Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
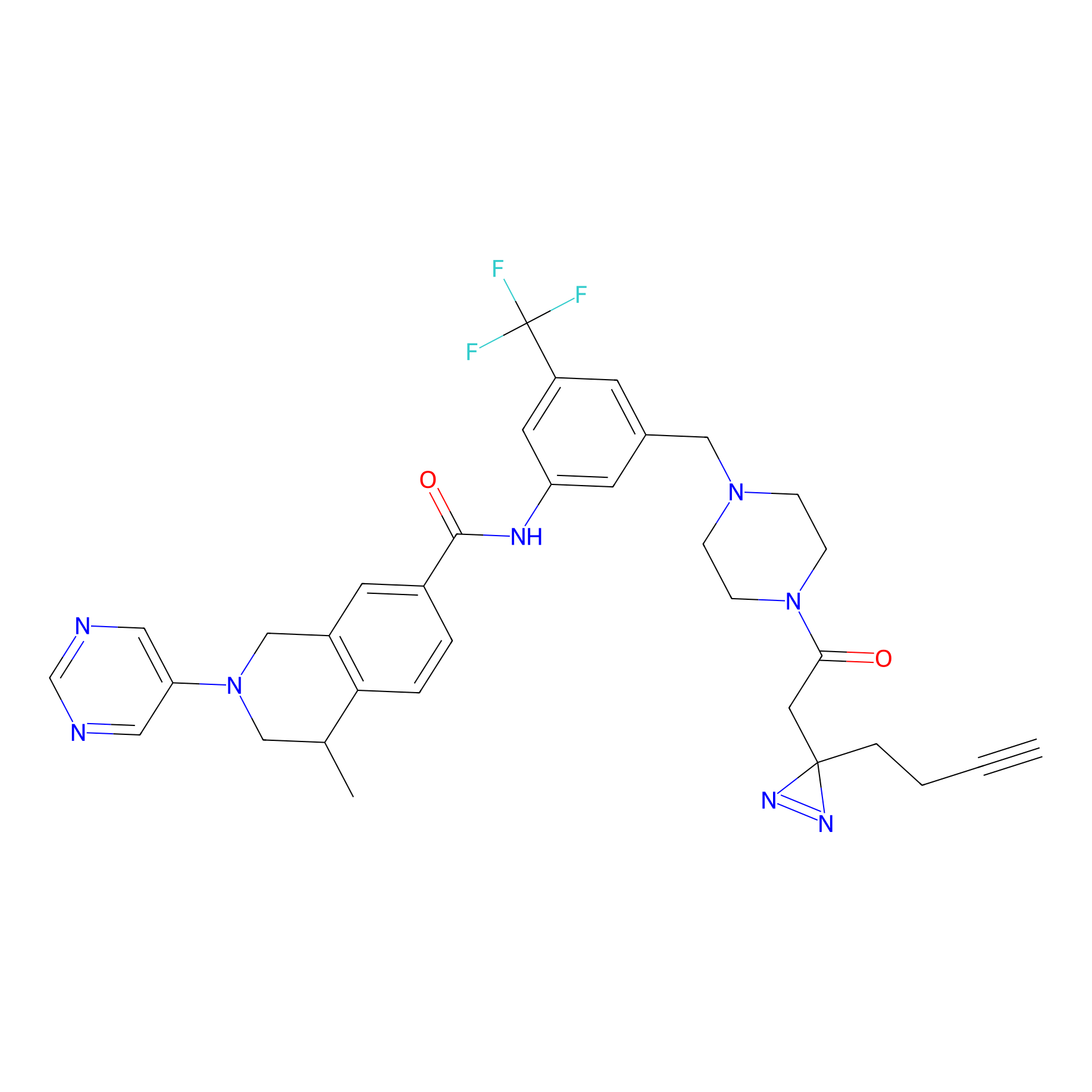
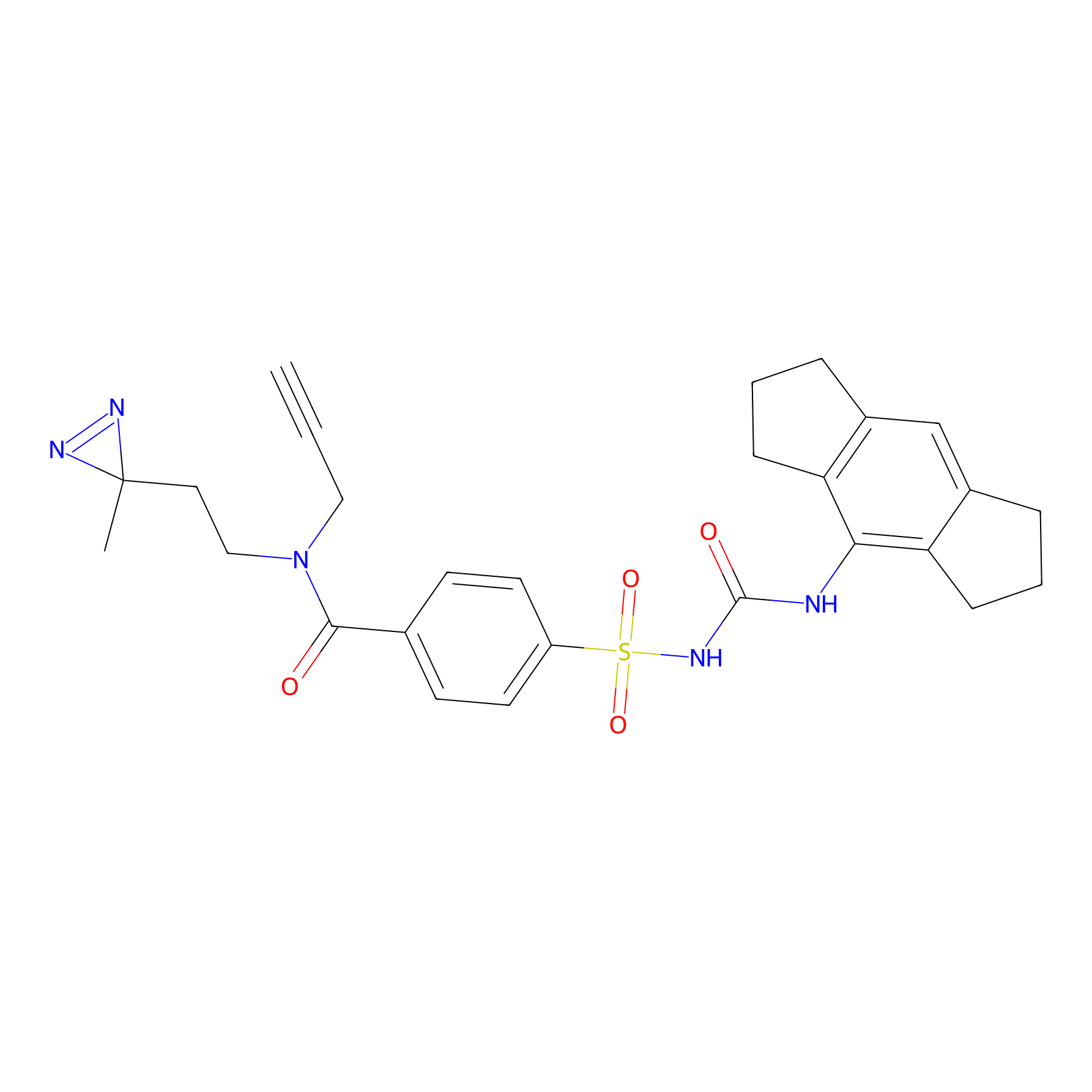
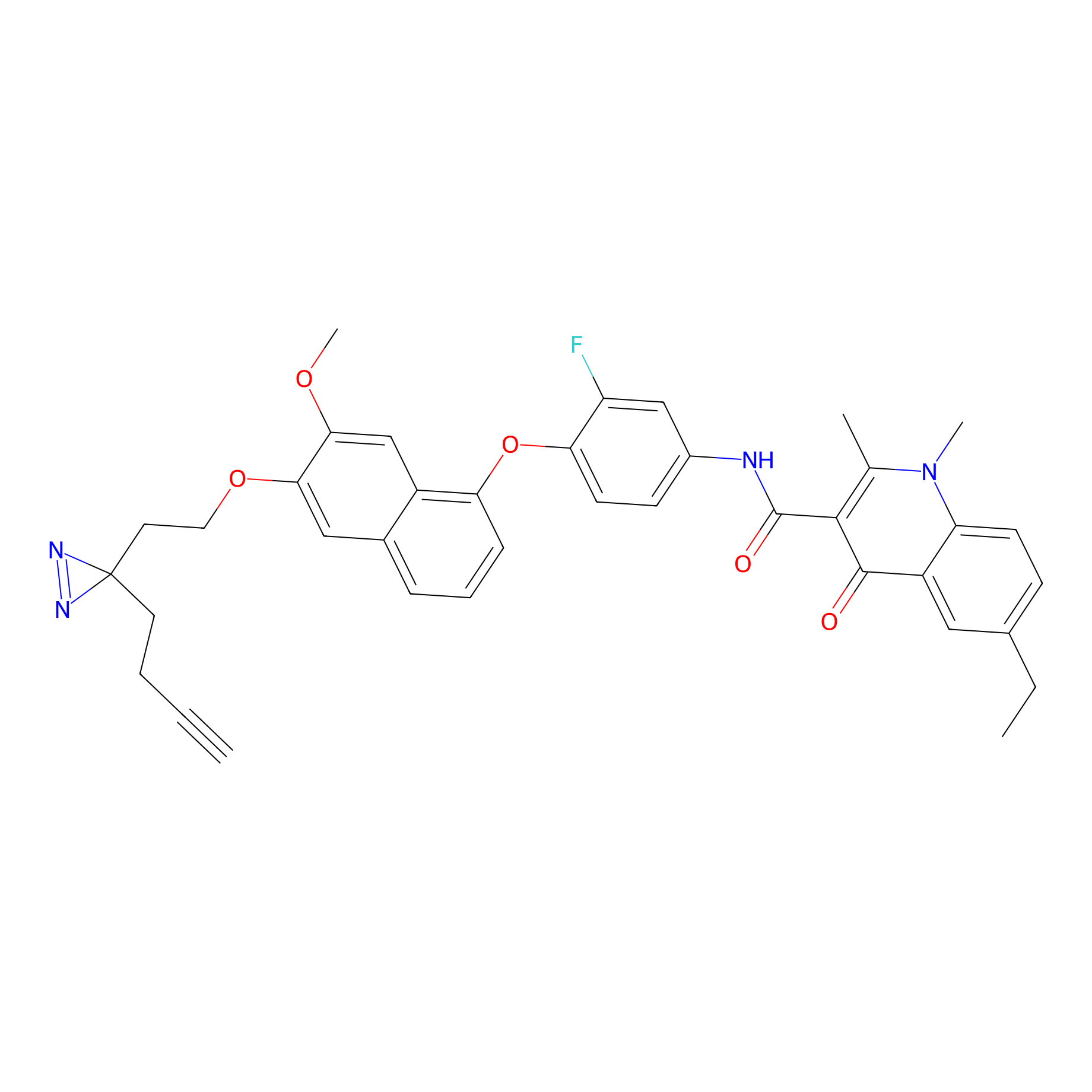
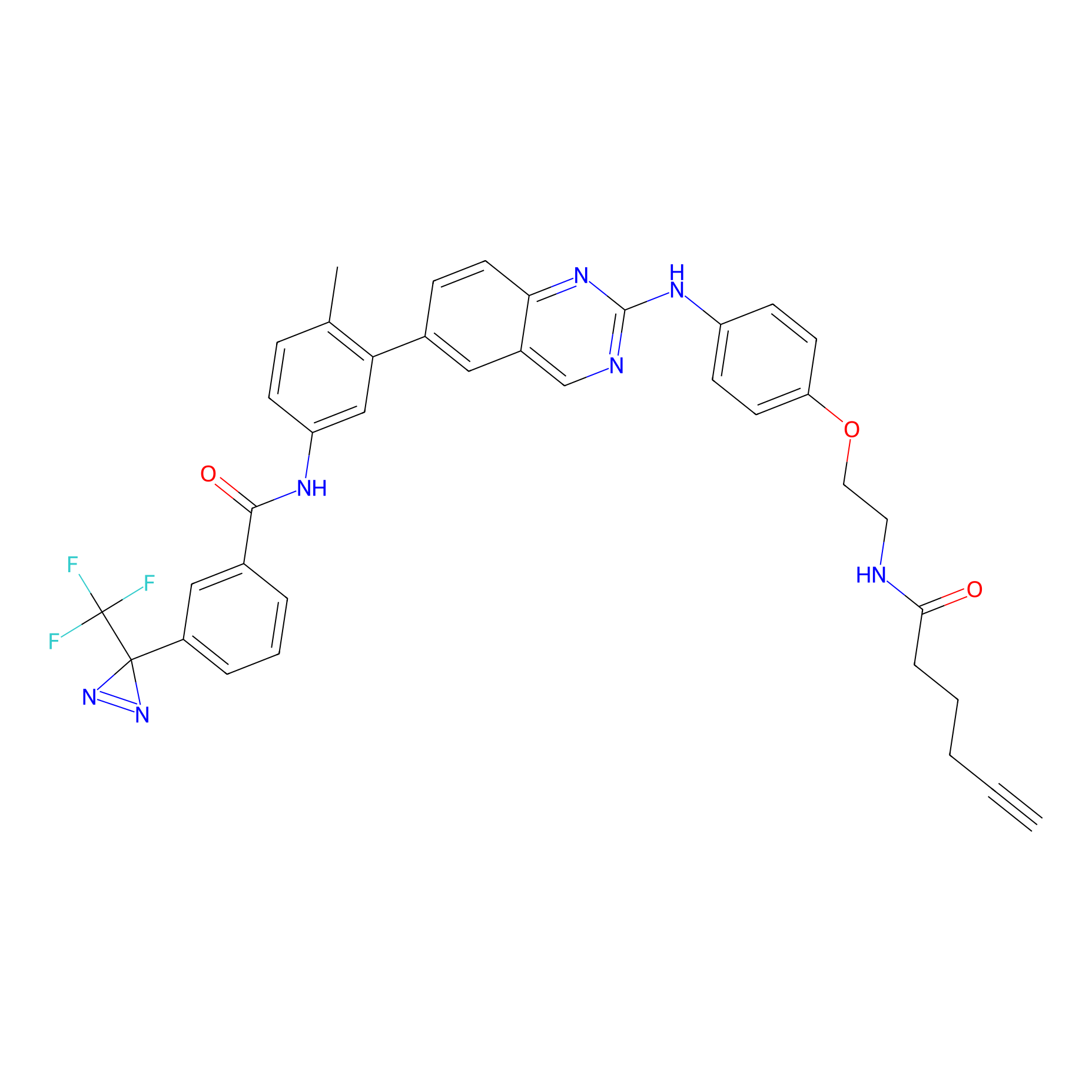
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
HDSF-alk Probe Info |
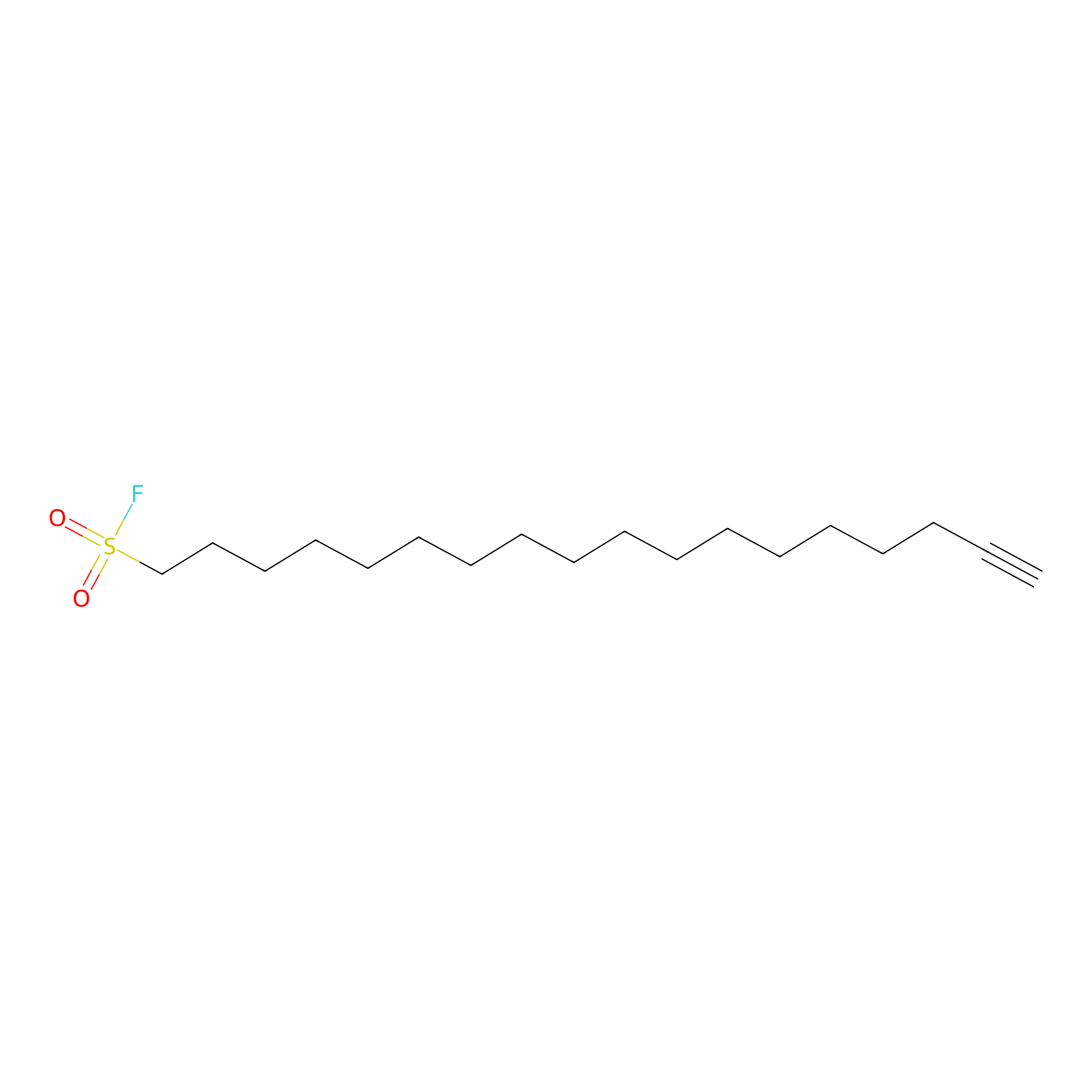 |
2.02 | LDD0197 | [1] | |
|
P8 Probe Info |
 |
4.25 | LDD0451 | [2] | |
|
CHEMBL5175495 Probe Info |
 |
9.49 | LDD0196 | [3] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [4] | |
|
TG42 Probe Info |
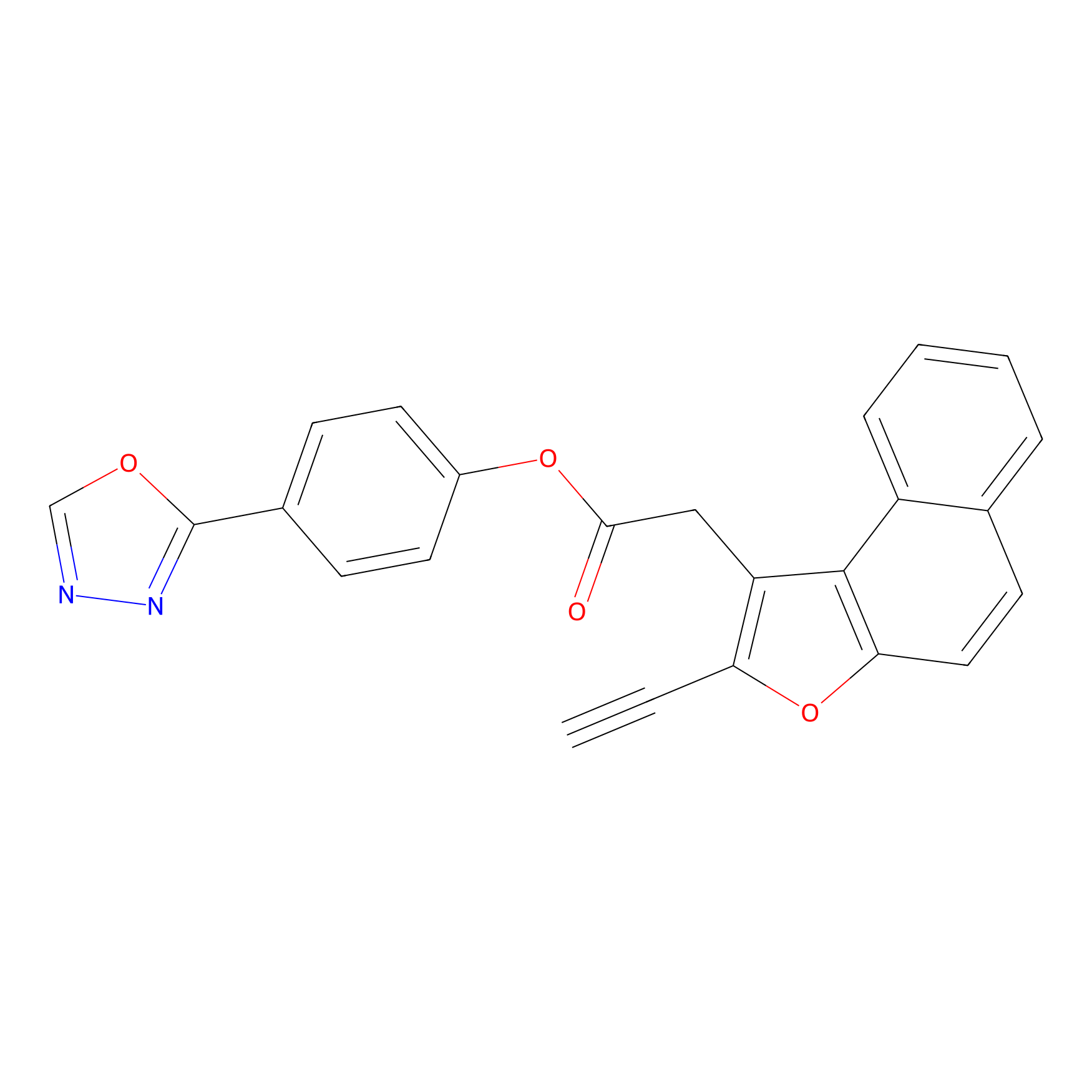 |
10.58 | LDD0326 | [5] | |
|
TH211 Probe Info |
 |
Y73(8.00) | LDD0260 | [6] | |
|
C-Sul Probe Info |
 |
6.24 | LDD0066 | [7] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [8] | |
|
ONAyne Probe Info |
 |
K138(1.04); K159(5.00) | LDD0274 | [9] | |
|
OPA-S-S-alkyne Probe Info |
 |
K176(0.75); K204(1.82) | LDD3494 | [10] | |
|
Probe 1 Probe Info |
 |
Y150(59.44) | LDD3495 | [11] | |
|
m-APA Probe Info |
 |
11.35 | LDD0403 | [12] | |
|
Jackson_1 Probe Info |
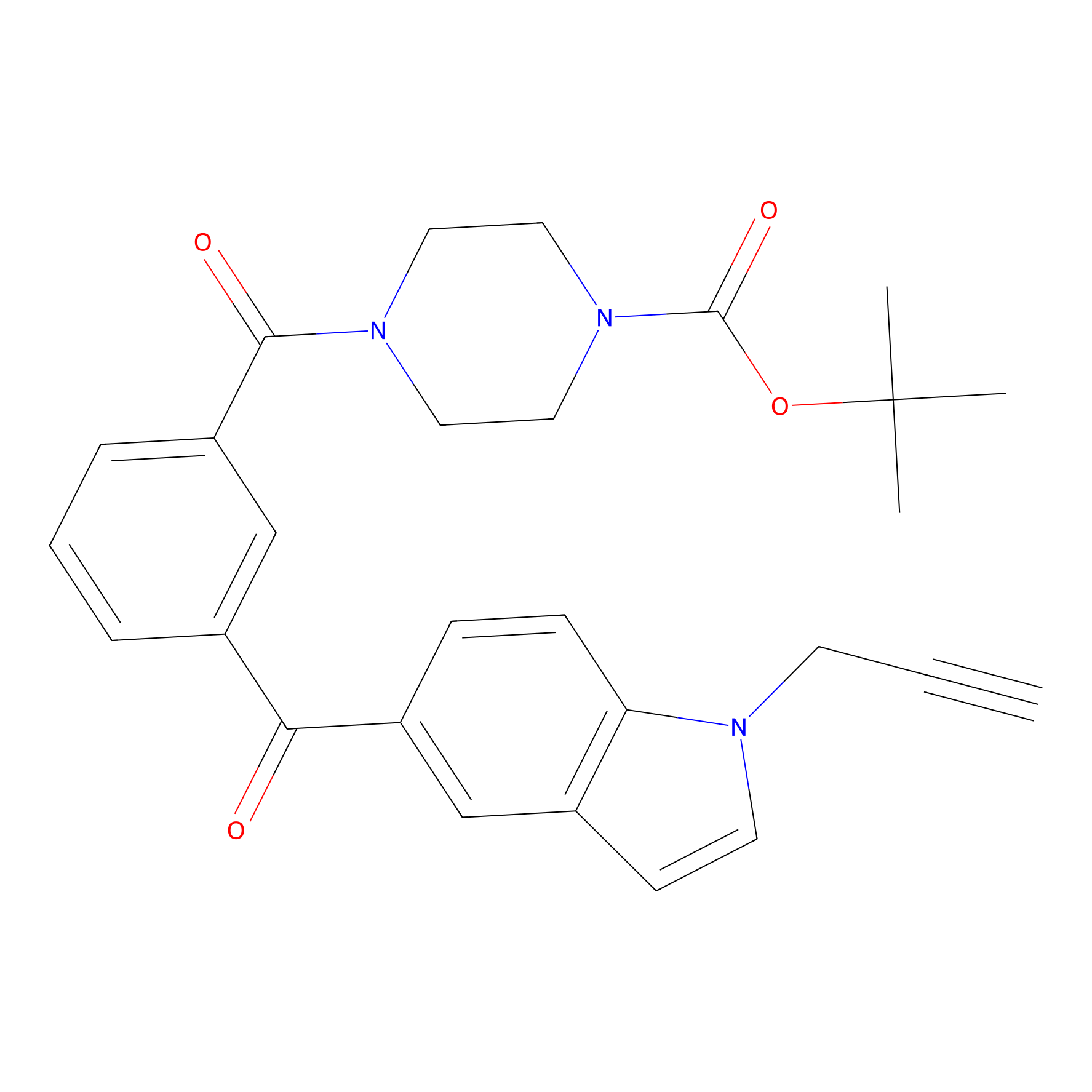 |
10.61 | LDD0120 | [13] | |
|
HPAP Probe Info |
 |
3.78 | LDD0064 | [14] | |
|
ATP probe Probe Info |
 |
K95(0.00); K204(0.00); K72(0.00); K214(0.00) | LDD0199 | [15] | |
|
ATP probe Probe Info |
 |
K95(0.00); K221(0.00); K192(0.00) | LDD0035 | [16] | |
|
NHS Probe Info |
 |
K201(0.00); K221(0.00); K176(0.00); K167(0.00) | LDD0010 | [17] | |
|
SF Probe Info |
 |
Y150(0.00); K192(0.00); K190(0.00) | LDD0028 | [18] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [17] | |
|
1c-yne Probe Info |
 |
K149(0.00); K167(0.00); K190(0.00) | LDD0228 | [19] | |
|
Acrolein Probe Info |
 |
H93(0.00); H230(0.00) | LDD0217 | [20] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [20] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0176 | 9im | MDA-MB-231 | 3.74 | LDD0441 | [27] |
| LDCM0156 | Aniline | NCI-H1299 | 11.35 | LDD0403 | [12] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [20] |
| LDCM0070 | Cisar_cp37 | MDA-MB-231 | 10.61 | LDD0120 | [13] |
| LDCM0017 | DFG-out-2 | A431 | 4.60 | LDD0075 | [28] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [20] |
| LDCM0109 | NEM | HeLa | H93(0.00); H230(0.00) | LDD0223 | [20] |
| LDCM0014 | Panhematin | hPBMC | 3.78 | LDD0064 | [14] |
| LDCM0019 | Staurosporine | Hep-G2 | 2.00 | LDD0083 | [29] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Mitochondrial fission 1 protein (FIS1) | FIS1 family | Q9Y3D6 | |||
| Complex I assembly factor TIMMDC1, mitochondrial (TIMMDC1) | Tim17/Tim22/Tim23 family | Q9NPL8 | |||
References