Details of the Target
General Information of Target
| Target ID | LDTP06905 | |||||
|---|---|---|---|---|---|---|
| Target Name | Acylglycerol kinase, mitochondrial (AGK) | |||||
| Gene Name | AGK | |||||
| Gene ID | 55750 | |||||
| Synonyms |
MULK; Acylglycerol kinase, mitochondrial; hAGK; EC 2.7.1.107; EC 2.7.1.138; EC 2.7.1.94; Multiple substrate lipid kinase; HsMuLK; MuLK; Multi-substrate lipid kinase |
|||||
| 3D Structure | ||||||
| Sequence |
MTVFFKTLRNHWKKTTAGLCLLTWGGHWLYGKHCDNLLRRAACQEAQVFGNQLIPPNAQV
KKATVFLNPAACKGKARTLFEKNAAPILHLSGMDVTIVKTDYEGQAKKLLELMENTDVII VAGGDGTLQEVVTGVLRRTDEATFSKIPIGFIPLGETSSLSHTLFAESGNKVQHITDATL AIVKGETVPLDVLQIKGEKEQPVFAMTGLRWGSFRDAGVKVSKYWYLGPLKIKAAHFFST LKEWPQTHQASISYTGPTERPPNEPEETPVQRPSLYRRILRRLASYWAQPQDALSQEVSP EVWKDVQLSTIELSITTRNNQLDPTSKEDFLNICIEPDTISKGDFITIGSRKVRNPKLHV EGTECLQASQCTLLIPEGAGGSFSIDSEEYEAMPVEVKLLPRKLQFFCDPRKREQMLTSP TQ |
|||||
| Target Type |
Literature-reported
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
AGK family
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Lipid kinase that can phosphorylate both monoacylglycerol and diacylglycerol to form lysophosphatidic acid (LPA) and phosphatidic acid (PA), respectively. Does not phosphorylate sphingosine. Phosphorylates ceramide. Phosphorylates 1,2-dioleoylglycerol more rapidly than 2,3-dioleoylglycerol. Independently of its lipid kinase activity, acts as a component of the TIM22 complex. The TIM22 complex mediates the import and insertion of multi-pass transmembrane proteins into the mitochondrial inner membrane by forming a twin-pore translocase that uses the membrane potential as the external driving force. In the TIM22 complex, required for the import of a subset of metabolite carriers into mitochondria, such as ANT1/SLC25A4 and SLC25A24, while it is not required for the import of TIMM23. Overexpression increases the formation and secretion of LPA, resulting in transactivation of EGFR and activation of the downstream MAPK signaling pathway, leading to increased cell growth.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CALU6 | SNV: p.L275M | DBIA Probe Info | |||
| GB2 | SNV: p.F238C | DBIA Probe Info | |||
| KATOIII | SNV: p.I119V | DBIA Probe Info | |||
| LN18 | SNV: p.T363R | DBIA Probe Info | |||
| SW480 | SNV: p.A250T | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
HDSF-alk Probe Info |
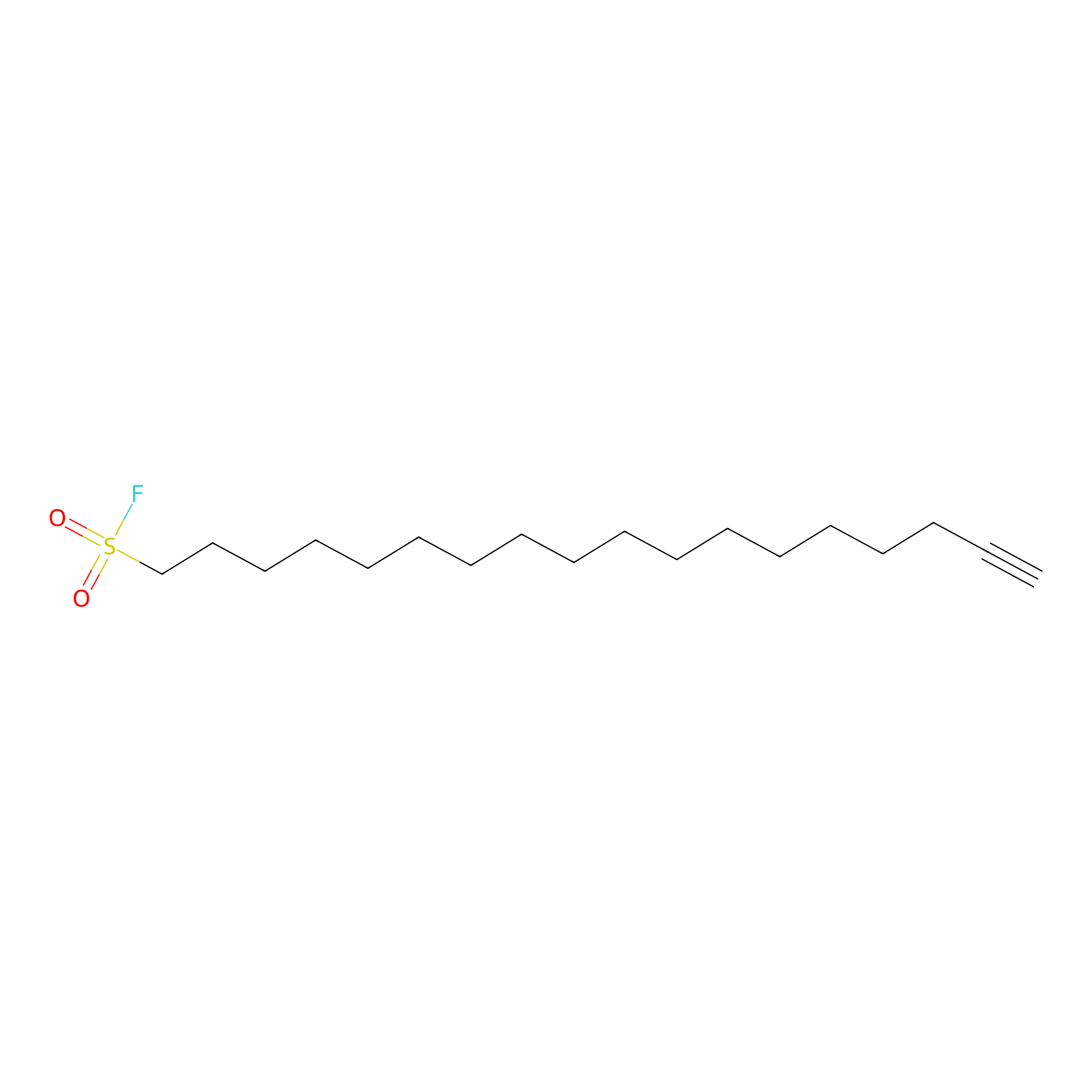 |
1.78 | LDD0197 | [1] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [2] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [2] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [3] | |
|
BTD Probe Info |
 |
C72(1.71) | LDD1700 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C34(2.14) | LDD0168 | [5] | |
|
HPAP Probe Info |
 |
3.04 | LDD0062 | [6] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [7] | |
|
DBIA Probe Info |
 |
C34(0.99) | LDD0078 | [8] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [9] | |
|
ATP probe Probe Info |
 |
K73(0.00); K75(0.00); K196(0.00); K199(0.00) | LDD0199 | [10] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [9] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C34(0.00); C43(0.00); C72(0.00); C408(0.00) | LDD0038 | [11] | |
|
IA-alkyne Probe Info |
 |
C408(0.00); C334(0.00); C72(0.00); C43(0.00) | LDD0032 | [12] | |
|
Lodoacetamide azide Probe Info |
 |
C408(0.00); C72(0.00); C334(0.00); C20(0.00) | LDD0037 | [11] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [13] | |
|
WYneN Probe Info |
 |
C72(0.00); C34(0.00) | LDD0021 | [14] | |
|
ENE Probe Info |
 |
C72(0.00); C34(0.00) | LDD0006 | [14] | |
|
IPM Probe Info |
 |
C43(0.00); C34(0.00); C72(0.00); C408(0.00) | LDD0005 | [14] | |
|
NHS Probe Info |
 |
K233(0.00); K6(0.00) | LDD0010 | [14] | |
|
PPMS Probe Info |
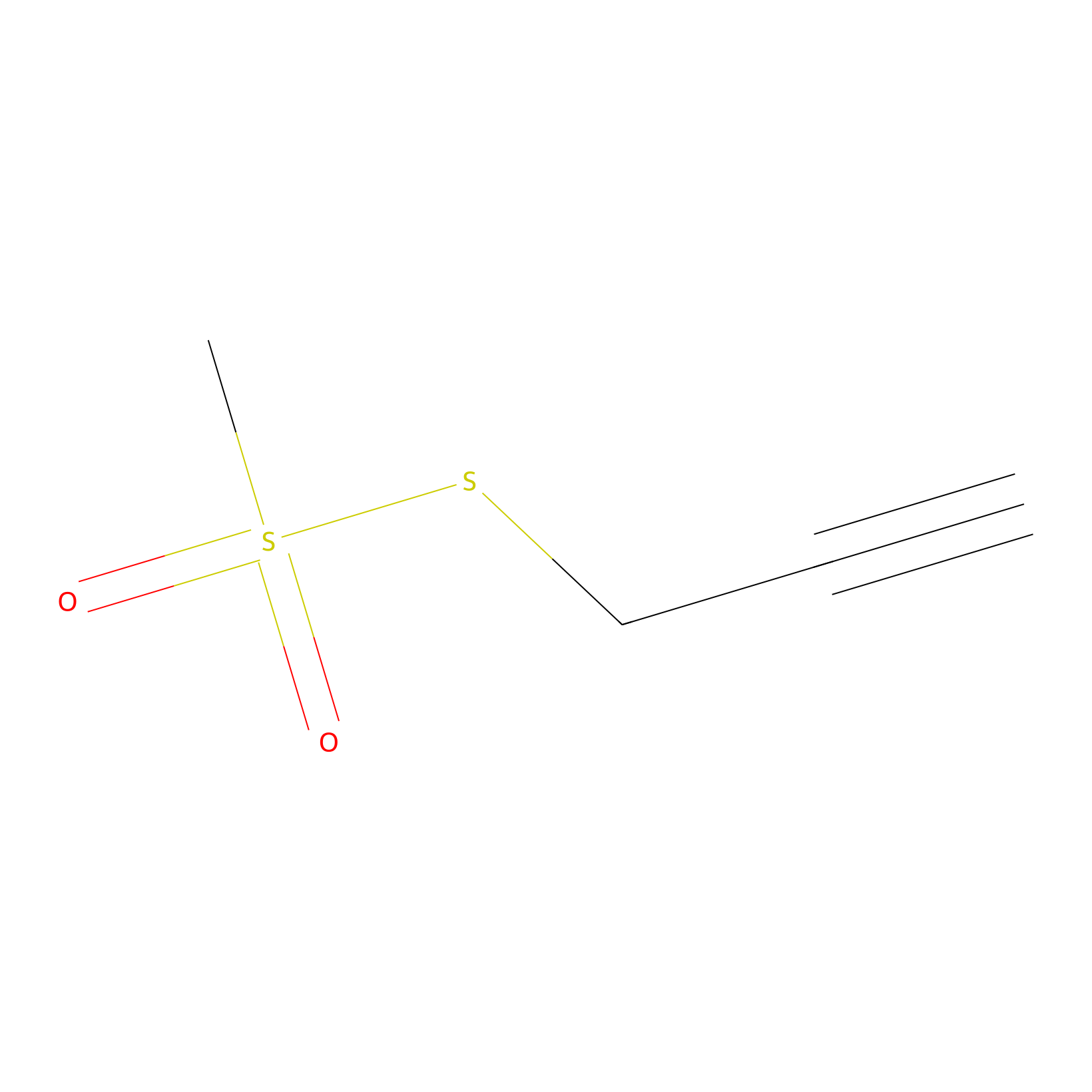 |
N.A. | LDD0008 | [14] | |
|
STPyne Probe Info |
 |
K233(0.00); K73(0.00) | LDD0009 | [14] | |
|
VSF Probe Info |
 |
C43(0.00); C334(0.00); C34(0.00) | LDD0007 | [14] | |
|
Phosphinate-6 Probe Info |
 |
C408(0.00); C72(0.00) | LDD0018 | [15] | |
|
1c-yne Probe Info |
 |
K223(0.00); K171(0.00) | LDD0228 | [16] | |
|
Acrolein Probe Info |
 |
C72(0.00); H162(0.00); C408(0.00); C334(0.00) | LDD0217 | [17] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [17] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [17] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [18] | |
|
NAIA_5 Probe Info |
 |
C43(0.00); C72(0.00); C34(0.00); C408(0.00) | LDD2223 | [19] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [20] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C040 Probe Info |
 |
7.57 | LDD1740 | [21] | |
|
C055 Probe Info |
 |
18.77 | LDD1752 | [21] | |
|
C106 Probe Info |
 |
17.03 | LDD1793 | [21] | |
|
C134 Probe Info |
 |
20.25 | LDD1816 | [21] | |
|
C161 Probe Info |
 |
10.13 | LDD1841 | [21] | |
|
C201 Probe Info |
 |
31.78 | LDD1877 | [21] | |
|
C210 Probe Info |
 |
61.82 | LDD1884 | [21] | |
|
C218 Probe Info |
 |
11.79 | LDD1892 | [21] | |
|
C228 Probe Info |
 |
14.83 | LDD1901 | [21] | |
|
C233 Probe Info |
 |
8.63 | LDD1906 | [21] | |
|
C246 Probe Info |
 |
12.82 | LDD1919 | [21] | |
|
C285 Probe Info |
 |
22.01 | LDD1955 | [21] | |
|
C289 Probe Info |
 |
27.47 | LDD1959 | [21] | |
|
C362 Probe Info |
 |
31.12 | LDD2023 | [21] | |
|
C388 Probe Info |
 |
43.41 | LDD2047 | [21] | |
|
C390 Probe Info |
 |
37.53 | LDD2049 | [21] | |
|
C403 Probe Info |
 |
15.45 | LDD2061 | [21] | |
|
C420 Probe Info |
 |
10.20 | LDD2075 | [21] | |
|
FFF probe11 Probe Info |
 |
16.55 | LDD0471 | [22] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [22] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [22] | |
|
FFF probe15 Probe Info |
 |
9.03 | LDD0478 | [22] | |
|
FFF probe2 Probe Info |
 |
20.00 | LDD0463 | [22] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [22] | |
|
FFF probe4 Probe Info |
 |
12.83 | LDD0466 | [22] | |
|
FFF probe6 Probe Info |
 |
6.64 | LDD0467 | [22] | |
|
JN0003 Probe Info |
 |
20.00 | LDD0469 | [22] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [23] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [24] | |
|
OEA-DA Probe Info |
 |
4.65 | LDD0046 | [25] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0068 | [26] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C34(1.05) | LDD2152 | [4] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C72(0.98) | LDD2131 | [4] |
| LDCM0025 | 4SU-RNA | HEK-293T | C34(2.14) | LDD0168 | [5] |
| LDCM0214 | AC1 | HCT 116 | C34(1.18); C408(0.75); C72(0.99) | LDD0531 | [8] |
| LDCM0215 | AC10 | HCT 116 | C408(0.67); C43(0.55); C72(0.79) | LDD0532 | [8] |
| LDCM0216 | AC100 | HCT 116 | C408(0.53); C43(0.42); C72(0.81) | LDD0533 | [8] |
| LDCM0217 | AC101 | HCT 116 | C408(0.54); C43(0.48); C72(0.73) | LDD0534 | [8] |
| LDCM0218 | AC102 | HCT 116 | C408(0.52); C43(0.40); C72(0.73) | LDD0535 | [8] |
| LDCM0219 | AC103 | HCT 116 | C408(0.39); C43(0.28); C72(0.63) | LDD0536 | [8] |
| LDCM0220 | AC104 | HCT 116 | C408(0.50); C43(0.29); C72(0.78) | LDD0537 | [8] |
| LDCM0221 | AC105 | HCT 116 | C408(0.43); C43(0.28); C72(0.75) | LDD0538 | [8] |
| LDCM0222 | AC106 | HCT 116 | C408(0.45); C43(0.30); C72(0.74) | LDD0539 | [8] |
| LDCM0223 | AC107 | HCT 116 | C408(0.44); C43(0.51); C72(0.67) | LDD0540 | [8] |
| LDCM0224 | AC108 | HCT 116 | C408(0.72); C43(0.31); C72(0.74) | LDD0541 | [8] |
| LDCM0225 | AC109 | HCT 116 | C408(0.63); C43(0.29); C72(0.85) | LDD0542 | [8] |
| LDCM0226 | AC11 | HCT 116 | C408(0.55); C43(0.63); C72(0.88) | LDD0543 | [8] |
| LDCM0227 | AC110 | HCT 116 | C408(0.54); C43(0.35); C72(0.74) | LDD0544 | [8] |
| LDCM0228 | AC111 | HCT 116 | C408(0.53); C43(0.39); C72(0.73) | LDD0545 | [8] |
| LDCM0229 | AC112 | HCT 116 | C408(0.46); C43(0.39); C72(0.72) | LDD0546 | [8] |
| LDCM0230 | AC113 | HCT 116 | C34(0.93); C408(1.04); C72(0.99) | LDD0547 | [8] |
| LDCM0231 | AC114 | HCT 116 | C34(0.95); C408(0.90); C72(0.87) | LDD0548 | [8] |
| LDCM0232 | AC115 | HCT 116 | C34(0.87); C408(0.92); C72(0.98) | LDD0549 | [8] |
| LDCM0233 | AC116 | HCT 116 | C34(1.01); C408(1.00); C72(0.74) | LDD0550 | [8] |
| LDCM0234 | AC117 | HCT 116 | C34(1.07); C408(0.85); C72(0.89) | LDD0551 | [8] |
| LDCM0235 | AC118 | HCT 116 | C34(1.16); C408(0.84); C72(0.93) | LDD0552 | [8] |
| LDCM0236 | AC119 | HCT 116 | C34(0.94); C408(1.12); C72(1.24) | LDD0553 | [8] |
| LDCM0237 | AC12 | HCT 116 | C408(0.94); C43(0.72); C72(0.96) | LDD0554 | [8] |
| LDCM0238 | AC120 | HCT 116 | C34(0.93); C408(0.96); C72(0.82) | LDD0555 | [8] |
| LDCM0239 | AC121 | HCT 116 | C34(1.47); C408(0.80); C72(1.13) | LDD0556 | [8] |
| LDCM0240 | AC122 | HCT 116 | C34(1.00); C408(0.89); C72(0.85) | LDD0557 | [8] |
| LDCM0241 | AC123 | HCT 116 | C34(1.21); C408(1.11); C72(0.98) | LDD0558 | [8] |
| LDCM0242 | AC124 | HCT 116 | C34(1.21); C408(0.85); C72(1.01) | LDD0559 | [8] |
| LDCM0243 | AC125 | HCT 116 | C34(1.04); C408(1.17); C72(0.89) | LDD0560 | [8] |
| LDCM0244 | AC126 | HCT 116 | C34(1.24); C408(0.91); C72(1.26) | LDD0561 | [8] |
| LDCM0245 | AC127 | HCT 116 | C34(1.27); C408(0.91); C72(1.14) | LDD0562 | [8] |
| LDCM0246 | AC128 | HCT 116 | C34(0.62); C408(0.89); C72(1.05) | LDD0563 | [8] |
| LDCM0247 | AC129 | HCT 116 | C34(0.72); C408(1.12); C72(1.21) | LDD0564 | [8] |
| LDCM0249 | AC130 | HCT 116 | C34(0.60); C408(0.89); C72(0.98) | LDD0566 | [8] |
| LDCM0250 | AC131 | HCT 116 | C34(1.34); C408(1.22); C72(1.04) | LDD0567 | [8] |
| LDCM0251 | AC132 | HCT 116 | C34(0.80); C408(0.97); C72(0.97) | LDD0568 | [8] |
| LDCM0252 | AC133 | HCT 116 | C34(0.97); C408(0.85); C72(1.01) | LDD0569 | [8] |
| LDCM0253 | AC134 | HCT 116 | C34(1.11); C408(0.84); C72(0.86) | LDD0570 | [8] |
| LDCM0254 | AC135 | HCT 116 | C34(0.72); C408(0.85); C72(1.04) | LDD0571 | [8] |
| LDCM0255 | AC136 | HCT 116 | C34(0.72); C408(0.88); C72(0.89) | LDD0572 | [8] |
| LDCM0256 | AC137 | HCT 116 | C34(0.94); C408(0.87); C72(0.92) | LDD0573 | [8] |
| LDCM0257 | AC138 | HCT 116 | C34(0.77); C408(0.84); C72(0.84) | LDD0574 | [8] |
| LDCM0258 | AC139 | HCT 116 | C34(0.92); C408(0.81); C72(0.92) | LDD0575 | [8] |
| LDCM0259 | AC14 | HCT 116 | C408(0.67); C43(0.59); C72(0.85) | LDD0576 | [8] |
| LDCM0260 | AC140 | HCT 116 | C34(0.76); C408(0.79); C72(0.81) | LDD0577 | [8] |
| LDCM0261 | AC141 | HCT 116 | C34(1.00); C408(0.84); C72(0.81) | LDD0578 | [8] |
| LDCM0262 | AC142 | HCT 116 | C34(0.99); C408(0.93); C72(0.93) | LDD0579 | [8] |
| LDCM0263 | AC143 | HCT 116 | C34(0.82); C408(0.72); C43(0.81); C72(0.89) | LDD0580 | [8] |
| LDCM0264 | AC144 | HCT 116 | C408(0.31); C72(1.09); C34(1.15); C43(1.16) | LDD0581 | [8] |
| LDCM0265 | AC145 | HCT 116 | C408(0.60); C34(1.10); C72(1.13); C43(1.25) | LDD0582 | [8] |
| LDCM0266 | AC146 | HCT 116 | C408(0.32); C72(1.13); C34(1.18); C43(1.31) | LDD0583 | [8] |
| LDCM0267 | AC147 | HCT 116 | C408(0.30); C43(0.98); C72(1.04); C34(1.10) | LDD0584 | [8] |
| LDCM0268 | AC148 | HCT 116 | C408(0.27); C72(1.13); C34(1.24); C43(1.25) | LDD0585 | [8] |
| LDCM0269 | AC149 | HCT 116 | C408(0.27); C34(1.09); C72(1.26); C43(1.63) | LDD0586 | [8] |
| LDCM0270 | AC15 | HCT 116 | C43(0.48); C408(0.75); C72(0.84) | LDD0587 | [8] |
| LDCM0271 | AC150 | HCT 116 | C408(0.60); C72(1.18); C34(1.22); C43(1.22) | LDD0588 | [8] |
| LDCM0272 | AC151 | HCT 116 | C408(0.70); C43(1.12); C34(1.12); C72(1.23) | LDD0589 | [8] |
| LDCM0273 | AC152 | HCT 116 | C408(0.36); C43(1.06); C34(1.22); C72(1.38) | LDD0590 | [8] |
| LDCM0274 | AC153 | HCT 116 | C408(0.27); C34(1.20); C72(1.44); C43(1.66) | LDD0591 | [8] |
| LDCM0621 | AC154 | HCT 116 | C34(1.17); C408(0.37); C43(1.17); C72(1.25) | LDD2158 | [8] |
| LDCM0622 | AC155 | HCT 116 | C34(1.28); C408(0.38); C43(1.31); C72(1.20) | LDD2159 | [8] |
| LDCM0623 | AC156 | HCT 116 | C34(1.15); C408(0.78); C43(1.24); C72(1.09) | LDD2160 | [8] |
| LDCM0624 | AC157 | HCT 116 | C34(1.18); C408(0.80); C43(1.23); C72(1.23) | LDD2161 | [8] |
| LDCM0276 | AC17 | HCT 116 | C408(0.90); C43(0.92) | LDD0593 | [8] |
| LDCM0277 | AC18 | HCT 116 | C43(0.53); C408(0.65) | LDD0594 | [8] |
| LDCM0278 | AC19 | HCT 116 | C43(0.71); C408(0.75) | LDD0595 | [8] |
| LDCM0279 | AC2 | HCT 116 | C408(0.78); C72(0.99); C34(1.07) | LDD0596 | [8] |
| LDCM0280 | AC20 | HCT 116 | C408(0.93); C43(1.22) | LDD0597 | [8] |
| LDCM0281 | AC21 | HCT 116 | C43(0.81); C408(0.99) | LDD0598 | [8] |
| LDCM0282 | AC22 | HCT 116 | C43(0.95); C408(1.35) | LDD0599 | [8] |
| LDCM0283 | AC23 | HCT 116 | C43(0.81); C408(0.97) | LDD0600 | [8] |
| LDCM0284 | AC24 | HCT 116 | C408(1.02); C43(1.03) | LDD0601 | [8] |
| LDCM0285 | AC25 | HCT 116 | C43(0.67); C408(0.96) | LDD0602 | [8] |
| LDCM0286 | AC26 | HCT 116 | C43(0.78); C408(0.82) | LDD0603 | [8] |
| LDCM0287 | AC27 | HCT 116 | C43(0.61); C408(0.83) | LDD0604 | [8] |
| LDCM0288 | AC28 | HCT 116 | C43(0.76); C408(0.93) | LDD0605 | [8] |
| LDCM0289 | AC29 | HCT 116 | C43(0.79); C408(0.93) | LDD0606 | [8] |
| LDCM0290 | AC3 | HCT 116 | C408(0.82); C72(1.03); C34(1.32) | LDD0607 | [8] |
| LDCM0291 | AC30 | HCT 116 | C408(0.59); C43(0.67) | LDD0608 | [8] |
| LDCM0292 | AC31 | HCT 116 | C43(0.53); C408(0.72) | LDD0609 | [8] |
| LDCM0293 | AC32 | HCT 116 | C43(0.51); C408(0.56) | LDD0610 | [8] |
| LDCM0294 | AC33 | HCT 116 | C43(0.54); C408(0.71) | LDD0611 | [8] |
| LDCM0295 | AC34 | HCT 116 | C43(0.50); C408(0.68) | LDD0612 | [8] |
| LDCM0296 | AC35 | HCT 116 | C72(1.10); C43(1.22); C34(1.36); C408(1.49) | LDD0613 | [8] |
| LDCM0297 | AC36 | HCT 116 | C43(0.73); C72(1.18); C408(1.31); C34(1.39) | LDD0614 | [8] |
| LDCM0298 | AC37 | HCT 116 | C43(0.89); C34(1.16); C72(1.18); C408(1.21) | LDD0615 | [8] |
| LDCM0299 | AC38 | HCT 116 | C43(0.96); C72(1.11); C34(1.13); C408(1.19) | LDD0616 | [8] |
| LDCM0300 | AC39 | HCT 116 | C43(0.82); C408(0.97); C72(1.06); C34(1.12) | LDD0617 | [8] |
| LDCM0301 | AC4 | HCT 116 | C408(0.71); C72(0.99); C34(1.53) | LDD0618 | [8] |
| LDCM0302 | AC40 | HCT 116 | C43(0.68); C408(0.95); C34(0.95); C72(1.00) | LDD0619 | [8] |
| LDCM0303 | AC41 | HCT 116 | C43(0.85); C408(1.07); C34(1.11); C72(1.16) | LDD0620 | [8] |
| LDCM0304 | AC42 | HCT 116 | C43(0.72); C408(0.97); C72(1.06); C34(1.12) | LDD0621 | [8] |
| LDCM0305 | AC43 | HCT 116 | C43(0.81); C408(0.93); C34(0.97); C72(1.04) | LDD0622 | [8] |
| LDCM0306 | AC44 | HCT 116 | C43(0.79); C34(1.03); C408(1.06); C72(1.20) | LDD0623 | [8] |
| LDCM0307 | AC45 | HCT 116 | C43(0.75); C34(0.90); C408(0.92); C72(0.99) | LDD0624 | [8] |
| LDCM0308 | AC46 | HCT 116 | C72(0.92); C408(0.96); C34(1.00); C43(1.02) | LDD0625 | [8] |
| LDCM0309 | AC47 | HCT 116 | C408(0.83); C43(0.92); C34(0.97); C72(1.06) | LDD0626 | [8] |
| LDCM0310 | AC48 | HCT 116 | C408(0.65); C43(0.74); C72(0.92); C34(1.02) | LDD0627 | [8] |
| LDCM0311 | AC49 | HCT 116 | C408(0.61); C43(0.78); C34(0.80); C72(0.81) | LDD0628 | [8] |
| LDCM0312 | AC5 | HCT 116 | C408(0.72); C72(1.01); C34(1.48) | LDD0629 | [8] |
| LDCM0313 | AC50 | HCT 116 | C408(0.62); C43(0.71); C34(0.83); C72(0.89) | LDD0630 | [8] |
| LDCM0314 | AC51 | HCT 116 | C72(0.87); C43(1.09); C34(1.28); C408(1.41) | LDD0631 | [8] |
| LDCM0315 | AC52 | HCT 116 | C72(0.90); C34(0.91); C43(0.99); C408(1.02) | LDD0632 | [8] |
| LDCM0316 | AC53 | HCT 116 | C408(0.71); C43(0.76); C34(0.77); C72(0.98) | LDD0633 | [8] |
| LDCM0317 | AC54 | HCT 116 | C408(0.80); C43(0.86); C34(0.89); C72(0.95) | LDD0634 | [8] |
| LDCM0318 | AC55 | HCT 116 | C408(0.64); C43(0.73); C34(0.81); C72(1.10) | LDD0635 | [8] |
| LDCM0319 | AC56 | HCT 116 | C408(0.68); C43(0.80); C34(0.82); C72(1.01) | LDD0636 | [8] |
| LDCM0320 | AC57 | HCT 116 | C43(0.53); C408(0.63); C72(0.85); C34(0.88) | LDD0637 | [8] |
| LDCM0321 | AC58 | HCT 116 | C43(0.51); C408(0.64); C72(0.65); C34(0.90) | LDD0638 | [8] |
| LDCM0322 | AC59 | HCT 116 | C43(0.50); C72(0.65); C408(0.70); C34(1.00) | LDD0639 | [8] |
| LDCM0323 | AC6 | HCT 116 | C408(0.66); C43(0.67); C72(0.94) | LDD0640 | [8] |
| LDCM0324 | AC60 | HCT 116 | C43(0.49); C72(0.53); C408(0.76); C34(0.89) | LDD0641 | [8] |
| LDCM0325 | AC61 | HCT 116 | C43(0.52); C72(0.61); C408(0.70); C34(0.96) | LDD0642 | [8] |
| LDCM0326 | AC62 | HCT 116 | C43(0.45); C72(0.63); C408(0.74); C34(0.88) | LDD0643 | [8] |
| LDCM0327 | AC63 | HCT 116 | C43(0.49); C72(0.50); C408(0.76); C34(0.91) | LDD0644 | [8] |
| LDCM0328 | AC64 | HCT 116 | C43(0.49); C72(0.66); C408(0.73); C34(0.95) | LDD0645 | [8] |
| LDCM0329 | AC65 | HCT 116 | C43(0.53); C72(0.65); C408(0.68); C34(0.92) | LDD0646 | [8] |
| LDCM0330 | AC66 | HCT 116 | C43(0.51); C72(0.62); C408(0.69); C34(0.91) | LDD0647 | [8] |
| LDCM0331 | AC67 | HCT 116 | C43(0.39); C72(0.49); C408(0.75); C34(0.78) | LDD0648 | [8] |
| LDCM0332 | AC68 | HCT 116 | C408(0.81); C72(1.03); C34(1.05) | LDD0649 | [8] |
| LDCM0333 | AC69 | HCT 116 | C408(0.75); C34(1.08); C72(1.15) | LDD0650 | [8] |
| LDCM0334 | AC7 | HCT 116 | C43(0.58); C408(0.79); C72(1.00) | LDD0651 | [8] |
| LDCM0335 | AC70 | HCT 116 | C408(0.73); C34(0.91); C72(1.11) | LDD0652 | [8] |
| LDCM0336 | AC71 | HCT 116 | C408(0.88); C34(1.06); C72(1.17) | LDD0653 | [8] |
| LDCM0337 | AC72 | HCT 116 | C408(0.83); C34(0.93); C72(1.04) | LDD0654 | [8] |
| LDCM0338 | AC73 | HCT 116 | C408(0.77); C72(1.06); C34(1.08) | LDD0655 | [8] |
| LDCM0339 | AC74 | HCT 116 | C408(0.68); C34(1.08); C72(1.11) | LDD0656 | [8] |
| LDCM0340 | AC75 | HCT 116 | C408(0.81); C34(0.84); C72(1.07) | LDD0657 | [8] |
| LDCM0341 | AC76 | HCT 116 | C408(0.79); C34(1.09); C72(1.11) | LDD0658 | [8] |
| LDCM0342 | AC77 | HCT 116 | C408(0.85); C34(0.98); C72(1.00) | LDD0659 | [8] |
| LDCM0343 | AC78 | HCT 116 | C408(0.91); C72(1.03); C34(1.13) | LDD0660 | [8] |
| LDCM0344 | AC79 | HCT 116 | C408(0.90); C72(1.04); C34(1.15) | LDD0661 | [8] |
| LDCM0345 | AC8 | HCT 116 | C43(0.61); C408(0.65); C72(0.90) | LDD0662 | [8] |
| LDCM0346 | AC80 | HCT 116 | C408(0.85); C34(1.01); C72(1.11) | LDD0663 | [8] |
| LDCM0347 | AC81 | HCT 116 | C34(0.96); C72(1.02); C408(1.17) | LDD0664 | [8] |
| LDCM0348 | AC82 | HCT 116 | C408(0.79); C72(1.06); C34(1.08) | LDD0665 | [8] |
| LDCM0349 | AC83 | HCT 116 | C43(0.41); C408(0.45); C34(0.88); C72(0.95) | LDD0666 | [8] |
| LDCM0350 | AC84 | HCT 116 | C43(0.45); C408(0.49); C34(0.78); C72(0.86) | LDD0667 | [8] |
| LDCM0351 | AC85 | HCT 116 | C43(0.53); C408(0.55); C34(0.70); C72(0.83) | LDD0668 | [8] |
| LDCM0352 | AC86 | HCT 116 | C43(0.50); C408(0.57); C72(0.80); C34(0.83) | LDD0669 | [8] |
| LDCM0353 | AC87 | HCT 116 | C408(0.84); C43(0.89); C72(0.94); C34(1.17) | LDD0670 | [8] |
| LDCM0354 | AC88 | HCT 116 | C408(0.62); C43(0.74); C72(0.86); C34(1.03) | LDD0671 | [8] |
| LDCM0355 | AC89 | HCT 116 | C43(0.44); C408(0.48); C72(0.76); C34(0.83) | LDD0672 | [8] |
| LDCM0357 | AC90 | HCT 116 | C72(0.81); C43(0.88); C408(0.98); C34(1.10) | LDD0674 | [8] |
| LDCM0358 | AC91 | HCT 116 | C43(0.43); C408(0.48); C34(0.74); C72(0.79) | LDD0675 | [8] |
| LDCM0359 | AC92 | HCT 116 | C43(0.42); C408(0.51); C34(0.75); C72(0.86) | LDD0676 | [8] |
| LDCM0360 | AC93 | HCT 116 | C43(0.57); C408(0.66); C34(0.84); C72(0.93) | LDD0677 | [8] |
| LDCM0361 | AC94 | HCT 116 | C34(0.72); C43(0.73); C408(0.80); C72(0.81) | LDD0678 | [8] |
| LDCM0362 | AC95 | HCT 116 | C408(0.69); C43(0.70); C72(0.82); C34(0.87) | LDD0679 | [8] |
| LDCM0363 | AC96 | HCT 116 | C43(0.43); C408(0.71); C34(0.79); C72(0.79) | LDD0680 | [8] |
| LDCM0364 | AC97 | HCT 116 | C43(0.38); C408(0.46); C34(0.78); C72(0.83) | LDD0681 | [8] |
| LDCM0365 | AC98 | HCT 116 | C43(0.21); C408(0.38); C72(0.58) | LDD0682 | [8] |
| LDCM0366 | AC99 | HCT 116 | C43(0.54); C408(0.63); C72(0.79) | LDD0683 | [8] |
| LDCM0545 | Acetamide | MDA-MB-231 | C34(0.94) | LDD2138 | [4] |
| LDCM0248 | AKOS034007472 | HCT 116 | C408(0.89); C43(0.70); C72(0.96) | LDD0565 | [8] |
| LDCM0356 | AKOS034007680 | HCT 116 | C408(0.64); C43(0.73); C72(0.88) | LDD0673 | [8] |
| LDCM0275 | AKOS034007705 | HCT 116 | C43(0.43); C408(0.65); C72(0.77) | LDD0592 | [8] |
| LDCM0020 | ARS-1620 | HCC44 | C34(0.99) | LDD0078 | [8] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C72(0.55); C34(0.79) | LDD2091 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | C334(0.00); C72(0.00); C408(0.00); H162(0.00) | LDD0222 | [17] |
| LDCM0632 | CL-Sc | Hep-G2 | C72(0.67) | LDD2227 | [19] |
| LDCM0367 | CL1 | HCT 116 | C408(1.00); C72(1.12); C34(1.19); C43(1.36) | LDD0684 | [8] |
| LDCM0368 | CL10 | HCT 116 | C43(0.79); C72(0.87); C34(0.92); C408(0.92) | LDD0685 | [8] |
| LDCM0369 | CL100 | HCT 116 | C408(0.76); C34(0.94); C72(0.96) | LDD0686 | [8] |
| LDCM0370 | CL101 | HCT 116 | C43(0.62); C408(0.68); C72(0.82) | LDD0687 | [8] |
| LDCM0371 | CL102 | HCT 116 | C43(0.65); C408(0.83); C72(1.00) | LDD0688 | [8] |
| LDCM0372 | CL103 | HCT 116 | C43(0.67); C408(0.87); C72(0.90) | LDD0689 | [8] |
| LDCM0373 | CL104 | HCT 116 | C43(0.69); C408(0.78); C72(0.86) | LDD0690 | [8] |
| LDCM0374 | CL105 | HCT 116 | C43(0.66); C408(0.73) | LDD0691 | [8] |
| LDCM0375 | CL106 | HCT 116 | C43(0.55); C408(0.67) | LDD0692 | [8] |
| LDCM0376 | CL107 | HCT 116 | C408(0.63); C43(0.73) | LDD0693 | [8] |
| LDCM0377 | CL108 | HCT 116 | C43(0.53); C408(0.60) | LDD0694 | [8] |
| LDCM0378 | CL109 | HCT 116 | C43(0.78); C408(0.80) | LDD0695 | [8] |
| LDCM0379 | CL11 | HCT 116 | C43(0.85); C34(0.94); C72(1.00); C408(1.04) | LDD0696 | [8] |
| LDCM0380 | CL110 | HCT 116 | C408(0.67); C43(0.75) | LDD0697 | [8] |
| LDCM0381 | CL111 | HCT 116 | C43(0.64); C408(0.74) | LDD0698 | [8] |
| LDCM0382 | CL112 | HCT 116 | C43(0.97); C408(1.02) | LDD0699 | [8] |
| LDCM0383 | CL113 | HCT 116 | C408(0.58); C43(0.61) | LDD0700 | [8] |
| LDCM0384 | CL114 | HCT 116 | C43(0.61); C408(0.81) | LDD0701 | [8] |
| LDCM0385 | CL115 | HCT 116 | C43(0.60); C408(0.74) | LDD0702 | [8] |
| LDCM0386 | CL116 | HCT 116 | C43(0.67); C408(0.75) | LDD0703 | [8] |
| LDCM0387 | CL117 | HCT 116 | C43(0.74); C408(0.92); C72(0.96); C34(1.01) | LDD0704 | [8] |
| LDCM0388 | CL118 | HCT 116 | C43(0.86); C408(0.96); C34(0.97); C72(1.09) | LDD0705 | [8] |
| LDCM0389 | CL119 | HCT 116 | C43(0.78); C408(0.91); C34(0.92); C72(1.00) | LDD0706 | [8] |
| LDCM0390 | CL12 | HCT 116 | C72(0.93); C43(0.96); C34(1.02); C408(1.06) | LDD0707 | [8] |
| LDCM0391 | CL120 | HCT 116 | C43(0.77); C408(1.05); C72(1.13); C34(1.18) | LDD0708 | [8] |
| LDCM0392 | CL121 | HCT 116 | C408(0.82); C43(0.84); C34(0.85); C72(0.94) | LDD0709 | [8] |
| LDCM0393 | CL122 | HCT 116 | C408(0.64); C43(0.87); C34(0.88); C72(0.95) | LDD0710 | [8] |
| LDCM0394 | CL123 | HCT 116 | C43(0.68); C408(0.71); C34(0.79); C72(0.94) | LDD0711 | [8] |
| LDCM0395 | CL124 | HCT 116 | C408(0.65); C43(0.69); C34(0.76); C72(0.92) | LDD0712 | [8] |
| LDCM0396 | CL125 | HCT 116 | C408(0.84); C72(0.85); C43(1.09); C34(1.22) | LDD0713 | [8] |
| LDCM0397 | CL126 | HCT 116 | C72(0.67); C408(0.75); C43(0.76); C34(0.94) | LDD0714 | [8] |
| LDCM0398 | CL127 | HCT 116 | C43(0.73); C408(0.80); C72(0.85); C34(1.00) | LDD0715 | [8] |
| LDCM0399 | CL128 | HCT 116 | C43(0.52); C72(0.56); C408(0.68); C34(0.95) | LDD0716 | [8] |
| LDCM0400 | CL13 | HCT 116 | C72(0.95); C43(1.03); C408(1.03); C34(1.04) | LDD0717 | [8] |
| LDCM0401 | CL14 | HCT 116 | C408(1.01); C34(1.07); C43(1.14); C72(1.14) | LDD0718 | [8] |
| LDCM0402 | CL15 | HCT 116 | C34(0.96); C43(0.97); C72(1.00); C408(1.18) | LDD0719 | [8] |
| LDCM0403 | CL16 | HCT 116 | C43(0.70); C408(0.82); C34(0.85); C72(0.95) | LDD0720 | [8] |
| LDCM0404 | CL17 | HCT 116 | C43(0.59); C34(0.78); C408(0.91); C72(1.03) | LDD0721 | [8] |
| LDCM0405 | CL18 | HCT 116 | C43(0.61); C34(0.79); C408(0.81); C72(0.97) | LDD0722 | [8] |
| LDCM0406 | CL19 | HCT 116 | C43(0.58); C408(0.76); C34(0.79); C72(1.02) | LDD0723 | [8] |
| LDCM0407 | CL2 | HCT 116 | C408(0.94); C34(1.15); C43(1.20); C72(1.21) | LDD0724 | [8] |
| LDCM0408 | CL20 | HCT 116 | C43(0.52); C34(0.73); C408(0.82); C72(0.98) | LDD0725 | [8] |
| LDCM0409 | CL21 | HCT 116 | C43(0.51); C34(0.74); C408(0.76); C72(0.96) | LDD0726 | [8] |
| LDCM0410 | CL22 | HCT 116 | C43(0.46); C408(0.71); C34(0.78); C72(1.15) | LDD0727 | [8] |
| LDCM0411 | CL23 | HCT 116 | C43(0.57); C408(0.72); C34(0.79); C72(1.04) | LDD0728 | [8] |
| LDCM0412 | CL24 | HCT 116 | C43(0.48); C408(0.65); C34(0.72); C72(0.99) | LDD0729 | [8] |
| LDCM0413 | CL25 | HCT 116 | C34(0.86); C408(0.64); C43(0.46); C72(0.97) | LDD0730 | [8] |
| LDCM0414 | CL26 | HCT 116 | C34(0.78); C408(0.72); C43(0.64); C72(0.98) | LDD0731 | [8] |
| LDCM0415 | CL27 | HCT 116 | C34(0.80); C408(0.74); C43(0.58); C72(1.06) | LDD0732 | [8] |
| LDCM0416 | CL28 | HCT 116 | C34(0.71); C408(0.69); C43(0.51); C72(0.98) | LDD0733 | [8] |
| LDCM0417 | CL29 | HCT 116 | C34(0.67); C408(0.77); C43(0.46); C72(1.00) | LDD0734 | [8] |
| LDCM0418 | CL3 | HCT 116 | C34(1.00); C408(1.10); C43(1.11); C72(1.10) | LDD0735 | [8] |
| LDCM0419 | CL30 | HCT 116 | C34(1.03); C408(0.83); C43(0.67); C72(1.14) | LDD0736 | [8] |
| LDCM0420 | CL31 | HCT 116 | C34(1.09); C408(0.75); C43(0.77); C72(0.85) | LDD0737 | [8] |
| LDCM0421 | CL32 | HCT 116 | C34(1.08); C408(0.59); C43(0.56); C72(0.88) | LDD0738 | [8] |
| LDCM0422 | CL33 | HCT 116 | C34(0.83); C408(0.61); C43(0.53); C72(0.79) | LDD0739 | [8] |
| LDCM0423 | CL34 | HCT 116 | C34(0.99); C408(0.51); C43(0.46); C72(0.82) | LDD0740 | [8] |
| LDCM0424 | CL35 | HCT 116 | C34(1.05); C408(0.61); C43(0.47); C72(0.82) | LDD0741 | [8] |
| LDCM0425 | CL36 | HCT 116 | C34(0.87); C408(0.60); C43(0.50); C72(0.82) | LDD0742 | [8] |
| LDCM0426 | CL37 | HCT 116 | C34(1.11); C408(0.60); C43(0.44); C72(0.78) | LDD0743 | [8] |
| LDCM0428 | CL39 | HCT 116 | C34(0.90); C408(0.60); C43(0.50); C72(0.84) | LDD0745 | [8] |
| LDCM0429 | CL4 | HCT 116 | C34(1.01); C408(1.00); C43(1.10); C72(0.96) | LDD0746 | [8] |
| LDCM0430 | CL40 | HCT 116 | C34(0.79); C408(0.72); C43(0.48); C72(0.83) | LDD0747 | [8] |
| LDCM0431 | CL41 | HCT 116 | C34(1.08); C408(0.63); C43(0.54); C72(0.86) | LDD0748 | [8] |
| LDCM0432 | CL42 | HCT 116 | C34(0.92); C408(0.65); C43(0.45); C72(0.86) | LDD0749 | [8] |
| LDCM0433 | CL43 | HCT 116 | C34(0.89); C408(0.55); C43(0.52); C72(0.85) | LDD0750 | [8] |
| LDCM0434 | CL44 | HCT 116 | C34(1.18); C408(0.67); C43(0.50); C72(0.82) | LDD0751 | [8] |
| LDCM0435 | CL45 | HCT 116 | C34(0.89); C408(0.59); C43(0.45); C72(0.81) | LDD0752 | [8] |
| LDCM0436 | CL46 | HCT 116 | C408(0.86); C72(1.11) | LDD0753 | [8] |
| LDCM0437 | CL47 | HCT 116 | C408(0.93); C72(1.18) | LDD0754 | [8] |
| LDCM0438 | CL48 | HCT 116 | C408(0.82); C72(1.11) | LDD0755 | [8] |
| LDCM0439 | CL49 | HCT 116 | C408(0.99); C72(1.13) | LDD0756 | [8] |
| LDCM0440 | CL5 | HCT 116 | C34(1.05); C408(1.11); C43(0.94); C72(1.09) | LDD0757 | [8] |
| LDCM0441 | CL50 | HCT 116 | C408(0.95); C72(1.03) | LDD0758 | [8] |
| LDCM0442 | CL51 | HCT 116 | C408(1.03); C72(1.09) | LDD0759 | [8] |
| LDCM0443 | CL52 | HCT 116 | C408(0.85); C72(1.01) | LDD0760 | [8] |
| LDCM0444 | CL53 | HCT 116 | C408(1.05); C72(1.02) | LDD0761 | [8] |
| LDCM0445 | CL54 | HCT 116 | C408(0.92); C72(1.04) | LDD0762 | [8] |
| LDCM0446 | CL55 | HCT 116 | C408(1.24); C72(1.11) | LDD0763 | [8] |
| LDCM0447 | CL56 | HCT 116 | C408(0.94); C72(1.08) | LDD0764 | [8] |
| LDCM0448 | CL57 | HCT 116 | C408(1.20); C72(1.06) | LDD0765 | [8] |
| LDCM0449 | CL58 | HCT 116 | C408(1.01); C72(1.06) | LDD0766 | [8] |
| LDCM0450 | CL59 | HCT 116 | C408(1.00); C72(1.08) | LDD0767 | [8] |
| LDCM0451 | CL6 | HCT 116 | C34(0.93); C408(1.00); C43(0.98); C72(0.94) | LDD0768 | [8] |
| LDCM0452 | CL60 | HCT 116 | C408(0.80); C72(1.05) | LDD0769 | [8] |
| LDCM0453 | CL61 | HCT 116 | C34(0.78); C408(0.95); C72(0.89) | LDD0770 | [8] |
| LDCM0454 | CL62 | HCT 116 | C34(0.99); C408(0.94); C72(0.91) | LDD0771 | [8] |
| LDCM0455 | CL63 | HCT 116 | C34(0.81); C408(0.74); C72(0.87) | LDD0772 | [8] |
| LDCM0456 | CL64 | HCT 116 | C34(0.69); C408(0.73); C72(0.81) | LDD0773 | [8] |
| LDCM0457 | CL65 | HCT 116 | C34(0.74); C408(0.83); C72(0.81) | LDD0774 | [8] |
| LDCM0458 | CL66 | HCT 116 | C34(0.53); C408(0.50); C72(0.78) | LDD0775 | [8] |
| LDCM0459 | CL67 | HCT 116 | C34(0.65); C408(0.65); C72(0.80) | LDD0776 | [8] |
| LDCM0460 | CL68 | HCT 116 | C34(0.60); C408(0.68); C72(0.78) | LDD0777 | [8] |
| LDCM0461 | CL69 | HCT 116 | C34(0.76); C408(0.87); C72(0.83) | LDD0778 | [8] |
| LDCM0462 | CL7 | HCT 116 | C34(1.12); C408(1.15); C43(1.00); C72(0.96) | LDD0779 | [8] |
| LDCM0463 | CL70 | HCT 116 | C34(0.65); C408(0.70); C72(0.86) | LDD0780 | [8] |
| LDCM0464 | CL71 | HCT 116 | C34(0.62); C408(0.70); C72(0.87) | LDD0781 | [8] |
| LDCM0465 | CL72 | HCT 116 | C34(0.70); C408(1.18); C72(0.94) | LDD0782 | [8] |
| LDCM0466 | CL73 | HCT 116 | C34(0.62); C408(0.82); C72(0.88) | LDD0783 | [8] |
| LDCM0467 | CL74 | HCT 116 | C34(0.70); C408(0.77); C72(0.84) | LDD0784 | [8] |
| LDCM0469 | CL76 | HCT 116 | C34(0.79); C408(0.81); C43(0.75); C72(0.91) | LDD0786 | [8] |
| LDCM0470 | CL77 | HCT 116 | C34(0.96); C408(0.97); C43(1.17); C72(1.02) | LDD0787 | [8] |
| LDCM0471 | CL78 | HCT 116 | C34(0.75); C408(0.73); C43(0.70); C72(0.92) | LDD0788 | [8] |
| LDCM0472 | CL79 | HCT 116 | C34(0.74); C408(0.71); C43(0.83); C72(0.85) | LDD0789 | [8] |
| LDCM0473 | CL8 | HCT 116 | C34(0.85); C408(1.12); C43(0.92); C72(0.95) | LDD0790 | [8] |
| LDCM0474 | CL80 | HCT 116 | C34(0.64); C408(0.86); C43(0.89); C72(0.95) | LDD0791 | [8] |
| LDCM0475 | CL81 | HCT 116 | C34(0.80); C408(0.81); C43(0.91); C72(0.93) | LDD0792 | [8] |
| LDCM0476 | CL82 | HCT 116 | C34(0.63); C408(0.92); C43(0.61); C72(0.91) | LDD0793 | [8] |
| LDCM0477 | CL83 | HCT 116 | C34(0.63); C408(0.69); C43(0.68); C72(0.87) | LDD0794 | [8] |
| LDCM0478 | CL84 | HCT 116 | C34(0.65); C408(0.59); C43(0.63); C72(0.82) | LDD0795 | [8] |
| LDCM0479 | CL85 | HCT 116 | C34(0.81); C408(0.75); C43(0.77); C72(0.90) | LDD0796 | [8] |
| LDCM0480 | CL86 | HCT 116 | C34(0.86); C408(0.91); C43(0.89); C72(0.95) | LDD0797 | [8] |
| LDCM0481 | CL87 | HCT 116 | C34(0.72); C408(0.90); C43(0.82); C72(0.93) | LDD0798 | [8] |
| LDCM0482 | CL88 | HCT 116 | C34(0.67); C408(0.79); C43(0.80); C72(0.86) | LDD0799 | [8] |
| LDCM0483 | CL89 | HCT 116 | C34(0.63); C408(0.62); C43(0.66); C72(0.86) | LDD0800 | [8] |
| LDCM0484 | CL9 | HCT 116 | C34(1.05); C408(0.99); C43(1.07); C72(1.05) | LDD0801 | [8] |
| LDCM0485 | CL90 | HCT 116 | C34(1.76); C408(1.36); C43(1.08); C72(0.95) | LDD0802 | [8] |
| LDCM0486 | CL91 | HCT 116 | C34(1.12); C408(0.83); C72(0.98) | LDD0803 | [8] |
| LDCM0487 | CL92 | HCT 116 | C34(1.10); C408(0.94); C72(1.00) | LDD0804 | [8] |
| LDCM0488 | CL93 | HCT 116 | C34(1.35); C408(0.81); C72(1.06) | LDD0805 | [8] |
| LDCM0489 | CL94 | HCT 116 | C34(1.10); C408(0.85); C72(0.90) | LDD0806 | [8] |
| LDCM0490 | CL95 | HCT 116 | C34(1.14); C408(0.74); C72(0.87) | LDD0807 | [8] |
| LDCM0491 | CL96 | HCT 116 | C34(1.24); C408(0.75); C72(0.89) | LDD0808 | [8] |
| LDCM0492 | CL97 | HCT 116 | C34(1.14); C408(0.67); C72(0.95) | LDD0809 | [8] |
| LDCM0493 | CL98 | HCT 116 | C34(1.08); C408(0.70); C72(0.94) | LDD0810 | [8] |
| LDCM0494 | CL99 | HCT 116 | C34(0.95); C408(0.70); C72(0.99) | LDD0811 | [8] |
| LDCM0495 | E2913 | HEK-293T | C408(1.16); C72(1.19); C43(0.96); C334(1.11) | LDD1698 | [27] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C408(7.00); C72(5.67) | LDD1702 | [4] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [7] |
| LDCM0625 | F8 | Ramos | C72(1.74); C43(0.57) | LDD2187 | [28] |
| LDCM0573 | Fragment11 | Ramos | C72(0.76); C43(8.73) | LDD2190 | [28] |
| LDCM0574 | Fragment12 | Ramos | C72(0.86) | LDD2191 | [28] |
| LDCM0576 | Fragment14 | Ramos | C72(2.23) | LDD2193 | [28] |
| LDCM0580 | Fragment21 | Ramos | C72(0.86) | LDD2195 | [28] |
| LDCM0582 | Fragment23 | Ramos | C72(0.78) | LDD2196 | [28] |
| LDCM0588 | Fragment30 | Ramos | C72(1.09) | LDD2199 | [28] |
| LDCM0468 | Fragment33 | HCT 116 | C34(0.66); C408(0.82); C72(0.89) | LDD0785 | [8] |
| LDCM0596 | Fragment38 | Ramos | C72(0.66) | LDD2203 | [28] |
| LDCM0566 | Fragment4 | Ramos | C72(1.92); C43(0.15) | LDD2184 | [28] |
| LDCM0427 | Fragment51 | HCT 116 | C34(0.98); C408(0.63); C43(0.43); C72(0.86) | LDD0744 | [8] |
| LDCM0614 | Fragment56 | Ramos | C72(1.34) | LDD2205 | [28] |
| LDCM0569 | Fragment7 | Ramos | C72(1.20) | LDD2186 | [28] |
| LDCM0107 | IAA | HeLa | H162(0.00); C334(0.00) | LDD0221 | [17] |
| LDCM0022 | KB02 | HCT 116 | C34(1.50); C72(1.09) | LDD0080 | [8] |
| LDCM0023 | KB03 | HCT 116 | C34(1.38); C72(1.04) | LDD0081 | [8] |
| LDCM0024 | KB05 | HCT 116 | C34(1.82); C72(1.00) | LDD0082 | [8] |
| LDCM0109 | NEM | HeLa | H162(0.00); H236(0.00) | LDD0223 | [17] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C34(0.99) | LDD2092 | [4] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C72(0.94) | LDD2093 | [4] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C34(0.97) | LDD2094 | [4] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C34(0.67) | LDD2096 | [4] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C72(0.71); C34(1.08) | LDD2098 | [4] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C72(0.98) | LDD2099 | [4] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C34(1.05) | LDD2100 | [4] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C34(1.11) | LDD2104 | [4] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C34(1.88) | LDD2105 | [4] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C72(1.12) | LDD2109 | [4] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C34(1.06) | LDD2111 | [4] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C72(0.74) | LDD2120 | [4] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C34(0.76) | LDD2122 | [4] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C72(1.02); C34(0.72) | LDD2123 | [4] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C72(0.66) | LDD2125 | [4] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C72(1.03) | LDD2127 | [4] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C72(0.80) | LDD2128 | [4] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C72(1.02) | LDD2129 | [4] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C72(1.57) | LDD2135 | [4] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C72(1.19) | LDD2136 | [4] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C72(0.97) | LDD2137 | [4] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C72(1.71) | LDD1700 | [4] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C72(0.77); C34(0.69) | LDD2140 | [4] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C34(0.85) | LDD2141 | [4] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C72(1.06) | LDD2143 | [4] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C72(2.77) | LDD2144 | [4] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C72(1.04) | LDD2146 | [4] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C72(1.89) | LDD2147 | [4] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C72(0.82) | LDD2148 | [4] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C72(0.41); C34(0.60) | LDD2150 | [4] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C20(1.03); C72(0.65) | LDD2206 | [29] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C72(1.01); C20(0.89); C72(0.76) | LDD2207 | [29] |
| LDCM0014 | Panhematin | HEK-293T | 3.04 | LDD0062 | [6] |
The Interaction Atlas With This Target
References
