Details of the Target
General Information of Target
| Target ID | LDTP03417 | |||||
|---|---|---|---|---|---|---|
| Target Name | Proteasome subunit beta type-4 (PSMB4) | |||||
| Gene Name | PSMB4 | |||||
| Gene ID | 5692 | |||||
| Synonyms |
PROS26; Proteasome subunit beta type-4; 26 kDa prosomal protein; HsBPROS26; PROS-26; Macropain beta chain; Multicatalytic endopeptidase complex beta chain; Proteasome beta chain; Proteasome chain 3; HsN3
|
|||||
| 3D Structure | ||||||
| Sequence |
MEAFLGSRSGLWAGGPAPGQFYRIPSTPDSFMDPASALYRGPITRTQNPMVTGTSVLGVK
FEGGVVIAADMLGSYGSLARFRNISRIMRVNNSTMLGASGDYADFQYLKQVLGQMVIDEE LLGDGHSYSPRAIHSWLTRAMYSRRSKMNPLWNTMVIGGYADGESFLGYVDMLGVAYEAP SLATGYGAYLAQPLLREVLEKQPVLSQTEARDLVERCMRVLYYRDARSYNRFQIATVTEK GVEIEGPLSTETNWDIAHMISGFE |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Peptidase T1B family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Non-catalytic component of the 20S core proteasome complex involved in the proteolytic degradation of most intracellular proteins. This complex plays numerous essential roles within the cell by associating with different regulatory particles. Associated with two 19S regulatory particles, forms the 26S proteasome and thus participates in the ATP-dependent degradation of ubiquitinated proteins. The 26S proteasome plays a key role in the maintenance of protein homeostasis by removing misfolded or damaged proteins that could impair cellular functions, and by removing proteins whose functions are no longer required. Associated with the PA200 or PA28, the 20S proteasome mediates ubiquitin-independent protein degradation. This type of proteolysis is required in several pathways including spermatogenesis (20S-PA200 complex) or generation of a subset of MHC class I-presented antigenic peptides (20S-PA28 complex). SMAD1/OAZ1/PSMB4 complex mediates the degradation of the CREBBP/EP300 repressor SNIP1.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
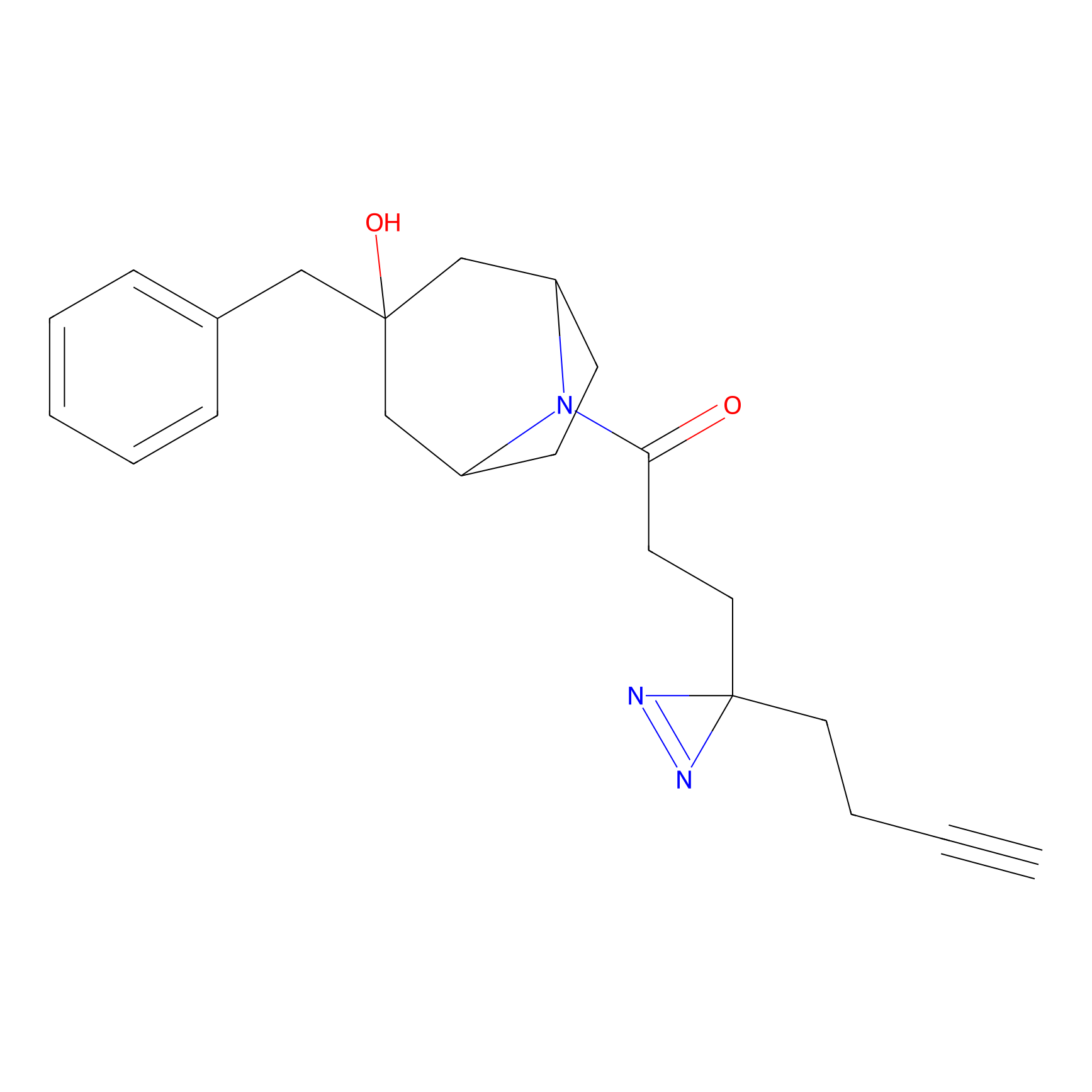
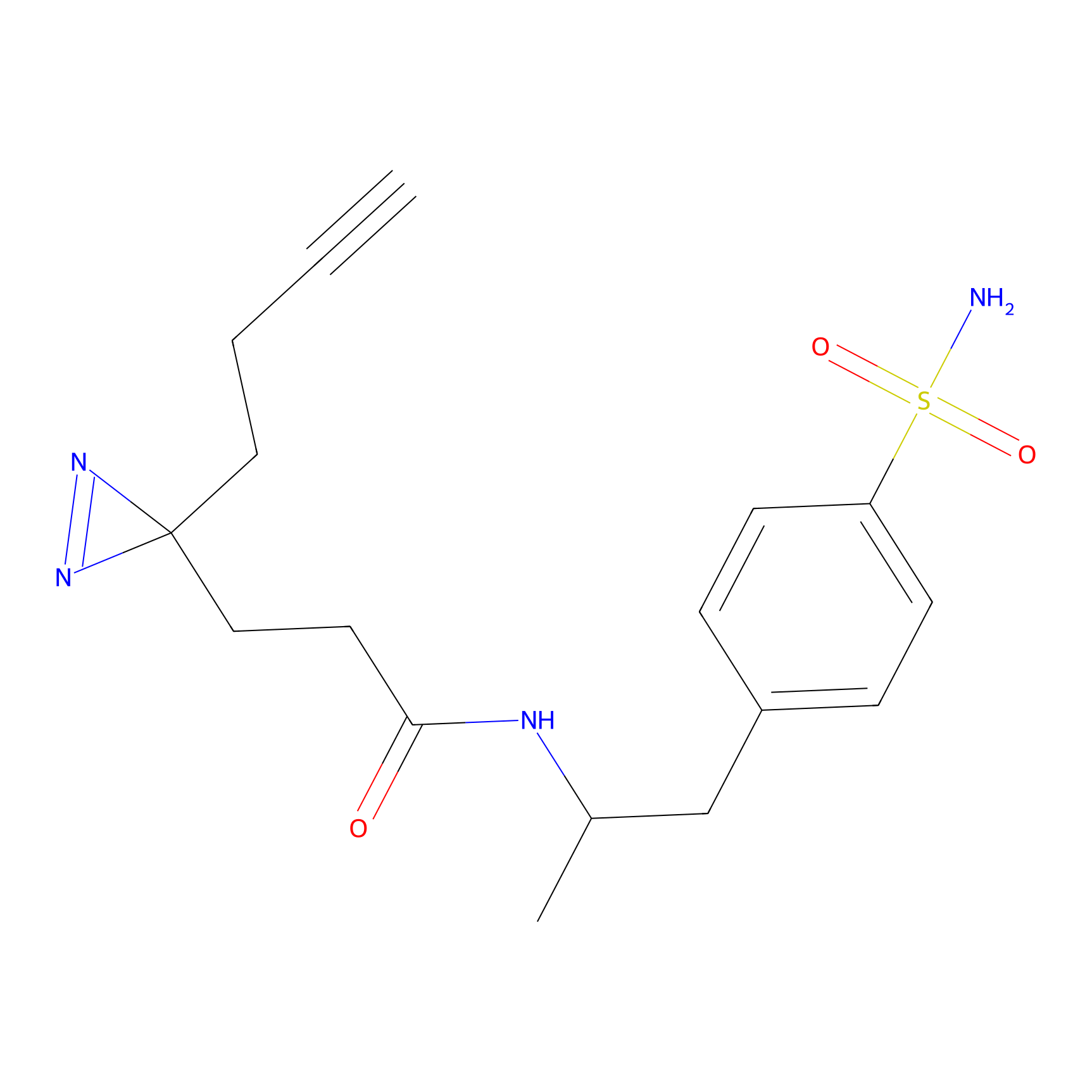
YN-1 Probe Info |
 |
100.00 | LDD0444 | [1] | |
|
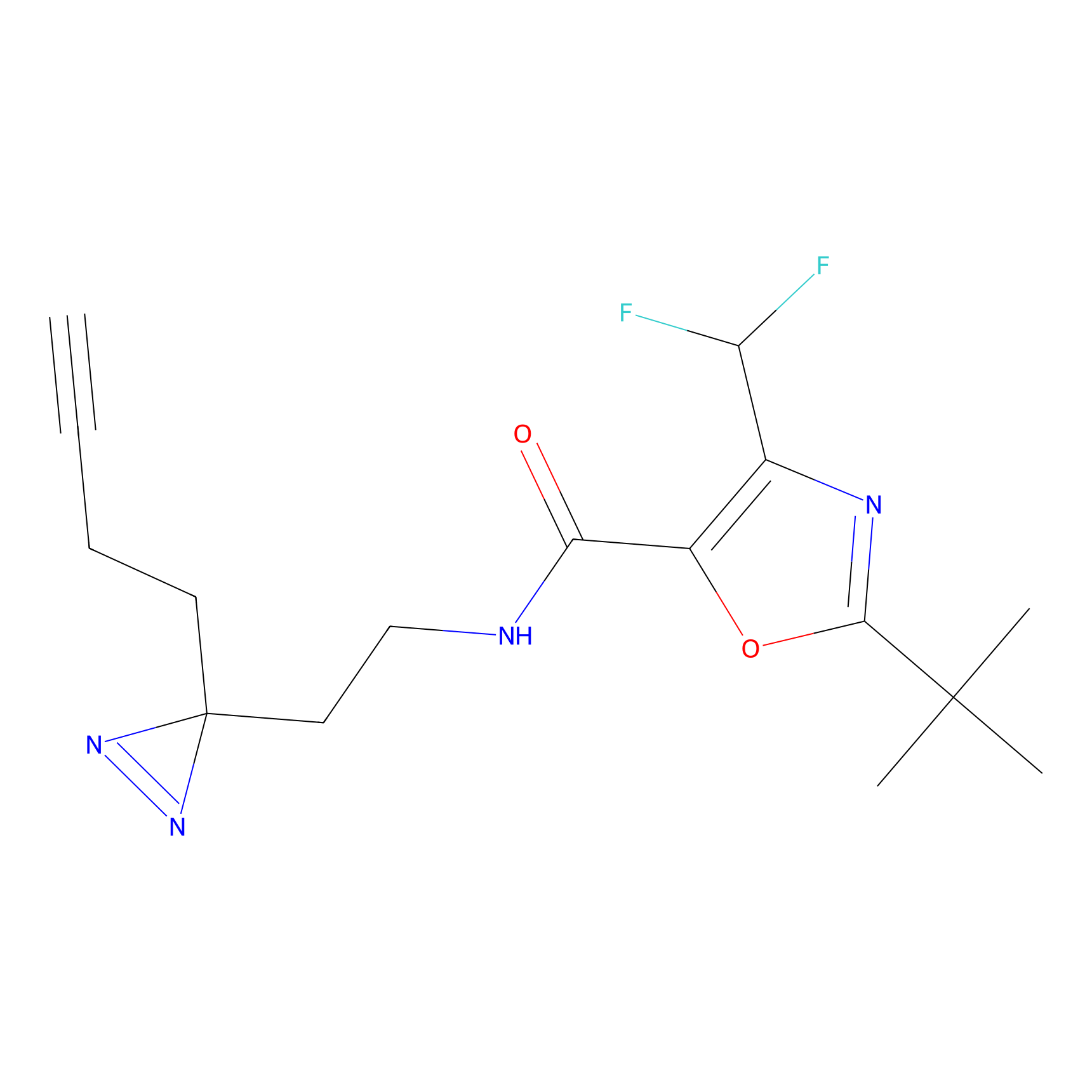
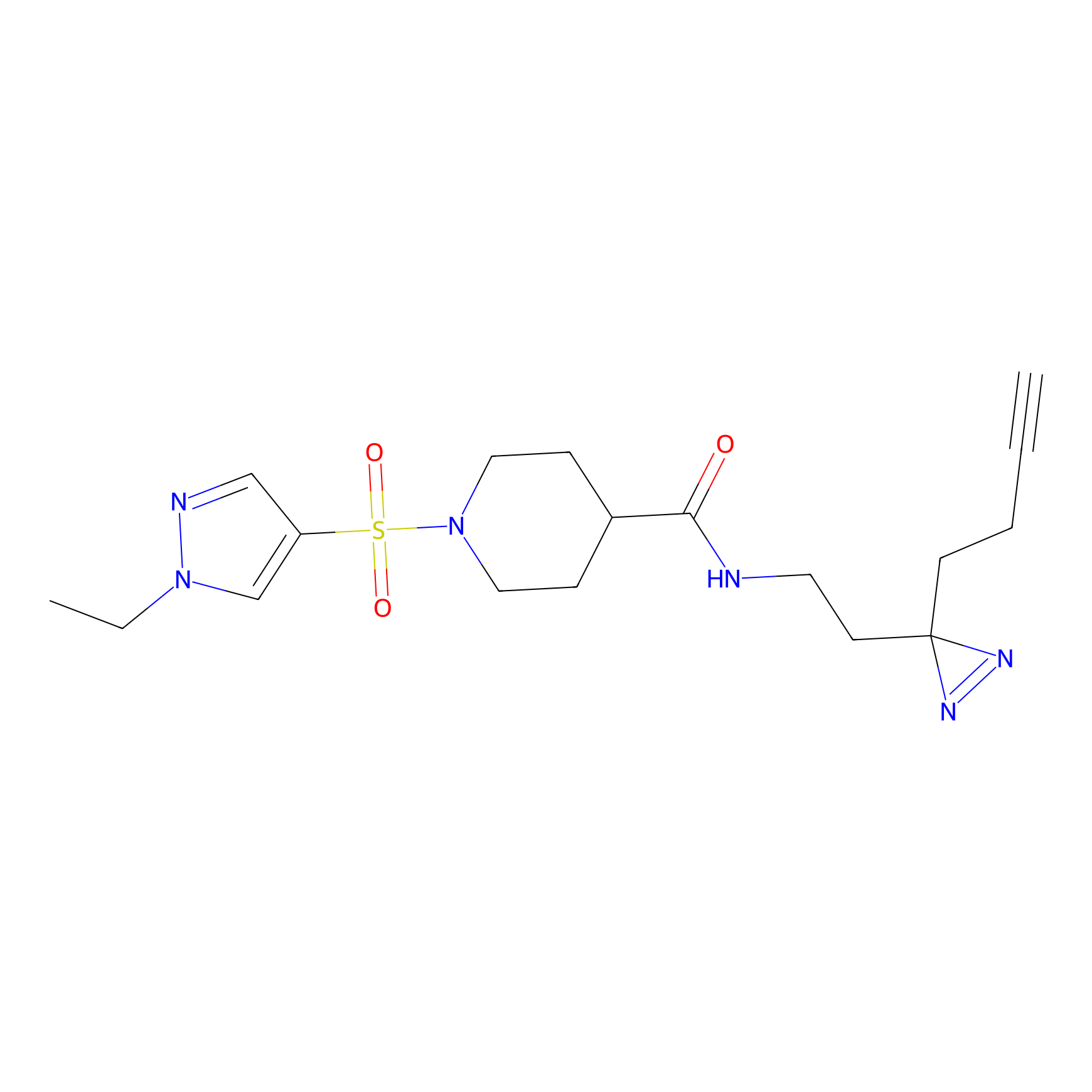
STPyne Probe Info |
 |
K109(8.90) | LDD2217 | [2] | |
|
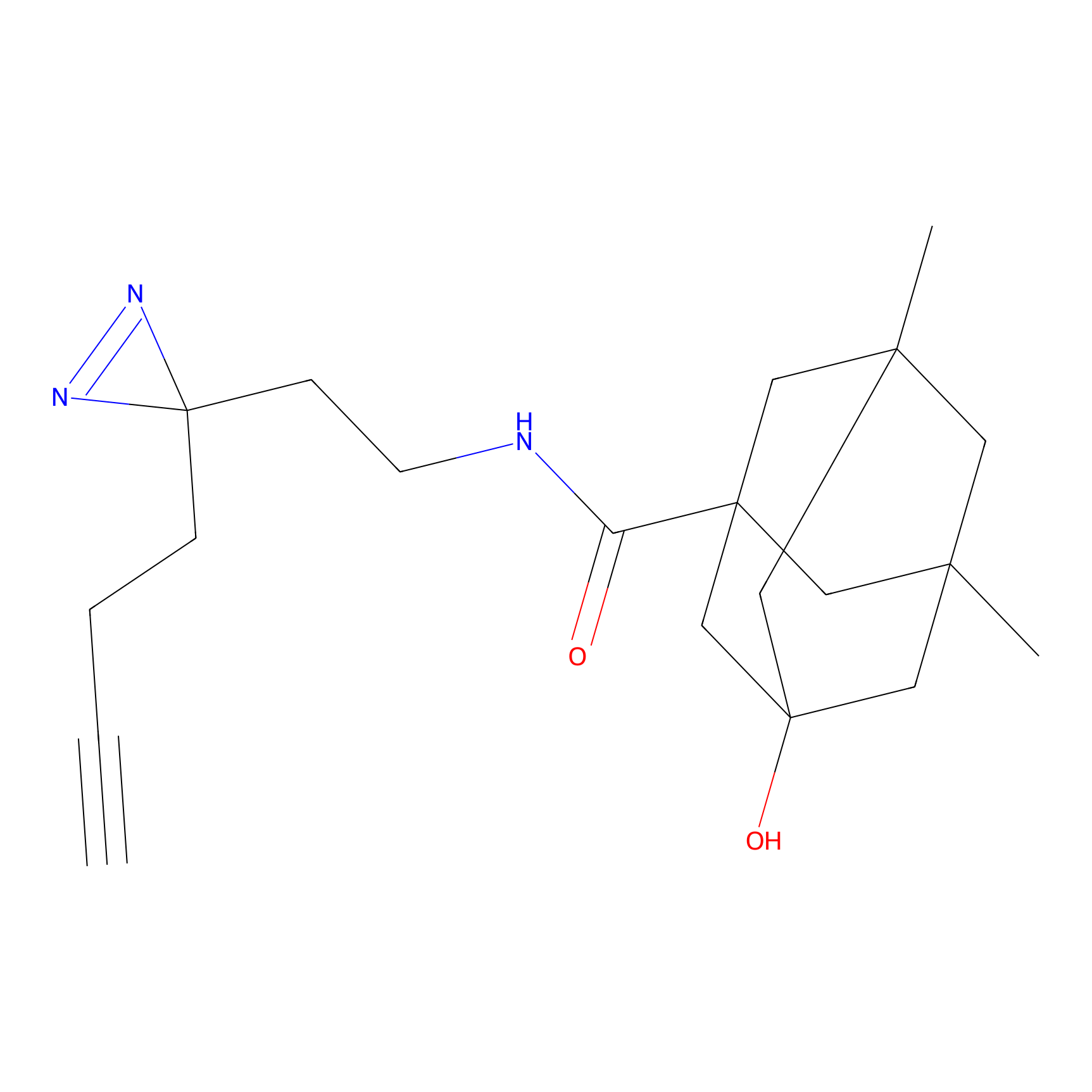
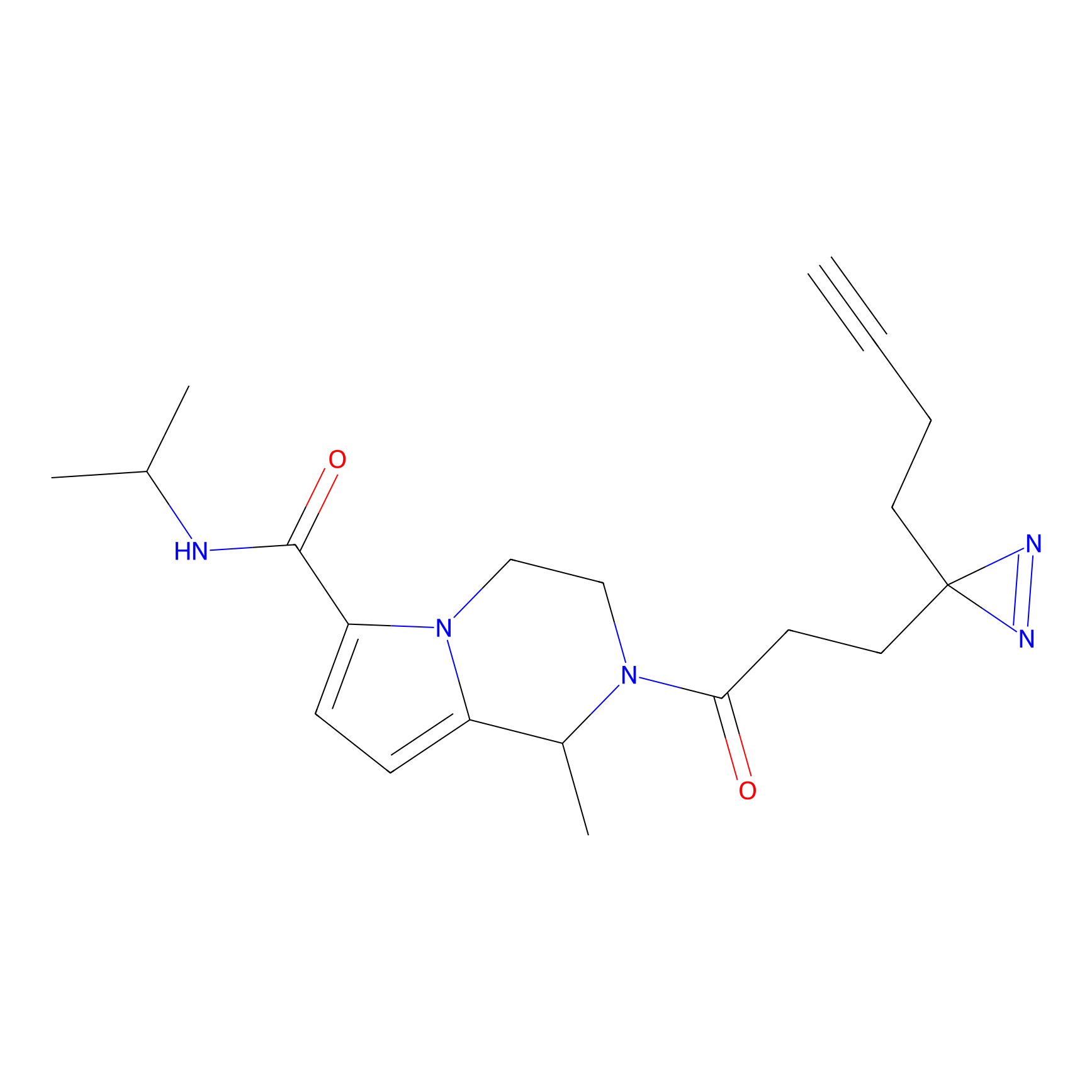
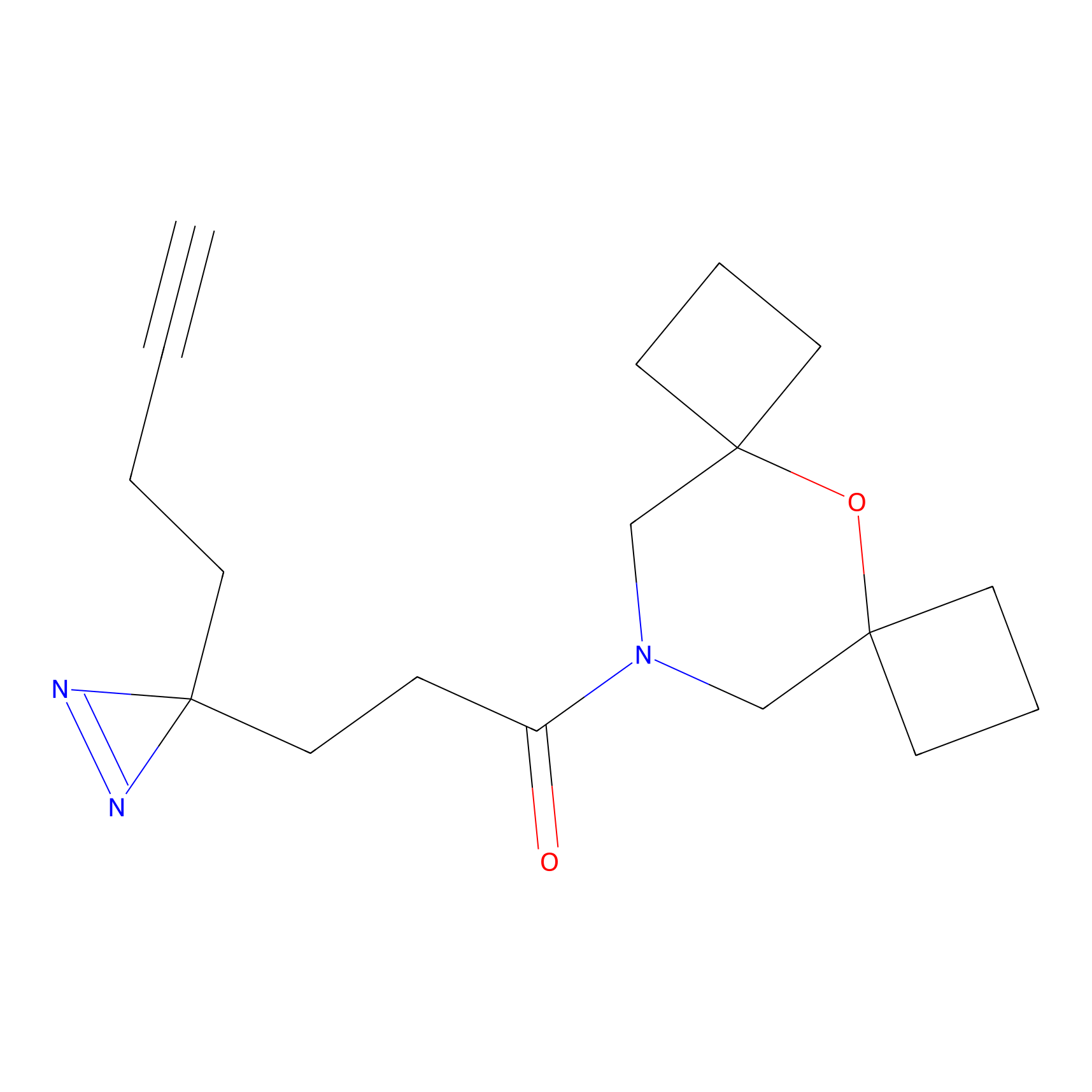
AZ-9 Probe Info |
 |
E239(1.42) | LDD2208 | [3] | |
|
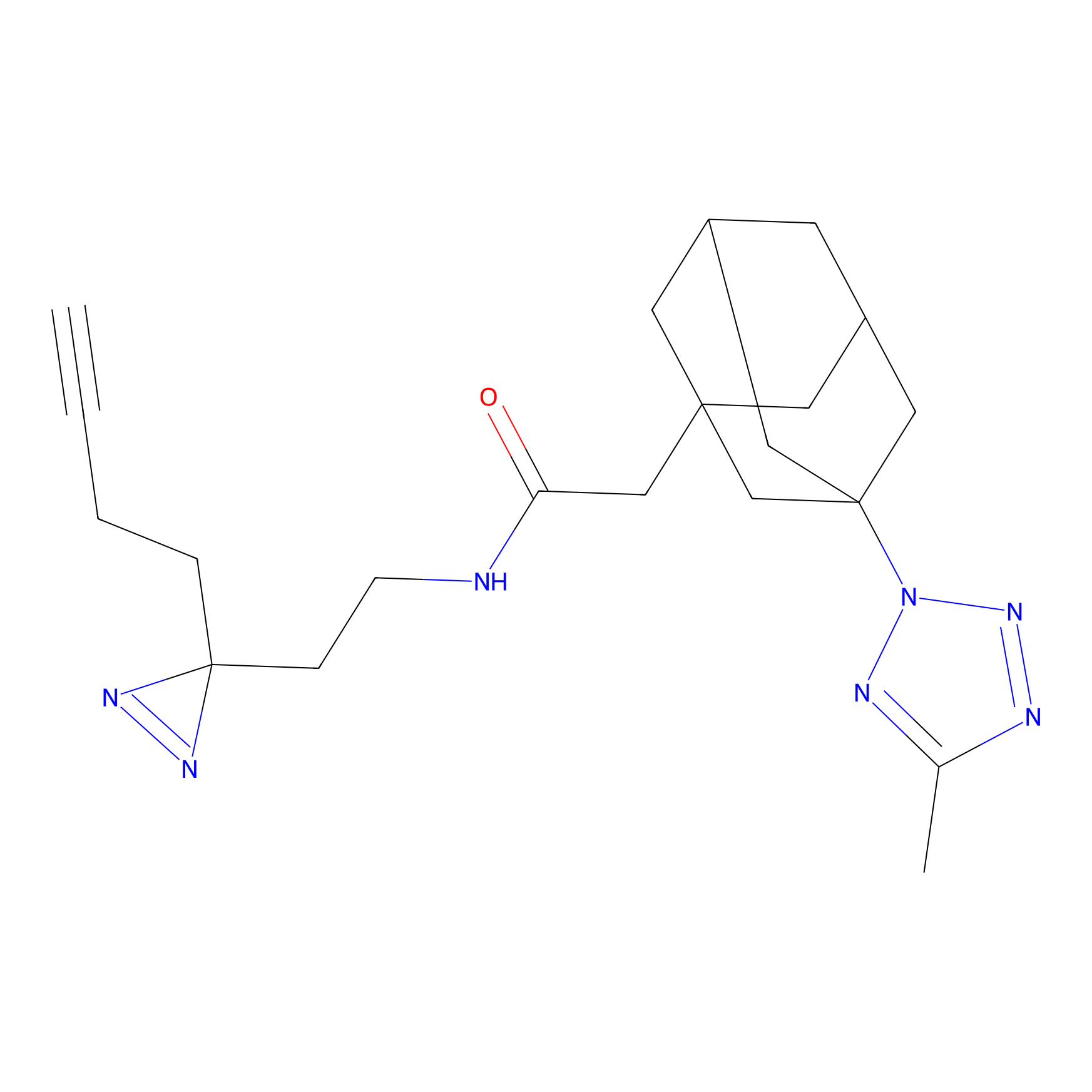
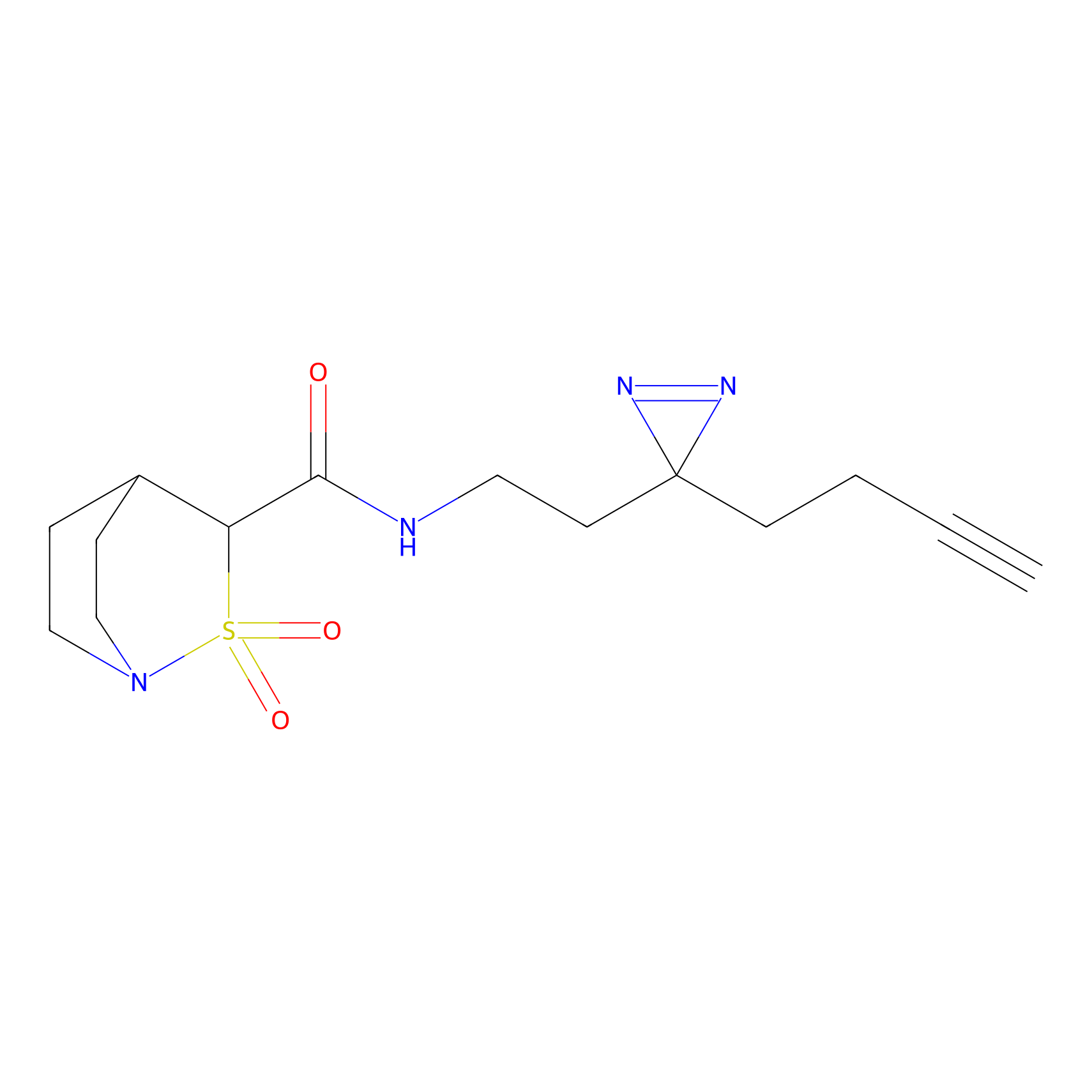
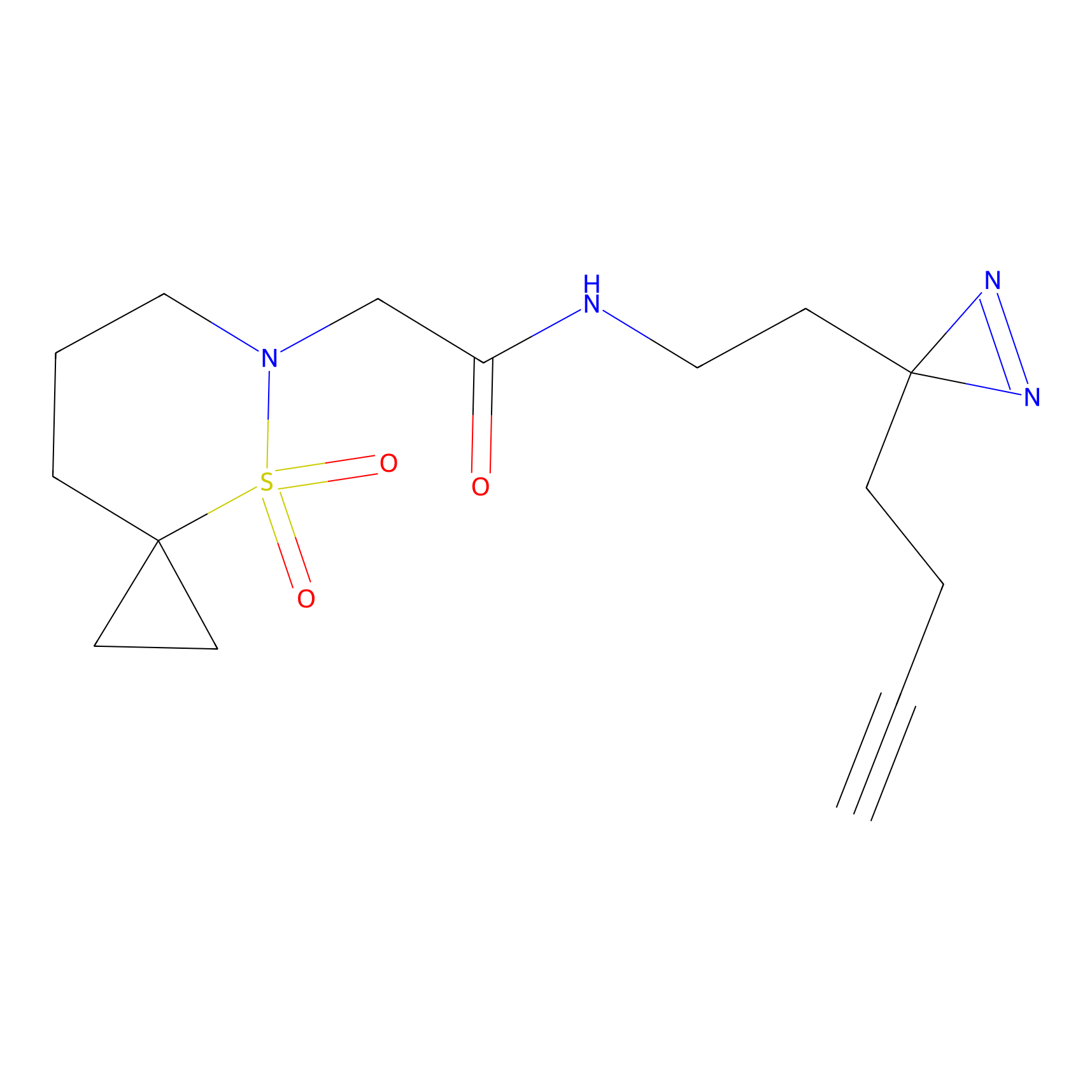
ONAyne Probe Info |
 |
N.A. | LDD0273 | [4] | |
|
OPA-S-S-alkyne Probe Info |
 |
K201(3.87) | LDD3494 | [5] | |
|
m-APA Probe Info |
 |
11.34 | LDD0403 | [6] | |
|
HHS-482 Probe Info |
 |
Y107(0.17) | LDD0286 | [7] | |
|
HHS-475 Probe Info |
 |
Y107(0.74); Y75(0.83) | LDD0264 | [8] | |
|
HHS-465 Probe Info |
 |
Y75(3.83) | LDD2237 | [9] | |
|
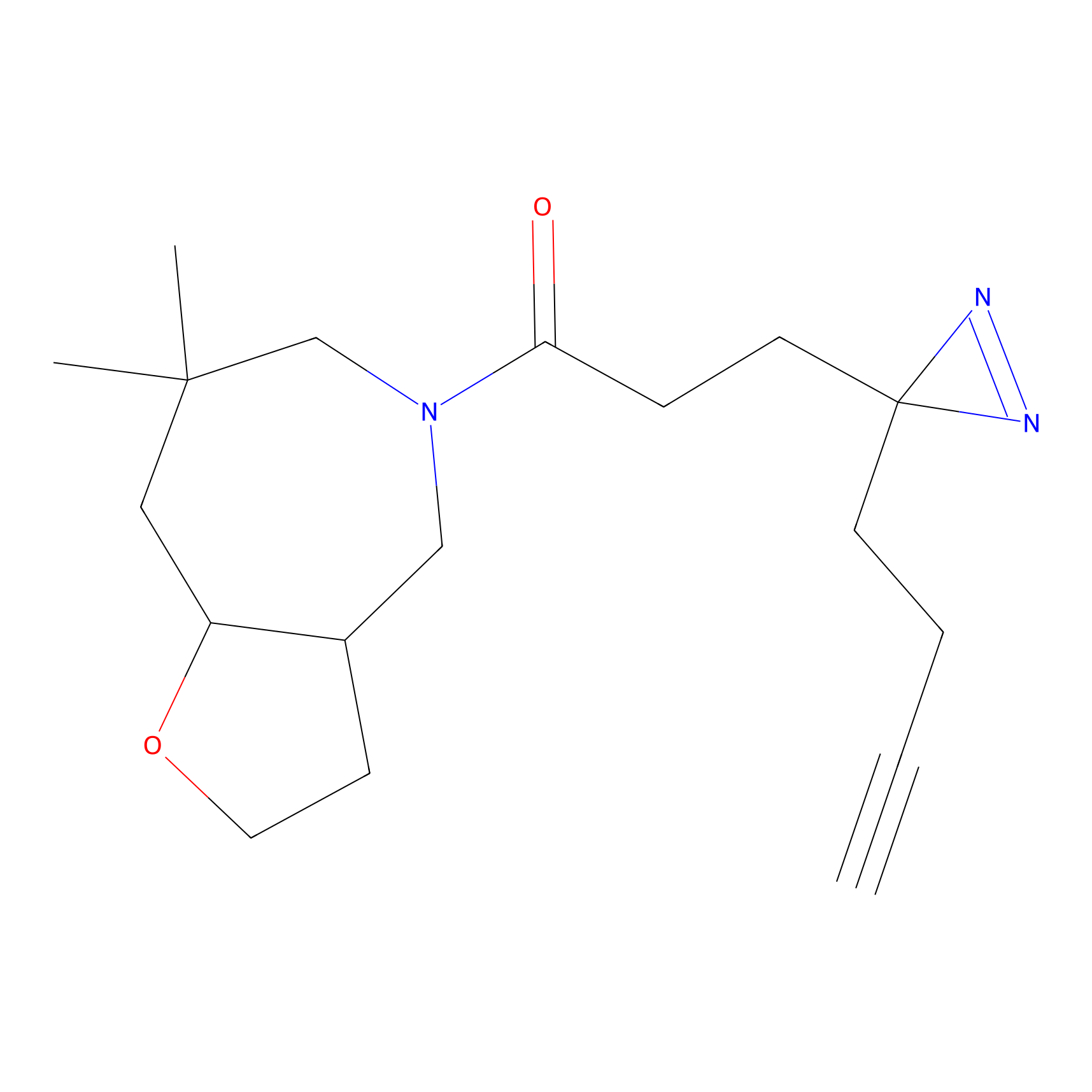
1d-yne Probe Info |
 |
N.A. | LDD0358 | [10] | |
|
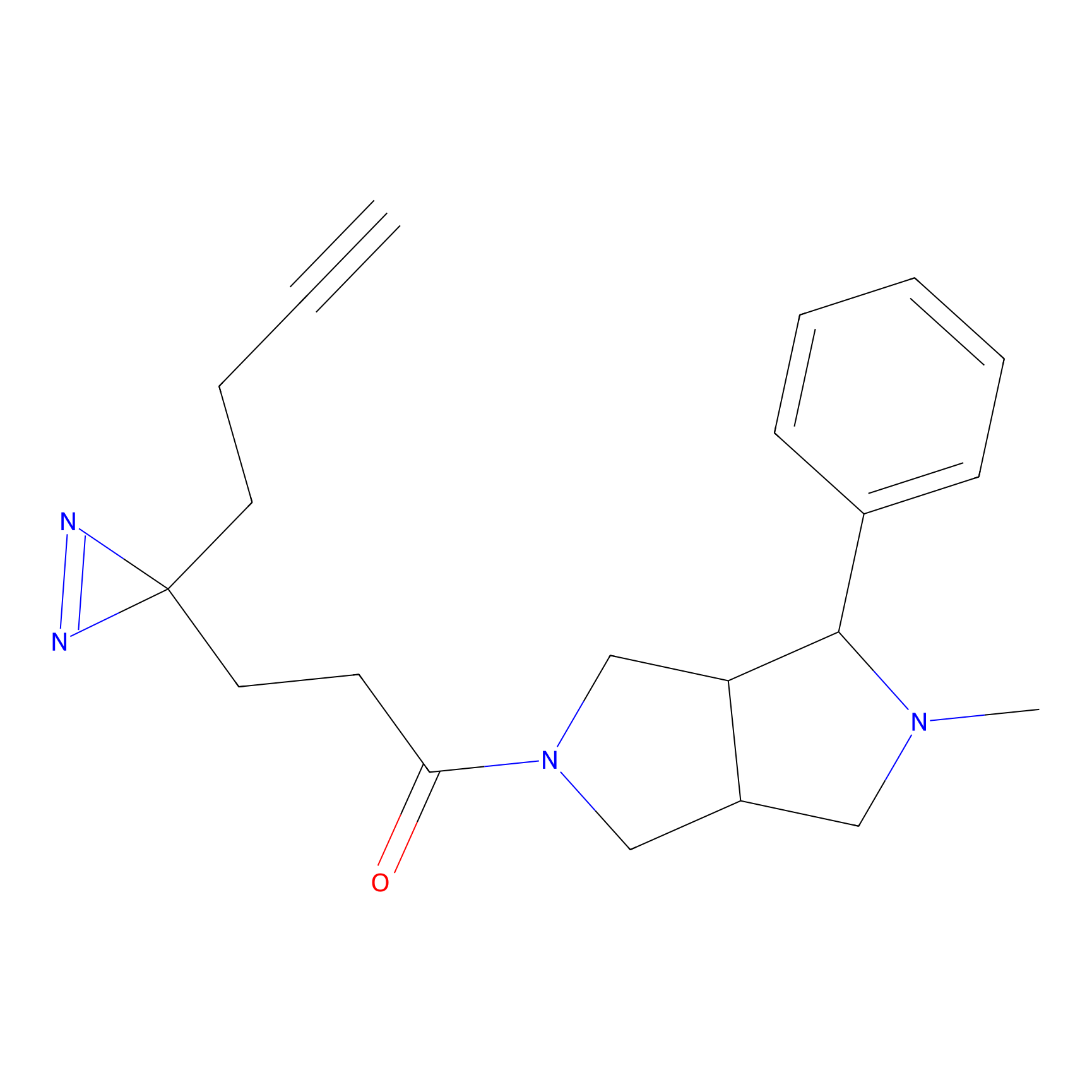
1c-yne Probe Info |
 |
N.A. | LDD0228 | [10] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 11.34 | LDD0403 | [6] |
| LDCM0116 | HHS-0101 | DM93 | Y107(0.74); Y75(0.83) | LDD0264 | [8] |
| LDCM0117 | HHS-0201 | DM93 | Y75(0.87); Y107(1.03) | LDD0265 | [8] |
| LDCM0118 | HHS-0301 | DM93 | Y75(0.91); Y107(1.21) | LDD0266 | [8] |
| LDCM0119 | HHS-0401 | DM93 | Y75(0.90); Y107(1.11) | LDD0267 | [8] |
| LDCM0120 | HHS-0701 | DM93 | Y75(1.03); Y107(1.19) | LDD0268 | [8] |
| LDCM0124 | JWB142 | DM93 | Y107(0.17) | LDD0286 | [7] |
| LDCM0126 | JWB150 | DM93 | Y107(2.34) | LDD0288 | [7] |
| LDCM0127 | JWB152 | DM93 | Y107(1.58) | LDD0289 | [7] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transcription factor
Other
References