Details of the Target
General Information of Target
| Target ID | LDTP08058 | |||||
|---|---|---|---|---|---|---|
| Target Name | Mitochondrial antiviral-signaling protein (MAVS) | |||||
| Gene Name | MAVS | |||||
| Gene ID | 57506 | |||||
| Synonyms |
IPS1; KIAA1271; VISA; Mitochondrial antiviral-signaling protein; MAVS; CARD adapter inducing interferon beta; Cardif; Interferon beta promoter stimulator protein 1; IPS-1; Putative NF-kappa-B-activating protein 031N; Virus-induced-signaling adapter; VISA
|
|||||
| 3D Structure | ||||||
| Sequence |
MPFAEDKTYKYICRNFSNFCNVDVVEILPYLPCLTARDQDRLRATCTLSGNRDTLWHLFN
TLQRRPGWVEYFIAALRGCELVDLADEVASVYQSYQPRTSDRPPDPLEPPSLPAERPGPP TPAAAHSIPYNSCREKEPSYPMPVQETQAPESPGENSEQALQTLSPRAIPRNPDGGPLES SSDLAALSPLTSSGHQEQDTELGSTHTAGATSSLTPSRGPVSPSVSFQPLARSTPRASRL PGPTGSVVSTGTSFSSSSPGLASAGAAEGKQGAESDQAEPIICSSGAEAPANSLPSKVPT TLMPVNTVALKVPANPASVSTVPSKLPTSSKPPGAVPSNALTNPAPSKLPINSTRAGMVP SKVPTSMVLTKVSASTVPTDGSSRNEETPAAPTPAGATGGSSAWLDSSSENRGLGSELSK PGVLASQVDSPFSGCFEDLAISASTSLGMGPCHGPEENEYKSEGTFGIHVAENPSIQLLE GNPGPPADPDGGPRPQADRKFQEREVPCHRPSPGALWLQVAVTGVLVVTLLVVLYRRRLH |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Mitochondrion outer membrane
|
|||||
| Function |
Adapter required for innate immune defense against viruses. Acts downstream of DHX33, RIGI and IFIH1/MDA5, which detect intracellular dsRNA produced during viral replication, to coordinate pathways leading to the activation of NF-kappa-B, IRF3 and IRF7, and to the subsequent induction of antiviral cytokines such as IFNB and RANTES (CCL5). Peroxisomal and mitochondrial MAVS act sequentially to create an antiviral cellular state. Upon viral infection, peroxisomal MAVS induces the rapid interferon-independent expression of defense factors that provide short-term protection, whereas mitochondrial MAVS activates an interferon-dependent signaling pathway with delayed kinetics, which amplifies and stabilizes the antiviral response. May activate the same pathways following detection of extracellular dsRNA by TLR3. May protect cells from apoptosis. Involved in NLRP3 inflammasome activation by mediating NLRP3 recruitment to mitochondria.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| IGR37 | SNV: p.T393I | DBIA Probe Info | |||
| IGR39 | SNV: p.T393I | DBIA Probe Info | |||
| KYSE30 | SNV: p.T445I | DBIA Probe Info | |||
| MCC13 | SNV: p.L190Q | DBIA Probe Info | |||
| MDAMB468 | SNV: p.V82I | DBIA Probe Info | |||
| SNU1 | SNV: p.V506L | DBIA Probe Info | |||
| SUPT1 | SNV: p.P493T | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
DAyne Probe Info |
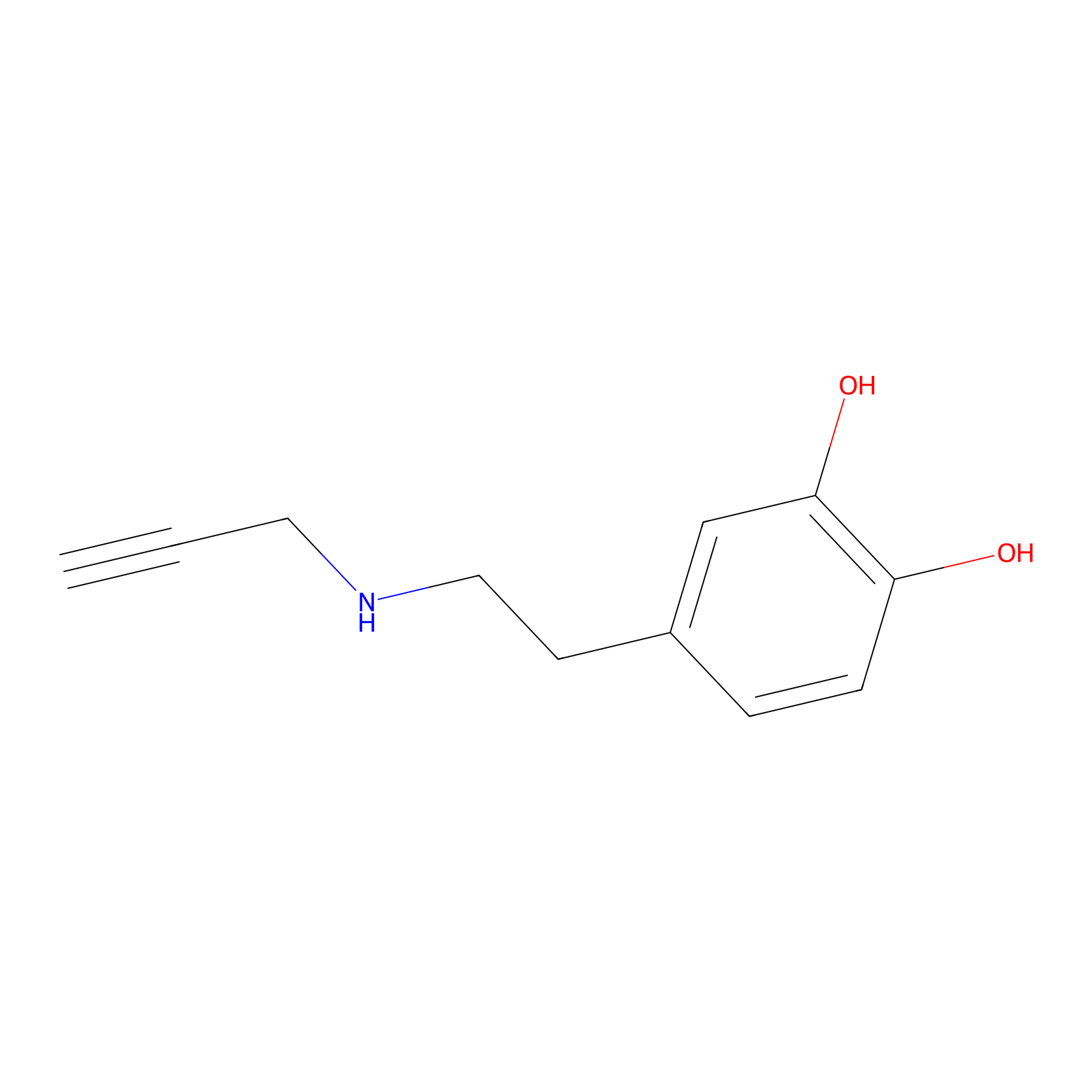 |
2.39 | LDD0261 | [2] | |
|
CHEMBL5175495 Probe Info |
 |
6.66 | LDD0196 | [3] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [4] | |
|
TG42 Probe Info |
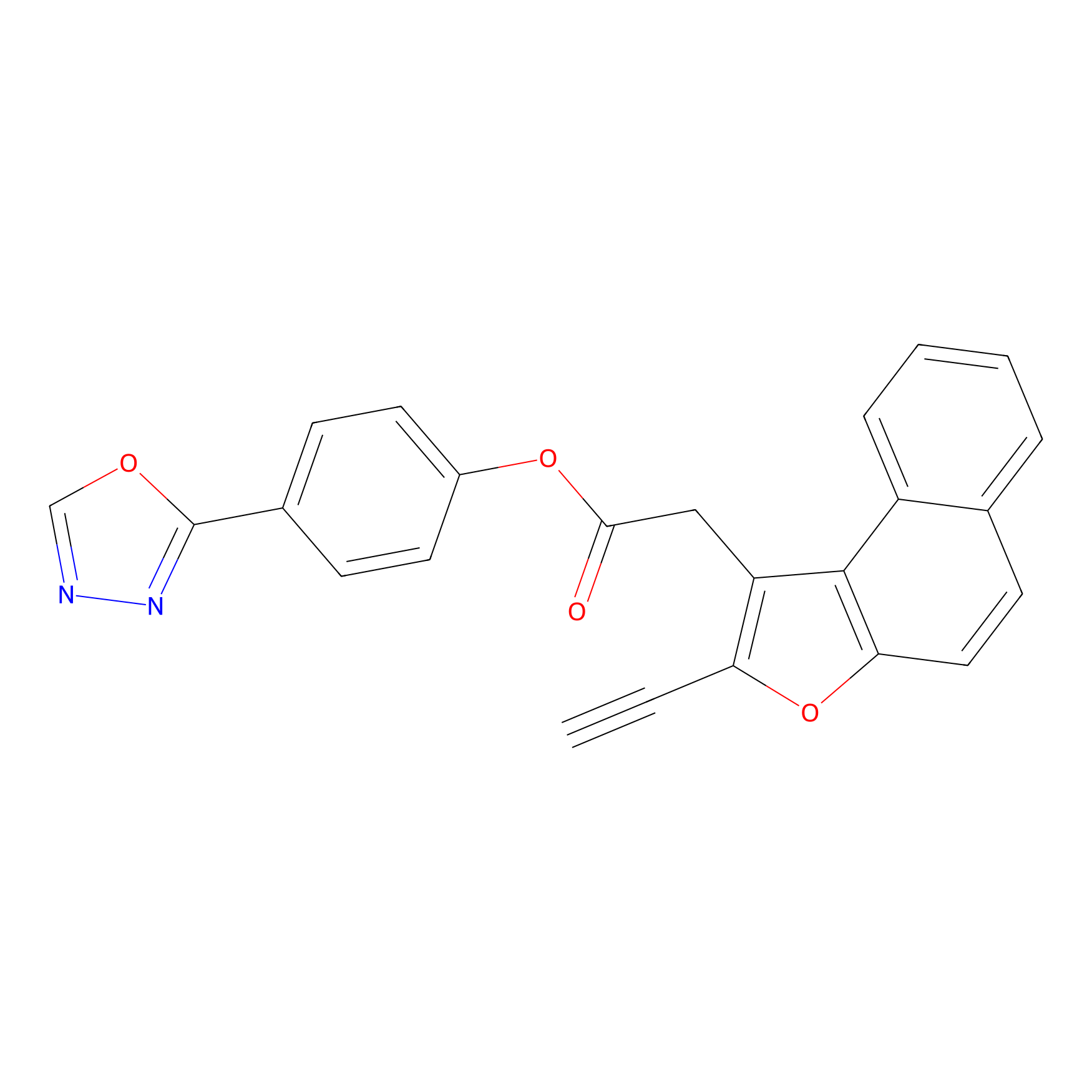 |
11.05 | LDD0043 | [5] | |
|
FBP2 Probe Info |
 |
3.13 | LDD0323 | [6] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [7] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [7] | |
|
STPyne Probe Info |
 |
K10(4.01); K362(10.00); K7(8.82) | LDD0277 | [8] | |
|
ONAyne Probe Info |
 |
K7(7.49) | LDD0275 | [8] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [9] | |
|
BTD Probe Info |
 |
C46(1.00) | LDD1700 | [10] | |
|
AHL-Pu-1 Probe Info |
 |
C452(2.96); C283(2.46) | LDD0168 | [11] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0222 | [12] | |
|
DBIA Probe Info |
 |
C283(1.99) | LDD0531 | [13] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C79(0.00); C46(0.00) | LDD0038 | [14] | |
|
IA-alkyne Probe Info |
 |
C20(0.00); C46(0.00); C79(0.00) | LDD0162 | [15] | |
|
Lodoacetamide azide Probe Info |
 |
C46(0.00); C79(0.00) | LDD0037 | [14] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [16] | |
|
DA-P3 Probe Info |
 |
N.A. | LDD0178 | [17] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [18] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [16] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [19] | |
|
Phosphinate-6 Probe Info |
 |
C33(0.00); C46(0.00) | LDD0018 | [20] | |
|
W1 Probe Info |
 |
C283(0.00); C133(0.00) | LDD0236 | [21] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [22] | |
|
NAIA_5 Probe Info |
 |
C46(0.00); C13(0.00); C79(0.00); C133(0.00) | LDD2223 | [23] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C231 Probe Info |
 |
15.56 | LDD1904 | [24] | |
|
C232 Probe Info |
 |
35.51 | LDD1905 | [24] | |
|
C233 Probe Info |
 |
13.09 | LDD1906 | [24] | |
|
C234 Probe Info |
 |
5.98 | LDD1907 | [24] | |
|
C235 Probe Info |
 |
25.28 | LDD1908 | [24] | |
|
C275 Probe Info |
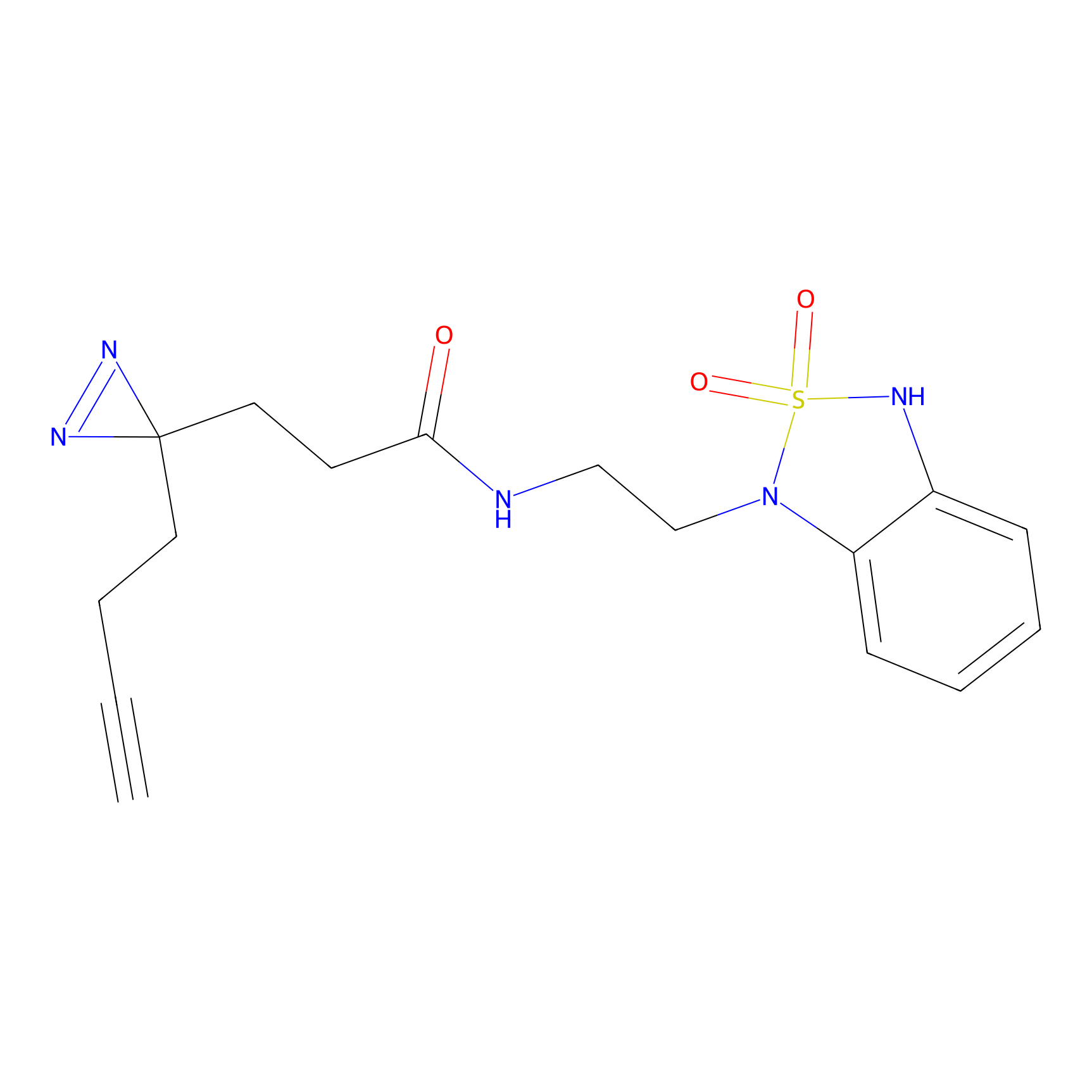 |
5.39 | LDD1945 | [24] | |
|
C289 Probe Info |
 |
25.28 | LDD1959 | [24] | |
|
C355 Probe Info |
 |
24.59 | LDD2016 | [24] | |
|
C383 Probe Info |
 |
11.88 | LDD2042 | [24] | |
|
FFF probe11 Probe Info |
 |
15.11 | LDD0471 | [25] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [25] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [25] | |
|
FFF probe2 Probe Info |
 |
20.00 | LDD0463 | [25] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [25] | |
|
FFF probe9 Probe Info |
 |
6.03 | LDD0470 | [25] | |
|
JN0003 Probe Info |
 |
17.16 | LDD0469 | [25] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [26] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C46(1.00) | LDD2142 | [10] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C46(0.94) | LDD2112 | [10] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C46(0.57) | LDD2117 | [10] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C46(0.95) | LDD2152 | [10] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C46(1.11) | LDD2103 | [10] |
| LDCM0025 | 4SU-RNA | HEK-293T | C452(2.96); C283(2.46) | LDD0168 | [11] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C452(5.35); C283(2.09) | LDD0169 | [11] |
| LDCM0214 | AC1 | HCT 116 | C283(1.99) | LDD0531 | [13] |
| LDCM0215 | AC10 | HCT 116 | C283(0.95) | LDD0532 | [13] |
| LDCM0216 | AC100 | HCT 116 | C283(1.17) | LDD0533 | [13] |
| LDCM0217 | AC101 | HCT 116 | C283(1.42) | LDD0534 | [13] |
| LDCM0218 | AC102 | HCT 116 | C283(1.12) | LDD0535 | [13] |
| LDCM0219 | AC103 | HCT 116 | C283(1.26) | LDD0536 | [13] |
| LDCM0220 | AC104 | HCT 116 | C283(1.42) | LDD0537 | [13] |
| LDCM0221 | AC105 | HCT 116 | C283(1.42) | LDD0538 | [13] |
| LDCM0222 | AC106 | HCT 116 | C283(1.32) | LDD0539 | [13] |
| LDCM0223 | AC107 | HCT 116 | C283(1.36) | LDD0540 | [13] |
| LDCM0224 | AC108 | HCT 116 | C283(0.97) | LDD0541 | [13] |
| LDCM0225 | AC109 | HCT 116 | C283(0.89) | LDD0542 | [13] |
| LDCM0226 | AC11 | HCT 116 | C283(1.05) | LDD0543 | [13] |
| LDCM0227 | AC110 | HCT 116 | C283(0.87) | LDD0544 | [13] |
| LDCM0228 | AC111 | HCT 116 | C283(1.03) | LDD0545 | [13] |
| LDCM0229 | AC112 | HCT 116 | C283(1.39) | LDD0546 | [13] |
| LDCM0230 | AC113 | HCT 116 | C283(0.99) | LDD0547 | [13] |
| LDCM0231 | AC114 | HCT 116 | C283(1.19) | LDD0548 | [13] |
| LDCM0232 | AC115 | HCT 116 | C283(1.08) | LDD0549 | [13] |
| LDCM0233 | AC116 | HCT 116 | C283(1.00) | LDD0550 | [13] |
| LDCM0234 | AC117 | HCT 116 | C283(0.99) | LDD0551 | [13] |
| LDCM0235 | AC118 | HCT 116 | C283(1.03) | LDD0552 | [13] |
| LDCM0236 | AC119 | HCT 116 | C283(1.06) | LDD0553 | [13] |
| LDCM0237 | AC12 | HCT 116 | C283(0.89) | LDD0554 | [13] |
| LDCM0238 | AC120 | HCT 116 | C283(1.08) | LDD0555 | [13] |
| LDCM0239 | AC121 | HCT 116 | C283(0.94) | LDD0556 | [13] |
| LDCM0240 | AC122 | HCT 116 | C283(1.00) | LDD0557 | [13] |
| LDCM0241 | AC123 | HCT 116 | C283(1.02) | LDD0558 | [13] |
| LDCM0242 | AC124 | HCT 116 | C283(0.97) | LDD0559 | [13] |
| LDCM0243 | AC125 | HCT 116 | C283(1.11) | LDD0560 | [13] |
| LDCM0244 | AC126 | HCT 116 | C283(1.00) | LDD0561 | [13] |
| LDCM0245 | AC127 | HCT 116 | C283(0.90) | LDD0562 | [13] |
| LDCM0246 | AC128 | HEK-293T | C283(0.97) | LDD0844 | [13] |
| LDCM0247 | AC129 | HEK-293T | C283(1.27) | LDD0845 | [13] |
| LDCM0249 | AC130 | HEK-293T | C283(1.20) | LDD0847 | [13] |
| LDCM0250 | AC131 | HEK-293T | C283(1.07) | LDD0848 | [13] |
| LDCM0251 | AC132 | HEK-293T | C283(1.16) | LDD0849 | [13] |
| LDCM0252 | AC133 | HEK-293T | C283(1.32) | LDD0850 | [13] |
| LDCM0253 | AC134 | HEK-293T | C283(1.32) | LDD0851 | [13] |
| LDCM0254 | AC135 | HEK-293T | C283(1.19) | LDD0852 | [13] |
| LDCM0255 | AC136 | HEK-293T | C283(1.32) | LDD0853 | [13] |
| LDCM0256 | AC137 | HEK-293T | C283(1.61) | LDD0854 | [13] |
| LDCM0257 | AC138 | HEK-293T | C283(1.81) | LDD0855 | [13] |
| LDCM0258 | AC139 | HEK-293T | C283(1.58) | LDD0856 | [13] |
| LDCM0259 | AC14 | HCT 116 | C283(0.90) | LDD0576 | [13] |
| LDCM0260 | AC140 | HEK-293T | C283(1.50) | LDD0858 | [13] |
| LDCM0261 | AC141 | HEK-293T | C283(1.67) | LDD0859 | [13] |
| LDCM0262 | AC142 | HEK-293T | C283(1.40) | LDD0860 | [13] |
| LDCM0263 | AC143 | HEK-293T | C283(1.10) | LDD0861 | [13] |
| LDCM0264 | AC144 | HEK-293T | C283(1.52) | LDD0862 | [13] |
| LDCM0265 | AC145 | HEK-293T | C283(1.40) | LDD0863 | [13] |
| LDCM0266 | AC146 | HEK-293T | C283(1.57) | LDD0864 | [13] |
| LDCM0267 | AC147 | HEK-293T | C283(1.66) | LDD0865 | [13] |
| LDCM0268 | AC148 | HEK-293T | C283(1.34) | LDD0866 | [13] |
| LDCM0269 | AC149 | HEK-293T | C283(1.06) | LDD0867 | [13] |
| LDCM0270 | AC15 | HCT 116 | C283(0.99) | LDD0587 | [13] |
| LDCM0271 | AC150 | HEK-293T | C283(1.20) | LDD0869 | [13] |
| LDCM0272 | AC151 | HEK-293T | C283(1.25) | LDD0870 | [13] |
| LDCM0273 | AC152 | HEK-293T | C283(1.19) | LDD0871 | [13] |
| LDCM0274 | AC153 | HEK-293T | C283(1.20) | LDD0872 | [13] |
| LDCM0621 | AC154 | HEK-293T | C283(1.28) | LDD2162 | [13] |
| LDCM0622 | AC155 | HEK-293T | C283(1.17) | LDD2163 | [13] |
| LDCM0623 | AC156 | HEK-293T | C283(1.55) | LDD2164 | [13] |
| LDCM0624 | AC157 | HEK-293T | C283(1.77) | LDD2165 | [13] |
| LDCM0276 | AC17 | HCT 116 | C283(1.05) | LDD0593 | [13] |
| LDCM0277 | AC18 | HCT 116 | C283(1.07) | LDD0594 | [13] |
| LDCM0278 | AC19 | HCT 116 | C283(1.07) | LDD0595 | [13] |
| LDCM0279 | AC2 | HCT 116 | C283(1.35) | LDD0596 | [13] |
| LDCM0280 | AC20 | HCT 116 | C283(0.96) | LDD0597 | [13] |
| LDCM0281 | AC21 | HCT 116 | C283(1.05) | LDD0598 | [13] |
| LDCM0282 | AC22 | HCT 116 | C283(0.94) | LDD0599 | [13] |
| LDCM0283 | AC23 | HCT 116 | C283(1.00) | LDD0600 | [13] |
| LDCM0284 | AC24 | HCT 116 | C283(1.12) | LDD0601 | [13] |
| LDCM0285 | AC25 | HCT 116 | C283(1.11) | LDD0602 | [13] |
| LDCM0286 | AC26 | HCT 116 | C283(1.30) | LDD0603 | [13] |
| LDCM0287 | AC27 | HCT 116 | C283(1.34) | LDD0604 | [13] |
| LDCM0288 | AC28 | HCT 116 | C283(1.34) | LDD0605 | [13] |
| LDCM0289 | AC29 | HCT 116 | C283(1.23) | LDD0606 | [13] |
| LDCM0290 | AC3 | HCT 116 | C283(1.11) | LDD0607 | [13] |
| LDCM0291 | AC30 | HCT 116 | C283(1.48) | LDD0608 | [13] |
| LDCM0292 | AC31 | HCT 116 | C283(1.27) | LDD0609 | [13] |
| LDCM0293 | AC32 | HCT 116 | C283(1.66) | LDD0610 | [13] |
| LDCM0294 | AC33 | HCT 116 | C283(1.48) | LDD0611 | [13] |
| LDCM0295 | AC34 | HCT 116 | C283(1.59) | LDD0612 | [13] |
| LDCM0296 | AC35 | HCT 116 | C283(0.88) | LDD0613 | [13] |
| LDCM0297 | AC36 | HCT 116 | C283(0.84) | LDD0614 | [13] |
| LDCM0298 | AC37 | HCT 116 | C283(0.77) | LDD0615 | [13] |
| LDCM0299 | AC38 | HCT 116 | C283(1.06) | LDD0616 | [13] |
| LDCM0300 | AC39 | HCT 116 | C283(0.93) | LDD0617 | [13] |
| LDCM0301 | AC4 | HCT 116 | C283(1.77) | LDD0618 | [13] |
| LDCM0302 | AC40 | HCT 116 | C283(0.92) | LDD0619 | [13] |
| LDCM0303 | AC41 | HCT 116 | C283(0.83) | LDD0620 | [13] |
| LDCM0304 | AC42 | HCT 116 | C283(0.99) | LDD0621 | [13] |
| LDCM0305 | AC43 | HCT 116 | C283(1.02) | LDD0622 | [13] |
| LDCM0306 | AC44 | HCT 116 | C283(0.98) | LDD0623 | [13] |
| LDCM0307 | AC45 | HCT 116 | C283(0.70) | LDD0624 | [13] |
| LDCM0308 | AC46 | HCT 116 | C283(0.89) | LDD0625 | [13] |
| LDCM0309 | AC47 | HCT 116 | C283(0.97) | LDD0626 | [13] |
| LDCM0310 | AC48 | HCT 116 | C283(1.11) | LDD0627 | [13] |
| LDCM0311 | AC49 | HCT 116 | C283(1.43) | LDD0628 | [13] |
| LDCM0312 | AC5 | HCT 116 | C283(1.31) | LDD0629 | [13] |
| LDCM0313 | AC50 | HCT 116 | C283(1.40) | LDD0630 | [13] |
| LDCM0314 | AC51 | HCT 116 | C283(0.82) | LDD0631 | [13] |
| LDCM0315 | AC52 | HCT 116 | C283(0.97) | LDD0632 | [13] |
| LDCM0316 | AC53 | HCT 116 | C283(1.44) | LDD0633 | [13] |
| LDCM0317 | AC54 | HCT 116 | C283(1.13) | LDD0634 | [13] |
| LDCM0318 | AC55 | HCT 116 | C283(1.33) | LDD0635 | [13] |
| LDCM0319 | AC56 | HCT 116 | C283(1.74) | LDD0636 | [13] |
| LDCM0320 | AC57 | HEK-293T | C283(0.83); C46(1.19) | LDD0918 | [13] |
| LDCM0321 | AC58 | HEK-293T | C283(1.14); C46(1.06) | LDD0919 | [13] |
| LDCM0322 | AC59 | HEK-293T | C283(1.18); C46(1.31) | LDD0920 | [13] |
| LDCM0323 | AC6 | HCT 116 | C283(0.99) | LDD0640 | [13] |
| LDCM0324 | AC60 | HEK-293T | C283(0.77); C46(1.24) | LDD0922 | [13] |
| LDCM0325 | AC61 | HEK-293T | C283(1.35); C46(1.48) | LDD0923 | [13] |
| LDCM0326 | AC62 | HEK-293T | C283(0.98); C46(1.19) | LDD0924 | [13] |
| LDCM0327 | AC63 | HEK-293T | C283(1.11); C46(1.28) | LDD0925 | [13] |
| LDCM0328 | AC64 | HEK-293T | C283(1.00); C46(1.28) | LDD0926 | [13] |
| LDCM0329 | AC65 | HEK-293T | C283(1.11); C46(1.34) | LDD0927 | [13] |
| LDCM0330 | AC66 | HEK-293T | C283(1.02); C46(1.17) | LDD0928 | [13] |
| LDCM0331 | AC67 | HEK-293T | C283(1.20); C46(1.31) | LDD0929 | [13] |
| LDCM0332 | AC68 | HEK-293T | C283(1.08) | LDD0930 | [13] |
| LDCM0333 | AC69 | HEK-293T | C283(1.07) | LDD0931 | [13] |
| LDCM0334 | AC7 | HCT 116 | C283(0.95) | LDD0651 | [13] |
| LDCM0335 | AC70 | HEK-293T | C283(1.55) | LDD0933 | [13] |
| LDCM0336 | AC71 | HEK-293T | C283(1.37) | LDD0934 | [13] |
| LDCM0337 | AC72 | HEK-293T | C283(1.05) | LDD0935 | [13] |
| LDCM0338 | AC73 | HEK-293T | C283(1.01) | LDD0936 | [13] |
| LDCM0339 | AC74 | HEK-293T | C283(1.32) | LDD0937 | [13] |
| LDCM0340 | AC75 | HEK-293T | C283(1.53) | LDD0938 | [13] |
| LDCM0341 | AC76 | HEK-293T | C283(1.22) | LDD0939 | [13] |
| LDCM0342 | AC77 | HEK-293T | C283(1.15) | LDD0940 | [13] |
| LDCM0343 | AC78 | HEK-293T | C283(1.25) | LDD0941 | [13] |
| LDCM0344 | AC79 | HEK-293T | C283(1.25) | LDD0942 | [13] |
| LDCM0345 | AC8 | HCT 116 | C283(0.94) | LDD0662 | [13] |
| LDCM0346 | AC80 | HEK-293T | C283(1.42) | LDD0944 | [13] |
| LDCM0347 | AC81 | HEK-293T | C283(1.30) | LDD0945 | [13] |
| LDCM0348 | AC82 | HEK-293T | C283(1.43) | LDD0946 | [13] |
| LDCM0349 | AC83 | HCT 116 | C283(1.73) | LDD0666 | [13] |
| LDCM0350 | AC84 | HCT 116 | C283(2.23) | LDD0667 | [13] |
| LDCM0351 | AC85 | HCT 116 | C283(1.10) | LDD0668 | [13] |
| LDCM0352 | AC86 | HCT 116 | C283(1.38) | LDD0669 | [13] |
| LDCM0353 | AC87 | HCT 116 | C283(1.15) | LDD0670 | [13] |
| LDCM0354 | AC88 | HCT 116 | C283(1.50) | LDD0671 | [13] |
| LDCM0355 | AC89 | HCT 116 | C283(1.41) | LDD0672 | [13] |
| LDCM0357 | AC90 | HCT 116 | C283(1.21) | LDD0674 | [13] |
| LDCM0358 | AC91 | HCT 116 | C283(1.59) | LDD0675 | [13] |
| LDCM0359 | AC92 | HCT 116 | C283(1.41) | LDD0676 | [13] |
| LDCM0360 | AC93 | HCT 116 | C283(1.44) | LDD0677 | [13] |
| LDCM0361 | AC94 | HCT 116 | C283(1.08) | LDD0678 | [13] |
| LDCM0362 | AC95 | HCT 116 | C283(1.43) | LDD0679 | [13] |
| LDCM0363 | AC96 | HCT 116 | C283(1.36) | LDD0680 | [13] |
| LDCM0364 | AC97 | HCT 116 | C283(1.81) | LDD0681 | [13] |
| LDCM0365 | AC98 | HCT 116 | C283(1.50) | LDD0682 | [13] |
| LDCM0366 | AC99 | HCT 116 | C283(0.85) | LDD0683 | [13] |
| LDCM0248 | AKOS034007472 | HCT 116 | C283(1.05) | LDD0565 | [13] |
| LDCM0356 | AKOS034007680 | HCT 116 | C283(0.93) | LDD0673 | [13] |
| LDCM0275 | AKOS034007705 | HCT 116 | C283(0.92) | LDD0592 | [13] |
| LDCM0156 | Aniline | NCI-H1299 | 11.98 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C133(0.91); C46(1.12) | LDD2171 | [13] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [12] |
| LDCM0632 | CL-Sc | Hep-G2 | C46(0.83) | LDD2227 | [23] |
| LDCM0367 | CL1 | HEK-293T | C283(1.02) | LDD0965 | [13] |
| LDCM0368 | CL10 | HEK-293T | C283(2.12) | LDD0966 | [13] |
| LDCM0369 | CL100 | HCT 116 | C283(1.49) | LDD0686 | [13] |
| LDCM0370 | CL101 | HCT 116 | C283(1.08) | LDD0687 | [13] |
| LDCM0371 | CL102 | HCT 116 | C283(0.91) | LDD0688 | [13] |
| LDCM0372 | CL103 | HCT 116 | C283(1.03) | LDD0689 | [13] |
| LDCM0373 | CL104 | HCT 116 | C283(0.94) | LDD0690 | [13] |
| LDCM0374 | CL105 | HCT 116 | C283(1.17) | LDD0691 | [13] |
| LDCM0375 | CL106 | HCT 116 | C283(1.23) | LDD0692 | [13] |
| LDCM0376 | CL107 | HCT 116 | C283(1.17) | LDD0693 | [13] |
| LDCM0377 | CL108 | HCT 116 | C283(1.33) | LDD0694 | [13] |
| LDCM0378 | CL109 | HCT 116 | C283(1.14) | LDD0695 | [13] |
| LDCM0379 | CL11 | HEK-293T | C283(1.89) | LDD0977 | [13] |
| LDCM0380 | CL110 | HCT 116 | C283(1.19) | LDD0697 | [13] |
| LDCM0381 | CL111 | HCT 116 | C283(1.16) | LDD0698 | [13] |
| LDCM0382 | CL112 | HCT 116 | C283(1.16) | LDD0699 | [13] |
| LDCM0383 | CL113 | HCT 116 | C283(1.34) | LDD0700 | [13] |
| LDCM0384 | CL114 | HCT 116 | C283(1.23) | LDD0701 | [13] |
| LDCM0385 | CL115 | HCT 116 | C283(1.52) | LDD0702 | [13] |
| LDCM0386 | CL116 | HCT 116 | C283(1.39) | LDD0703 | [13] |
| LDCM0387 | CL117 | HCT 116 | C283(1.12) | LDD0704 | [13] |
| LDCM0388 | CL118 | HCT 116 | C283(0.84) | LDD0705 | [13] |
| LDCM0389 | CL119 | HCT 116 | C283(0.85) | LDD0706 | [13] |
| LDCM0390 | CL12 | HEK-293T | C283(1.07) | LDD0988 | [13] |
| LDCM0391 | CL120 | HCT 116 | C283(1.04) | LDD0708 | [13] |
| LDCM0392 | CL121 | HCT 116 | C283(1.40) | LDD0709 | [13] |
| LDCM0393 | CL122 | HCT 116 | C283(1.08) | LDD0710 | [13] |
| LDCM0394 | CL123 | HCT 116 | C283(1.15) | LDD0711 | [13] |
| LDCM0395 | CL124 | HCT 116 | C283(1.34) | LDD0712 | [13] |
| LDCM0396 | CL125 | HEK-293T | C283(1.12); C46(1.23) | LDD0994 | [13] |
| LDCM0397 | CL126 | HEK-293T | C283(1.29); C46(1.14) | LDD0995 | [13] |
| LDCM0398 | CL127 | HEK-293T | C283(0.99); C46(1.22) | LDD0996 | [13] |
| LDCM0399 | CL128 | HEK-293T | C283(0.94); C46(0.91) | LDD0997 | [13] |
| LDCM0400 | CL13 | HEK-293T | C283(1.04) | LDD0998 | [13] |
| LDCM0401 | CL14 | HEK-293T | C283(1.04) | LDD0999 | [13] |
| LDCM0402 | CL15 | HEK-293T | C283(1.45) | LDD1000 | [13] |
| LDCM0403 | CL16 | HCT 116 | C283(0.57) | LDD0720 | [13] |
| LDCM0404 | CL17 | HCT 116 | C283(0.69) | LDD0721 | [13] |
| LDCM0405 | CL18 | HCT 116 | C283(0.79) | LDD0722 | [13] |
| LDCM0406 | CL19 | HCT 116 | C283(0.75) | LDD0723 | [13] |
| LDCM0407 | CL2 | HEK-293T | C283(1.13) | LDD1005 | [13] |
| LDCM0408 | CL20 | HCT 116 | C283(0.68) | LDD0725 | [13] |
| LDCM0409 | CL21 | HCT 116 | C283(0.87) | LDD0726 | [13] |
| LDCM0410 | CL22 | HCT 116 | C283(1.02) | LDD0727 | [13] |
| LDCM0411 | CL23 | HCT 116 | C283(1.23) | LDD0728 | [13] |
| LDCM0412 | CL24 | HCT 116 | C283(0.86) | LDD0729 | [13] |
| LDCM0413 | CL25 | HCT 116 | C283(0.75) | LDD0730 | [13] |
| LDCM0414 | CL26 | HCT 116 | C283(0.73) | LDD0731 | [13] |
| LDCM0415 | CL27 | HCT 116 | C283(0.81) | LDD0732 | [13] |
| LDCM0416 | CL28 | HCT 116 | C283(0.69) | LDD0733 | [13] |
| LDCM0417 | CL29 | HCT 116 | C283(0.58) | LDD0734 | [13] |
| LDCM0418 | CL3 | HEK-293T | C283(1.12) | LDD1016 | [13] |
| LDCM0419 | CL30 | HCT 116 | C283(0.86) | LDD0736 | [13] |
| LDCM0420 | CL31 | HEK-293T | C283(1.26); C46(0.90) | LDD1018 | [13] |
| LDCM0421 | CL32 | HEK-293T | C283(1.02); C46(0.86) | LDD1019 | [13] |
| LDCM0422 | CL33 | HEK-293T | C283(1.42); C46(0.95) | LDD1020 | [13] |
| LDCM0423 | CL34 | HEK-293T | C283(1.15); C46(1.25) | LDD1021 | [13] |
| LDCM0424 | CL35 | HEK-293T | C283(1.23); C46(0.98) | LDD1022 | [13] |
| LDCM0425 | CL36 | HEK-293T | C283(1.07); C46(0.98) | LDD1023 | [13] |
| LDCM0426 | CL37 | HEK-293T | C283(1.27); C46(1.19) | LDD1024 | [13] |
| LDCM0428 | CL39 | HEK-293T | C283(1.77); C46(0.90) | LDD1026 | [13] |
| LDCM0429 | CL4 | HEK-293T | C283(1.23) | LDD1027 | [13] |
| LDCM0430 | CL40 | HEK-293T | C283(1.15); C46(1.10) | LDD1028 | [13] |
| LDCM0431 | CL41 | HEK-293T | C283(1.41); C46(0.76) | LDD1029 | [13] |
| LDCM0432 | CL42 | HEK-293T | C283(1.60); C46(1.12) | LDD1030 | [13] |
| LDCM0433 | CL43 | HEK-293T | C283(1.47); C46(1.03) | LDD1031 | [13] |
| LDCM0434 | CL44 | HEK-293T | C283(1.20); C46(1.16) | LDD1032 | [13] |
| LDCM0435 | CL45 | HEK-293T | C283(1.43); C46(1.07) | LDD1033 | [13] |
| LDCM0436 | CL46 | HEK-293T | C283(1.23) | LDD1034 | [13] |
| LDCM0437 | CL47 | HEK-293T | C283(1.07) | LDD1035 | [13] |
| LDCM0438 | CL48 | HEK-293T | C283(1.11) | LDD1036 | [13] |
| LDCM0439 | CL49 | HEK-293T | C283(1.24) | LDD1037 | [13] |
| LDCM0440 | CL5 | HEK-293T | C283(1.18) | LDD1038 | [13] |
| LDCM0441 | CL50 | HEK-293T | C283(1.43) | LDD1039 | [13] |
| LDCM0442 | CL51 | HEK-293T | C283(1.13) | LDD1040 | [13] |
| LDCM0443 | CL52 | HEK-293T | C283(0.95) | LDD1041 | [13] |
| LDCM0444 | CL53 | HEK-293T | C283(1.00) | LDD1042 | [13] |
| LDCM0445 | CL54 | HEK-293T | C283(1.16) | LDD1043 | [13] |
| LDCM0446 | CL55 | HEK-293T | C283(1.10) | LDD1044 | [13] |
| LDCM0447 | CL56 | HEK-293T | C283(0.93) | LDD1045 | [13] |
| LDCM0448 | CL57 | HEK-293T | C283(1.10) | LDD1046 | [13] |
| LDCM0449 | CL58 | HEK-293T | C283(0.94) | LDD1047 | [13] |
| LDCM0450 | CL59 | HEK-293T | C283(1.20) | LDD1048 | [13] |
| LDCM0451 | CL6 | HEK-293T | C283(1.19) | LDD1049 | [13] |
| LDCM0452 | CL60 | HEK-293T | C283(1.34) | LDD1050 | [13] |
| LDCM0453 | CL61 | HCT 116 | C283(0.98) | LDD0770 | [13] |
| LDCM0454 | CL62 | HCT 116 | C283(1.03) | LDD0771 | [13] |
| LDCM0455 | CL63 | HCT 116 | C283(1.12) | LDD0772 | [13] |
| LDCM0456 | CL64 | HCT 116 | C283(1.22) | LDD0773 | [13] |
| LDCM0457 | CL65 | HCT 116 | C283(1.01) | LDD0774 | [13] |
| LDCM0458 | CL66 | HCT 116 | C283(1.66) | LDD0775 | [13] |
| LDCM0459 | CL67 | HCT 116 | C283(1.40) | LDD0776 | [13] |
| LDCM0460 | CL68 | HCT 116 | C283(1.12) | LDD0777 | [13] |
| LDCM0461 | CL69 | HCT 116 | C283(0.88) | LDD0778 | [13] |
| LDCM0462 | CL7 | HEK-293T | C283(1.25) | LDD1060 | [13] |
| LDCM0463 | CL70 | HCT 116 | C283(0.99) | LDD0780 | [13] |
| LDCM0464 | CL71 | HCT 116 | C283(1.35) | LDD0781 | [13] |
| LDCM0465 | CL72 | HCT 116 | C283(1.22) | LDD0782 | [13] |
| LDCM0466 | CL73 | HCT 116 | C283(1.13) | LDD0783 | [13] |
| LDCM0467 | CL74 | HCT 116 | C283(1.12) | LDD0784 | [13] |
| LDCM0469 | CL76 | HCT 116 | C283(1.08) | LDD0786 | [13] |
| LDCM0470 | CL77 | HCT 116 | C283(1.02) | LDD0787 | [13] |
| LDCM0471 | CL78 | HCT 116 | C283(0.83) | LDD0788 | [13] |
| LDCM0472 | CL79 | HCT 116 | C283(1.15) | LDD0789 | [13] |
| LDCM0473 | CL8 | HEK-293T | C283(1.47) | LDD1071 | [13] |
| LDCM0474 | CL80 | HCT 116 | C283(1.04) | LDD0791 | [13] |
| LDCM0475 | CL81 | HCT 116 | C283(1.04) | LDD0792 | [13] |
| LDCM0476 | CL82 | HCT 116 | C283(0.96) | LDD0793 | [13] |
| LDCM0477 | CL83 | HCT 116 | C283(0.99) | LDD0794 | [13] |
| LDCM0478 | CL84 | HCT 116 | C283(0.93) | LDD0795 | [13] |
| LDCM0479 | CL85 | HCT 116 | C283(1.05) | LDD0796 | [13] |
| LDCM0480 | CL86 | HCT 116 | C283(1.02) | LDD0797 | [13] |
| LDCM0481 | CL87 | HCT 116 | C283(1.01) | LDD0798 | [13] |
| LDCM0482 | CL88 | HCT 116 | C283(0.99) | LDD0799 | [13] |
| LDCM0483 | CL89 | HCT 116 | C283(0.82) | LDD0800 | [13] |
| LDCM0484 | CL9 | HEK-293T | C283(1.25) | LDD1082 | [13] |
| LDCM0485 | CL90 | HCT 116 | C283(1.15) | LDD0802 | [13] |
| LDCM0486 | CL91 | HCT 116 | C283(1.17) | LDD0803 | [13] |
| LDCM0487 | CL92 | HCT 116 | C283(1.02) | LDD0804 | [13] |
| LDCM0488 | CL93 | HCT 116 | C283(1.11) | LDD0805 | [13] |
| LDCM0489 | CL94 | HCT 116 | C283(1.11) | LDD0806 | [13] |
| LDCM0490 | CL95 | HCT 116 | C283(1.04) | LDD0807 | [13] |
| LDCM0491 | CL96 | HCT 116 | C283(0.99) | LDD0808 | [13] |
| LDCM0492 | CL97 | HCT 116 | C283(1.29) | LDD0809 | [13] |
| LDCM0493 | CL98 | HCT 116 | C283(1.12) | LDD0810 | [13] |
| LDCM0494 | CL99 | HCT 116 | C283(1.36) | LDD0811 | [13] |
| LDCM0189 | Compound 16 | HEK-293T | 3.84 | LDD0492 | [25] |
| LDCM0185 | Compound 17 | HEK-293T | 5.36 | LDD0512 | [25] |
| LDCM0191 | Compound 21 | HEK-293T | 7.04 | LDD0508 | [25] |
| LDCM0190 | Compound 34 | HEK-293T | 11.59 | LDD0497 | [25] |
| LDCM0192 | Compound 35 | HEK-293T | 4.71 | LDD0491 | [25] |
| LDCM0193 | Compound 36 | HEK-293T | 7.72 | LDD0511 | [25] |
| LDCM0028 | Dobutamine | HEK-293T | 4.80 | LDD0180 | [17] |
| LDCM0027 | Dopamine | HEK-293T | 6.43 | LDD0179 | [17] |
| LDCM0495 | E2913 | HEK-293T | C46(1.01); C133(1.05); C283(0.95); C33(0.80) | LDD1698 | [27] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C46(4.74); C133(1.14); C508(0.80) | LDD1702 | [10] |
| LDCM0625 | F8 | Ramos | C46(1.28); C133(1.36) | LDD2187 | [28] |
| LDCM0572 | Fragment10 | Ramos | C46(7.78); C133(1.73) | LDD2189 | [28] |
| LDCM0573 | Fragment11 | Ramos | C46(0.50); C133(0.27) | LDD2190 | [28] |
| LDCM0574 | Fragment12 | Ramos | C46(9.62); C133(0.58) | LDD2191 | [28] |
| LDCM0575 | Fragment13 | Ramos | C46(1.22); C133(0.86) | LDD2192 | [28] |
| LDCM0576 | Fragment14 | Ramos | C46(2.87); C133(0.98) | LDD2193 | [28] |
| LDCM0579 | Fragment20 | Ramos | C133(1.46) | LDD2194 | [28] |
| LDCM0580 | Fragment21 | Ramos | C46(1.48); C133(0.80) | LDD2195 | [28] |
| LDCM0582 | Fragment23 | Ramos | C46(1.29); C133(0.39) | LDD2196 | [28] |
| LDCM0578 | Fragment27 | Ramos | C46(1.26); C133(0.87) | LDD2197 | [28] |
| LDCM0586 | Fragment28 | Ramos | C46(1.14); C133(1.20) | LDD2198 | [28] |
| LDCM0588 | Fragment30 | Ramos | C46(1.66); C133(0.68) | LDD2199 | [28] |
| LDCM0589 | Fragment31 | Ramos | C46(1.89); C133(0.87) | LDD2200 | [28] |
| LDCM0590 | Fragment32 | Ramos | C46(8.28); C133(1.10) | LDD2201 | [28] |
| LDCM0468 | Fragment33 | HCT 116 | C283(1.24) | LDD0785 | [13] |
| LDCM0596 | Fragment38 | Ramos | C46(2.01); C133(1.01) | LDD2203 | [28] |
| LDCM0566 | Fragment4 | Ramos | C46(1.74); C133(0.71) | LDD2184 | [28] |
| LDCM0427 | Fragment51 | HEK-293T | C283(1.25); C46(0.98) | LDD1025 | [13] |
| LDCM0610 | Fragment52 | Ramos | C46(2.18); C133(1.17) | LDD2204 | [28] |
| LDCM0614 | Fragment56 | Ramos | C46(1.78); C133(0.73) | LDD2205 | [28] |
| LDCM0569 | Fragment7 | Ramos | C46(2.36); C133(1.08) | LDD2186 | [28] |
| LDCM0571 | Fragment9 | Ramos | C133(0.82) | LDD2188 | [28] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [9] |
| LDCM0022 | KB02 | HEK-293T | C133(0.91); C283(1.01); C435(0.98); C452(0.98) | LDD1492 | [27] |
| LDCM0023 | KB03 | HEK-293T | C133(0.96); C283(1.04); C435(1.11); C452(1.11) | LDD1497 | [27] |
| LDCM0024 | KB05 | HMCB | C46(3.02); C283(1.48); C133(2.31) | LDD3312 | [29] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C46(1.39) | LDD2102 | [10] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C46(0.46) | LDD2121 | [10] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C46(0.51) | LDD2089 | [10] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C46(1.55); C133(1.31) | LDD2090 | [10] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C46(1.24); C133(0.81) | LDD2092 | [10] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C46(1.08) | LDD2093 | [10] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C46(1.62); C133(0.94) | LDD2094 | [10] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C46(0.62) | LDD2097 | [10] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C133(1.10) | LDD2098 | [10] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C46(0.68) | LDD2099 | [10] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C46(0.80) | LDD2100 | [10] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C46(0.94) | LDD2101 | [10] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C46(0.80) | LDD2107 | [10] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C46(0.66) | LDD2109 | [10] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C46(0.80) | LDD2111 | [10] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C46(0.60) | LDD2115 | [10] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C46(0.64) | LDD2123 | [10] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C46(0.63) | LDD2125 | [10] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C46(0.71) | LDD2129 | [10] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C46(0.68) | LDD2133 | [10] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C46(0.91) | LDD2135 | [10] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C46(0.91) | LDD2136 | [10] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C46(0.82) | LDD2137 | [10] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C46(1.00) | LDD1700 | [10] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C46(0.49) | LDD2140 | [10] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C46(4.43) | LDD2145 | [10] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C46(0.59) | LDD2146 | [10] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C46(0.70) | LDD2148 | [10] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C46(1.06) | LDD2206 | [30] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C46(0.88); C46(0.27) | LDD2207 | [30] |
| LDCM0029 | Quercetin | HEK-293T | 7.90 | LDD0181 | [17] |
| LDCM0131 | RA190 | MM1.R | C283(1.07) | LDD0304 | [31] |
| LDCM0021 | THZ1 | HCT 116 | C133(0.91); C46(1.12) | LDD2173 | [13] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Interferon regulatory factor 5 (IRF5) | IRF family | Q13568 | |||
| Signal transducer and activator of transcription 1-alpha/beta (STAT1) | Transcription factor STAT family | P42224 | |||
Other
References
