Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DA-P2 Probe Info |
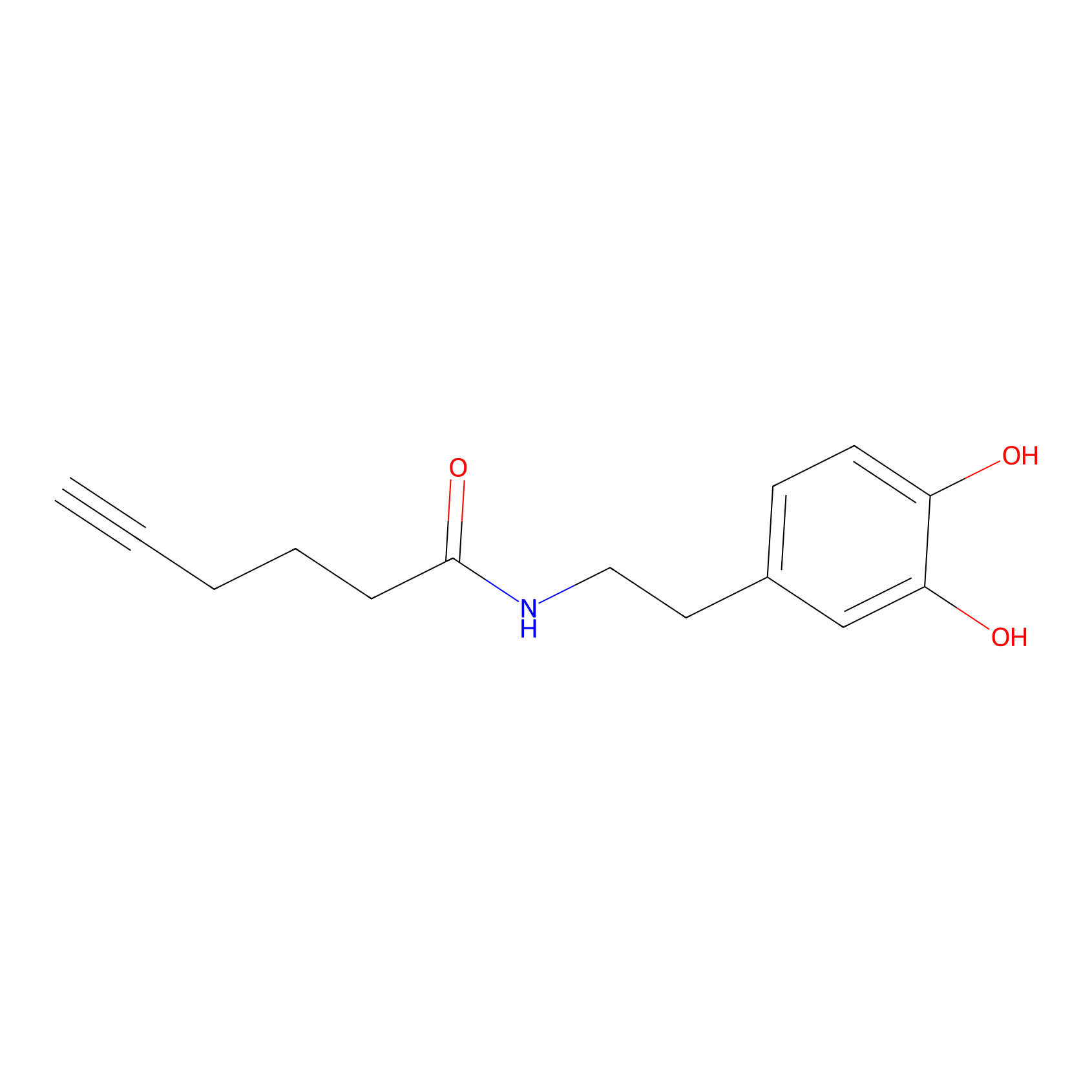 |
6.81 | LDD0348 | [1] | |
|
HRP Probe Info |
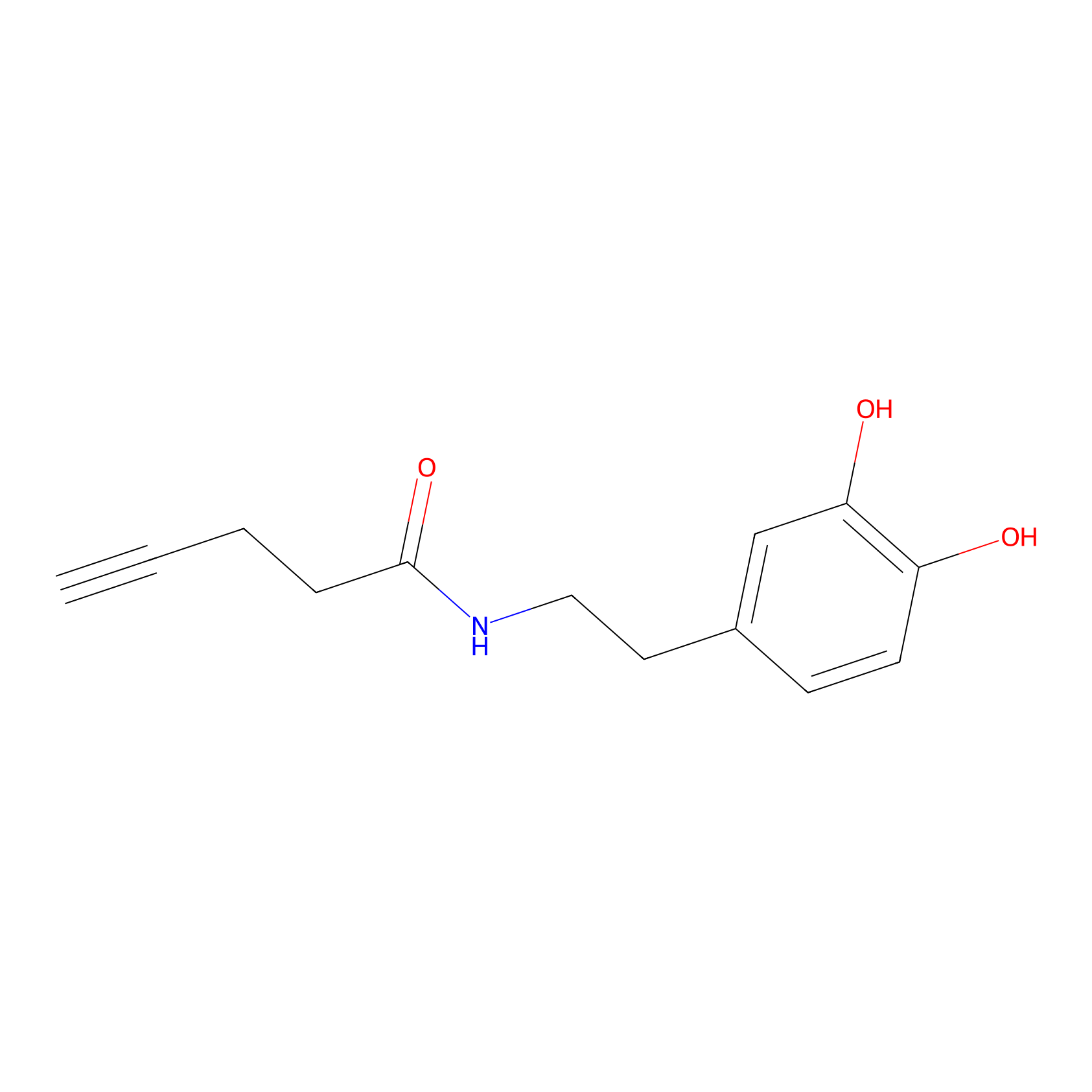 |
4.91 | LDD0347 | [1] | |
|
FBP2 Probe Info |
 |
5.55 | LDD0317 | [2] | |
|
Alkylaryl probe 2 Probe Info |
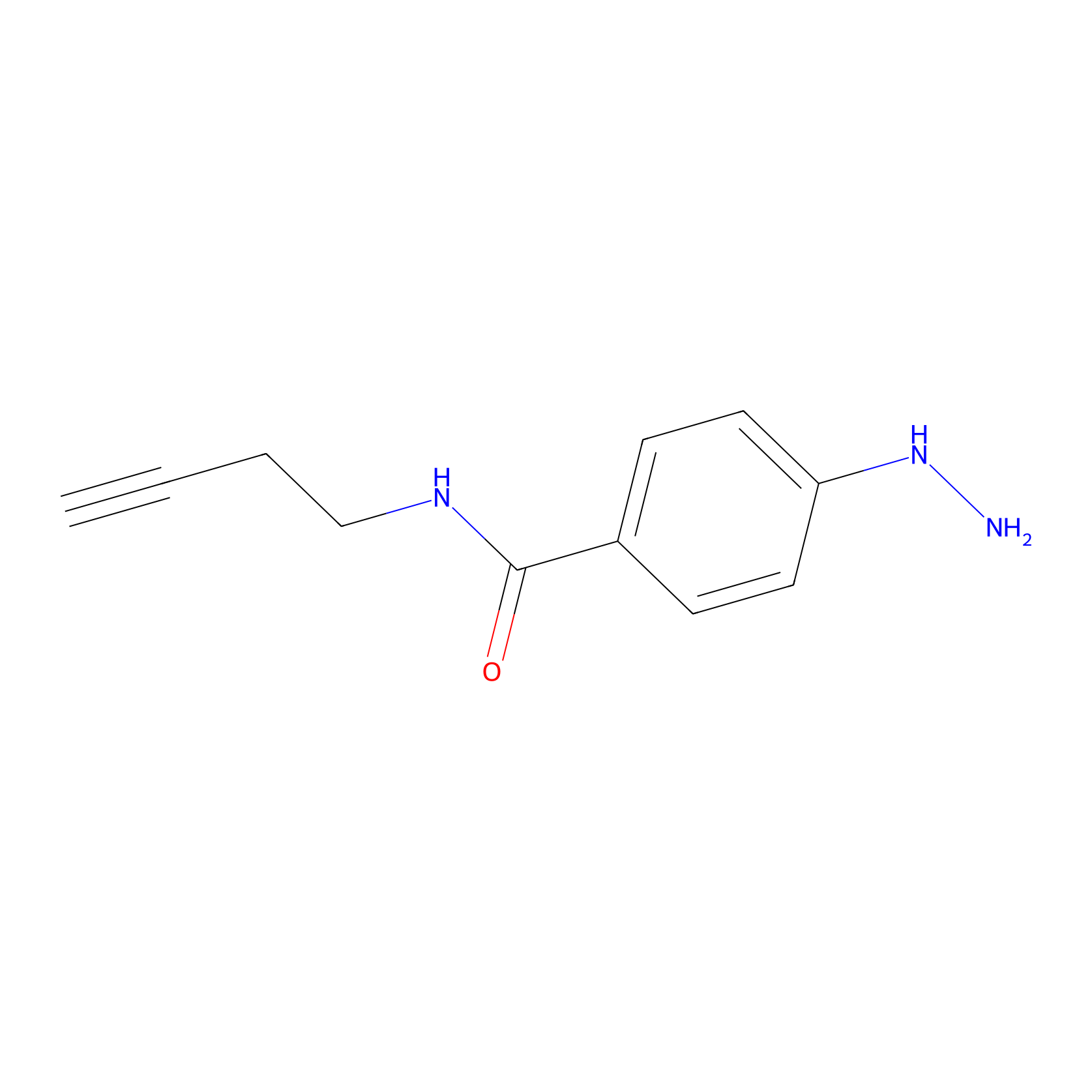 |
5.00 | LDD0390 | [3] | |
|
Sulforaphane-probe2 Probe Info |
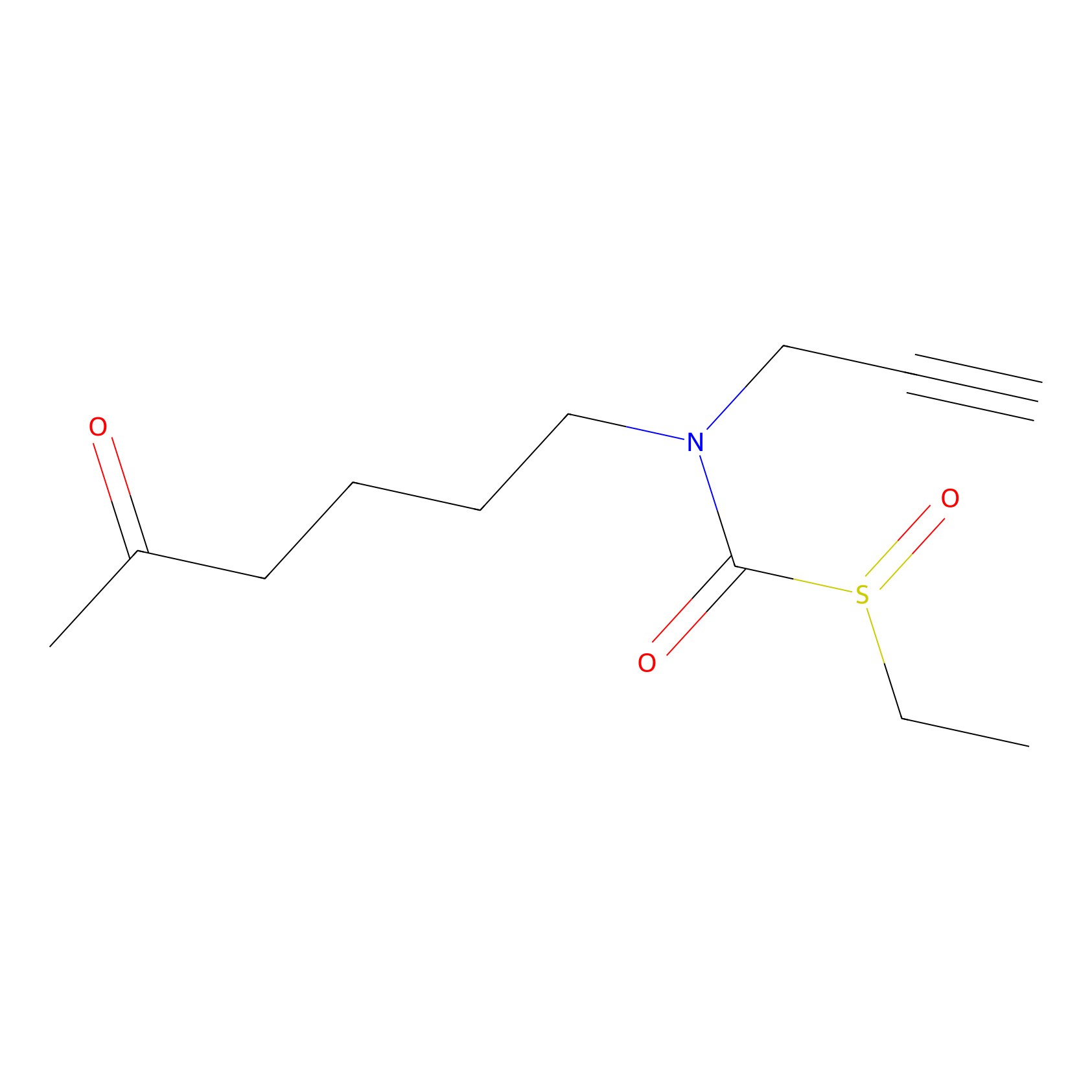 |
2.61 | LDD0160 | [4] | |
|
Johansson_61 Probe Info |
 |
_(20.00) | LDD1490 | [5] | |
|
DA-P3 Probe Info |
 |
11.97 | LDD0183 | [1] | |
|
AOyne Probe Info |
 |
6.10 | LDD0443 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C063 Probe Info |
 |
15.78 | LDD1760 | [7] | |
|
C070 Probe Info |
 |
25.11 | LDD1766 | [7] | |
|
C112 Probe Info |
 |
33.36 | LDD1799 | [7] | |
|
C153 Probe Info |
 |
24.08 | LDD1834 | [7] | |
|
C159 Probe Info |
 |
62.25 | LDD1839 | [7] | |
|
C160 Probe Info |
 |
11.63 | LDD1840 | [7] | |
|
C161 Probe Info |
 |
17.15 | LDD1841 | [7] | |
|
C183 Probe Info |
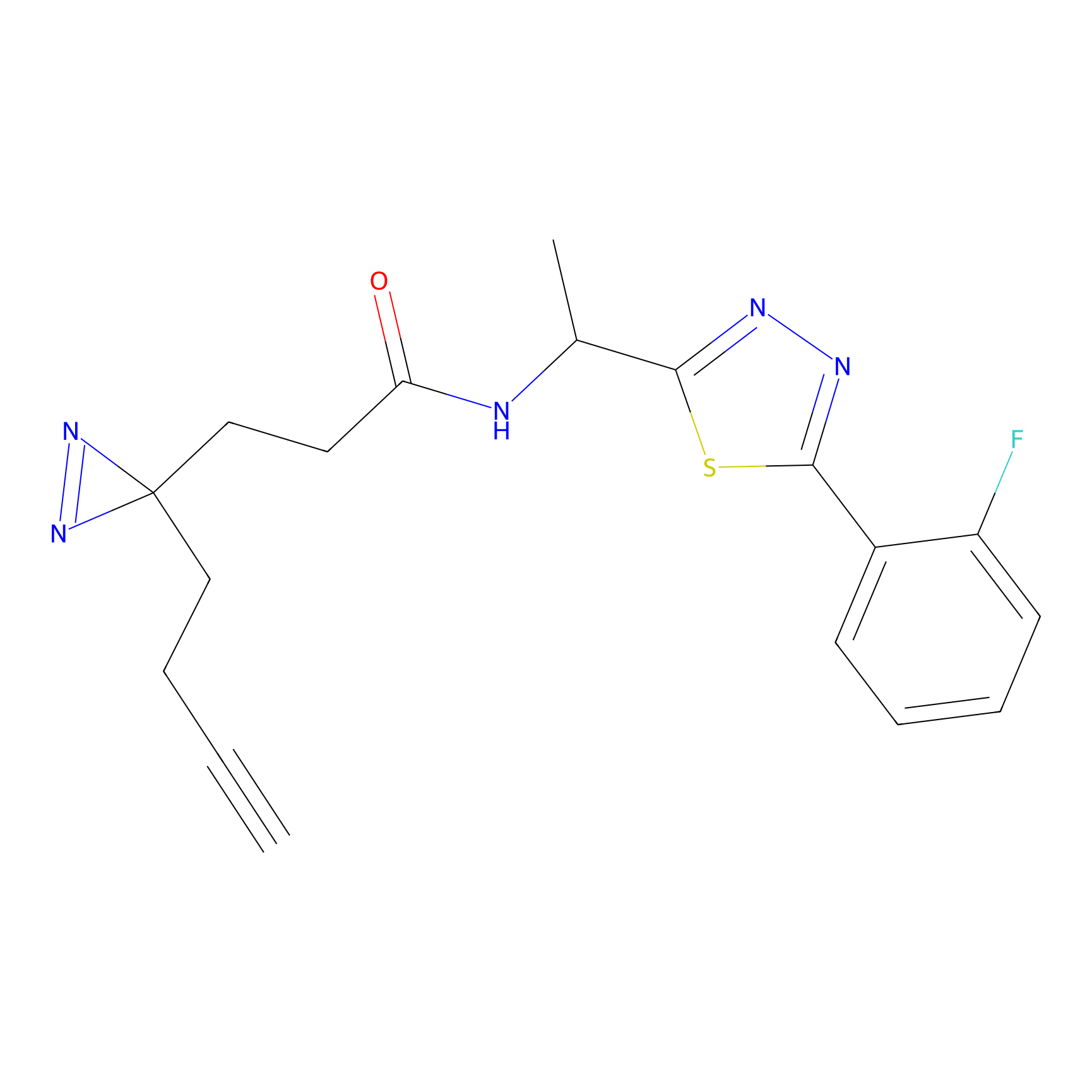 |
5.13 | LDD1861 | [7] | |
|
C218 Probe Info |
 |
13.55 | LDD1892 | [7] | |
|
C219 Probe Info |
 |
10.48 | LDD1893 | [7] | |
|
C220 Probe Info |
 |
18.38 | LDD1894 | [7] | |
|
C231 Probe Info |
 |
17.03 | LDD1904 | [7] | |
|
C233 Probe Info |
 |
9.99 | LDD1906 | [7] | |
|
C235 Probe Info |
 |
27.28 | LDD1908 | [7] | |
|
C355 Probe Info |
 |
30.27 | LDD2016 | [7] | |
|
C356 Probe Info |
 |
11.31 | LDD2017 | [7] | |
|
C399 Probe Info |
 |
9.51 | LDD2058 | [7] | |
|
C403 Probe Info |
 |
26.35 | LDD2061 | [7] | |
|
C407 Probe Info |
 |
44.32 | LDD2064 | [7] | |
|
C413 Probe Info |
 |
24.59 | LDD2069 | [7] | |
|
C429 Probe Info |
 |
13.09 | LDD2084 | [7] | |
|
C430 Probe Info |
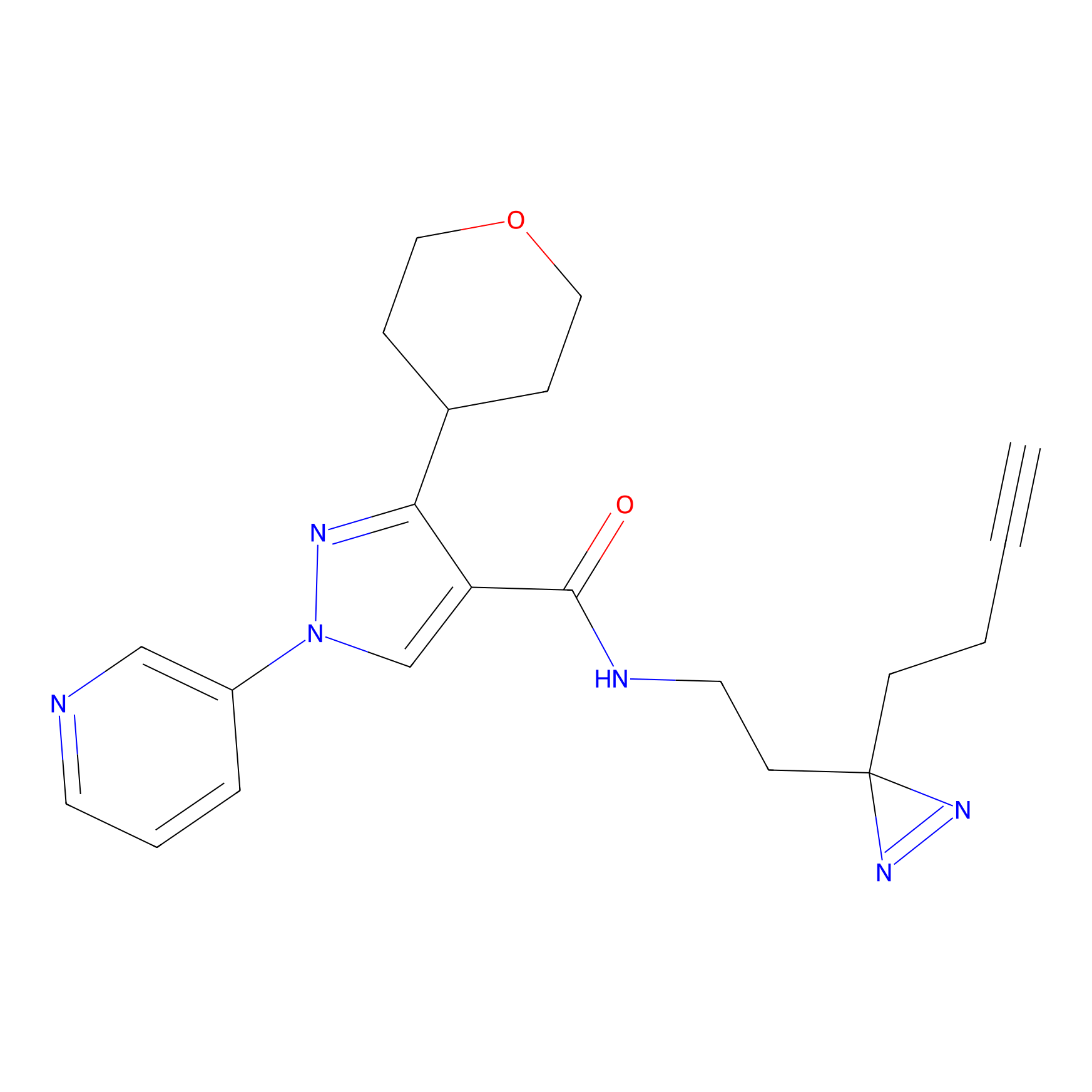 |
7.36 | LDD2085 | [7] | |
|
C431 Probe Info |
 |
15.24 | LDD2086 | [7] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [8] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [8] | |
|
FFF probe14 Probe Info |
 |
15.97 | LDD0477 | [8] | |
|
FFF probe15 Probe Info |
 |
20.00 | LDD0478 | [8] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [8] | |
|
FFF probe9 Probe Info |
 |
20.00 | LDD0470 | [8] | |
|
JN0003 Probe Info |
 |
20.00 | LDD0469 | [8] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [9] | |
|
OEA-DA Probe Info |
 |
9.70 | LDD0046 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 11.97 | LDD0183 | [1] |
| LDCM0617 | Fragment63-S | Jurkat | _(20.00) | LDD1490 | [5] |
| LDCM0032 | Oleacein | HEK-293T | 9.88 | LDD0184 | [1] |
| LDCM0099 | Phenelzine | HEK-293T | 5.00 | LDD0390 | [3] |
| LDCM0003 | Sulforaphane | MDA-MB-231 | 2.61 | LDD0160 | [4] |
References
