Details of the Target
General Information of Target
| Target ID | LDTP06293 | |||||
|---|---|---|---|---|---|---|
| Target Name | Metal cation symporter ZIP14 (SLC39A14) | |||||
| Gene Name | SLC39A14 | |||||
| Gene ID | 23516 | |||||
| Synonyms |
KIAA0062; ZIP14; Metal cation symporter ZIP14; LIV-1 subfamily of ZIP zinc transporter 4; LZT-Hs4; Solute carrier family 39 member 14; Zrt- and Irt-like protein 14; ZIP-14 |
|||||
| 3D Structure | ||||||
| Sequence |
MKLLLLHPAFQSCLLLTLLGLWRTTPEAHASSLGAPAISAASFLQDLIHRYGEGDSLTLQ
QLKALLNHLDVGVGRGNVTQHVQGHRNLSTCFSSGDLFTAHNFSEQSRIGSSELQEFCPT ILQQLDSRACTSENQENEENEQTEEGRPSAVEVWGYGLLCVTVISLCSLLGASVVPFMKK TFYKRLLLYFIALAIGTLYSNALFQLIPEAFGFNPLEDYYVSKSAVVFGGFYLFFFTEKI LKILLKQKNEHHHGHSHYASESLPSKKDQEEGVMEKLQNGDLDHMIPQHCSSELDGKAPM VDEKVIVGSLSVQDLQASQSACYWLKGVRYSDIGTLAWMITLSDGLHNFIDGLAIGASFT VSVFQGISTSVAILCEEFPHELGDFVILLNAGMSIQQALFFNFLSACCCYLGLAFGILAG SHFSANWIFALAGGMFLYISLADMFPEMNEVCQEDERKGSILIPFIIQNLGLLTGFTIMV VLTMYSGQIQIG |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
ZIP transporter (TC 2.A.5) family
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
Electroneutral transporter of the plasma membrane mediating the cellular uptake of the divalent metal cations zinc, manganese and iron that are important for tissue homeostasis, metabolism, development and immunity. Functions as an energy-dependent symporter, transporting through the membranes an electroneutral complex composed of a divalent metal cation and two bicarbonate anions. Beside these endogenous cellular substrates, can also import cadmium a non-essential metal which is cytotoxic and carcinogenic. Controls the cellular uptake by the intestinal epithelium of systemic zinc, which is in turn required to maintain tight junctions and the intestinal permeability. Modifies the activity of zinc-dependent phosphodiesterases, thereby indirectly regulating G protein-coupled receptor signaling pathways important for gluconeogenesis and chondrocyte differentiation. Regulates insulin receptor signaling, glucose uptake, glycogen synthesis and gluconeogenesis in hepatocytes through the zinc-dependent intracellular catabolism of insulin. Through zinc cellular uptake also plays a role in the adaptation of cells to endoplasmic reticulum stress. Major manganese transporter of the basolateral membrane of intestinal epithelial cells, it plays a central role in manganese systemic homeostasis through intestinal manganese uptake. Also involved in manganese extracellular uptake by cells of the blood-brain barrier. May also play a role in manganese and zinc homeostasis participating in their elimination from the blood through the hepatobiliary excretion. Also functions in the extracellular uptake of free iron. May also function intracellularly and mediate the transport from endosomes to cytosol of iron endocytosed by transferrin. Plays a role in innate immunity by regulating the expression of cytokines by activated macrophages.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
7.19 | LDD0402 | [1] | |
|
AZ-5 Probe Info |
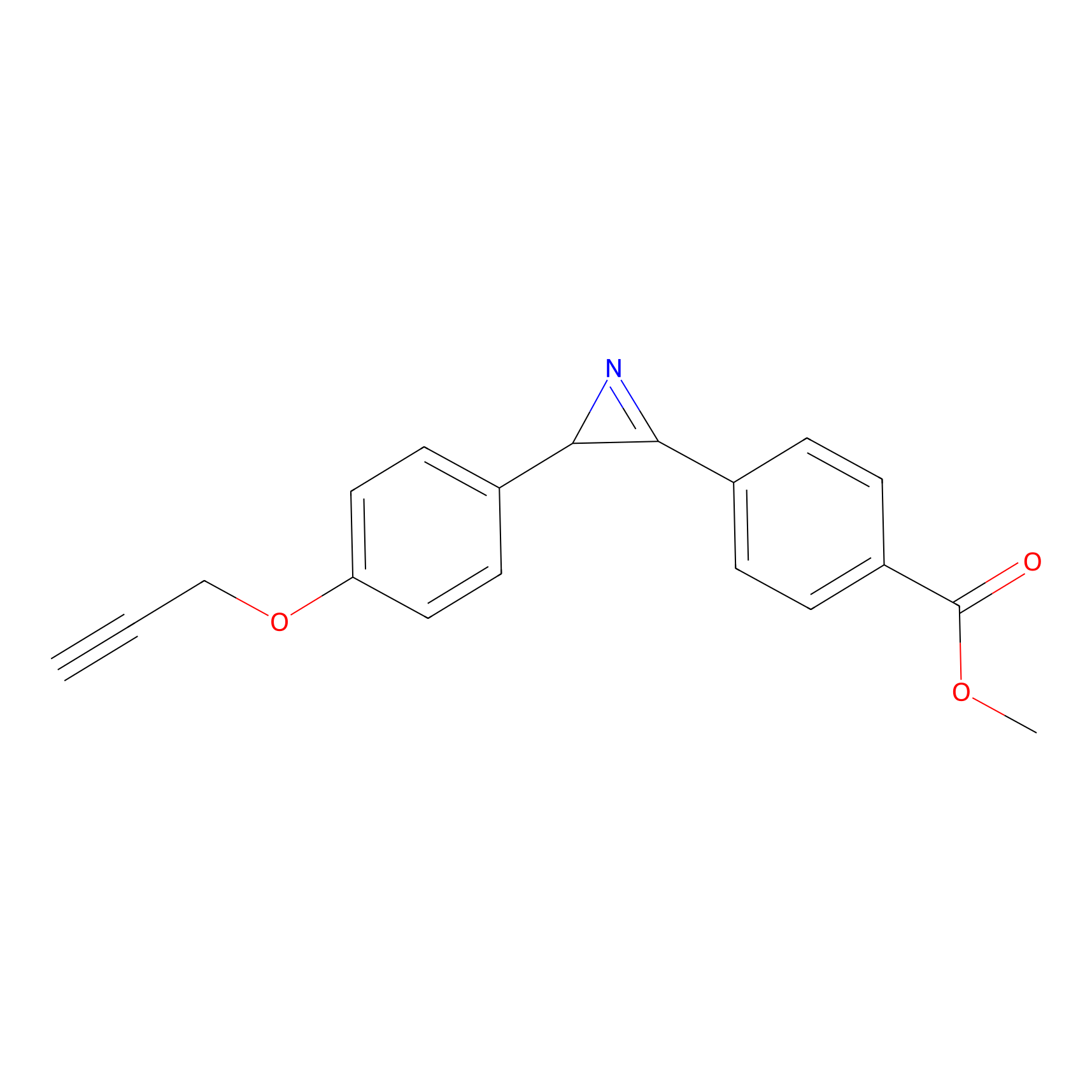 |
10.00 | LDD0394 | [2] | |
|
IPM Probe Info |
 |
N.A. | LDD0241 | [3] | |
|
Johansson_61 Probe Info |
 |
_(20.00) | LDD1485 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C290(5.39) | LDD0171 | [5] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0226 | [6] | |
|
DBIA Probe Info |
 |
C322(8.04) | LDD0080 | [7] | |
|
CY-1 Probe Info |
 |
N.A. | LDD0246 | [8] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [9] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [9] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [9] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [10] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [3] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C004 Probe Info |
 |
6.68 | LDD1714 | [12] | |
|
C022 Probe Info |
 |
8.63 | LDD1728 | [12] | |
|
C053 Probe Info |
 |
5.78 | LDD1751 | [12] | |
|
C091 Probe Info |
 |
10.48 | LDD1782 | [12] | |
|
C092 Probe Info |
 |
18.13 | LDD1783 | [12] | |
|
C106 Probe Info |
 |
16.91 | LDD1793 | [12] | |
|
C108 Probe Info |
 |
7.94 | LDD1795 | [12] | |
|
C112 Probe Info |
 |
24.76 | LDD1799 | [12] | |
|
C134 Probe Info |
 |
20.53 | LDD1816 | [12] | |
|
C143 Probe Info |
 |
12.38 | LDD1825 | [12] | |
|
C153 Probe Info |
 |
14.62 | LDD1834 | [12] | |
|
C160 Probe Info |
 |
13.09 | LDD1840 | [12] | |
|
C161 Probe Info |
 |
12.47 | LDD1841 | [12] | |
|
C170 Probe Info |
 |
8.46 | LDD1850 | [12] | |
|
C201 Probe Info |
 |
34.78 | LDD1877 | [12] | |
|
C208 Probe Info |
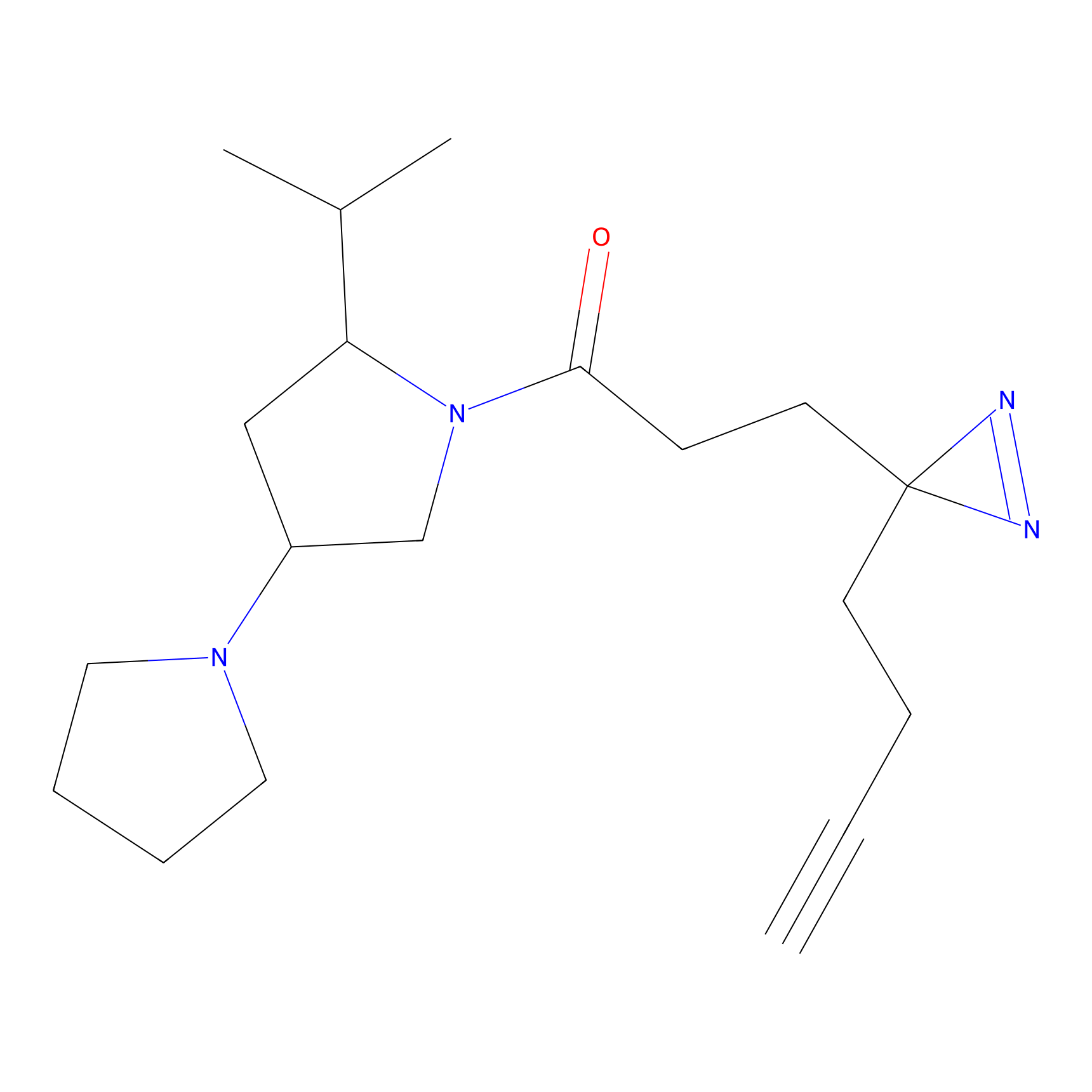 |
5.06 | LDD1883 | [12] | |
|
C218 Probe Info |
 |
13.93 | LDD1892 | [12] | |
|
C219 Probe Info |
 |
8.69 | LDD1893 | [12] | |
|
C220 Probe Info |
 |
18.38 | LDD1894 | [12] | |
|
C225 Probe Info |
 |
5.28 | LDD1898 | [12] | |
|
C228 Probe Info |
 |
15.67 | LDD1901 | [12] | |
|
C310 Probe Info |
 |
7.62 | LDD1977 | [12] | |
|
C349 Probe Info |
 |
9.06 | LDD2010 | [12] | |
|
C350 Probe Info |
 |
24.93 | LDD2011 | [12] | |
|
C356 Probe Info |
 |
9.92 | LDD2017 | [12] | |
|
C390 Probe Info |
 |
23.26 | LDD2049 | [12] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C290(5.39) | LDD0171 | [5] |
| LDCM0214 | AC1 | HCT 116 | C322(1.08) | LDD0531 | [7] |
| LDCM0215 | AC10 | HCT 116 | C322(1.03) | LDD0532 | [7] |
| LDCM0216 | AC100 | HCT 116 | C322(0.90) | LDD0533 | [7] |
| LDCM0217 | AC101 | HCT 116 | C322(0.81) | LDD0534 | [7] |
| LDCM0218 | AC102 | HCT 116 | C322(0.87) | LDD0535 | [7] |
| LDCM0219 | AC103 | HCT 116 | C322(1.11) | LDD0536 | [7] |
| LDCM0220 | AC104 | HCT 116 | C322(1.14) | LDD0537 | [7] |
| LDCM0221 | AC105 | HCT 116 | C322(0.90) | LDD0538 | [7] |
| LDCM0222 | AC106 | HCT 116 | C322(0.87) | LDD0539 | [7] |
| LDCM0223 | AC107 | HCT 116 | C322(0.54) | LDD0540 | [7] |
| LDCM0224 | AC108 | HCT 116 | C322(1.28) | LDD0541 | [7] |
| LDCM0225 | AC109 | HCT 116 | C322(1.13) | LDD0542 | [7] |
| LDCM0226 | AC11 | HCT 116 | C322(1.00) | LDD0543 | [7] |
| LDCM0227 | AC110 | HCT 116 | C322(1.47) | LDD0544 | [7] |
| LDCM0228 | AC111 | HCT 116 | C322(1.42) | LDD0545 | [7] |
| LDCM0229 | AC112 | HCT 116 | C322(1.06) | LDD0546 | [7] |
| LDCM0230 | AC113 | HCT 116 | C322(0.94) | LDD0547 | [7] |
| LDCM0231 | AC114 | HCT 116 | C322(1.09) | LDD0548 | [7] |
| LDCM0232 | AC115 | HCT 116 | C322(1.17) | LDD0549 | [7] |
| LDCM0233 | AC116 | HCT 116 | C322(1.13) | LDD0550 | [7] |
| LDCM0234 | AC117 | HCT 116 | C322(1.06) | LDD0551 | [7] |
| LDCM0235 | AC118 | HCT 116 | C322(0.99) | LDD0552 | [7] |
| LDCM0236 | AC119 | HCT 116 | C322(1.03) | LDD0553 | [7] |
| LDCM0237 | AC12 | HCT 116 | C322(0.99) | LDD0554 | [7] |
| LDCM0238 | AC120 | HCT 116 | C322(1.04) | LDD0555 | [7] |
| LDCM0239 | AC121 | HCT 116 | C322(0.89) | LDD0556 | [7] |
| LDCM0240 | AC122 | HCT 116 | C322(0.86) | LDD0557 | [7] |
| LDCM0241 | AC123 | HCT 116 | C322(0.90) | LDD0558 | [7] |
| LDCM0242 | AC124 | HCT 116 | C322(0.91) | LDD0559 | [7] |
| LDCM0243 | AC125 | HCT 116 | C322(1.07) | LDD0560 | [7] |
| LDCM0244 | AC126 | HCT 116 | C322(1.24) | LDD0561 | [7] |
| LDCM0245 | AC127 | HCT 116 | C322(1.09) | LDD0562 | [7] |
| LDCM0246 | AC128 | HEK-293T | C322(0.95) | LDD0844 | [7] |
| LDCM0247 | AC129 | HEK-293T | C322(0.94) | LDD0845 | [7] |
| LDCM0249 | AC130 | HEK-293T | C322(1.14) | LDD0847 | [7] |
| LDCM0250 | AC131 | HEK-293T | C322(0.96) | LDD0848 | [7] |
| LDCM0251 | AC132 | HEK-293T | C322(1.16) | LDD0849 | [7] |
| LDCM0252 | AC133 | HEK-293T | C322(1.36) | LDD0850 | [7] |
| LDCM0253 | AC134 | HEK-293T | C322(1.23) | LDD0851 | [7] |
| LDCM0254 | AC135 | HEK-293T | C322(1.20) | LDD0852 | [7] |
| LDCM0255 | AC136 | HEK-293T | C322(1.07) | LDD0853 | [7] |
| LDCM0256 | AC137 | HEK-293T | C322(1.42) | LDD0854 | [7] |
| LDCM0257 | AC138 | HEK-293T | C322(1.43) | LDD0855 | [7] |
| LDCM0258 | AC139 | HEK-293T | C322(1.26) | LDD0856 | [7] |
| LDCM0259 | AC14 | HCT 116 | C322(0.85) | LDD0576 | [7] |
| LDCM0260 | AC140 | HEK-293T | C322(1.26) | LDD0858 | [7] |
| LDCM0261 | AC141 | HEK-293T | C322(1.44) | LDD0859 | [7] |
| LDCM0262 | AC142 | HEK-293T | C322(1.04) | LDD0860 | [7] |
| LDCM0263 | AC143 | HCT 116 | C322(1.08) | LDD0580 | [7] |
| LDCM0264 | AC144 | HCT 116 | C322(1.45) | LDD0581 | [7] |
| LDCM0265 | AC145 | HCT 116 | C322(1.21) | LDD0582 | [7] |
| LDCM0266 | AC146 | HCT 116 | C322(1.22) | LDD0583 | [7] |
| LDCM0267 | AC147 | HCT 116 | C322(1.34) | LDD0584 | [7] |
| LDCM0268 | AC148 | HCT 116 | C322(1.81) | LDD0585 | [7] |
| LDCM0269 | AC149 | HCT 116 | C322(1.54) | LDD0586 | [7] |
| LDCM0270 | AC15 | HCT 116 | C322(0.98) | LDD0587 | [7] |
| LDCM0271 | AC150 | HCT 116 | C322(1.04) | LDD0588 | [7] |
| LDCM0272 | AC151 | HCT 116 | C322(1.07) | LDD0589 | [7] |
| LDCM0273 | AC152 | HCT 116 | C322(1.50) | LDD0590 | [7] |
| LDCM0274 | AC153 | HCT 116 | C322(2.05) | LDD0591 | [7] |
| LDCM0621 | AC154 | HCT 116 | C322(1.28) | LDD2158 | [7] |
| LDCM0622 | AC155 | HCT 116 | C322(1.28) | LDD2159 | [7] |
| LDCM0623 | AC156 | HCT 116 | C322(1.11) | LDD2160 | [7] |
| LDCM0624 | AC157 | HCT 116 | C322(1.37) | LDD2161 | [7] |
| LDCM0276 | AC17 | HCT 116 | C322(1.20) | LDD0593 | [7] |
| LDCM0277 | AC18 | HCT 116 | C322(1.27) | LDD0594 | [7] |
| LDCM0278 | AC19 | HCT 116 | C322(1.90) | LDD0595 | [7] |
| LDCM0279 | AC2 | HCT 116 | C322(0.97) | LDD0596 | [7] |
| LDCM0280 | AC20 | HCT 116 | C322(1.56) | LDD0597 | [7] |
| LDCM0281 | AC21 | HCT 116 | C322(1.04) | LDD0598 | [7] |
| LDCM0282 | AC22 | HCT 116 | C322(1.07) | LDD0599 | [7] |
| LDCM0283 | AC23 | HCT 116 | C322(1.53) | LDD0600 | [7] |
| LDCM0284 | AC24 | HCT 116 | C322(1.18) | LDD0601 | [7] |
| LDCM0285 | AC25 | PaTu 8988t | C322(0.96) | LDD1164 | [7] |
| LDCM0286 | AC26 | PaTu 8988t | C322(0.94) | LDD1165 | [7] |
| LDCM0287 | AC27 | PaTu 8988t | C322(1.24) | LDD1166 | [7] |
| LDCM0288 | AC28 | PaTu 8988t | C322(1.37) | LDD1167 | [7] |
| LDCM0289 | AC29 | PaTu 8988t | C322(1.10) | LDD1168 | [7] |
| LDCM0290 | AC3 | HCT 116 | C322(0.88) | LDD0607 | [7] |
| LDCM0291 | AC30 | PaTu 8988t | C322(1.06) | LDD1170 | [7] |
| LDCM0292 | AC31 | PaTu 8988t | C322(1.07) | LDD1171 | [7] |
| LDCM0293 | AC32 | PaTu 8988t | C322(1.00) | LDD1172 | [7] |
| LDCM0294 | AC33 | PaTu 8988t | C322(1.55) | LDD1173 | [7] |
| LDCM0295 | AC34 | PaTu 8988t | C322(1.26) | LDD1174 | [7] |
| LDCM0296 | AC35 | HCT 116 | C322(0.92) | LDD0613 | [7] |
| LDCM0297 | AC36 | HCT 116 | C322(0.71) | LDD0614 | [7] |
| LDCM0298 | AC37 | HCT 116 | C322(0.90) | LDD0615 | [7] |
| LDCM0299 | AC38 | HCT 116 | C322(0.95) | LDD0616 | [7] |
| LDCM0300 | AC39 | HCT 116 | C322(0.81) | LDD0617 | [7] |
| LDCM0301 | AC4 | HCT 116 | C322(1.07) | LDD0618 | [7] |
| LDCM0302 | AC40 | HCT 116 | C322(1.16) | LDD0619 | [7] |
| LDCM0303 | AC41 | HCT 116 | C322(0.81) | LDD0620 | [7] |
| LDCM0304 | AC42 | HCT 116 | C322(0.93) | LDD0621 | [7] |
| LDCM0305 | AC43 | HCT 116 | C322(1.00) | LDD0622 | [7] |
| LDCM0306 | AC44 | HCT 116 | C322(0.94) | LDD0623 | [7] |
| LDCM0307 | AC45 | HCT 116 | C322(1.03) | LDD0624 | [7] |
| LDCM0308 | AC46 | HCT 116 | C322(0.92) | LDD0625 | [7] |
| LDCM0309 | AC47 | HCT 116 | C322(1.12) | LDD0626 | [7] |
| LDCM0310 | AC48 | HCT 116 | C322(0.98) | LDD0627 | [7] |
| LDCM0311 | AC49 | HCT 116 | C322(1.47) | LDD0628 | [7] |
| LDCM0312 | AC5 | HCT 116 | C322(0.90) | LDD0629 | [7] |
| LDCM0313 | AC50 | HCT 116 | C322(1.28) | LDD0630 | [7] |
| LDCM0314 | AC51 | HCT 116 | C322(0.97) | LDD0631 | [7] |
| LDCM0315 | AC52 | HCT 116 | C322(1.13) | LDD0632 | [7] |
| LDCM0316 | AC53 | HCT 116 | C322(1.13) | LDD0633 | [7] |
| LDCM0317 | AC54 | HCT 116 | C322(1.23) | LDD0634 | [7] |
| LDCM0318 | AC55 | HCT 116 | C322(1.42) | LDD0635 | [7] |
| LDCM0319 | AC56 | HCT 116 | C322(1.84) | LDD0636 | [7] |
| LDCM0320 | AC57 | HCT 116 | C322(1.13) | LDD0637 | [7] |
| LDCM0321 | AC58 | HCT 116 | C322(1.53) | LDD0638 | [7] |
| LDCM0322 | AC59 | HCT 116 | C322(0.99) | LDD0639 | [7] |
| LDCM0323 | AC6 | HCT 116 | C322(1.13) | LDD0640 | [7] |
| LDCM0324 | AC60 | HCT 116 | C322(1.07) | LDD0641 | [7] |
| LDCM0325 | AC61 | HCT 116 | C322(0.93) | LDD0642 | [7] |
| LDCM0326 | AC62 | HCT 116 | C322(1.41) | LDD0643 | [7] |
| LDCM0327 | AC63 | HCT 116 | C322(1.10) | LDD0644 | [7] |
| LDCM0328 | AC64 | HCT 116 | C322(1.32) | LDD0645 | [7] |
| LDCM0329 | AC65 | HCT 116 | C322(1.29) | LDD0646 | [7] |
| LDCM0330 | AC66 | HCT 116 | C322(1.25) | LDD0647 | [7] |
| LDCM0331 | AC67 | HCT 116 | C322(1.37) | LDD0648 | [7] |
| LDCM0332 | AC68 | HCT 116 | C322(1.03) | LDD0649 | [7] |
| LDCM0333 | AC69 | HCT 116 | C322(1.27) | LDD0650 | [7] |
| LDCM0334 | AC7 | HCT 116 | C322(0.93) | LDD0651 | [7] |
| LDCM0335 | AC70 | HCT 116 | C322(1.21) | LDD0652 | [7] |
| LDCM0336 | AC71 | HCT 116 | C322(1.07) | LDD0653 | [7] |
| LDCM0337 | AC72 | HCT 116 | C322(1.30) | LDD0654 | [7] |
| LDCM0338 | AC73 | HCT 116 | C322(1.72) | LDD0655 | [7] |
| LDCM0339 | AC74 | HCT 116 | C322(1.65) | LDD0656 | [7] |
| LDCM0340 | AC75 | HCT 116 | C322(1.89) | LDD0657 | [7] |
| LDCM0341 | AC76 | HCT 116 | C322(1.33) | LDD0658 | [7] |
| LDCM0342 | AC77 | HCT 116 | C322(1.18) | LDD0659 | [7] |
| LDCM0343 | AC78 | HCT 116 | C322(1.17) | LDD0660 | [7] |
| LDCM0344 | AC79 | HCT 116 | C322(1.27) | LDD0661 | [7] |
| LDCM0345 | AC8 | HCT 116 | C322(1.03) | LDD0662 | [7] |
| LDCM0346 | AC80 | HCT 116 | C322(1.33) | LDD0663 | [7] |
| LDCM0347 | AC81 | HCT 116 | C322(0.66) | LDD0664 | [7] |
| LDCM0348 | AC82 | HCT 116 | C322(1.83) | LDD0665 | [7] |
| LDCM0349 | AC83 | HCT 116 | C322(1.48) | LDD0666 | [7] |
| LDCM0350 | AC84 | HCT 116 | C322(1.23) | LDD0667 | [7] |
| LDCM0351 | AC85 | HCT 116 | C322(1.12) | LDD0668 | [7] |
| LDCM0352 | AC86 | HCT 116 | C322(1.07) | LDD0669 | [7] |
| LDCM0353 | AC87 | HCT 116 | C322(1.08) | LDD0670 | [7] |
| LDCM0354 | AC88 | HCT 116 | C322(1.12) | LDD0671 | [7] |
| LDCM0355 | AC89 | HCT 116 | C322(1.38) | LDD0672 | [7] |
| LDCM0357 | AC90 | HCT 116 | C322(1.13) | LDD0674 | [7] |
| LDCM0358 | AC91 | HCT 116 | C322(1.57) | LDD0675 | [7] |
| LDCM0359 | AC92 | HCT 116 | C322(1.29) | LDD0676 | [7] |
| LDCM0360 | AC93 | HCT 116 | C322(1.24) | LDD0677 | [7] |
| LDCM0361 | AC94 | HCT 116 | C322(1.48) | LDD0678 | [7] |
| LDCM0362 | AC95 | HCT 116 | C322(1.13) | LDD0679 | [7] |
| LDCM0363 | AC96 | HCT 116 | C322(1.65) | LDD0680 | [7] |
| LDCM0364 | AC97 | HCT 116 | C322(1.71) | LDD0681 | [7] |
| LDCM0365 | AC98 | HCT 116 | C322(0.95) | LDD0682 | [7] |
| LDCM0366 | AC99 | HCT 116 | C322(0.84) | LDD0683 | [7] |
| LDCM0248 | AKOS034007472 | HCT 116 | C322(0.80) | LDD0565 | [7] |
| LDCM0356 | AKOS034007680 | HCT 116 | C322(0.92) | LDD0673 | [7] |
| LDCM0275 | AKOS034007705 | HCT 116 | C322(1.41) | LDD0592 | [7] |
| LDCM0020 | ARS-1620 | HCC44 | C322(1.16) | LDD2171 | [7] |
| LDCM0367 | CL1 | HCT 116 | C322(1.75) | LDD0684 | [7] |
| LDCM0368 | CL10 | HCT 116 | C322(1.47) | LDD0685 | [7] |
| LDCM0369 | CL100 | HCT 116 | C322(1.05) | LDD0686 | [7] |
| LDCM0370 | CL101 | HCT 116 | C322(0.90) | LDD0687 | [7] |
| LDCM0371 | CL102 | HCT 116 | C322(0.92) | LDD0688 | [7] |
| LDCM0372 | CL103 | HCT 116 | C322(1.33) | LDD0689 | [7] |
| LDCM0373 | CL104 | HCT 116 | C322(1.14) | LDD0690 | [7] |
| LDCM0374 | CL105 | HCT 116 | C322(1.30) | LDD0691 | [7] |
| LDCM0375 | CL106 | HCT 116 | C322(1.16) | LDD0692 | [7] |
| LDCM0376 | CL107 | HCT 116 | C322(1.32) | LDD0693 | [7] |
| LDCM0377 | CL108 | HCT 116 | C322(2.21) | LDD0694 | [7] |
| LDCM0378 | CL109 | HCT 116 | C322(1.20) | LDD0695 | [7] |
| LDCM0379 | CL11 | HCT 116 | C322(1.49) | LDD0696 | [7] |
| LDCM0380 | CL110 | HCT 116 | C322(1.33) | LDD0697 | [7] |
| LDCM0381 | CL111 | HCT 116 | C322(1.13) | LDD0698 | [7] |
| LDCM0382 | CL112 | PaTu 8988t | C322(0.90) | LDD1261 | [7] |
| LDCM0383 | CL113 | PaTu 8988t | C322(0.86) | LDD1262 | [7] |
| LDCM0384 | CL114 | PaTu 8988t | C322(1.49) | LDD1263 | [7] |
| LDCM0385 | CL115 | PaTu 8988t | C322(0.86) | LDD1264 | [7] |
| LDCM0386 | CL116 | PaTu 8988t | C322(1.11) | LDD1265 | [7] |
| LDCM0387 | CL117 | HCT 116 | C322(1.50) | LDD0704 | [7] |
| LDCM0388 | CL118 | HCT 116 | C322(0.84) | LDD0705 | [7] |
| LDCM0389 | CL119 | HCT 116 | C322(1.10) | LDD0706 | [7] |
| LDCM0390 | CL12 | HCT 116 | C322(2.37) | LDD0707 | [7] |
| LDCM0391 | CL120 | HCT 116 | C322(0.78) | LDD0708 | [7] |
| LDCM0392 | CL121 | HCT 116 | C322(1.06) | LDD0709 | [7] |
| LDCM0393 | CL122 | HCT 116 | C322(1.12) | LDD0710 | [7] |
| LDCM0394 | CL123 | HCT 116 | C322(1.76) | LDD0711 | [7] |
| LDCM0395 | CL124 | HCT 116 | C322(1.27) | LDD0712 | [7] |
| LDCM0396 | CL125 | HCT 116 | C322(0.77) | LDD0713 | [7] |
| LDCM0397 | CL126 | HCT 116 | C322(1.13) | LDD0714 | [7] |
| LDCM0398 | CL127 | HCT 116 | C322(1.19) | LDD0715 | [7] |
| LDCM0399 | CL128 | HCT 116 | C322(1.18) | LDD0716 | [7] |
| LDCM0400 | CL13 | HCT 116 | C322(1.55) | LDD0717 | [7] |
| LDCM0401 | CL14 | HCT 116 | C322(1.61) | LDD0718 | [7] |
| LDCM0402 | CL15 | HCT 116 | C322(1.48) | LDD0719 | [7] |
| LDCM0403 | CL16 | HCT 116 | C322(1.29) | LDD0720 | [7] |
| LDCM0404 | CL17 | HCT 116 | C322(1.82) | LDD0721 | [7] |
| LDCM0405 | CL18 | HCT 116 | C322(1.35) | LDD0722 | [7] |
| LDCM0406 | CL19 | HCT 116 | C322(1.32) | LDD0723 | [7] |
| LDCM0407 | CL2 | HCT 116 | C322(1.01) | LDD0724 | [7] |
| LDCM0408 | CL20 | HCT 116 | C322(1.35) | LDD0725 | [7] |
| LDCM0409 | CL21 | HCT 116 | C322(2.01) | LDD0726 | [7] |
| LDCM0410 | CL22 | HCT 116 | C322(2.14) | LDD0727 | [7] |
| LDCM0411 | CL23 | HCT 116 | C322(1.59) | LDD0728 | [7] |
| LDCM0412 | CL24 | HCT 116 | C322(1.82) | LDD0729 | [7] |
| LDCM0413 | CL25 | HCT 116 | C322(1.74) | LDD0730 | [7] |
| LDCM0414 | CL26 | HCT 116 | C322(1.44) | LDD0731 | [7] |
| LDCM0415 | CL27 | HCT 116 | C322(1.13) | LDD0732 | [7] |
| LDCM0416 | CL28 | HCT 116 | C322(1.49) | LDD0733 | [7] |
| LDCM0417 | CL29 | HCT 116 | C322(1.35) | LDD0734 | [7] |
| LDCM0418 | CL3 | HCT 116 | C322(1.22) | LDD0735 | [7] |
| LDCM0419 | CL30 | HCT 116 | C322(1.35) | LDD0736 | [7] |
| LDCM0420 | CL31 | HCT 116 | C322(0.74) | LDD0737 | [7] |
| LDCM0421 | CL32 | HCT 116 | C322(0.75) | LDD0738 | [7] |
| LDCM0422 | CL33 | HCT 116 | C322(0.87) | LDD0739 | [7] |
| LDCM0423 | CL34 | HCT 116 | C322(1.08) | LDD0740 | [7] |
| LDCM0424 | CL35 | HCT 116 | C322(1.00) | LDD0741 | [7] |
| LDCM0425 | CL36 | HCT 116 | C322(0.98) | LDD0742 | [7] |
| LDCM0426 | CL37 | HCT 116 | C322(1.08) | LDD0743 | [7] |
| LDCM0428 | CL39 | HCT 116 | C322(0.94) | LDD0745 | [7] |
| LDCM0429 | CL4 | HCT 116 | C322(1.67) | LDD0746 | [7] |
| LDCM0430 | CL40 | HCT 116 | C322(0.91) | LDD0747 | [7] |
| LDCM0431 | CL41 | HCT 116 | C322(0.91) | LDD0748 | [7] |
| LDCM0432 | CL42 | HCT 116 | C322(1.14) | LDD0749 | [7] |
| LDCM0433 | CL43 | HCT 116 | C322(1.11) | LDD0750 | [7] |
| LDCM0434 | CL44 | HCT 116 | C322(0.88) | LDD0751 | [7] |
| LDCM0435 | CL45 | HCT 116 | C322(0.79) | LDD0752 | [7] |
| LDCM0436 | CL46 | PaTu 8988t | C322(1.11) | LDD1315 | [7] |
| LDCM0437 | CL47 | PaTu 8988t | C322(1.28) | LDD1316 | [7] |
| LDCM0438 | CL48 | PaTu 8988t | C322(1.37) | LDD1317 | [7] |
| LDCM0439 | CL49 | PaTu 8988t | C322(1.21) | LDD1318 | [7] |
| LDCM0440 | CL5 | HCT 116 | C322(1.09) | LDD0757 | [7] |
| LDCM0441 | CL50 | PaTu 8988t | C322(1.43) | LDD1320 | [7] |
| LDCM0442 | CL51 | PaTu 8988t | C322(1.38) | LDD1321 | [7] |
| LDCM0443 | CL52 | PaTu 8988t | C322(1.26) | LDD1322 | [7] |
| LDCM0444 | CL53 | PaTu 8988t | C322(1.75) | LDD1323 | [7] |
| LDCM0445 | CL54 | PaTu 8988t | C322(1.24) | LDD1324 | [7] |
| LDCM0446 | CL55 | PaTu 8988t | C322(1.23) | LDD1325 | [7] |
| LDCM0447 | CL56 | PaTu 8988t | C322(0.77) | LDD1326 | [7] |
| LDCM0448 | CL57 | PaTu 8988t | C322(1.88) | LDD1327 | [7] |
| LDCM0449 | CL58 | PaTu 8988t | C322(1.54) | LDD1328 | [7] |
| LDCM0450 | CL59 | PaTu 8988t | C322(1.68) | LDD1329 | [7] |
| LDCM0451 | CL6 | HCT 116 | C322(1.45) | LDD0768 | [7] |
| LDCM0452 | CL60 | PaTu 8988t | C322(1.51) | LDD1331 | [7] |
| LDCM0453 | CL61 | HCT 116 | C322(0.80) | LDD0770 | [7] |
| LDCM0454 | CL62 | HCT 116 | C322(0.88) | LDD0771 | [7] |
| LDCM0455 | CL63 | HCT 116 | C322(0.86) | LDD0772 | [7] |
| LDCM0456 | CL64 | HCT 116 | C322(1.14) | LDD0773 | [7] |
| LDCM0457 | CL65 | HCT 116 | C322(0.88) | LDD0774 | [7] |
| LDCM0458 | CL66 | HCT 116 | C322(0.98) | LDD0775 | [7] |
| LDCM0459 | CL67 | HCT 116 | C322(0.98) | LDD0776 | [7] |
| LDCM0460 | CL68 | HCT 116 | C322(0.93) | LDD0777 | [7] |
| LDCM0461 | CL69 | HCT 116 | C322(0.88) | LDD0778 | [7] |
| LDCM0462 | CL7 | HCT 116 | C322(1.67) | LDD0779 | [7] |
| LDCM0463 | CL70 | HCT 116 | C322(1.10) | LDD0780 | [7] |
| LDCM0464 | CL71 | HCT 116 | C322(0.83) | LDD0781 | [7] |
| LDCM0465 | CL72 | HCT 116 | C322(0.72) | LDD0782 | [7] |
| LDCM0466 | CL73 | HCT 116 | C322(1.30) | LDD0783 | [7] |
| LDCM0467 | CL74 | HCT 116 | C322(1.06) | LDD0784 | [7] |
| LDCM0469 | CL76 | HCT 116 | C322(1.27) | LDD0786 | [7] |
| LDCM0470 | CL77 | HCT 116 | C322(3.88) | LDD0787 | [7] |
| LDCM0471 | CL78 | HCT 116 | C322(0.89) | LDD0788 | [7] |
| LDCM0472 | CL79 | HCT 116 | C322(1.14) | LDD0789 | [7] |
| LDCM0473 | CL8 | HCT 116 | C322(1.65) | LDD0790 | [7] |
| LDCM0474 | CL80 | HCT 116 | C322(1.30) | LDD0791 | [7] |
| LDCM0475 | CL81 | HCT 116 | C322(0.99) | LDD0792 | [7] |
| LDCM0476 | CL82 | HCT 116 | C322(1.38) | LDD0793 | [7] |
| LDCM0477 | CL83 | HCT 116 | C322(1.33) | LDD0794 | [7] |
| LDCM0478 | CL84 | HCT 116 | C322(1.42) | LDD0795 | [7] |
| LDCM0479 | CL85 | HCT 116 | C322(0.97) | LDD0796 | [7] |
| LDCM0480 | CL86 | HCT 116 | C322(0.95) | LDD0797 | [7] |
| LDCM0481 | CL87 | HCT 116 | C322(1.29) | LDD0798 | [7] |
| LDCM0482 | CL88 | HCT 116 | C322(1.32) | LDD0799 | [7] |
| LDCM0483 | CL89 | HCT 116 | C322(1.71) | LDD0800 | [7] |
| LDCM0484 | CL9 | HCT 116 | C322(1.35) | LDD0801 | [7] |
| LDCM0485 | CL90 | HCT 116 | C322(0.99) | LDD0802 | [7] |
| LDCM0486 | CL91 | HCT 116 | C322(0.87) | LDD0803 | [7] |
| LDCM0487 | CL92 | HCT 116 | C322(1.11) | LDD0804 | [7] |
| LDCM0488 | CL93 | HCT 116 | C322(1.16) | LDD0805 | [7] |
| LDCM0489 | CL94 | HCT 116 | C322(1.29) | LDD0806 | [7] |
| LDCM0490 | CL95 | HCT 116 | C322(1.21) | LDD0807 | [7] |
| LDCM0491 | CL96 | HCT 116 | C322(0.98) | LDD0808 | [7] |
| LDCM0492 | CL97 | HCT 116 | C322(1.18) | LDD0809 | [7] |
| LDCM0493 | CL98 | HCT 116 | C322(1.02) | LDD0810 | [7] |
| LDCM0494 | CL99 | HCT 116 | C322(1.07) | LDD0811 | [7] |
| LDCM0495 | E2913 | HEK-293T | C290(0.90); C118(0.94); C322(1.03) | LDD1698 | [14] |
| LDCM0468 | Fragment33 | HCT 116 | C322(1.04) | LDD0785 | [7] |
| LDCM0427 | Fragment51 | HCT 116 | C322(1.00) | LDD0744 | [7] |
| LDCM0615 | Fragment63-R | Jurkat | _(7.16) | LDD1487 | [4] |
| LDCM0569 | Fragment7 | Jurkat | _(20.00) | LDD1485 | [4] |
| LDCM0022 | KB02 | HCT 116 | C322(8.04) | LDD0080 | [7] |
| LDCM0023 | KB03 | HCT 116 | C322(7.01) | LDD0081 | [7] |
| LDCM0024 | KB05 | HCT 116 | C322(9.64) | LDD0082 | [7] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0226 | [6] |
| LDCM0021 | THZ1 | HCT 116 | C322(1.16) | LDD2173 | [7] |
The Interaction Atlas With This Target
References
