Details of the Target
General Information of Target
| Target ID | LDTP03327 | |||||
|---|---|---|---|---|---|---|
| Target Name | ATP synthase subunit alpha, mitochondrial (ATP5F1A) | |||||
| Gene Name | ATP5F1A | |||||
| Gene ID | 498 | |||||
| Synonyms |
ATP5A; ATP5A1; ATP5AL2; ATPM; ATP synthase subunit alpha, mitochondrial; ATP synthase F1 subunit alpha |
|||||
| 3D Structure | ||||||
| Sequence |
MLSVRVAAAVVRALPRRAGLVSRNALGSSFIAARNFHASNTHLQKTGTAEMSSILEERIL
GADTSVDLEETGRVLSIGDGIARVHGLRNVQAEEMVEFSSGLKGMSLNLEPDNVGVVVFG NDKLIKEGDIVKRTGAIVDVPVGEELLGRVVDALGNAIDGKGPIGSKTRRRVGLKAPGII PRISVREPMQTGIKAVDSLVPIGRGQRELIIGDRQTGKTSIAIDTIINQKRFNDGSDEKK KLYCIYVAIGQKRSTVAQLVKRLTDADAMKYTIVVSATASDAAPLQYLAPYSGCSMGEYF RDNGKHALIIYDDLSKQAVAYRQMSLLLRRPPGREAYPGDVFYLHSRLLERAAKMNDAFG GGSLTALPVIETQAGDVSAYIPTNVISITDGQIFLETELFYKGIRPAINVGLSVSRVGSA AQTRAMKQVAGTMKLELAQYREVAAFAQFGSDLDAATQQLLSRGVRLTELLKQGQYSPMA IEEQVAVIYAGVRGYLDKLEPSKITKFENAFLSHVVSQHQALLGTIRADGKISEQSDAKL KEIVTNFLAGFEA |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
ATPase alpha/beta chains family
|
|||||
| Subcellular location |
Mitochondrion
|
|||||
| Function |
Mitochondrial membrane ATP synthase (F(1)F(0) ATP synthase or Complex V) produces ATP from ADP in the presence of a proton gradient across the membrane which is generated by electron transport complexes of the respiratory chain. F-type ATPases consist of two structural domains, F(1) - containing the extramembraneous catalytic core, and F(0) - containing the membrane proton channel, linked together by a central stalk and a peripheral stalk. During catalysis, ATP synthesis in the catalytic domain of F(1) is coupled via a rotary mechanism of the central stalk subunits to proton translocation. Subunits alpha and beta form the catalytic core in F(1). Rotation of the central stalk against the surrounding alpha(3)beta(3) subunits leads to hydrolysis of ATP in three separate catalytic sites on the beta subunits. Subunit alpha does not bear the catalytic high-affinity ATP-binding sites. Binds the bacterial siderophore enterobactin and can promote mitochondrial accumulation of enterobactin-derived iron ions.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| COLO678 | SNV: p.A282G | DBIA Probe Info | |||
| JURKAT | SNV: p.R405H | . | |||
| RL952 | SNV: p.T278M | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P2 Probe Info |
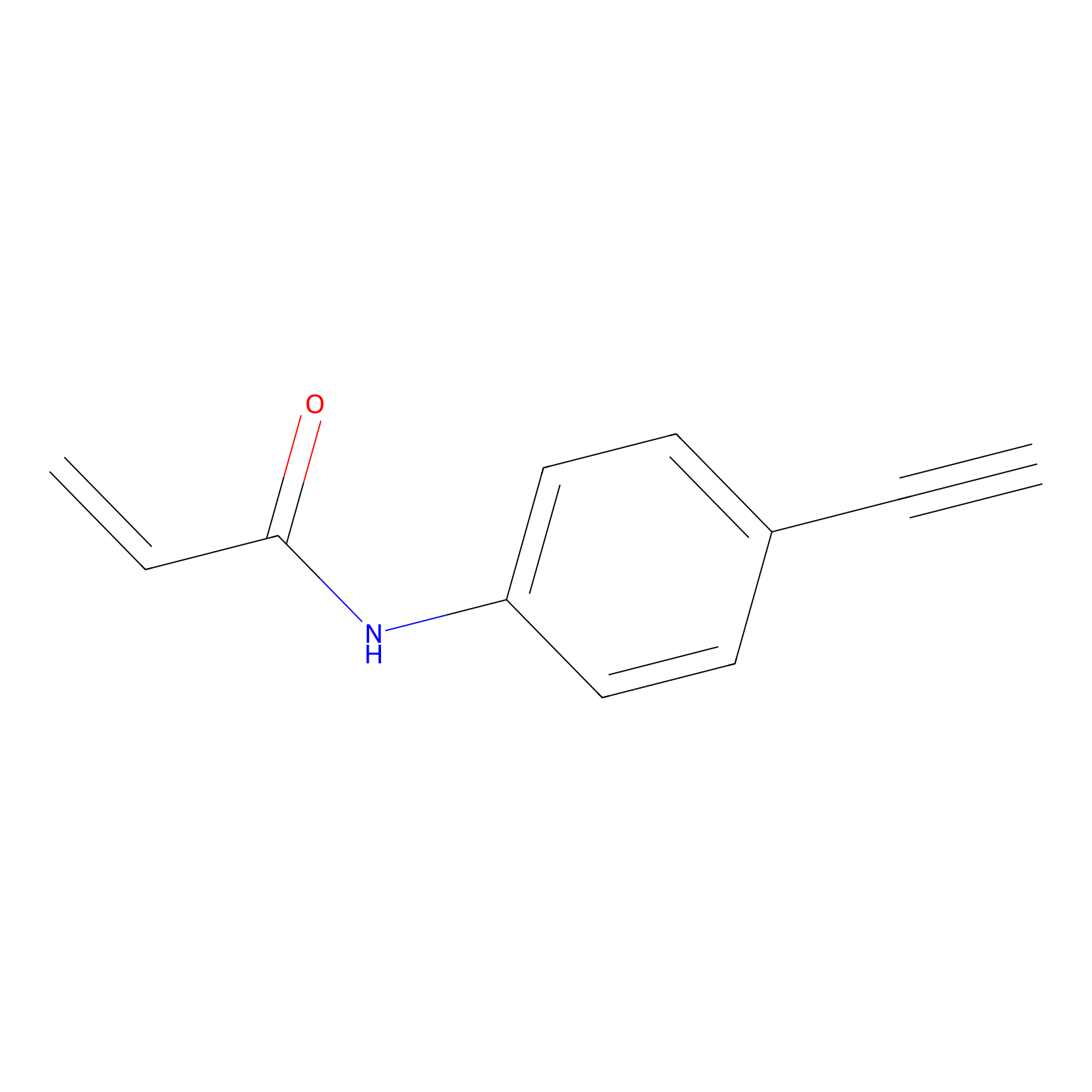 |
2.40 | LDD0449 | [1] | |
|
P8 Probe Info |
 |
3.96 | LDD0451 | [1] | |
|
A-EBA Probe Info |
 |
3.08 | LDD0215 | [2] | |
|
N1 Probe Info |
 |
5.23 | LDD0242 | [3] | |
|
C-Sul Probe Info |
 |
3.44 | LDD0066 | [4] | |
|
TH211 Probe Info |
 |
Y311(6.56) | LDD0257 | [5] | |
|
TH214 Probe Info |
 |
Y311(19.27); Y343(6.16) | LDD0258 | [5] | |
|
TH216 Probe Info |
 |
Y321(15.84); Y337(14.12); Y343(12.02) | LDD0259 | [5] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [6] | |
|
1oxF11yne Probe Info |
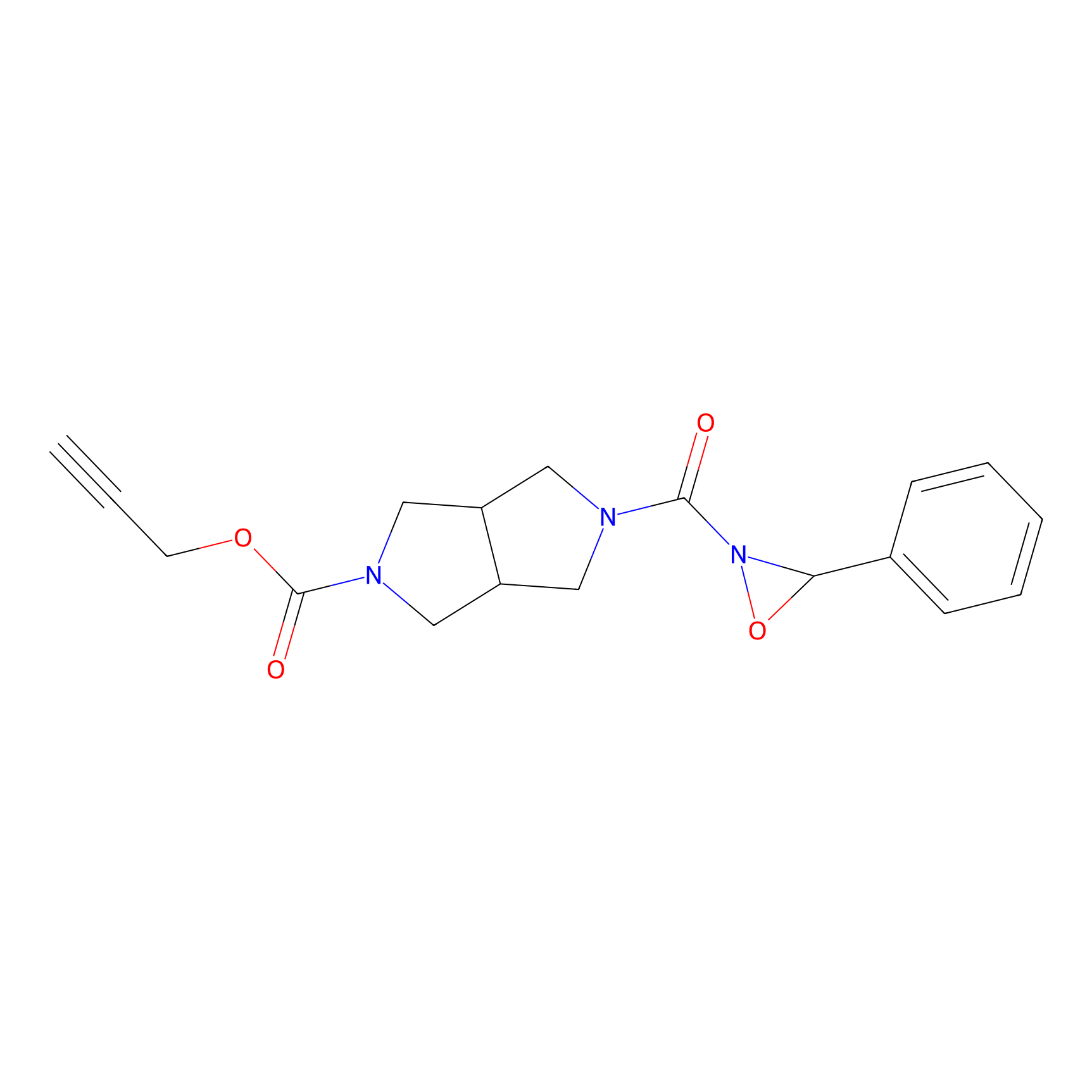 |
N.A. | LDD0193 | [7] | |
|
AZ-9 Probe Info |
 |
E69(10.00); E50(10.00); E57(10.00) | LDD2209 | [8] | |
|
ONAyne Probe Info |
 |
K230(0.00); K194(0.00); K132(0.00); K175(0.00) | LDD0273 | [9] | |
|
IPM Probe Info |
 |
C244(0.00); C294(0.00) | LDD0241 | [10] | |
|
OPA-S-S-alkyne Probe Info |
 |
K434(1.73); K132(1.76); K194(1.81); K167(2.05) | LDD3494 | [11] | |
|
Probe 1 Probe Info |
 |
Y321(87.68); Y337(7.70); Y343(10.59); Y440(33.95) | LDD3495 | [12] | |
|
THZ1-DTB Probe Info |
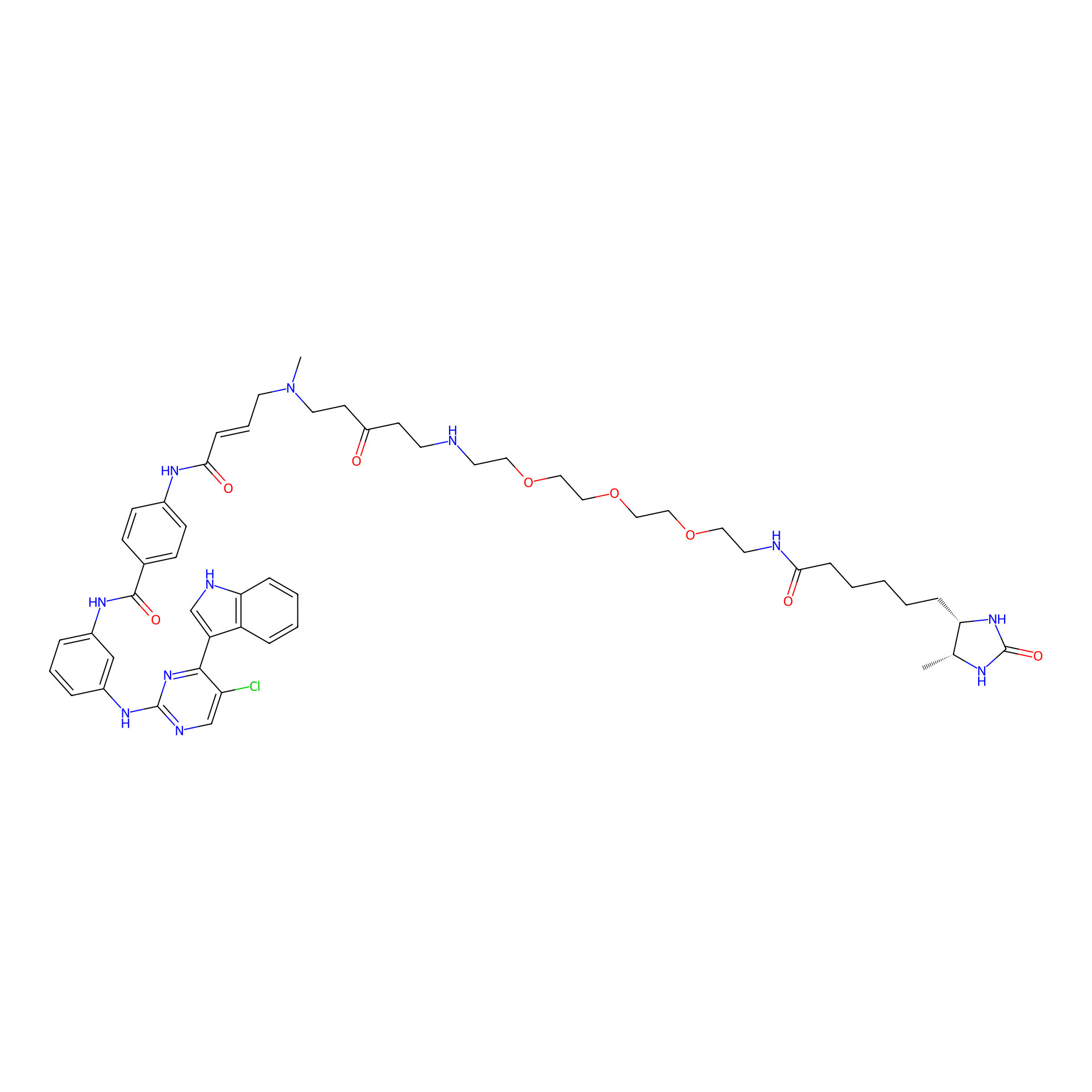 |
C294(1.03) | LDD0460 | [13] | |
|
EA-probe Probe Info |
 |
C294(1.26) | LDD2210 | [14] | |
|
HHS-482 Probe Info |
 |
Y337(0.62); Y343(0.85) | LDD0285 | [15] | |
|
DBIA Probe Info |
 |
C294(1.50) | LDD0078 | [16] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [17] | |
|
AMP probe Probe Info |
 |
N.A. | LDD0200 | [18] | |
|
ATP probe Probe Info |
 |
K531(0.00); K427(0.00); K506(0.00); K498(0.00) | LDD0199 | [18] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [17] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C244(0.00); C294(0.00) | LDD0038 | [19] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [20] | |
|
Lodoacetamide azide Probe Info |
 |
C244(0.00); C294(0.00) | LDD0037 | [19] | |
|
ATP probe Probe Info |
 |
K230(0.00); K167(0.00); K161(0.00); K427(0.00) | LDD0035 | [21] | |
|
NHS Probe Info |
 |
K230(0.00); K539(0.00); K434(0.00); K506(0.00) | LDD0010 | [22] | |
|
SF Probe Info |
 |
K261(0.00); K531(0.00); K175(0.00); Y337(0.00) | LDD0028 | [23] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [22] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [24] | |
|
1c-yne Probe Info |
 |
K503(0.00); K175(0.00); K305(0.00); K218(0.00) | LDD0228 | [25] | |
|
1d-yne Probe Info |
 |
K472(0.00); K161(0.00) | LDD0357 | [25] | |
|
Acrolein Probe Info |
 |
H514(0.00); H519(0.00); H306(0.00); C244(0.00) | LDD0217 | [26] | |
|
Crotonaldehyde Probe Info |
 |
H306(0.00); H345(0.00); C244(0.00) | LDD0219 | [26] | |
|
W1 Probe Info |
 |
D197(0.00); S198(0.00) | LDD0236 | [10] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [27] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [28] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [27] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [27] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [29] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C218 Probe Info |
 |
15.78 | LDD1892 | [30] | |
|
C305 Probe Info |
 |
12.73 | LDD1974 | [30] | |
|
C326 Probe Info |
 |
9.71 | LDD1990 | [30] | |
|
C413 Probe Info |
 |
20.97 | LDD2069 | [30] | |
|
FFF probe11 Probe Info |
 |
5.01 | LDD0471 | [31] | |
|
FFF probe13 Probe Info |
 |
13.79 | LDD0475 | [31] | |
|
FFF probe14 Probe Info |
 |
12.38 | LDD0477 | [31] | |
|
FFF probe2 Probe Info |
 |
6.38 | LDD0463 | [31] | |
|
FFF probe3 Probe Info |
 |
14.46 | LDD0464 | [31] | |
|
JN0003 Probe Info |
 |
10.29 | LDD0469 | [31] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0139 | [32] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [33] | |
|
BD-F Probe Info |
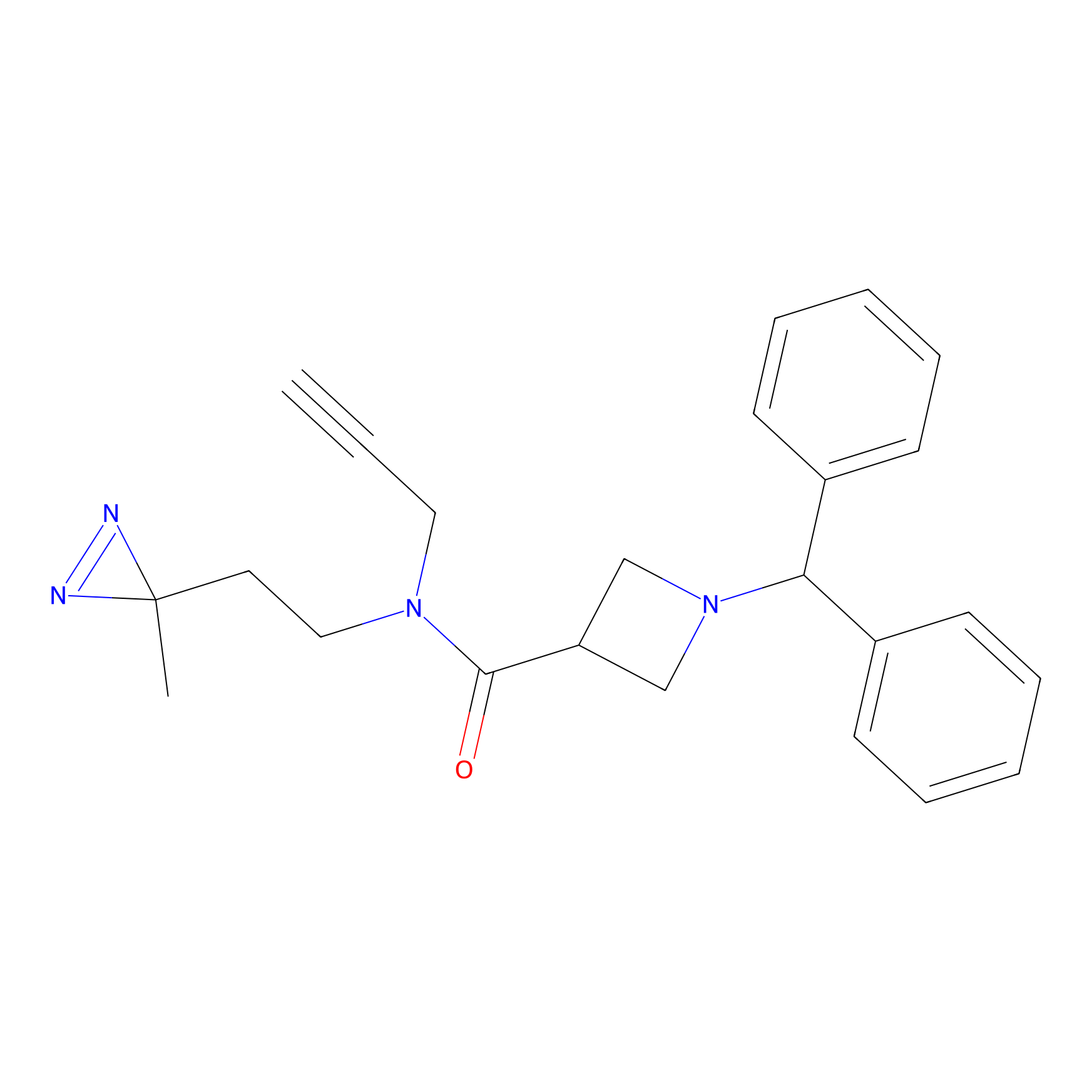 |
D452(0.00); V443(0.00); Q448(0.00); A444(0.00) | LDD0024 | [34] | |
|
LD-F Probe Info |
 |
N.A. | LDD0015 | [34] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [35] | |
|
OEA-DA Probe Info |
 |
5.38 | LDD0046 | [36] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0069 | [37] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HCT 116 | C244(0.77); C294(0.86) | LDD0531 | [16] |
| LDCM0215 | AC10 | HCT 116 | C244(1.03); C294(0.67) | LDD0532 | [16] |
| LDCM0216 | AC100 | HCT 116 | C244(1.00); C294(0.80) | LDD0533 | [16] |
| LDCM0217 | AC101 | HCT 116 | C244(0.96); C294(1.02) | LDD0534 | [16] |
| LDCM0218 | AC102 | HCT 116 | C244(0.96); C294(1.30) | LDD0535 | [16] |
| LDCM0219 | AC103 | HCT 116 | C244(0.94); C294(1.21) | LDD0536 | [16] |
| LDCM0220 | AC104 | HCT 116 | C244(1.11); C294(1.16) | LDD0537 | [16] |
| LDCM0221 | AC105 | HCT 116 | C244(0.95); C294(1.33) | LDD0538 | [16] |
| LDCM0222 | AC106 | HCT 116 | C244(1.05); C294(1.21) | LDD0539 | [16] |
| LDCM0223 | AC107 | HCT 116 | C244(1.08); C294(1.49) | LDD0540 | [16] |
| LDCM0224 | AC108 | HCT 116 | C244(1.18); C294(0.72) | LDD0541 | [16] |
| LDCM0225 | AC109 | HCT 116 | C244(1.16); C294(0.64) | LDD0542 | [16] |
| LDCM0226 | AC11 | HCT 116 | C244(1.08); C294(0.59) | LDD0543 | [16] |
| LDCM0227 | AC110 | HCT 116 | C244(1.17); C294(0.82) | LDD0544 | [16] |
| LDCM0228 | AC111 | HCT 116 | C244(0.99); C294(0.92) | LDD0545 | [16] |
| LDCM0229 | AC112 | HCT 116 | C244(0.86); C294(1.02) | LDD0546 | [16] |
| LDCM0230 | AC113 | HCT 116 | C244(1.08); C294(0.91) | LDD0547 | [16] |
| LDCM0231 | AC114 | HCT 116 | C244(0.89); C294(0.65) | LDD0548 | [16] |
| LDCM0232 | AC115 | HCT 116 | C244(0.79); C294(0.65) | LDD0549 | [16] |
| LDCM0233 | AC116 | HCT 116 | C244(0.88); C294(0.50) | LDD0550 | [16] |
| LDCM0234 | AC117 | HCT 116 | C244(1.06); C294(0.63) | LDD0551 | [16] |
| LDCM0235 | AC118 | HCT 116 | C244(1.12); C294(0.55) | LDD0552 | [16] |
| LDCM0236 | AC119 | HCT 116 | C244(0.94); C294(0.44) | LDD0553 | [16] |
| LDCM0237 | AC12 | HCT 116 | C244(1.25); C294(0.74) | LDD0554 | [16] |
| LDCM0238 | AC120 | HCT 116 | C244(0.92); C294(0.52) | LDD0555 | [16] |
| LDCM0239 | AC121 | HCT 116 | C244(1.13); C294(0.45) | LDD0556 | [16] |
| LDCM0240 | AC122 | HCT 116 | C244(0.91); C294(0.48) | LDD0557 | [16] |
| LDCM0241 | AC123 | HCT 116 | C244(1.61); C294(0.56) | LDD0558 | [16] |
| LDCM0242 | AC124 | HCT 116 | C244(1.07); C294(0.61) | LDD0559 | [16] |
| LDCM0243 | AC125 | HCT 116 | C244(0.93); C294(0.49) | LDD0560 | [16] |
| LDCM0244 | AC126 | HCT 116 | C244(0.84); C294(0.50) | LDD0561 | [16] |
| LDCM0245 | AC127 | HCT 116 | C244(0.87); C294(0.64) | LDD0562 | [16] |
| LDCM0246 | AC128 | HCT 116 | C244(1.18) | LDD0563 | [16] |
| LDCM0247 | AC129 | HCT 116 | C244(1.26) | LDD0564 | [16] |
| LDCM0249 | AC130 | HCT 116 | C244(1.08) | LDD0566 | [16] |
| LDCM0250 | AC131 | HCT 116 | C244(1.24) | LDD0567 | [16] |
| LDCM0251 | AC132 | HCT 116 | C244(1.16) | LDD0568 | [16] |
| LDCM0252 | AC133 | HCT 116 | C244(0.98) | LDD0569 | [16] |
| LDCM0253 | AC134 | HCT 116 | C244(0.89) | LDD0570 | [16] |
| LDCM0254 | AC135 | HCT 116 | C244(0.93) | LDD0571 | [16] |
| LDCM0255 | AC136 | HCT 116 | C244(0.95) | LDD0572 | [16] |
| LDCM0256 | AC137 | HCT 116 | C244(1.16) | LDD0573 | [16] |
| LDCM0257 | AC138 | HCT 116 | C244(1.09) | LDD0574 | [16] |
| LDCM0258 | AC139 | HCT 116 | C244(0.88) | LDD0575 | [16] |
| LDCM0259 | AC14 | HCT 116 | C244(1.22); C294(0.50) | LDD0576 | [16] |
| LDCM0260 | AC140 | HCT 116 | C244(0.85) | LDD0577 | [16] |
| LDCM0261 | AC141 | HCT 116 | C244(0.92) | LDD0578 | [16] |
| LDCM0262 | AC142 | HCT 116 | C244(1.12) | LDD0579 | [16] |
| LDCM0263 | AC143 | HCT 116 | C244(0.77); C294(0.68) | LDD0580 | [16] |
| LDCM0264 | AC144 | HCT 116 | C244(0.82); C294(0.83) | LDD0581 | [16] |
| LDCM0265 | AC145 | HCT 116 | C294(0.75); C244(0.90) | LDD0582 | [16] |
| LDCM0266 | AC146 | HCT 116 | C244(0.74); C294(0.83) | LDD0583 | [16] |
| LDCM0267 | AC147 | HCT 116 | C294(0.78); C244(0.83) | LDD0584 | [16] |
| LDCM0268 | AC148 | HCT 116 | C294(0.57); C244(0.64) | LDD0585 | [16] |
| LDCM0269 | AC149 | HCT 116 | C294(0.63); C244(0.64) | LDD0586 | [16] |
| LDCM0270 | AC15 | HCT 116 | C294(0.58); C244(1.26) | LDD0587 | [16] |
| LDCM0271 | AC150 | HCT 116 | C294(0.84); C244(0.92) | LDD0588 | [16] |
| LDCM0272 | AC151 | HCT 116 | C244(0.90); C294(1.06) | LDD0589 | [16] |
| LDCM0273 | AC152 | HCT 116 | C244(0.71); C294(0.83) | LDD0590 | [16] |
| LDCM0274 | AC153 | HCT 116 | C294(0.50); C244(0.53) | LDD0591 | [16] |
| LDCM0621 | AC154 | HCT 116 | C244(0.82); C294(0.97) | LDD2158 | [16] |
| LDCM0622 | AC155 | HCT 116 | C244(0.80); C294(0.86) | LDD2159 | [16] |
| LDCM0623 | AC156 | HCT 116 | C244(0.87); C294(1.14) | LDD2160 | [16] |
| LDCM0624 | AC157 | HCT 116 | C244(0.89); C294(1.75) | LDD2161 | [16] |
| LDCM0276 | AC17 | HCT 116 | C244(1.13); C294(1.71) | LDD0593 | [16] |
| LDCM0277 | AC18 | HCT 116 | C294(0.99); C244(1.07) | LDD0594 | [16] |
| LDCM0278 | AC19 | HCT 116 | C244(1.04); C294(1.55) | LDD0595 | [16] |
| LDCM0279 | AC2 | HCT 116 | C244(0.63); C294(0.76) | LDD0596 | [16] |
| LDCM0280 | AC20 | HCT 116 | C244(1.02); C294(1.43) | LDD0597 | [16] |
| LDCM0281 | AC21 | HCT 116 | C244(0.87); C294(1.08) | LDD0598 | [16] |
| LDCM0282 | AC22 | HCT 116 | C244(0.92); C294(1.41) | LDD0599 | [16] |
| LDCM0283 | AC23 | HCT 116 | C244(1.01); C294(1.52) | LDD0600 | [16] |
| LDCM0284 | AC24 | HCT 116 | C244(1.15); C294(1.77) | LDD0601 | [16] |
| LDCM0285 | AC25 | HCT 116 | C244(1.16); C294(1.32) | LDD0602 | [16] |
| LDCM0286 | AC26 | HCT 116 | C294(0.91); C244(0.92) | LDD0603 | [16] |
| LDCM0287 | AC27 | HCT 116 | C244(1.04); C294(1.17) | LDD0604 | [16] |
| LDCM0288 | AC28 | HCT 116 | C244(0.98); C294(1.00) | LDD0605 | [16] |
| LDCM0289 | AC29 | HCT 116 | C244(0.85); C294(1.21) | LDD0606 | [16] |
| LDCM0290 | AC3 | HCT 116 | C244(0.59); C294(0.73) | LDD0607 | [16] |
| LDCM0291 | AC30 | HCT 116 | C244(0.77); C294(1.14) | LDD0608 | [16] |
| LDCM0292 | AC31 | HCT 116 | C244(0.98); C294(1.02) | LDD0609 | [16] |
| LDCM0293 | AC32 | HCT 116 | C244(0.74); C294(0.81) | LDD0610 | [16] |
| LDCM0294 | AC33 | HCT 116 | C294(0.99); C244(1.14) | LDD0611 | [16] |
| LDCM0295 | AC34 | HCT 116 | C294(1.14); C244(1.33) | LDD0612 | [16] |
| LDCM0296 | AC35 | HCT 116 | C294(1.17); C244(1.37) | LDD0613 | [16] |
| LDCM0297 | AC36 | HCT 116 | C294(0.70); C244(1.12) | LDD0614 | [16] |
| LDCM0298 | AC37 | HCT 116 | C294(0.54); C244(1.13) | LDD0615 | [16] |
| LDCM0299 | AC38 | HCT 116 | C294(0.69); C244(1.17) | LDD0616 | [16] |
| LDCM0300 | AC39 | HCT 116 | C294(0.50); C244(1.06) | LDD0617 | [16] |
| LDCM0301 | AC4 | HCT 116 | C244(0.63); C294(0.75) | LDD0618 | [16] |
| LDCM0302 | AC40 | HCT 116 | C294(0.68); C244(0.87) | LDD0619 | [16] |
| LDCM0303 | AC41 | HCT 116 | C294(0.58); C244(1.06) | LDD0620 | [16] |
| LDCM0304 | AC42 | HCT 116 | C294(0.67); C244(1.06) | LDD0621 | [16] |
| LDCM0305 | AC43 | HCT 116 | C294(0.70); C244(1.16) | LDD0622 | [16] |
| LDCM0306 | AC44 | HCT 116 | C294(0.66); C244(1.11) | LDD0623 | [16] |
| LDCM0307 | AC45 | HCT 116 | C294(0.61); C244(0.91) | LDD0624 | [16] |
| LDCM0308 | AC46 | HCT 116 | C294(0.90); C244(0.92) | LDD0625 | [16] |
| LDCM0309 | AC47 | HCT 116 | C294(0.71); C244(0.86) | LDD0626 | [16] |
| LDCM0310 | AC48 | HCT 116 | C294(0.64); C244(1.00) | LDD0627 | [16] |
| LDCM0311 | AC49 | HCT 116 | C294(0.80); C244(1.03) | LDD0628 | [16] |
| LDCM0312 | AC5 | HCT 116 | C294(0.55); C244(0.58) | LDD0629 | [16] |
| LDCM0313 | AC50 | HCT 116 | C294(0.76); C244(0.81) | LDD0630 | [16] |
| LDCM0314 | AC51 | HCT 116 | C294(0.77); C244(1.22) | LDD0631 | [16] |
| LDCM0315 | AC52 | HCT 116 | C294(0.77); C244(0.96) | LDD0632 | [16] |
| LDCM0316 | AC53 | HCT 116 | C294(0.88); C244(0.91) | LDD0633 | [16] |
| LDCM0317 | AC54 | HCT 116 | C244(0.85); C294(0.88) | LDD0634 | [16] |
| LDCM0318 | AC55 | HCT 116 | C244(0.79); C294(0.84) | LDD0635 | [16] |
| LDCM0319 | AC56 | HCT 116 | C294(0.67); C244(0.69) | LDD0636 | [16] |
| LDCM0320 | AC57 | HCT 116 | C294(0.67); C244(1.28) | LDD0637 | [16] |
| LDCM0321 | AC58 | HCT 116 | C294(0.81); C244(1.09) | LDD0638 | [16] |
| LDCM0322 | AC59 | HCT 116 | C294(0.65); C244(1.25) | LDD0639 | [16] |
| LDCM0323 | AC6 | HCT 116 | C294(0.62); C244(0.96) | LDD0640 | [16] |
| LDCM0324 | AC60 | HCT 116 | C294(0.67); C244(1.29) | LDD0641 | [16] |
| LDCM0325 | AC61 | HCT 116 | C294(0.79); C244(1.58) | LDD0642 | [16] |
| LDCM0326 | AC62 | HCT 116 | C294(0.81); C244(1.18) | LDD0643 | [16] |
| LDCM0327 | AC63 | HCT 116 | C294(0.76); C244(1.55) | LDD0644 | [16] |
| LDCM0328 | AC64 | HCT 116 | C294(0.64); C244(1.30) | LDD0645 | [16] |
| LDCM0329 | AC65 | HCT 116 | C294(0.66); C244(1.65) | LDD0646 | [16] |
| LDCM0330 | AC66 | HCT 116 | C294(0.83); C244(1.78) | LDD0647 | [16] |
| LDCM0331 | AC67 | HCT 116 | C294(0.35); C244(1.06) | LDD0648 | [16] |
| LDCM0332 | AC68 | HCT 116 | C294(0.61); C244(0.84) | LDD0649 | [16] |
| LDCM0333 | AC69 | HCT 116 | C294(0.76); C244(0.79) | LDD0650 | [16] |
| LDCM0334 | AC7 | HCT 116 | C294(0.68); C244(1.14) | LDD0651 | [16] |
| LDCM0335 | AC70 | HCT 116 | C244(0.49); C294(0.50) | LDD0652 | [16] |
| LDCM0336 | AC71 | HCT 116 | C294(0.70); C244(1.06) | LDD0653 | [16] |
| LDCM0337 | AC72 | HCT 116 | C244(0.64); C294(0.87) | LDD0654 | [16] |
| LDCM0338 | AC73 | HCT 116 | C244(0.39); C294(0.49) | LDD0655 | [16] |
| LDCM0339 | AC74 | HCT 116 | C294(0.43); C244(0.50) | LDD0656 | [16] |
| LDCM0340 | AC75 | HCT 116 | C244(0.38); C294(0.41) | LDD0657 | [16] |
| LDCM0341 | AC76 | HCT 116 | C294(0.63); C244(0.75) | LDD0658 | [16] |
| LDCM0342 | AC77 | HCT 116 | C294(0.56); C244(0.62) | LDD0659 | [16] |
| LDCM0343 | AC78 | HCT 116 | C294(0.85); C244(0.96) | LDD0660 | [16] |
| LDCM0344 | AC79 | HCT 116 | C244(0.73); C294(0.75) | LDD0661 | [16] |
| LDCM0345 | AC8 | HCT 116 | C294(0.84); C244(1.10) | LDD0662 | [16] |
| LDCM0346 | AC80 | HCT 116 | C294(0.60); C244(0.79) | LDD0663 | [16] |
| LDCM0347 | AC81 | HCT 116 | C294(0.67); C244(0.90) | LDD0664 | [16] |
| LDCM0348 | AC82 | HCT 116 | C294(0.21); C244(0.40) | LDD0665 | [16] |
| LDCM0349 | AC83 | HCT 116 | C244(0.39); C294(0.67) | LDD0666 | [16] |
| LDCM0350 | AC84 | HCT 116 | C244(0.50); C294(0.93) | LDD0667 | [16] |
| LDCM0351 | AC85 | HCT 116 | C244(0.75); C294(0.95) | LDD0668 | [16] |
| LDCM0352 | AC86 | HCT 116 | C244(0.67); C294(0.94) | LDD0669 | [16] |
| LDCM0353 | AC87 | HCT 116 | C244(0.77); C294(1.22) | LDD0670 | [16] |
| LDCM0354 | AC88 | HCT 116 | C244(0.69); C294(0.90) | LDD0671 | [16] |
| LDCM0355 | AC89 | HCT 116 | C244(0.56); C294(1.16) | LDD0672 | [16] |
| LDCM0357 | AC90 | HCT 116 | C244(0.79); C294(1.19) | LDD0674 | [16] |
| LDCM0358 | AC91 | HCT 116 | C244(0.38); C294(0.86) | LDD0675 | [16] |
| LDCM0359 | AC92 | HCT 116 | C244(0.43); C294(0.97) | LDD0676 | [16] |
| LDCM0360 | AC93 | HCT 116 | C244(0.73); C294(1.01) | LDD0677 | [16] |
| LDCM0361 | AC94 | HCT 116 | C244(0.91); C294(1.78) | LDD0678 | [16] |
| LDCM0362 | AC95 | HCT 116 | C244(0.73); C294(2.51) | LDD0679 | [16] |
| LDCM0363 | AC96 | HCT 116 | C244(0.59); C294(1.25) | LDD0680 | [16] |
| LDCM0364 | AC97 | HCT 116 | C244(0.45); C294(1.04) | LDD0681 | [16] |
| LDCM0365 | AC98 | HCT 116 | C294(0.36); C244(0.91) | LDD0682 | [16] |
| LDCM0366 | AC99 | HCT 116 | C244(1.05); C294(1.61) | LDD0683 | [16] |
| LDCM0248 | AKOS034007472 | HCT 116 | C244(1.25); C294(0.54) | LDD0565 | [16] |
| LDCM0356 | AKOS034007680 | HCT 116 | C294(0.61); C244(1.08) | LDD0673 | [16] |
| LDCM0275 | AKOS034007705 | HCT 116 | C294(0.47); C244(0.85) | LDD0592 | [16] |
| LDCM0020 | ARS-1620 | HCC44 | C294(1.50) | LDD0078 | [16] |
| LDCM0108 | Chloroacetamide | HeLa | H514(0.00); H519(0.00); H306(0.00); C244(0.00) | LDD0222 | [26] |
| LDCM0632 | CL-Sc | Hep-G2 | C244(1.30); C244(0.73) | LDD2227 | [28] |
| LDCM0367 | CL1 | HCT 116 | C244(1.02); C294(1.22) | LDD0684 | [16] |
| LDCM0368 | CL10 | HCT 116 | C244(0.88); C294(0.93) | LDD0685 | [16] |
| LDCM0369 | CL100 | HCT 116 | C294(0.74); C244(0.86) | LDD0686 | [16] |
| LDCM0370 | CL101 | HCT 116 | C294(0.50); C244(1.08) | LDD0687 | [16] |
| LDCM0371 | CL102 | HCT 116 | C294(0.68); C244(1.28) | LDD0688 | [16] |
| LDCM0372 | CL103 | HCT 116 | C294(0.87); C244(1.16) | LDD0689 | [16] |
| LDCM0373 | CL104 | HCT 116 | C294(0.86); C244(1.10) | LDD0690 | [16] |
| LDCM0374 | CL105 | HCT 116 | C294(0.89); C244(0.92) | LDD0691 | [16] |
| LDCM0375 | CL106 | HCT 116 | C244(0.79); C294(0.95) | LDD0692 | [16] |
| LDCM0376 | CL107 | HCT 116 | C294(0.81); C244(1.00) | LDD0693 | [16] |
| LDCM0377 | CL108 | HCT 116 | C244(0.91); C294(1.04) | LDD0694 | [16] |
| LDCM0378 | CL109 | HCT 116 | C244(0.89); C294(1.20) | LDD0695 | [16] |
| LDCM0379 | CL11 | HCT 116 | C244(0.88); C294(0.89) | LDD0696 | [16] |
| LDCM0380 | CL110 | HCT 116 | C294(0.95); C244(0.95) | LDD0697 | [16] |
| LDCM0381 | CL111 | HCT 116 | C244(0.93); C294(1.35) | LDD0698 | [16] |
| LDCM0382 | CL112 | HCT 116 | C294(0.83); C244(1.02) | LDD0699 | [16] |
| LDCM0383 | CL113 | HCT 116 | C244(0.92); C294(1.16) | LDD0700 | [16] |
| LDCM0384 | CL114 | HCT 116 | C244(0.98); C294(1.07) | LDD0701 | [16] |
| LDCM0385 | CL115 | HCT 116 | C244(1.01); C294(1.04) | LDD0702 | [16] |
| LDCM0386 | CL116 | HCT 116 | C294(0.71); C244(1.19) | LDD0703 | [16] |
| LDCM0387 | CL117 | HCT 116 | C294(0.53); C244(0.73) | LDD0704 | [16] |
| LDCM0388 | CL118 | HCT 116 | C294(0.71); C244(1.24) | LDD0705 | [16] |
| LDCM0389 | CL119 | HCT 116 | C294(0.84); C244(0.97) | LDD0706 | [16] |
| LDCM0390 | CL12 | HCT 116 | C244(0.64); C294(0.86) | LDD0707 | [16] |
| LDCM0391 | CL120 | HCT 116 | C294(0.53); C244(1.08) | LDD0708 | [16] |
| LDCM0392 | CL121 | HCT 116 | C294(0.65); C244(1.09) | LDD0709 | [16] |
| LDCM0393 | CL122 | HCT 116 | C294(0.67); C244(0.96) | LDD0710 | [16] |
| LDCM0394 | CL123 | HCT 116 | C294(0.81); C244(0.83) | LDD0711 | [16] |
| LDCM0395 | CL124 | HCT 116 | C244(0.76); C294(0.81) | LDD0712 | [16] |
| LDCM0396 | CL125 | HCT 116 | C294(0.85); C244(1.32) | LDD0713 | [16] |
| LDCM0397 | CL126 | HCT 116 | C294(0.77); C244(1.52) | LDD0714 | [16] |
| LDCM0398 | CL127 | HCT 116 | C294(0.90); C244(1.49) | LDD0715 | [16] |
| LDCM0399 | CL128 | HCT 116 | C294(0.64); C244(1.04) | LDD0716 | [16] |
| LDCM0400 | CL13 | HCT 116 | C244(1.02); C294(1.17) | LDD0717 | [16] |
| LDCM0401 | CL14 | HCT 116 | C244(0.95); C294(1.19) | LDD0718 | [16] |
| LDCM0402 | CL15 | HCT 116 | C294(0.92); C244(1.05) | LDD0719 | [16] |
| LDCM0403 | CL16 | HCT 116 | C244(0.73); C294(1.06) | LDD0720 | [16] |
| LDCM0404 | CL17 | HCT 116 | C244(0.92); C294(0.98) | LDD0721 | [16] |
| LDCM0405 | CL18 | HCT 116 | C244(1.02); C294(1.46) | LDD0722 | [16] |
| LDCM0406 | CL19 | HCT 116 | C244(0.89); C294(1.07) | LDD0723 | [16] |
| LDCM0407 | CL2 | HCT 116 | C244(1.05); C294(1.08) | LDD0724 | [16] |
| LDCM0408 | CL20 | HCT 116 | C244(0.69); C294(1.20) | LDD0725 | [16] |
| LDCM0409 | CL21 | HCT 116 | C294(0.81); C244(0.85) | LDD0726 | [16] |
| LDCM0410 | CL22 | HCT 116 | C244(0.76); C294(0.83) | LDD0727 | [16] |
| LDCM0411 | CL23 | HCT 116 | C244(0.78); C294(1.16) | LDD0728 | [16] |
| LDCM0412 | CL24 | HCT 116 | C244(0.68); C294(1.03) | LDD0729 | [16] |
| LDCM0413 | CL25 | HCT 116 | C244(0.69); C294(0.78) | LDD0730 | [16] |
| LDCM0414 | CL26 | HCT 116 | C244(0.73); C294(1.24) | LDD0731 | [16] |
| LDCM0415 | CL27 | HCT 116 | C244(0.91); C294(0.96) | LDD0732 | [16] |
| LDCM0416 | CL28 | HCT 116 | C244(0.75); C294(0.96) | LDD0733 | [16] |
| LDCM0417 | CL29 | HCT 116 | C244(0.71); C294(0.81) | LDD0734 | [16] |
| LDCM0418 | CL3 | HCT 116 | C244(1.03); C294(1.22) | LDD0735 | [16] |
| LDCM0419 | CL30 | HCT 116 | C244(0.85); C294(1.07) | LDD0736 | [16] |
| LDCM0420 | CL31 | HCT 116 | C244(0.87); C294(1.27) | LDD0737 | [16] |
| LDCM0421 | CL32 | HCT 116 | C244(0.86); C294(1.39) | LDD0738 | [16] |
| LDCM0422 | CL33 | HCT 116 | C244(0.91); C294(1.65) | LDD0739 | [16] |
| LDCM0423 | CL34 | HCT 116 | C244(0.99); C294(1.10) | LDD0740 | [16] |
| LDCM0424 | CL35 | HCT 116 | C244(0.76); C294(1.61) | LDD0741 | [16] |
| LDCM0425 | CL36 | HCT 116 | C244(1.51); C294(1.61) | LDD0742 | [16] |
| LDCM0426 | CL37 | HCT 116 | C244(0.64); C294(1.53) | LDD0743 | [16] |
| LDCM0428 | CL39 | HCT 116 | C244(0.74); C294(1.19) | LDD0745 | [16] |
| LDCM0429 | CL4 | HCT 116 | C244(0.92); C294(1.07) | LDD0746 | [16] |
| LDCM0430 | CL40 | HCT 116 | C244(0.76); C294(1.56) | LDD0747 | [16] |
| LDCM0431 | CL41 | HCT 116 | C244(1.33); C294(1.47) | LDD0748 | [16] |
| LDCM0432 | CL42 | HCT 116 | C244(1.28); C294(0.97) | LDD0749 | [16] |
| LDCM0433 | CL43 | HCT 116 | C244(0.87); C294(1.73) | LDD0750 | [16] |
| LDCM0434 | CL44 | HCT 116 | C244(0.97); C294(1.65) | LDD0751 | [16] |
| LDCM0435 | CL45 | HCT 116 | C244(0.92); C294(1.06) | LDD0752 | [16] |
| LDCM0436 | CL46 | HCT 116 | C244(0.91); C294(1.69) | LDD0753 | [16] |
| LDCM0437 | CL47 | HCT 116 | C244(0.91); C294(1.14) | LDD0754 | [16] |
| LDCM0438 | CL48 | HCT 116 | C244(0.98); C294(1.45) | LDD0755 | [16] |
| LDCM0439 | CL49 | HCT 116 | C244(1.30); C294(2.10) | LDD0756 | [16] |
| LDCM0440 | CL5 | HCT 116 | C244(1.05); C294(1.19) | LDD0757 | [16] |
| LDCM0441 | CL50 | HCT 116 | C244(0.89); C294(1.32) | LDD0758 | [16] |
| LDCM0442 | CL51 | HCT 116 | C244(0.92); C294(1.45) | LDD0759 | [16] |
| LDCM0443 | CL52 | HCT 116 | C244(0.71); C294(1.20) | LDD0760 | [16] |
| LDCM0444 | CL53 | HCT 116 | C244(0.79); C294(2.12) | LDD0761 | [16] |
| LDCM0445 | CL54 | HCT 116 | C244(0.82); C294(1.34) | LDD0762 | [16] |
| LDCM0446 | CL55 | HCT 116 | C244(1.02); C294(2.11) | LDD0763 | [16] |
| LDCM0447 | CL56 | HCT 116 | C244(1.03); C294(1.12) | LDD0764 | [16] |
| LDCM0448 | CL57 | HCT 116 | C244(1.16); C294(1.39) | LDD0765 | [16] |
| LDCM0449 | CL58 | HCT 116 | C244(1.08); C294(1.49) | LDD0766 | [16] |
| LDCM0450 | CL59 | HCT 116 | C244(0.99); C294(1.28) | LDD0767 | [16] |
| LDCM0451 | CL6 | HCT 116 | C244(1.02); C294(1.00) | LDD0768 | [16] |
| LDCM0452 | CL60 | HCT 116 | C244(0.92); C294(1.99) | LDD0769 | [16] |
| LDCM0453 | CL61 | HCT 116 | C244(1.04); C294(0.69) | LDD0770 | [16] |
| LDCM0454 | CL62 | HCT 116 | C244(1.02); C294(0.73) | LDD0771 | [16] |
| LDCM0455 | CL63 | HCT 116 | C244(0.89); C294(0.43) | LDD0772 | [16] |
| LDCM0456 | CL64 | HCT 116 | C244(1.06); C294(0.74) | LDD0773 | [16] |
| LDCM0457 | CL65 | HCT 116 | C244(1.15); C294(0.86) | LDD0774 | [16] |
| LDCM0458 | CL66 | HCT 116 | C244(1.16); C294(0.45) | LDD0775 | [16] |
| LDCM0459 | CL67 | HCT 116 | C244(1.00); C294(0.43) | LDD0776 | [16] |
| LDCM0460 | CL68 | HCT 116 | C244(0.81); C294(0.44) | LDD0777 | [16] |
| LDCM0461 | CL69 | HCT 116 | C244(0.93); C294(0.73) | LDD0778 | [16] |
| LDCM0462 | CL7 | HCT 116 | C244(0.94); C294(0.95) | LDD0779 | [16] |
| LDCM0463 | CL70 | HCT 116 | C244(1.00); C294(0.66) | LDD0780 | [16] |
| LDCM0464 | CL71 | HCT 116 | C244(0.87); C294(0.41) | LDD0781 | [16] |
| LDCM0465 | CL72 | HCT 116 | C244(0.91); C294(0.86) | LDD0782 | [16] |
| LDCM0466 | CL73 | HCT 116 | C244(0.88); C294(0.70) | LDD0783 | [16] |
| LDCM0467 | CL74 | HCT 116 | C244(0.92); C294(0.66) | LDD0784 | [16] |
| LDCM0469 | CL76 | HCT 116 | C244(0.83); C294(1.05) | LDD0786 | [16] |
| LDCM0470 | CL77 | HCT 116 | C244(0.66); C294(2.31) | LDD0787 | [16] |
| LDCM0471 | CL78 | HCT 116 | C244(0.84); C294(0.83) | LDD0788 | [16] |
| LDCM0472 | CL79 | HCT 116 | C244(0.77); C294(1.14) | LDD0789 | [16] |
| LDCM0473 | CL8 | HCT 116 | C244(0.85); C294(1.05) | LDD0790 | [16] |
| LDCM0474 | CL80 | HCT 116 | C244(0.90); C294(1.46) | LDD0791 | [16] |
| LDCM0475 | CL81 | HCT 116 | C244(0.83); C294(0.90) | LDD0792 | [16] |
| LDCM0476 | CL82 | HCT 116 | C244(1.12); C294(0.76) | LDD0793 | [16] |
| LDCM0477 | CL83 | HCT 116 | C244(0.66); C294(0.83) | LDD0794 | [16] |
| LDCM0478 | CL84 | HCT 116 | C244(0.71); C294(0.62) | LDD0795 | [16] |
| LDCM0479 | CL85 | HCT 116 | C244(0.80); C294(0.97) | LDD0796 | [16] |
| LDCM0480 | CL86 | HCT 116 | C244(1.03); C294(1.03) | LDD0797 | [16] |
| LDCM0481 | CL87 | HCT 116 | C244(0.85); C294(0.91) | LDD0798 | [16] |
| LDCM0482 | CL88 | HCT 116 | C244(1.00); C294(0.77) | LDD0799 | [16] |
| LDCM0483 | CL89 | HCT 116 | C244(0.90); C294(0.55) | LDD0800 | [16] |
| LDCM0484 | CL9 | HCT 116 | C244(1.01); C294(1.14) | LDD0801 | [16] |
| LDCM0485 | CL90 | HCT 116 | C244(0.90); C294(0.81) | LDD0802 | [16] |
| LDCM0486 | CL91 | HCT 116 | C244(0.53); C294(0.57) | LDD0803 | [16] |
| LDCM0487 | CL92 | HCT 116 | C244(0.66); C294(0.54) | LDD0804 | [16] |
| LDCM0488 | CL93 | HCT 116 | C244(0.74); C294(0.60) | LDD0805 | [16] |
| LDCM0489 | CL94 | HCT 116 | C244(0.78); C294(0.49) | LDD0806 | [16] |
| LDCM0490 | CL95 | HCT 116 | C244(0.68); C294(0.45) | LDD0807 | [16] |
| LDCM0491 | CL96 | HCT 116 | C244(0.52); C294(0.49) | LDD0808 | [16] |
| LDCM0492 | CL97 | HCT 116 | C244(0.67); C294(0.64) | LDD0809 | [16] |
| LDCM0493 | CL98 | HCT 116 | C244(0.51); C294(0.58) | LDD0810 | [16] |
| LDCM0494 | CL99 | HCT 116 | C244(0.79); C294(0.68) | LDD0811 | [16] |
| LDCM0495 | E2913 | HEK-293T | C244(1.06); C294(1.03) | LDD1698 | [38] |
| LDCM0175 | Ethacrynic acid | HeLa | C294(1.26) | LDD2210 | [14] |
| LDCM0572 | Fragment10 | MDA-MB-231 | C294(3.67) | LDD1389 | [39] |
| LDCM0573 | Fragment11 | MDA-MB-231 | C294(1.55) | LDD1391 | [39] |
| LDCM0574 | Fragment12 | MDA-MB-231 | C294(2.60) | LDD1393 | [39] |
| LDCM0575 | Fragment13 | MDA-MB-231 | C294(1.18) | LDD1395 | [39] |
| LDCM0576 | Fragment14 | MDA-MB-231 | C294(1.81) | LDD1397 | [39] |
| LDCM0579 | Fragment20 | MDA-MB-231 | C294(20.00) | LDD1402 | [39] |
| LDCM0580 | Fragment21 | MDA-MB-231 | C294(1.54) | LDD1404 | [39] |
| LDCM0581 | Fragment22 | MDA-MB-231 | C294(1.13) | LDD1406 | [39] |
| LDCM0582 | Fragment23 | MDA-MB-231 | C294(2.16) | LDD1408 | [39] |
| LDCM0585 | Fragment26 | Ramos | C294(1.05) | LDD1412 | [39] |
| LDCM0578 | Fragment27 | MDA-MB-231 | C294(1.02) | LDD1401 | [39] |
| LDCM0586 | Fragment28 | MDA-MB-231 | C294(1.17) | LDD1415 | [39] |
| LDCM0587 | Fragment29 | MDA-MB-231 | C294(1.25) | LDD1417 | [39] |
| LDCM0588 | Fragment30 | Ramos | C294(1.63) | LDD1420 | [39] |
| LDCM0589 | Fragment31 | MDA-MB-231 | C294(3.21) | LDD1421 | [39] |
| LDCM0590 | Fragment32 | MDA-MB-231 | C294(1.63) | LDD1423 | [39] |
| LDCM0468 | Fragment33 | HCT 116 | C244(1.06); C294(0.45) | LDD0785 | [16] |
| LDCM0592 | Fragment34 | Ramos | C294(1.17) | LDD1428 | [39] |
| LDCM0595 | Fragment37 | Ramos | C294(1.01) | LDD1432 | [39] |
| LDCM0596 | Fragment38 | MDA-MB-231 | C294(0.96) | LDD1433 | [39] |
| LDCM0566 | Fragment4 | MDA-MB-231 | C294(2.00) | LDD1378 | [39] |
| LDCM0598 | Fragment40 | Ramos | C294(0.68) | LDD1437 | [39] |
| LDCM0599 | Fragment41 | MDA-MB-231 | C294(1.01) | LDD1438 | [39] |
| LDCM0600 | Fragment42 | Ramos | C294(1.04) | LDD1440 | [39] |
| LDCM0601 | Fragment43 | MDA-MB-231 | C294(1.98) | LDD1441 | [39] |
| LDCM0602 | Fragment44 | MDA-MB-231 | C294(0.93) | LDD1443 | [39] |
| LDCM0604 | Fragment46 | MDA-MB-231 | C294(0.76) | LDD1445 | [39] |
| LDCM0605 | Fragment47 | MDA-MB-231 | C294(1.08) | LDD1446 | [39] |
| LDCM0606 | Fragment48 | MDA-MB-231 | C294(1.02) | LDD1447 | [39] |
| LDCM0607 | Fragment49 | MDA-MB-231 | C294(2.02) | LDD1448 | [39] |
| LDCM0427 | Fragment51 | HCT 116 | C244(0.91); C294(0.92) | LDD0744 | [16] |
| LDCM0610 | Fragment52 | MDA-MB-231 | C294(1.32) | LDD1452 | [39] |
| LDCM0611 | Fragment53 | MDA-MB-231 | C294(1.00) | LDD1454 | [39] |
| LDCM0612 | Fragment54 | MDA-MB-231 | C294(1.00) | LDD1456 | [39] |
| LDCM0613 | Fragment55 | MDA-MB-231 | C294(1.40) | LDD1457 | [39] |
| LDCM0614 | Fragment56 | MDA-MB-231 | C294(1.00) | LDD1458 | [39] |
| LDCM0568 | Fragment6 | MDA-MB-231 | C294(1.27) | LDD1382 | [39] |
| LDCM0570 | Fragment8 | MDA-MB-231 | C294(2.16) | LDD1385 | [39] |
| LDCM0571 | Fragment9 | MDA-MB-231 | C294(3.16) | LDD1387 | [39] |
| LDCM0107 | IAA | HeLa | H519(0.00); H306(0.00); H514(0.00); H345(0.00) | LDD0221 | [26] |
| LDCM0123 | JWB131 | DM93 | Y337(0.62); Y343(0.85) | LDD0285 | [15] |
| LDCM0124 | JWB142 | DM93 | Y337(1.42); Y343(0.82) | LDD0286 | [15] |
| LDCM0125 | JWB146 | DM93 | Y337(0.76); Y343(1.44) | LDD0287 | [15] |
| LDCM0126 | JWB150 | DM93 | Y337(4.40); Y343(2.68) | LDD0288 | [15] |
| LDCM0127 | JWB152 | DM93 | Y337(1.42); Y343(1.80) | LDD0289 | [15] |
| LDCM0128 | JWB198 | DM93 | Y337(1.03); Y343(1.65) | LDD0290 | [15] |
| LDCM0129 | JWB202 | DM93 | Y337(0.74) | LDD0291 | [15] |
| LDCM0130 | JWB211 | DM93 | Y337(1.00); Y343(0.95) | LDD0292 | [15] |
| LDCM0022 | KB02 | HCT 116 | C244(1.50) | LDD0080 | [16] |
| LDCM0023 | KB03 | HCT 116 | C244(1.82) | LDD0081 | [16] |
| LDCM0024 | KB05 | HCT 116 | C244(2.37) | LDD0082 | [16] |
| LDCM0109 | NEM | HeLa | H519(0.00); H514(0.00); H306(0.00); H345(0.00) | LDD0223 | [26] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C294(1.39) | LDD2206 | [40] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C294(1.03) | LDD2207 | [40] |
| LDCM0021 | THZ1 | HeLa S3 | C294(1.03) | LDD0460 | [13] |
| LDCM0110 | W12 | Hep-G2 | K427(0.56); Q428(0.56) | LDD0237 | [10] |
| LDCM0111 | W14 | Hep-G2 | Q536(0.61) | LDD0238 | [10] |
| LDCM0112 | W16 | Hep-G2 | R262(0.89) | LDD0239 | [10] |
| LDCM0113 | W17 | Hep-G2 | D197(0.56); S198(0.56); R262(0.57); K261(0.59) | LDD0240 | [10] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Other
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Artenimol | . | DB11638 | |||
Investigative
References
