Details of the Target
General Information of Target
| Target ID | LDTP13486 | |||||
|---|---|---|---|---|---|---|
| Target Name | Long-chain-fatty-acid--CoA ligase 6 (ACSL6) | |||||
| Gene Name | ACSL6 | |||||
| Gene ID | 23305 | |||||
| Synonyms |
ACS2; FACL6; KIAA0837; LACS5; Long-chain-fatty-acid--CoA ligase 6; EC 6.2.1.3; Arachidonate--CoA ligase; EC 6.2.1.15; Long-chain acyl-CoA synthetase 6; LACS 6 |
|||||
| 3D Structure | ||||||
| Sequence |
MEDIQTNAELKSTQEQSVPAESAAVLNDYSLTKSHEMENVDSGEGPANEDEDIGDDSMKV
KDEYSERDENVLKSEPMGNAEEPEIPYSYSREYNEYENIKLERHVVSFDSSRPTSGKMNC DVCGLSCISFNVLMVHKRSHTGERPFQCNQCGASFTQKGNLLRHIKLHTGEKPFKCHLCN YACQRRDALTGHLRTHSVEKPYKCEFCGRSYKQRSSLEEHKERCRTFLQSTDPGDTASAE ARHIKAEMGSERALVLDRLASNVAKRKSSMPQKFIGEKRHCFDVNYNSSYMYEKESELIQ TRMMDQAINNAISYLGAEALRPLVQTPPAPTSEMVPVISSMYPIALTRAEMSNGAPQELE KKSIHLPEKSVPSERGLSPNNSGHDSTDTDSNHEERQNHIYQQNHMVLSRARNGMPLLKE VPRSYELLKPPPICPRDSVKVINKEGEVMDVYRCDHCRVLFLDYVMFTIHMGCHGFRDPF ECNMCGYRSHDRYEFSSHIARGEHRALLK |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
ATP-dependent AMP-binding enzyme family
|
|||||
| Subcellular location |
Mitochondrion outer membrane
|
|||||
| Function |
Catalyzes the conversion of long-chain fatty acids to their active form acyl-CoA for both synthesis of cellular lipids, and degradation via beta-oxidation. Plays an important role in fatty acid metabolism in brain and the acyl-CoAs produced may be utilized exclusively for the synthesis of the brain lipid.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
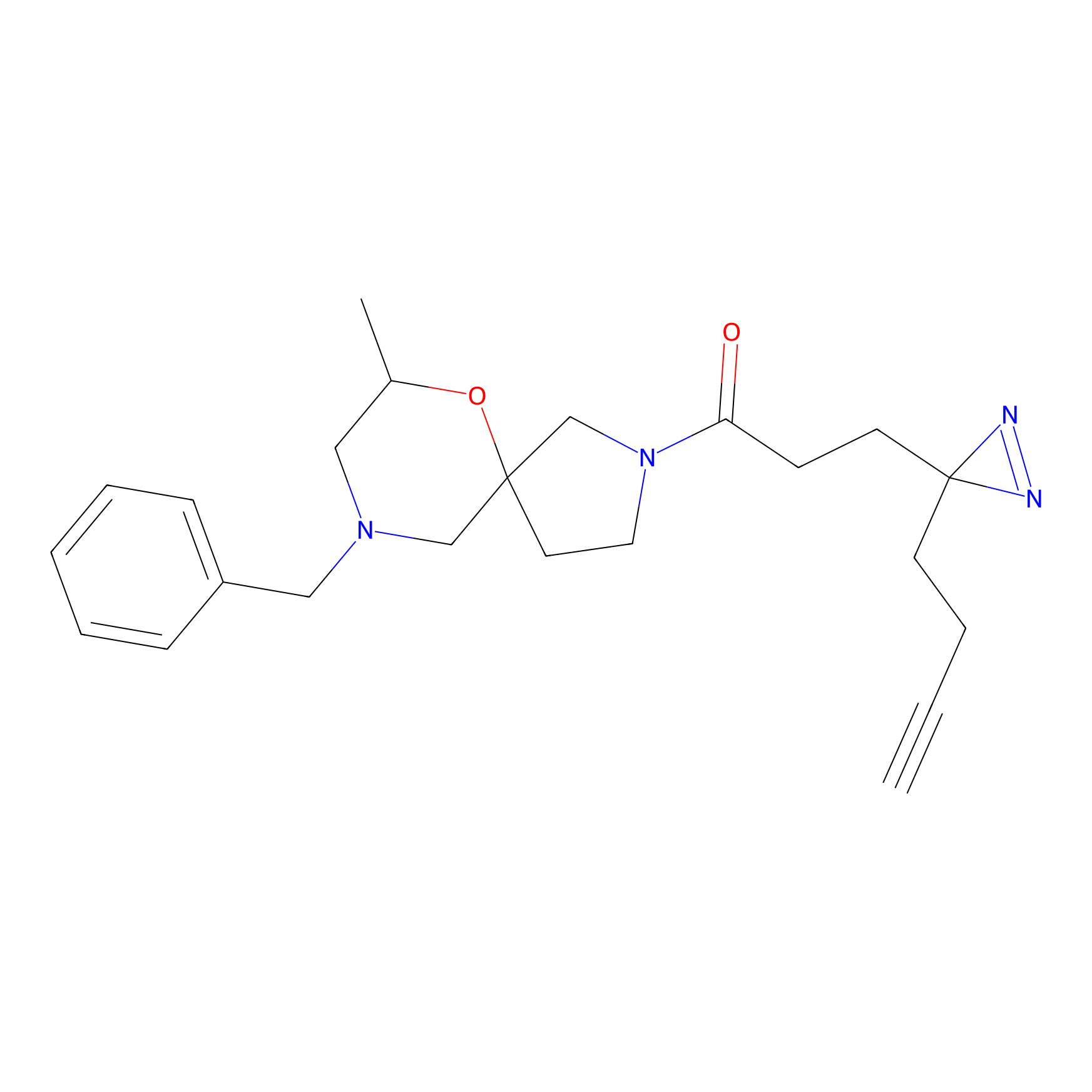
|
DBIA Probe Info |
 |
C167(1.38); C650(1.40); C452(2.12) | LDD3430 | [1] | |
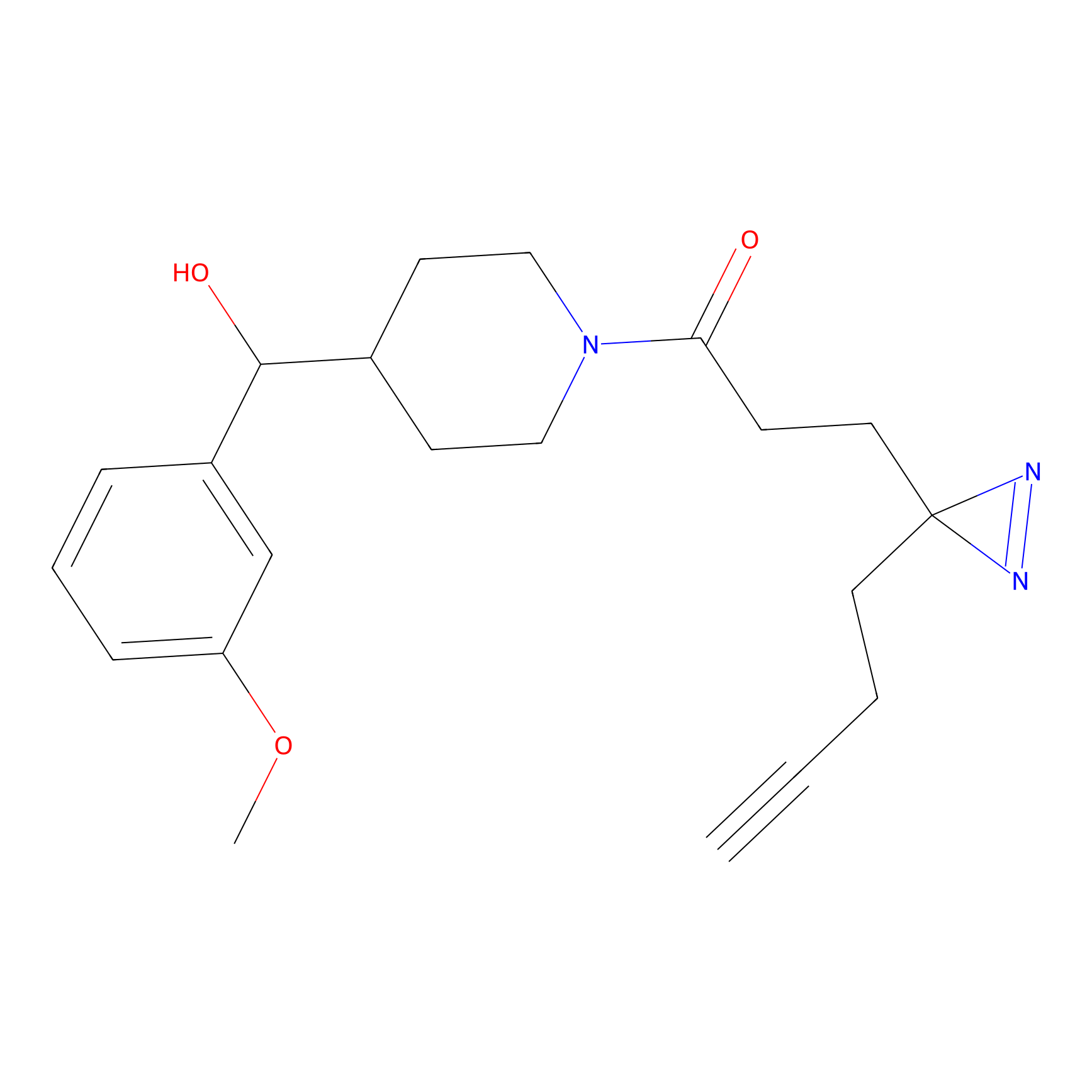
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [2] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0371 | CL102 | HEK-293T | C502(0.87) | LDD1575 | [5] |
| LDCM0375 | CL106 | HEK-293T | C502(1.00) | LDD1579 | [5] |
| LDCM0380 | CL110 | HEK-293T | C502(0.84) | LDD1584 | [5] |
| LDCM0384 | CL114 | HEK-293T | C502(0.83) | LDD1588 | [5] |
| LDCM0388 | CL118 | HEK-293T | C502(0.94) | LDD1592 | [5] |
| LDCM0393 | CL122 | HEK-293T | C502(0.92) | LDD1597 | [5] |
| LDCM0397 | CL126 | HEK-293T | C502(0.97) | LDD1601 | [5] |
| LDCM0401 | CL14 | HEK-293T | C502(1.00) | LDD1605 | [5] |
| LDCM0407 | CL2 | HEK-293T | C502(1.05) | LDD1611 | [5] |
| LDCM0414 | CL26 | HEK-293T | C502(0.94) | LDD1618 | [5] |
| LDCM0441 | CL50 | HEK-293T | C502(0.89) | LDD1645 | [5] |
| LDCM0454 | CL62 | HEK-293T | C502(0.92) | LDD1657 | [5] |
| LDCM0467 | CL74 | HEK-293T | C502(1.00) | LDD1670 | [5] |
| LDCM0480 | CL86 | HEK-293T | C502(0.84) | LDD1683 | [5] |
| LDCM0493 | CL98 | HEK-293T | C502(0.87) | LDD1696 | [5] |
| LDCM0427 | Fragment51 | HEK-293T | C502(0.92) | LDD1631 | [5] |
| LDCM0022 | KB02 | A101D | C167(1.14); C384(1.23); C80(1.20); C534(1.30) | LDD2250 | [1] |
| LDCM0023 | KB03 | A101D | C167(1.38); C384(3.20); C80(1.67); C534(2.05) | LDD2667 | [1] |
| LDCM0024 | KB05 | SK-MEL-5 | C167(1.38); C650(1.40); C452(2.12) | LDD3430 | [1] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Protein-L-isoaspartate O-methyltransferase domain-containing protein 2 (PCMTD2) | L-isoaspartyl/D-aspartyl protein methyltransferase family | Q9NV79 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Zinc finger protein 655 (ZNF655) | Krueppel C2H2-type zinc-finger protein family | Q8N720 | |||
References