50(0.69) | LDD0697 | [16] |
| LDCM0381 | CL111 | HCT 116 | C50(0.75) | LDD0698 | [16] |
| LDCM0382 | CL112 | HCT 116 | C50(1.05) | LDD0699 | [16] |
| LDCM0383 | CL113 | HCT 116 | C50(0.78) | LDD0700 | [16] |
| LDCM0384 | CL114 | HCT 116 | C50(1.00) | LDD0701 | [16] |
| LDCM0385 | CL115 | HCT 116 | C50(0.87) | LDD0702 | [16] |
| LDCM0386 | CL116 | HCT 116 | C50(0.84) | LDD0703 | [16] |
| LDCM0387 | CL117 | HCT 116 | C50(0.84) | LDD0704 | [16] |
| LDCM0388 | CL118 | HCT 116 | C50(1.16) | LDD0705 | [16] |
| LDCM0389 | CL119 | HCT 116 | C50(1.15) | LDD0706 | [16] |
| LDCM0390 | CL12 | HCT 116 | C50(0.92) | LDD0707 | [16] |
| LDCM0391 | CL120 | HCT 116 | C50(0.90) | LDD0708 | [16] |
| LDCM0392 | CL121 | HCT 116 | C50(1.04) | LDD0709 | [16] |
| LDCM0393 | CL122 | HCT 116 | C50(0.95) | LDD0710 | [16] |
| LDCM0394 | CL123 | HCT 116 | C50(1.06) | LDD0711 | [16] |
| LDCM0395 | CL124 | HCT 116 | C50(0.99) | LDD0712 | [16] |
| LDCM0396 | CL125 | HCT 116 | C50(1.39) | LDD0713 | [16] |
| LDCM0397 | CL126 | HCT 116 | C50(1.06) | LDD0714 | [16] |
| LDCM0398 | CL127 | HCT 116 | C50(1.13) | LDD0715 | [16] |
| LDCM0399 | CL128 | HCT 116 | C50(0.88) | LDD0716 | [16] |
| LDCM0400 | CL13 | HCT 116 | C50(1.27) | LDD0717 | [16] |
| LDCM0401 | CL14 | HCT 116 | C50(1.06) | LDD0718 | [16] |
| LDCM0402 | CL15 | HCT 116 | C50(1.11) | LDD0719 | [16] |
| LDCM0403 | CL16 | HCT 116 | C50(0.76) | LDD0720 | [16] |
| LDCM0404 | CL17 | HEK-293T | C50(0.55) | LDD1608 | [45] |
| LDCM0405 | CL18 | HEK-293T | C50(0.63) | LDD1609 | [45] |
| LDCM0406 | CL19 | HEK-293T | C50(0.58) | LDD1610 | [45] |
| LDCM0407 | CL2 | HCT 116 | C50(1.04) | LDD0724 | [16] |
| LDCM0408 | CL20 | HEK-293T | C50(0.65) | LDD1612 | [45] |
| LDCM0409 | CL21 | HEK-293T | C50(0.67) | LDD1613 | [45] |
| LDCM0410 | CL22 | HEK-293T | C50(0.61) | LDD1614 | [45] |
| LDCM0411 | CL23 | HEK-293T | C50(0.83) | LDD1615 | [45] |
| LDCM0412 | CL24 | HEK-293T | C50(0.53) | LDD1616 | [45] |
| LDCM0413 | CL25 | HEK-293T | C50(0.90) | LDD1617 | [45] |
| LDCM0414 | CL26 | HEK-293T | C50(0.95) | LDD1618 | [45] |
| LDCM0415 | CL27 | HEK-293T | C50(0.86) | LDD1619 | [45] |
| LDCM0416 | CL28 | HEK-293T | C50(0.90) | LDD1620 | [45] |
| LDCM0417 | CL29 | HEK-293T | C50(0.83) | LDD1621 | [45] |
| LDCM0418 | CL3 | HCT 116 | C50(1.21) | LDD0735 | [16] |
| LDCM0419 | CL30 | HEK-293T | C50(0.79) | LDD1623 | [45] |
| LDCM0420 | CL31 | HCT 116 | C50(0.80) | LDD0737 | [16] |
| LDCM0421 | CL32 | HCT 116 | C50(0.68) | LDD0738 | [16] |
| LDCM0422 | CL33 | HCT 116 | C50(0.67) | LDD0739 | [16] |
| LDCM0423 | CL34 | HCT 116 | C50(0.54) | LDD0740 | [16] |
| LDCM0424 | CL35 | HCT 116 | C50(0.56) | LDD0741 | [16] |
| LDCM0425 | CL36 | HCT 116 | C50(0.57) | LDD0742 | [16] |
| LDCM0426 | CL37 | HCT 116 | C50(0.53) | LDD0743 | [16] |
| LDCM0428 | CL39 | HCT 116 | C50(0.62) | LDD0745 | [16] |
| LDCM0429 | CL4 | HCT 116 | C50(1.02) | LDD0746 | [16] |
| LDCM0430 | CL40 | HCT 116 | C50(0.63) | LDD0747 | [16] |
| LDCM0431 | CL41 | HCT 116 | C50(0.70) | LDD0748 | [16] |
| LDCM0432 | CL42 | HCT 116 | C50(0.59) | LDD0749 | [16] |
| LDCM0433 | CL43 | HCT 116 | C50(0.51) | LDD0750 | [16] |
| LDCM0434 | CL44 | HCT 116 | C50(0.68) | LDD0751 | [16] |
| LDCM0435 | CL45 | HCT 116 | C50(0.65) | LDD0752 | [16] |
| LDCM0436 | CL46 | HCT 116 | C50(1.36) | LDD0753 | [16] |
| LDCM0437 | CL47 | HCT 116 | C50(1.35) | LDD0754 | [16] |
| LDCM0438 | CL48 | HCT 116 | C50(1.58) | LDD0755 | [16] |
| LDCM0439 | CL49 | HCT 116 | C50(1.12) | LDD0756 | [16] |
| LDCM0440 | CL5 | HCT 116 | C50(1.56) | LDD0757 | [16] |
| LDCM0441 | CL50 | HCT 116 | C50(1.26) | LDD0758 | [16] |
| LDCM0442 | CL51 | HCT 116 | C50(1.52) | LDD0759 | [16] |
| LDCM0443 | CL52 | HCT 116 | C50(1.20) | LDD0760 | [16] |
| LDCM0444 | CL53 | HCT 116 | C50(1.05) | LDD0761 | [16] |
| LDCM0445 | CL54 | HCT 116 | C50(1.27) | LDD0762 | [16] |
| LDCM0446 | CL55 | HCT 116 | C50(1.16) | LDD0763 | [16] |
| LDCM0447 | CL56 | HCT 116 | C50(0.88) | LDD0764 | [16] |
| LDCM0448 | CL57 | HCT 116 | C50(1.20) | LDD0765 | [16] |
| LDCM0449 | CL58 | HCT 116 | C50(1.32) | LDD0766 | [16] |
| LDCM0450 | CL59 | HCT 116 | C50(1.08) | LDD0767 | [16] |
| LDCM0451 | CL6 | HCT 116 | C50(1.07) | LDD0768 | [16] |
| LDCM0452 | CL60 | HCT 116 | C50(1.29) | LDD0769 | [16] |
| LDCM0453 | CL61 | PaTu 8988t | C50(0.86) | LDD1332 | [16] |
| LDCM0454 | CL62 | PaTu 8988t | C50(2.15) | LDD1333 | [16] |
| LDCM0455 | CL63 | PaTu 8988t | C50(1.63) | LDD1334 | [16] |
| LDCM0456 | CL64 | PaTu 8988t | C50(1.07) | LDD1335 | [16] |
| LDCM0457 | CL65 | PaTu 8988t | C50(1.03) | LDD1336 | [16] |
| LDCM0458 | CL66 | PaTu 8988t | C50(0.87) | LDD1337 | [16] |
| LDCM0459 | CL67 | PaTu 8988t | C50(0.45) | LDD1338 | [16] |
| LDCM0460 | CL68 | PaTu 8988t | C50(0.28) | LDD1339 | [16] |
| LDCM0461 | CL69 | PaTu 8988t | C50(0.42) | LDD1340 | [16] |
| LDCM0462 | CL7 | HCT 116 | C50(1.44) | LDD0779 | [16] |
| LDCM0463 | CL70 | PaTu 8988t | C50(0.45) | LDD1342 | [16] |
| LDCM0464 | CL71 | PaTu 8988t | C50(1.76) | LDD1343 | [16] |
| LDCM0465 | CL72 | PaTu 8988t | C50(4.06) | LDD1344 | [16] |
| LDCM0466 | CL73 | PaTu 8988t | C50(0.49) | LDD1345 | [16] |
| LDCM0467 | CL74 | PaTu 8988t | C50(1.92) | LDD1346 | [16] |
| LDCM0469 | CL76 | HCT 116 | C50(0.59) | LDD0786 | [16] |
| LDCM0470 | CL77 | HCT 116 | C50(0.49) | LDD0787 | [16] |
| LDCM0471 | CL78 | HCT 116 | C50(0.55) | LDD0788 | [16] |
| LDCM0472 | CL79 | HCT 116 | C50(0.35) | LDD0789 | [16] |
| LDCM0473 | CL8 | HCT 116 | C50(0.64) | LDD0790 | [16] |
| LDCM0474 | CL80 | HCT 116 | C50(1.26) | LDD0791 | [16] |
| LDCM0475 | CL81 | HCT 116 | C50(0.42) | LDD0792 | [16] |
| LDCM0476 | CL82 | HCT 116 | C50(0.14) | LDD0793 | [16] |
| LDCM0477 | CL83 | HCT 116 | C50(0.25) | LDD0794 | [16] |
| LDCM0478 | CL84 | HCT 116 | C50(0.09) | LDD0795 | [16] |
| LDCM0479 | CL85 | HCT 116 | C50(0.62) | LDD0796 | [16] |
| LDCM0480 | CL86 | HCT 116 | C50(1.24) | LDD0797 | [16] |
| LDCM0481 | CL87 | HCT 116 | C50(0.50) | LDD0798 | [16] |
| LDCM0482 | CL88 | HCT 116 | C50(0.24) | LDD0799 | [16] |
| LDCM0483 | CL89 | HCT 116 | C50(0.05) | LDD0800 | [16] |
| LDCM0484 | CL9 | HCT 116 | C50(1.06) | LDD0801 | [16] |
| LDCM0485 | CL90 | HCT 116 | C50(0.74) | LDD0802 | [16] |
| LDCM0486 | CL91 | HCT 116 | C50(0.93) | LDD0803 | [16] |
| LDCM0487 | CL92 | HCT 116 | C50(1.26) | LDD0804 | [16] |
| LDCM0488 | CL93 | HCT 116 | C50(2.15) | LDD0805 | [16] |
| LDCM0489 | CL94 | HCT 116 | C50(1.95) | LDD0806 | [16] |
| LDCM0490 | CL95 | HCT 116 | C50(1.67) | LDD0807 | [16] |
| LDCM0491 | CL96 | HCT 116 | C50(1.57) | LDD0808 | [16] |
| LDCM0492 | CL97 | HCT 116 | C50(1.53) | LDD0809 | [16] |
| LDCM0493 | CL98 | HCT 116 | C50(1.37) | LDD0810 | [16] |
| LDCM0494 | CL99 | HCT 116 | C50(1.69) | LDD0811 | [16] |
| LDCM0495 | E2913 | HEK-293T | C50(0.82) | LDD1698 | [45] |
| LDCM0625 | F8 | Ramos | C50(0.54) | LDD2187 | [46] |
| LDCM0572 | Fragment10 | Ramos | C50(0.68) | LDD2189 | [46] |
| LDCM0573 | Fragment11 | Ramos | C50(0.03) | LDD2190 | [46] |
| LDCM0574 | Fragment12 | Ramos | C50(0.68) | LDD2191 | [46] |
| LDCM0575 | Fragment13 | Ramos | C50(1.27) | LDD2192 | [46] |
| LDCM0576 | Fragment14 | Ramos | C50(0.82) | LDD2193 | [46] |
| LDCM0579 | Fragment20 | Ramos | C50(0.54) | LDD2194 | [46] |
| LDCM0580 | Fragment21 | Ramos | C50(0.93) | LDD2195 | [46] |
| LDCM0582 | Fragment23 | Ramos | C50(1.44) | LDD2196 | [46] |
| LDCM0578 | Fragment27 | Ramos | C50(1.31) | LDD2197 | [46] |
| LDCM0586 | Fragment28 | Ramos | C50(0.65) | LDD2198 | [46] |
| LDCM0588 | Fragment30 | Ramos | C50(1.51) | LDD2199 | [46] |
| LDCM0589 | Fragment31 | Ramos | C50(1.29) | LDD2200 | [46] |
| LDCM0590 | Fragment32 | Ramos | C50(0.71) | LDD2201 | [46] |
| LDCM0468 | Fragment33 | PaTu 8988t | C50(0.88) | LDD1347 | [16] |
| LDCM0596 | Fragment38 | Ramos | C50(1.24) | LDD2203 | [46] |
| LDCM0566 | Fragment4 | Ramos | C50(0.68) | LDD2184 | [46] |
| LDCM0427 | Fragment51 | HCT 116 | C50(0.47) | LDD0744 | [16] |
| LDCM0610 | Fragment52 | Ramos | C50(1.63) | LDD2204 | [46] |
| LDCM0614 | Fragment56 | Ramos | C50(1.24) | LDD2205 | [46] |
| LDCM0569 | Fragment7 | Ramos | C50(0.61) | LDD2186 | [46] |
| LDCM0571 | Fragment9 | Ramos | C50(0.55) | LDD2188 | [46] |
| LDCM0116 | HHS-0101 | DM93 | Y331(0.71); Y174(0.74); Y347(0.74); Y336(0.75) | LDD0264 | [14] |
| LDCM0117 | HHS-0201 | DM93 | Y234(0.71); Y331(0.72); Y347(0.77); Y174(0.78) | LDD0265 | [14] |
| LDCM0118 | HHS-0301 | DM93 | Y262(0.05); Y247(0.35); Y244(0.48); Y331(0.54) | LDD0266 | [14] |
| LDCM0119 | HHS-0401 | DM93 | Y244(0.75); Y331(0.79); Y336(0.83); Y347(0.83) | LDD0267 | [14] |
| LDCM0120 | HHS-0701 | DM93 | Y262(0.09); Y234(0.68); Y174(0.76); Y347(0.80) | LDD0268 | [14] |
| LDCM0107 | IAA | HeLa | H163(0.00); H180(0.00); C50(0.00); H175(0.00) | LDD0221 | [32] |
| LDCM0123 | JWB131 | DM93 | Y174(1.01); Y234(0.87); Y331(0.80); Y336(0.81) | LDD0285 | [13] |
| LDCM0124 | JWB142 | DM93 | Y174(0.76); Y234(0.61); Y331(0.57); Y336(0.51) | LDD0286 | [13] |
| LDCM0125 | JWB146 | DM93 | Y174(0.97); Y234(1.13); Y331(1.50); Y336(1.01) | LDD0287 | [13] |
| LDCM0126 | JWB150 | DM93 | Y174(2.77); Y234(2.21); Y331(1.72); Y336(2.75) | LDD0288 | [13] |
| LDCM0127 | JWB152 | DM93 | Y174(2.14); Y234(1.87); Y331(2.71); Y336(3.04) | LDD0289 | [13] |
| LDCM0128 | JWB198 | DM93 | Y174(1.16); Y234(0.98); Y331(1.01); Y336(0.95) | LDD0290 | [13] |
| LDCM0129 | JWB202 | DM93 | Y174(0.73); Y234(0.41); Y331(0.36); Y336(0.33) | LDD0291 | [13] |
| LDCM0130 | JWB211 | DM93 | Y174(1.00); Y234(0.88); Y331(0.98); Y336(0.93) | LDD0292 | [13] |
| LDCM0022 | KB02 | HEK-293T | C50(1.01) | LDD1492 | [45] |
| LDCM0023 | KB03 | HEK-293T | C50(0.98) | LDD1497 | [45] |
| LDCM0024 | KB05 | COLO792 | C50(1.34) | LDD3310 | [47] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C50(0.94) | LDD2121 | [12] |
| LDCM0109 | NEM | HeLa | H163(0.00); H180(0.00); C50(0.00); H108(0.00) | LDD0223 | [32] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C50(1.38) | LDD2089 | [12] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C50(3.59) | LDD2090 | [12] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C50(2.50) | LDD2092 | [12] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C50(2.44) | LDD2094 | [12] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C50(0.07) | LDD2096 | [12] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C50(2.47) | LDD2098 | [12] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C50(0.41) | LDD2104 | [12] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C50(1.12) | LDD2107 | [12] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C50(0.26) | LDD2108 | [12] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C50(0.44) | LDD2109 | [12] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C50(0.51) | LDD2110 | [12] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C50(0.14) | LDD2116 | [12] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C50(1.00) | LDD2120 | [12] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C50(0.19) | LDD2124 | [12] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C50(0.73) | LDD2125 | [12] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C50(0.11) | LDD2126 | [12] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C50(0.74) | LDD2127 | [12] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C50(0.36) | LDD2128 | [12] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C50(1.01) | LDD2129 | [12] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C50(0.91) | LDD2135 | [12] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C50(0.89) | LDD2137 | [12] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C50(2.07) | LDD1700 | [12] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C50(0.52) | LDD2140 | [12] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C50(0.29) | LDD2141 | [12] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C50(0.35) | LDD2143 | [12] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C50(2.83) | LDD2144 | [12] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C50(0.43) | LDD2145 | [12] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C50(0.73) | LDD2146 | [12] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C50(0.93) | LDD2147 | [12] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C50(0.13) | LDD2149 | [12] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C50(1.41) | LDD2150 | [12] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C50(5.42) | LDD2153 | [12] |
| LDCM0131 | RA190 | MM1.R | C50(1.26) | LDD0304 | [48] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| LINE-1 type transposase domain-containing protein 1 (L1TD1) | Transposase 22 family | Q5T7N2 | |||
Other
References
