Details of the Target
General Information of Target
| Target ID | LDTP06633 | |||||
|---|---|---|---|---|---|---|
| Target Name | Reticulon-1 (RTN1) | |||||
| Gene Name | RTN1 | |||||
| Gene ID | 6252 | |||||
| Synonyms |
NSP; Reticulon-1; Neuroendocrine-specific protein |
|||||
| 3D Structure | ||||||
| Sequence |
MAAPGDPQDELLPLAGPGSQWLRHRGEGENEAVTPKGATPAPQAGEPSPGLGARAREAAS
REAGSGPARQSPVAMETASTGVAGVSSAMDHTFSTTSKDGEGSCYTSLISDICYPPQEDS TYFTGILQKENGHVTISESPEELGTPGPSLPDVPGIESRGLFSSDSGIEMTPAESTEVNK ILADPLDQMKAEAYKYIDITRPEEVKHQEQHHPELEDKDLDFKNKDTDISIKPEGVREPD KPAPVEGKIIKDHLLEESTFAPYIDDLSEEQRRAPQITTPVKITLTEIEPSVETTTQEKT PEKQDICLKPSPDTVPTVTVSEPEDDSPGSITPPSSGTEPSAAESQGKGSISEDELITAI KEAKGLSYETAENPRPVGQLADRPEVKARSGPPTIPSPLDHEASSAESGDSEIELVSEDP MAAEDALPSGYVSFGHVGGPPPSPASPSIQYSILREEREAELDSELIIESCDASSASEES PKREQDSPPMKPSALDAIREETGVRAEERAPSRRGLAEPGSFLDYPSTEPQPGPELPPGD GALEPETPMLPRKPEEDSSSNQSPAATKGPGPLGPGAPPPLLFLNKQKAIDLLYWRDIKQ TGIVFGSFLLLLFSLTQFSVVSVVAYLALAALSATISFRIYKSVLQAVQKTDEGHPFKAY LELEITLSQEQIQKYTDCLQFYVNSTLKELRRLFLVQDLVDSLKFAVLMWLLTYVGALFN GLTLLLMAVVSMFTLPVVYVKHQAQIDQYLGLVRTHINAVVAKIQAKIPGAKRHAE |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function | Inhibits amyloid precursor protein processing, probably by blocking BACE1 activity. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| A673 | SNV: p.E362Q | . | |||
| DU145 | SNV: p.G66S; p.V761E | . | |||
| HCT116 | SNV: p.A510V | . | |||
| HEC1 | SNV: p.H253Y | . | |||
| IGR1 | SNV: p.L285P | . | |||
| IPC298 | SNV: p.G391R | . | |||
| MFE296 | Deletion: p.E27RfsTer7 | . | |||
| NCIH358 | SNV: p.P148H | . | |||
| OVK18 | Deletion: p.L399WfsTer34 | . | |||
| PC9 | SNV: p.P538H | . | |||
| SF295 | SNV: p.T635A | . | |||
| SHP77 | Substitution: p.P280S | DBIA Probe Info | |||
| SKOV3 | SNV: p.G37E | . | |||
| TE4 | SNV: p.R499L | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
FBPP2 Probe Info |
 |
3.25 | LDD0318 | [2] | |
|
DA-P2 Probe Info |
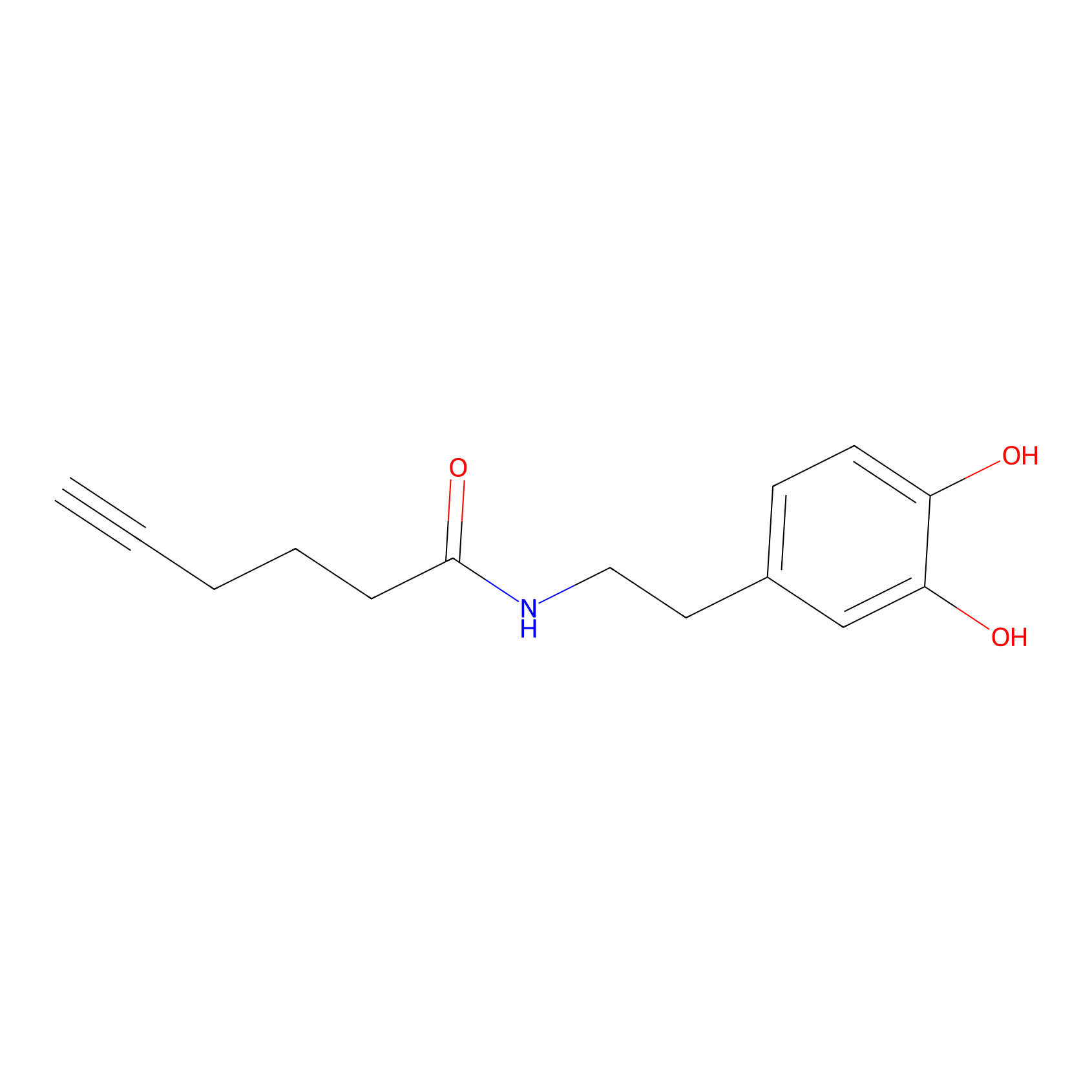 |
52.30 | LDD0348 | [3] | |
|
HRP Probe Info |
 |
35.42 | LDD0347 | [3] | |
|
SAA-alkyne Probe Info |
 |
1.10 | LDD0252 | [4] | |
|
FBP2 Probe Info |
 |
4.52 | LDD0317 | [2] | |
|
DBIA Probe Info |
 |
C678(4.56) | LDD3344 | [5] | |
|
Sulforaphane-probe2 Probe Info |
 |
1.58 | LDD0160 | [6] | |
|
DA-P3 Probe Info |
 |
6.84 | LDD0180 | [3] | |
|
HPAP Probe Info |
 |
5.14 | LDD0064 | [7] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [8] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 12.37 | LDD0403 | [1] |
| LDCM0087 | Capsaicin | HEK-293T | 23.11 | LDD0185 | [3] |
| LDCM0028 | Dobutamine | HEK-293T | 6.84 | LDD0180 | [3] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 10.17 | LDD0183 | [3] |
| LDCM0022 | KB02 | BRX211 | C678(5.64) | LDD2268 | [5] |
| LDCM0023 | KB03 | BRX211 | C678(12.42) | LDD2685 | [5] |
| LDCM0024 | KB05 | NCI-H146 | C678(4.56) | LDD3344 | [5] |
| LDCM0032 | Oleacein | HEK-293T | 4.67 | LDD0184 | [3] |
| LDCM0014 | Panhematin | hPBMC | 5.14 | LDD0064 | [7] |
| LDCM0029 | Quercetin | HEK-293T | 21.12 | LDD0181 | [3] |
| LDCM0003 | Sulforaphane | MDA-MB-231 | 1.58 | LDD0160 | [6] |
The Interaction Atlas With This Target
References