Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
HDSF-alk Probe Info |
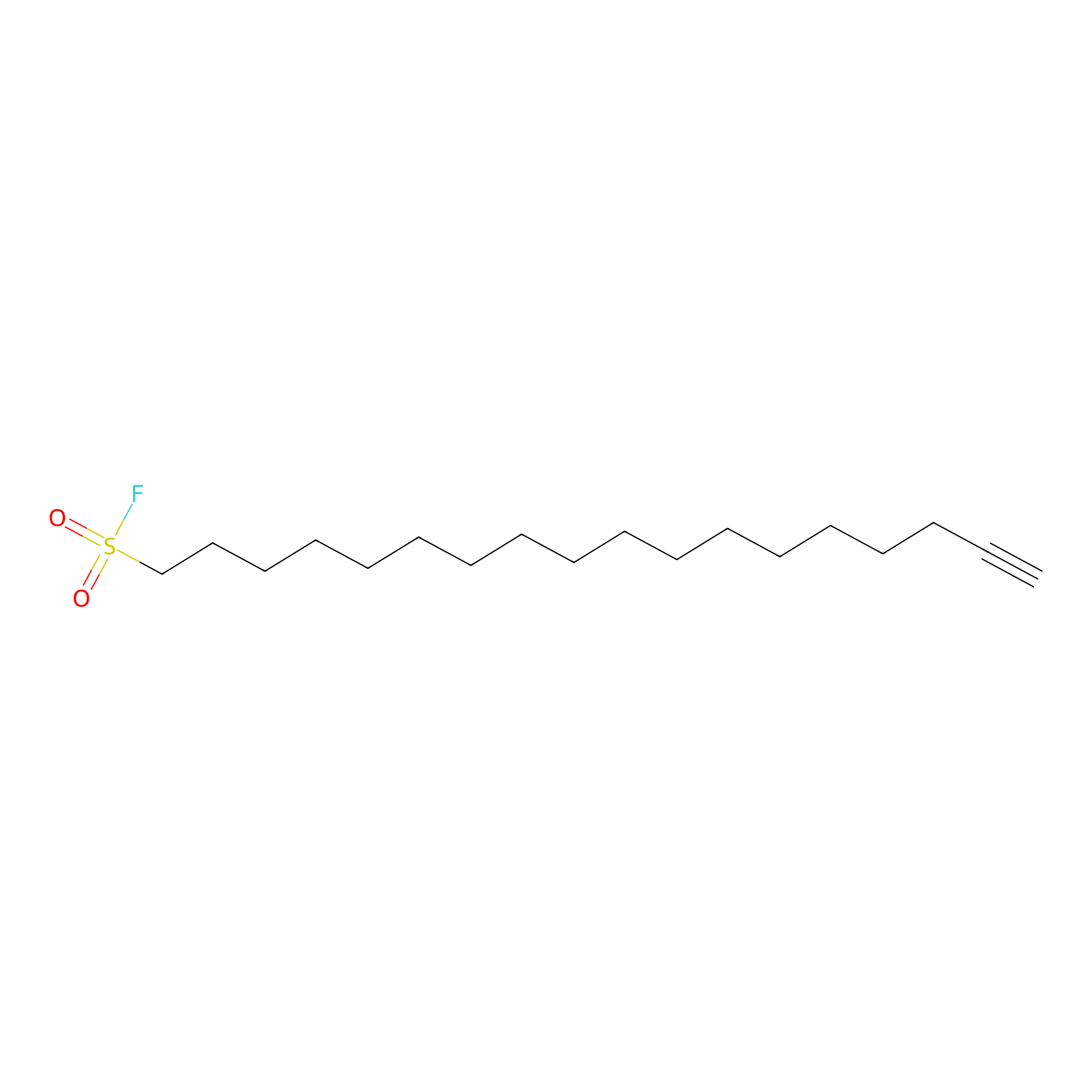 |
1.92 | LDD0197 | [2] | |
|
P8 Probe Info |
 |
10.00 | LDD0451 | [3] | |
|
AZ-9 Probe Info |
 |
2.32 | LDD0393 | [4] | |
|
FBPP2 Probe Info |
 |
3.11 | LDD0318 | [5] | |
|
CHEMBL5175495 Probe Info |
 |
13.39 | LDD0196 | [6] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [7] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [7] | |
|
TH211 Probe Info |
 |
Y77(5.33) | LDD0260 | [8] | |
|
C-Sul Probe Info |
 |
3.59 | LDD0066 | [9] | |
|
FBP2 Probe Info |
 |
3.45 | LDD0317 | [5] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [10] | |
|
ONAyne Probe Info |
 |
K216(0.00); K89(0.00); K250(0.00); K296(0.00) | LDD0273 | [11] | |
|
OPA-S-S-alkyne Probe Info |
 |
K200(2.34); K218(2.79); K216(2.79); K262(3.03) | LDD3494 | [12] | |
|
Probe 1 Probe Info |
 |
Y248(29.89) | LDD3495 | [13] | |
|
Jackson_14 Probe Info |
 |
2.13 | LDD0123 | [14] | |
|
ATP probe Probe Info |
 |
K89(0.00); K262(0.00) | LDD0199 | [15] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [16] | |
|
NHS Probe Info |
 |
K262(0.00); K200(0.00); K6(0.00); K244(0.00) | LDD0010 | [17] | |
|
SF Probe Info |
 |
K216(0.00); K296(0.00); K89(0.00) | LDD0028 | [18] | |
|
STPyne Probe Info |
 |
K262(0.00); K200(0.00) | LDD0009 | [17] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [19] | |
|
1c-yne Probe Info |
 |
K200(0.00); K147(0.00); K24(0.00); K262(0.00) | LDD0228 | [20] | |
|
Acrolein Probe Info |
 |
H69(0.00); H47(0.00) | LDD0217 | [21] | |
|
Crotonaldehyde Probe Info |
 |
H47(0.00); H69(0.00) | LDD0219 | [21] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DR-1 Probe Info |
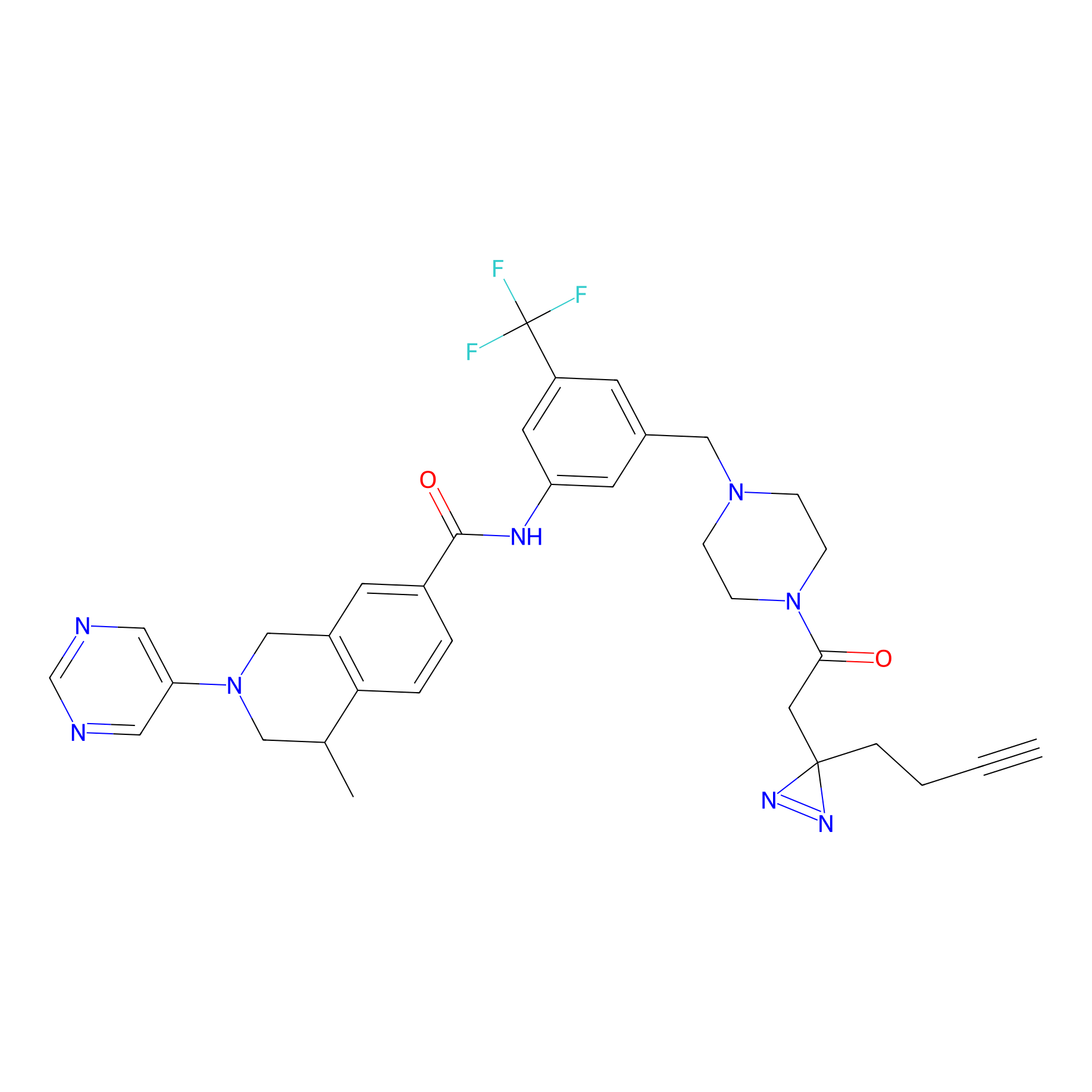 |
7.81 | LDD0398 | [22] | |
|
ILS-1 PP Probe Info |
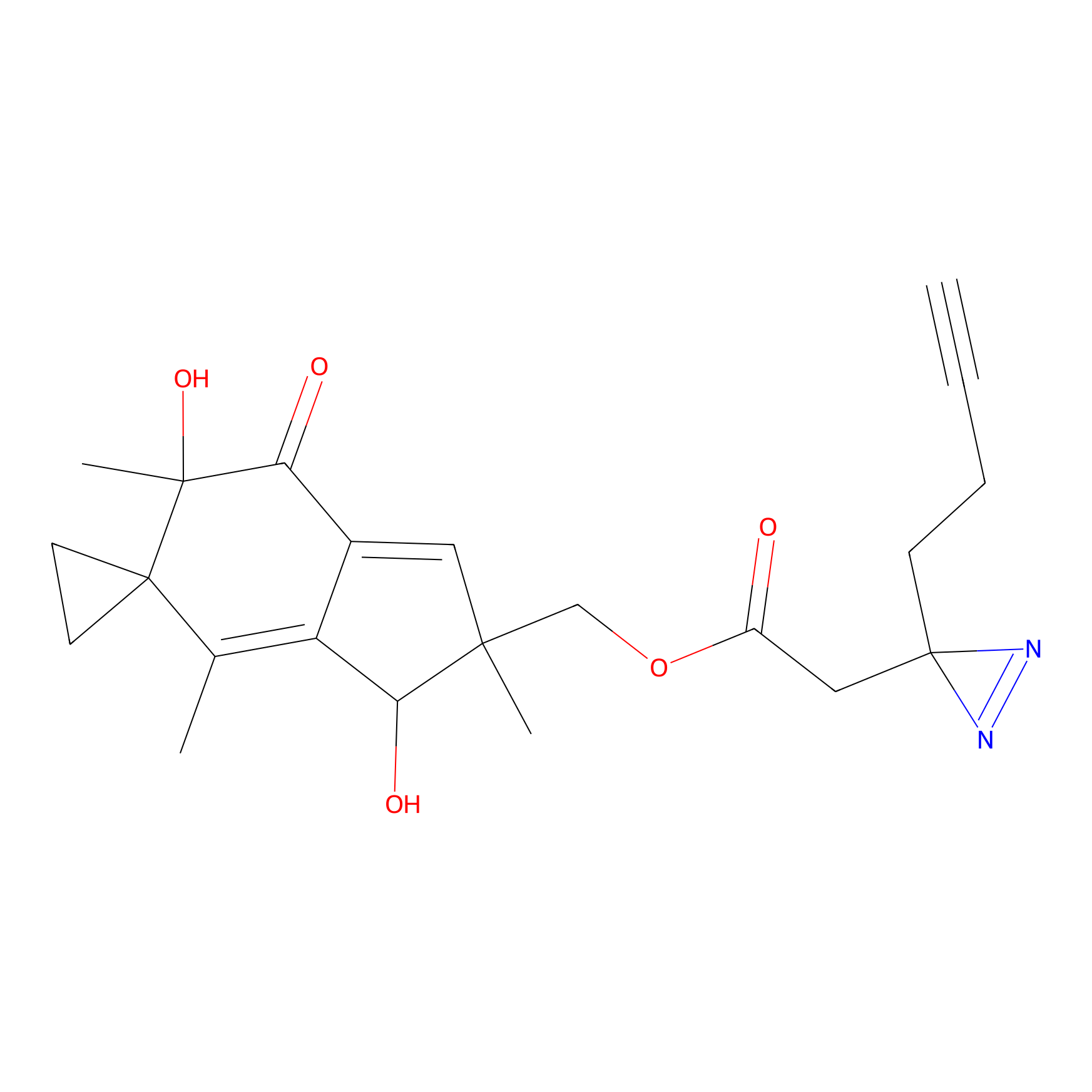 |
2.68 | LDD0417 | [23] | |
|
C003 Probe Info |
 |
11.96 | LDD1713 | [24] | |
|
C017 Probe Info |
 |
6.23 | LDD1725 | [24] | |
|
C040 Probe Info |
 |
15.78 | LDD1740 | [24] | |
|
C055 Probe Info |
 |
15.03 | LDD1752 | [24] | |
|
C056 Probe Info |
 |
19.70 | LDD1753 | [24] | |
|
C063 Probe Info |
 |
12.55 | LDD1760 | [24] | |
|
C094 Probe Info |
 |
33.36 | LDD1785 | [24] | |
|
C112 Probe Info |
 |
39.12 | LDD1799 | [24] | |
|
C143 Probe Info |
 |
12.21 | LDD1825 | [24] | |
|
C161 Probe Info |
 |
11.39 | LDD1841 | [24] | |
|
C169 Probe Info |
 |
20.97 | LDD1849 | [24] | |
|
C201 Probe Info |
 |
35.51 | LDD1877 | [24] | |
|
C210 Probe Info |
 |
58.49 | LDD1884 | [24] | |
|
C228 Probe Info |
 |
17.15 | LDD1901 | [24] | |
|
C231 Probe Info |
 |
22.47 | LDD1904 | [24] | |
|
C235 Probe Info |
 |
22.63 | LDD1908 | [24] | |
|
C238 Probe Info |
 |
19.43 | LDD1911 | [24] | |
|
C246 Probe Info |
 |
13.00 | LDD1919 | [24] | |
|
C278 Probe Info |
 |
49.52 | LDD1948 | [24] | |
|
C285 Probe Info |
 |
20.11 | LDD1955 | [24] | |
|
C289 Probe Info |
 |
32.45 | LDD1959 | [24] | |
|
C349 Probe Info |
 |
12.47 | LDD2010 | [24] | |
|
C350 Probe Info |
 |
30.48 | LDD2011 | [24] | |
|
C355 Probe Info |
 |
23.92 | LDD2016 | [24] | |
|
C356 Probe Info |
 |
7.89 | LDD2017 | [24] | |
|
C362 Probe Info |
 |
51.27 | LDD2023 | [24] | |
|
C388 Probe Info |
 |
69.55 | LDD2047 | [24] | |
|
C390 Probe Info |
 |
41.93 | LDD2049 | [24] | |
|
C407 Probe Info |
 |
11.24 | LDD2064 | [24] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [25] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [25] | |
|
FFF probe14 Probe Info |
 |
16.43 | LDD0477 | [25] | |
|
FFF probe2 Probe Info |
 |
20.00 | LDD0463 | [25] | |
|
FFF probe3 Probe Info |
 |
19.73 | LDD0464 | [25] | |
|
FFF probe4 Probe Info |
 |
14.34 | LDD0466 | [25] | |
|
FFF probe6 Probe Info |
 |
8.25 | LDD0467 | [25] | |
|
FFF probe9 Probe Info |
 |
5.92 | LDD0470 | [25] | |
|
JN0003 Probe Info |
 |
20.00 | LDD0469 | [25] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [26] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [26] | |
|
TM-F Probe Info |
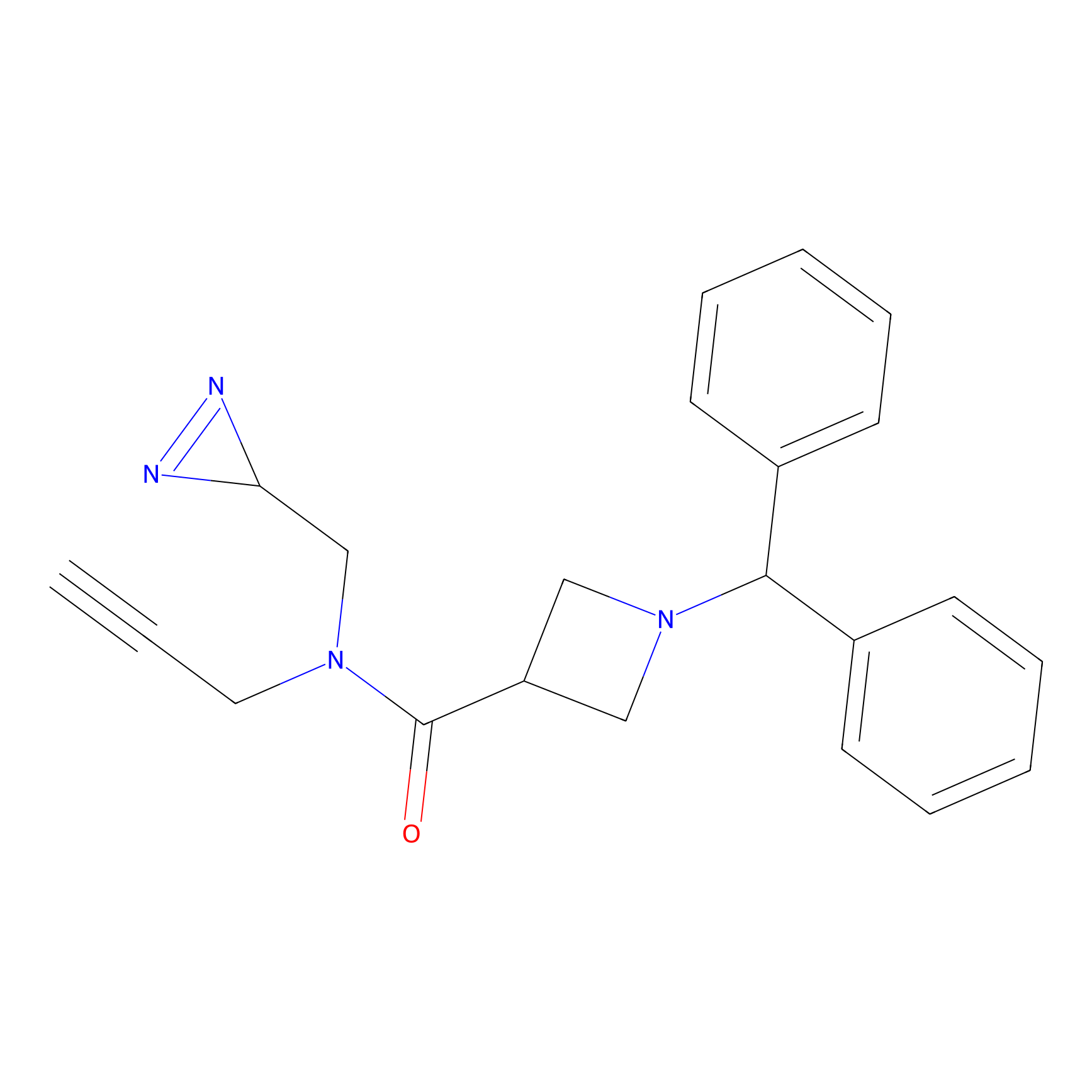 |
A65(0.00); G67(0.00); H69(0.00) | LDD0020 | [27] | |
|
Photoindomethacin Probe Info |
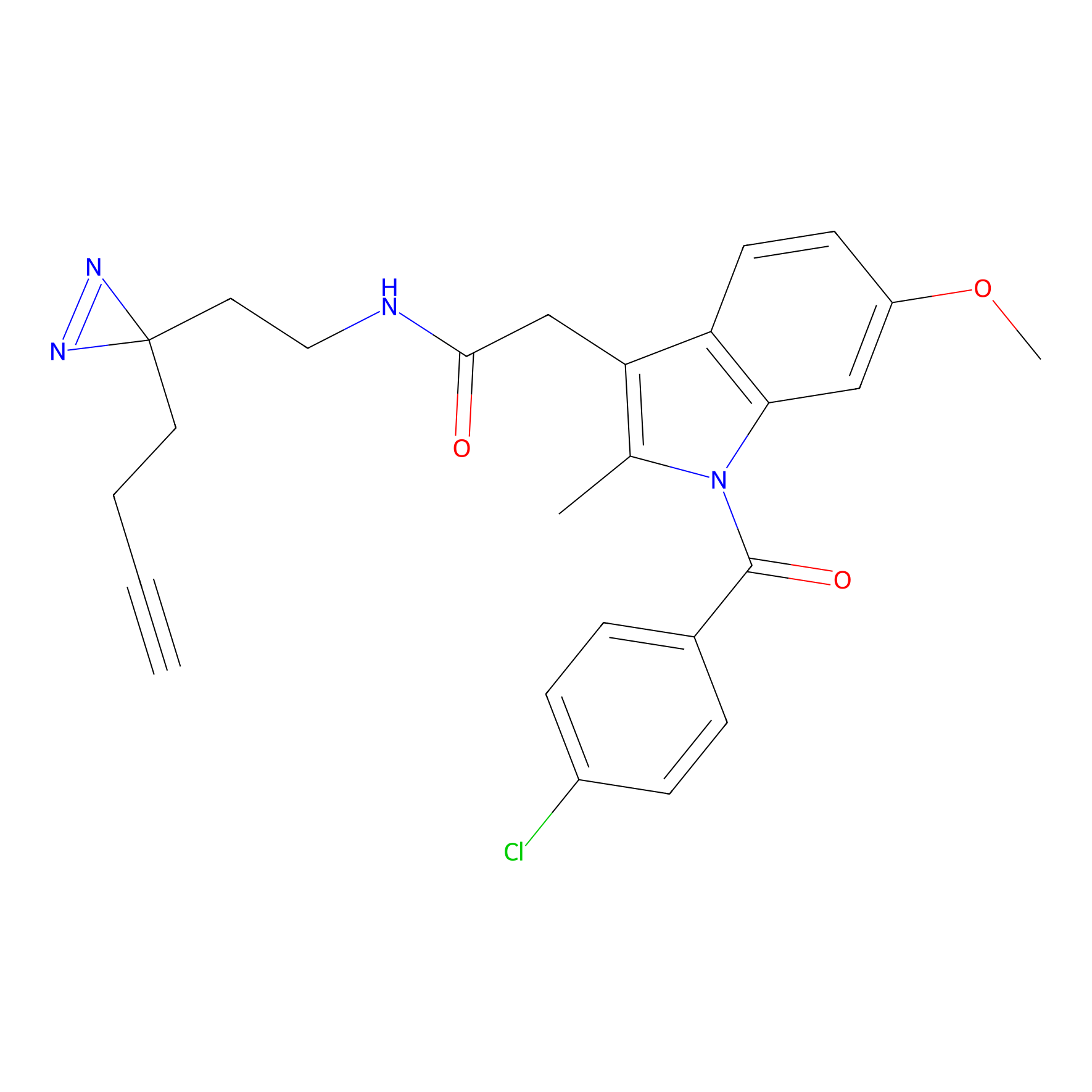 |
N.A. | LDD0154 | [28] | |
|
OEA-DA Probe Info |
 |
6.48 | LDD0046 | [29] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Dual specificity protein phosphatase 6 (DUSP6) | Protein-tyrosine phosphatase family | Q16828 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Prohibitin 1 (PHB1) | Prohibitin family | P35232 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Estrogen receptor (ESR1) | Nuclear hormone receptor family | P03372 | |||
References
