Details of the Target
General Information of Target
| Target ID | LDTP09515 | |||||
|---|---|---|---|---|---|---|
| Target Name | Importin-4 (IPO4) | |||||
| Gene Name | IPO4 | |||||
| Gene ID | 79711 | |||||
| Synonyms |
IMP4B; RANBP4; Importin-4; Imp4; Importin-4b; Imp4b; Ran-binding protein 4; RanBP4 |
|||||
| 3D Structure | ||||||
| Sequence |
MESAGLEQLLRELLLPDTERIRRATEQLQIVLRAPAALPALCDLLASAADPQIRQFAAVL
TRRRLNTRWRRLAAEQRESLKSLILTALQRETEHCVSLSLAQLSATIFRKEGLEAWPQLL QLLQHSTHSPHSPEREMGLLLLSVVVTSRPEAFQPHHRELLRLLNETLGEVGSPGLLFYS LRTLTTMAPYLSTEDVPLARMLVPKLIMAMQTLIPIDEAKACEALEALDELLESEVPVIT PYLSEVLTFCLEVARNVALGNAIRIRILCCLTFLVKVKSKALLKNRLLPPLLHTLFPIVA AEPPPGQLDPEDQDSEEEELEIELMGETPKHFAVQVVDMLALHLPPEKLCPQLMPMLEEA LRSESPYQRKAGLLVLAVLSDGAGDHIRQRLLPPLLQIVCKGLEDPSQVVRNAALFALGQ FSENLQPHISSYSREVMPLLLAYLKSVPLGHTHHLAKACYALENFVENLGPKVQPYLPEL MECMLQLLRNPSSPRAKELAVSALGAIATAAQASLLPYFPAIMEHLREFLLTGREDLQPV QIQSLETLGVLARAVGEPMRPLAEECCQLGLGLCDQVDDPDLRRCTYSLFAALSGLMGEG LAPHLEQITTLMLLSLRSTEGIVPQYDGSSSFLLFDDESDGEEEEELMDEDVEEEDDSEI SGYSVENAFFDEKEDTCAAVGEISVNTSVAFLPYMESVFEEVFKLLECPHLNVRKAAHEA LGQFCCALHKACQSCPSEPNTAALQAALARVVPSYMQAVNRERERQVVMAVLEALTGVLR SCGTLTLKPPGRLAELCGVLKAVLQRKTACQDTDEEEEEEDDDQAEYDAMLLEHAGEAIP ALAAAAGGDSFAPFFAGFLPLLVCKTKQGCTVAEKSFAVGTLAETIQGLGAASAQFVSRL LPVLLSTAQEADPEVRSNAIFGMGVLAEHGGHPAQEHFPKLLGLLFPLLARERHDRVRDN ICGALARLLMASPTRKPEPQVLAALLHALPLKEDLEEWVTIGRLFSFLYQSSPDQVIDVA PELLRICSLILADNKIPPDTKAALLLLLTFLAKQHTDSFQAALGSLPVDKAQELQAVLGL S |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Importin beta family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Nuclear transport receptor that mediates nuclear import of proteins, such as histones, RPS3A, TNP2 and VDR. Serves as receptor for nuclear localization signals (NLS) in cargo substrates. Is thought to mediate docking of the importin/substrate complex to the nuclear pore complex (NPC) through binding to nucleoporin and the complex is subsequently translocated through the pore by an energy requiring, Ran-dependent mechanism. At the nucleoplasmic side of the NPC, Ran binds to the importin, the importin/substrate complex dissociates and importin is re-exported from the nucleus to the cytoplasm where GTP hydrolysis releases Ran. The directionality of nuclear import is thought to be conferred by an asymmetric distribution of the GTP- and GDP-bound forms of Ran between the cytoplasm and nucleus. Mediates the nuclear import of the histone H3-H4 dimer when in complex with ASF1 (ASF1A or ASF1B). Mediates the ligand-independent nuclear import of vitamin D receptor (VDR). In vitro, mediates the nuclear import of human cytomegalovirus UL84 by recognizing a non-classical NLS.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| AN3CA | SNV: p.S492R | DBIA Probe Info | |||
| CAL78 | SNV: p.A500S | DBIA Probe Info | |||
| COLO792 | SNV: p.G848E | DBIA Probe Info | |||
| HCT116 | Deletion: p.P790LfsTer116 | . | |||
| HDMYZ | SNV: p.C95Y | DBIA Probe Info | |||
| IGR1 | SNV: p.A34P | DBIA Probe Info | |||
| IGROV1 | Deletion: p.E136DfsTer30 | DBIA Probe Info | |||
| MCC13 | SNV: p.H1055Y | DBIA Probe Info | |||
| MCC26 | SNV: p.S493C | DBIA Probe Info | |||
| MDAMB468 | SNV: p.H710Q | DBIA Probe Info | |||
| MOLT4 | SNV: p.E91K; p.A115V; p.E682G | IA-alkyne Probe Info | |||
| OCIAML2 | SNV: p.V31F | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
A-EBA Probe Info |
 |
3.73 | LDD0215 | [2] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [3] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [4] | |
|
STPyne Probe Info |
 |
K788(10.00) | LDD0277 | [5] | |
|
Probe 1 Probe Info |
 |
Y367(6.25); Y755(14.39) | LDD3495 | [6] | |
|
JZ128-DTB Probe Info |
 |
C725(0.00); C42(0.00); C726(0.00); C732(0.00) | LDD0462 | [7] | |
|
DA-P3 Probe Info |
 |
10.11 | LDD0183 | [8] | |
|
AHL-Pu-1 Probe Info |
 |
C735(3.45); C725(3.14) | LDD0168 | [9] | |
|
Alkyne-RA190 Probe Info |
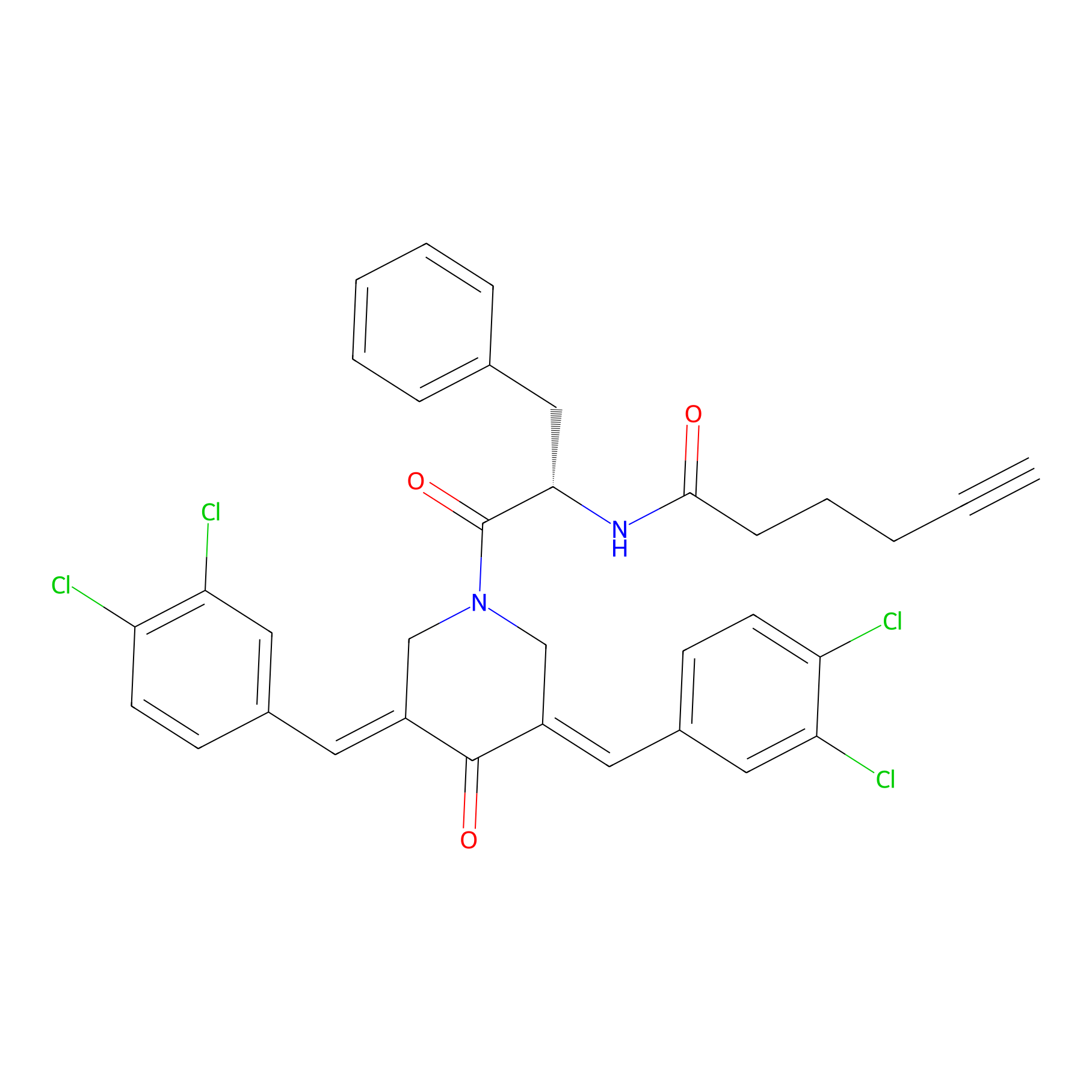 |
2.63 | LDD0300 | [10] | |
|
EA-probe Probe Info |
 |
C350(2.23); C42(1.10) | LDD2210 | [11] | |
|
DBIA Probe Info |
 |
C797(14.05); C708(17.13); C782(5.61); C459(4.74) | LDD0209 | [12] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [13] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C459(0.00); C42(0.00); C95(0.00); C797(0.00) | LDD0038 | [14] | |
|
IA-alkyne Probe Info |
 |
C962(0.00); C797(0.00) | LDD0032 | [15] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [16] | |
|
Lodoacetamide azide Probe Info |
 |
C459(0.00); C42(0.00); C962(0.00); C797(0.00) | LDD0037 | [14] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [17] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [18] | |
|
NAIA_4 Probe Info |
 |
C42(0.00); C400(0.00); C459(0.00) | LDD2226 | [19] | |
|
TFBX Probe Info |
 |
C797(0.00); C732(0.00); C962(0.00); C708(0.00) | LDD0027 | [18] | |
|
WYneN Probe Info |
 |
C782(0.00); C708(0.00); C459(0.00); C400(0.00) | LDD0021 | [17] | |
|
WYneO Probe Info |
 |
C708(0.00); C797(0.00) | LDD0022 | [17] | |
|
aHNE Probe Info |
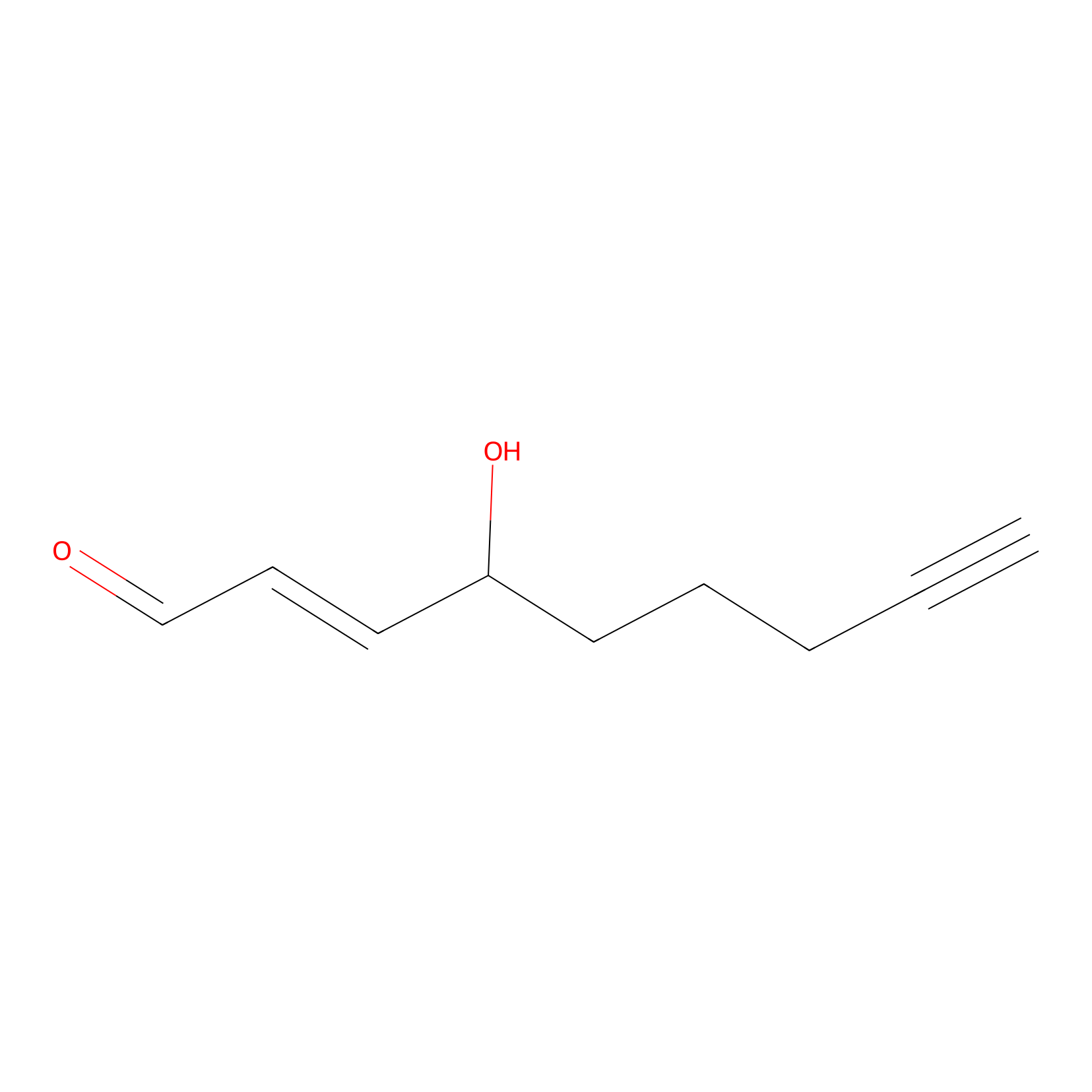 |
N.A. | LDD0001 | [17] | |
|
Compound 10 Probe Info |
 |
C459(0.00); C782(0.00); C797(0.00); C870(0.00) | LDD2216 | [20] | |
|
ENE Probe Info |
 |
C962(0.00); C708(0.00); C797(0.00) | LDD0006 | [17] | |
|
IPM Probe Info |
 |
C735(0.00); C400(0.00); C870(0.00); C726(0.00) | LDD0005 | [17] | |
|
VSF Probe Info |
 |
C400(0.00); C962(0.00); C735(0.00); C797(0.00) | LDD0007 | [17] | |
|
Phosphinate-6 Probe Info |
 |
C870(0.00); C708(0.00); C725(0.00) | LDD0018 | [21] | |
|
1c-yne Probe Info |
 |
K370(0.00); K875(0.00) | LDD0228 | [22] | |
|
Acrolein Probe Info |
 |
C962(0.00); C459(0.00); C797(0.00); C708(0.00) | LDD0217 | [23] | |
|
Methacrolein Probe Info |
 |
C708(0.00); C962(0.00); C797(0.00); C735(0.00) | LDD0218 | [23] | |
|
W1 Probe Info |
 |
C400(0.00); C962(0.00); C797(0.00); C708(0.00) | LDD0236 | [24] | |
|
AOyne Probe Info |
 |
5.20 | LDD0443 | [25] | |
|
NAIA_5 Probe Info |
 |
C726(0.00); C725(0.00); C459(0.00); C42(0.00) | LDD2223 | [19] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C040 Probe Info |
 |
8.00 | LDD1740 | [26] | |
|
C055 Probe Info |
 |
14.03 | LDD1752 | [26] | |
|
C094 Probe Info |
 |
22.78 | LDD1785 | [26] | |
|
C100 Probe Info |
 |
5.28 | LDD1789 | [26] | |
|
C112 Probe Info |
 |
17.63 | LDD1799 | [26] | |
|
C141 Probe Info |
 |
11.63 | LDD1823 | [26] | |
|
C143 Probe Info |
 |
18.77 | LDD1825 | [26] | |
|
C158 Probe Info |
 |
23.59 | LDD1838 | [26] | |
|
C163 Probe Info |
 |
5.90 | LDD1843 | [26] | |
|
C166 Probe Info |
 |
6.73 | LDD1846 | [26] | |
|
C187 Probe Info |
 |
15.56 | LDD1865 | [26] | |
|
C201 Probe Info |
 |
35.02 | LDD1877 | [26] | |
|
C270 Probe Info |
 |
7.57 | LDD1940 | [26] | |
|
C277 Probe Info |
 |
11.79 | LDD1947 | [26] | |
|
C278 Probe Info |
 |
47.50 | LDD1948 | [26] | |
|
C296 Probe Info |
 |
23.43 | LDD1966 | [26] | |
|
C343 Probe Info |
 |
13.00 | LDD2005 | [26] | |
|
C350 Probe Info |
 |
22.78 | LDD2011 | [26] | |
|
C376 Probe Info |
 |
6.82 | LDD2036 | [26] | |
|
C403 Probe Info |
 |
23.92 | LDD2061 | [26] | |
|
C407 Probe Info |
 |
12.47 | LDD2064 | [26] | |
|
FFF probe11 Probe Info |
 |
5.53 | LDD0471 | [27] | |
|
FFF probe13 Probe Info |
 |
10.18 | LDD0475 | [27] | |
|
FFF probe14 Probe Info |
 |
15.87 | LDD0477 | [27] | |
|
FFF probe2 Probe Info |
 |
6.18 | LDD0463 | [27] | |
|
FFF probe3 Probe Info |
 |
5.89 | LDD0464 | [27] | |
|
FFF probe4 Probe Info |
 |
8.78 | LDD0466 | [27] | |
|
JN0003 Probe Info |
 |
5.78 | LDD0469 | [27] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [28] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [28] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [29] | |
|
Staurosporine capture compound Probe Info |
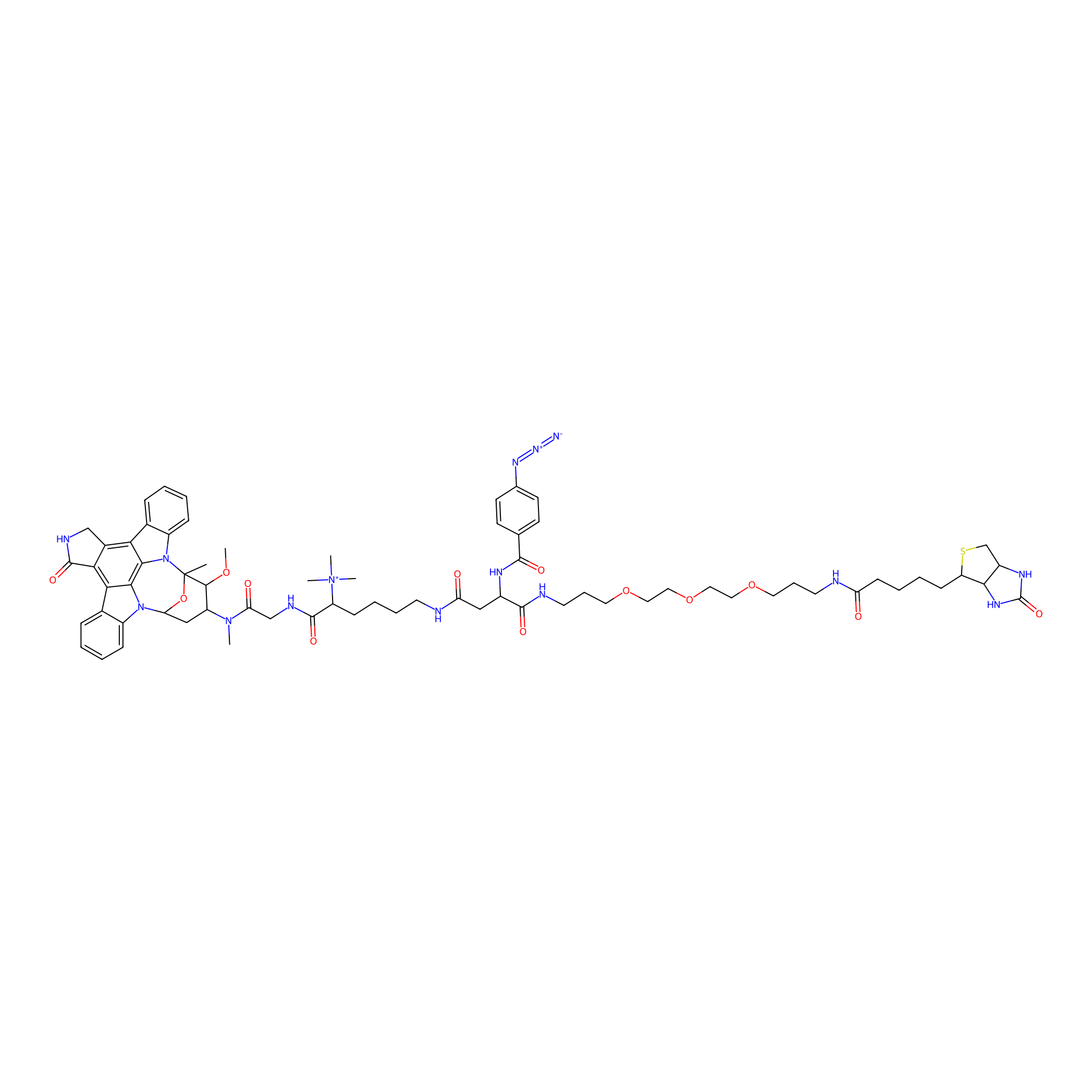 |
N.A. | LDD0083 | [30] | |
|
OEA-DA Probe Info |
 |
18.55 | LDD0046 | [31] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C962(0.96); C708(0.84); C797(0.47) | LDD2142 | [32] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C782(1.01); C797(0.88) | LDD2112 | [32] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C962(1.16); C782(0.82); C708(0.60) | LDD2095 | [32] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C782(0.79); C708(0.92) | LDD2130 | [32] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C782(0.85); C708(0.99); C797(1.37); C725(0.99) | LDD2117 | [32] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C782(1.32); C708(1.01); C797(1.68); C870(1.85) | LDD2152 | [32] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C708(1.17) | LDD2103 | [32] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C782(0.66); C708(0.74) | LDD2132 | [32] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C782(0.56); C708(0.82) | LDD2131 | [32] |
| LDCM0025 | 4SU-RNA | HEK-293T | C735(3.45); C725(3.14) | LDD0168 | [9] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C735(3.51); C725(2.17); C708(2.08); C350(2.89) | LDD0169 | [9] |
| LDCM0561 | Abegg_cp(-)-10 | HeLa | C350(2.32) | LDD0312 | [18] |
| LDCM0214 | AC1 | HEK-293T | C870(1.05); C962(0.98); C782(1.05); C708(0.93) | LDD1507 | [33] |
| LDCM0215 | AC10 | HEK-293T | C870(1.01); C962(1.00); C782(0.97); C708(0.92) | LDD1508 | [33] |
| LDCM0226 | AC11 | HEK-293T | C870(0.98); C962(0.90); C782(1.01); C708(0.89) | LDD1509 | [33] |
| LDCM0237 | AC12 | HEK-293T | C870(0.98); C962(0.88); C782(1.00); C708(0.91) | LDD1510 | [33] |
| LDCM0259 | AC14 | HEK-293T | C870(1.00); C962(0.94); C782(1.00); C708(0.91) | LDD1512 | [33] |
| LDCM0270 | AC15 | HEK-293T | C870(1.01); C962(1.00); C782(1.01); C708(0.95) | LDD1513 | [33] |
| LDCM0276 | AC17 | HEK-293T | C870(1.00); C962(0.92); C782(0.99); C708(0.97) | LDD1515 | [33] |
| LDCM0277 | AC18 | HEK-293T | C870(0.97); C962(1.07); C782(1.01); C708(0.99) | LDD1516 | [33] |
| LDCM0278 | AC19 | HEK-293T | C870(1.46); C962(1.05); C782(1.29); C708(1.04) | LDD1517 | [33] |
| LDCM0279 | AC2 | HEK-293T | C870(1.00); C962(0.98); C782(1.01); C708(0.94) | LDD1518 | [33] |
| LDCM0280 | AC20 | HEK-293T | C870(1.02); C962(0.94); C782(0.99); C708(0.93) | LDD1519 | [33] |
| LDCM0281 | AC21 | HEK-293T | C870(1.04); C962(0.99); C782(1.00); C708(1.05) | LDD1520 | [33] |
| LDCM0282 | AC22 | HEK-293T | C870(1.00); C962(0.94); C782(0.98); C708(1.03) | LDD1521 | [33] |
| LDCM0283 | AC23 | HEK-293T | C870(1.04); C962(1.05); C782(0.97); C708(0.92) | LDD1522 | [33] |
| LDCM0284 | AC24 | HEK-293T | C870(1.01); C962(1.01); C782(0.99); C708(0.90) | LDD1523 | [33] |
| LDCM0285 | AC25 | HEK-293T | C870(1.02); C962(0.99); C782(0.97); C708(0.94) | LDD1524 | [33] |
| LDCM0286 | AC26 | HEK-293T | C870(1.01); C962(1.06); C782(1.02); C708(0.95) | LDD1525 | [33] |
| LDCM0287 | AC27 | HEK-293T | C870(1.02); C962(0.89); C782(1.00); C708(0.91) | LDD1526 | [33] |
| LDCM0288 | AC28 | HEK-293T | C870(1.05); C962(0.91); C782(0.98); C708(0.87) | LDD1527 | [33] |
| LDCM0289 | AC29 | HEK-293T | C870(1.00); C962(1.13); C782(0.99); C708(0.95) | LDD1528 | [33] |
| LDCM0290 | AC3 | HEK-293T | C870(0.99); C962(0.90); C782(1.02); C708(0.90) | LDD1529 | [33] |
| LDCM0291 | AC30 | HEK-293T | C870(1.05); C962(0.90); C782(1.00); C708(0.99) | LDD1530 | [33] |
| LDCM0292 | AC31 | HEK-293T | C870(1.01); C962(0.93); C782(0.98); C708(0.92) | LDD1531 | [33] |
| LDCM0293 | AC32 | HEK-293T | C870(0.97); C962(0.99); C782(0.97); C708(0.95) | LDD1532 | [33] |
| LDCM0294 | AC33 | HEK-293T | C870(0.98); C962(0.94); C782(0.95); C708(0.92) | LDD1533 | [33] |
| LDCM0295 | AC34 | HEK-293T | C870(0.99); C962(1.01); C782(1.01); C708(0.92) | LDD1534 | [33] |
| LDCM0296 | AC35 | HEK-293T | C870(0.93); C962(0.91); C782(1.01); C708(0.92) | LDD1535 | [33] |
| LDCM0297 | AC36 | HEK-293T | C870(0.94); C962(0.86); C782(0.99); C708(0.90) | LDD1536 | [33] |
| LDCM0298 | AC37 | HEK-293T | C870(0.97); C962(1.00); C782(0.91); C708(0.94) | LDD1537 | [33] |
| LDCM0299 | AC38 | HEK-293T | C870(0.95); C962(0.95); C782(0.97); C708(0.89) | LDD1538 | [33] |
| LDCM0300 | AC39 | HEK-293T | C870(0.97); C962(0.94); C782(0.99); C708(0.92) | LDD1539 | [33] |
| LDCM0301 | AC4 | HEK-293T | C870(0.98); C962(0.90); C782(0.98); C708(0.93) | LDD1540 | [33] |
| LDCM0302 | AC40 | HEK-293T | C870(0.96); C962(1.00); C782(0.97); C708(1.02) | LDD1541 | [33] |
| LDCM0303 | AC41 | HEK-293T | C870(0.99); C962(0.96); C782(0.94); C708(0.93) | LDD1542 | [33] |
| LDCM0304 | AC42 | HEK-293T | C870(1.09); C962(1.06); C782(1.00); C708(0.95) | LDD1543 | [33] |
| LDCM0305 | AC43 | HEK-293T | C870(0.92); C962(0.98); C782(0.95); C708(0.89) | LDD1544 | [33] |
| LDCM0306 | AC44 | HEK-293T | C870(0.93); C962(0.88); C782(0.99); C708(0.91) | LDD1545 | [33] |
| LDCM0307 | AC45 | HEK-293T | C870(1.01); C962(0.94); C782(0.94); C708(0.91) | LDD1546 | [33] |
| LDCM0308 | AC46 | HEK-293T | C870(0.95); C962(0.93); C782(1.03); C708(0.89) | LDD1547 | [33] |
| LDCM0309 | AC47 | HEK-293T | C870(0.93); C962(0.96); C782(0.99); C708(0.94) | LDD1548 | [33] |
| LDCM0310 | AC48 | HEK-293T | C870(0.94); C962(0.86); C782(0.93); C708(0.95) | LDD1549 | [33] |
| LDCM0311 | AC49 | HEK-293T | C870(1.00); C962(0.93); C782(1.02); C708(0.96) | LDD1550 | [33] |
| LDCM0312 | AC5 | HEK-293T | C870(1.02); C962(0.90); C782(0.98); C708(0.94) | LDD1551 | [33] |
| LDCM0313 | AC50 | HEK-293T | C870(0.99); C962(0.99); C782(1.01); C708(0.91) | LDD1552 | [33] |
| LDCM0314 | AC51 | HEK-293T | C870(0.93); C962(1.04); C782(1.01); C708(0.96) | LDD1553 | [33] |
| LDCM0315 | AC52 | HEK-293T | C870(0.94); C962(0.91); C782(1.00); C708(0.97) | LDD1554 | [33] |
| LDCM0316 | AC53 | HEK-293T | C870(0.97); C962(1.00); C782(0.95); C708(0.94) | LDD1555 | [33] |
| LDCM0317 | AC54 | HEK-293T | C870(0.97); C962(0.93); C782(1.02); C708(0.98) | LDD1556 | [33] |
| LDCM0318 | AC55 | HEK-293T | C870(0.97); C962(0.98); C782(0.99); C708(0.97) | LDD1557 | [33] |
| LDCM0319 | AC56 | HEK-293T | C870(0.95); C962(0.96); C782(1.00); C708(0.98) | LDD1558 | [33] |
| LDCM0320 | AC57 | HEK-293T | C870(1.04); C962(0.96); C782(1.00); C708(0.92) | LDD1559 | [33] |
| LDCM0321 | AC58 | HEK-293T | C870(1.05); C962(0.96); C782(1.01); C708(0.95) | LDD1560 | [33] |
| LDCM0322 | AC59 | HEK-293T | C870(0.98); C962(0.99); C782(0.98); C708(0.93) | LDD1561 | [33] |
| LDCM0323 | AC6 | HEK-293T | C870(1.05); C962(0.90); C782(0.99); C708(0.99) | LDD1562 | [33] |
| LDCM0324 | AC60 | HEK-293T | C870(1.00); C962(0.87); C782(0.99); C708(1.00) | LDD1563 | [33] |
| LDCM0325 | AC61 | HEK-293T | C870(1.07); C962(0.96); C782(1.01); C708(0.94) | LDD1564 | [33] |
| LDCM0326 | AC62 | HEK-293T | C870(1.04); C962(0.95); C782(0.98); C708(0.96) | LDD1565 | [33] |
| LDCM0327 | AC63 | HEK-293T | C870(1.08); C962(1.02); C782(0.99); C708(0.98) | LDD1566 | [33] |
| LDCM0328 | AC64 | HEK-293T | C870(0.95); C962(0.96); C782(0.97); C708(0.97) | LDD1567 | [33] |
| LDCM0334 | AC7 | HEK-293T | C870(0.98); C962(0.94); C782(0.99); C708(0.88) | LDD1568 | [33] |
| LDCM0345 | AC8 | HEK-293T | C870(0.99); C962(0.95); C782(1.02); C708(0.98) | LDD1569 | [33] |
| LDCM0545 | Acetamide | MDA-MB-231 | C782(0.85); C708(0.53) | LDD2138 | [32] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C962(1.19); C782(1.00); C870(0.84); C1027(1.09) | LDD2113 | [32] |
| LDCM0248 | AKOS034007472 | HEK-293T | C870(1.02); C962(1.04); C782(1.03); C708(0.91) | LDD1511 | [33] |
| LDCM0356 | AKOS034007680 | HEK-293T | C870(1.02); C962(0.92); C782(0.98); C708(0.93) | LDD1570 | [33] |
| LDCM0275 | AKOS034007705 | HEK-293T | C870(1.04); C962(1.00); C782(1.04); C708(0.98) | LDD1514 | [33] |
| LDCM0156 | Aniline | NCI-H1299 | C708(0.00); C95(0.00) | LDD0404 | [1] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C782(0.96); C726(1.56) | LDD2091 | [32] |
| LDCM0108 | Chloroacetamide | HeLa | C962(0.00); C797(0.00) | LDD0222 | [23] |
| LDCM0632 | CL-Sc | Hep-G2 | C782(5.02); C95(2.19); C459(1.54); C708(1.47) | LDD2227 | [19] |
| LDCM0367 | CL1 | HEK-293T | C870(0.98); C962(1.02); C782(1.11); C708(1.13) | LDD1571 | [33] |
| LDCM0368 | CL10 | HEK-293T | C870(1.65); C962(1.11); C782(1.17); C708(3.33) | LDD1572 | [33] |
| LDCM0369 | CL100 | HEK-293T | C870(1.07); C962(0.95); C782(1.05); C350(1.08) | LDD1573 | [33] |
| LDCM0370 | CL101 | HEK-293T | C870(0.97); C962(1.03); C782(1.01); C708(1.00) | LDD1574 | [33] |
| LDCM0371 | CL102 | HEK-293T | C870(1.14); C962(1.04); C782(0.96); C708(1.66) | LDD1575 | [33] |
| LDCM0372 | CL103 | HEK-293T | C870(1.04); C962(0.89); C782(0.99); C708(0.97) | LDD1576 | [33] |
| LDCM0373 | CL104 | HEK-293T | C870(0.99); C962(0.99); C782(1.01); C350(1.03) | LDD1577 | [33] |
| LDCM0374 | CL105 | HEK-293T | C870(0.98); C962(1.14); C782(1.01); C708(1.13) | LDD1578 | [33] |
| LDCM0375 | CL106 | HEK-293T | C870(1.01); C962(1.03); C782(1.01); C708(1.26) | LDD1579 | [33] |
| LDCM0376 | CL107 | HEK-293T | C870(0.97); C962(0.89); C782(1.00); C708(0.94) | LDD1580 | [33] |
| LDCM0377 | CL108 | HEK-293T | C870(0.95); C962(0.97); C782(0.97); C350(0.99) | LDD1581 | [33] |
| LDCM0378 | CL109 | HEK-293T | C870(1.06); C962(1.05); C782(1.07); C708(1.13) | LDD1582 | [33] |
| LDCM0379 | CL11 | HEK-293T | C870(1.06); C962(1.12); C782(1.15); C708(1.43) | LDD1583 | [33] |
| LDCM0380 | CL110 | HEK-293T | C870(1.22); C962(1.12); C782(0.99); C708(1.71) | LDD1584 | [33] |
| LDCM0381 | CL111 | HEK-293T | C870(1.10); C962(0.91); C782(0.94); C708(1.35) | LDD1585 | [33] |
| LDCM0382 | CL112 | HEK-293T | C870(1.03); C962(1.07); C782(0.95); C350(0.97) | LDD1586 | [33] |
| LDCM0383 | CL113 | HEK-293T | C870(0.91); C962(0.96); C782(0.96); C708(0.93) | LDD1587 | [33] |
| LDCM0384 | CL114 | HEK-293T | C870(1.10); C962(1.07); C782(1.01); C708(1.87) | LDD1588 | [33] |
| LDCM0385 | CL115 | HEK-293T | C870(0.94); C962(0.95); C782(0.94); C708(0.96) | LDD1589 | [33] |
| LDCM0386 | CL116 | HEK-293T | C870(0.92); C962(0.93); C782(0.96); C350(1.12) | LDD1590 | [33] |
| LDCM0387 | CL117 | HEK-293T | C870(0.94); C962(0.97); C782(0.95); C708(0.97) | LDD1591 | [33] |
| LDCM0388 | CL118 | HEK-293T | C870(0.96); C962(0.99); C782(0.94); C708(0.88) | LDD1592 | [33] |
| LDCM0389 | CL119 | HEK-293T | C870(0.95); C962(0.94); C782(0.97); C708(0.94) | LDD1593 | [33] |
| LDCM0390 | CL12 | HEK-293T | C870(1.12); C962(1.07); C782(1.16); C708(1.33) | LDD1594 | [33] |
| LDCM0391 | CL120 | HEK-293T | C870(0.99); C962(0.96); C782(1.00); C350(1.05) | LDD1595 | [33] |
| LDCM0392 | CL121 | HEK-293T | C870(0.89); C962(1.01); C782(1.01); C708(1.03) | LDD1596 | [33] |
| LDCM0393 | CL122 | HEK-293T | C870(1.00); C962(1.10); C782(0.95); C708(1.09) | LDD1597 | [33] |
| LDCM0394 | CL123 | HEK-293T | C870(1.22); C962(1.41); C782(1.07); C708(1.65) | LDD1598 | [33] |
| LDCM0395 | CL124 | HEK-293T | C870(0.95); C962(1.20); C782(1.02); C350(1.24) | LDD1599 | [33] |
| LDCM0396 | CL125 | HEK-293T | C870(1.05); C962(0.91); C782(1.00); C708(0.92) | LDD1600 | [33] |
| LDCM0397 | CL126 | HEK-293T | C870(1.07); C962(1.07); C782(0.91); C708(1.14) | LDD1601 | [33] |
| LDCM0398 | CL127 | HEK-293T | C870(1.03); C962(0.98); C782(0.94); C708(0.93) | LDD1602 | [33] |
| LDCM0399 | CL128 | HEK-293T | C870(0.94); C962(0.95); C782(1.02); C350(1.02) | LDD1603 | [33] |
| LDCM0400 | CL13 | HEK-293T | C870(0.96); C962(0.92); C782(1.13); C708(1.11) | LDD1604 | [33] |
| LDCM0401 | CL14 | HEK-293T | C870(1.00); C962(0.98); C782(0.91); C708(1.03) | LDD1605 | [33] |
| LDCM0402 | CL15 | HEK-293T | C870(1.39); C962(1.25); C782(1.20); C708(2.27) | LDD1606 | [33] |
| LDCM0403 | CL16 | HEK-293T | C870(0.98); C962(0.96); C782(1.05); C350(1.09) | LDD1607 | [33] |
| LDCM0404 | CL17 | HEK-293T | C870(1.35); C962(1.12); C782(1.02); C708(2.09) | LDD1608 | [33] |
| LDCM0405 | CL18 | HEK-293T | C870(1.05); C962(1.01); C782(1.02); C708(0.95) | LDD1609 | [33] |
| LDCM0406 | CL19 | HEK-293T | C870(1.02); C962(0.99); C782(1.11); C708(0.98) | LDD1610 | [33] |
| LDCM0407 | CL2 | HEK-293T | C870(1.08); C962(0.98); C782(0.97); C708(1.01) | LDD1611 | [33] |
| LDCM0408 | CL20 | HEK-293T | C870(0.99); C962(0.98); C782(1.07); C708(0.92) | LDD1612 | [33] |
| LDCM0409 | CL21 | HEK-293T | C870(1.36); C962(1.57); C782(1.12); C708(2.13) | LDD1613 | [33] |
| LDCM0410 | CL22 | HEK-293T | C870(0.96); C962(0.99); C782(1.09); C708(1.03) | LDD1614 | [33] |
| LDCM0411 | CL23 | HEK-293T | C870(1.05); C962(1.19); C782(1.16); C708(1.02) | LDD1615 | [33] |
| LDCM0412 | CL24 | HEK-293T | C870(1.02); C962(1.08); C782(1.08); C708(1.11) | LDD1616 | [33] |
| LDCM0413 | CL25 | HEK-293T | C870(1.08); C962(1.30); C782(1.13); C708(1.35) | LDD1617 | [33] |
| LDCM0414 | CL26 | HEK-293T | C870(0.98); C962(1.05); C782(0.96); C708(0.93) | LDD1618 | [33] |
| LDCM0415 | CL27 | HEK-293T | C870(0.97); C962(0.99); C782(1.02); C708(0.94) | LDD1619 | [33] |
| LDCM0416 | CL28 | HEK-293T | C870(1.03); C962(1.09); C782(1.08); C350(1.03) | LDD1620 | [33] |
| LDCM0417 | CL29 | HEK-293T | C870(0.99); C962(0.96); C782(1.01); C708(0.93) | LDD1621 | [33] |
| LDCM0418 | CL3 | HEK-293T | C870(1.01); C962(0.96); C782(1.03); C708(1.01) | LDD1622 | [33] |
| LDCM0419 | CL30 | HEK-293T | C870(1.02); C962(1.06); C782(1.06); C708(0.93) | LDD1623 | [33] |
| LDCM0420 | CL31 | HEK-293T | C870(1.03); C962(1.00); C782(0.99); C708(1.00) | LDD1624 | [33] |
| LDCM0421 | CL32 | HEK-293T | C870(0.98); C962(0.90); C782(1.05); C708(0.92) | LDD1625 | [33] |
| LDCM0422 | CL33 | HEK-293T | C870(1.62); C962(1.33); C782(1.11); C708(2.45) | LDD1626 | [33] |
| LDCM0423 | CL34 | HEK-293T | C870(1.01); C962(1.00); C782(1.12); C708(1.29) | LDD1627 | [33] |
| LDCM0424 | CL35 | HEK-293T | C870(0.96); C962(1.14); C782(1.08); C708(1.14) | LDD1628 | [33] |
| LDCM0425 | CL36 | HEK-293T | C870(0.99); C962(1.02); C782(1.10); C708(1.26) | LDD1629 | [33] |
| LDCM0426 | CL37 | HEK-293T | C870(1.08); C962(1.11); C782(1.05); C708(1.03) | LDD1630 | [33] |
| LDCM0428 | CL39 | HEK-293T | C870(1.00); C962(0.90); C782(0.94); C708(1.14) | LDD1632 | [33] |
| LDCM0429 | CL4 | HEK-293T | C870(1.01); C962(0.99); C782(1.09); C350(1.14) | LDD1633 | [33] |
| LDCM0430 | CL40 | HEK-293T | C870(0.99); C962(1.04); C782(1.06); C350(0.98) | LDD1634 | [33] |
| LDCM0431 | CL41 | HEK-293T | C870(1.14); C962(1.02); C782(1.01); C708(1.30) | LDD1635 | [33] |
| LDCM0432 | CL42 | HEK-293T | C870(1.02); C962(1.06); C782(1.01); C708(1.01) | LDD1636 | [33] |
| LDCM0433 | CL43 | HEK-293T | C870(0.98); C962(0.94); C782(1.02); C708(1.05) | LDD1637 | [33] |
| LDCM0434 | CL44 | HEK-293T | C870(1.08); C962(1.04); C782(1.02); C708(0.95) | LDD1638 | [33] |
| LDCM0435 | CL45 | HEK-293T | C870(1.13); C962(1.10); C782(0.97); C708(1.64) | LDD1639 | [33] |
| LDCM0436 | CL46 | HEK-293T | C870(1.03); C962(1.00); C782(1.07); C708(1.20) | LDD1640 | [33] |
| LDCM0437 | CL47 | HEK-293T | C870(0.99); C962(1.05); C782(1.08); C708(1.17) | LDD1641 | [33] |
| LDCM0438 | CL48 | HEK-293T | C870(0.98); C962(1.05); C782(1.12); C708(1.27) | LDD1642 | [33] |
| LDCM0439 | CL49 | HEK-293T | C870(0.96); C962(0.95); C782(1.02); C708(0.98) | LDD1643 | [33] |
| LDCM0440 | CL5 | HEK-293T | C870(1.03); C962(1.02); C782(0.98); C708(0.99) | LDD1644 | [33] |
| LDCM0441 | CL50 | HEK-293T | C870(1.00); C962(1.13); C782(0.97); C708(1.29) | LDD1645 | [33] |
| LDCM0443 | CL52 | HEK-293T | C870(0.95); C962(0.95); C782(1.03); C350(1.06) | LDD1646 | [33] |
| LDCM0444 | CL53 | HEK-293T | C870(1.13); C962(1.04); C782(0.99); C708(1.60) | LDD1647 | [33] |
| LDCM0445 | CL54 | HEK-293T | C870(1.21); C962(1.65); C782(1.21); C708(1.56) | LDD1648 | [33] |
| LDCM0446 | CL55 | HEK-293T | C870(0.93); C962(0.91); C782(0.97); C708(0.95) | LDD1649 | [33] |
| LDCM0447 | CL56 | HEK-293T | C870(0.92); C962(0.91); C782(0.98); C708(1.11) | LDD1650 | [33] |
| LDCM0448 | CL57 | HEK-293T | C870(1.16); C962(1.22); C782(1.05); C708(1.61) | LDD1651 | [33] |
| LDCM0449 | CL58 | HEK-293T | C870(0.98); C962(1.06); C782(1.04); C708(1.20) | LDD1652 | [33] |
| LDCM0450 | CL59 | HEK-293T | C870(0.92); C962(0.96); C782(1.08); C708(1.13) | LDD1653 | [33] |
| LDCM0451 | CL6 | HEK-293T | C870(1.25); C962(1.72); C782(1.07); C708(1.50) | LDD1654 | [33] |
| LDCM0452 | CL60 | HEK-293T | C870(0.97); C962(1.06); C782(1.05); C708(1.31) | LDD1655 | [33] |
| LDCM0453 | CL61 | HEK-293T | C870(0.91); C962(0.93); C782(0.98); C708(0.89) | LDD1656 | [33] |
| LDCM0454 | CL62 | HEK-293T | C870(0.99); C962(1.03); C782(0.90); C708(0.88) | LDD1657 | [33] |
| LDCM0455 | CL63 | HEK-293T | C870(0.94); C962(0.97); C782(0.94); C708(0.87) | LDD1658 | [33] |
| LDCM0456 | CL64 | HEK-293T | C870(1.13); C962(1.17); C782(1.07); C350(1.23) | LDD1659 | [33] |
| LDCM0457 | CL65 | HEK-293T | C870(1.00); C962(0.93); C782(1.03); C708(0.95) | LDD1660 | [33] |
| LDCM0458 | CL66 | HEK-293T | C870(0.98); C962(0.97); C782(0.98); C708(1.11) | LDD1661 | [33] |
| LDCM0459 | CL67 | HEK-293T | C870(0.97); C962(0.93); C782(0.99); C708(0.86) | LDD1662 | [33] |
| LDCM0460 | CL68 | HEK-293T | C870(1.01); C962(0.94); C782(0.99); C708(1.03) | LDD1663 | [33] |
| LDCM0461 | CL69 | HEK-293T | C870(1.01); C962(1.14); C782(0.98); C708(1.14) | LDD1664 | [33] |
| LDCM0462 | CL7 | HEK-293T | C870(1.02); C962(0.97); C782(1.05); C708(0.97) | LDD1665 | [33] |
| LDCM0463 | CL70 | HEK-293T | C870(0.99); C962(0.97); C782(1.09); C708(0.97) | LDD1666 | [33] |
| LDCM0464 | CL71 | HEK-293T | C870(0.98); C962(1.09); C782(1.04); C708(1.14) | LDD1667 | [33] |
| LDCM0465 | CL72 | HEK-293T | C870(1.01); C962(1.09); C782(1.10); C708(1.00) | LDD1668 | [33] |
| LDCM0466 | CL73 | HEK-293T | C870(0.97); C962(1.04); C782(1.03); C708(1.36) | LDD1669 | [33] |
| LDCM0467 | CL74 | HEK-293T | C870(0.99); C962(0.99); C782(0.95); C708(0.91) | LDD1670 | [33] |
| LDCM0469 | CL76 | HEK-293T | C870(0.91); C962(0.94); C782(1.03); C350(1.06) | LDD1672 | [33] |
| LDCM0470 | CL77 | HEK-293T | C870(1.16); C962(0.99); C782(0.98); C708(1.65) | LDD1673 | [33] |
| LDCM0471 | CL78 | HEK-293T | C870(0.97); C962(1.00); C782(1.04); C708(0.92) | LDD1674 | [33] |
| LDCM0472 | CL79 | HEK-293T | C870(0.96); C962(0.98); C782(1.04); C708(0.97) | LDD1675 | [33] |
| LDCM0473 | CL8 | HEK-293T | C870(2.49); C962(2.26); C782(1.72); C708(3.38) | LDD1676 | [33] |
| LDCM0474 | CL80 | HEK-293T | C870(0.98); C962(0.98); C782(1.03); C708(0.89) | LDD1677 | [33] |
| LDCM0475 | CL81 | HEK-293T | C870(1.02); C962(0.99); C782(0.99); C708(0.85) | LDD1678 | [33] |
| LDCM0476 | CL82 | HEK-293T | C870(0.98); C962(0.95); C782(1.13); C708(0.99) | LDD1679 | [33] |
| LDCM0477 | CL83 | HEK-293T | C870(1.00); C962(1.09); C782(1.07); C708(1.19) | LDD1680 | [33] |
| LDCM0478 | CL84 | HEK-293T | C870(1.08); C962(1.12); C782(1.15); C708(1.65) | LDD1681 | [33] |
| LDCM0479 | CL85 | HEK-293T | C870(1.05); C962(1.00); C782(0.99); C708(0.88) | LDD1682 | [33] |
| LDCM0480 | CL86 | HEK-293T | C870(1.02); C962(1.18); C782(0.89); C708(0.99) | LDD1683 | [33] |
| LDCM0481 | CL87 | HEK-293T | C870(1.01); C962(0.97); C782(0.94); C708(0.99) | LDD1684 | [33] |
| LDCM0482 | CL88 | HEK-293T | C870(1.01); C962(0.93); C782(1.05); C350(1.10) | LDD1685 | [33] |
| LDCM0483 | CL89 | HEK-293T | C870(1.01); C962(0.98); C782(0.93); C708(0.89) | LDD1686 | [33] |
| LDCM0484 | CL9 | HEK-293T | C870(1.12); C962(1.08); C782(0.96); C708(1.16) | LDD1687 | [33] |
| LDCM0485 | CL90 | HEK-293T | C870(1.89); C962(1.56); C782(1.12); C708(2.15) | LDD1688 | [33] |
| LDCM0486 | CL91 | HEK-293T | C870(0.98); C962(0.73); C782(0.96); C708(0.97) | LDD1689 | [33] |
| LDCM0487 | CL92 | HEK-293T | C870(1.05); C962(0.92); C782(1.00); C708(1.25) | LDD1690 | [33] |
| LDCM0488 | CL93 | HEK-293T | C870(1.14); C962(1.11); C782(1.00); C708(1.39) | LDD1691 | [33] |
| LDCM0489 | CL94 | HEK-293T | C870(0.96); C962(0.92); C782(1.02); C708(1.13) | LDD1692 | [33] |
| LDCM0490 | CL95 | HEK-293T | C870(1.66); C962(1.68); C782(1.32); C708(4.14) | LDD1693 | [33] |
| LDCM0491 | CL96 | HEK-293T | C870(1.08); C962(1.17); C782(1.11); C708(1.53) | LDD1694 | [33] |
| LDCM0492 | CL97 | HEK-293T | C870(1.03); C962(1.32); C782(1.10); C708(1.27) | LDD1695 | [33] |
| LDCM0493 | CL98 | HEK-293T | C870(0.98); C962(0.97); C782(0.95); C708(1.15) | LDD1696 | [33] |
| LDCM0494 | CL99 | HEK-293T | C870(0.96); C962(0.91); C782(0.95); C708(1.22) | LDD1697 | [33] |
| LDCM0191 | Compound 21 | HEK-293T | 3.37 | LDD0508 | [27] |
| LDCM0634 | CY-0357 | Hep-G2 | C708(0.59) | LDD2228 | [19] |
| LDCM0495 | E2913 | HEK-293T | C870(0.92); C962(0.88); C782(0.86); C708(1.29) | LDD1698 | [33] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C782(4.52); C400(3.76) | LDD1702 | [32] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 10.11 | LDD0183 | [8] |
| LDCM0175 | Ethacrynic acid | HeLa | C350(2.23); C42(1.10) | LDD2210 | [11] |
| LDCM0625 | F8 | Ramos | C708(0.87); C400(0.74); C962(1.32); C42(1.04) | LDD2187 | [34] |
| LDCM0572 | Fragment10 | Ramos | C400(0.49); C962(0.63); C42(0.63) | LDD2189 | [34] |
| LDCM0573 | Fragment11 | Ramos | C400(1.87); C962(0.16); C42(0.16) | LDD2190 | [34] |
| LDCM0574 | Fragment12 | Ramos | C400(0.66); C962(0.63); C42(0.86) | LDD2191 | [34] |
| LDCM0575 | Fragment13 | Ramos | C708(1.36); C400(0.97); C962(0.85); C42(0.97) | LDD2192 | [34] |
| LDCM0576 | Fragment14 | Ramos | C708(0.55); C400(1.21); C962(0.61); C42(1.34) | LDD2193 | [34] |
| LDCM0579 | Fragment20 | Ramos | C400(0.50); C962(0.58); C42(0.71) | LDD2194 | [34] |
| LDCM0580 | Fragment21 | Ramos | C708(1.48); C400(1.40); C962(1.15); C42(1.12) | LDD2195 | [34] |
| LDCM0582 | Fragment23 | Ramos | C708(0.59); C400(0.95); C962(0.86); C42(1.21) | LDD2196 | [34] |
| LDCM0578 | Fragment27 | Ramos | C708(1.17); C400(0.94); C962(0.81); C42(1.23) | LDD2197 | [34] |
| LDCM0586 | Fragment28 | Ramos | C708(0.96); C400(0.68); C962(0.73); C42(0.54) | LDD2198 | [34] |
| LDCM0588 | Fragment30 | Ramos | C708(6.63); C400(0.89); C962(1.17); C42(1.02) | LDD2199 | [34] |
| LDCM0589 | Fragment31 | Ramos | C708(1.30); C400(0.62); C962(1.10); C42(1.63) | LDD2200 | [34] |
| LDCM0590 | Fragment32 | Ramos | C400(0.47); C962(0.68); C42(1.29) | LDD2201 | [34] |
| LDCM0468 | Fragment33 | HEK-293T | C870(0.97); C962(0.95); C782(0.95); C708(0.98) | LDD1671 | [33] |
| LDCM0596 | Fragment38 | Ramos | C708(1.31); C400(0.45); C962(0.89); C42(0.89) | LDD2203 | [34] |
| LDCM0566 | Fragment4 | Ramos | C400(0.91); C962(1.05); C42(0.70) | LDD2184 | [34] |
| LDCM0427 | Fragment51 | HEK-293T | C870(1.03); C962(1.03); C782(0.98); C708(0.94) | LDD1631 | [33] |
| LDCM0610 | Fragment52 | Ramos | C708(1.04); C400(1.36); C962(0.83); C42(1.33) | LDD2204 | [34] |
| LDCM0614 | Fragment56 | Ramos | C400(1.11); C962(0.88); C42(1.58) | LDD2205 | [34] |
| LDCM0569 | Fragment7 | Ramos | C708(4.56); C400(0.64); C42(0.47) | LDD2186 | [34] |
| LDCM0571 | Fragment9 | Ramos | C400(0.52); C962(0.51); C42(1.29) | LDD2188 | [34] |
| LDCM0107 | IAA | HeLa | C797(0.00); C962(0.00) | LDD0221 | [23] |
| LDCM0179 | JZ128 | PC-3 | C725(0.00); C42(0.00); C726(0.00); C732(0.00) | LDD0462 | [7] |
| LDCM0022 | KB02 | HEK-293T | C870(0.81); C962(1.00); C782(0.97); C708(0.74) | LDD1492 | [33] |
| LDCM0023 | KB03 | Jurkat | C797(14.05); C708(17.13); C782(5.61); C459(4.74) | LDD0209 | [12] |
| LDCM0024 | KB05 | G361 | C459(4.33); C350(3.69); C400(5.19); C870(5.69) | LDD3311 | [35] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C708(1.26); C797(1.14) | LDD2102 | [32] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C782(1.03); C708(0.88) | LDD2121 | [32] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C782(1.11); C708(0.82) | LDD2089 | [32] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C962(1.14); C708(1.27); C797(1.45) | LDD2090 | [32] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C782(1.05); C708(0.93) | LDD2092 | [32] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C782(1.03); C708(1.22); C797(1.24) | LDD2093 | [32] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C782(1.20); C708(1.07); C725(1.12) | LDD2094 | [32] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C782(0.54); C708(0.13); C870(0.39) | LDD2096 | [32] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C962(0.92); C782(1.00); C797(1.05); C725(1.54) | LDD2097 | [32] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C782(1.07); C708(1.36); C797(0.90) | LDD2098 | [32] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C962(1.09); C782(1.64); C708(0.86); C797(1.12) | LDD2099 | [32] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C708(1.14) | LDD2100 | [32] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C962(1.61); C782(0.90); C708(0.53); C797(0.78) | LDD2101 | [32] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C962(0.92); C782(0.55); C708(0.94) | LDD2104 | [32] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C782(1.07); C708(1.13); C797(1.30) | LDD2105 | [32] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C962(0.81); C708(0.98); C797(0.55) | LDD2106 | [32] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C962(1.02); C782(1.03); C708(1.06); C797(1.24) | LDD2107 | [32] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C962(0.80); C708(1.05) | LDD2108 | [32] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C782(0.63); C708(0.89); C870(0.76); C725(0.66) | LDD2109 | [32] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C708(2.66); C797(1.09) | LDD2110 | [32] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C782(1.07); C708(1.03); C797(1.36); C870(0.88) | LDD2111 | [32] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C782(0.62); C708(0.68) | LDD2114 | [32] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C962(0.89); C708(0.56); C797(0.43); C870(0.50) | LDD2115 | [32] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C782(0.84); C708(0.25); C725(0.32) | LDD2116 | [32] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C962(0.17); C782(0.73); C708(0.23); C725(0.25) | LDD2118 | [32] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C962(2.15); C782(1.67); C708(2.21); C797(1.80) | LDD2119 | [32] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C782(0.99); C708(0.85); C797(0.65); C726(0.85) | LDD2120 | [32] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C962(0.12); C782(0.69); C708(0.25); C870(0.17) | LDD2122 | [32] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C782(0.99); C708(1.00); C797(0.96); C726(0.69) | LDD2123 | [32] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C962(0.11); C782(0.56); C708(0.40) | LDD2124 | [32] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C782(0.96); C708(0.93); C870(0.78); C725(0.99) | LDD2125 | [32] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C962(0.12); C782(0.56); C708(0.28); C870(0.45) | LDD2126 | [32] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C962(1.44); C782(1.19); C708(0.97); C797(1.03) | LDD2127 | [32] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C782(1.23); C797(0.74) | LDD2128 | [32] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C962(1.03); C782(1.34); C708(4.09); C797(1.45) | LDD2129 | [32] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C782(0.93); C708(0.56); C797(0.61); C725(0.85) | LDD2133 | [32] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C782(0.54); C708(0.46); C797(0.43); C1027(0.62) | LDD2134 | [32] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C962(1.66); C782(1.92); C708(0.85); C797(2.03) | LDD2135 | [32] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C962(1.16); C782(1.54); C708(1.10); C797(0.66) | LDD2136 | [32] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C962(1.10); C782(1.07); C708(0.96); C797(0.96) | LDD2137 | [32] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C708(3.89); C782(3.35); C797(2.87); C870(2.00) | LDD1700 | [32] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C782(1.00); C708(0.95); C726(1.08); C870(0.78) | LDD2140 | [32] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C782(0.74); C797(0.59); C870(0.81) | LDD2141 | [32] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C782(0.89); C708(1.07); C726(0.69); C725(0.90) | LDD2143 | [32] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C962(2.45); C708(2.52); C797(1.41); C726(1.04) | LDD2144 | [32] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C708(4.08) | LDD2145 | [32] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C782(1.11); C708(1.08); C797(0.90); C726(1.11) | LDD2146 | [32] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C708(1.30) | LDD2147 | [32] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C962(0.80); C782(0.46); C797(0.47); C725(0.52) | LDD2148 | [32] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C782(0.68); C708(0.22) | LDD2149 | [32] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C962(0.52); C782(0.77); C708(0.58); C797(0.76) | LDD2150 | [32] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C962(0.17); C782(0.70); C708(0.39) | LDD2151 | [32] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C962(1.25); C708(2.32); C797(2.16) | LDD2153 | [32] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C42(5.08); C95(0.90); C708(0.75) | LDD2206 | [36] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C95(0.99); C708(0.27) | LDD2207 | [36] |
| LDCM0131 | RA190 | MM1.R | 2.63 | LDD0300 | [10] |
| LDCM0019 | Staurosporine | Hep-G2 | N.A. | LDD0083 | [30] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| CCAAT/enhancer-binding protein delta (CEBPD) | BZIP family | P49716 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| T-complex protein 11-like protein 1 (TCP11L1) | TCP11 family | Q9NUJ3 | |||
| Hepatocyte growth factor-regulated tyrosine kinase substrate (HGS) | . | O14964 | |||
References
