Details of the Target
General Information of Target
| Target ID | LDTP08981 | |||||
|---|---|---|---|---|---|---|
| Target Name | Cytochrome c oxidase assembly protein COX18, mitochondrial (COX18) | |||||
| Gene Name | COX18 | |||||
| Gene ID | 285521 | |||||
| Synonyms |
OXA1L2; Cytochrome c oxidase assembly protein COX18, mitochondrial; COX18Hs; Cytochrome c oxidase assembly protein 18 |
|||||
| 3D Structure | ||||||
| Sequence |
MLCRLGGRWLRPLPALQLWARDLPLAPVPTSGAKRPTLPVWAVAPVSAVHANGWYEALAA
SSPVRVAEEVLLGVHAATGLPWWGSILLSTVALRGAVTLPLAAYQHYILAKVENLQPEIK TIARHLNQEVAVRANQLGWSKRDARLTYLKNMRRLISELYVRDNCHPFKATVLVWIQLPM WIFMSFALRNLSTGAAHSEGFSVQEQLATGGILWFPDLTAPDSTWILPISVGVINLLIVE ICALQKIGMSRFQTYITYFVRAMSVLMIPIAATVPSSIVLYWLCSSFVGLSQNLLLRSPG FRQLCRIPSTKSDSETPYKDIFAAFNTKFISRK |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
OXA1/ALB3/YidC family
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Mitochondrial membrane insertase required for the translocation of the C-terminus of cytochrome c oxidase subunit II (MT-CO2/COX2) across the mitochondrial inner membrane. Plays a role in MT-CO2/COX2 maturation following the COX20-mediated stabilization of newly synthesized MT-CO2/COX2 protein and before the action of the metallochaperones SCO1/2. Essential for the assembly and stability of the mitochondrial respiratory chain complex IV (also known as cytochrome c oxidase).
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STPyne Probe Info |
 |
K141(3.03); K328(0.90) | LDD0277 | [1] | |
|
Curcusone 37 Probe Info |
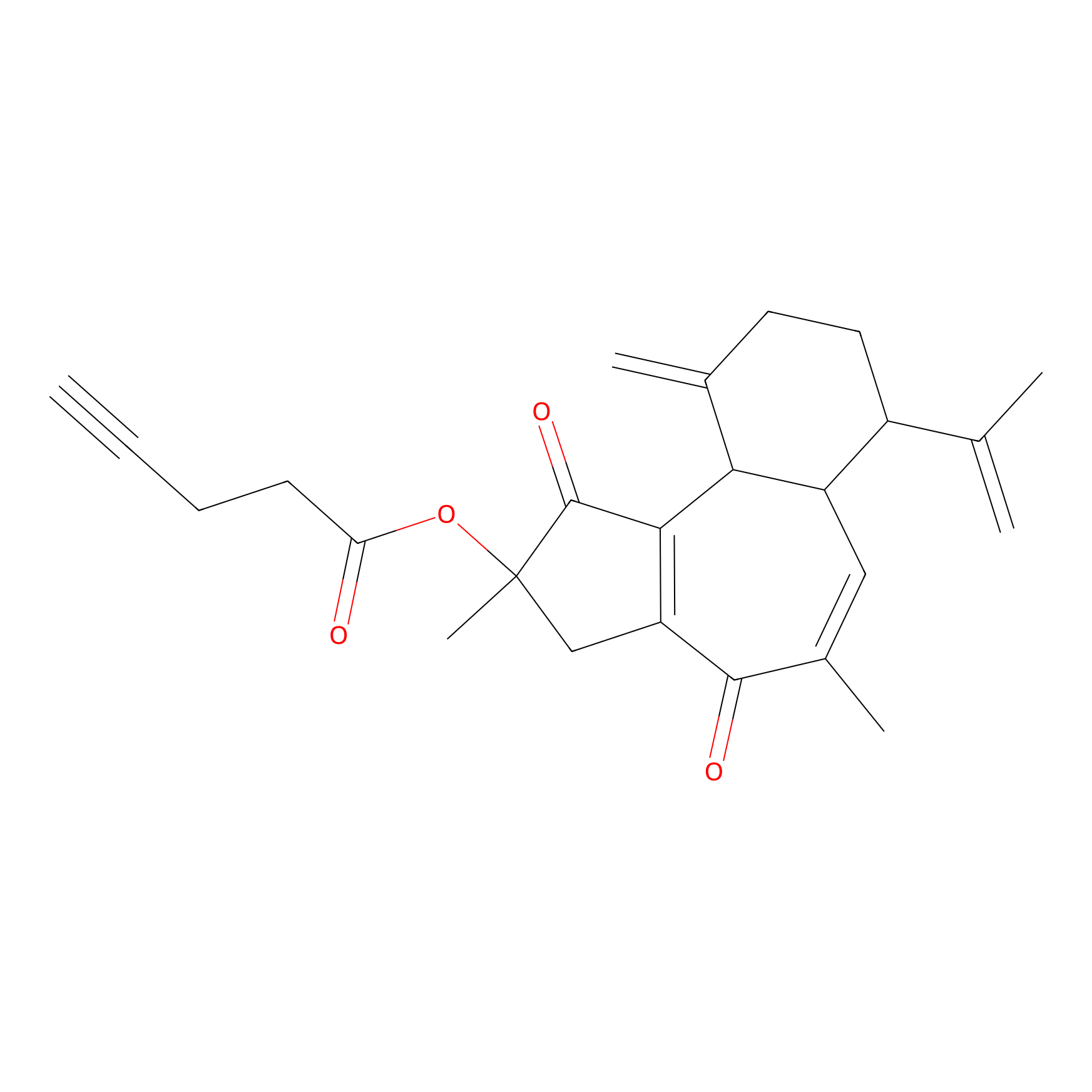 |
3.88 | LDD0188 | [2] | |
|
DBIA Probe Info |
 |
C165(0.86) | LDD1492 | [3] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0233 | [4] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [5] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C165(0.95) | LDD1507 | [3] |
| LDCM0259 | AC14 | HEK-293T | C165(1.02) | LDD1512 | [3] |
| LDCM0276 | AC17 | HEK-293T | C165(0.95) | LDD1515 | [3] |
| LDCM0282 | AC22 | HEK-293T | C165(0.95) | LDD1521 | [3] |
| LDCM0285 | AC25 | HEK-293T | C165(0.88) | LDD1524 | [3] |
| LDCM0291 | AC30 | HEK-293T | C165(0.93) | LDD1530 | [3] |
| LDCM0294 | AC33 | HEK-293T | C165(1.09) | LDD1533 | [3] |
| LDCM0299 | AC38 | HEK-293T | C165(0.98) | LDD1538 | [3] |
| LDCM0303 | AC41 | HEK-293T | C165(1.06) | LDD1542 | [3] |
| LDCM0308 | AC46 | HEK-293T | C165(0.94) | LDD1547 | [3] |
| LDCM0311 | AC49 | HEK-293T | C165(0.94) | LDD1550 | [3] |
| LDCM0317 | AC54 | HEK-293T | C165(0.93) | LDD1556 | [3] |
| LDCM0320 | AC57 | HEK-293T | C165(1.09) | LDD1559 | [3] |
| LDCM0323 | AC6 | HEK-293T | C165(1.00) | LDD1562 | [3] |
| LDCM0326 | AC62 | HEK-293T | C165(0.92) | LDD1565 | [3] |
| LDCM0356 | AKOS034007680 | HEK-293T | C165(0.99) | LDD1570 | [3] |
| LDCM0368 | CL10 | HEK-293T | C165(1.92) | LDD1572 | [3] |
| LDCM0404 | CL17 | HEK-293T | C165(1.49) | LDD1608 | [3] |
| LDCM0410 | CL22 | HEK-293T | C165(1.08) | LDD1614 | [3] |
| LDCM0417 | CL29 | HEK-293T | C165(0.92) | LDD1621 | [3] |
| LDCM0423 | CL34 | HEK-293T | C165(1.09) | LDD1627 | [3] |
| LDCM0431 | CL41 | HEK-293T | C165(1.06) | LDD1635 | [3] |
| LDCM0436 | CL46 | HEK-293T | C165(1.02) | LDD1640 | [3] |
| LDCM0440 | CL5 | HEK-293T | C165(1.20) | LDD1644 | [3] |
| LDCM0444 | CL53 | HEK-293T | C165(1.29) | LDD1647 | [3] |
| LDCM0449 | CL58 | HEK-293T | C165(1.17) | LDD1652 | [3] |
| LDCM0457 | CL65 | HEK-293T | C165(1.02) | LDD1660 | [3] |
| LDCM0463 | CL70 | HEK-293T | C165(1.02) | LDD1666 | [3] |
| LDCM0470 | CL77 | HEK-293T | C165(1.23) | LDD1673 | [3] |
| LDCM0476 | CL82 | HEK-293T | C165(0.99) | LDD1679 | [3] |
| LDCM0483 | CL89 | HEK-293T | C165(1.08) | LDD1686 | [3] |
| LDCM0489 | CL94 | HEK-293T | C165(1.06) | LDD1692 | [3] |
| LDCM0033 | Curcusone1d | MCF-7 | 3.88 | LDD0188 | [2] |
| LDCM0022 | KB02 | HEK-293T | C165(0.86) | LDD1492 | [3] |
| LDCM0023 | KB03 | HEK-293T | C165(0.91) | LDD1497 | [3] |
| LDCM0024 | KB05 | HEK-293T | C165(0.68) | LDD1502 | [3] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0233 | [4] |
References